Allium ducissae (A. subgen. Polyprason, Amaryllidaceae) a New Species from the Central Apennines (Italy)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Morphological Analyses
2.2. Chromosome Count
2.3. Scanning Electron Microscopic Analyses
2.4. Molecular Analyses
3. Results
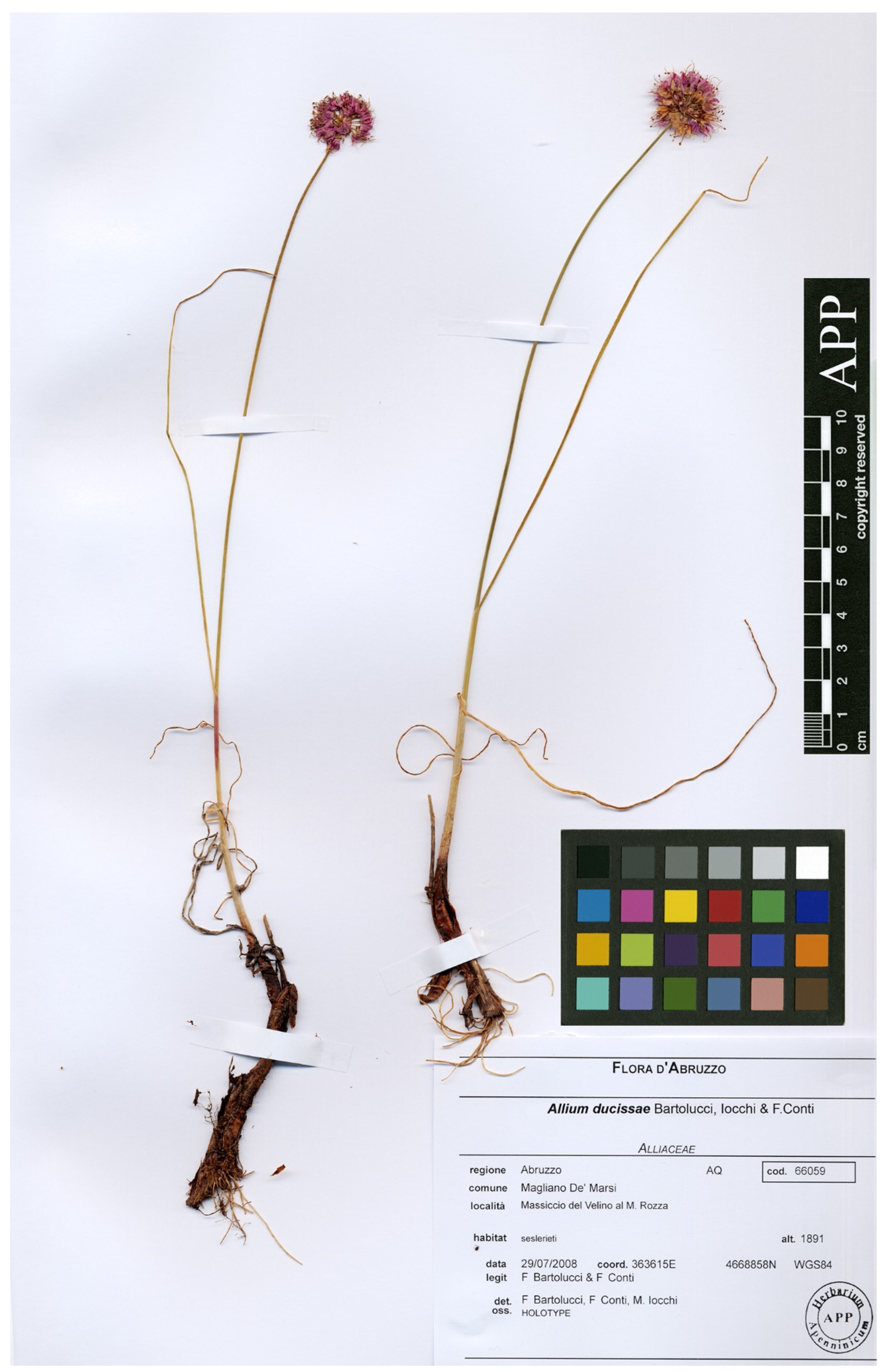
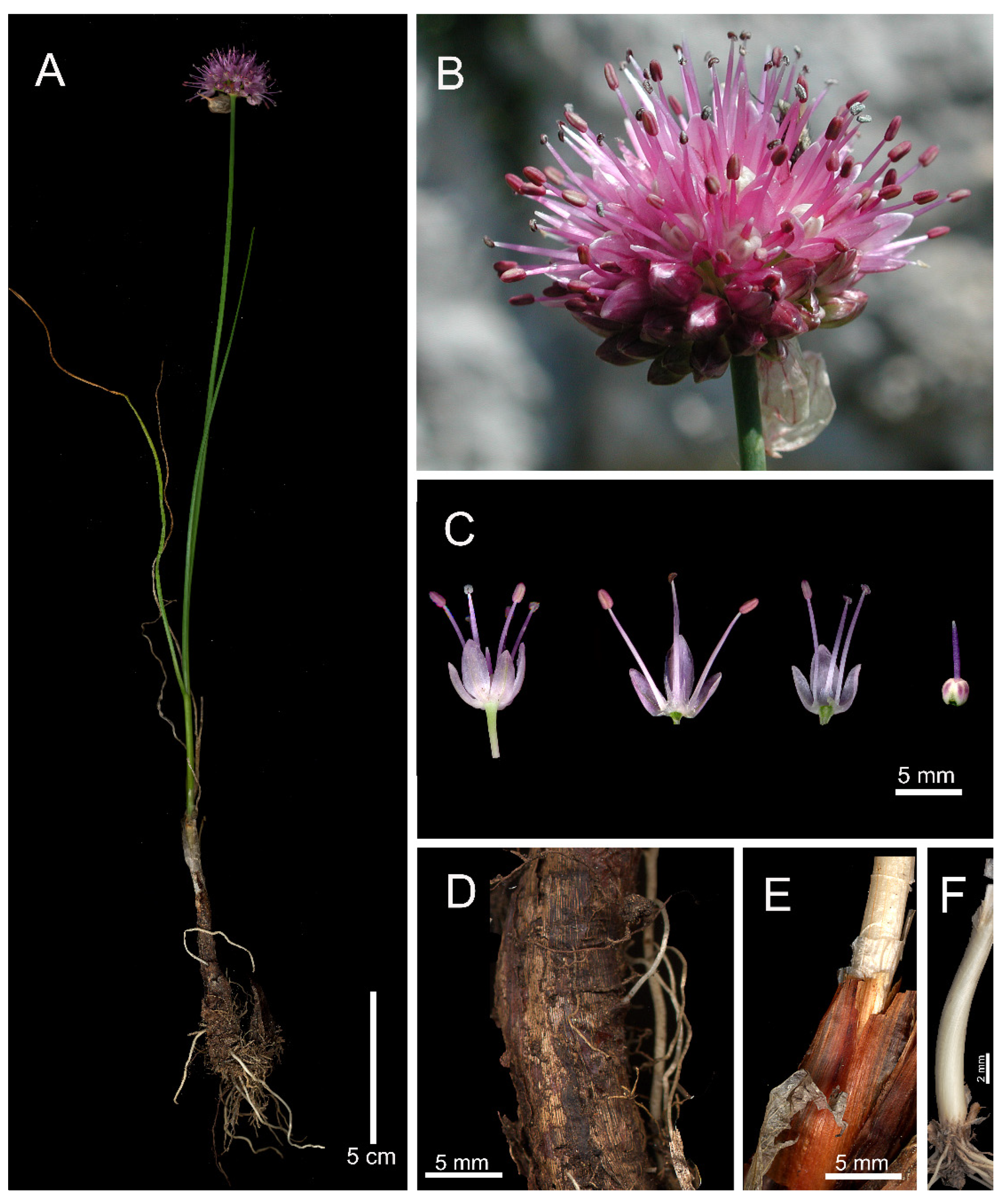
3.1. Taxonomic Treatment
3.2. Morphological Analyses
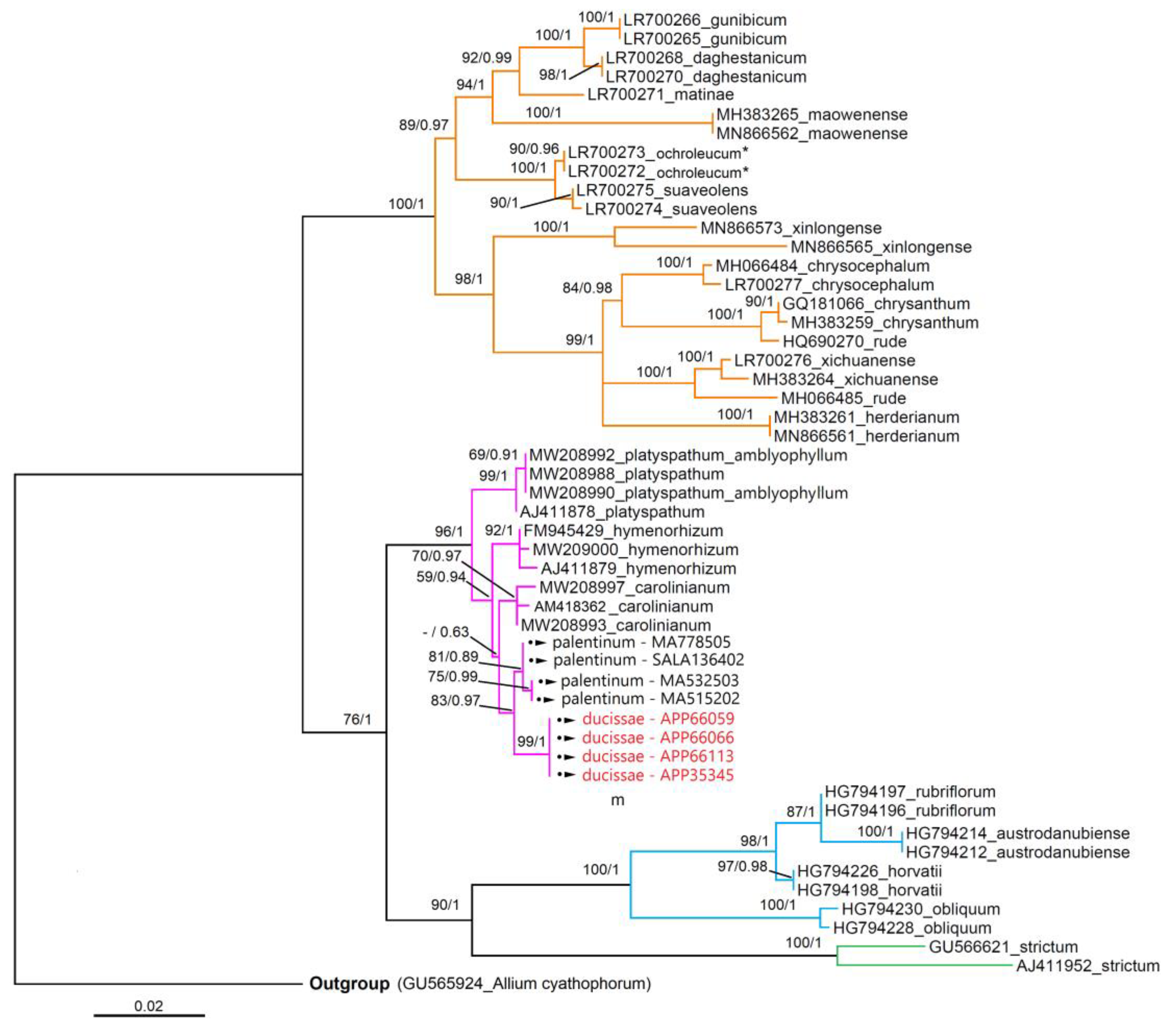
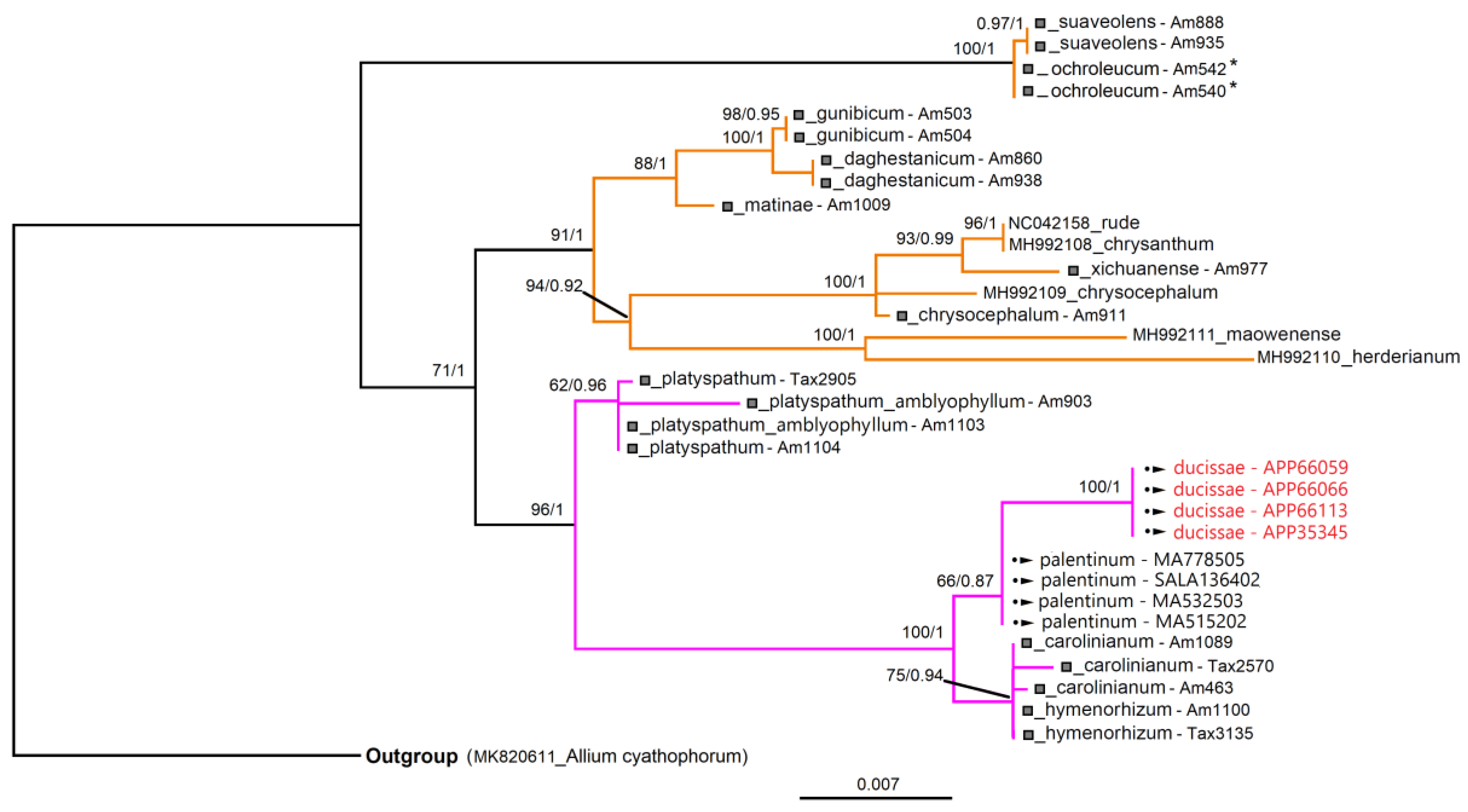
3.3. Molecular Analyses
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Khassanov, F.O. Taxonomical and ethnobotanical aspects of Allium species from Middle Asia with particular reference to subgenus Allium. In The Allium Genomes. Compendium of Plant Genomes; Shigyo, M., Khar, A., Abdelrahman, M., Eds.; Springer: Cham, Switzerland, 2018; pp. 11–21. [Google Scholar]
- POWO. Plants of the World Online. Facilitated by the Royal Botanic Gardens, Kew. Available online: http://www.plantsoftheworldonline.org/ (accessed on 3 November 2021).
- De Sarker, D.; Johnson, M.A.T.; Reynolds, A.; Brandham, P.E. Cytology of the highly polyploid disjunct species, Allium dregeanum(Alliaceae), and of some Eurasian relatives. Bot. J. Linn. Soc. 1997, 124, 361–373. [Google Scholar] [CrossRef][Green Version]
- Friesen, N.; Fritsch, R.M.; Blattner, F. Phylogeny and new intrageneric classification of Allium (Alliaceae) based on nuclear ribosomal DNA its sequences. Aliso 2006, 22, 372–395. [Google Scholar] [CrossRef]
- Li, Q.-Q.; Zhou, S.-D.; He, X.-J.; Yu, Y.; Zhang, Y.-C.; Wei, X.-Q. Phylogeny and biogeography of Allium (Amaryllidaceae: Allieae) based on nuclear ribosomal internal transcribed spacer and chloroplast rps16 sequences, focusing on the inclusion of species endemic to China. Ann. Bot. 2010, 106, 709–733. [Google Scholar] [CrossRef] [PubMed]
- Fritsch, R.M.; Blattner, F.; Gurushidze, M. New classification of Allium L. subg. Melanocrommyum (Webb & Berthel.) Rouy (Alliaceae) based on molecular and morphological characters. Phyton (Horn) 2010, 49, 145–220. [Google Scholar]
- Huang, D.-Q.; Yang, J.-T.; Zhou, C.-J.; Zhou, S.-D.; He, X.-J. Phylogenetic reappraisal of Allium subgenus Cyathophora (Amaryllidaceae) and related taxa, with a proposal of two new sections. J. Plant Res. 2014, 127, 275–286. [Google Scholar] [CrossRef]
- Brullo, S.; Brullo, C.; Cambria, S.; Giusso del Galdo, G.; Salmeri, C. Allium albanicum (Amaryllidaceae), a new species from balkans and its relationships with A. meteoricum Heldr. & Hausskn. ex Halácsy. PhytoKeys 2019, 119, 117–136. [Google Scholar]
- Bartolucci, F.; Peruzzi, L.; Galasso, G.; Albano, A.; Alessandrini, A.; Ardenghi, N.M.G.; Astuti, G.; Bacchetta, G.; Ballelli, S.; Banfi, E.; et al. An updated checklist of the vascular flora native to Italy. Plant Biosyst. 2018, 152, 179–303. [Google Scholar] [CrossRef]
- Galasso, G.; Conti, F.; Peruzzi, L.; Ardenghi, N.M.G.; Banfi, E.; Celesti-Grapow, L.; Albano, A.; Alessandrini, A.; Bacchetta, G.; Ballelli, S.; et al. An updated checklist of the vascular flora alien to Italy. Plant Biosyst. 2018, 152, 556–592. [Google Scholar] [CrossRef]
- Petriccione, B. Flora e Vegetazione del Massiccio del Monte Velino (Appennino centrale). Collana Verde 1993, 92, 3–261. [Google Scholar]
- Dal Vesco, G.; Garbari, F.; Giordani, A. Il genere Allium L. (Alliaceae) in Italia. XX. Allium strictum Schrader. Webbia 2003, 58, 401–410. [Google Scholar] [CrossRef]
- Iocchi, M.; Bartolucci, F.; Carotenuto, L.; Valfré, D.; Cutini, M.; Theurillat, J.P. Note floristiche per la Riserva Naturale Regionale delle “Montagne della Duchessa” (Lazio Nord-Orientale). Inf. Bot. Ital. 2011, 42, 503–508. [Google Scholar]
- Brullo, S.; Guarino, R. Allium L. In Flora d’Italia, 2nd ed.; Pignatti, S., Guarino, R., La Rosa, M., Eds.; Edagricole: Bologna, Italy, 2017; Volume 1, pp. 238–269. [Google Scholar]
- De Santis, E.; Soldati, R. Lucoli e i suoi Fiori; CM Grafica: L’Aquila, Italy, 2019; p. 357. [Google Scholar]
- Schrader, H.A. Hortus Gottingensis. Seu Plantae Novae et Rariores Horti Regii Botanici Gottingensis; Dieterich: Göttingen, Germany, 1809. [Google Scholar]
- Kamelin, R.V. Florogeneticheskij Analiz Estestvennoj Flory Gornoj Srednej Azii; Nauka: Leningrad, Russia, 1973; p. 354. [Google Scholar]
- Friesen, N. The genus Allium L. in the flora of Mongolia. Feddes Repert. 1995, 106, 59–81. [Google Scholar] [CrossRef]
- Sinitsyna, T.A. Genus Allium L. (Alliaceae) in Siberia. Vavilovia 2019, 2, 3–22. [Google Scholar] [CrossRef]
- Anzalone, B.; Iberite, M.; Lattanzi, E. La flora vascolare del Lazio. Inf. Bot. Ital. 2010, 42, 187–317. [Google Scholar]
- Bartolucci, F.; Iocchi, M.; Thurillat, J.-P.; Conti, F. Indagini preliminari sulle popolazioni di Allium strictum (Amaryllidaceae) dell’Appennino Centrale. Not. Soc. Bot. Ital. 2020, 4, 139–140. [Google Scholar]
- Losa, T.M.; Montserrat, P. Nueva Aportación al Estudio de la Flora de los Montes Cántabro-Leoneses. An. Inst. Bot. A. J. Cavanilles 1954, 11, 385–462. [Google Scholar]
- Pastor, J.; Valdes, B. Revision del genero Allium (Liliaceae) en la Peninsula Iberica e Islas Baleares; Secretariado de Publicaciones de la Universidad de Sevilla: Sevilla, Spain, 1983. [Google Scholar]
- Aedo, C. Allium L. In Flora Iberica. Plantas Vasculares de la Península Ibérica e Islas Baleares; Castroviejo, S., Ed.; CSIC: Madrid, Spain, 2013; Volume XX, pp. 220–273. [Google Scholar]
- Friesen, N.; Abbasi, N.; Murtazaliev, R.; Fritsch, R. Allium matinae, a new species from Northwestern Iran. Phytotaxa 2020, 433, 181–189. [Google Scholar] [CrossRef]
- Bartolucci, F.; Iocchi, M.; De Castro, O.; Conti, F. Typification and correct place of publication of the name Allium kermesinum (Amaryllidaceae). Novon 2017, in press. [Google Scholar]
- Stearn, W.T. Allium L. In Flora Europaea; Tutin, T.G., Heywood, V.H., Burges, N.A., Valentine, D.H., Walters, S.M., Webb, D.A., Eds.; Cambridge University Press: Cambridge, UK, 1980; Volume 5, pp. 49–69. [Google Scholar]
- Miceli, P.; Garbari, F.; Charpin, A. Sur quelques Allium de la section Rhizirideum G. Don ex Koch. Candollea 1987, 42, 627–643. [Google Scholar]
- Tison, J.-M.; de Foucault, B. Flora Gallica. Flore de France; Biotope Éditons: Mèze, France, 2014. [Google Scholar]
- Thiers, B. Index Herbariorum: A Global Directory of Public Herbaria and Associated Staff. New York Botanical Garden’s Virtual Herbarium. Available online: http://sweetgum.nybg.org/science/ih/ (accessed on 26 June 2020).
- IBM. IBM SPSS Statistics for Windows; Version 25.0; IBM Corp.: Armonk, NY, USA, 2017. [Google Scholar]
- Hammer, Ø.H.D.; Ryan, P.D. PAST: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol. Electron. 2001, 4, 1–9. [Google Scholar]
- Hammer, Ø.H.D. PAST Version 4.04; Natural History Museum: Oslo, Norway, 2020; Available online: http://folk.uio.no/ohammer/past/ (accessed on 20 December 2021).
- Barthlott, W. Epidermal and Seed Surface Characters of Plants: Systematic Applicability and Some Evolutionary Aspects. Nord. J. Bot. 1981, 1, 345–355. [Google Scholar] [CrossRef]
- Baasanmunkh, S.; Lee, J.K.; Jang, J.E.; Park, M.S.; Friesen, N.; Chung, S.; Choi, H.J. Seed morphology of Allium L. (Amaryllidaceae) from Central Asian countries and its taxonomic implications. Plants 2020, 9, 1239. [Google Scholar] [CrossRef] [PubMed]
- De Castro, O.; Geraci, A.; Mannino, A.M.; Mormile, N.; Santangelo, A.; Troia, A. A Contribution to the characterization of Ruppia drepanensis (Ruppiaceae), a key species of threatened Mediterranean wetlands. Ann. Mo. Bot. Gard. 2021, 106, 1–9. [Google Scholar] [CrossRef]
- Fritsch, R.M.; Friesen, N. Allium oreotadzhikorum and Allium vallivanchense, two new species of Allium Subg. Polyprason (Alliaceae) from the Central Asian Republic Tajikistan. Feddes Repert. 2009, 120, 221–231. [Google Scholar] [CrossRef]
- Seregin, A.P.; Anačkov, G.; Friesen, N. Molecular and morphological revision of the Allium saxatile group (Amaryllidaceae): Geographical isolation as the driving force of underestimated speciation. Bot. J. Linn. Soc. 2015, 178, 67–101. [Google Scholar] [CrossRef]
- Xie, D.-F.; Xie, F.-M.; Jia, S.-B.; Li, H.; Yang, X.; Zhang, X.-Y.; Zhou, S.-D.; He, X.-J. Allium xinlongense (Amaryllidaceae, Allioideae), a new species from Western Sichuan. Phytotaxa 2020, 432, 274–282. [Google Scholar] [CrossRef]
- Friesen, N.V.P.; Osmonaly, B.; Sitpayeva, G.; Luferov, A.; Shmakov, A. Allium toksanbaicum (Amaryllidaceae), a new species from Southeast Kazakhstan. Phytotaxa 2021, 494, 251–267. [Google Scholar] [CrossRef]
- Li, M.-J.; Yu, H.-X.; Guo, X.-L.; He, X.-J. Out of the Qinghai–Tibetan Plateau and rapid radiation across Eurasia for Allium section Daghestanica (Amaryllidaceae). AoB PLANTS 2021, 13, plab017. [Google Scholar] [CrossRef]
- Aceto, S.; Caputo, P.; Cozzolino, S.; Gaudio, L.; Moretti, A. Phylogeny and evolution of Orchis and allied genera based on ITS DNA Variation: Morphological gaps and molecular continuity. Mol. Phylogenet. Evolut. 1999, 13, 67–76. [Google Scholar] [CrossRef]
- De Castro, O.; Di Maio, A.; Lozada García, J.A.; Piacenti, D.; Vázquez-Torres, M.; De Luca, P. Plastid DNA sequencing and nuclear SNP genotyping help resolve the puzzle of Central American Platanus. Ann. Bot. 2013, 112, 589–602. [Google Scholar] [CrossRef][Green Version]
- White, T.J.B.T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Gelfand, D.H., Sninsky, J.J., White, T.J., Innis, M.A., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar] [CrossRef]
- Taberlet, P.; Gielly, L.; Pautou, G.; Bouvet, J. Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol. Biol. 1991, 17, 1105–1109. [Google Scholar] [CrossRef]
- Shaw, J.; Lickey, E.B.; Schilling, E.E.; Small, R.L. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: The tortoise and the hare III. Am. J. Bot. 2007, 94, 275–288. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A User-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior summarization in bayesian phylogenetics using tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef]
- Kozlov, A.M.; Darriba, D.; Flouri, T.; Morel, B.; Stamatakis, A. RAxML-NG: A Fast, Scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 2019, 35, 4453–4455. [Google Scholar] [CrossRef]
- Fernandez Casas, J. Números cromosómicos de plantas Españolas, IV. An. Inst. Bot. A. J. Cavanilles 1977, 34, 335–349. [Google Scholar]
- IUCN. Guidelines for Using the IUCN Red List Categories and Criteria. Version 14. Prepared by the Standards and Petitions Subcommittee, Cambridge. Available online: https://www.iucnredlist.org/resources/redlistguidelines (accessed on 26 November 2021).
- QGIS Development Team. QGIS Geographic Information System. Open Source Geospatial Foundation Project. 2020.
- Celep, F.; Koyuncu, M.; Fritsch, R.M.; Kahraman, A.; Doan, M. Taxonomic importance of seed morphology in Allium (Amaryllidaceae). Syst. Bot. 2012, 37, 893–912. [Google Scholar] [CrossRef]
- Lin, C.-Y.; Tan, D.-Y. Seed Testa Micromorphology of thirty-eight species of Allium (Amaryllidaceae) from Central Asia, and its taxonomic implications. Nord. J. Bot. 2017, 35, 189–200. [Google Scholar] [CrossRef]
- Seregin, A.P. Allium marmoratum (Amaryllidaceae), a new species of section Falcatifolia from Chimgan Massif, Eastern Uzbekistan. Phytotaxa 2015, 205, 211–214. [Google Scholar] [CrossRef]
- Seregin, A.P.; Friesen, N. Molecular and morphological revision reveals different evolution patterns in Allium sect. Oreiprason and sect. Falcatifolia (Amaryllidaceae). In Semicentenary after Konstantin Meyer: XIII Moscow Symposium on Plant Phylogeny: Proceedings of the International Conference; Timonin, A.C., Ed.; MAKS Press: Moscow, Russia, 2015; pp. 353–356. [Google Scholar] [CrossRef]
- Conti, F.; Bartolucci, F. The Vascular Flora of the National Park of Abruzzo, Lazio and Molise (Central Italy): An Annotated Checklist; Geobotany Studies Series; Springer: Berlin/Heidelberg, Germany, 2015; p. 254. [Google Scholar]
- Conti, F.; Bartolucci, F. The vascular flora of Gran Sasso and Monti della Laga National Park (Central Italy). Phytotaxa 2016, 256, 1–119. [Google Scholar] [CrossRef]
- Conti, F.; Ciaschetti, G.; Martino, L.D.; Bartolucci, F. An annotated checklist of the vascular flora of Majella National Park (Central Italy). Phytotaxa 2019, 412, 1–90. [Google Scholar] [CrossRef]
- Favarger, C. Endemism in the montane floras of Europe. In Taxonomy, Phytogeography and Evolution; Valentine, D.H., Ed.; Academic Press: London, UK; New York, NY, USA, 1972; pp. 191–204. [Google Scholar]
- Favarger, C. Endémisme, biosystématique et conservation du patrimoin génétique. Atti Inst. Bot. Lab. Crittogam. Univ. Pavia 1986, 5, 5–14. [Google Scholar]
- Tammaro, F. Diversità floristica delle montagne Abruzzesi. Ann. Musei Civici-Rovereto. Sez. Archeol. Storia, Sci. Nat. 2000, 14, 147–176. [Google Scholar]
- Conti, F. La flora ipsofila dell’Appennino Centrale: Ricchezza ed endemiti. Inf. Bot. Ital. 2004, 35, 383–386. [Google Scholar]
- Conti, F.; Bartolucci, F.; Di Martino, L.; Manzi, A. La Flora Endemica Minacciata Delle Montagne Italiane; CAI-Club Alpino Italiano: Milano, Italy, 2018; Volume 33, p. 487. [Google Scholar]
- Conti, F.; Bartolucci, F. Anthyllis apennina (Fabaceae), a new species from Central Apennine (Italy). PhytoKeys 2021, 176, 111–129. [Google Scholar] [CrossRef]
- Conti, F.; Bracchetti, L.; Uzunov, D.; Bartolucci, F. A new subspecies of Corydalis densiflora (Papaveraceae) from the Apennines (Italy). Willdenowia 2019, 49, 53–64. [Google Scholar] [CrossRef]
- Conti, F.; Uzunov, D. Crepis magellensis F. Conti & Uzunov (Asteraceae), a new species from Central Apennine (Abruzzo, Italy). Candollea 2011, 66, 81–86. [Google Scholar] [CrossRef]
- Peruzzi, L.; Bartolucci, F.; Frignani, F.; Minutillo, F. Gagea tisoniana, a new species of Gagea Salisb. sect. Gagea (Liliaceae) from Central Italy. Bot. J. Linn. Soc. 2007, 155, 337–347. [Google Scholar] [CrossRef]
- Conti, F. Morphometric study and taxonomy of Genista pulchella Vis. s.l. (Fabaceae), a South European species. Bot. J. Linn. Soc. 2007, 153, 245–254. [Google Scholar] [CrossRef]
- Conti, F. A new species of Lathyrus L. (Fabaceae) from Central Apennine (Italy). Plant Biosyst. 2010, 144, 814–818. [Google Scholar] [CrossRef]
- Conti, F.; Pennesi, R.; Uzunov, D.; Bracchetti, L.; Bartolucci, F. A new species of Oxytropis (Fabaceae) from Central Apennines (Italy). Phytotaxa 2018, 336, 69–81. [Google Scholar] [CrossRef]
- Conti, F.; Peruzzi, L. Pinguicula (Lentibulariaceae) in Central Italy: Taxonomic study. Ann. Bot. Fenn. 2006, 43, 321–337. [Google Scholar]
- Conti, F.; Martino, L.D.; Bartolucci, F. Poa magellensis (Poaceae), a new species from Central Apennine (Italy). PhytoKeys 2020, 144, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Conti, F.; Proietti, E.; Ogwu, M.C.; Gubellini, L.; Bartolucci, F. Re-evaluation of Senecio apenninus (Asteraceae, Senecioneae). Willdenowia 2019, 49, 329–341. [Google Scholar] [CrossRef]
- Conti, F.; Bartolucci, F. Ranunculus giordanoi sp. nov. from the R. auricomus complex (ranunculaceae), Central Apennines (Italy). Nord. J. Bot. 2017, 35, 322–327. [Google Scholar] [CrossRef]
- Dunkel, F.G. The Ranunculus auricomus L. complex (Ranunculaceae) in Central and Southern Italy, with additions to north italian taxa. Webbia 2011, 66, 165–193. [Google Scholar] [CrossRef]
- Gallo, L.; Conti, F. On the true identity of the plants recently referred to Sedum nevadense (Crassulaceae) in Abruzzo (Italy). Phytotaxa 2015, 239, 43–54. [Google Scholar] [CrossRef]
- Orsenigo, S.; Montagnani, C.; Fenu, G.; Gargano, D.; Peruzzi, L.; Abeli, T.; Alessandrini, A.; Bacchetta, G.; Bartolucci, F.; Bovio, M.; et al. Red Listing plants under full national responsibility: Extinction risk and threats in the vascular flora endemic to Italy. Biol. Conserv. 2018, 224, 213–222. [Google Scholar] [CrossRef]
- Abeli, T.; Albani Rocchetti, G.; Barina, Z.; Bazos, I.; Draper, D.; Grillas, P.; Iriondo, J.M.; Laguna, E.; Moreno-Saiz, J.C.; Bartolucci, F. Seventeen ‘extinct’ plant species back to conservation attention in Europe. Nat. Plants 2021, 7, 282–286. [Google Scholar] [CrossRef] [PubMed]



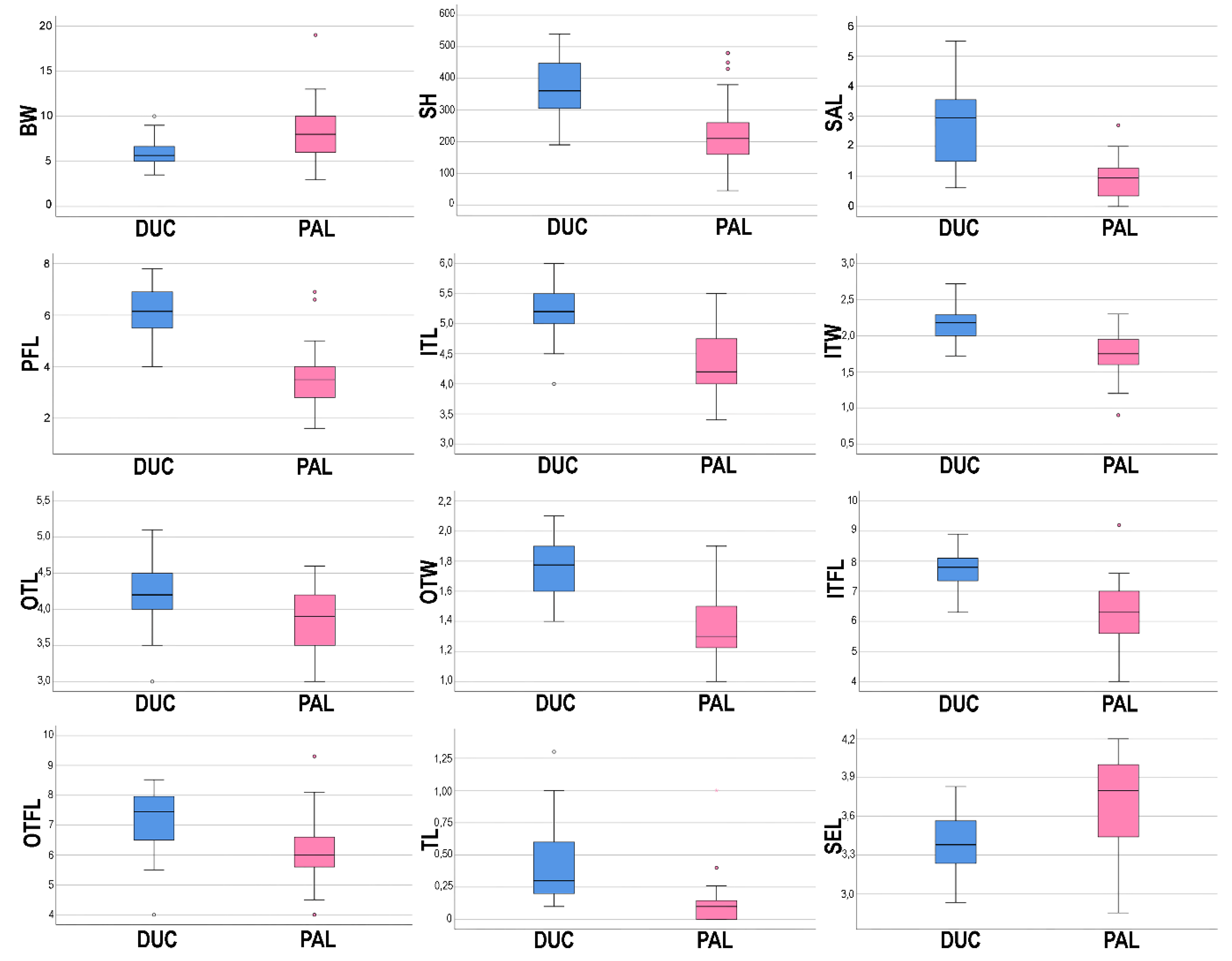
| Abbreviations | Characters | |
|---|---|---|
| 1 | BL | Bulb length (mm) |
| 2 | BW | Bulb width (mm) |
| 3 | SH | Stem height (mm) |
| 4 | SW | Stem width under the inflorescence (mm) |
| 5 | LW | Leaf max width (mm) |
| 6 | SAL | Spathe appendage length (mm) |
| 7 | ID | Inflorescence diameter (mm) |
| 8 | PFL | Pedicel flower length (mm) |
| 9 | ITL | Inner tepal length (mm) |
| 10 | ITW | Inner tepal width (mm) |
| 11 | OTL | Outer tepal length (mm) |
| 12 | OTW | Outer tepal width (mm) |
| 13 | ITFL | Inner filament length (mm) |
| 14 | OTFL | Outer filament length (mm) |
| 15 | IFTL | Tooth of inner filament length (mm) |
| 16 | AL | Anther length (mm) |
| 17 | OL | Ovary length (mm) |
| 18 | SL | Style length (mm) |
| 19 | SEL | Seed length (mm) |
| 20 | NL | Numbers of leaves |
| 21 | NF | Numbers of flowers |
| Character | Allium ducissae | Allium palentinum | |||
|---|---|---|---|---|---|
| 1 | BL | 62.1 ± 14.5 | (35)50.5–70(95.8) | 57.3 ± 14.7 | (33)45–69.1(86.5) |
| 2 | BW | 5.93 ± 1.45 | (3.5)5–6.68(10) | 8.44 ± 2.97 | (3)6–10(19) |
| 3 | SH | 364.81 ± 86.67 | (190)302.5–448.75(540) | 226.73 ± 104.8 | (45)157–260(480) |
| 4 | SW | 1.13 ± 0.27 | (0.8)0.9–1.3(1.8) | 1.29 ± 0.33 | (0.7)1–1.6(2) |
| 5 | LW | 1.95 ± 0.5 | (1)1.5–2.2(3.5) | 2.16 ± 0.78 | (1.1)1.5–2.8(4.6) |
| 6 | SAL | 2.72 ± 1.31 | (0.62)1.5–3.58(5.5) | 0.88 ± 0.69 | (0)0.33–1.29(2.7) |
| 7 | ID | 20.98 ± 4.5 | (15)16.88–23.36(32) | 18.23 ± 3.73 | (13)15–20(27) |
| 8 | PFL | 6.15 ± 0.83 | (4)5.5–6.9(7.8) | 3.51 ± 1.11 | (1.6)2.75–4(6.9) |
| 9 | ITL | 5.25 ± 0.45 | (4)5–5.5(6) | 4.34 ± 0.52 | (3.4)4–4.9(5.5) |
| 10 | ITW | 2.16 ± 0.24 | (1.72)2–2.29(2.72) | 1.74 ± 0.3 | (0.9)1.6–2(2.3) |
| 11 | OTL | 4.25 ± 0.42 | (3)4–4.5(5.1) | 3.84 ± 0.42 | (3)3.5–4.2(4.6) |
| 12 | OTW | 1.73 ± 0.17 | (1.4)1.6–1.9(2.1) | 1.37 ± 0.23 | (1)1.2–1.5(1.9) |
| 13 | ITFL | 7.77 ± 0.59 | (6.3)7.33–8.15(8.9) | 6.33 ± 1.03 | (4)5.55–7(9.2) |
| 14 | OTFL | 7.15 ± 0.99 | (4)6.5–7.98(8.52) | 6.12 ± 1.13 | (4)5.55–6.7(9.3) |
| 15 | IFTL | 0.41 ± 0.29 | (0.1)0.2–0.6(1.3) | 0.18 ± 0.04 | (0.02)0.1–0.19(1) |
| 16 | AL | 1.15 ± 0.25 | (0.7)0.93–1.3(1.6) | 1.13 ± 0.22 | (0.8)1–1.3(1.5) |
| 17 | OL | 1.92 ± 0.13 | (1.6)1.8–2(2.2) | 1.83 ± 0.3 | (1.2)1.63–2(2.5) |
| 18 | SL | 4.28 ± 1.44 | (1.66)3–5.52(6.64) | 4.01 ± 1.19 | (2)3–5(6.6) |
| 19 | SEL | 3.39 ± 0.2 | (2.93)3.23–3.57(3.83) | 3.68 ± 0.38 | (2.85)3.41–4(4.2) |
| 20 | NL | (2)3–4(6) | (2)3–4(5) | ||
| 21 | NF | (14)22.25–36.5(60) | (6)14–30(60) | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bartolucci, F.; Iocchi, M.; De Castro, O.; Conti, F. Allium ducissae (A. subgen. Polyprason, Amaryllidaceae) a New Species from the Central Apennines (Italy). Plants 2022, 11, 426. https://doi.org/10.3390/plants11030426
Bartolucci F, Iocchi M, De Castro O, Conti F. Allium ducissae (A. subgen. Polyprason, Amaryllidaceae) a New Species from the Central Apennines (Italy). Plants. 2022; 11(3):426. https://doi.org/10.3390/plants11030426
Chicago/Turabian StyleBartolucci, Fabrizio, Marco Iocchi, Olga De Castro, and Fabio Conti. 2022. "Allium ducissae (A. subgen. Polyprason, Amaryllidaceae) a New Species from the Central Apennines (Italy)" Plants 11, no. 3: 426. https://doi.org/10.3390/plants11030426
APA StyleBartolucci, F., Iocchi, M., De Castro, O., & Conti, F. (2022). Allium ducissae (A. subgen. Polyprason, Amaryllidaceae) a New Species from the Central Apennines (Italy). Plants, 11(3), 426. https://doi.org/10.3390/plants11030426







