Population Structure of Modern Winter Wheat Accessions from Central Asia
Abstract
1. Introduction
2. Results
2.1. Genetic Diversity and Genetic Distances in Winter Wheat Populations
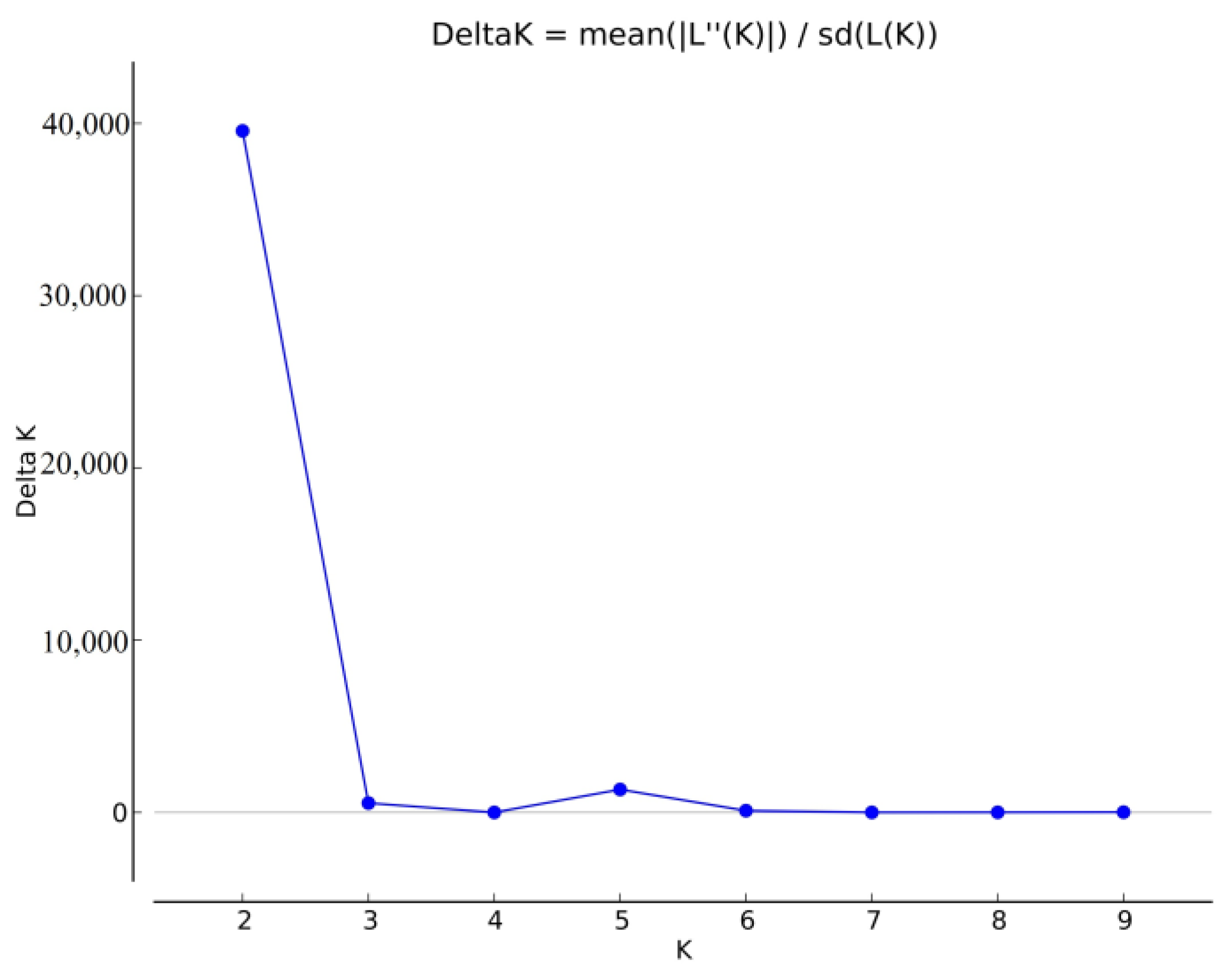
2.2. Population Structure of the Winter Wheat Accessions from Central Asia
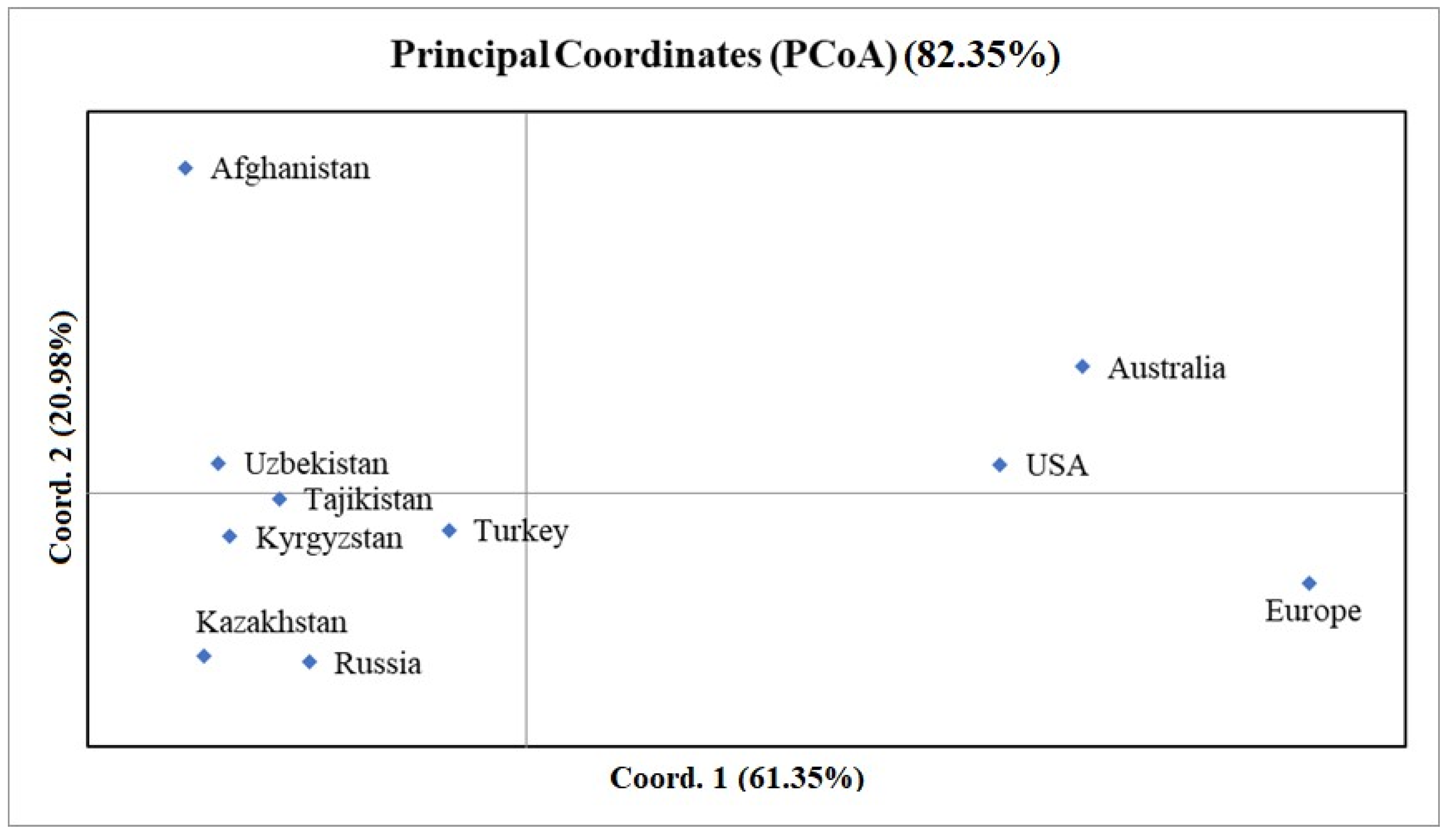
2.3. The Evaluation of the Winter Wheat Collection Based on Chromosomes Using Principal Coordinate Analysis
2.4. The Selection of SNP Markers Associated with the Regional Adaptation of Winter Wheat in Central Asia
3. Materials and Methods
3.1. Winter Wheat Collection
3.2. Genotyping Collection
3.3. Statistics Analysis
4. Discussion
4.1. Comparative Population Structures of Spring and Winter Types of Wheat in Central Asia
4.2. Patterns of PCoA Plots Using SNPs on Individual Wheat Chromosomes
4.3. Identification of Genes Associated with Plant Adaptation in the Central Asian Region
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Food and Agriculture Organization of the United Nations. Available online: https://www.fao.org/faostat/en/#home (accessed on 25 February 2023).
- Morgounov, A.; Gómez-Becerra, H.F.; Abugalieva, A.; Dzhunusova, M.; Yessimbekova, M.; Muminjanov, H.; Zelrnskiy, Y.; Cakmak, I. Iron and zinc grain density in common wheat grown in Central Asia. Euphytica 2007, 155, 193–203. [Google Scholar] [CrossRef]
- Li, X.; Liu, Y. The conversion of spring wheat into winter wheat and vice versa: False claimor Lamarckian inheritance? J. Biosci. 2010, 35, 321–325. [Google Scholar] [CrossRef]
- Trevaskis, B.; Hemming, M.N.; Dennis, E.S.; Peacock, W.J. The molecular basis of vernalization-induced flowering in cereals. Trends Plant Sci. 2007, 12, 352–357. [Google Scholar] [CrossRef]
- Milec, Z.; Strejčková, B.; Šafář, J. Contemplation on wheat vernalization. Front. Plant Sci. 2022, 13, 1093792. [Google Scholar] [CrossRef]
- Morgounov, A.; Rosseeva, L.; Koyshibayev, M. Leaf rust of spring wheat in Northern Kazakhstan and Siberia: Incidence, virulence, and breeding for resistance. Aust. J. Agric. Res. 2007, 58, 847. [Google Scholar] [CrossRef]
- Abugaliyeva, A.I.; Morgounov, A.I. Genetic potential of winter wheat grain quality in Central Asia. Int. J. Environ. Sci. Educ. 2016, 11, 4869–4884. [Google Scholar]
- Bureau of National Statistics Agency for Strategic Planning and Reforms of the Republic of Kazakhstan. Available online: https://new.stat.gov.kz/en/ (accessed on 20 February 2023).
- Official Information Source of the Prime Minister of the Republic of Kazakhstan. Available online: https://primeminister.kz/en/news/reviews/itogi-razvitiya-sfery-selskogo-hozyaystva-za-2021-god-i-plany-na-predstoyashchiy-period-22422 (accessed on 20 March 2023).
- Tolstoy, S.P. Ancient Khorezm; Moscow State University: Moscow, Russia, 1948; p. 269. (In Russian) [Google Scholar]
- Vavilov, N.L. Studies on the Origin of Cultivated Plants; Institute of Applied Botany and Plant Breeding: Leningrad, Russia, 1926; p. 78. [Google Scholar]
- Udachin, R.; Shahmedov, I. Wheats of Central Asia; FAN, USSR: Leningrad, Russia, 1984; p. 135. (In Russian) [Google Scholar]
- Morgounov, A.I. Wheat and wheat breeding in the former USSR. CIMMYT 1992, 13, 35. [Google Scholar]
- Rong, X.J. The Silk Road and Cultural Interaction between East and West; Peking University Press: Beijing, China, 2015. [Google Scholar]
- Wang, G.; Chen, Q.; Yang, Y.; Duan, Y.; Yang, Y. Exchanges of economic plants along the land silk road. BMC Plant Biol. 2022, 22, 619. [Google Scholar] [CrossRef]
- Martynov, S.P.; Dobrovotvorskaya, T.V.; Morvounov, A.I.; Urazaliev, R.A.; Absattarova, A.S. Genealogical analysis of diversity of spring bread wheat cultivars released in Kazakhstan from 1929–2004. Acta Agron. Hung. 2005, 53, 261–272. [Google Scholar] [CrossRef]
- Liefert, W.M.; Liefert, O.; Vocke, G.; Allen, E.W. Former Soviet Union region to play larger role in meeting World wheat needs. Amber Waves 2010, 8, 12–19. [Google Scholar] [CrossRef]
- Rasheed, A.; Hao, Y.; Xia, X.; Khan, A.; Xu, Y.; Varshney, R.K.; He, Z. Crop breeding chips and genotyping platforms: Progress, challenges, and perspectives. Mol. Plant 2017, 10, 1047–1064. [Google Scholar] [CrossRef]
- Appels, R.; Eversole, K.; Feuille, C.; Keller, B.; Rogers, J.; Stein, N.; The International Wheat Genome Sequencing Consortium (IWGSC). Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 2018, 361, eaar7191. [Google Scholar] [CrossRef]
- Cavanagh, C.R.; Chao, S.; Wang, S.; Huang, B.E.; Stephen, S.; Kiani, S.; Forrest, K.; Saintenac, C.; Brown-Guedira, G.L.; Akhunova, A.; et al. Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc. Natl. Acad. Sci. USA 2013, 110, 8057–8062. [Google Scholar] [CrossRef]
- Boeven, P.H.; Longin, C.F.H.; Leiser, W.L.; Kollers, S.; Ebmeyer, E.; Würschum, T. Genetic architecture of male floral traits required for hybrid wheat breeding. Theor. Appl. Genet. 2016, 129, 2343–2357. [Google Scholar] [CrossRef]
- Wang, S.; Wong, D.; Forrest, K.; Allen, A.; Chao, S.; Huang, B.E.; Maccaferri, M.; Salvi, S.; Milner, S.G.; Cattivelli, L.; et al. Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol. J. 2014, 12, 787–796. [Google Scholar] [CrossRef]
- Allen, A.M.; Winfield, M.O.; Burridge, A.J.; Downie, R.C.; Benbow, H.R.; Barker, G.L.; Wilkinson, P.A.; Coghill, J.; Waterfall, C.; Davassi, A.; et al. Characterization of a Wheat Breeders’ Array suitable for high-throughput SNP genotyping of global accessions of hexaploid bread wheat (Triticum aestivum). Plant Biotechnol. J. 2017, 15, 390–401. [Google Scholar] [CrossRef]
- Winfield, M.O.; Allen, A.M.; Burridge, A.J.; Barker, G.L.; Benbow, H.R.; Wilkinson, P.A.; Coghill, J.; Waterfall, C.; Davassi, A.; Scopes, G.; et al. High-density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol. J. 2016, 14, 1195–1206. [Google Scholar] [CrossRef]
- Turuspekov, Y.; Plieske, J.; Ganal, M.; Akhunov, E.; Abugalieva, S. Phylogenetic analysis of wheat cultivars in Kazakhstan based on the wheat 90 K single nucleotide polymorphism array. Plant Genet. Resour. 2017, 15, 29–35. [Google Scholar] [CrossRef]
- Ensembl Plants. Available online: http://plants.ensembl.org/Triticum_aestivum/Info/Index (accessed on 6 February 2022).
- Jain, M.; Nijhawan, A.; Arora, R.; Agarwal, P.; Ray, S.; Sharma, P.; Kapoor, S.; Tyagi, A.; Khurana, J.P. F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol. 2007, 143, 1467–1483. [Google Scholar] [CrossRef]
- Hong, M.J.; Kim, D.Y.; Kang, S.Y.; Kim, D.S.; Kim, J.B.; Seo, Y.W. Wheat F-box protein recruits proteins and regulates their abundance during wheat spike development. Mol. Biol. Rep. 2012, 39, 9681–9696. [Google Scholar] [CrossRef]
- Zoltowski, B.D.; Imaizumi, T. Structure and function of the ZTL/FKF1/LKP2 group proteins in Arabidopsis. Enzymes 2014, 35, 213–239. [Google Scholar]
- Song, X.J.; Huang, W.; Shi, M.; Zhu, M.Z.; Lin, H.X. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat. Genet. 2007, 39, 623–630. [Google Scholar] [CrossRef]
- Muehlbauer, F.J.; Burnell, D.G.; Bogyo, T.P.; Bogyo, M.T. Simulated Comparisons of Single Seed Descent and Bulk Population Breeding Methods 1. Crop. Sci. 1981, 21, 572–577. [Google Scholar] [CrossRef]
- Wilkinson, P.A.; Winfield, M.O.; Barker, G.L.A.; Allen, A.M.; Burridge, A.; Coghill, J.A.; Burridge, A.; Edwards, K.J. CerealsDB 2.0: An integrated resource for plant breeders and scientists. BMC Bioinform. 2012, 13, 219. [Google Scholar] [CrossRef]
- Miyagawa, T.; Nishida, N.; Ohashi, J.; Kimura, R.; Fujimoto, A.; Kawashima, M.; Koike, A.; Sasaki, T.; Tanii, H.; Otowa, T.; et al. Appropriate data cleaning methods for genome-wide association study. J. Hum. Genet. 2008, 53, 886–893. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Thorndike, R.L. Who belongs in the family? Psychometrika 1953, 18, 267–276. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes. 2012, 6, 288–295. [Google Scholar] [CrossRef]
- Quisenberry, K.S.; Reitz, L.P. Turkey wheat: The cornerstone of an empire. Agric. Hist. 1974, 48, 98–110. [Google Scholar]
- Olmstead, A.L.; Rhode, P.W. The red queen and the hard reds: Productivity growth in American wheat, 1800–1940. J. Econ. Hist. 2002, 62, 929–966. [Google Scholar] [CrossRef]
- Shegebaev, O.S. Scientific Support for Spring Wheat Production in Kazakhstan. Spring wheat in Kazakstan: Current status and future directions. In Proceedings of the Kazakhstan—CIMMYT Conference, Akmola, Kazakhstan, 22–24 September 1997; pp. 24–29. [Google Scholar]
- Charmet, G. Wheat domestication: Lessons for the future. C. R. Biol. 2011, 334, 212–220. [Google Scholar] [CrossRef]
- de Sousa, T.; Ribeiro, M.; Sabença, C.; Igrejas, G. The 10,000-year success story of wheat. Foods 2021, 10, 2124. [Google Scholar] [CrossRef]
- Almerekova, S.; Genievskaya, Y.; Abugalieva, S.; Sato, K.; Turuspekov, Y. Population structure and genetic diversity of two-rowed barley accessions from Kazakhstan based on SNP genotyping data. Plants 2021, 10, 2025. [Google Scholar] [CrossRef]
- Morgil, H.; Gercek, Y.C.; Tulum, I. Single nucleotide polymorphisms (SNPs) in plant genetics and breeding. In The Recent Topics in Genetic Polymorphisms; InTech Open: Rijeka, Croatia, 2020; pp. 800–825. [Google Scholar]
- Dzhunusova, M.; Ten, D.; Aubekerova, N.; Morgounov, A. history of wheat breeding in Kyrgyzstan. In The World Wheat Book: A history of Wheat Breeding; Bonjean, A.P., Angus, W.J., van Ginkel, M., Eds.; Intercept: Andover, UK, 2014; Volume 3, pp. 235–247. [Google Scholar]
- Ramadan, A.; Nemoto, K.; Seki, M.; Shinozaki, K.; Takeda, H.; Takahashi, H.; Sawasaki, T. Wheat germ-based protein libraries for the functional characterisation of the Arabidopsis E2 ubiquitin conjugating enzymes and the RING-type E3 ubiquitin ligase enzymes. BMC Plant Biol. 2015, 15, 275. [Google Scholar] [CrossRef]
- Xu, M.; Jin, P.; Liu, T.; Gao, S.; Zhang, T.; Zhang, F.; Han, X.; He, L.; Chen, J.; Yang, J. Genome-wide identification and characterization of UBP gene family in wheat (Triticum aestivum L.). PeerJ 2021, 9, e11594. [Google Scholar] [CrossRef]


| Groups | n | Ne | uh | % P |
|---|---|---|---|---|
| Europe | 440 | 1.486 ± 0.003 | 0.292 ± 0.002 | 99.26% |
| Australia | 16 | 1.468 ± 0.003 | 0.297 ± 0.002 | 82.68% |
| USA | 19 | 1.516 ± 0.003 | 0.325 ± 0.002 | 87.45% |
| Afghanistan | 32 | 1.420 ± 0.003 | 0.259 ± 0.002 | 82.87% |
| Turkey | 19 | 1.484 ± 0.003 | 0.308 ± 0.002 | 88.88% |
| Russia | 26 | 1.406 ± 0.003 | 0.258 ± 0.002 | 86.09% |
| Kazakhstan | 52 | 1.325 ± 0.003 | 0.212 ± 0.002 | 86.73% |
| Kyrgyzstan | 27 | 1.434 ± 0.003 | 0.270 ± 0.002 | 83.08% |
| Tajikistan | 9 | 1.418 ± 0.003 | 0.280 ± 0.002 | 69.71% |
| Uzbekistan | 27 | 1.460 ± 0.003 | 0.282 ± 0.002 | 87.78% |
| Total | 667 | 1.428 ± 0.001 | 0.267 ± 0.001 | 84.94 ± 2.31% |
| Origins | EU | AUS | USA | AFG | TUR | RUS | KAZ | KGS | TAJ |
|---|---|---|---|---|---|---|---|---|---|
| AUS | 0.084 | ||||||||
| USA | 0.064 | 0.041 | |||||||
| AFG | 0.280 | 0.183 | 0.168 | ||||||
| TUR | 0.131 | 0.091 | 0.067 | 0.082 | |||||
| RUS | 0.180 | 0.154 | 0.111 | 0.127 | 0.021 | ||||
| KAZ | 0.216 | 0.181 | 0.134 | 0.122 | 0.026 | 0.020 | |||
| KGS | 0.197 | 0.142 | 0.111 | 0.073 | 0.010 | 0.018 | 0.015 | ||
| TAJ | 0.196 | 0.129 | 0.104 | 0.077 | 0.012 | 0.026 | 0.042 | 0.012 | |
| UZB | 0.207 | 0.135 | 0.111 | 0.050 | 0.010 | 0.027 | 0.028 | 0.003 | 0.007 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Amalova, A.; Yermekbayev, K.; Griffiths, S.; Winfield, M.O.; Morgounov, A.; Abugalieva, S.; Turuspekov, Y. Population Structure of Modern Winter Wheat Accessions from Central Asia. Plants 2023, 12, 2233. https://doi.org/10.3390/plants12122233
Amalova A, Yermekbayev K, Griffiths S, Winfield MO, Morgounov A, Abugalieva S, Turuspekov Y. Population Structure of Modern Winter Wheat Accessions from Central Asia. Plants. 2023; 12(12):2233. https://doi.org/10.3390/plants12122233
Chicago/Turabian StyleAmalova, Akerke, Kanat Yermekbayev, Simon Griffiths, Mark Owen Winfield, Alexey Morgounov, Saule Abugalieva, and Yerlan Turuspekov. 2023. "Population Structure of Modern Winter Wheat Accessions from Central Asia" Plants 12, no. 12: 2233. https://doi.org/10.3390/plants12122233
APA StyleAmalova, A., Yermekbayev, K., Griffiths, S., Winfield, M. O., Morgounov, A., Abugalieva, S., & Turuspekov, Y. (2023). Population Structure of Modern Winter Wheat Accessions from Central Asia. Plants, 12(12), 2233. https://doi.org/10.3390/plants12122233








