Dosage-Dependent Gynoecium Development and Gene Expression in Brassica napus-Orychophragmus violaceus Addition Lines
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
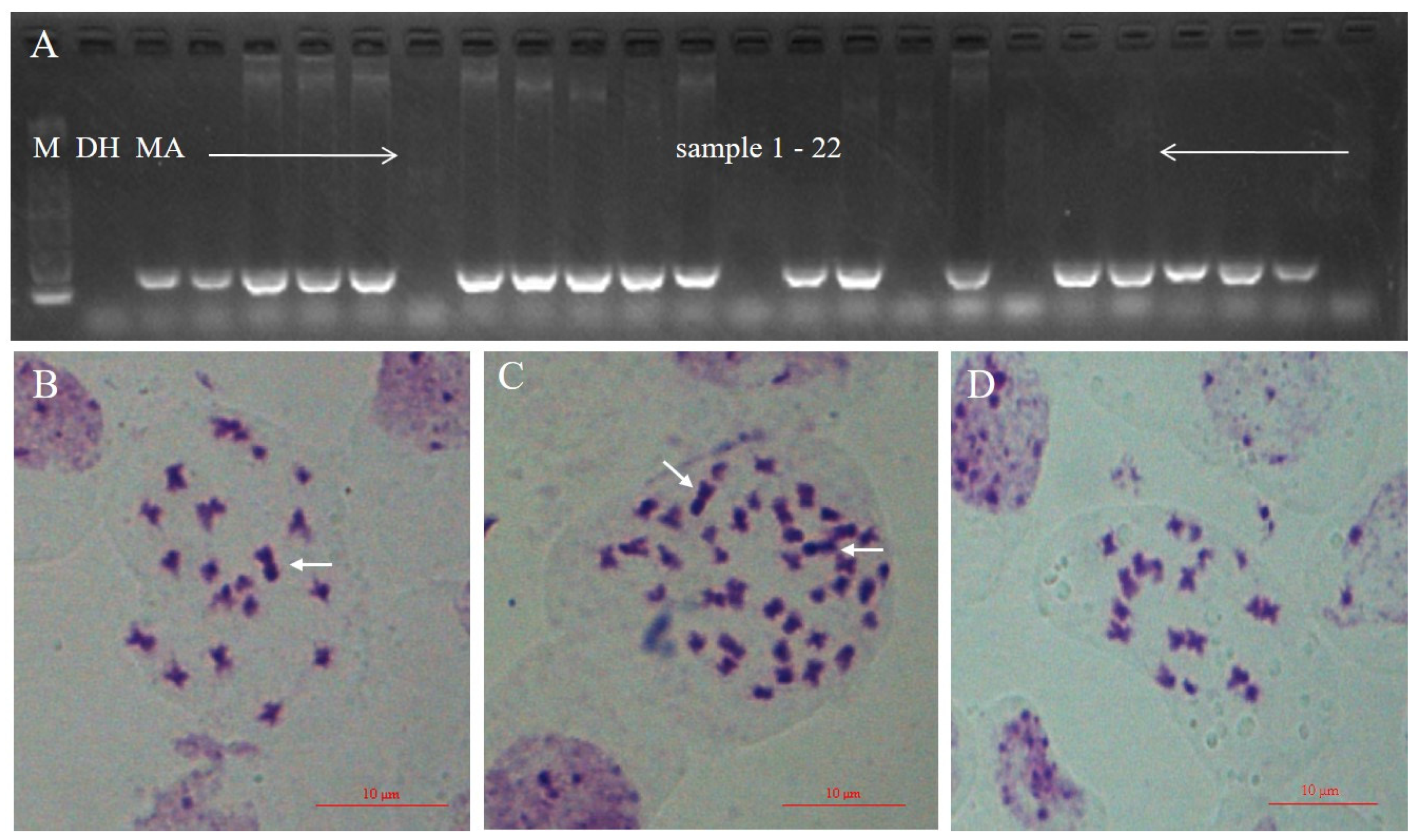
2.2. Cytological Observation and GISH Analysis
2.3. DNA Extraction and Probe Labelling
2.4. RNA Extraction, cDNA Libraries Preparation, and Sequencing
2.5. Full-Length Transcriptome Analysis
2.6. Reads Mapping and Differentially Expressed Genes (DEGs) Analysis
2.7. Quantitative Real-Time PCR (qRT-PCR) Analysis
3. Results
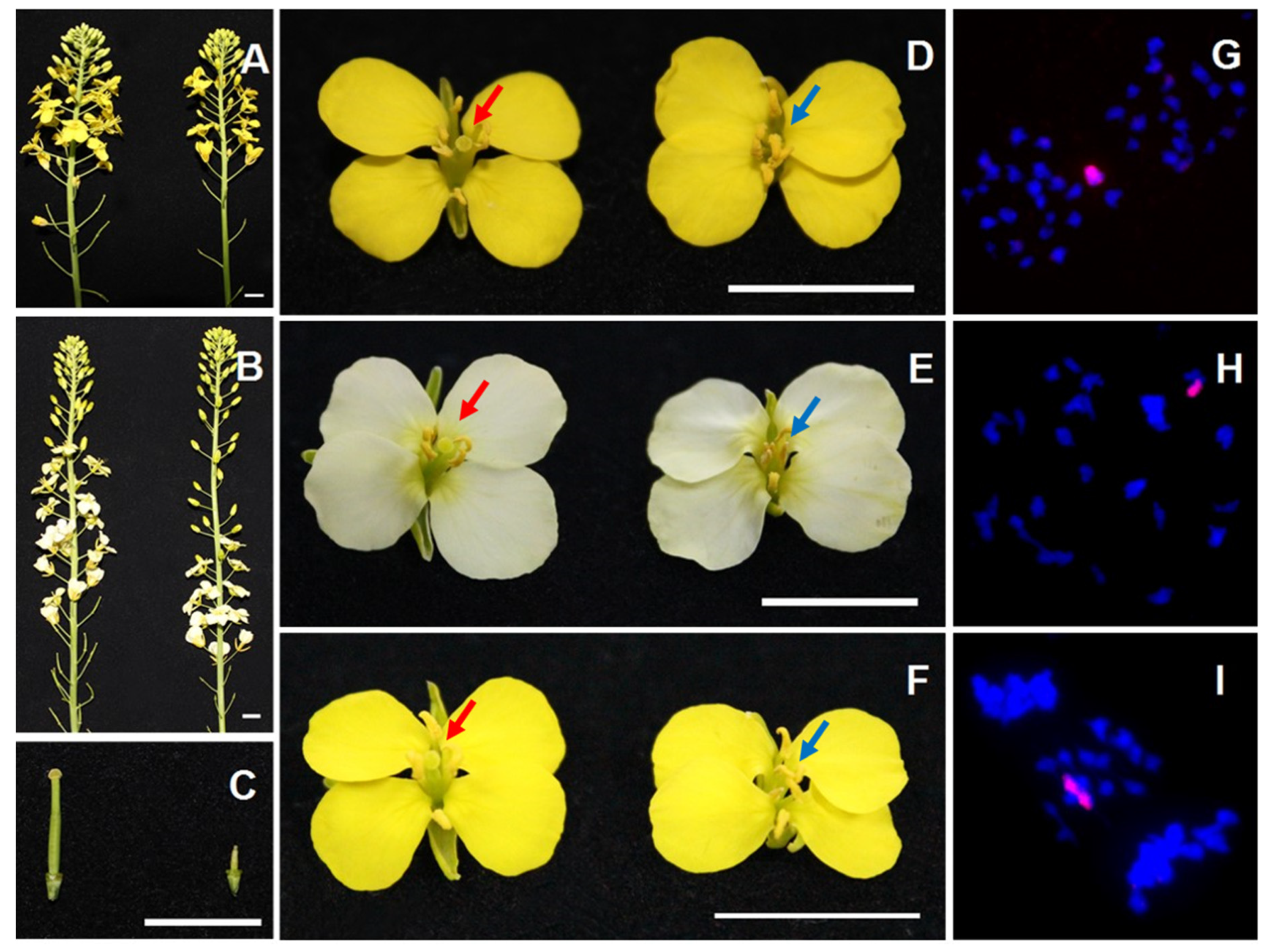
3.1. Complete Female Sterility in Different Brassica Species Caused by Single O. violaceus Chromosome
3.2. Dosage Effect of Alien Chromosome on Gynoecium Development
3.3. RNA-seq Analysis of O. violaceus and Additions
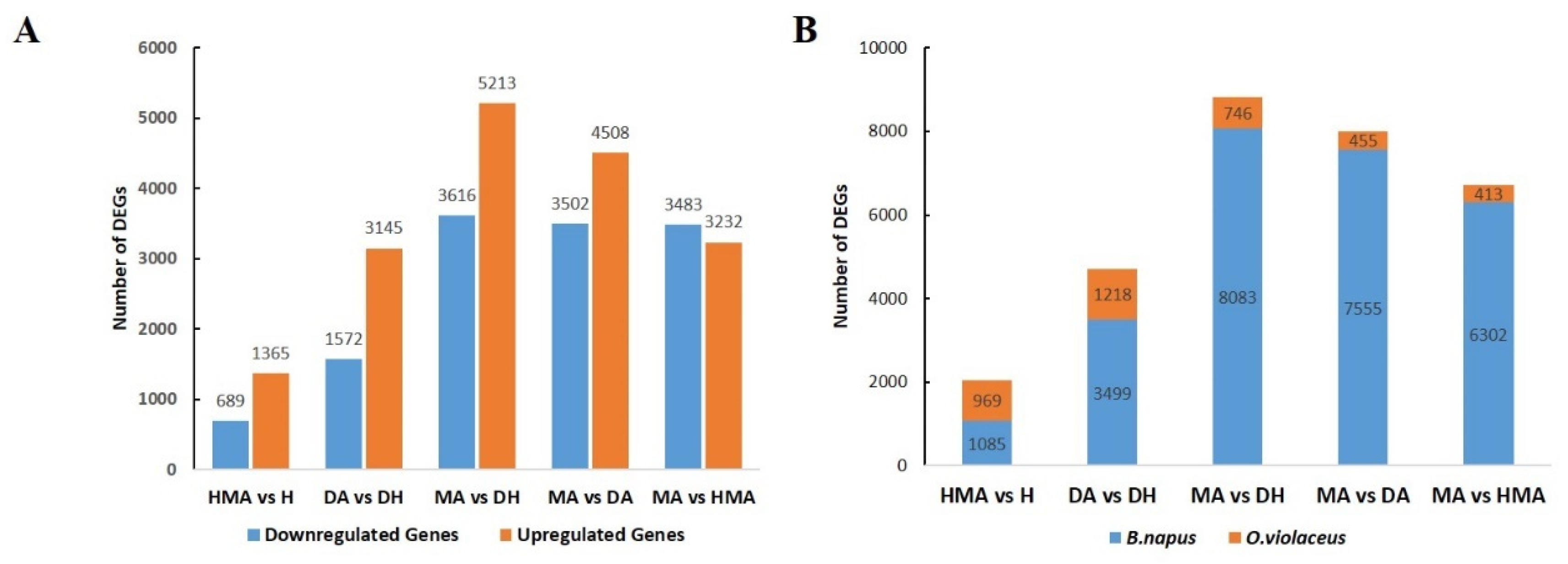
3.4. Gene Expression Changes in Additions Induced by the O. violaceus Chromosome
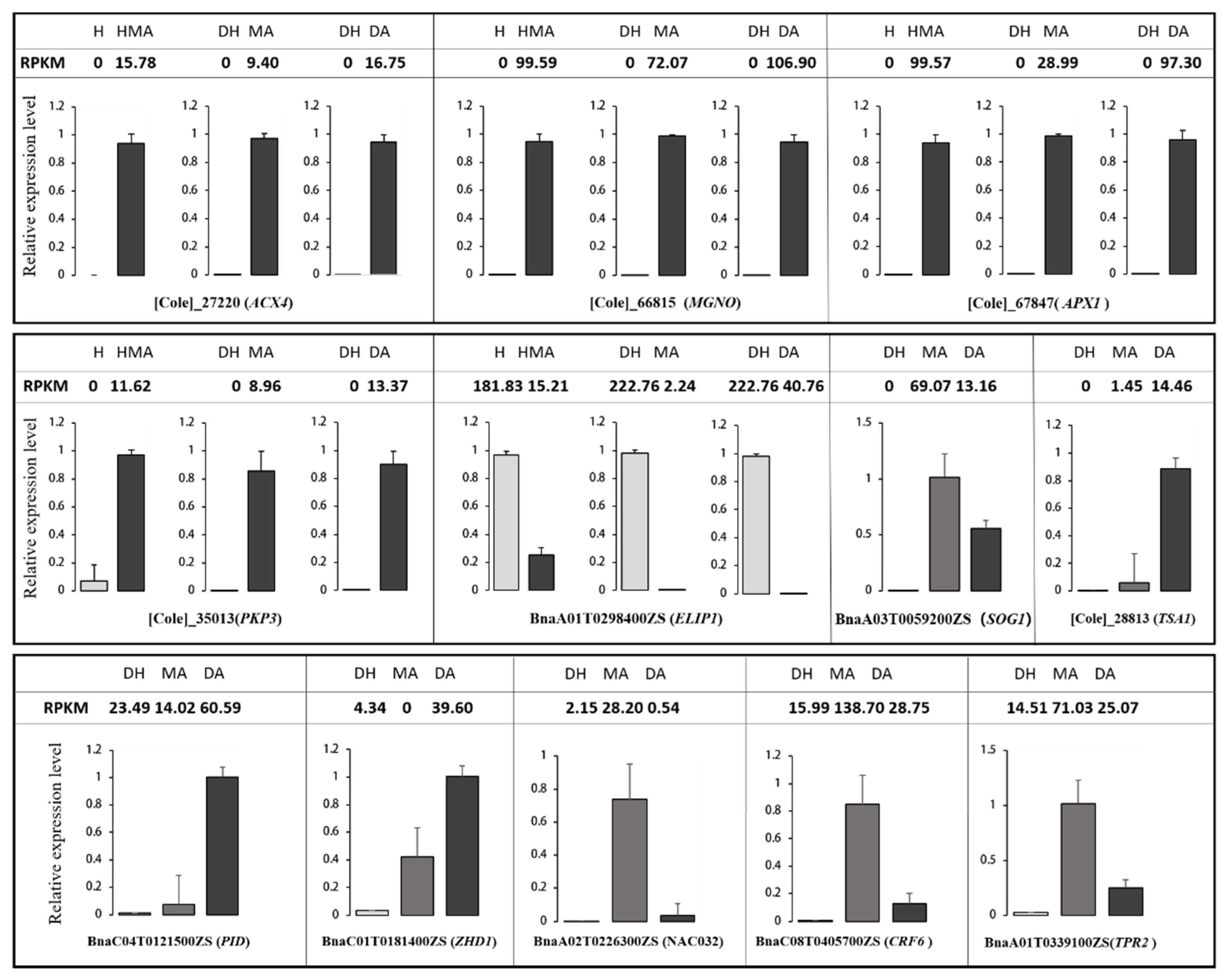
3.5. DEGs Related to the Pistil Development
3.6. Down-Regulation Genes for Brassinosteroid Biosynthesis in MA
4. Discussion
4.1. Genetic Effects of Alien Chromosomes in Different Brassica Species and Dosage Effect of Alien Chromosome on Gynoecium Development
4.2. Global Gene Expression Changes upon the Alien Chromosome and Genome Ploidy
4.3. Gene Activation and Silencing Related to Female Sterility in Additions
4.4. Dosage-Dependent Gene Expression for BR Biosynthesis and Pistil Growth
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Scutt, C.P.; Vinauger-Douard, M.; Fourquin, C.; Finet, C.; Dumas, C. An evolutionary perspective on the regulation of carpel development. J. Exp. Bot. 2006, 57, 2143–2152. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smyth, D.R.; Bowman, J.L.; Meyerowitz, E.M. Early flower development in Arabidopsis. Plant Cell 1990, 2, 755–767. [Google Scholar] [PubMed] [Green Version]
- Alvarez-Buylla, A.; Elena, R.; Cador, C.; Folter, D.; Alvarez-Buylla, E.R.; Benítez, M.; Corvera-Poiré, A.; Cador, C.; De Folter, S. Flower development. In The Arabidopsis Book; American Society of Plant Biologists: Rockville, MD, USA, 2010; Volume e0127. [Google Scholar]
- Reyes-Olalde, J.I.; de Folter, S. Control of stem cell activity in the carpel margin meristem (CMM) in Arabidopsis. Plant Reprod. 2019, 32, 123–136. [Google Scholar] [CrossRef]
- Schneitz, K.; Hülskamp, M.; Pruitt, R.E. Wild-type ovule development in Arabidopsis thaliana: A light microscope study of cleared whole-mount tissue. Plant J. 1995, 7, 731–749. [Google Scholar] [CrossRef]
- Drews, G.N.; Koltunow, A.M. The female gametophyte. In The Arabidopsis Book; American Society of Plant Biologists: Rockville, MD, USA, 2011. [Google Scholar] [CrossRef]
- Zúñiga-Mayo, V.M.; Gómez-Felipe, A.; Herrera-Ubaldo, H.; De Folter, S. Gynoecium development: Networks in Arabidopsis and beyond. J. Exp. Bot. 2019, 70, 1447–1460. [Google Scholar] [CrossRef]
- Cucinotta, M.; Di Marzo, M.; Guazzotti, A.; de Folter, S.; Kater, M.M.; Colombo, L. Gynoecium size and ovule number are interconnected traits that impact seed yield. J. Exp. Bot. 2020, 71, 2479–2489. [Google Scholar] [CrossRef]
- Ouyang, Y.D.; Chen, J.J.; Ding, J.H.; Zhang, Q.F. Advances in the understanding of inter-subspecific hybrid sterility and wide-compatibility in rice. Chin. Sci. Bull. 2009, 54, 2332–2341. [Google Scholar] [CrossRef]
- Schuster, S.C. Next-generation sequencing transforms today’s biology. Nat. Methods 2008, 5, 16–18. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef]
- Yue, L.; Twell, D.; Kuang, Y.; Liao, J.; Zhou, X. Transcriptome analysis of Hamelia patens (Rubiaceae) anthers reveals candidate genes for tapetum and pollen wall development. Front. Plant Sci. 2017, 7, 1991. [Google Scholar] [CrossRef] [Green Version]
- Marie, K.L.; Sassano, A.; Yang, H.H.; Michalowski, A.M.; Michael, H.T.; Guo, T.; Tsai, Y.C.; Weissman, A.M.; Lee, M.P.; Jenkins, L.M.; et al. Melanoblast transcriptome analysis reveals pathways promoting melanoma metastasis. Nat. Commun. 2020, 11, 333. [Google Scholar] [CrossRef] [Green Version]
- Schmidt, A.; Wuest, S.E.; Vijverberg, K.; Baroux, C.; Kleen, D.; Grossniklaus, U. Transcriptome analysis of the arabidopsis megaspore mother cell uncovers the importance of RNA helicases for plant germline development. PLoS Biol. 2011, 9, e1001155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, L.; Wu, Y.; Yu, M.; Mao, B.; Zhao, B.; Wang, J. Genome-wide transcriptome analysis of female-sterile rice ovule shed light on its abortive mechanism. Planta 2016, 244, 1011–1028. [Google Scholar] [CrossRef] [PubMed]
- Shi, T.; Gao, Z.; Wang, L.; Zhang, Z.; Zhuang, W.; Sun, H.; Zhong, W. Identification of Differentially-Expressed Genes Associated with Pistil Abortion in Japanese Apricot by Genome-Wide Transcriptional Analysis. PLoS ONE 2012, 7, e47810. [Google Scholar] [CrossRef] [Green Version]
- Huang, S.; Liu, Z.; Li, C.; Yao, R.; Li, D.; Hou, L.; Li, X.; Liu, W.; Feng, H. Transcriptome analysis of a female-sterile mutant (Fsm) in Chinese cabbage (Brassica campestris ssp. pekinensis). Front. Plant Sci. 2017, 8, 546. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fu, W.; Zhao, Z.; Ge, X.; Ding, L.; Li, Z. Anatomy and transcript profiling of gynoecium development in female sterile Brassica napus mediated by one alien chromosome from Orychophragmus violaceus. BMC Genom. 2014, 15, 61. [Google Scholar] [CrossRef] [Green Version]
- Teng, C.; Du, D.; Xiao, L.; Yu, Q.; Shang, G.; Zhao, Z. Mapping and identifying a candidate gene (BNMFS) for female-male sterility through whole-genome resequencing and rna-seq in rapeseed (brassica napus L.). Front. Plant Sci. 2017, 8, 2086. [Google Scholar] [CrossRef]
- Alagna, F.; Cirilli, M.; Galla, G.; Carbone, F.; Daddiego, L.; Facella, P.; Lopez, L.; Colao, C.; Mariotti, R.; Cultrera, N.; et al. Transcript analysis and regulative events during flower development in olive (Olea europaea L.). PLoS ONE 2016, 11, e0152943. [Google Scholar] [CrossRef] [Green Version]
- Zhang, S.; Shi, Q.; Albrecht, U.; Shatters, R.G.; Stange, R.; McCollum, G.; Zhang, S.; Fan, C.; Stover, E. Comparative transcriptome analysis during early fruit development between three seedy citrus genotypes and their seedless mutants. Hortic. Res. 2017, 4, 17041. [Google Scholar] [CrossRef]
- Bräuning, S.; Catanach, A.; Lord, J.M.; Bicknell, R.; Macknight, R.C. Comparative transcriptome analysis of the wild-type model apomict Hieracium praealtum and its loss of parthenogenesis (lop) mutant. BMC Plant Biol. 2018, 18, 206. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Robinson-Beers, K.; Pruitt, R.E.; Gasser, C.S. Ovule development in wild-type Arabidopsis and two female-sterile mutants. Plant Cell 1992, 4, 1237–1249. [Google Scholar] [CrossRef]
- Shi, D.Q.; Yang, W.C. Ovule development in Arabidopsis: Progress and challenge. Curr. Opin. Plant Biol. 2011, 14, 74–80. [Google Scholar] [CrossRef]
- Figueiredo, D.D.; Köhler, C. Bridging the generation gap: Communication between maternal sporophyte, female gametophyte and fertilization products. Curr. Opin. Plant Biol. 2016, 29, 16–20. [Google Scholar] [CrossRef]
- Raymer, P.L. Canola: An emerging oilseed crop. Trends New Crop. New Uses 2002, 1, 122–126. [Google Scholar]
- Cartea, M.E.; Lema, M.; Francisco, M.; Velasco, P. Basic information on vegetable Brassica crops. Genet. Genom. Breed. Veg. Brassicas 2011, 1–33. [Google Scholar]
- Xin, Q.; Shen, Y.; Li, X.; Lu, W.; Wang, X.; Han, X.; Dong, F.; Wan, L.; Yang, G.; Hong, D.; et al. MS5 mediates early meiotic progression and its natural variants may have applications for hybrid production in Brassica napus. Plant Cell 2016, 28, 1263–1278. [Google Scholar] [CrossRef] [Green Version]
- Li, C.H.; Fu, S.X.; Chen, X.J.; Qi, C.K. Phenotypic characterization and genetic analysis of a partially female-sterile mutant in Brassica napus. Plant Sci. 2012, 185, 112–117. [Google Scholar] [CrossRef]
- Ding, L.; Zhao, Z.G.; Ge, X.H.; Li, Z.Y. Intergeneric addition and substitution of Brassica napus with different chromosomes from Orychophragmus violaceus: Phenotype and cytology. Sci. Hortic. 2013, 164, 303–309. [Google Scholar] [CrossRef]
- Zhao, Z.G.; Hu, T.T.; Ge, X.H.; Du, X.Z.; Ding, L.; Li, Z.Y. Production and characterization of intergeneric somatic hybrids between Brassica napus and Orychophragmus violaceus and their backcrossing progenies. Plant Cell Rep. 2008, 27, 1611–1621. [Google Scholar] [CrossRef]
- Ge, X.H.; Wang, J.; Li, Z.Y. Different genome-specific chromosome stabilities in synthetic Brassica allohexaploids revealed by wide crosses with Orychophragmus. Ann. Bot. 2009, 104, 19–31. [Google Scholar] [CrossRef] [Green Version]
- Cui, C.; Ge, X.; Gautam, M.; Kang, L.; Li, Z. Cytoplasmic and genomic effects on meiotic pairing in Brassica hybrids and allotetraploids from pair crosses of three cultivated diploids. Genetics 2012, 191, 725–738. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.P.; Liu, Z.W.; Tu, J.X.; Chen, B.Y.; Fu, T.D. Improvement of Microspores Culture Techniques in Brassica napus. L (In Chinese with English abstract). Yi Chuan 2003, 25, 4–7. [Google Scholar]
- Li, Z.; Liu, H.L.; Luo, P. Production and cytogenetics of intergeneric hybrids between Brassica napus and Orychophragmus violaceus. Theor. Appl. Genet. 1995, 91, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Zhong, X.B.; De Jong, J.H.; Zabel, P. Preparation of tomato meiotic pachytene and mitotic metaphase chromosomes suitable for fluorescence in situ hybridization (FISH). Chromosome Res. 1996, 4, 24–28. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.H.; Li, Z.Y. Intra- and intergenomic homology of B-genome chromosomes in trigenomic combinations of the cultivated Brassica species revealed by GISH analysis. Chromosome Res. 2007, 15, 849–861. [Google Scholar] [CrossRef]
- Leitch, A.R.; Schwarzacher, T.; Jackson, D.; Leitch, I.J. Microscopy Handbook No. 27. In Situ Hybridization: A Practical Guide; BIOS Scientific: Abingdon, UK, 1994. [Google Scholar]
- Dellaporta, S.L.; Wood, J.; Hicks, J.B. A plant DNA minipreparation: Version II. Plant Mol. Biol. Report. 1983, 1, 19–21. [Google Scholar] [CrossRef]
- Song, J.M.; Guan, Z.; Hu, J.; Guo, C.; Yang, Z.; Wang, S.; Liu, D.; Wang, B.; Lu, S.; Zhou, R.; et al. Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat. Plants 2020, 6, 34–45. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Coate, J.E.; Doyle, J.J. Quantifying whole transcriptome size, a prerequisite for understanding transcriptome evolution across species: An example from a plant allopolyploid. Genome Biol. Evol. 2010, 2, 534–546. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S.; Sato, Y.; Furumichi, M.; Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 2012, 40, D109–D114. [Google Scholar] [CrossRef] [Green Version]
- Yang, H.; Liu, J.; Huang, S.; Guo, T.; Deng, L.; Hua, W. Selection and evaluation of novel reference genes for quantitative reverse transcription PCR (qRT-PCR) based on genome and transcriptome data in Brassica napus L. Gene 2014, 538, 113–122. [Google Scholar] [CrossRef]
- Miller, T.E.; Hutchinson, J.; Chapman, V. Investigation of a preferentially transmitted Aegilops sharonensis chromosome in wheat. Theor. Appl. Genet. 1982, 61, 27–33. [Google Scholar] [CrossRef]
- Wang, Y.; Sonntag, K.; Rudloff, E.; Wehling, P.; Snowdon, R.J. GISH analysis of disomic Brassica napus-Crambe abyssinica chromosome addition lines produced by microspore culture from monosomic addition lines. Plant Cell Rep. 2006, 25, 35–40. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, A.; Lu, L.; Chen, L.; Hu, F.; Wang, Y. Disomic Alien Addition Lines Generated through Microspore Culture from Progeny of Intergeneric Hybrids between Brassica napus and Sinapis alba (In Chinese with English abstract). Acta Agron. Sin. 2006, 32, 1764–1766. [Google Scholar]
- Shi, X.; Ng, D.W.K.; Zhang, C.; Comai, L.; Ye, W.; Chen, Z.J. Cis- and trans-regulatory divergence between progenitor species determines gene-expression novelty in Arabidopsis allopolyploids. Nat. Commun. 2012, 3, 950. [Google Scholar] [CrossRef] [PubMed]
- Birchler, J.A.; Veitia, R.A. The gene balance hypothesis: From classical genetics to modern genomics. Plant Cell 2007, 19, 395–402. [Google Scholar] [CrossRef] [Green Version]
- Huettel, B.; Kreil, D.P.; Matzke, M.; Matzke, A.J.M. Effects of aneuploidy on genome structure, expression, and interphase organization in Arabidopsis thaliana. PLoS Genet. 2008, 4, e1000226. [Google Scholar] [CrossRef] [Green Version]
- Bryant, N.; Lloyd, J.; Sweeney, C.; Myouga, F.; Meinke, D. Identification of nuclear genes encoding chloroplast-localized proteins required for embryo development in Arabidopsis. Plant. Physiol. 2011, 155, 1678–1689. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sørmo, C.G.; Brembu, T.; Winge, P.; Bones, A.M. Arabidopsis thaliana MIRO1 and MIRO2 GTPases are unequally redundant in pollen tube growth and fusion of polar nuclei during female gametogenesis. PLoS ONE 2011, 6, e18530. [Google Scholar] [CrossRef] [Green Version]
- Paolo, D.; Rotasperti, L.; Schnittger, A.; Masiero, S.; Colombo, L.; Mizzotti, C. The arabidopsis mads-domain transcription factor seedstick controls seed size via direct activation of e2fa. Plants 2021, 10, 192. [Google Scholar] [CrossRef] [PubMed]
- Nemhauser, J.L.; Feldman, L.J.; Zambryski, P.C. Auxin and ETTIN in Arabidopsis gynoecium morphogenesis. Development 2000, 127, 3877–3888. [Google Scholar] [CrossRef]
- Wu, M.F.; Tian, Q.; Reed, J.W. Arabidopis microRNA 167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar] [CrossRef] [Green Version]
- Bencivenga, S.; Simonini, S.; Colombo, L. The Transcription Factors BEL1 and SPL Are Required for Cytokinin and Auxin Signaling During Ovule Development in Arabidopsis. Plant Cell 2012, 24, 2886–2897. [Google Scholar] [CrossRef] [Green Version]
- Ceccato, L.; Masiero, S.; Sinha Roy, D.; Bencivenga, S.; Roig-Villanova, I.; Ditengou, F.A.; Palme, K.; Simon, R.; Colombo, L. Maternal Control of PIN1 Is Required for Female Gametophyte Development in Arabidopsis. PLoS ONE 2013, 8, e66148. [Google Scholar] [CrossRef] [Green Version]
- Panoli, A.; Martin, M.V.; Alandete-Saez, M.; Simon, M.; Neff, C.; Swarup, R.; Bellido, A.; Yuan, L.; Pagnussat, G.C.; Sundaresan, V. Auxin import and local auxin biosynthesis are required for mitotic divisions, cell expansion and cell specification during female gametophyte development in Arabidopsis thaliana. PLoS ONE 2015, 10, e0126164. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Guo, X.; Xiao, Q.; Zhu, J.; Cheung, A.Y.; Yuan, L.; Vierling, E.; Xu, S. Auxin efflux controls orderly nucellar degeneration and expansion of the female gametophyte in Arabidopsis. New Phytol. 2021, 230, 2261–2274. [Google Scholar] [CrossRef]
- Takahashi, N.; Ogita, N.; Takahashi, T.; Taniguchi, S.; Tanaka, M.; Seki, M.; Umeda, M. A regulatory module controlling stress-induced cell cycle arrest in arabidopsis. eLife 2019, 8, e43944. [Google Scholar] [CrossRef] [PubMed]
- Berg, M.; Rogers, R.; Muralla, R.; Meinke, D. Requirement of aminoacyl-tRNA synthetases for gametogenesis and embryo development in Arabidopsis. Plant J. 2005, 44, 866–878. [Google Scholar] [CrossRef]
- Ishikawa, T.; Machida, C.; Yoshioka, Y.; Ueda, T.; Nakano, A.; Machida, Y. EMBRYO YELLOW gene, encoding a subunit of the conserved oligomeric Golgi complex, is required for appropriate cell expansion and meristem organization in Arabidopsis thaliana. Genes Cells 2008, 13, 521–535. [Google Scholar] [CrossRef]
- Schneitz, K.; Hülskamp, M.; Kopczak, S.D.; Pruitt, R.E. Dissection of sexual organ ontogenesis: A genetic analysis of ovule development in Arabidopsis thaliana. Development 1997, 124, 1367–1376. [Google Scholar] [CrossRef]
- Villanueva, J.M.; Broadhvest, J.; Hauser, B.A.; Meister, R.J.; Schneitz, K.; Gasser, C.S. INNER NO OUTER regulates abaxial-adaxial patterning in Arabidopsis ovules. Genes Dev. 1999, 13, 3160–3169. [Google Scholar] [CrossRef] [Green Version]
- Sieber, P.; Gheyselinck, J.; Gross-Hardt, R.; Laux, T.; Grossniklaus, U.; Schneitz, K. Pattern formation during early ovule development in Arabidopsis thaliana. Dev. Biol. 2004, 273, 321–334. [Google Scholar] [CrossRef] [Green Version]
- Vijayan, A.; Tofanelli, R.; Strauss, S.; Cerrone, L.; Wolny, A.; Strohmeier, J.; Kreshuk, A.; Hamprecht, F.A.; Smith, R.S.; Schneitz, K. A digital 3D reference atlas reveals cellular growth patterns shaping the arabidopsis ovule. eLife 2021, 10, e63262. [Google Scholar] [CrossRef] [PubMed]
- Jia, D.; Chen, L.; Yin, G.; Yang, X.; Gao, Z.; Guo, Y.; Sun, Y.; Tang, W. Brassinosteroids regulate outer ovule integument growth in part via the control of INNER NO OUTER by BRASSINOZOLE-RESISTANT family transcription factors. J. Integr. Plant Biol. 2020, 62, 1093–1111. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cai, B.; Wang, T.; Fu, W.; Harun, A.; Ge, X.; Li, Z. Dosage-Dependent Gynoecium Development and Gene Expression in Brassica napus-Orychophragmus violaceus Addition Lines. Plants 2021, 10, 1766. https://doi.org/10.3390/plants10091766
Cai B, Wang T, Fu W, Harun A, Ge X, Li Z. Dosage-Dependent Gynoecium Development and Gene Expression in Brassica napus-Orychophragmus violaceus Addition Lines. Plants. 2021; 10(9):1766. https://doi.org/10.3390/plants10091766
Chicago/Turabian StyleCai, Bowei, Tai Wang, Wenqin Fu, Arrashid Harun, Xianhong Ge, and Zaiyun Li. 2021. "Dosage-Dependent Gynoecium Development and Gene Expression in Brassica napus-Orychophragmus violaceus Addition Lines" Plants 10, no. 9: 1766. https://doi.org/10.3390/plants10091766
APA StyleCai, B., Wang, T., Fu, W., Harun, A., Ge, X., & Li, Z. (2021). Dosage-Dependent Gynoecium Development and Gene Expression in Brassica napus-Orychophragmus violaceus Addition Lines. Plants, 10(9), 1766. https://doi.org/10.3390/plants10091766






