Study and Physical Mapping of the Species-Specific Tandem Repeat CS-237 Linked with 45S Ribosomal DNA Intergenic Spacer in Cannabis sativa L.
Abstract
:1. Introduction
2. Results
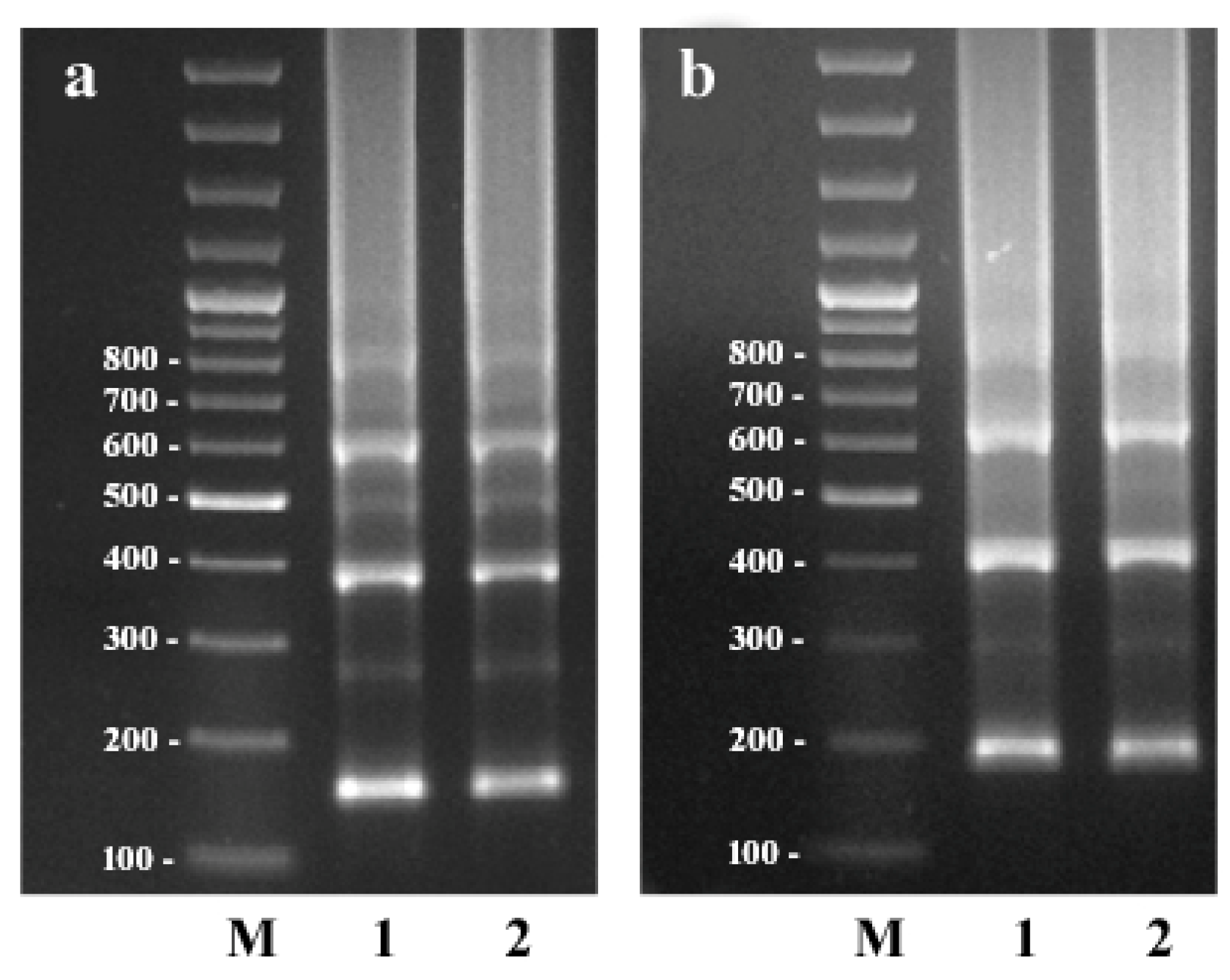
2.1. Search and Amplification of the CS-237 Repeat
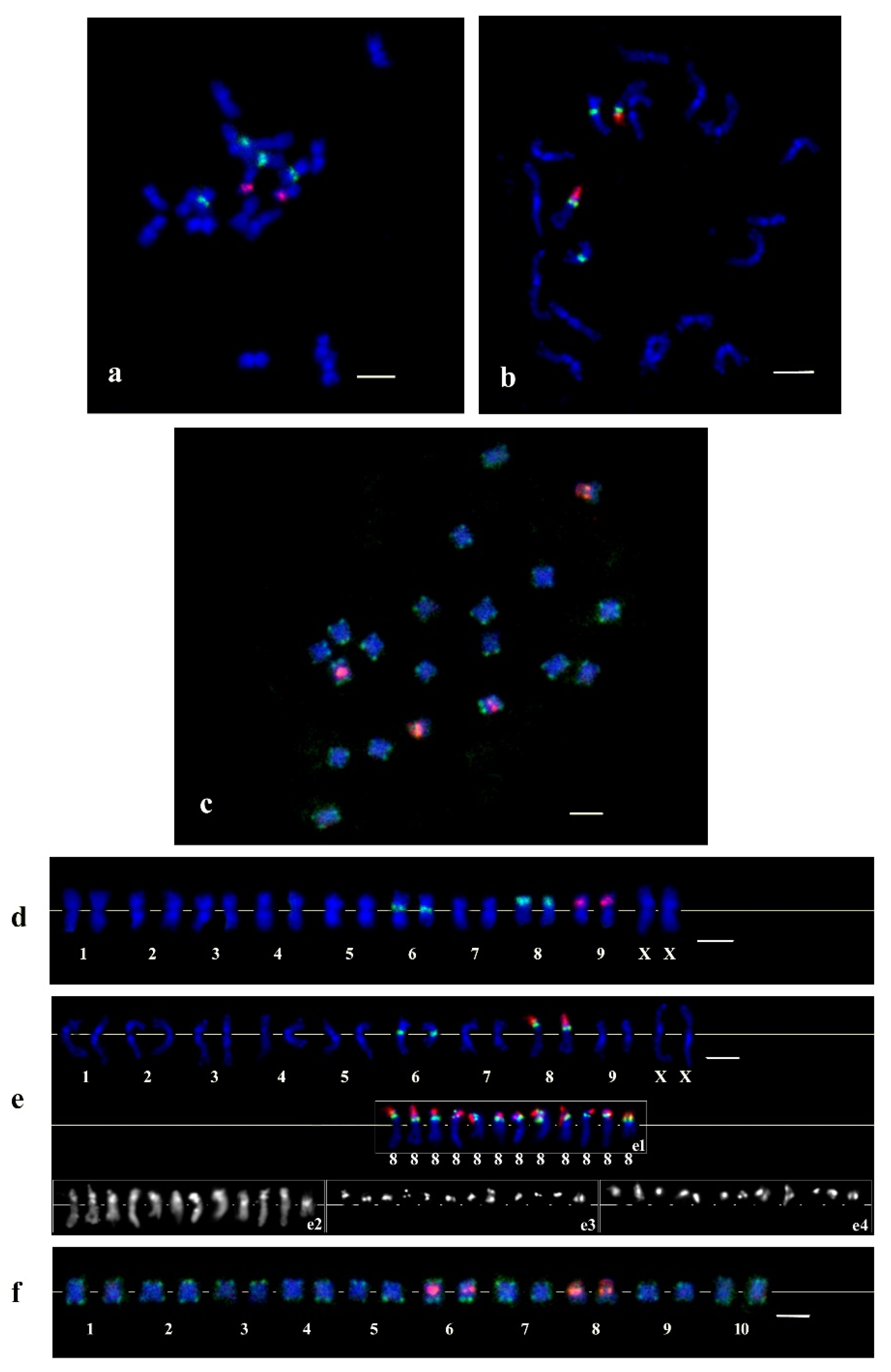
2.2. Physical Mapping of the CS-237 Repeat with Other Cytogenetic Markers on C. sativa Chromosomes
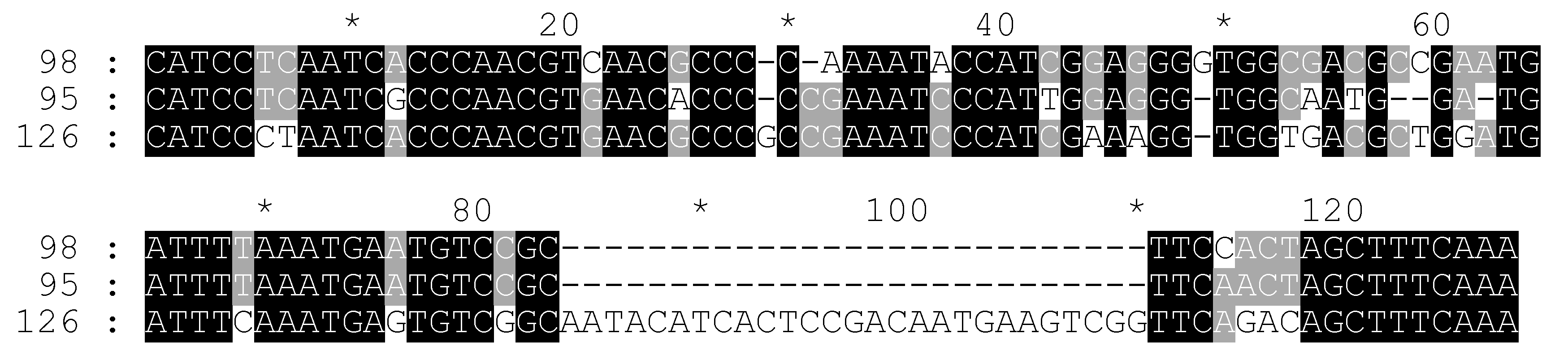
2.3. Molecular Search of the CS-237 in 45S rDNA
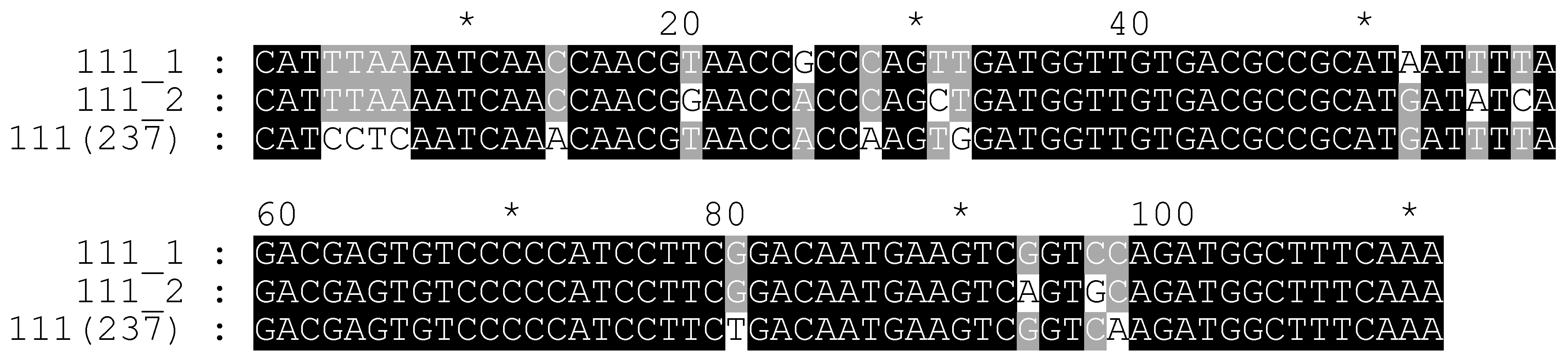
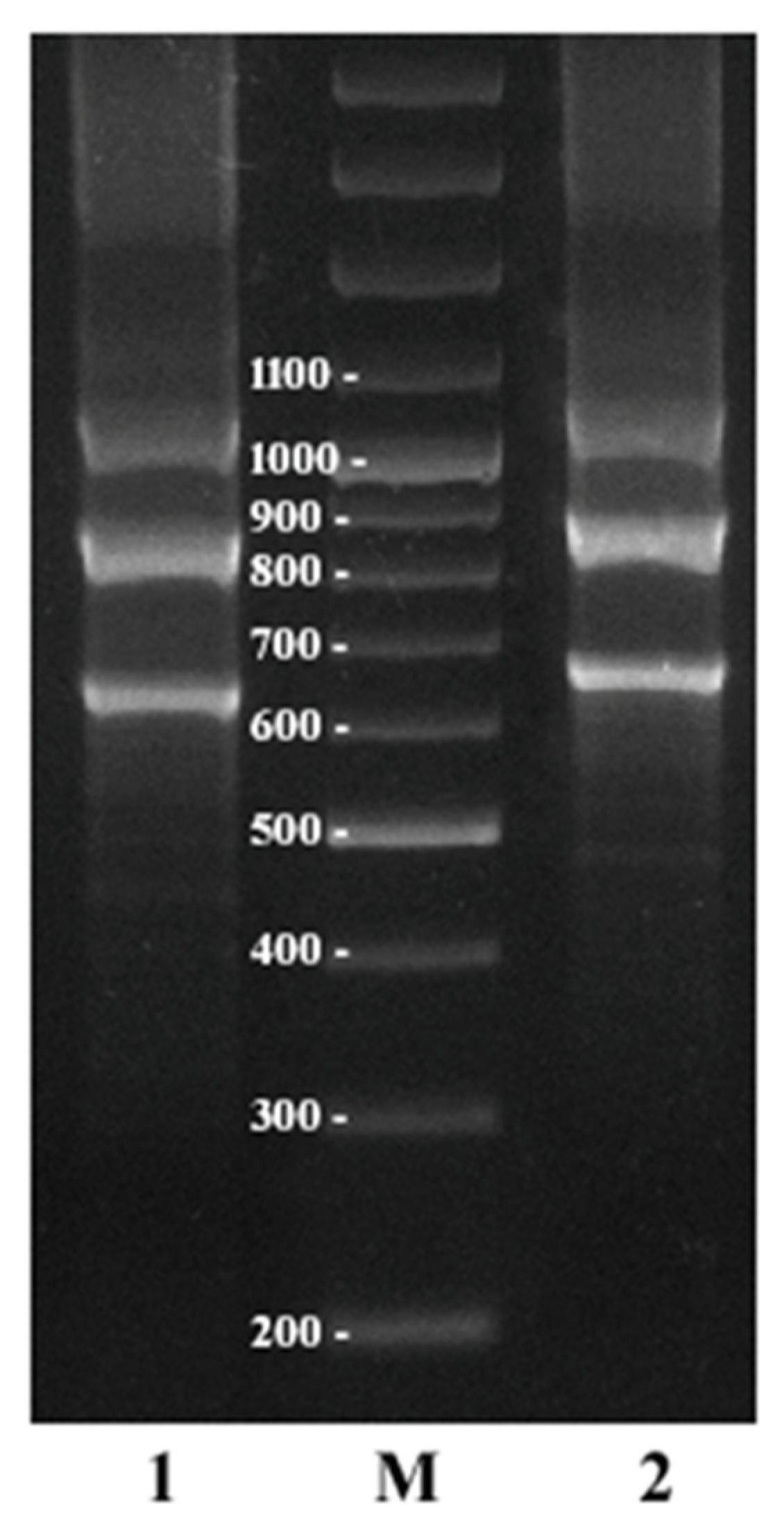
2.4. Additional Study of C. sativa IGS Variants
2.5. Bioinformatical Analysis of CS-237 and delCS-237 Repeats Localization in Chromosomes
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Plant Material and Bioinformatic Analysis
5.2. Chromosome Preparation
5.3. DNA Isolation
5.4. PCR Analysis
5.5. DNA Probes and Fluorescent In Situ Hybridization (FISH)
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Russo, E. Hemp for headache: An in-depth historical and scientific review of cannabis in migraine treatment. J. Cannabis Ther. 2001, 1, 21–92. [Google Scholar] [CrossRef]
- Merlin, M.D. Archaeological evidence for the tradition of psychoactive plant use in the old world. Econ. Bot. 2003, 57, 295–323. [Google Scholar] [CrossRef]
- Cherney, J.H.; Small, E. Industrial hemp in North America: Production, politics and potential. Agronomy 2016, 6, 58. [Google Scholar] [CrossRef] [Green Version]
- Crini, G.; Lichtfouse, E.; Chanet, G.; Morin-Crini, N. Applications of hemp in textiles, paper industry, insulation and building materials, horticulture, animal nutrition, food and beverages, nutraceuticals, cosmetics and hygiene, medicine, agrochemistry, energy production and environment: A review. Environ. Chem. Lett. 2020, 18, 1451–1476. [Google Scholar] [CrossRef]
- Van der Werf, H.M.G.; Mathijssen, E.W.J.M.; Haverkort, A.J. The potential of hemp (Cannabis sativa L.) for sustainable fiber production: A crop physiological appraisal. Ann. Appl. Biol. 1996, 129, 109–123. [Google Scholar] [CrossRef]
- Kubitzki, K. Cannabaceae. In Flowering Plants · Dicotyledons. The Families and Genera of Vascular Plants; Kubitzki, K., Rohwer, J.G., Bittrich, V., Eds.; Springer: Berlin/Heidelberg, Germany, 1993; Volume 2, pp. 204–206. [Google Scholar] [CrossRef]
- Ming, R.; Bendahmane, A.; Renner, S.S. Sex chromosomes in land plants. Annu. Rev. Plant Biol. 2011, 62, 485–514. [Google Scholar] [CrossRef] [Green Version]
- Matsunaga, S.; Kawano, S. Sex determination by sex chromosomes in dioecious plants. Plant Biol. 2001, 3, 481–488. [Google Scholar] [CrossRef]
- Divashuk, M.G.; Alexandrov, O.S.; Kroupin, P.Y.; Karlov, G.I. Molecular cytogenetic mapping of Humulus lupulus sex chromosomes. Cytogenet. Genome Res. 2011, 134, 213–219. [Google Scholar] [CrossRef]
- Alexandrov, O.S.; Divashuk, M.G.; Yakovin, N.A.; Karlov, G.I. Sex chromosome differentiation in Humulus japonicus Siebold & Zuccarini, 1846 (Cannabaceae) revealed by fluorescence in situ hybridization of subtelomeric repeat. Comp. Cytogen. 2012, 6, 239–247. [Google Scholar] [CrossRef]
- Grabowska-Joachimiak, A.; Mosiolek, M.; Lech, A.; Goralski, G. C-banding/DAPI and in situ hybridization reflect karyotype structure and sex chromosome differentiation in Humulus japonicus Siebold&Zucc. Cytogenet. Genome Res. 2011, 132, 203–211. [Google Scholar] [CrossRef]
- Hesami, M.; Pepe, M.; Alizadeh, M.; Rakei, A.; Baiton, A.; Jones, A.M.P. Recent advances in cannabis biotechnology. Ind. Crops Prod. 2000, 158, 113026. [Google Scholar] [CrossRef]
- Guerriero, G.; Behr, M.; Legay, S.; Mangeot-Peter, L.; Zorzan, S.; Ghoniem, M.; Hausman, J.F. Transcriptomic profiling of hemp bast fibres at different developmental stages. Sci. Rep. 2017, 7, 4961. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kovalchuk, I.; Pellino, M.; Rigault, P.; Van Velzen, R.; Ebersbach, J.; Ashnest, J.R.; Mau, M.; Schranz, M.E.; Alcorn, J.; Laprairie, R.B.; et al. The genomics of Cannabis and its close relatives. Annu. Rev. Plant Biol. 2020, 71, 713–739. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Divashuk, M.G.; Alexandrov, O.S.; Razumova, O.V.; Kirov, I.V.; Karlov, G.I. Molecular cytogenetic characterization of the dioecious Cannabis sativa with an XY chromosome sex determination system. PLoS ONE 2014, 9, e85118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gerlach, W.L.; Bedbrook, J.R. Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res. 1979, 7, 1869–1885. [Google Scholar] [CrossRef]
- Pontvianne, F.; Abou-Ellail, M.; Douet, J.; Comella, P.; Matia, I.; Chandrasekhara, C.; Debures, A.; Blevins, T.; Cooke, R.; Medina, F.J.; et al. Nucleolin is required for DNA methylation state and the expression of rRNA gene variants in Arabidopsis thaliana. PLoS Genet. 2010, 6, e1001225. [Google Scholar] [CrossRef] [Green Version]
- Besse, P. Nuclear ribosomal RNA genes: ITS region. In Molecular Plant Taxonomy. Methods in Molecular Biology (Methods and Protocols); Besse, P., Ed.; Humana Press: Totowa, NJ, USA, 2014; Volume 1115, pp. 141–149. [Google Scholar] [CrossRef]
- Kim, K.; Lee, S.C.; Lee, J.; Lee, H.O.; Joh, H.J.; Kim, N.H.; Park, H.S.; Yang, T.J. Comprehensive survey of genetic diversity in chloroplast genomes and 45S nrDNAs within Panax ginseng species. PLoS ONE 2015, 10, e0117159. [Google Scholar] [CrossRef]
- Chang, K.D.; Fang, S.A.; Chang, F.C.; Chung, M.C. Chromosomal conservation and sequence diversity of ribosomal RNA genes of two distant Oryza species. Genomics 2010, 96, 181–190. [Google Scholar] [CrossRef] [Green Version]
- Bauer, N.; Horvat, T.; Birus, I.; Vicić, V.; Zoldos, V. Nucleotide sequence, structural organization and length heterogeneity of ribosomal DNA intergenic spacer in Quercus petraea (Matt.) Liebl. and Q. robur L. Mol. Genet. Genom. 2009, 281, 207–221. [Google Scholar] [CrossRef]
- Yang, K.; Robin, A.H.K.; Yi, G.-E.; Lee, J.; Chung, M.-Y.; Yang, T.-J.; Nou, I.-S. Diversity and inheritance of intergenic spacer sequences of 45S ribosomal DNA among accessions of Brassica oleracea L. var. capitata. Int. J. Mol. Sci. 2015, 16, 28783–28799. [Google Scholar] [CrossRef] [Green Version]
- Almeida, C.; Fonsêca, A.; dos Santos, K.G.; Mosiolek, M.; Pedrosa-Harand, A. Contrasting evolution of a satellite DNA and its ancestral IGS rDNA in Phaseolus (Fabaceae). Genome 2012, 55, 683–689. [Google Scholar] [CrossRef] [PubMed]
- Jo, S.H.; Koo, D.H.; Kim, J.F.; Hur, C.G.; Lee, S.; Yang, T.J.; Kwon, S.Y.; Choi, D. Evolution of ribosomal DNA-derived satellite repeat in tomato genome. BMC Plant Biol. 2009, 9, 42. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raina, S.N.; Sharma, S.; Sasakuma, T.; Kishii, M.; Vaishnavi, S. Novel repeated DNA sequences in safflower (Carthamus tinctorius L.) (Asteraceae): Cloning, sequencing, and physical mapping by fluorescence in situ hybridization. J. Hered. 2005, 96, 424–429. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Charlesworth, B.; Sniegowski, P.; Stephan, W. The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 1994, 371, 215–220. [Google Scholar] [CrossRef]
- Garrido-Ramos, M.A. Satellite DNA: An evolving topic. Genes 2017, 8, 230. [Google Scholar] [CrossRef]
- Ruiz-Ruano, F.J.; López-León, M.D.; Cabrero, J.; Camacho, J.P.M. High-throughput analysis of the satellitome illuminates satellite DNA evolution. Sci. Rep. 2016, 6, 28333. [Google Scholar] [CrossRef] [Green Version]
- Alexandrov, O.S.; Karlov, G.I. Molecular cytogenetic analysis and genomic organization of major DNA repeats in castor bean (Ricinus communis L.). Mol. Genet. Genom. 2016, 291, 775–787. [Google Scholar] [CrossRef]
- Lee, Y.I.; Yap, J.W.; Izan, S.; Leitch, I.J.; Fay, M.F.; Lee, Y.C.; Hidalgo, O.; Dodsworth, S.; Smulders, M.J.M.; Gravendeel, B.; et al. Satellite DNA in Paphiopedilum subgenus Parvisepalum as revealed by high-throughput sequencing and fluorescent in situ hybridization. BMC Genom. 2018, 19, 578. [Google Scholar] [CrossRef] [Green Version]
- Meštrović, N.; Mravinac, B.; Pavlek, M.; Vojvoda-Zeljko, T.; Šatović, E.; Plohl, M. Structural and functional liaisons between transposable elements and satellite DNAs. Chromosome Res. 2015, 23, 583–596. [Google Scholar] [CrossRef]
- Ruiz-Ruano, F.J.; Castillo-Martínez, J.; Cabrero, J.; Gómez, R.; Camacho, J.P.M.; López-León, M.D. High-throughput analysis of satellite DNA in the grasshopper Pyrgomorpha conica reveals abundance of homologous and heterologous higher-order repeats. Chromosoma 2018, 127, 323–340. [Google Scholar] [CrossRef]
- Willard, H.F.; Waye, J.S. Chromosome-specific subsets of human alpha satellite DNA: Analysis of sequence divergence within and between chromosomal subsets and evidence for an ancestral pentameric repeat. J. Mol. Evol. 1987, 25, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Warburton, P.E.; Willard, H.F. Genomic analysis of sequence variation in tandemly repeated DNA. Evidence for localized homogeneous sequence domains within arrays of alpha-satellite DNA. J. Mol. Biol. 1990, 216, 3–16. [Google Scholar] [CrossRef]
- Plohl, M.; Meštrović, N.; Mravinac, B. Satellite DNA evolution. In Repetitive DNA; Garrido-Ramos, M.A., Ed.; Karger: Basel, Switzerland, 2012; Volume 7, pp. 126–152. [Google Scholar] [CrossRef]
- Cooney, C.A.; Matthews, H.R. The isolation of satellite DNA by density gradient centrifugation. In Nucleic Acids. Methods in Molecular Biology; Walker, J.M., Ed.; Humana Press: Clifton, NJ, USA, 1984; Volume 2, pp. 21–29. [Google Scholar] [CrossRef]
- Hemleben, V.; Beridze, T.G.; Bakhman, L.; Kovarik, J.; Torrez, R. Satellite DNA. Usp. Biol. Khim. 2003, 43, 267–306. (In Russian) [Google Scholar]
- Van Dijk, E.L.; Auger, H.; Jaszczyszyn, Y.; Thermes, C. Ten years of next-generation sequencing technology. Trends Genet. 2014, 30, 418–426. [Google Scholar] [CrossRef] [PubMed]
- Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 1999, 27, 573–580. [Google Scholar] [CrossRef] [Green Version]
- Novák, P.; Neumann, P.; Pech, J.; Steinhaisl, J.; Macas, J. RepeatExplorer: A Galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 2013, 29, 792–793. [Google Scholar] [CrossRef] [Green Version]
- The Cannabis Genome Browser. Available online: http://genome.ccbr.utoronto.ca/index.html?org=C.+sativa&db=canSat3&hgsid=259020 (accessed on 30 March 2022).
- Hsieh, H.M.; Hou, R.J.; Chen, K.F.; Tsai, L.C.; Liu, S.W.; Liu, K.L.; Linacre, A.; Lee, J.C. Establishing the rDNA IGS structure of Cannabis sativa. J. Forensic Sci. 2004, 49, 477–480. [Google Scholar] [CrossRef]
- Garrido-Ramos, M.A. Satellite DNA in plants: More than just rubbish. Cytogenet. Genome Res. 2015, 146, 153–170. [Google Scholar] [CrossRef]
- Gatto, K.P.; Mattos, J.V.; Seger, K.R.; Lourenço, L.B. Sex chromosome differentiation in the frog genus Pseudis involves satellite DNA and chromosome rearrangements. Front. Genet. 2018, 9, 301. [Google Scholar] [CrossRef]
- Muirhead, C.A.; Presgraves, D.C. Satellite DNA-mediated diversification of a sex-ratio meiotic drive gene family in Drosophila. Nat. Ecol. Evol. 2021, 5, 1604–1612. [Google Scholar] [CrossRef]
- Hobza, R.; Kubat, Z.; Cegan, R.; Jesionek, W.; Vyskot, B.; Kejnovsky, E. Impact of repetitive DNA on sex chromosome evolution in plants. Chromosome Res. 2015, 23, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Puterova, J.; Razumova, O.; Martinek, T.; Alexandrov, O.; Divashuk, M.; Kubat, Z.; Hobza, R.; Karlov, G.; Kejnovsky, E. Satellite DNA and transposable elements in seabuckthorn (Hippophae rhamnoides), a dioecious plant with small Y and large X chromosomes. Genome Biol. Evol. 2017, 9, 197–212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prentout, D.; Razumova, O.; Rhoné, B.; Badouin, H.; Henri, H.; Feng, C.; Käfer, J.; Karlov, G.; Marais, G.A. An efficient RNA-seq-based segregation analysis identifies the sex chromosomes of Cannabis sativa. Genome Res. 2020, 30, 164–172. [Google Scholar] [CrossRef] [PubMed]
- Fry, K.; Salser, W. Nucleotide sequences of HS-alpha satellite DNA from kangaroo rat Dipodomys ordii and characterization of similar sequences in other rodents. Cell 1977, 12, 1069–1084. [Google Scholar] [CrossRef]
- Ugarković, D.; Plohl, M. Variation in satellite DNA profiles—Causes and effects. EMBO J. 2002, 21, 5955–5959. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- King, K.; Jobst, J.; Hemleben, V. Differential homogenization and amplification of two satellite DNAs in the genus Cucurbita (Cucurbitaceae). J. Mol. Evol. 1995, 41, 996–1005. [Google Scholar] [CrossRef]
- Vershinin, A.V.; Alkhimova, E.G.; Heslop-Harrison, J.S. Molecular diversification of tandemly organized DNA sequences and heterochromatic chromosome regions in some Triticeae species. Chromosome Res. 1996, 4, 517–525. [Google Scholar] [CrossRef]
- Ross, R.; Hankeln, T.; Schmidt, E.R. Complex evolution of tandem-repetitive DNA in the Chironomus thummi species group. J. Mol. Evol. 1997, 44, 321–326. [Google Scholar] [CrossRef]
- Landais, I.; Chavigny, P.; Castagnone, C.; Pizzol, J.; Abad, P.; Vanlerberghe-Masutti, F. Characterization of a highly conserved satellite DNA from the parasitoid wasp Trichogramma brassicae. Gene 2000, 255, 65–73. [Google Scholar] [CrossRef]
- Maggini, F.; Cremonini, R.; Zolfino, C.; Tucci, G.F.; D’Ovidio, R.; Delre, V.; DePace, C.; Scarascia Mugnozza, G.T.; Cionini, P.G. Structure and chromosomal localization of DNA sequences related to ribosomal subrepeats in Vicia faba. Chromosoma 1991, 100, 229–234. [Google Scholar] [CrossRef]
- Falquet, J.; Creusot, F.; Dron, M. Molecular analysis of Phaseolus vulgaris rDNA unit and characterization of a satellite DNA homologous to IGS subrepeats. Plant Physiol. Biochem. 1997, 35, 611–622. [Google Scholar]
- Schmidt, E.R. Clustered and interspersed repetitive DNA sequence family of Chironomus: The nucleotide sequence of the Cla-elements and of various flanking sequences. J. Mol. Biol. 1984, 178, 1–15. [Google Scholar] [CrossRef]
- Macas, J.; Navrátilová, A.; Mészáros, T. Sequence subfamilies of satellite repeats related to rDNA intergenic spacer are differentially amplified on Vicia sativa chromosomes. Chromosoma 2003, 112, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.Y.; Skalicka, K.; Koukalova, B.; Volkov, R.A.; Matyasek, R.; Hemleben, V.; Leitch, A.R.; Kovarik, A. Dynamic changes in the distribution of a satellite homologous to intergenic 26–18S rDNA spacer in the evolution of Nicotiana. Genetics 2004, 166, 1935–1946. [Google Scholar] [CrossRef]
- Zerega, N.J.; Clement, W.L.; Datwyler, S.L.; Weiblen, G.D. Biogeography and divergence times in the mulberry family (Moraceae). Mol. Phylogenet. Evol. 2005, 37, 402–416. [Google Scholar] [CrossRef] [Green Version]
- GeneDoc: Analysis and Visualization of Genetic Variation. Available online: http://www.nrbsc.org/gfx/genedoc/ebinet.htm (accessed on 17 March 2022).
- Braich, S.; Baillie, R.; Spangenberg, G.; Cogan, N. A new and improved genome sequence of Cannabis sativa. Gigabyte 2020, 10, 1–13. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T. BLAST+: Architecture and applications. BMC Bioinform. 2008, 10, 421. [Google Scholar] [CrossRef] [Green Version]
- Gao, S.; Wang, B.; Xie, S.; Xu, X.; Zhang, J.; Pei, L.; Yu, Y.; Yang, W.; Zhang, Y. A high-quality reference genome of wild Cannabis sativa. Hortic. Res. 2020, 7, 73. [Google Scholar] [CrossRef]
- Wu, T.; Watanabe, C. GMAP: A genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 2005, 21, 1859–1875. [Google Scholar] [CrossRef] [Green Version]
- Kirov, I.; Divashuk, M.; Van Laere, K.; Soloviev, A.; Khrustaleva, L. An easy “SteamDrop” method for high quality plant chromosome preparation. Mol. Cytogenet. 2014, 7, 21. [Google Scholar] [CrossRef] [Green Version]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissue. Focus 1990, 12, 13–15. [Google Scholar]
- Primer3 Web Service. Available online: https://primer3.ut.ee/ (accessed on 30 March 2022).
- Campell, B.R.; Song, Y.; Posch, T.E.; Cullis, C.A. Town CD Sequence and organization of 5S ribosomal RNA-encoding genes of Arabidopsis thaliana. Gene 1992, 112, 225–228. [Google Scholar] [CrossRef]
- Karlov, G.I.; Danilova, T.V.; Horlemann, C.; Weber, G. Molecular cytogenetic in hop (Humulus lupulus L.) and identification of sex chromosomes by DAPI-banding. Euphytica 2003, 132, 185–190. [Google Scholar] [CrossRef]
- Levan, A.; Fredga, K.; Sandberg, A. Nomenclature for centromeric position on chromosomes. Hereditas 1964, 52, 201–220. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alexandrov, O.S.; Romanov, D.V.; Divashuk, M.G.; Razumova, O.V.; Ulyanov, D.S.; Karlov, G.I. Study and Physical Mapping of the Species-Specific Tandem Repeat CS-237 Linked with 45S Ribosomal DNA Intergenic Spacer in Cannabis sativa L. Plants 2022, 11, 1396. https://doi.org/10.3390/plants11111396
Alexandrov OS, Romanov DV, Divashuk MG, Razumova OV, Ulyanov DS, Karlov GI. Study and Physical Mapping of the Species-Specific Tandem Repeat CS-237 Linked with 45S Ribosomal DNA Intergenic Spacer in Cannabis sativa L. Plants. 2022; 11(11):1396. https://doi.org/10.3390/plants11111396
Chicago/Turabian StyleAlexandrov, Oleg S., Dmitry V. Romanov, Mikhail G. Divashuk, Olga V. Razumova, Daniil S. Ulyanov, and Gennady I. Karlov. 2022. "Study and Physical Mapping of the Species-Specific Tandem Repeat CS-237 Linked with 45S Ribosomal DNA Intergenic Spacer in Cannabis sativa L." Plants 11, no. 11: 1396. https://doi.org/10.3390/plants11111396
APA StyleAlexandrov, O. S., Romanov, D. V., Divashuk, M. G., Razumova, O. V., Ulyanov, D. S., & Karlov, G. I. (2022). Study and Physical Mapping of the Species-Specific Tandem Repeat CS-237 Linked with 45S Ribosomal DNA Intergenic Spacer in Cannabis sativa L. Plants, 11(11), 1396. https://doi.org/10.3390/plants11111396






