Development and Characterization of 20 Genomic SSR Markers for Ornamental Cultivars of Weigela
Abstract
1. Introduction
2. Results and Discussion
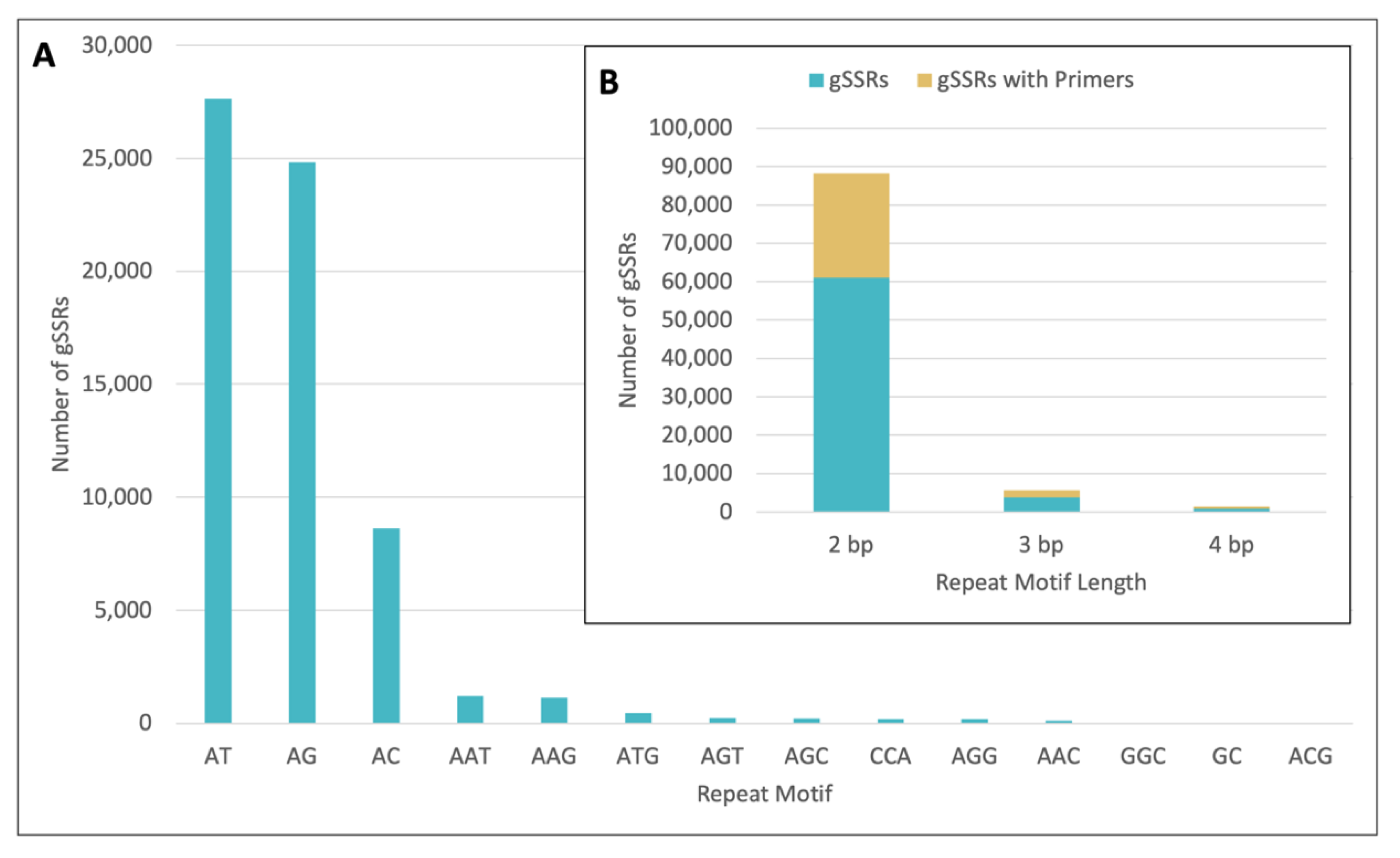
2.1. gSSR Marker Development
2.2. gSSR Characteristics
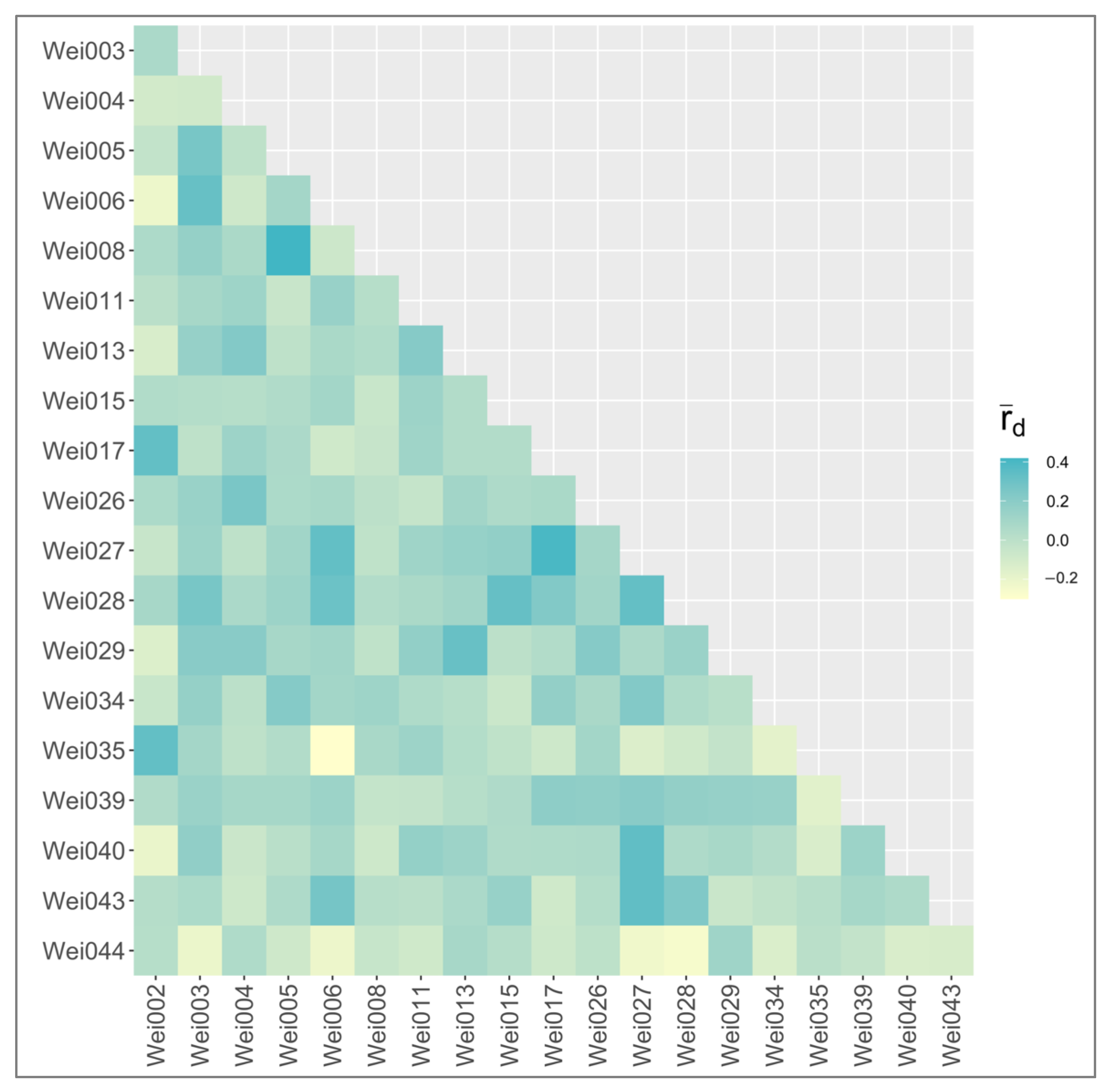
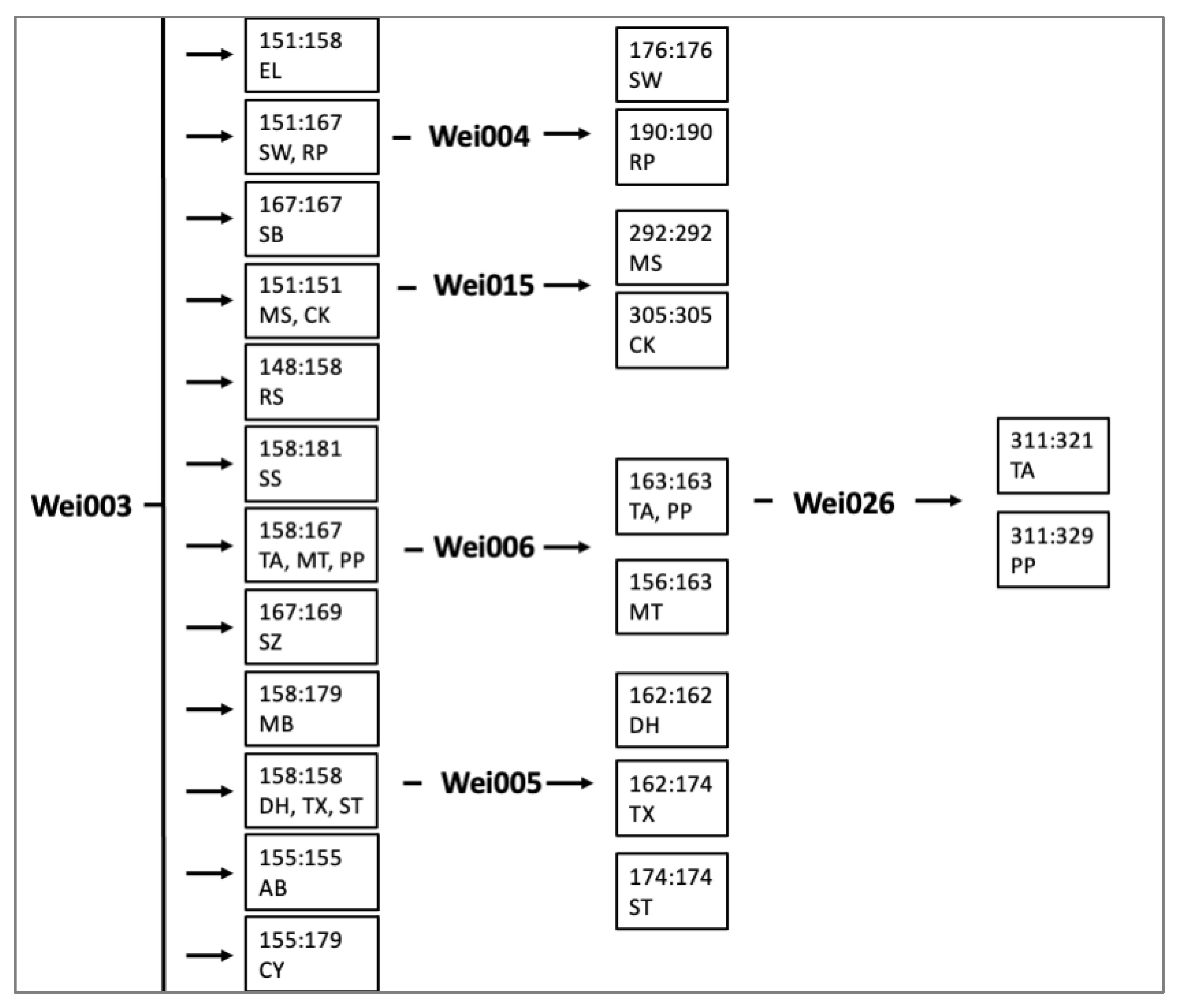
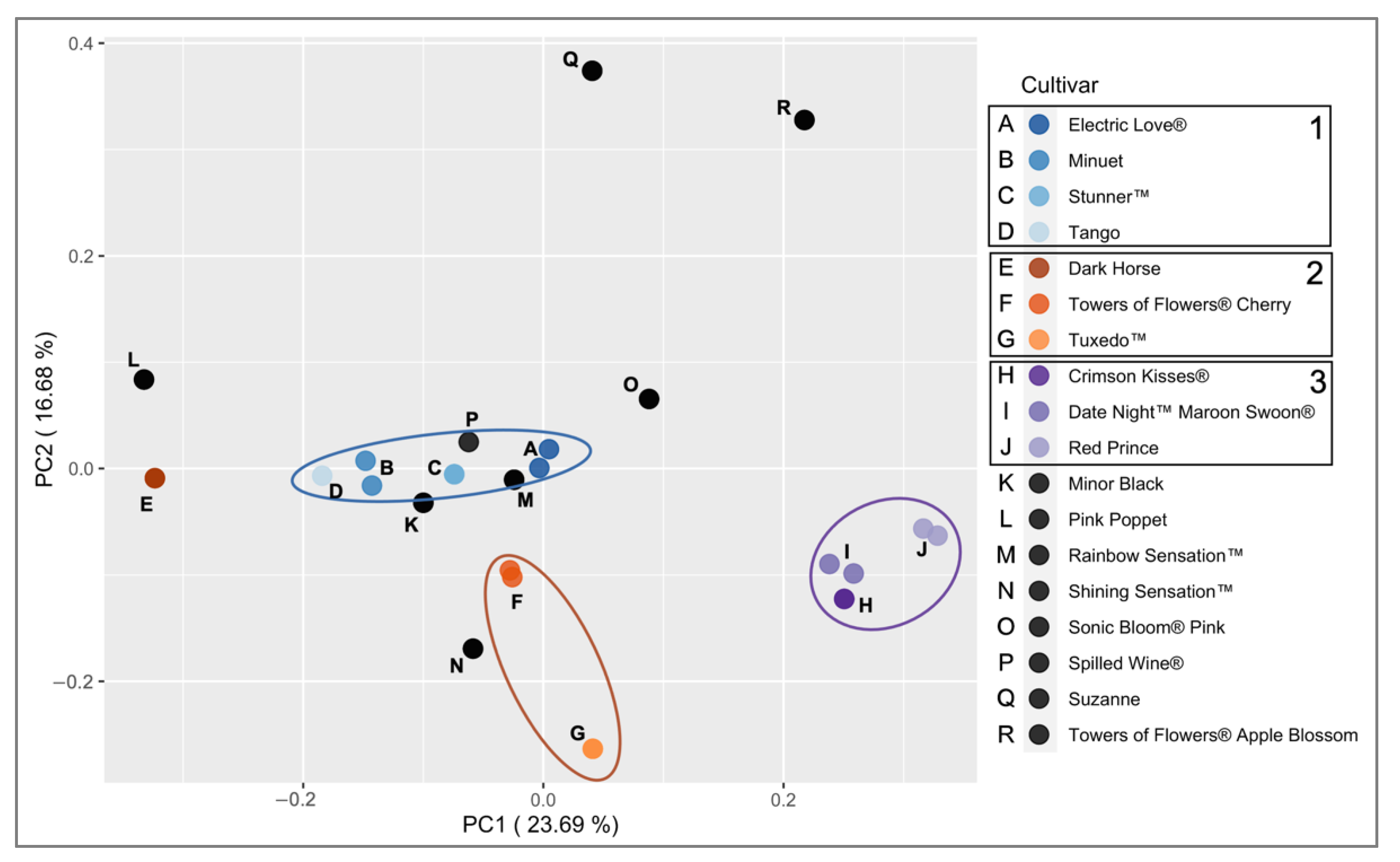
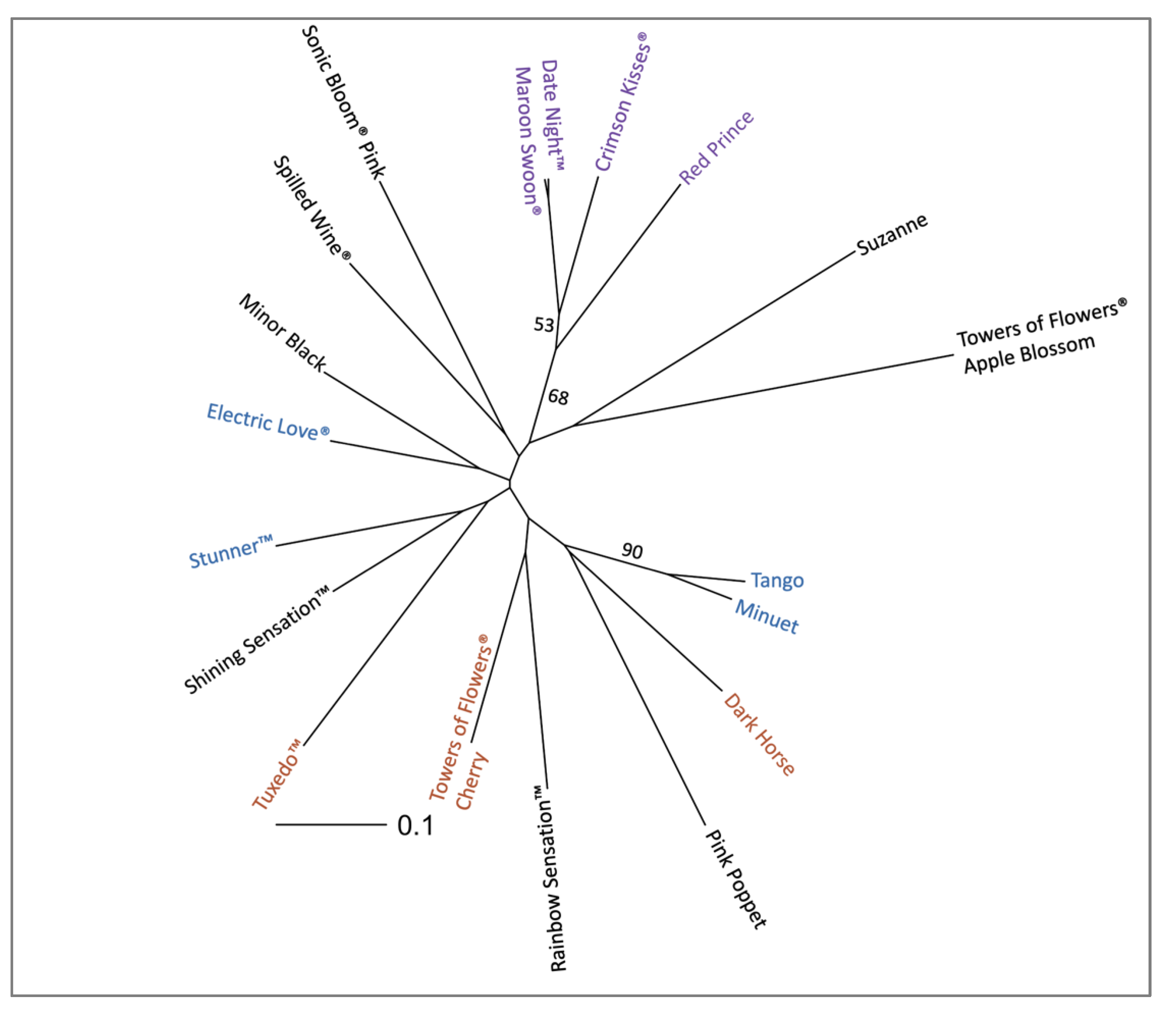
2.3. Applications with Weigela Cultivars
3. Materials and Methods
3.1. Plant Material and DNA Extraction
3.2. DNA Sequencing and Primer Design
3.3. Primer Screening and Optimization
3.4. gSSR Data Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hoffman, M. Cultivar classification of Weigela. In Proceedings of the V International Symposium on the Taxonomy of Cultivated Plants 799, Wageningen, The Netherlands, 15 October 2007; pp. 31–38. [Google Scholar]
- Duron, M.; Decourtye, L. In vitro variation in Weigela. In Somaclonal Variation in Crop Improvement I; Springer: Berlin/Heidelberg, Germany, 1990; pp. 606–623. [Google Scholar]
- Kim, Y.-D.; Kim, S.-H. Phylogeny of Weigela and Diervilla (Caprifoliaceae) based on nuclear rDNA ITS sequences: Biogeographic and taxonomic implications. J. Plant Res. 1999, 112, 331–341. [Google Scholar] [CrossRef]
- Pfarr, E.; Rothleutner, J. Weigela species and cultivar genome size and ploidy estimations: Shrub breeding©. In Proceedings of the 2015 Annual Meeting of the International Plant Propagators’ Society 1140, Bellefonte, PA, USA, 1 January 2015; pp. 221–224. [Google Scholar]
- Christian, T.; Elliott, A. Weigela—Trees and Shrubs Online. Available online: https://treesandshrubsonline.org/articles/weigela/ (accessed on 17 January 2022).
- USDA-NASS. Census of Horticultural Specialities: Table Deciduous Shrubs; USDA-NASS: Washington, DC, USA, 2020. [Google Scholar]
- Howard, R.A. A check-list of cultivar names in Weigela. Arnoldia 1965, 25, 49–69. [Google Scholar]
- Touchell, D.; Viloria, Z.; Ranney, T. Intergeneric hybrids between Weigela and Diervilla (Caprifoliaceae). HortScience 2006, 41, 1008D–1008. [Google Scholar] [CrossRef]
- Fan, W.-B.; Wu, Y.; Yang, J.; Shahzad, K.; Li, Z.-H. Comparative chloroplast genomics of Dipsacales species: Insights into sequence variation, adaptive evolution, and phylogenetic relationships. Front. Plant Sci. 2018, 9, 689. [Google Scholar] [CrossRef]
- Yamada, T.; Maki, M. Isolation and characterization of microsatellite loci from Weigela coraeensis (Caprifoliaceae). Mol. Ecol. Resour. 2008, 8, 155–157. [Google Scholar] [CrossRef]
- Yamada, T.; Maki, M. Impact of geographical isolation on genetic differentiation in insular and mainland populations of Weigela coraeensis (Caprifoliaceae) on Honshu and the Izu Islands. J. Biogeogr. 2012, 39, 901–917. [Google Scholar] [CrossRef]
- Valdes, A.M.; Slatkin, M.; Freimer, N.B. Allele frequencies at microsatellite loci: The stepwise mutation model revisited. Genetics 1993, 133, 737–749. [Google Scholar] [CrossRef]
- Tautz, D.; Renz, M. Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Res. 1984, 12, 4127–4138. [Google Scholar] [CrossRef]
- Zalapa, J.E.; Cuevas, H.; Zhu, H.; Steffan, S.; Senalik, D.; Zeldin, E.; McCown, B.; Harbut, R.; Simon, P. Using next-generation sequencing approaches to isolate simple sequence repeat (SSR) loci in the plant sciences. Am. J. Bot. 2012, 99, 193–208. [Google Scholar] [CrossRef]
- Kalia, R.K.; Rai, M.K.; Kalia, S.; Singh, R.; Dhawan, A.K. Microsatellite markers: An overview of the recent progress in plants. Euphytica 2011, 177, 309–334. [Google Scholar] [CrossRef]
- Tautz, D. Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res. 1989, 17, 6463–6471. [Google Scholar] [CrossRef] [PubMed]
- Hoshino, A.A.; Bravo, J.P.; Nobile, P.M.; Morelli, K.A. Microsatellites as Tools for Genetic Diversity Analysis; IntechOpen: London, England, 2012. [Google Scholar]
- Peakall, R.; Gilmore, S.; Keys, W.; Morgante, M.; Rafalski, A. Cross-species amplification of soybean (Glycine max) simple sequence repeats (SSRs) within the genus and other legume genera: Implications for the transferability of SSRs in plants. Mol. Biol. Evol. 1998, 15, 1275–1287. [Google Scholar] [CrossRef] [PubMed]
- Senan, S.; Kizhakayil, D.; Sasikumar, B.; Sheeja, T.E. Methods for development of microsatellite markers: An overview. Not. Sci. Biol. 2014, 6, 1–13. [Google Scholar] [CrossRef]
- Tabbasam, N.; Zafar, Y. Mehboob-ur-Rahman. Pros and cons of using genomic SSRs and EST-SSRs for resolving phylogeny of the genus Gossypium. Plant Syst. Evol. 2014, 300, 559–575. [Google Scholar] [CrossRef][Green Version]
- Reed, S.M.; Rinehart, T.A. Simple-sequence repeat marker analysis of genetic relationships within Hydrangea paniculata. HortScience 2009, 44, 27–31. [Google Scholar] [CrossRef]
- Hamm, T.P.; Nowicki, M.; Boggess, S.L.; Klingeman, W.E.; Hadziabdic, D.; Huff, M.L.; Staton, M.E.; Trigiano, R.N. Development and characterization of 15 novel genomic SSRs for Viburnum farreri. Plants 2021, 10, 487. [Google Scholar] [CrossRef]
- Nowicki, M.; Schilling, E.E.; Boggess, S.L.; Houston, L.C.; Huff, M.L.; Staton, M.E.; Lampley, J.A.; Trigiano, R.N. Development and characterization of genic microsatellites for the ornamental plant Green and Gold (Chrysogonum virginianum). HortScience 2019, 54, 395–400. [Google Scholar] [CrossRef]
- Somers, D.J.; Isaac, P.; Edwards, K. A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor. Appl. Genet. 2004, 109, 1105–1114. [Google Scholar] [CrossRef]
- Wang, X.; Wadl, P.A.; Rinehart, T.A.; Scheffler, B.E.; Windham, M.T.; Spiers, J.M.; Johnson, D.H.; Trigiano, R.N. A linkage map for flowering dogwood (Cornus florida L.) based on microsatellite markers. Euphytica 2009, 165, 165–175. [Google Scholar] [CrossRef]
- Carter, A.H.; Chen, X.; Garland-Campbell, K.; Kidwell, K. Identifying QTL for high-temperature adult-plant resistance to stripe rust (Puccinia striiformis f. sp. tritici) in the spring wheat (Triticum aestivum L.) cultivar ‘Louise’. Theor. Appl. Genet. 2009, 119, 1119–1128. [Google Scholar] [CrossRef]
- Brbaklić, L.; Trkulja, D.; Kondić-Špika, A.; Treskić, S.; Kobiljski, B. Detection of QTLs for important agronomical traits in hexaploid wheat using association analysis. Czech J. Genet. Plant Breed. 2013, 49, 1–8. [Google Scholar] [CrossRef]
- Ercisli, S.; Ipek, A.; Barut, E. SSR marker-based DNA fingerprinting and cultivar identification of olives (Olea europaea). Biochem. Genet. 2011, 49, 555. [Google Scholar] [CrossRef] [PubMed]
- Tóth, G.; Gáspári, Z.; Jurka, J. Microsatellites in different eukaryotic genomes: Survey and analysis. Genome Res. 2000, 10, 967–981. [Google Scholar] [CrossRef] [PubMed]
- Agapow, P.-M.; Burt, A. Indices of multilocus linkage disequilibrium. Mol. Ecol. Notes 2001, 1, 101–102. [Google Scholar] [CrossRef]
- Svejda, F. Minuet Weigela. Can. J. Plant Sci. 1982, 62, 249–250. [Google Scholar] [CrossRef]
- Svejda, F. Weigela cultivars Tango and Polka. HortScience 1988, 23, 787–788. [Google Scholar]
- Verhoef, G. Plant Named ‘ZR1’. USPP30065, 8 January 2019. [Google Scholar]
- Verhoef, G. Weigela Plant Named ‘Spring2’. USPP30185P2, 12 February 2019. [Google Scholar]
- Moore, P.R. Weigela Plant Named ‘Dark Horse’. USPP14381P3, 16 December 2003. [Google Scholar]
- Verhoef, G. Weigela Plant Named ‘Velda’. USPP26842P3, 21 June 2016. [Google Scholar]
- Valin, C. Weigela Plant Named ‘TMWG16-04’. USPP31915P2, 30 June 2020. [Google Scholar]
- Weigle, J.; Stephens, L. ‘Red Prince’ Weigela. HortScience 1991, 26, 218–219. [Google Scholar] [CrossRef]
- Verhoef, G. Weigela Plant Named ‘Slingco 1’. USPP23654P2, 11 June 2013. [Google Scholar]
- Verhoef, G. Weigela Plant Named ‘Slingco2’. US2016/0135346P1, 12 May 2016. [Google Scholar]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 28 May 2022).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Simpson, J.T.; Wong, K.; Jackman, S.D.; Schein, J.E.; Jones, S.J.; Birol, I. ABySS: A parallel assembler for short read sequence data. Genome Res. 2009, 19, 1117–1123. [Google Scholar] [CrossRef]
- Morgulis, A.; Gertz, E.M.; Schäffer, A.A.; Agarwala, R. A fast and symmetric DUST implementation to mask low-complexity DNA sequences. J. Comput. Biol. 2006, 13, 1028–1040. [Google Scholar] [CrossRef]
- Staton, M.E.; Ficklin, S. Finding SSRs—Findssrs_altered.pl. Available online: https://github.com/statonlab/Finding-SSRs/blob/master/findSSRs_altered.pl (accessed on 9 May 2022).
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed]
- Don, R.H.; Cox, P.T.; Wainwright, B.J.; Baker, K.; Mattick, J.S. ‘Touchdown’ PCR to circumvent spurious priming during gene amplification. Nucleic Acids Res. 1991, 19, 4008. [Google Scholar] [CrossRef] [PubMed]
- Amos, W.; Hoffman, J.I.; Frodsham, A.; Zhang, L.; Best, S.; Hill, A.V.S. Automated binning of microsatellite alleles: Problems and solutions. Mol. Ecol. Notes 2007, 7, 10–14. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021. [Google Scholar]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2014, 2, e281. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: Berlin/Heidelberg, Germany, 2016. [Google Scholar]
- Chessel, D.; Dufour, A.B.; Thioulouse, J. The ade4 package—I: One-table methods. R News 2004, 4, 5–10. [Google Scholar]
| Locus | GenBank Accession | Primer Sequences | Repeat Motif | Allele Size Range (bp) | Number of Alleles | Missing Data (%) |
|---|---|---|---|---|---|---|
| Wei002 | OM158066 | F: ACATCATCATCACTTGGGTGG | [GAAA]6 | 135–150 | 5 | 17.65 |
| R: ACCAGCACTTTCAATCTTCC | ||||||
| Wei003 | OM158067 | F: ACATTCCAAGGCCACAAATGC | [TC]9 | 148–181 | 8 | 0 |
| R: GTGGTCCTTGAATGTGTTTCATACG | ||||||
| Wei004 | OM158068 | F: TGCACCTCAAATGAGACACC | [CT]8 | 150–202 | 6 | 0 |
| R: AGGAGGTGGAGGAGACAAGG | ||||||
| Wei005 | OM158069 | F: TCGCTGATCGTTTGGAGTCC | [TCT]11 | 159–192 | 5 | 0 |
| R: AGCAAAGTAACCCTAGACAGG | ||||||
| Wei006 | OM158070 | F: TCCTTCAATGGAGAGAGCCC | [ATAG]8 | 156–175 | 4 | 0 |
| R: CAGAACTTGAATTCATGTTGTTGCC | ||||||
| Wei008 | OM158071 | F: GCGTGACAAATTGCTTACTTGG | [TATC]8 | 174–201 | 6 | 11.76 |
| R: ACATGGTTCAACAGTCTCCC | ||||||
| Wei011 | OM158072 | F: TTTCTCAGCAACCAAACCGC | [ACAT]6 | 237–249 | 4 | 0 |
| R: AAAGCACAAGCACAGAAGGG | ||||||
| Wei013 | OM158073 | F: CCTCAAGAAGAAAGCGCTGC | [GAA]10 | 280–297 | 5 | 0 |
| R: ACCAGAGAAATGTGTAACGC | ||||||
| Wei015 | OM158074 | F: GAGTGCCAATAGCCAAACCC | [AT]9 | 292–325 | 6 | 0 |
| R: TCGAAAGGTGGACCAACTGG | ||||||
| Wei017 | OM158075 | F: TCTTCTCATTTGGTGTAGCG | [AAT]11 | 201–217 | 4 | 5.88 |
| R: CGCCACCAATTTGGGTAACG | ||||||
| Wei026 | OM158076 | F: GTGATTGAGTTTGAGCCGCC | [TA]13 | 304–329 | 6 | 0 |
| R: CATGCACCACACCTTCATGC | ||||||
| Wei027 | OM158077 | F: GTAAACCAATCAGGCGCACC | [CAT]7 | 340–362 | 7 | 0 |
| R: GCACAAGCAAAGAAGCCGG | ||||||
| Wei028 | OM158078 | F: GAACCACAACTCAAGCTCCG | [TATG]7 | 308–340 | 6 | 0 |
| R: TCCGACGATTATGCTCACCG | ||||||
| Wei029 | OM158079 | F: CAATACGAGAAGTGACGCTACC | [TC]10 | 348–361 | 6 | 0 |
| R: CCCTTGCATTAAGAGGGTGC | ||||||
| Wei034 | OM158080 | F: TCATGCAGTCTAAGCCCACC | [CATA]6 | 383–432 | 6 | 0 |
| R: GGTTACCGCGTGAAGTATGC | ||||||
| Wei035 | OM158081 | F: TCCACTTCAACCTGAGCTGC | [AG]8 | 388–404 | 7 | 23.53 |
| R: TTAGTGACCACATCGTGACG | ||||||
| Wei039 | OM158082 | F: TTTCTGCCTAACCAAATCAGCC | [AG]9 | 149–197 | 7 | 8.82 |
| R: CTCACACGTACCACTCTAGCC | ||||||
| Wei040 | OM158083 | F: TTCAGTTGAAAGACCAACCG | [GA]8 | 171–221 | 5 | 0 |
| R: CACATTGAGAGAGCAATAATTTCCC | ||||||
| Wei043 | OM158084 | F: AAGAGTTCCGTCCATGCTGG | [AG]9 | 266–286 | 6 | 0 |
| R: GCGAACAAGCTCAATTCCGG | ||||||
| Wei044 | OM158085 | F: TCTAAAGCTCCATGGTCGGC | [AC]10 | 299–301 | 2 | 29.41 |
| R: AGTGCCAATAGCCAATACCC |
| Group a | Taxa | Nursery b | US Patent Number/Publication | Breeding Information c | Private Alleles (n) |
|---|---|---|---|---|---|
| 1 | W. ‘ZR1’ Electric Love® | Pope’s Creekside Nursery | USPP30065 | W. ‘Tango’ × Open pollinated | 0 |
| 1 | W. ‘Minuet’ | Nature Hills | Svejda, 1982 d | W. ‘Purpurea’ × W. ‘Dropmore Pink’ | 2 |
| 1 | W. ‘Spring 2’ Stunner™ | Wayside Gardens | USPP30185 | Open Pollinated W. ‘Tango’ | 0 |
| 1 | W. ‘Tango’ | Pixie Gardens | Svejda, 1988 e | W. ‘Minuet’ × W. ‘Nana variegata’ | 1 |
| 2 | W. ‘Dark Horse’ | Nature Hills | USPP14381 | W. ‘Victoria’ g × W. ‘Foliis Purpureis’ | 1 |
| 2 | W. ‘TMWG16-04’ Towers of Flowers® Cherry | Wayside Gardens | USPP31915 | Open pollination W. florida ‘WG13003’ × W. florida ‘Alexandra’? h | 1 |
| 2 | W. ‘Velda’ Tuxedo™ | Wayside Gardens | USPP26842 | W. ‘Milk and Honey’ × W. florida ‘Alexandra’ | 3 |
| 3 | W. ‘Slingco 1’ Crimson Kisses® | Arbor Foundation | USPP23654 | W. ‘Evita’ × W. ‘Red Prince’ | 1 |
| 3 | W. ‘Slingco 2’ Date Night™ Maroon Swoon® | Pope’s Creekside Nursery | USPP26841 | Open pollinated W. ‘Red Prince’ | 2 |
| 3 | W. ‘Red Prince’ | Nature Hills | Weigle, 1991 f | W. ‘Vanicek’ × W. ISU41 | 3 |
| N/A | W. ‘Verweig 3’ Minor Black | Wayside Gardens | N/A | unknown | 0 |
| N/A | W. ‘Pink Poppet’ | Nature Hills | US 20030033647 P1 | W. florida ‘Venusta’ × W. hybrida ‘Eva Rathke’ | 6 |
| N/A | W. ‘Kolmagira’ Rainbow Sensation™ | Nature Hills | USPP20384 | Cross-pollination of unnamed seedling | 2 |
| N/A | W. ‘Bokrashine’ Shining Sensation™ | Nature Hills | N/A | unknown | 2 |
| N/A | W. ‘Bokrasopin’ Sonic Bloom® Pink | Proven Winners | USPP24572 | Open Pollinated W. hybrida 93118 | 1 |
| N/A | W. ‘Bokraspiwi’ Spilled Wine® | Pope’s Creekside Nursery | USPP23781 | Open Pollinated W. hybrida 93115 | 2 |
| N/A | W. ‘Suzanne’ | Nature Hills | N/A | unknown | 5 |
| N/A | W. ‘TMWG16-02’ Towers of Flowers® Apple Blossom | Wayside Gardens | USPP32237 | Open Pollinated W. florida ‘WG13001’ | 4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hamm, T.P.; Boggess, S.L.; Kandel, J.S.; Staton, M.E.; Huff, M.L.; Hadziabdic, D.; Shoemaker, D.; Adamczyk Jr., J.J.; Nowicki, M.; Trigiano, R.N. Development and Characterization of 20 Genomic SSR Markers for Ornamental Cultivars of Weigela. Plants 2022, 11, 1444. https://doi.org/10.3390/plants11111444
Hamm TP, Boggess SL, Kandel JS, Staton ME, Huff ML, Hadziabdic D, Shoemaker D, Adamczyk Jr. JJ, Nowicki M, Trigiano RN. Development and Characterization of 20 Genomic SSR Markers for Ornamental Cultivars of Weigela. Plants. 2022; 11(11):1444. https://doi.org/10.3390/plants11111444
Chicago/Turabian StyleHamm, Trinity P., Sarah L. Boggess, Jinita Sthapit Kandel, Margaret E. Staton, Matthew L. Huff, Denita Hadziabdic, DeWayne Shoemaker, John J. Adamczyk Jr., Marcin Nowicki, and Robert N. Trigiano. 2022. "Development and Characterization of 20 Genomic SSR Markers for Ornamental Cultivars of Weigela" Plants 11, no. 11: 1444. https://doi.org/10.3390/plants11111444
APA StyleHamm, T. P., Boggess, S. L., Kandel, J. S., Staton, M. E., Huff, M. L., Hadziabdic, D., Shoemaker, D., Adamczyk Jr., J. J., Nowicki, M., & Trigiano, R. N. (2022). Development and Characterization of 20 Genomic SSR Markers for Ornamental Cultivars of Weigela. Plants, 11(11), 1444. https://doi.org/10.3390/plants11111444








