Diversity and Relationships among Neglected Apricot (Prunus armeniaca L.) Landraces Using Morphological Traits and SSR Markers: Implications for Agro-Biodiversity Conservation
Abstract
:1. Introduction
2. Results
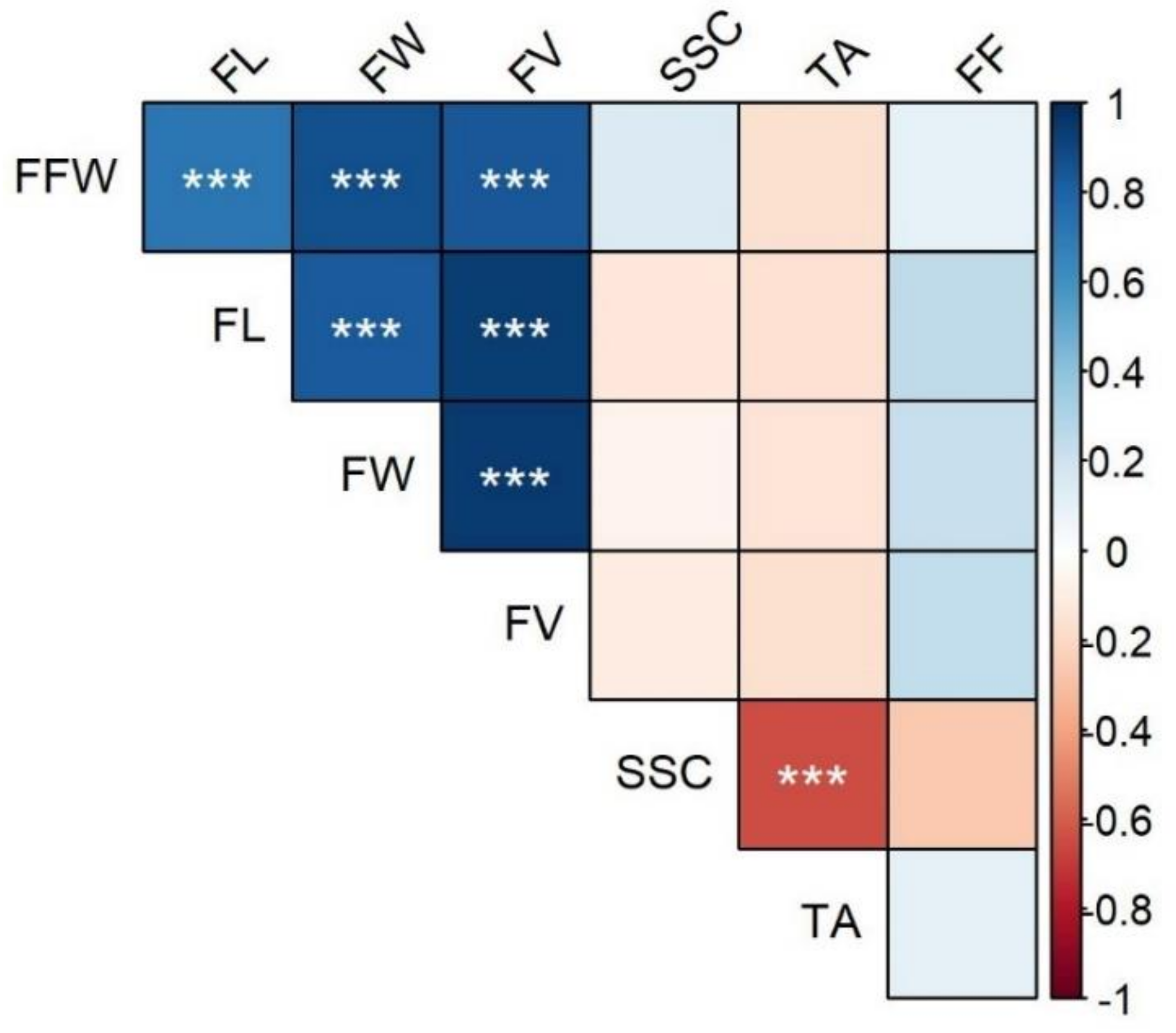
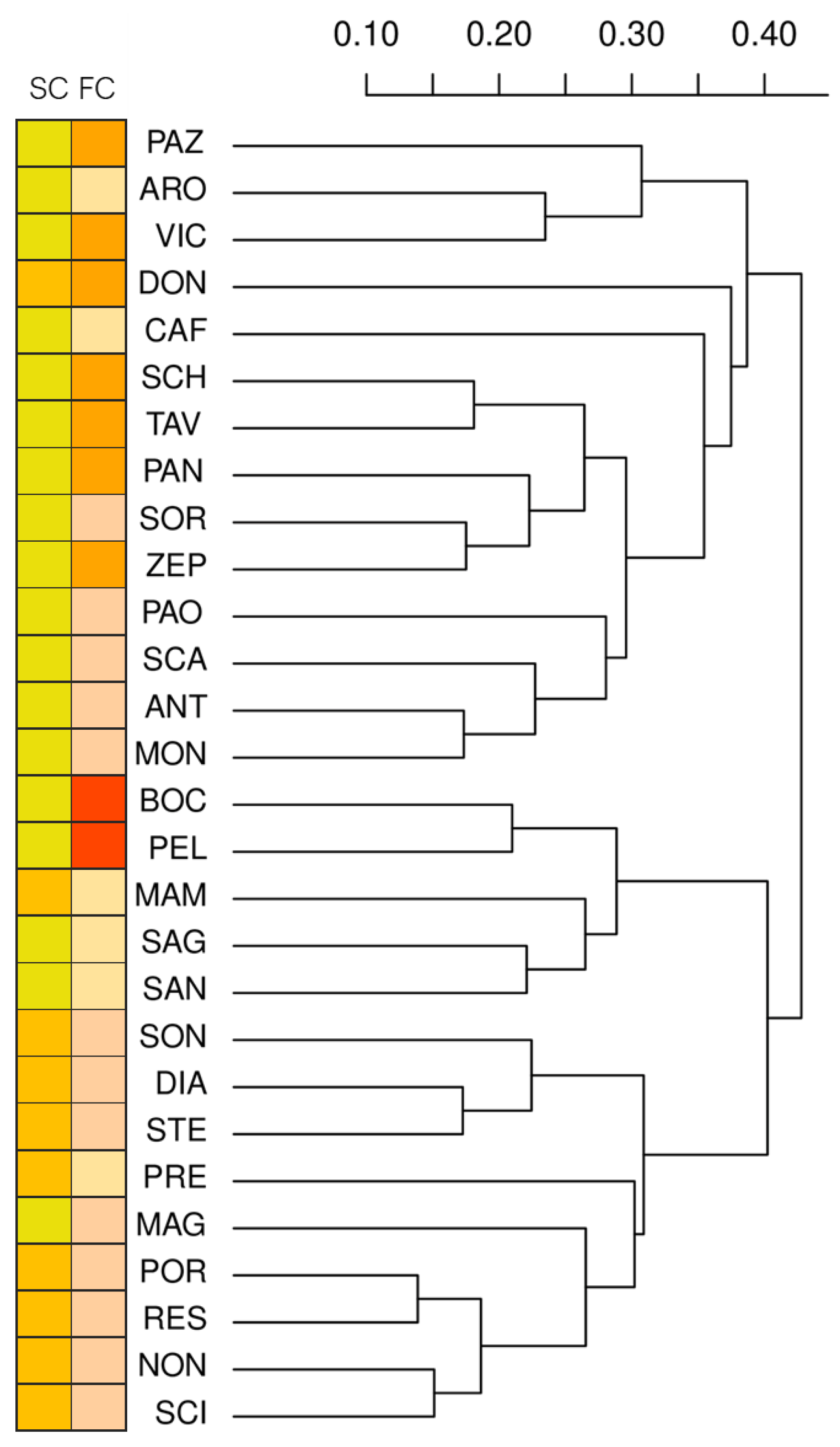
2.1. Morphological Analysis
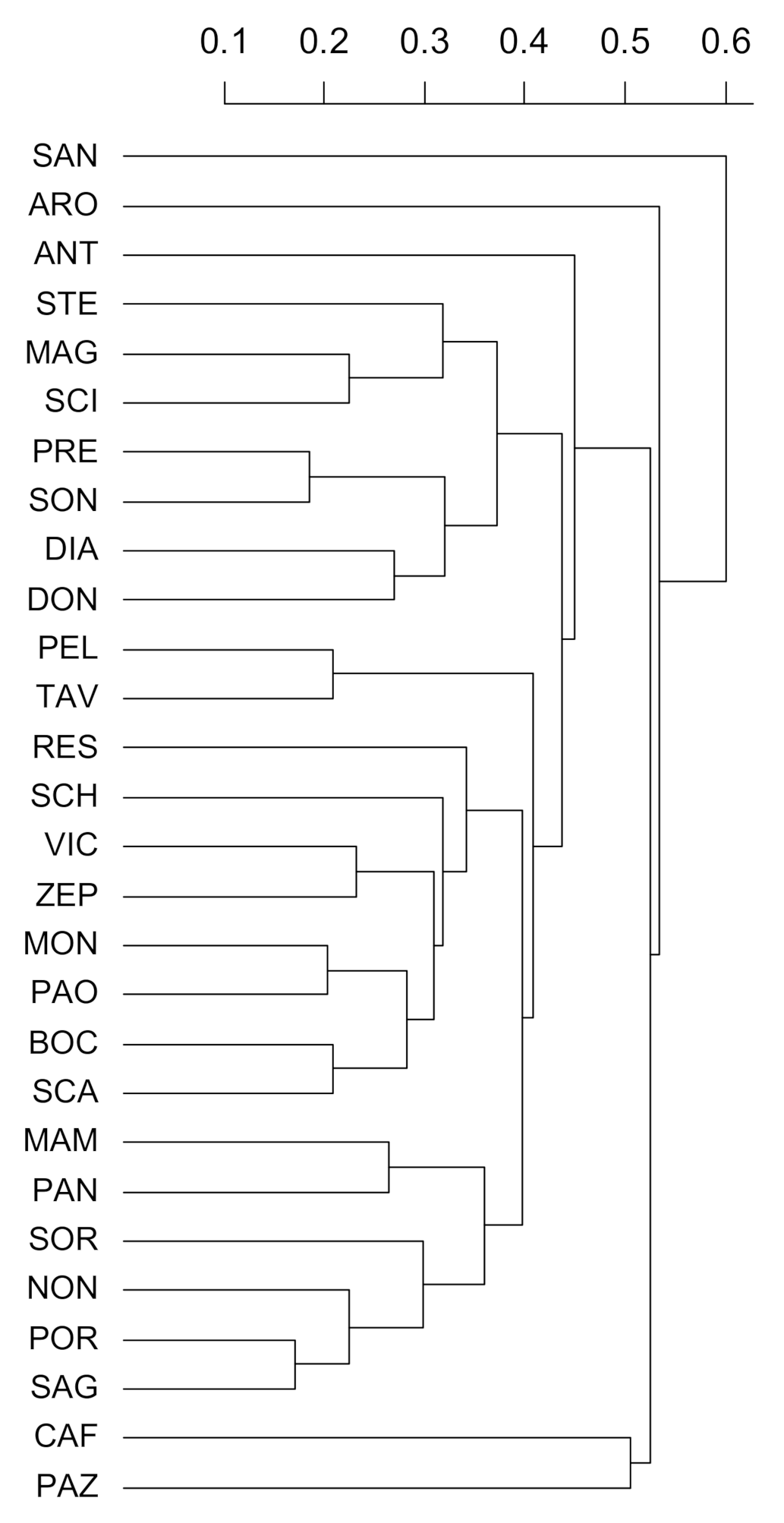
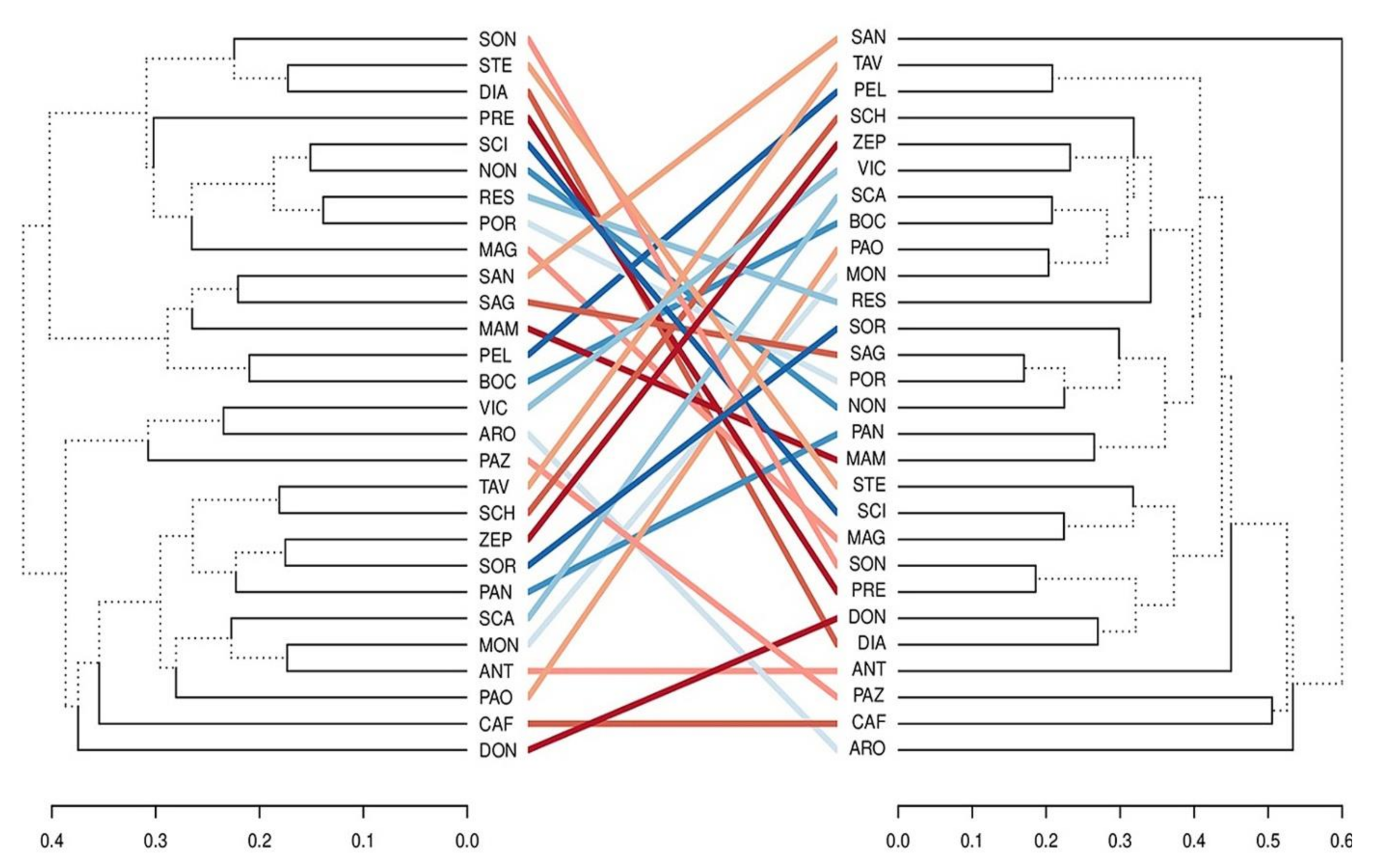
2.2. Analysis of Genetic Diversity by SSR Markers
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Analysis of Morphological Data
4.3. DNA Isolation and SSR Analysis
4.4. Molecular Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sinha, N.; Sidhu, J.; Barta, J.; Wu, J.; Cano, M.P. Handbook of Fruits and Fruit Processing; John Wiley & Sons: Hoboken, NJ, USA, 2012. [Google Scholar]
- Faust, M.; Suranyi, D.; Nyujto, F. Origin and dissemination of apricot. Hortic. Rev. Westport N. Y. 1998, 22, 225–260. [Google Scholar]
- Yu, Y.; Fu, J.; Xu, Y.; Zhang, J.; Ren, F.; Zhao, H.; Tian, S.; Guo, W.; Tu, X.; Zhao, J. Genome re-sequencing reveals the evolutionary history of peach fruit edibility. Nat. Commun. 2018, 9, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ledbetter, C.A. Apricots. In Temperate Fruit Crop Breeding: Germplasm to Genomics; Hancock, J.F., Ed.; Springer: Dordrecht, The Netherlands, 2008; pp. 39–82. [Google Scholar] [CrossRef]
- FAO. Available online: http://www.fao.org/ (accessed on 1 March 2021).
- Istat. Available online: http://dati.istat.it/ (accessed on 1 March 2021).
- Pugliano, G.; Forlani, M. Two-year observations on the biology and fructification of apricot. Acta Hortic. 2019, 192, 383–400. [Google Scholar]
- Jashemski, W.F. Ancient Roman gardens in Campania and Tunisia: A comparison of the evidence. J. Gard. His. 1996, 16, 231–243. [Google Scholar] [CrossRef]
- Basile, B.; Cirillo, C.; Santin, A.; Forlani, M. Fruit quality of Vesuvian apricots harvested at different ripening stages after a cold-storage period. In Proceedings of the V International Postharvest Symposium, Verona, Italy, 6–11 June 2005; Volume 682, pp. 1443–1450. [Google Scholar]
- Cirillo, C.; Basile, B.; Hernandez, G.; Pannico, A.; Giaccone, M.; Forlani, M. Influence of fruiting shoot on flowering pattern and fruit quality of Vesuvian apricot cultivars. Acta Hortic. 2010, 862, 557–564. [Google Scholar] [CrossRef]
- Mennone, C. Effect of climate on apricot productivity. Informatore Agrario 2016, 72, 47–50. [Google Scholar]
- Campoy, J.A.; Audergon, J.M.; Ruiz, D.; Martínez-Gómez, P. Genomic designing for new climate-resilient apricot varieties. In Genomic Designing of Climate-Smart Fruit Crops; Kole, C., Ed.; Springer: Cham, Switzerland, 2019; pp. 73–89. [Google Scholar]
- Pennone, F.; Abbate, V. Prospettive per la valorizzazione della coltura dell’albicocco nel Mezzogiorno attraverso il miglioramento genetico. Rivista di Frutticoltura e di Ortofloricoltura 2004, 66, 36–39. [Google Scholar]
- Campoy, J.A.; Ruiz, D.; Egea, J. Dormancy in temperate fruit trees in a global warming context: A review. Sci. Hortic. 2011, 130, 357–372. [Google Scholar] [CrossRef]
- Bartolini, S.; Massai, R.; Viti, R. The influence of autumn-winter temperatures on endodormancy release and blooming performance of apricot (Prunus armeniaca L.) in central Italy based on long-term observations. J. Hortic. Sci. Biotechnol. 2020, 95, 794–803. [Google Scholar] [CrossRef]
- Cirillo, O. Fashion and tourism in Campania in the middle of the twentieth century: A story with many protagonists. Almatour. J. Tour. Cult. Territ. Dev. 2018, 9, 23–46. [Google Scholar]
- Mainolfi, A.; Abbate, V.; Buccheri, M.; Damiano, C. Molecular characterization of local ecotypes and commercial varieties of apricot (Prunus armeniaca L.) [Campania]. Italus Hortus 2006, 13, 262–265. [Google Scholar]
- Enjolras, G.; Aubert, M. Short food supply chains and the issue of sustainability: A case study of French fruit producers. Int. J. Retail Distrib. Manag. 2018, 46, 194–209. [Google Scholar] [CrossRef] [Green Version]
- Rao, R.; Bencivenni, M.; Mura, L.; Araujo-Burgos, T.; Corrado, G. Molecular characterisation of Vesuvian apricot cultivars: Implications for the certification and authentication of protected plant material. J. Hortic. Sci. Biotechnol. 2010, 85, 42–347. [Google Scholar] [CrossRef]
- Rao, R.; Bencivenni, M.; Corrado, G.; Basile, B.; Forlani, M. Molecular characterization of apricot varieties included in the “Albicocca Vesuviana” PGI Regulation. Acta Hortic. 2010, 862, 61–66. [Google Scholar] [CrossRef]
- Krishna, V.V.; Pascual, U. Can greening markets help conserve landraces in situ? In Agrobiodiversity Conservation and Economic Development; Routledge: London, UK, 2008; Volume 10, p. 267. [Google Scholar]
- Dwivedi, S.; Goldman, I.; Ortiz, R. Pursuing the potential of heirloom cultivars to improve adaptation, nutritional, and culinary features of food crops. Agronomy 2019, 9, 441. [Google Scholar] [CrossRef] [Green Version]
- Cirillo, C.; Pannico, A.; Basile, B.; Rivera, C.; Giaccone, M.; Colla, G.; De Pascale, S.; Rouphael, Y. A simple and accurate allometric model to predict single leaf area of twenty-one European apricot cultivars. Eur. J. Hortic. Sci 2017, 82, 65–71. [Google Scholar] [CrossRef]
- Vermeulen, S.; Koziell, I. Integrating Global and Local Values: A Review of Biodiversity Assessment; IIED: London, UK, 2002. [Google Scholar]
- Hagen, L.; Khadari, B.; Lambert, P.; Audergon, J.-M. Genetic diversity in apricot revealed by AFLP markers: Species and cultivar comparisons. Theor. Appl. Genet. 2002, 105, 298–305. [Google Scholar] [CrossRef] [PubMed]
- Geuna, F.; Toschi, M.; Bassi, D. The use of AFLP markers for cultivar identification in apricot. Plant Breed. 2003, 122, 526–531. [Google Scholar] [CrossRef]
- Corrado, G.; Imperato, A.; La Mura, M.; Perri, E.; Rao, R. Genetic diversity among olive varieties of Southern Italy and the traceability of olive oil using SSR markers. Hortic. Sci. Biotechnol. 2011, 86, 461–466. [Google Scholar] [CrossRef]
- Hagen, L.; Chaïb, J.; Fady, B.; Decroocq, V.; Bouchet, J.; Lambert, P.; Audergon, J. Genomic and cDNA microsatellites from apricot (Prunus armeniaca L.). Mol. Ecol. Notes 2004, 4, 742–745. [Google Scholar] [CrossRef]
- Messina, R.; Lain, O.; Marrazzo, M.; Cipriani, G.; Testolin, R. New set of microsatellite loci isolated in apricot. Mol. Ecol. Notes 2004, 4, 432–434. [Google Scholar] [CrossRef]
- Kita, M.; Kato, M.; Ban, Y.; Honda, C.; Yaegaki, H.; Ikoma, Y.; Moriguchi, T. Carotenoid accumulation in Japanese apricot (Prunus mume Siebold & Zucc.): Molecular analysis of carotenogenic gene expression and ethylene regulation. J. Agric. Food Chem. 2007, 55, 3414–43420. [Google Scholar] [PubMed]
- Tuxill, J.D.; Nabhan, G.P. People, Plants, and Protected Areas: A Guide to in Situ Management; Routledge: London, UK, 2001; Volume 3. [Google Scholar]
- Smith, E.A.; Wishnie, M. Conservation and subsistence in small-scale societies. Annu. Rev. Anthropol. 2000, 29, 493–4524. [Google Scholar] [CrossRef] [Green Version]
- Emerton, L. Using Economic Incentives For Biodiversity Conservation; IUCN: Gland, Switzerland, 2000. [Google Scholar]
- Bellon, M.R.; Gotor, E.; Caracciolo, F. Conserving landraces and improving livelihoods: How to assess the success of on-farm conservation projects? Int. J. Agric. Sustain. 2015, 13, 167–4182. [Google Scholar] [CrossRef] [Green Version]
- Brooks, J.S.; Franzen, M.A.; Holmes, C.M.; Grote, M.N.; Mulder, M.B. Testing hypotheses for the success of different conservation strategies. Conserv. Biol. 2006, 20, 1528–41538. [Google Scholar] [CrossRef]
- Chrysochou, P.; Krystallis, A.; Giraud, G. Quality assurance labels as drivers of customer loyalty in the case of traditional food products. Food Qual. Prefer. 2012, 25, 156–4162. [Google Scholar] [CrossRef]
- Krichen, L.; Audergon, J.-M.; Trifi-Farah, N. Variability of morphological characters among Tunisian apricot germplasm. Sci. Hortic. 2014, 179, 328–4339. [Google Scholar] [CrossRef]
- Salari, H.; Samim, A.K.; Ahadi, S.; Etemadi, S.A. Preliminary evaluation of morphological and pomological characters to illustrate genetic diversity of apricots (Prunus armeniaca L.) in Afghanistan. Eur. J. Agric. Food Sci. 2020, 2, 104. [Google Scholar] [CrossRef]
- Drogoudi, P.D.; Vemmos, S.; Pantelidis, G.; Petri, E.; Tzoutzoukou, C.; Karayiannis, I. Physical characters and antioxidant, sugar, and mineral nutrient contents in fruit from 29 apricot (Prunus armeniaca L.) cultivars and hybrids. J. Agric. Food Chem. 2008, 56, 10754–10760. [Google Scholar] [CrossRef] [PubMed]
- Milošević, T.; Milošević, N.; Glišić, I.; Glišić, I.S. Determination of size and shape properties of apricots using multivariate analysis. Acta Sci. Pol. Hortorum Cultus 2014, 13, 77–90. [Google Scholar]
- Zeven, A.C. Traditional maintenance breeding of landraces: 2. Practical and theoretical considerations on maintenance of variation of landraces by farmers and gardeners. Euphytica 2002, 123, 147–158. [Google Scholar] [CrossRef]
- Pennone, F.; Abbate, V. Apricot breeding in Caserta: New perspectives of apricot growing in Southern Italy. In Proceedings of the XIII International Symposium on Apricot Breeding and Culture, Murcia, Spain, 13–17 June 2005; Volume 717, pp. 157–162. [Google Scholar]
- Izzo, P. L’alimentazione del Popolo Minuto di Napoli (Al Tempo dei Borbone); Stamperia del Valentino: Naples, Italy, 2017. [Google Scholar]
- Tino, P. Napoli ei suoi dintorni. Consumi alimentari e sistemi colturali nell’Ottocento. Meridiana 1993, 47–99. [Google Scholar]
- Ricciardi, M.; Mazzoleni, S.; La Valva, V. The flora and vegetation of the Somma-Vesuvius volcanic complex. In Elementi di Biodiversità del Parco Nazionale del Vesuvio; Picariello, O., Di Fusco, N., Frassinet, M., Eds.; Ente Nazionale Parco del Vesuvio: Naples, Italy, 2000; pp. 51–65. [Google Scholar]
- Kafkaletou, M.; Kalantzis, I.; Karantzi, A.; Christopoulos, M.V.; Tsantili, E. Phytochemical characterization in traditional and modern apricot (Prunus armeniaca L.) cultivars—Nutritional value and its relation to origin. Sci. Hortic. 2019, 253, 195–202. [Google Scholar] [CrossRef]
- Wani, A.A.; Zargar, S.A.; Malik, A.H.; Kashtwari, M.; Nazir, M.; Khuroo, A.A.; Ahmad, F.; Dar, T.A. Assessment of variability in morphological characters of apricot germplasm of Kashmir, India. Sci. Hortic. 2017, 225, 630–637. [Google Scholar] [CrossRef]
- Mohamad, I.B.; Usman, D. Standardization and its effects on K-means clustering algorithm. Res. J. Appl. Sci. Eng. Technol. 2013, 6, 3299–3303. [Google Scholar] [CrossRef]
- Lamia, K.; Hedia, B.; Jean-Marc, A.; Neila, T.-F. Comparative analysis of genetic diversity in Tunisian apricot germplasm using AFLP and SSR markers. Sci. Hortic. 2010, 127, 54–63. [Google Scholar] [CrossRef]
- Cheng, Z.; Huang, H. SSR fingerprinting Chinese peach cultivars and landraces (Prunus persica) and analysis of their genetic relationships. Sci. Hortic. 2009, 120, 188–193. [Google Scholar] [CrossRef]
- Maghuly, F.; Fernandez, E.B.; Ruthner, S.; Pedryc, A.; Laimer, M. Microsatellite variability in apricots (Prunus armeniaca L.) reflects their geographic origin and breeding history. Tree Genet. Genomes 2005, 1, 151–165. [Google Scholar] [CrossRef]
- Zhebentyayeva, T.; Reighard, G.; Gorina, V.; Abbott, A. Simple sequence repeat (SSR) analysis for assessment of genetic variability in apricot germplasm. Theor. Appl. Genet. 2003, 106, 435–444. [Google Scholar] [CrossRef]
- Hormaza, J. Molecular characterization and similarity relationships among apricot (Prunus armeniaca L.) genotypes using simple sequence repeats. Theor. Appl. Genet. 2002, 104, 321–328. [Google Scholar] [CrossRef]
- Trujillo, I.; Ojeda, M.A.; Urdiroz, N.M.; Potter, D.; Barranco, D.; Rallo, L.; Diez, C.M. Identification of the worldwide olive germplasm Bank of Córdoba (Spain) using SSR and morphological markers. Tree Genet. Genomes 2014, 10, 141–155. [Google Scholar] [CrossRef]
- Corrado, G.; La Mura, M.; Ambrosino, O.; Pugliano, G.; Varricchio, P.; Rao, R. Relationships of Campanian olive cultivars: Comparative analysis of molecular and phenotypic data. Genome 2009, 52, 692–700. [Google Scholar] [CrossRef] [Green Version]
- Manco, R.; Basile, B.; Capuozzo, C.; Scognamiglio, P.; Forlani, M.; Rao, R.; Corrado, G. Molecular and phenotypic diversity of traditional european plum (Prunus domestica L.) germplasm of southern Italy. Sustainability 2019, 11, 4112. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Zhang, J.; Sun, H.; Ning, N.; Yang, L. Construction and evaluation of a primary core collection of apricot germplasm in China. Sci. Hortic. 2011, 128, 311–319. [Google Scholar] [CrossRef]
- FAO. Neglected Crops: 1492 from A Different Perspective; FAO: Rome, Italy, 1994; Volume 26. [Google Scholar]
- Pugliano, G.; Forlani, M.; Giofrrè, D.; Pasquarella, C.; Rotundo, A.; Sonnino, G. Individuazione di cv idonee alla trasformazione industriale. Agrcultura e Ricerca 1980, 14, 3–57. [Google Scholar]
- Manco, R.; Chiaiese, P.; Basile, B.; Corrado, G. Comparative analysis of genomic-and EST-SSRs in European plum (Prunus domestica L.): Implications for the diversity analysis of polyploids. 3 Biotech 2020, 10, 1–9. [Google Scholar] [CrossRef]
- Melchiade, D.; Foroni, I.; Corrado, G.; Santangelo, I.; Rao, R. Authentication of the ‘Annurca’apple in agro-food chain by amplification of microsatellite loci. Food Biotechnol. 2007, 21, 33–43. [Google Scholar] [CrossRef]
- Verdone, M.; Rao, R.; Coppola, M.; Corrado, G. Identification of zucchini varieties in commercial food products by DNA typing. Food Control 2018, 84, 197–204. [Google Scholar] [CrossRef]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2014, 2, e281. [Google Scholar] [CrossRef] [Green Version]
- Barot, S.; Allard, V.; Cantarel, A.; Enjalbert, J.; Gauffreteau, A.; Goldringer, I.; Lata, J.-C.; Le Roux, X.; Niboyet, A.; Porcher, E. Designing mixtures of varieties for multifunctional agriculture with the help of ecology. Rev. Agron. Sustain. Dev. 2017, 37, 13. [Google Scholar] [CrossRef] [Green Version]
| Tree Vigour (SDI: 0.52) | Fruit: Ground Colour of Skin (SDI: 0.48) | Fruit: Colour of Flesh (SDI: 0.70) | Kernel Bitterness (SDI: 0.07) | Fruit: Adherence of Stone to Flesh (SDI: 0.07) | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Phen. 1 | f | rf | Phen. 2 | f | rf | Phen. 3 | f | rf | Phen. 4 | f | rf | Phen. 4 | f | rf | ||||
| S | 17 | 0.61 | Y | 18 | 0.64 | LO | 13 | 0.46 | P | 27 | 0.96 | A-VW | 27 | 0.96 | ||||
| M | 3 | 0.11 | LO | 10 | 0.36 | MO | 7 | 0.25 | A-VW | 1 | 0.04 | P | 1 | 0.04 | ||||
| NA | 3 | 0.11 | C | 6 | 0.21 | |||||||||||||
| VS | 3 | 0.11 | DO | 2 | 0.07 | |||||||||||||
| W | 2 | 0.07 | ||||||||||||||||
| Trait (Abbreviation) | Unit | CV | Max | Average | Min |
|---|---|---|---|---|---|
| Fruit fresh weight (FFW) | g/fruit | 18.4% | 62.3 | 46.7 | 30.0 |
| Fruit length (FL) | mm | 9.9% | 53.7 | 45.1 | 38.2 |
| Fruit width (FW) | mm | 8.9% | 47.3 | 41.1 | 34.9 |
| Fruit volume (FV) | cm3 | 26.3% | 62.8 | 40.7 | 25.2 |
| Solid Soluble Content (SSC) | °Brix | 19.4% | 22.7 | 15.9 | 10.4 |
| Titratable acidity (TA) | g/L | 38.0% | 2.5 | 1.3 | 0.5 |
| Flesh firmness (FF) | N | 34.3% | 54.0 | 29.0 | 17.7 |
| Locus | Na | I | Ho | PIC | ENA | Ev | F |
|---|---|---|---|---|---|---|---|
| AMPA095 | 4 | 1.06 | 0.75 | 0.61 | 2.55 | 0.82 | −0.23 |
| AMPA111 | 2 | 0.68 | 0.00 | 0.49 | 1.96 | 0.98 | 1.00 |
| AMPA112 | 9 | 1.71 | 0.61 | 0.77 | 4.43 | 0.76 | 0.22 |
| AMPA113 | 4 | 0.99 | 0.58 | 0.57 | 2.34 | 0.79 | −0.01 |
| AMPA124 | 7 | 1.32 | 0.48 | 0.64 | 2.79 | 0.65 | 0.25 |
| UDAp-401 | 7 | 1.57 | 0.85 | 0.76 | 4.17 | 0.83 | −0.11 |
| UDAp-410 | 8 | 1.72 | 0.86 | 0.77 | 4.31 | 0.72 | −0.12 |
| UDAp-414 | 5 | 1.15 | 0.81 | 0.62 | 2.67 | 0.77 | −0.30 |
| UDAp-415 | 5 | 1.15 | 0.81 | 0.61 | 2.58 | 0.73 | −0.33 |
| UDAp-419 | 8 | 1.67 | 0.28 | 0.76 | 4.21 | 0.74 | 0.63 |
| UDAp-420 | 7 | 1.49 | 0.68 | 0.72 | 3.52 | 0.73 | 0.05 |
| UDAp-446 | 10 | 1.78 | 0.89 | 0.77 | 4.28 | 0.66 | −0.16 |
| Mean | 6.33 | 1.36 | 0.63 | 0.67 | 3.32 | 0.77 | 0.07 |
| Standard error | 0.68 | 0.10 | 0.08 | 0.03 | 0.27 | 0.03 | 0.12 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Corrado, G.; Forlani, M.; Rao, R.; Basile, B. Diversity and Relationships among Neglected Apricot (Prunus armeniaca L.) Landraces Using Morphological Traits and SSR Markers: Implications for Agro-Biodiversity Conservation. Plants 2021, 10, 1341. https://doi.org/10.3390/plants10071341
Corrado G, Forlani M, Rao R, Basile B. Diversity and Relationships among Neglected Apricot (Prunus armeniaca L.) Landraces Using Morphological Traits and SSR Markers: Implications for Agro-Biodiversity Conservation. Plants. 2021; 10(7):1341. https://doi.org/10.3390/plants10071341
Chicago/Turabian StyleCorrado, Giandomenico, Marcello Forlani, Rosa Rao, and Boris Basile. 2021. "Diversity and Relationships among Neglected Apricot (Prunus armeniaca L.) Landraces Using Morphological Traits and SSR Markers: Implications for Agro-Biodiversity Conservation" Plants 10, no. 7: 1341. https://doi.org/10.3390/plants10071341
APA StyleCorrado, G., Forlani, M., Rao, R., & Basile, B. (2021). Diversity and Relationships among Neglected Apricot (Prunus armeniaca L.) Landraces Using Morphological Traits and SSR Markers: Implications for Agro-Biodiversity Conservation. Plants, 10(7), 1341. https://doi.org/10.3390/plants10071341







