Impacts of Chronic Habitat Fragmentation on Genetic Diversity of Natural Populations of Prunus persica in China
Abstract
1. Introduction
2. Results
2.1. Characteristics of 23 nSSR Loci
2.2. Genetic Diversity
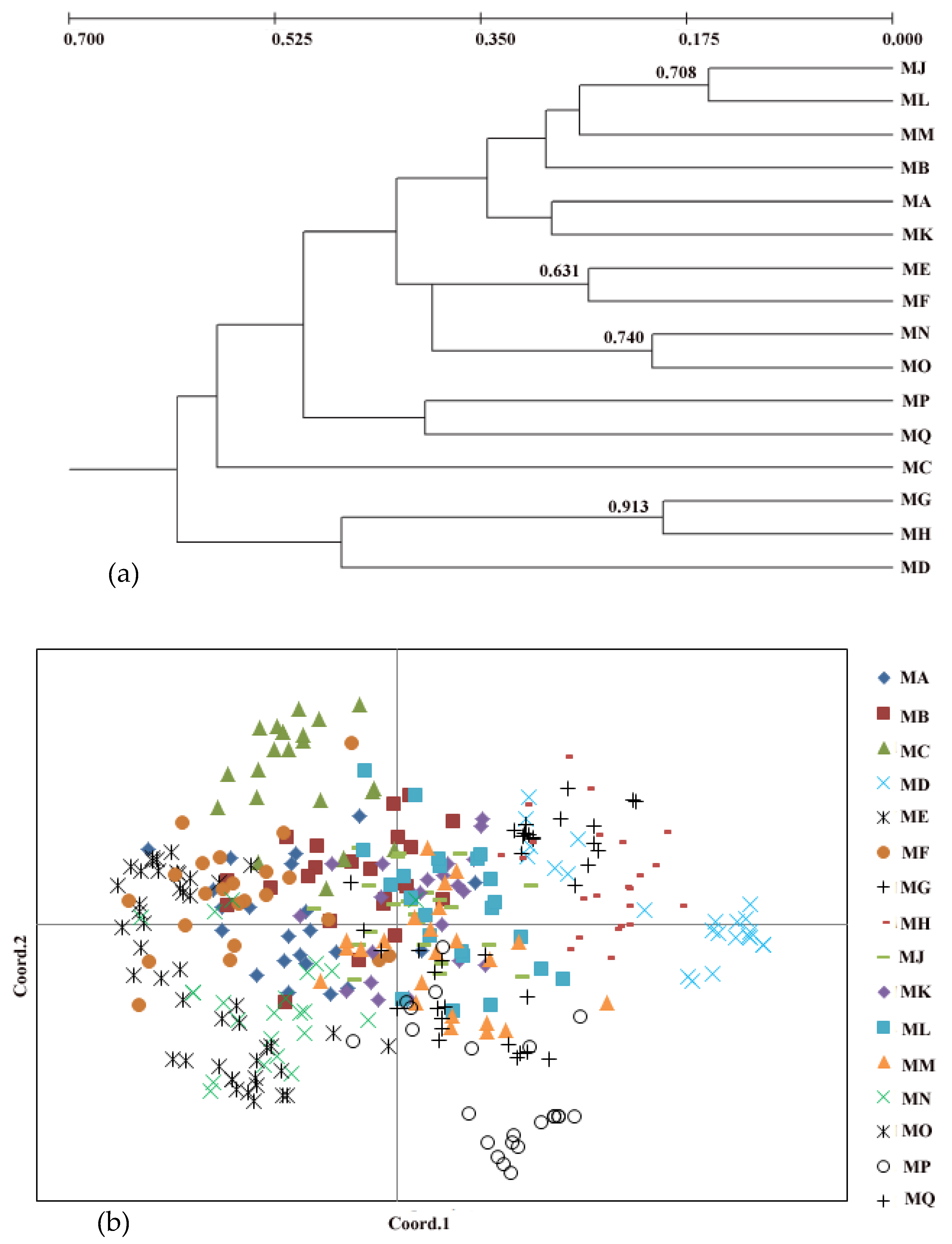
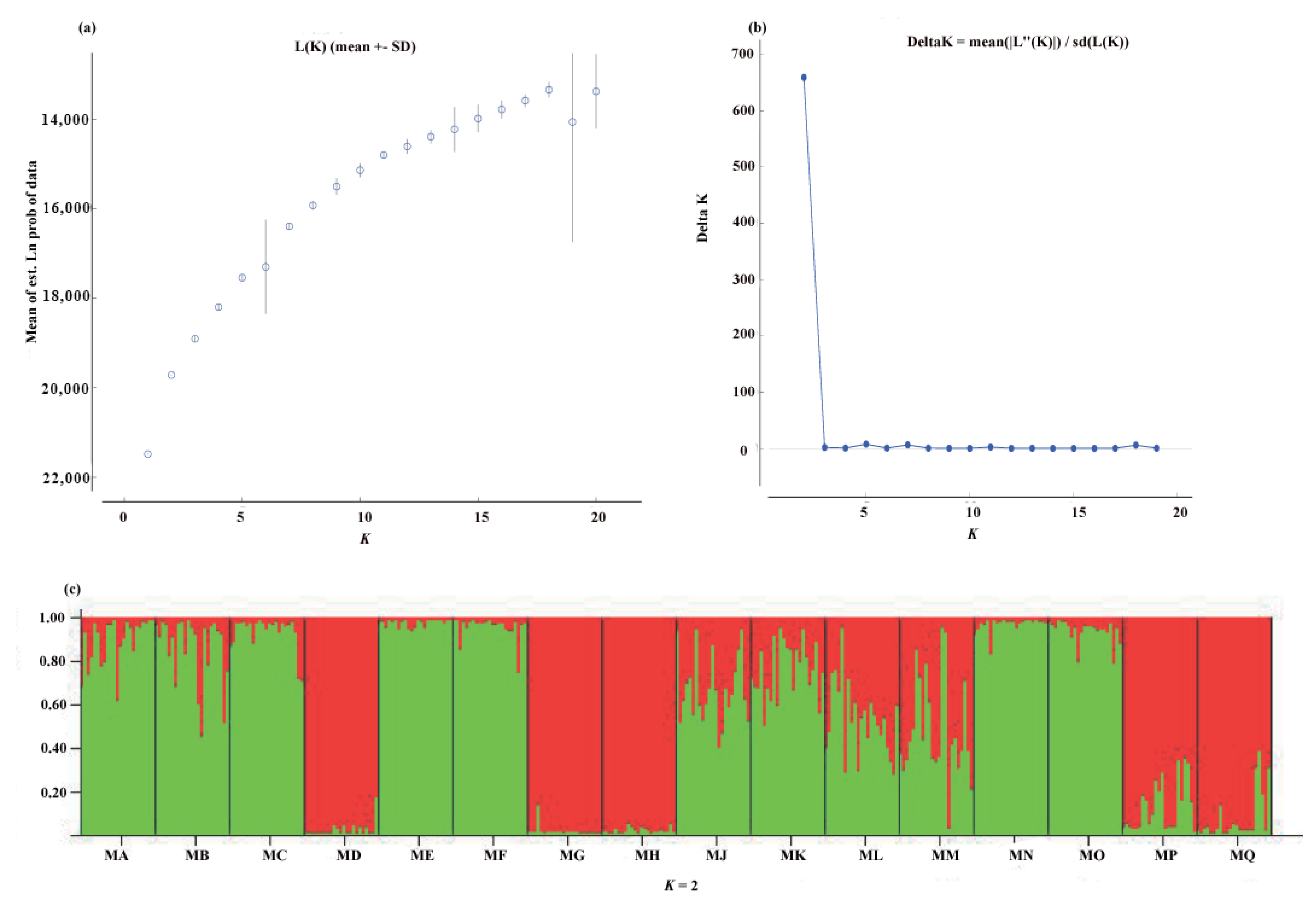
2.3. Population Structure
2.4. Mutation–Drift Equilibrium
2.5. Historical Gene Flow vs. Contemporary Gene Flow
3. Discussion
3.1. Genetic Diversity of Wild Populations
3.2. Genetic Structure
3.3. Implication for Genetic Conservation
4. Materials and Methods
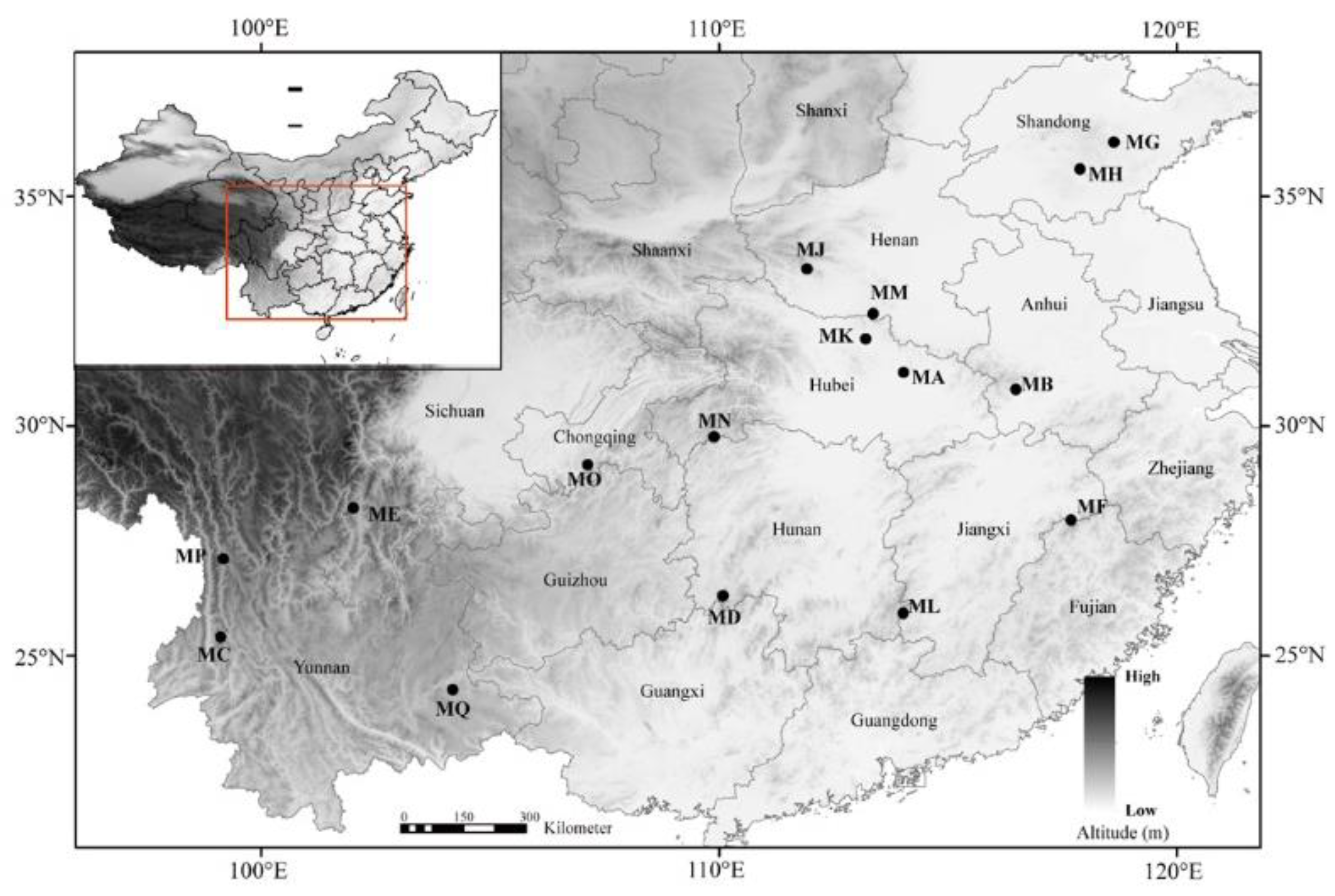
4.1. Plant Material
4.2. Microsatellite Genotyping
4.3. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Young, A.; Boyle, T.; Brown, T. The population genetic consequences of habitat fragmentation for plants. Trends Ecol. Evol. 1996, 11, 413–418. [Google Scholar] [CrossRef]
- Frankham, R.; Ballou, J.D.; Briscoe, D.A. Introduction to Conservation Genetics; Cambridge University Press: Cambridge, UK, 2002. [Google Scholar]
- Lande, R. Genetics and demography in biological conservation. Science 1988, 241, 1455–1460. [Google Scholar] [CrossRef] [PubMed]
- Yao, X.; Ye, Q.; Kang, M.; Huang, H. Microsatellite analysis reveals interpopulation differentiation and gene flow in the endangered tree Changiostyrax dolichocarpa (Styracaceae) with fragmented distribution in central China. New Phytol. 2007, 176, 472–480. [Google Scholar] [CrossRef] [PubMed]
- Honnay, O.; Jacquemyn, H. Susceptibility of common and rare plant species to the genetic consequences of habitat fragmentation. Conserv. Biol. 2007, 21, 823–831. [Google Scholar] [CrossRef] [PubMed]
- Young, A.G.; Clarke, G.M. Genetics, Demography and Viability of Fragmented Populations; Cambridge University Press: Cambridge, UK, 2000. [Google Scholar]
- Galeuchet, D.J.; Perret, C.; Fischer, M. Microsatellite variation and structure of 28 populations of the common wetland plant, Lychnis flos-cuculi L., in a fragmented landscape. Mol. Ecol. 2005, 14, 991–1000. [Google Scholar] [CrossRef] [PubMed]
- Bacles, C.F.E.; Burczyk, J.B.; Lowe, A.J.; Ennos, A.R. Historical and contemporary mating patterns in remnant populations of the forest tree Fraxinus excelsior L. Evolution 2005, 59, 979–990. [Google Scholar] [CrossRef]
- Browne, L.; Ottewell, K.; Karubian, J. Short-term genetic consequences of habitat loss and fragmentation for the neotropical palm Oenocarpus bataua. Heredity 2015, 115, 389–395. [Google Scholar] [CrossRef]
- Su, T.; Wilf, P.; Huang, Y.J.; Zhang, S.T.; Zhou, Z.K. Peaches preceded humans: Fossil evidence from SW China. Sci. Rep. 2015, 5, 16794. [Google Scholar] [CrossRef]
- Faust, M.; Timon, B. Origin and dissemination of Peach. Hortic. Rev. 2010, 17, 331–379. [Google Scholar]
- Yu, Y.; Fu, J.; Xu, Y.; Zhang, J.; Ren, F.; Zhao, H.; Tian, S.; Guo, W.; Tu, X.; Zhao, J.; et al. Genome re-sequencing reveals the evolutionary history of peach fruit edibility. Nat. Commun. 2018, 9, 5404. [Google Scholar] [CrossRef]
- Wang, L.; Zhu, G.; Fang, W.; Cao, K.; Wang, X.; Chen, C.; Zhao, P.; Wang, X. Peach Genetic Resource in China; China Agriculture Press Co., Ltd.: Beijing, China, 2012. [Google Scholar]
- Lu, L.T. Rosaceae (volume 3), Prunoideae. In Flora Reipublicae Popularis Sinicae; Yu, T.T., Lu, L.T., Ku, T.C., Li, C.L., Chen, S.X., Eds.; Science Press: Beijing, China, 1986. [Google Scholar]
- Cheng, Z.P. Molecular Biological Study on Phylogeny of Subgenus Amygdalus and Genetic Diversity of Prunus persica. Ph.D. Thesis, University of Chinese Academy of Sciences, Beijing, China, 2007. (In Chinese). [Google Scholar]
- Li, Y.; Cao, K.; Zhu, G.; Fang, W.; Chen, C.; Wang, X.; Zhao, P.; Guo, J.; Ding, T.; Guan, L.; et al. Genomic analyses of an extensive collection of wild and cultivated accessions provide new insights into peach breeding history. Genome Biol. 2019, 20, 36. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.J.; Peng, M.; Guan, F.C. Amplified fragment length polymorphism analysis of genetic diversity and relationships of wild and cultivated peach (Prunus persica L.). HortScience 2015, 50, 44–50. [Google Scholar] [CrossRef]
- Schlötterer, C. Evolutionary dynamics of microsatellite DNA. Chromosoma 2000, 109, 365–371. [Google Scholar] [CrossRef] [PubMed]
- Nybom, H. Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol. Ecol. 2004, 13, 1143–1155. [Google Scholar] [CrossRef]
- Guan, L.P.; Cao, K.; Li, Y.; Guo, J.; Xu, Q.; Wang, L.R. Detection and application of genome-wide variations in peach for association and genetic relationship analysis. BMC Genet. 2019, 20, 101. [Google Scholar] [CrossRef]
- Cheng, Z.; Gasic, K.; Wang, Z. Population structure, linkage disequilibrium and selective loci in natural populations of Prunus davidiana. J. Agric. Sci. 2013, 5, 69–85. [Google Scholar] [CrossRef][Green Version]
- Cheng, Z.; Huang, H. SSR fingerprinting Chinese peach cultivars and landraces (Prunus persica) and analysis of their genetic relationships. Sci. Hortic. 2009, 120, 188–193. [Google Scholar] [CrossRef]
- Aranzana, M.J.; Garcia-Mas, J.; Carbó, J.; Arús, P. Development and variability analysis of microsatellite markers in peach. Plant Breed. 2002, 121, 87–92. [Google Scholar] [CrossRef]
- Cao, K.; Wang, L.R.; Zhu, G.R.; Fang, W.C.; Chen, C.W.; Luo, J. Genetic diversity, linkage disequilibrium, and association mapping analyses of peach (Prunus persica) landraces in China. Tree Genet. Genomes 2012, 8, 975–990. [Google Scholar] [CrossRef]
- Testolin, R.; Marrazzo, T.; Cipriani, G.; Quarta, R.; Verde, I.; Dettori, M.; Pancaldi, M.; Sansavini, S. Microsatellite DNA in peach (Prunus persica L. Batsch) and its use in fingerprinting and testing the genetic origin of cultivars. Genome 2000, 43, 512–520. [Google Scholar] [CrossRef]
- Khadivi-Khub, A.; Yarahmadi, M.; Jannatizadeh, A.; Ebrahimi, A. Genetic relationships and diversity of common apricot (Prunus armeniaca L.) based on simple sequence repeat (SSR) markers. Biochem. Syst. Ecol. 2015, 61, 366–371. [Google Scholar] [CrossRef]
- Sosinski, B.; Gannavarapu, M.; Hager, L.D.; Beck, L.E.; King, G.J.; Ryder, C.D. Characterization of microsatellite markers in peach [Prunus persica (L.) Batsch]. Theor. Appl. Genet. 2000, 101, 421–428. [Google Scholar] [CrossRef]
- Barać, G.; Ognjanov, V.; Vidaković, D.O.; Dorić, D.; Ljubojević, M.; Dulić, J.; Miodragović, M.; Gašić, K. Genetic diversity and population structure of European ground cherry (Prunus fruticosa Pall.) using SSR markers. Sci. Hortic. 2017, 224, 374–383. [Google Scholar] [CrossRef]
- Bouhadida, M.; Casas, A.M.; Moreno, M.A.; Gogorcena, Y. Molecular characterization of Miraflores peach variety and relatives using SSRs. Sci. Hortic. 2007, 111, 140–145. [Google Scholar] [CrossRef]
- García-Gómez, B.; Razi, M.; Salzar, J.A.; Prudencio, A.S.; Ruiz, D.; Dondini, L.; Martínez-Gómez, P. Comparative Analysis of SSR markers developed in exon, intron, and intergenic regions and distributed in regions controlling fruit quality traits in Prunus Species: Genetic diversity and association studies. Plant Mol. Biol. Rep. 2018, 36, 23–35. [Google Scholar] [CrossRef]
- Bouhadida, M.; Moreno, M.A.; Gonzalo, M.J.; Alonso, J.M. Genetic variability of introduced and local Spanish peach cultivars determined by SSR markers. Tree Genet. Genomes 2011, 7, 257–270. [Google Scholar] [CrossRef]
- Guidugli, M.C.; Nazareno, A.G.; Feres, J.M.; Contel, E.P.B.; Mestriner, M.A.; Alzate-Marin, A.L. Small but not isolated: A population genetic survey of the tropical tree Cariniana estrellensis (Lecythidaceae) in a highly fragmented habitat. Heredity 2016, 116, 339–347. [Google Scholar] [CrossRef]
- Scorza, R.; Mehlenbacher, S.A.; Lightner, G.W. Inbreeding and coancestry of freestone peach cultivars of the eastern United States and implications for peach germplasm improvement. J. Am. Soc. Hortic. Sci. 1985, 110, 547–552. [Google Scholar]
- Quesada, M.; Herrerias-Diego, Y.; Lobo, J.A.; Sanchez-Montoya, G.; Rosas, F.; Aguilar, R. Long-term effects of habitat fragmentation on mating patterns and gene flow of a tropical dry forest tree, Ceiba aesculifolia (Malvaceae: Bombacoideae). Am. J. Bot. 2013, 100, 1095–1101. [Google Scholar] [CrossRef]
- Li, Y.; Wang, L. Genetic resources, breeding programs in China, and gene mining of peach: A review. Hortic. Plant J. 2020, 6, 205–215. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Goudet, J. FSTAT (Version 1.2): A computer program to calculate F-Statistics. J. Hered. 1995, 86, 485–486. [Google Scholar] [CrossRef]
- Raymond, M.; Rousset, F. GENEPOP (Version 1.2): Population genetics software for exact tests and ecumenicism. J. Hered. 1995, 86, 248–249. [Google Scholar] [CrossRef]
- Peakall, R.O.D.; Smouse, P.E. genalex 6: Genetic analysis in Excel. population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Wright, S. The genetical structure of populations. Ann. Eugen. 1951, 15, 323–354. [Google Scholar] [CrossRef]
- Weir, B.S.; Cockerham, C.C. Estimating F-Statistics for the analysis of population structure. Evolution 1984, 38, 1358–1370. [Google Scholar]
- Schneider, S.D.; Roessli, D.; Excoffier, L. Arlequin Version 2.0: A Software for Population Genetic Analysis; University of Geneva: Geneva, Switzerland, 2000; Volume 2, Available online: https://www.botanicalgarden.cn/source/ (accessed on 11 April 2022).
- Miller, M.P. Tools for Population Genetic Analysis (TFPGA) 1.3: A Windows Program for the Analysis of Allozyme and Molecular Population Genetic Data. Computer Software Distributed by Author r. 1997. Available online: https://www.ccg.unam.mx/~vinuesa/tlem09/docs/TFPGADOC.PDF (accessed on 26 May 2022).
- Nei, M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 1978, 89, 583–590. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Piry, S.; Luikart, G.; Cornuet, J.M. Computer note. BOTTLENECK: A computer program for detecting recent reductions in the effective size using allele frequency data. J. Hered. 1999, 90, 502–503. [Google Scholar] [CrossRef]
- Beerli, P.; Palczewski, M. Unified framework to evaluate panmixia and migration direction among multiple sampling locations. Genetics 2010, 185, 313–326. [Google Scholar] [CrossRef] [PubMed]
- Wilson, G.A.; Rannala, B. Bayesian inference of recent migration rates using multilocus genotypes. Genetics 2003, 163, 1177–1191. [Google Scholar] [CrossRef] [PubMed]

| Population Code | Population Locality | Altitude (m) | Latitude (N) | Longitude (E) | N | A | HO | HE | FIS |
|---|---|---|---|---|---|---|---|---|---|
| MA | Xiaogan, Hubei Province | 250–500 | 31°10′ | 114°03′ | 23 | 4.0 | 0.240 | 0.590 | 0.593 ** |
| MK | Suizhou, Hubei Province | 800–1025 | 31°54′ | 113°13′ | 23 | 3.5 | 0.219 | 0.502 | 0.564 ** |
| MJ | Nanyang, Henan Province | 450–600 | 33°25′ | 111°56′ | 23 | 3.8 | 0.325 | 0.544 | 0.402 * |
| MM | Xingyang, Henan Province | 400–600 | 32°27′ | 113°23′ | 23 | 4.0 | 0.285 | 0.569 | 0.498 ** |
| MD | Shaoyang, Hunan Province | 450–650 | 26°18′ | 110°06′ | 23 | 2.5 | 0.115 | 0.372 | 0.690 ** |
| MN | Sangzhi, Hunan Province | 530–620 | 29°46′ | 109°54′ | 23 | 2.6 | 0.144 | 0.386 | 0.628 ** |
| ME | Mianning, Sichuan Province | 1910 | 28°13′ | 102°01′ | 23 | 3.0 | 0.142 | 0.409 | 0.653 ** |
| MG | Qingzhou, Shandong Province | 1032 | 36°11′ | 118°38′ | 23 | 2.0 | 0.083 | 0.140 | 0.407 * |
| MH | Mengyin, Shandong Province | 450 | 35°36′ | 117°54′ | 23 | 3.2 | 0.217 | 0.438 | 0.504 ** |
| MB | Anqing, Anhui Province | 400–650 | 30°48′ | 116°30′ | 23 | 4.3 | 0.229 | 0.568 | 0.597 ** |
| ML | Shangyou, Jiangxi Province | 600–650 | 25°55′ | 114°02′ | 23 | 3.7 | 0.240 | 0.523 | 0.541 ** |
| MF | Qianshan, Jiangxi Province | 587 | 27°57′ | 117°42′ | 23 | 3.7 | 0.166 | 0.510 | 0.674 ** |
| MO | Nanchuan, Chongqing City | 730–850 | 29°09′ | 107°09′ | 23 | 2.8 | 0.146 | 0.375 | 0.612 ** |
| MC | Baoshan, Yunnan Province | 1200 | 25°24′ | 99°08′ | 23 | 2.8 | 0.091 | 0.387 | 0.765 ** |
| MP | Weixi, Yunnan Province | 2100 | 27°06′ | 99°11′ | 23 | 2.6 | 0.113 | 0.381 | 0.703 ** |
| MQ | Qiubei, Yunnan Province | 1580 | 24°15′ | 104°12′ | 23 | 2.3 | 0.206 | 0.376 | 0.452 ** |
| Average | 23 | 3.2 | 0.185 | 0.442 |
| Source of Variation | d.f. | Sum of Squares | Variance Components | Percentage of Variation | p-Value |
|---|---|---|---|---|---|
| Among populations | 15 | 1777.571 | 2.40156 Va | 32.1 | p < 0.001 |
| Among individuals | 352 | 2827.565 | 2.95189 Vb | 39.45 | p < 0.001 |
| Within individuals | 368 | 783.500 | 2.12908 Vc | 28.45 | p < 0.001 |
| Total | 735 | 5838.636 | 7.48253 |
| Population | Mutation–Drift Test | |||
|---|---|---|---|---|
| IAM | TPM | SMM | Mode Shift | |
| MA | 0.001 ** | 0.038 * | 0.753 | L-shaped |
| MB | 0.002 ** | 0.329 | 0.052 | L-shaped |
| MC | 0.033 * | 0.368 | 0.674 | L-shaped |
| MD | 0.002 ** | 0.064 | 0.701 | L-shaped |
| ME | 0.048 * | 0.388 | 0.841 | L-shaped |
| MF | 0.018 * | 0.410 | 0.463 | L-shaped |
| MG | 0.039 * | 0.016 * | 0.006 * | L-shaped |
| MH | 0.026 * | 0.257 | 0.609 | L-shaped |
| MJ | 0.001 ** | 0.016 * | 0.975 | L-shaped |
| MK | 0.006 ** | 0.151 | 0.890 | L-shaped |
| ML | 0.004 ** | 0.079 | 1.000 | L-shaped |
| MM | 0.000 ** | 0.005 ** | 0.974 | L-shaped |
| MN | 0.024 * | 0.216 | 0.812 | L-shaped |
| MO | 0.076 | 0.701 | 0.349 | L-shaped |
| MP | 0.005 ** | 0.143 | 0.956 | L-shaped |
| MQ | 0.000 ** | 0.002 ** | 0.087 | L-shaped |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, Q.; Xu, Q.; Pan, J.; Yao, X.; Cheng, Z. Impacts of Chronic Habitat Fragmentation on Genetic Diversity of Natural Populations of Prunus persica in China. Plants 2022, 11, 1458. https://doi.org/10.3390/plants11111458
Jiang Q, Xu Q, Pan J, Yao X, Cheng Z. Impacts of Chronic Habitat Fragmentation on Genetic Diversity of Natural Populations of Prunus persica in China. Plants. 2022; 11(11):1458. https://doi.org/10.3390/plants11111458
Chicago/Turabian StyleJiang, Quan, Qiang Xu, Junfeng Pan, Xiaohong Yao, and Zhongping Cheng. 2022. "Impacts of Chronic Habitat Fragmentation on Genetic Diversity of Natural Populations of Prunus persica in China" Plants 11, no. 11: 1458. https://doi.org/10.3390/plants11111458
APA StyleJiang, Q., Xu, Q., Pan, J., Yao, X., & Cheng, Z. (2022). Impacts of Chronic Habitat Fragmentation on Genetic Diversity of Natural Populations of Prunus persica in China. Plants, 11(11), 1458. https://doi.org/10.3390/plants11111458






