Genetic Diversity Assessment of the International Maize and Wheat Improvement Center and Chinese Wheat Core Germplasms by Non-Denaturing Fluorescence In Situ Hybridization
Abstract
:1. Introduction
2. Results
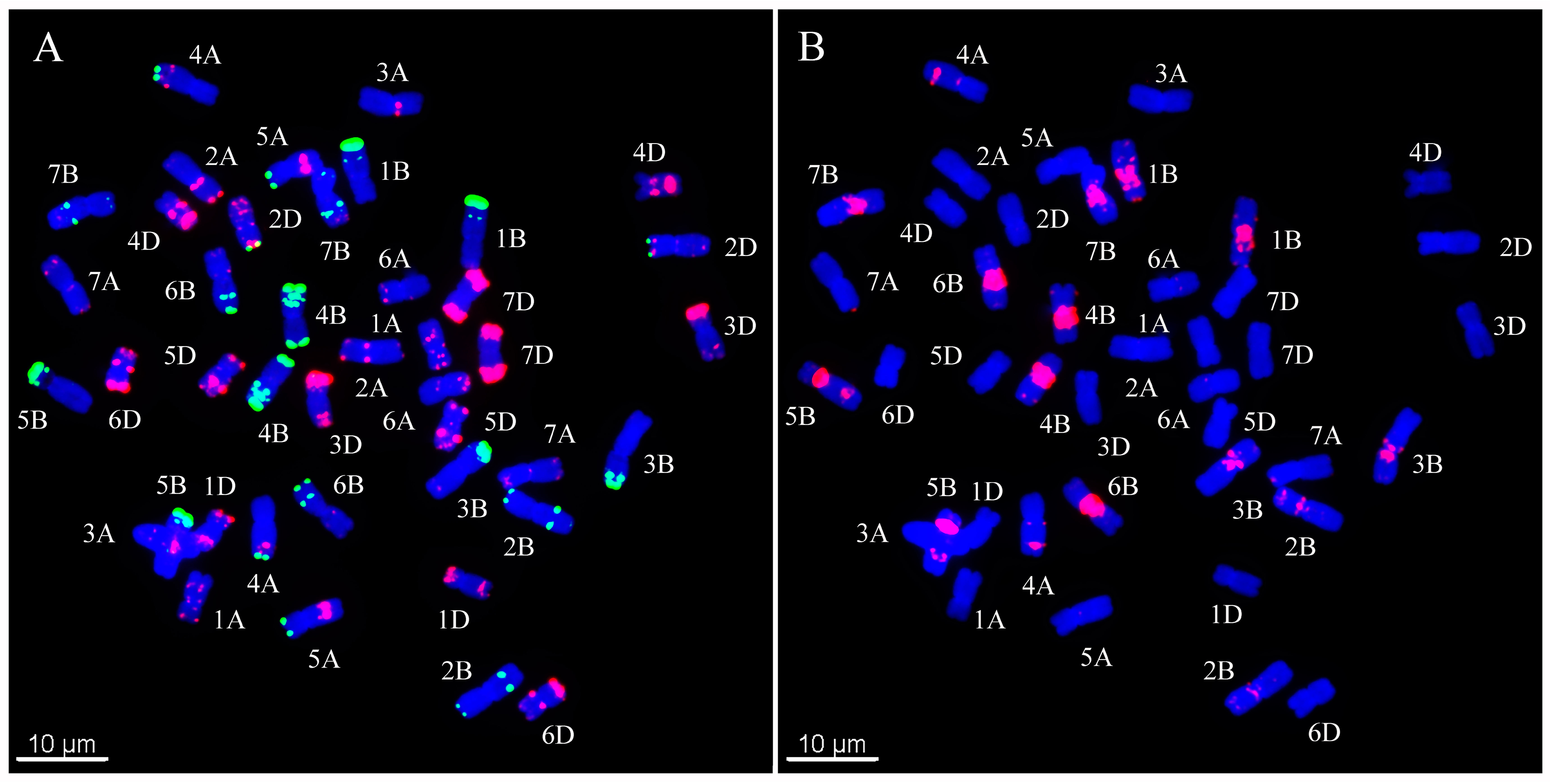
2.1. Construction of Wheat Karyotype by ND-FISH
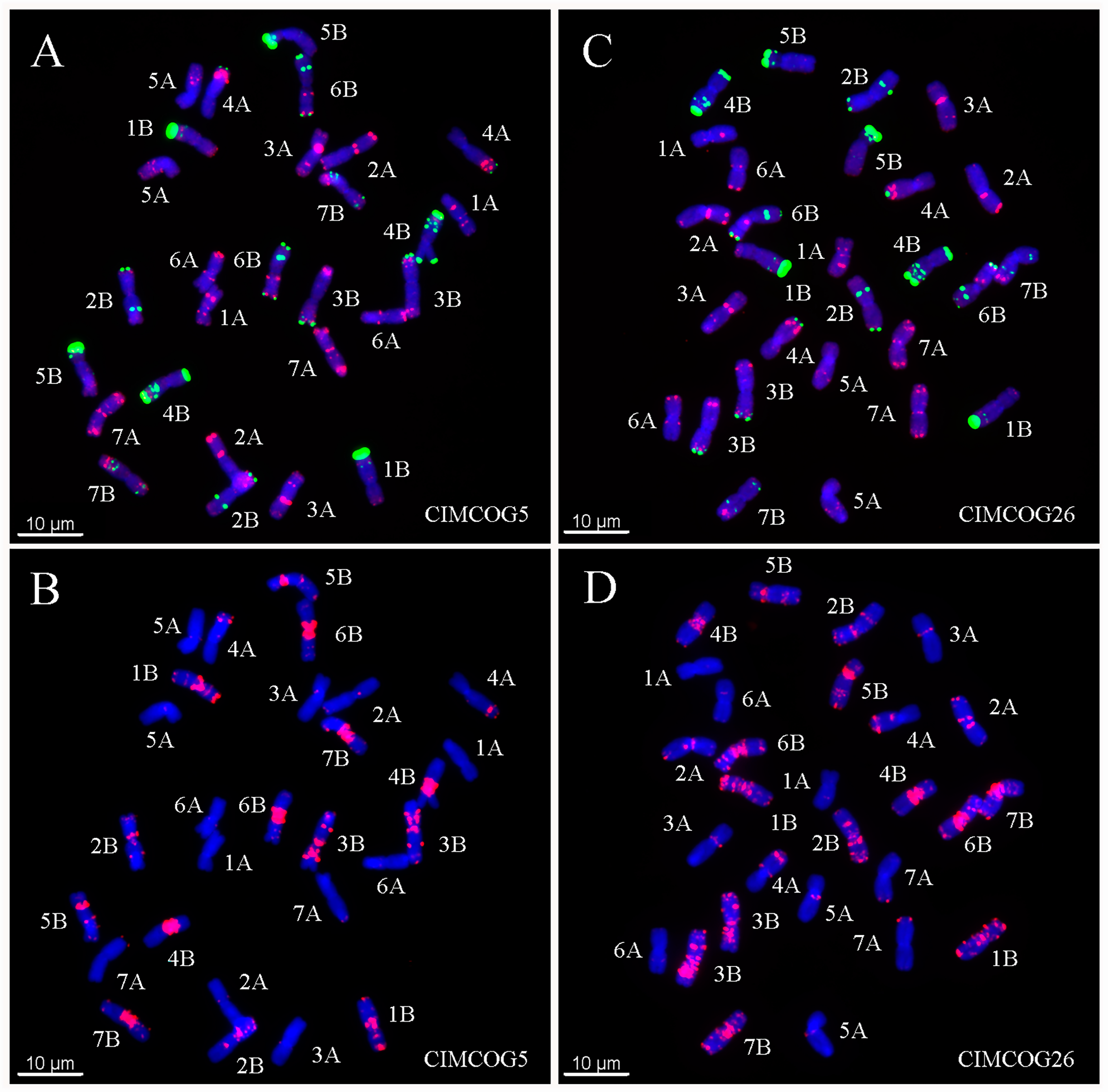
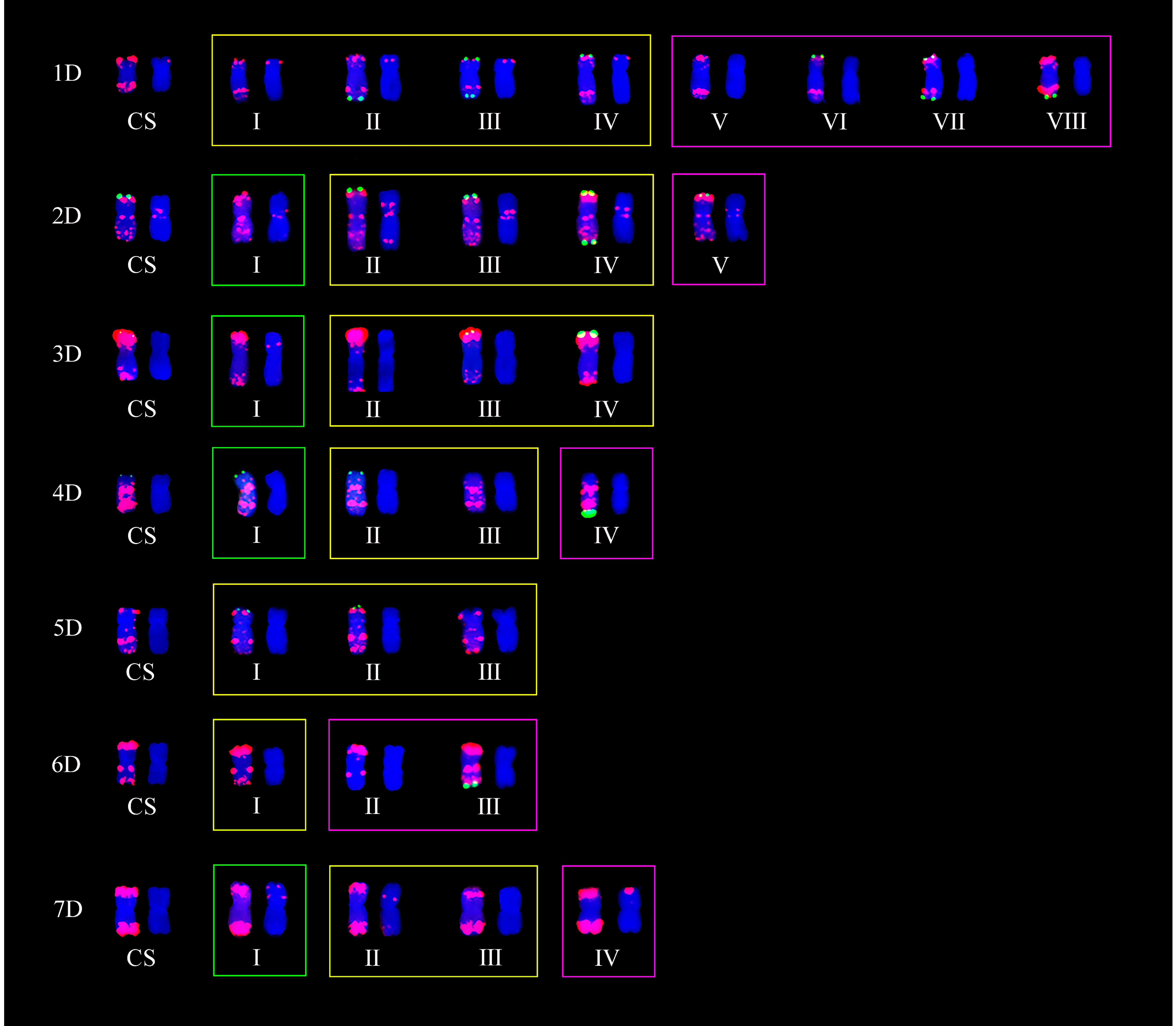
2.2. ND-FISH Polymorphisms of A-, B-, and D-Genome Chromosomes
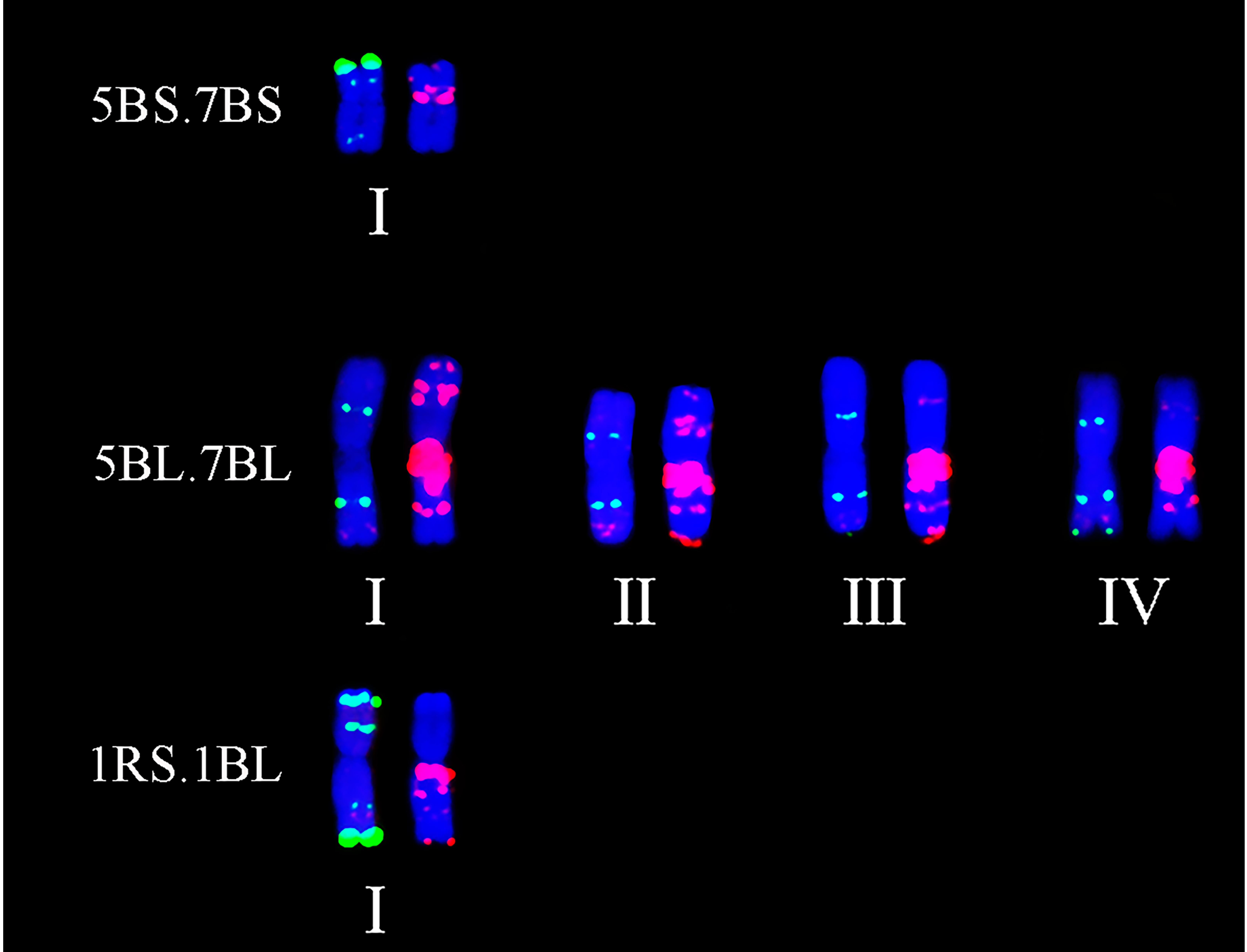
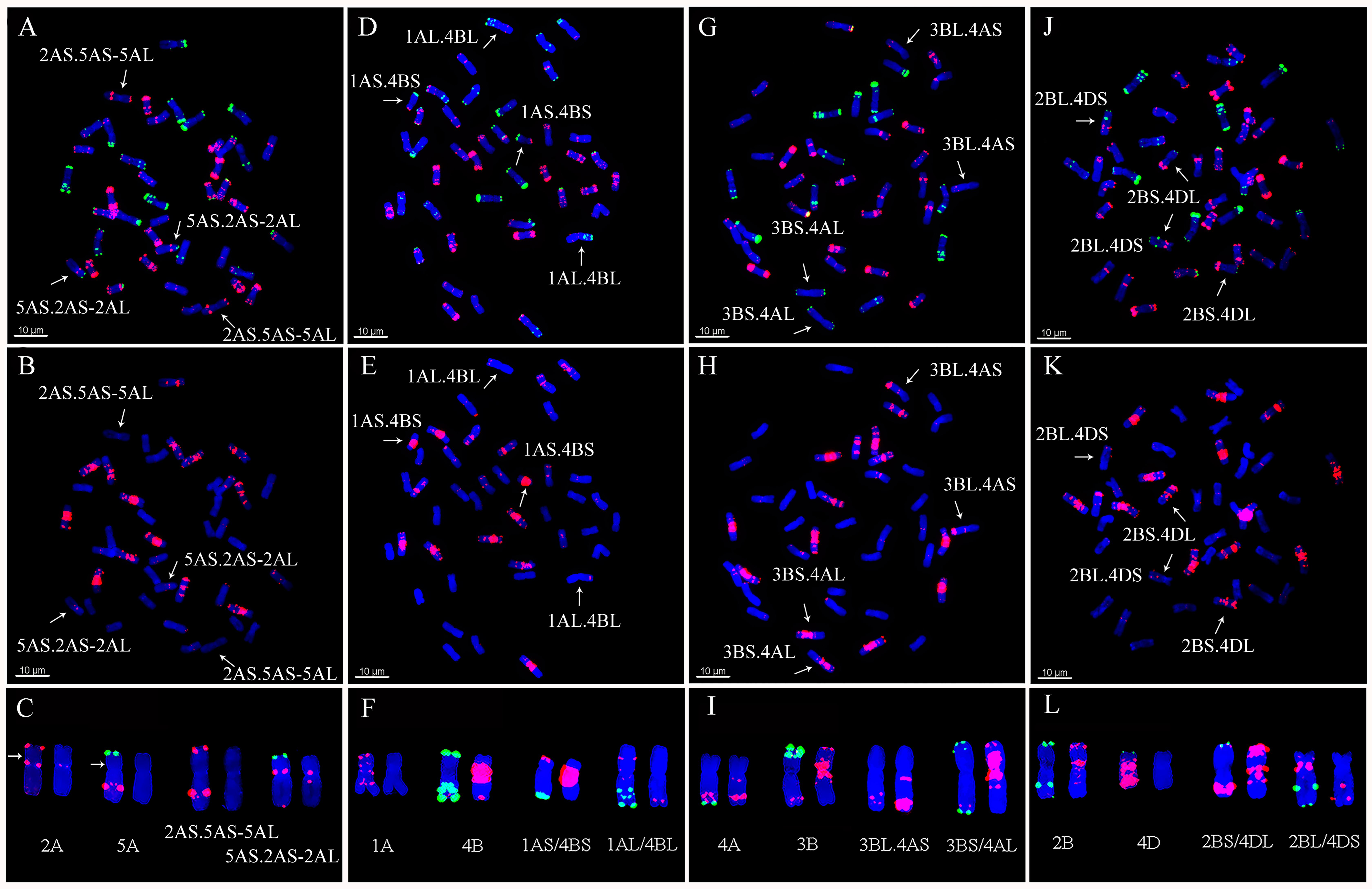
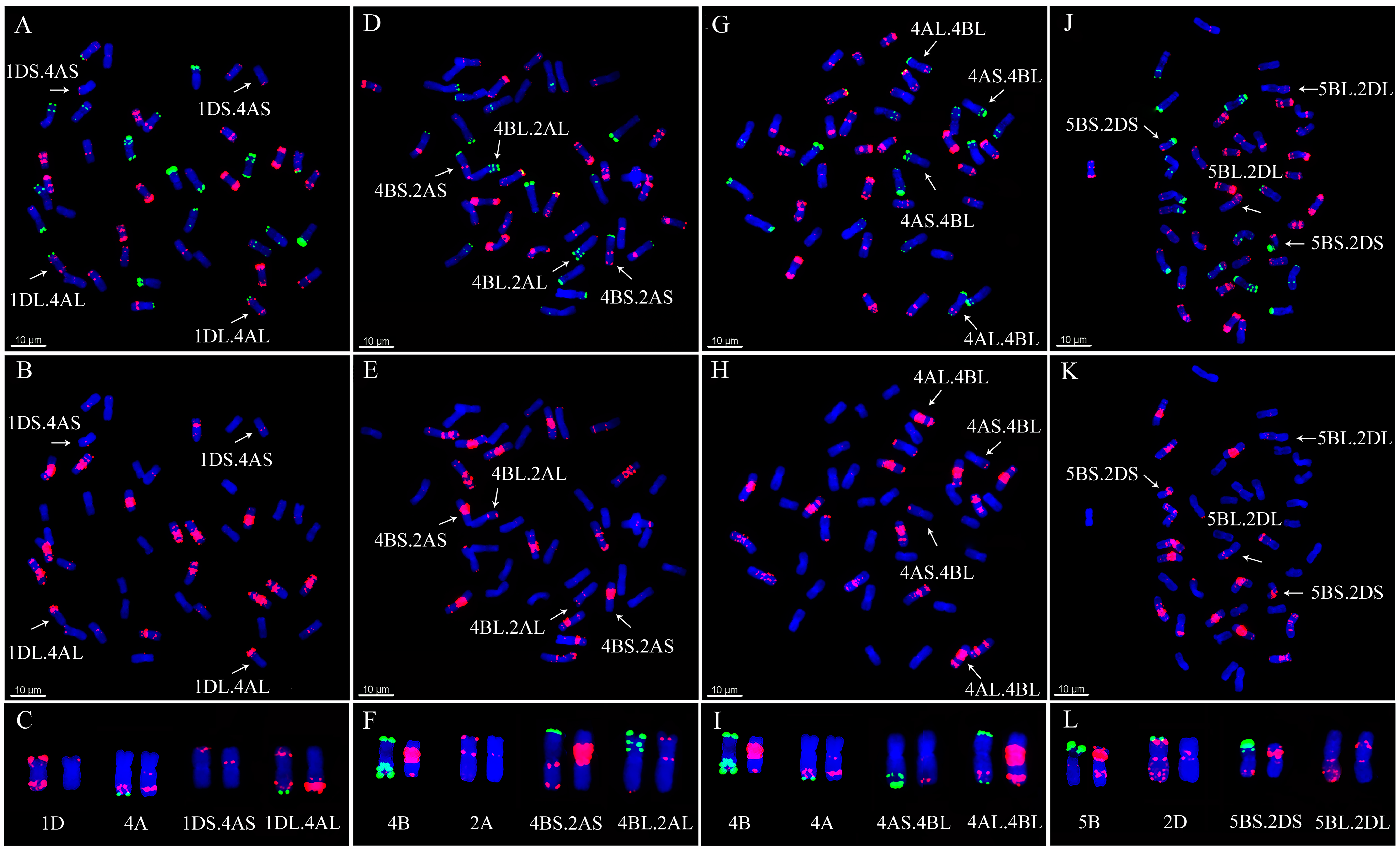
2.3. Detection of Chromosomal Translocations
2.4. Distributions of Different Chromosomal Types between the Two Wheat Germplasm Resources
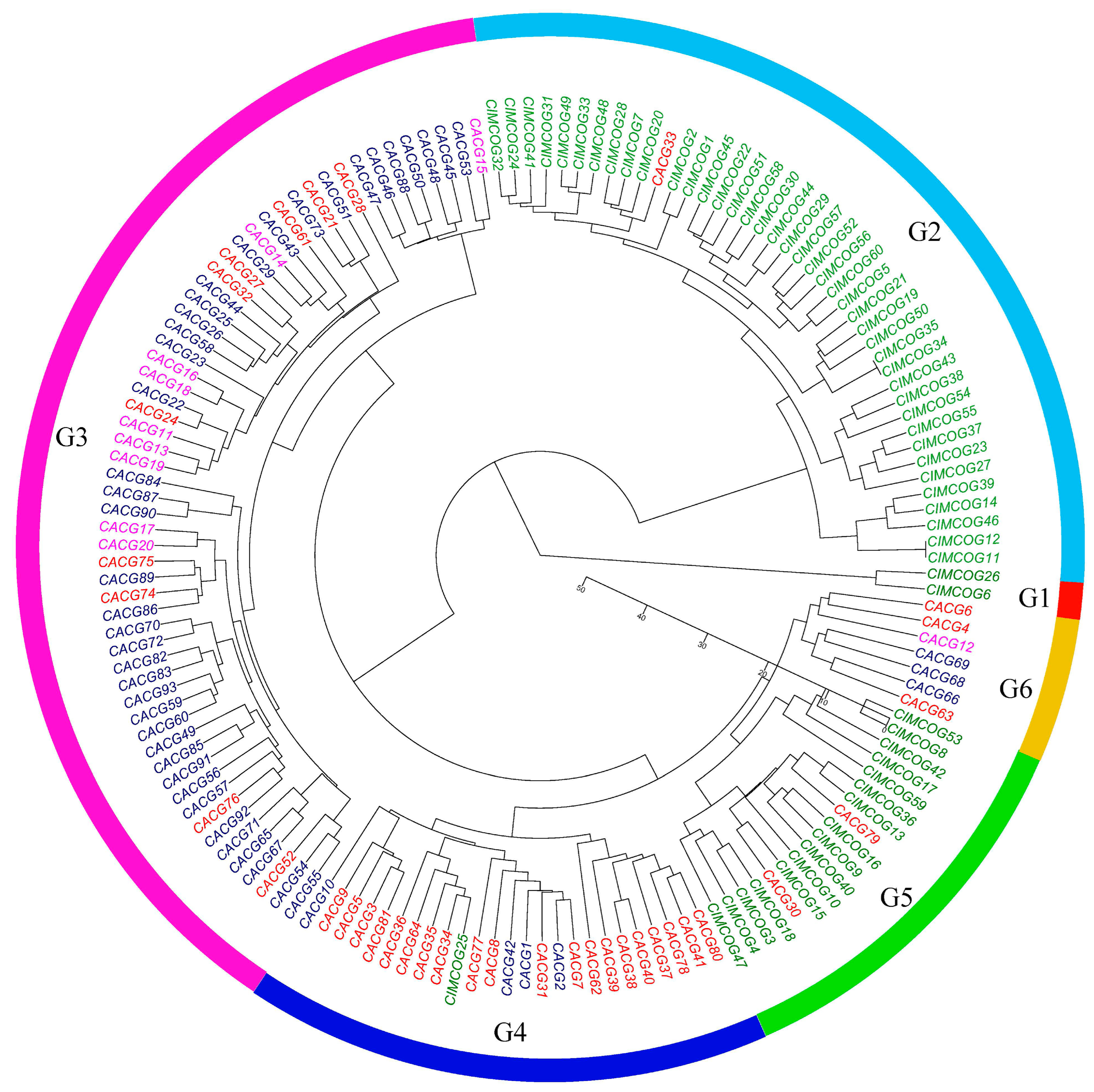
2.5. Genetic Relationships Revealed by FISH Patterns
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. ND-FISH Analysis
4.3. Data Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hedden, P. The genes of the Green Revolution. Trends Genet. 2003, 19, 5–9. [Google Scholar] [CrossRef]
- Yu, Y.H.; Wu, Y.T.; Zeng, X.; Yuan, L.P. Present situation of utilization on rice dwarf gene resources and its research advances in molecular biology. Hunan Agric. Sci. 2007, 5, 20–24. [Google Scholar] [CrossRef]
- Zhu, H.; Niu, Y.L.; Wang, P.; Quan, J.L.; Chen, Y.F. Molecular detection and genetic effect research of dwarf dene in wheat. Mol. Plant Breed. 2018, 16, 1855–1861. [Google Scholar] [CrossRef]
- Wen, H.Q.; Pei, Z.Y.; Hou, M.L.; Zhang, L.S.; Cheng, T.L.; Li, X.; Zhu, M. Research progress and application of Taigu male-sterile wheat in Shanxi province. J. Shanxi Agric. Sci. 2015, 43, 1715–1717. [Google Scholar] [CrossRef]
- Zhang, Y.; Yang, W.Y.; Zou, Y.C. Characterisation of the new wheat cultivar Chuanmai42 derived from CIMMYT hexaploidy wheat. Southwest China J. Agric. Sci. 2007, 20, 199–202. [Google Scholar] [CrossRef]
- Ban, T.; Suenaga, K. Genetic analysis of resistance to Fusarium head blight caused by Fusarium graminearum in Chinese wheat cultivar Sumai 3 and the Japanese cultivar Saikai 165. Euphytica 2000, 113, 87–99. [Google Scholar] [CrossRef]
- Bai, G.; Shaner, G. Management and resistance in wheat and barley to fusarium head blight. Annu. Rev. Phytopathol. 2004, 42, 135–161. [Google Scholar] [CrossRef]
- Talha, M.; Swati, H.; Jaiswal, J.P. Marker assisted detection of underutilized potential Yr genes in elite wheat breeding lines. SABRAO J. Breed. Genet. 2016, 48, 309–317. [Google Scholar]
- Tanksley, S.D.; McCouch, S.R. Seed banks and molecular maps: Unlocking genetic potential from the wild. Science 1997, 277, 1063–1066. [Google Scholar] [CrossRef] [Green Version]
- Lopes, M.S.; El-Basyoni, I.; Baenziger, P.S.; Singh, S.; Royo, C.; Ozbek, K.; Aktas, H.; Ozer, E.; Ozdemir, F.; Manickavelu, A.; et al. Exploiting genetic diversity from landraces in wheat breeding for adaptation to climate change. J. Exp. Bot. 2015, 66, 3477–3486. [Google Scholar] [CrossRef]
- Autrique, E.; Nachit, M.M.; Monneveux, P.; Tanksley, S.D.; Sorrells, M.E. Genetic diversity in durum wheat based on RFLPs, morphophysiological traits, and coefficient of parentage. Crop Sci. 1996, 36, 735–742. [Google Scholar] [CrossRef]
- Reif, J.C.; Zhang, P.; Dreisigacker, S.; Warburton, M.L.; Van Ginkel, M.; Hoisington, D.; Bohn, M.; Melchinger, A.E. Wheat genetic diversity trends during domestication and breeding. Theor. Appl. Genet. 2005, 110, 859–864. [Google Scholar] [CrossRef] [PubMed]
- Bhatta, M.; Morgounov, A.; Belamkar, V.; Poland, J.; Baenziger, P.S. Unlocking the novel genetic diversity and population structure of synthetic hexaploid wheat. BMC Genom. 2018, 19, 591. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, C.L.; Hu, S.L.; Gardner, C.; Lübberstedt, T. Emerging avenues for utilization of exotic germplasm. Trends Plant Sci. 2017, 22, 624–637. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zou, Y.C.; Zhang, Y.; Yang, W.Y.; Yang, E.N.; Huang, G. Reviews and prospects of Sichuan-CIMMYT wheat shuttle breeding program. Southwest China J. Agric. Sci. 2003, 16, 102–106. [Google Scholar] [CrossRef]
- Branlard, G.; Dardevet, M.; Saccomano, R.; Lagoutte, F.; Gourdon, J. Genetic diversity of wheat storage proteins and bread wheat quality. Euphytica 2001, 119, 59–67. [Google Scholar] [CrossRef]
- Hao, C.Y.; Dong, Y.C.; Wang, L.F.; You, G.X.; Zhang, H.N.; Ge, H.M.; Jia, J.Z.; Zhang, X.Y. Genetic diversity and construction of core collection in Chinese wheat genetic resources. Chin. Sci. Bull. 2008, 53, 1518–1526. [Google Scholar] [CrossRef] [Green Version]
- Fu, Y.-B.; Somers, D.J. Genome-wide reduction of genetic diversity in wheat breeding. Crop Sci. 2009, 49, 161–168. [Google Scholar] [CrossRef]
- Bhatta, M.; Shamanin, V.; Shepelev, S.; Baenziger, P.S.; Pozherukova, V.; Pototskaya, I.; Morgounov, A. Genetic diversity and population structure analysis of synthetic and bread wheat accessions in Western Siberia. J. Appl. Genet. 2019, 60, 283–289. [Google Scholar] [CrossRef]
- Adonina, I.G.; Leonova, I.N.; Badaeva, E.D.; Salina, E.A. Genotyping of hexaploid wheat varieties from different Russian regions. Russ. J. Genet. Appl. Res. 2017, 7, 6–13. [Google Scholar] [CrossRef]
- Ceoloni, C.; Del Signore, G.; Ercoli, L.; Donini, P. Locating the alien chromatin segment in common wheat—Aegilops longissima mildew resistant transfers. Hereditas 1992, 116, 239–245. [Google Scholar] [CrossRef]
- Kuzmanović, L.; Rossini, F.; Ruggeri, R.; Pagnotta, M.A.; Ceoloni, C. Engineered durum wheat germplasm with multiple alien introgressions: Agronomic and quality performance. Agronomy 2020, 10, 486. [Google Scholar] [CrossRef] [Green Version]
- Cuadrado, A.; Vitellozzi, F.; Jouve, N.; Ceoloni, C. Fluorescence in situ hybridization with multiple repeated DNA probes applied to the analysis of wheat-rye chromosome pairing. Theor. Appl. Genet. 1997, 94, 347–355. [Google Scholar] [CrossRef]
- Tang, Z.X.; Yang, Z.J.; Fu, S.L. Oligonucleotides replacing the roles of repetitive sequences pAs1, pSc119.2, pTa-535, pTa71, CCS1, and pAWRC.1 for FISH analysis. J. Appl. Genet. 2014, 55, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.L.; Chen, L.; Wang, Y.Y.; Li, M.; Yang, Z.J.; Qiu, L.; Yan, B.J.; Ren, Z.L.; Tang, Z.X. Oligonucleotide probes for ND-FISH analysis to identify rye and wheat chromosomes. Sci. Rep. 2015, 5, 10552. [Google Scholar] [CrossRef] [Green Version]
- Jiang, M.; Xaio, Z.Q.; Fu, S.L.; Tang, Z.X. FISH karyotype of 85 common wheat cultivars/lines displayed by ND-FISH using oligonucleotide probes. Cereal Res. Commun. 2017, 45, 549–563. [Google Scholar] [CrossRef] [Green Version]
- Guo, J.; Gao, D.; Gong, W.P.; Li, H.S.; Li, J.B.; Li, G.R.; Song, J.M.; Liu, J.J.; Yang, Z.J.; Liu, C. Genetic diversity in common wheat lines revealed by fluorescence in situ hybridization. Plant Syst. Evol. 2019, 305, 247–254. [Google Scholar] [CrossRef]
- Ren, T.H.; He, M.J.; Sun, Z.X.; Tian, F.Q.; Luo, P.G.; Tang, Z.X.; Fu, S.L.; Yan, B.J.; Ren, Z.L.; Li, Z. The polymorphisms of oligonucleotide probes in wheat cultivars determined by ND-FISH. Molecules 2019, 24, 1126. [Google Scholar] [CrossRef] [Green Version]
- Hu, Z.L.; Luo, J.T.; Wan, L.R.; Luo, J.; Li, Y.Z.; Fu, S.L.; Liu, D.C.; Hao, M.; Tang, Z.X. Chromosomes polymorphisms of Sichuan wheat cultivars displayed by ND-FISH landmarks. Cereal Res. Commun. 2021, 50, 253–262. [Google Scholar] [CrossRef]
- Fleitas, M.C.; Mondal, S.; Gerard, G.S.; Hernández-Espinosa, N.; Singh, R.P.; Crossa, J.; Guzmán, C. Identification of CIMMYT spring bread wheat germplasm maintaining superior grain yield and quality under heat-stress. J. Cereal Sci. 2020, 93, 102981. [Google Scholar] [CrossRef]
- Singh, V.K.; Singh, P.K.; Harikrishna; Mathuria, R.C.; Gogoi, R.; Aggarwal, R. Assessment of slow rusting resistance components to stripe rust pathogen in some exotic wheat germplasm. Indian Phytopathol. 2017, 70, 52–57. [Google Scholar] [CrossRef] [Green Version]
- Piñera-Chavez, F.J.; Berry, P.M.; Foulkes, M.J.; Molero, G.; Reynolds, M.P. Avoiding lodging in irrigated spring wheat. II. Genetic variation of stem and root structural properties. Field Crop. Res. 2016, 196, 64–74. [Google Scholar] [CrossRef]
- Yang, Y.M.; Qu, J.P.; Zhang, J.; Xiang, S.J.; Li, J.; Wan, H.S. Seed storability of CIMMYT core wheat germplasm panel and their haplotypes in lipoxygenase locus. Agric. Sci. 2021, 12, 977–989. [Google Scholar] [CrossRef]
- Zhang, J.; Xiang, S.J.; Wan, H.S. Negative association between seed dormancy and seed longevity in bread wheat. Am. J. Plant Sci. 2021, 12, 347–365. [Google Scholar] [CrossRef]
- Mahdavi, S.; Arzani, A.; Maibody, S.A.M.M.; Kadivar, M. Grain quality of wheat genotypes under heat stress. Preprints 2021, 2021090447. [Google Scholar]
- Turuspekov, Y.; Baibulatova, A.; Yermekbayev, K.; Tokhetova, L.; Chudinov, V.; Sereda, G.; Ganal, M.; Griffiths, S.; Abugalieva, S. GWAS for plant growth stages and yield components in spring wheat (Triticum aestivum L.) harvested in three regions of Kazakhstan. BMC Plant Biol. 2017, 17, 190–200. [Google Scholar] [CrossRef] [Green Version]
- Huang, L.; Liu, T.G.; Liu, B.; Gao, L.; Luo, P.G.; Chen, W.Q. Resistance evaluation of 197 Chinese wheat core germplasms to a new stripe rust race, CYR34. Plant Prot. 2019, 45, 148–154. [Google Scholar] [CrossRef]
- Qiu, Y.C.; Li, J.J.; Rong, Z.X.; Zhu, H.R. Identification and analysis of China wheat micro-core germplasms resistance to stem rust, stripe rust and powdery mildew. J. Triticeae Crop. 2015, 35, 268–273. [Google Scholar] [CrossRef]
- Yang, X.; Tan, B.W.; Yang, Y.L.; Zhang, X.H.; Zhu, W.; Xu, L.L.; Wang, Y.; Zeng, J.; Fan, X.; Sha, L.N.; et al. Genetic diversity of Asian and European common wheat lines assessed by fluorescence in situ hybridization. Genome 2021, 64, 959–968. [Google Scholar] [CrossRef]
- Huang, X.Q.; Zeller, F.J.; Hsam, S.L.K.; Wenzel, G.; Mohler, V. Chromosomal location of AFLP markers in common wheat (Triticum aestivum L.) utilizing nulli-tetrasomic stocks. Genome 2000, 43, 298–305. [Google Scholar] [CrossRef]
- Huang, X.Q.; Börner, A.; Röder, M.S.; Ganal, M.W. Assessing genetic diversity of wheat (Triticum aestivum L.) germplasm using microsatellite markers. Theor. Appl. Genet. 2002, 105, 699–707. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, N.H.; Backes, G.; Stougaard, J.; Andersen, S.U.; Jahoor, A. Genetic diversity and population structure analysis of European hexaploid bread wheat (Triticum aestivum L.) varieties. PLoS ONE 2014, 9, e94000. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eltaher, S.; Sallam, A.; Belamkar, V.; Emara, H.A.; Nower, A.A.; Salem, K.F.M.; Poland, J.; Baenziger, P.S. Genetic diversity and population structure of F3:6 Nebraska winter wheat genotypes using genotyping-by-sequencing. Front. Genet. 2018, 9, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Mourad, A.M.I.; Belamkar, V.; Baenziger, P.S. Molecular genetic analysis of spring wheat core collection using genetic diversity, population structure, and linkage disequilibrium. BMC Genom. 2020, 21, 434–445. [Google Scholar] [CrossRef] [PubMed]
- Wingen, L.U.; West, C.; Leverington-Waite, M.; Collier, S.; Orford, S.; Goram, R.; Yang, C.-Y.; King, J.; Allen, A.M.; Burridge, A.; et al. Wheat landrace genome diversity. Genetics 2017, 205, 1657–1676. [Google Scholar] [CrossRef] [Green Version]
- Zhao, C.H.; Cui, F.; Wang, X.Q.; Shan, S.C.; Li, X.F.; Bao, Y.G.; Wang, H.G. Effects of 1BL/1RS translocation in wheat on agronomic performance and quality characteristics. Field Crop. Res. 2012, 127, 79–84. [Google Scholar] [CrossRef]
- Qi, Z.J.; Liu, D.J.; Chen, P.D.; Li, Q.Q. Molecular cytogenetic analysis of winter wheat germplasm Aimengniu. Acta Bot. Sin. 2001, 5, 469–474. [Google Scholar]
- Qi, Z.J.; Liu, D.J.; Chen, P.D.; Wang, S.L.; Li, S.S.; Li, Q.Q. Genetic transmission characteristics of the new wheat-rye complex translocation discovered in winter wheat germplasm Aimengniu. Acta Agron. Sin. 2001, 5, 582–589. [Google Scholar]
- Riley, R.; Coucoli, H.; Chapman, V. Chromosomal interchanges and the phylogeny of wheat. Heredity 1967, 22, 233–247. [Google Scholar] [CrossRef]
- Badaeva, E.D.; Dedkova, O.S.; Gay, G.; Pukhalskyi, V.A.; Zelenin, A.V.; Bernard, S.; Bernard, M. Chromosomal rearrangements in wheat: Their types and distribution. Genome 2007, 50, 907–926. [Google Scholar] [CrossRef]
- Walkowiak, S.; Gao, L.L.; Monat, C.; Haberer, G.; Kassa, M.T.; Brinton, J.; Ramirez-Gonzalez, R.H.; Kolodziej, M.C.; Delorean, E.; Thambugala, D.; et al. Multiple wheat genomes reveal global variation in modern breeding. Nature 2020, 588, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.Y.; Li, G.R.; Wan, H.S.; Li, L.P.; Li, J.; Yang, W.Y.; Pu, Z.J.; Yang, Z.J.; Yang, E.N. Identification of QTLs for stripe rust resistance in a recombinant inbred line population. Int. J. Mol. Sci. 2019, 20, 3410. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, M.Y.; Yang, Z.J.; Yang, W.Y.; Yang, E.N. Development and identification of new wheat varieties (lines) with multiple translocation chromosomes via cytogenetic method. J. Triticeae Crop. 2018, 38, 127–133. [Google Scholar] [CrossRef]
- Kawahara, T.; Taketa, S. Fixation of translocation 2A·4B infers the monophyletic origin of Ethiopian tetraploid wheat. Theor. Appl. Genet. 2000, 101, 705–710. [Google Scholar] [CrossRef]
- Lage, J.; Warburton, M.L.; Crossa, J.; Skovmand, B.; Andersen, S.B. Assessment of genetic diversity in synthetic hexaploid wheats and their Triticum dicoccum and Aegilops tauschii parents using AFLPs and agronomic traits. Euphytica 2003, 134, 305–317. [Google Scholar] [CrossRef]
- Zhang, P.; Dreisigacker, S.; Melchinger, A.E.; Reif, J.C.; Kazi, A.M.; Van Ginkel, M.; Hoisington, D.; Warburton, M.L. Quantifying novel sequence variation and selective advantage in synthetic hexaploid wheats and their backcross-derived lines using SSR markers. Mol. Breed. 2005, 15, 1–10. [Google Scholar] [CrossRef]
- Huang, J.K.; Xiang, C.; Wang, Y.Q. The Impact of CIMMYT Wheat Germplasm on Wheat Productivity in China; Center for Chinese Agricultural Policy (CCAP), Chinese Academy of Science (CAS): Beijing, China, 2015. [Google Scholar]
- Han, F.P.; Lamb, J.C.; Birchler, J.A. High frequency of centromere inactivation resulting in stable dicentric chromosomes of maize. Proc. Natl. Acad. Sci. USA 2006, 103, 3238–3243. [Google Scholar] [CrossRef] [Green Version]


| Number | Name | No. of Chr | Translocation |
|---|---|---|---|
| CIMCOG1 | ATTILA | 2n = 42 | |
| CIMCOG2 | ATTILA∗2/PBW65 | 2n = 42 | |
| CIMCOG3 | ATTILA∗2/PBW65∗2/5/REH/HARE//2∗BCN/3/CROC_1/AE.SQUARROSA (213)//PGO/4/HUITES | 2n = 42 | |
| CIMCOG4 | ATTILA//PGO/SERI/3/PASTOR | 2n = 42 | |
| CIMCOG5 | BABAX/LR42//BABAX/3/ER2000 | 2n = 42 | |
| CIMCOG6 | BABAX/LR42//BABAX/3/VORB | 2n = 28 | |
| CIMCOG7 | BACANORA T 88 | 2n = 42 | |
| CIMCOG8 | BAVIACORA M 92 | 2n = 42 | 5BS.7BS, 5BL.7BL |
| CIMCOG9 | BCN/RIALTO | 2n = 42 | |
| CIMCOG10 | BCN/WBLL1 | 2n = 42 | |
| CIMCOG11 | BECARD | 2n = 42 | |
| CIMCOG12 | BECARD/KACHU | 2n = 42 | |
| CIMCOG13 | BRBT1∗2/KIRITATI | 2n = 42 | |
| CIMCOG14 | C80.1/3∗BATAVIA//2∗WBLL1/5/REH/HARE//2∗BCN/3/CROC_1/AE.SQUARROSA (213)//PGO/4/HUITES | 2n = 42 | 1RS.1BL |
| CIMCOG15 | SAUAL/4/CROC_1/AE.SQUARROSA (205)//KAUZ/3/ATTILA/5/SAUAL | 2n = 42 | |
| CIMCOG16 | SAUAL/WHEAR//SAUAL | 2n = 42 | |
| CIMCOG17 | CHIR3/4/SIREN//ALTAR 84/AE.SQUARROSA (205)/3/3∗BUC/5/PFAU/WEAVER | 2n = 42 | 1RS.1BL, 5BS.7BS, 5BL.7BL |
| CIMCOG18 | CHWL86/6/FILIN/IRENA/5/CNDO/R143//ENTE/MEXI_2/3/AEGILOPS SQUARROSA (TAUS)/4/WEAVER | 2n = 42 | |
| CIMCOG19 | CMH79A.955/4/AGA/3/4∗SN64/CNO67//INIA66/5/NAC/6/RIALTO | 2n = 42 | |
| CIMCOG20 | CIRNO C 2008 | 2n = 42 | |
| CIMCOG21 | CNDO/R143//ENTE/MEXI_2/3/AEGILOPS SQUARROSA (TAUS)/4/OCI/5/PASTOR/6/TEMPORALERA M 87/ROMO96 | 2n = 42 | |
| CIMCOG22 | CNDO/R143//ENTE/MEXI_2/3/AEGILOPS SQUARROSA (TAUS)/4/WEAVER/5/2∗KAUZ | 2n = 42 | |
| CIMCOG23 | CNO79//PF70354/MUS/3/PASTOR/4/BAV92∗2/5/FH6-1-7 | 2n = 42 | |
| CIMCOG24 | CROC_1/AE.SQUARROSA (205)//BORL95/3/PRL/SARA//TSI/VEE#5/4/FRET2 | 2n = 42 | |
| CIMCOG25 | GK ARON/AG SECO 7846//2180/4/2∗MILAN/KAUZ//PRINIA/3/BAV92 | 2n = 42 | |
| CIMCOG26 | KINGBIRD #1//INQALAB 91∗2/TUKURU | 2n = 28 | |
| CIMCOG27 | KFA/3/PFAU/WEAVER//BRAMBLING/4/PFAU/WEAVER∗2//BRAMBLING | 2n = 42 | |
| CIMCOG28 | MEX94.27.1.20/3/SOKOLL//ATTILA/3∗BCN | 2n = 42 | |
| CIMCOG29 | MILAN/KAUZ//PRINIA/3/BAV92 | 2n = 42 | |
| CIMCOG30 | MUNAL #1 | 2n = 42 | |
| CIMCOG31 | ATTILA/PASTOR | 2n = 42 | |
| CIMCOG32 | OASIS/5∗BORL95/5/CNDO/R143//ENTE/MEXI75/3/AE.SQ/4/2∗OCI | 2n = 42 | |
| CIMCOG33 | MISR 1 | 2n = 42 | |
| CIMCOG34 | OASIS/SKAUZ//4∗BCN/3/2∗PASTOR/5/FRET2∗2/4/SNI/TRAP#1/3/KAUZ∗2/TRAP//KAUZ/6/SAUAL #1 | 2n = 42 | |
| CIMCOG35 | PANDORA//WBLL1∗2/BRAMBLING | 2n = 42 | |
| CIMCOG36 | PASTOR/3/URES/JUN//KAUZ/4/WBLL1 | 2n = 42 | |
| CIMCOG37 | PAVON F 76 | 2n = 42 | |
| CIMCOG38 | PBW343∗2/KUKUNA∗2//FRTL/PIFED | 2n = 42 | 1RS.1BL |
| CIMCOG39 | PFAU/SERI.1B//AMAD/3/WAXWING | 2n = 42 | |
| CIMCOG40 | ARMENT//2∗SOOTY_9/RASCON_37/4/CNDO/PRIMADUR//HAI-OU_17/3/SNITAN | 2n = 42 | |
| CIMCOG41 | QUAIU #3//MILAN/AMSEL | 2n = 42 | |
| CIMCOG42 | RL6043/4∗NAC//2∗PASTOR | 2n = 42 | 5BS.7BS, 5BL.7BL |
| CIMCOG43 | ROLF07∗2/5/REH/HARE//2∗BCN/3/CROC_1/AE.SQUARROSA (213)//PGO/4/HUITES | 2n = 42 | 1RS.1BL |
| CIMCOG44 | SERI M 82 | 2n = 42 | |
| CIMCOG45 | SIETE CERROS T66 | 2n = 42 | 1RS.1BL |
| CIMCOG46 | SOKOLL∗2/3/BABAX/LR42//BABAX | 2n = 42 | 1RS.1BL |
| CIMCOG47 | SOKOLL//PBW343∗2/KUKUNA/3/ATTILA/PASTOR | 2n = 42 | |
| CIMCOG48 | TACUPETO F2001/SAUAL/4/BABAX/LR42//BABAX∗2/3/KURUKU | 2n = 42 | |
| CIMCOG49 | TACUPETO F2001/BRAMBLING∗2//KACHU | 2n = 42 | |
| CIMCOG50 | TC870344/GUI//TEMPORALERA M 87/AGR/3/2∗WBLL1 | 2n = 42 | |
| CIMCOG51 | TRAP#1/BOW/3/VEE/PJN//2∗TUI/4/BAV92/RAYON/5/KACHU #1 | 2n = 42 | |
| CIMCOG52 | TRCH/SRTU//KACHU | 2n = 42 | |
| CIMCOG53 | UP2338∗2/4/SNI/TRAP#1/3/KAUZ∗2/TRAP//KAUZ/5/MILAN/KAUZ//CHIL/CHUM18/6/UP2338∗2/4/SNI/TRAP#1/3/KAUZ∗2/TRAP//KAUZ | 2n = 42 | 5BS.7BS, 5BL.7BL |
| CIMCOG54 | W15.92/4/PASTOR//HXL7573/2∗BAU/3/WBLL1 | 2n = 42 | |
| CIMCOG55 | BECARD | 2n = 42 | |
| CIMCOG56 | WBLL1∗2/4/BABAX/LR42//BABAX/3/BABAX/LR42//BABAX | 2n = 42 | |
| CIMCOG57 | WBLL1∗2/KURUKU∗2/5/REH/HARE//2∗BCN/3/CROC_1/AE.SQUARROSA (213)//PGO/4/HUITES | 2n = 42 | |
| CIMCOG58 | WBLL1∗2/TUKURU∗2/4/CROC_1/AE.SQUARROSA (205)//BORL95/3/2∗MILAN | 2n = 42 | |
| CIMCOG59 | WHEAR/SOKOLL | 2n = 42 | 1RS.1BL, 5BS.7BS, 5BL.7BL |
| CIMCOG60 | YAV_3/SCO//JO69/CRA/3/YAV79/4/AE.SQUARROSA (498)/5/LINE 1073/6/KAUZ∗2/4/CAR//KAL/BB/3/NAC/5/KAUZ/7/KRONSTAD F2004/8/KAUZ/PASTOR//PBW343 | 2n = 42 |
| Number | Name | Variety Type | No. of Chr | Translocation | Number | Name | Variety Type | No. of Chr | Translocation |
|---|---|---|---|---|---|---|---|---|---|
| CACG1 | Daqingmai | landrace | 2n = 42 | CACG48 | Baihuomia | landrace | 2n = 42 | ||
| CACG2 | Guangmai | landrace | 2n = 42 | CACG49 | Sanyuehuang | landrace | 2n = 42 | ||
| CACG3 | Xinkehan9 | breeding variety | 2n = 42 | CACG50 | Hongqiangchang | landrace | 2n = 42 | ||
| CACG4 | Kefeng3 | breeding variety | 2n = 42 | 2AS.5AS-5AL/5AS.2AS-2AL | CACG51 | Quzimai | landrace | 2n = 42 | |
| CACG5 | Kelao4 | breeding variety | 2n = 42 | CACG52 | Pingyuan50 | breeding variety | 2n = 42 | ||
| CACG6 | Xinshuguang1 | breeding variety | 2n = 42 | 1AS.4BS, 1AL.4BL | CACG53 | Baibiansui | landrace | 2n = 42 | |
| CACG7 | Dongnong101 | breeding variety | 2n = 42 | CACG54 | Gejiaxiang | landrace | 2n = 42 | ||
| CACG8 | Xinshuguang6 | breeding variety | 2n = 42 | CACG55 | Dachunbaisilengmai | landrace | 2n = 42 | ||
| CACG9 | Jichun1016 | breeding variety | 2n = 42 | CACG56 | Zhahong | landrace | 2n = 42 | ||
| CACG10 | Chixiaomai | landrace | 2n = 42 | CACG57 | Motuoxiaomai | landrace | 2n = 42 | ||
| CACG11 | st2422/464 | introduced variety | 2n = 42 | CACG58 | Bianbachunmai-6 | landrace | 2n = 42 | ||
| CACG12 | Orofen | introduced variety | 2n = 42 | 3BS.4AL, 3BL.4AS | CACG59 | Baimangxiaomai | landrace | 2n = 42 | |
| CACG13 | Nonglin10 | introduced variety | 2n = 42 | CACG60 | Wujiangzhuo | landrace | 2n = 42 | ||
| CACG14 | introduced variety | 2n = 42 | CACG61 | Kangdingxiaomai | breeding variety | 2n = 42 | |||
| CACG15 | Early premiun | introduced variety | 2n = 42 | CACG62 | Zangdong4 | breeding variety | 2n = 42 | ||
| CACG16 | Triumph | introduced variety | 2n = 42 | CACG63 | Rikaze54 | breeding variety | 2n = 42 | ||
| CACG17 | Lovrin | introduced variety | 2n = 42 | 1RS.1BL | CACG64 | Rikaze8 | breeding variety | 2n = 42 | |
| CACG18 | introduced variety | 2n = 42 | CACG65 | Yizhimai | landrace | 2n = 42 | |||
| CACG19 | Tanori | introduced variety | 2n = 42 | CACG66 | Dabaimai | landrace | 2n = 42 | ||
| CACG20 | Atlas66 | introduced variety | 2n = 42 | CACG67 | Laohan | landrace | 2n = 42 | ||
| CACG21 | Gansu96 | breeding variety | 2n = 42 | CACG68 | Huoliyan | landrace | 2n = 42 | 4BS.2AS, 4BL.2AL | |
| CACG22 | Chaoanxiaomai | landrace | 2n = 42 | CACG69 | Shanmai | landrace | 2n = 42 | 4AS.4BL, 4AL.4BS | |
| CACG23 | Chike | landrace | 2n = 42 | 2BL.4DS, 2BS.4DL | CACG70 | Hongtuzi | landrace | 2n = 42 | |
| CACG24 | Songruimai4 | breeding variety | 2n = 42 | CACG71 | Baidatou | landrace | 2n = 42 | ||
| CACG25 | Shengen | landrace | 2n = 42 | CACG72 | Jinhuangmai | landrace | 2n = 42 | ||
| CACG26 | Shanglinxiaomai | landrace | 2n = 42 | CACG73 | Huzhuhong | landrace | 2n = 42 | ||
| CACG27 | Pingyang27 | breeding variety | 2n = 42 | CACG74 | Jinmai4 | breeding variety | 2n = 42 | ||
| CACG28 | Jinan2 | breeding variety | 2n = 42 | 1DL.4AL, 1DS.4AS | CACG75 | Dingxi24 | breeding variety | 2n = 42 | |
| CACG29 | Qubao | landrace | 2n = 42 | CACG76 | Ning10 | breeding variety | 2n = 42 | ||
| CACG30 | Bainong3217 | breeding variety | 2n = 42 | CACG77 | Fan6 | breeding variety | 2n = 42 | ||
| CACG31 | Yannong15 | breeding variety | 2n = 42 | CACG78 | Guinong10 | breeding variety | 2n = 42 | 1RS.1BL | |
| CACG32 | Xinong6028 | breeding variety | 2n = 42 | CACG79 | Yunmai34 | breeding variety | 2n = 42 | 1RS.1BL | |
| CACG33 | Jibei2 | breeding variety | 2n = 42 | 1RS.1BL | CACG80 | Xingyi4 | breeding variety | 2n = 42 | 1RS.1BL |
| CACG34 | Neichan5 | breeding variety | 2n = 42 | CACG81 | Fengmai11 | breeding variety | 2n = 42 | ||
| CACG35 | Zhengzhou6 | breeding variety | 2n = 42 | CACG82 | Tongjiabaxiaomai | landrace | 2n = 42 | ||
| CACG36 | Jinan17 | breeding variety | 2n = 42 | CACG83 | Honghuamai | landrace | 2n = 42 | ||
| CACG37 | Shannong7859 | breeding variety | 2n = 42 | 1RS.1BL | CACG84 | Baimaizi | landrace | 2n = 42 | 5BS.2DS, 5BL.2DL |
| CACG38 | Aifeng3 | breeding variety | 2n = 42 | CACG85 | Chengduguangtou | landrace | 2n = 42 | ||
| CACG39 | Lumai1 | breeding variety | 2n = 42 | 1RS.7DS, 1BL.7DL | CACG86 | Jiangmai | landrace | 2n = 42 | |
| CACG40 | Wenmai6 | breeding variety | 2n = 42 | CACG87 | Baihuamai | landrace | 2n = 42 | ||
| CACG41 | Laizhou953 | breeding variety | 2n = 42 | 1RS.1BL | CACG88 | Huanxiangmai | landrace | 2n = 42 | |
| CACG42 | Baimangmai | landrace | 2n = 42 | CACG89 | Hanzhongbai | landrace | 2n = 42 | ||
| CACG43 | Huangguaxian | landrace | 2n = 42 | CACG90 | Xiaosanyuehuang | landrace | 2n = 42 | ||
| CACG44 | Banjiemang | landrace | 2n = 42 | CACG91 | Suotiaohongmai | landrace | 2n = 42 | ||
| CACG45 | Quanguding | landrace | 2n = 42 | CACG92 | Hongxumai | landrace | 2n = 42 | ||
| CACG46 | Xishanbiansui | landrace | 2n = 42 | CACG93 | Zimai | landrace | 2n = 42 | ||
| CACG47 | Honggoudou | landrace | 2n = 42 | CACG94 | Chinese Spring | landrace | 2n = 42 | Reference genome |
| Chr | No. of Types | Type | CIMCOG Wheat Lines | CACG Wheat Lines | Chr | No. of Types | Type | CIMCOG Wheat Lines | CACG Wheat Lines | Chr | No. of Types | Type | CIMCOG Wheat Lines | CACG Wheat Lines | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No. of Lines | Percent (%) | No. of Lines | Percent (%) | No. of Lines | Percent (%) | No. of Lines | Percent (%) | No. of Lines | Percent (%) | No. of Lines | Percent (%) | |||||||||
| 1A | 2 | I | 48 | 80.00 | 1B | 8 | I | 2 | 3.33 | 1D | 8 | I | 27 | 45.00 | 60 | 64.52 | ||||
| II | 12 | 20.00 | 92 | 98.92 | II | 5 | 8.33 | 41 | 44.09 | II | 21 | 35.00 | 6 | 6.45 | ||||||
| 2A | 13 | I | 13 | 21.67 | III | 35 | 58.33 | 8 | 8.60 | III | 5 | 8.33 | 3 | 3.23 | ||||||
| II | 10 | 16.67 | 14 | 15.05 | IV | 11 | 18.33 | 24 | 25.81 | IV | 5 | 8.33 | 4 | 4.30 | ||||||
| III | 34 | 56.67 | 49 | 52.69 | V | 2 | 2.15 | V | 16 | 17.20 | ||||||||||
| IV | 2 | 3.33 | 4 | 4.30 | VI | 4 | 4.30 | VI | 1 | 1.08 | ||||||||||
| V | 1 | 1.67 | 3 | 3.23 | VII | 4 | 4.30 | VII | 1 | 1.08 | ||||||||||
| VI | 2 | 2.15 | VIII | 2 | 2.15 | VIII | 1 | 1.08 | ||||||||||||
| VII | 2 | 2.15 | 2B | 10 | I | 6 | 10.00 | 2D | 5 | I | 1 | 1.67 | ||||||||
| VIII | 4 | 4.30 | II | 19 | 31.67 | 5 | 5.38 | II | 14 | 23.33 | 24 | 25.81 | ||||||||
| IX | 4 | 4.30 | III | 12 | 20.00 | 24 | 25.81 | III | 40 | 66.67 | 51 | 54.84 | ||||||||
| X | 4 | 4.30 | IV | 11 | 18.33 | 7 | 7.53 | IV | 3 | 5.00 | 3 | 3.23 | ||||||||
| XI | 1 | 1.08 | V | 12 | 20.00 | 2 | 2.15 | V | 14 | 15.05 | ||||||||||
| XII | 2 | 2.15 | VI | 19 | 20.43 | 3D | 4 | I | 4 | 6.67 | ||||||||||
| XIII | 2 | 2.15 | VII | 7 | 7.53 | II | 42 | 70.00 | 37 | 39.78 | ||||||||||
| 3A | 4 | I | 60 | 100.00 | 80 | 86.02 | VIII | 8 | 8.60 | III | 11 | 18.33 | 55 | 59.14 | ||||||
| II | 7 | 7.53 | IX | 17 | 18.28 | IV | 1 | 1.67 | 1 | 1.08 | ||||||||||
| III | 5 | 5.38 | X | 3 | 3.23 | 4D | 4 | I | 6 | 10.00 | ||||||||||
| IV | 1 | 1.08 | 3B | 12 | I | 2 | 3.33 | II | 47 | 78.33 | 48 | 51.61 | ||||||||
| 4A | 10 | I | 20 | 33.33 | II | 16 | 26.67 | 8 | 8.60 | III | 5 | 8.33 | 43 | 46.24 | ||||||
| II | 24 | 40.00 | III | 25 | 41.67 | 6 | 6.45 | IV | 1 | 1.08 | ||||||||||
| III | 1 | 1.67 | IV | 6 | 10.00 | 4 | 4.30 | 5D | 3 | I | 50 | 83.33 | 17 | 18.28 | ||||||
| IV | 1 | 1.67 | V | 3 | 5.00 | 21 | 22.58 | II | 5 | 8.33 | 33 | 35.48 | ||||||||
| V | 1 | 1.67 | 6 | 6.45 | VI | 5 | 8.33 | 10 | 10.75 | III | 3 | 5.00 | 43 | 46.24 | ||||||
| VI | 7 | 11.67 | 22 | 23.66 | VII | 3 | 5.00 | 3 | 3.23 | 6D | 3 | I | 58 | 96.67 | 77 | 82.80 | ||||
| VII | 4 | 6.67 | 6 | 6.45 | VIII | 6 | 6.45 | II | 10 | 10.75 | ||||||||||
| VIII | 2 | 3.33 | 24 | 25.81 | IX | 3 | 3.23 | III | 6 | 6.45 | ||||||||||
| IX | 31 | 33.33 | X | 10 | 10.75 | 7D | 4 | I | 4 | 6.67 | ||||||||||
| X | 1 | 1.08 | XI | 13 | 13.98 | II | 46 | 76.67 | 81 | 87.10 | ||||||||||
| 5A | 9 | I | 6 | 10.00 | XII | 8 | 8.60 | III | 8 | 13.33 | 10 | 10.75 | ||||||||
| II | 4 | 6.67 | 4B | 5 | I | 48 | 80.00 | 44 | 47.31 | IV | 1 | 1.08 | ||||||||
| III | 12 | 20.00 | 8 | 8.60 | II | 10 | 16.67 | 37 | 39.78 | 5BS/7BS | 1 | I | 5 | 8.33 | ||||||
| IV | 31 | 51.67 | 39 | 41.94 | III | 2 | 3.33 | 1 | 1.08 | 5BL/7BL | 4 | I | 2 | 3.33 | ||||||
| V | 4 | 6.67 | 7 | 7.53 | IV | 7 | 7.53 | II | 1 | 1.67 | ||||||||||
| VI | 3 | 5.00 | 22 | 23.66 | V | 1 | 1.08 | III | 1 | 1.67 | ||||||||||
| VII | 6 | 6.45 | 5B | 6 | I | 2 | 3.33 | IV | 1 | 1.67 | ||||||||||
| VIII | 9 | 9.68 | II | 36 | 60.00 | 30 | 32.26 | 1RS/1BL | 1 | I | 7 | 11.67 | 7 | 7.53 | ||||||
| IX | 1 | 1.08 | III | 15 | 25.00 | 23 | 24.73 | 1RS.7DS, 1BL.7DL | 1 | 1.08 | ||||||||||
| 6A | 2 | I | 46 | 76.67 | 31 | 33.33 | IV | 1 | 1.67 | 2 | 2.15 | 2AS.5AS-5AL/5AS.2AS-2AL | 1 | 1.08 | ||||||
| II | 14 | 23.33 | 62 | 66.67 | V | 1 | 1.67 | 4 | 4.30 | 1AS.4BS, 1AL.4BL | 1 | 1.08 | ||||||||
| 7A | 7 | I | 38 | 63.33 | 22 | 23.66 | VI | 33 | 35.48 | 3BS.4AL, 3BL.4AS | 1 | 1.08 | ||||||||
| II | 10 | 16.67 | 14 | 15.05 | 6B | 13 | I | 7 | 11.67 | 4BS.2AS, 4BL.2AL | 1 | 1.08 | ||||||||
| III | 8 | 13.33 | 4 | 4.30 | II | 1 | 1.67 | 4AS.4BL,4AL.4BS | 1 | 1.08 | ||||||||||
| IV | 4 | 6.67 | 30 | 32.26 | III | 1 | 1.67 | 2BL.4DS, 2BS.4DL | 1 | 1.08 | ||||||||||
| V | 15 | 16.13 | IV | 31 | 51.67 | 20 | 21.51 | 5BS.2DS, 5BL.2DL | 1 | 1.08 | ||||||||||
| VI | 3 | 3.23 | V | 14 | 23.33 | 10 | 10.75 | 1DL.4AL, 1DS.4AS | 1 | 1.08 | ||||||||||
| VII | 5 | 5.38 | VI | 2 | 3.33 | 3 | 3.23 | |||||||||||||
| VII | 2 | 3.33 | 12 | 12.90 | ||||||||||||||||
| VIII | 2 | 3.33 | 24 | 25.81 | ||||||||||||||||
| IX | 7 | 7.53 | ||||||||||||||||||
| X | 9 | 9.68 | ||||||||||||||||||
| XI | 6 | 6.45 | ||||||||||||||||||
| XII | 1 | 1.08 | ||||||||||||||||||
| XIII | 1 | 1.08 | ||||||||||||||||||
| 7B | 11 | I | 7 | 11.67 | ||||||||||||||||
| II | 3 | 5.00 | ||||||||||||||||||
| III | 3 | 5.00 | ||||||||||||||||||
| IV | 20 | 33.33 | 5 | 5.38 | ||||||||||||||||
| V | 15 | 25.00 | 44 | 47.31 | ||||||||||||||||
| VI | 1 | 1.67 | 10 | 10.75 | ||||||||||||||||
| VII | 1 | 1.67 | 20 | 21.51 | ||||||||||||||||
| VIII | 1 | 1.67 | 4 | 4.30 | ||||||||||||||||
| IX | 4 | 6.67 | 5 | 5.38 | ||||||||||||||||
| X | 3 | 3.23 | ||||||||||||||||||
| XI | 2 | 2.15 | ||||||||||||||||||
| Name of Probe | Sequence and Fluorochrome Label |
|---|---|
| Oligo-pSc119.2-1 | 6-FAM-5′CCGTT TTGTG GACTA TTACT CACCG CTTTG GGGTC CCATA GCTAT3′ |
| Oligo-pTa535-1 | Tamra-5′AAAAA CTTGA CGCAC GTCAC GTACA AATTG GACAA ACTCT TTCGG AGTAT CAGGG TTTC3′ |
| Oligo-(GAA)7 | Cy5-5′GAAGAAGAAGAAGAAGAAGAA3′ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, M.; Yang, Z.; Yang, W.; Yang, E. Genetic Diversity Assessment of the International Maize and Wheat Improvement Center and Chinese Wheat Core Germplasms by Non-Denaturing Fluorescence In Situ Hybridization. Plants 2022, 11, 1403. https://doi.org/10.3390/plants11111403
Yang M, Yang Z, Yang W, Yang E. Genetic Diversity Assessment of the International Maize and Wheat Improvement Center and Chinese Wheat Core Germplasms by Non-Denaturing Fluorescence In Situ Hybridization. Plants. 2022; 11(11):1403. https://doi.org/10.3390/plants11111403
Chicago/Turabian StyleYang, Manyu, Zujun Yang, Wuyun Yang, and Ennian Yang. 2022. "Genetic Diversity Assessment of the International Maize and Wheat Improvement Center and Chinese Wheat Core Germplasms by Non-Denaturing Fluorescence In Situ Hybridization" Plants 11, no. 11: 1403. https://doi.org/10.3390/plants11111403
APA StyleYang, M., Yang, Z., Yang, W., & Yang, E. (2022). Genetic Diversity Assessment of the International Maize and Wheat Improvement Center and Chinese Wheat Core Germplasms by Non-Denaturing Fluorescence In Situ Hybridization. Plants, 11(11), 1403. https://doi.org/10.3390/plants11111403







