Utilization of a Wheat50K SNP Microarray-Derived High-Density Genetic Map for QTL Mapping of Plant Height and Grain Traits in Wheat
Abstract
1. Introduction
2. Results
2.1. Agronomic Traits Analysis
2.2. Construction of a Genetic Map
2.2.1. Description and Illustration of a Genetic Map
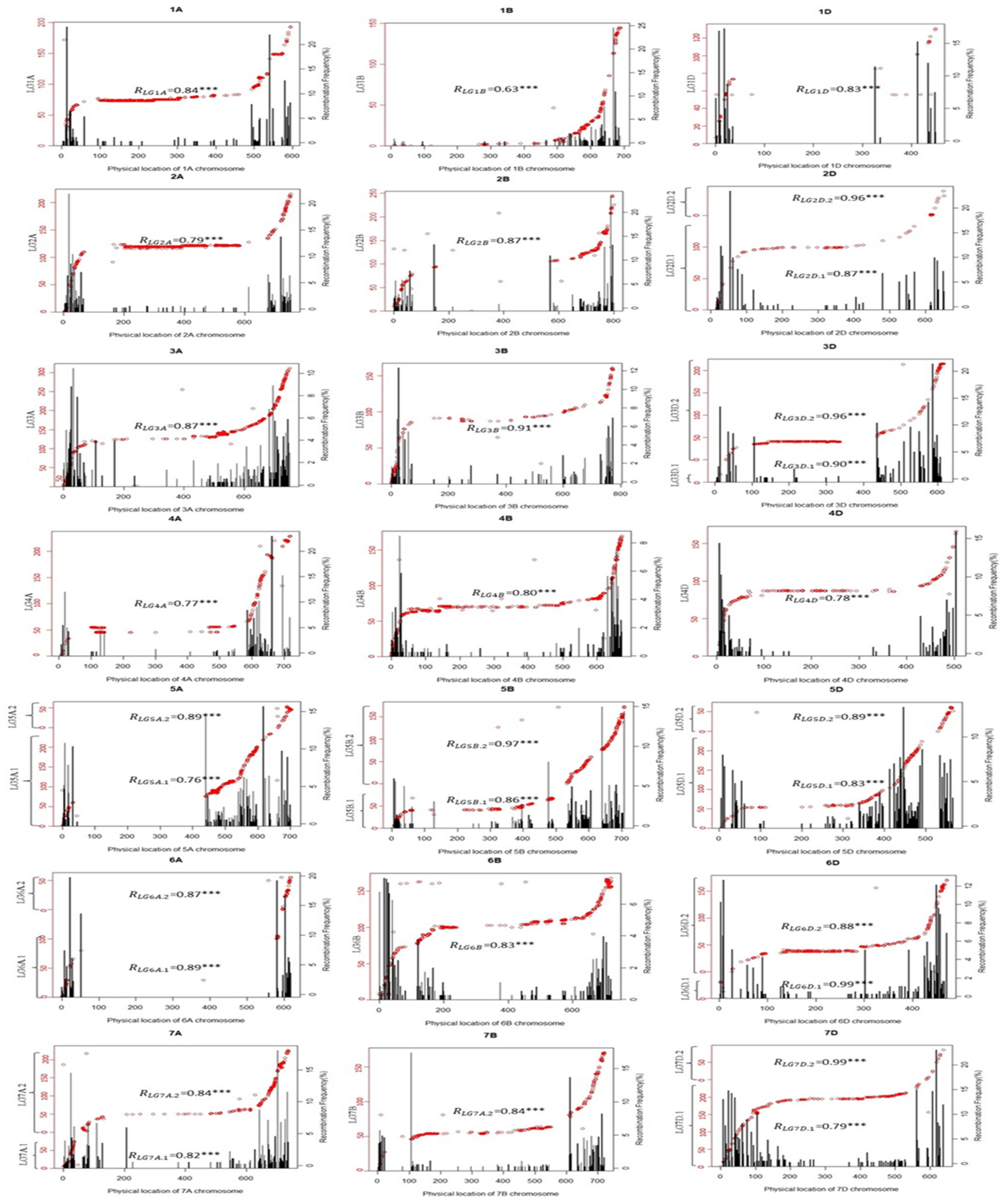
2.2.2. Collinearity Analysis of the Genetic Map
2.2.3. Effects of 1B/1R on Traits Related to Plant Height and TGW
2.3. QTL Mapping Analysis
3. Discussion
3.1. The Impact of Linkage Map on QTL Mapping
3.2. Comparison with Previous Research Results
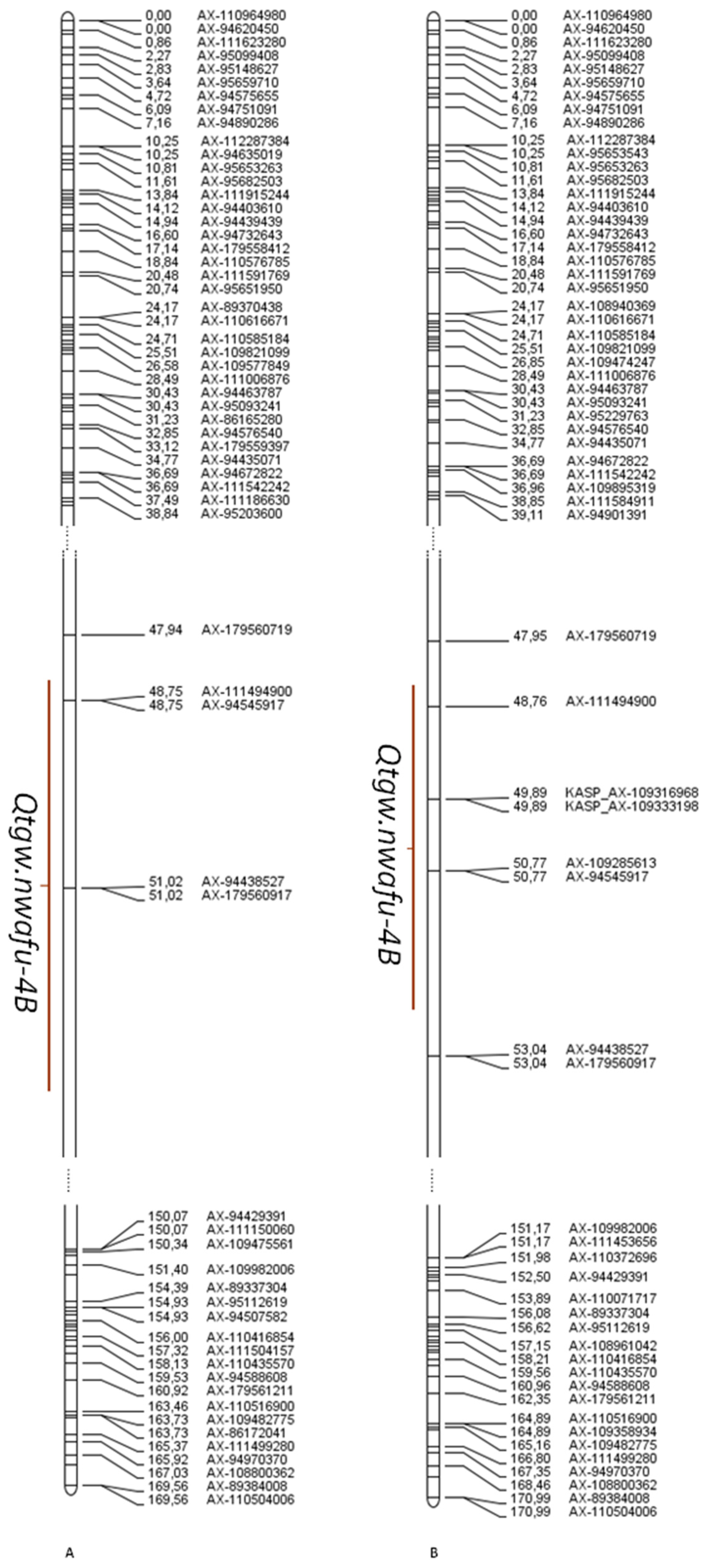
3.3. Qtgw.Nwafu-4B Molecular Marker Development
4. Materials and Methods
4.1. Plant Materials, Experimental Design, and Investigation of Agronomic Traits
4.2. Construction and Evaluation of Genetic Maps
4.3. Identification of 1BL/1RS Translocation
4.4. Detection of Quantitative Loci
4.5. Breeding Molecular Marker Development
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bailey-Serres, J.; Parker, J.E.; Ainsworth, E.A.; Oldroyd, G.E.D.; Schroeder, J.I. Genetic strategies for improving crop yields. Nature 2019, 575, 109–118. [Google Scholar] [CrossRef]
- Hanif, M.; Gao, F.; Liu, J.; Wen, W.; Zhang, Y.; Rasheed, A.; Xia, X.; He, Z.; Cao, S. TaTGW6-A1, an ortholog of rice TGW6, is associated with grain weight and yield in bread wheat. Mol. Breed. 2016, 36, 1. [Google Scholar] [CrossRef]
- Kumari, S.; Jaiswal, V.; Mishra, V.K.; Paliwal, R.; Balyan, H.S.; Gupta, P.K. QTL mapping for some grain traits in bread wheat (Triticum aestivum L.). Physiol. Mol. Biol. Plants 2018, 24, 909–920. [Google Scholar] [CrossRef]
- Xie, Q.; Mayes, S.; Sparkes, D.L. Carpel size, grain filling, and morphology determine individual grain weight in wheat. J. Exp. Bot. 2015, 66, 6715–6730. [Google Scholar] [CrossRef]
- Guan, P.; Di, N.; Mu, Q.; Shen, X.; Wang, Y.; Wang, X.; Yu, K.; Song, W.; Chen, Y.; Xin, M.; et al. Use of near-isogenic lines to precisely map and validate a major QTL for grain weight on chromosome 4AL in bread wheat (Triticum aestivum L.). Theor. Appl. Genet. 2019, 132, 2367–2379. [Google Scholar] [CrossRef]
- Zhai, H.; Feng, Z.; Li, J.; Liu, X.; Xiao, S.; Ni, Z.; Sun, Q. QTL Analysis of Spike Morphological Traits and Plant Height in Winter Wheat (Triticum aestivum L.) Using a High-Density SNP and SSR-Based Linkage Map. Front. Plant Sci. 2016, 7, 1617. [Google Scholar] [CrossRef]
- Jiang, Q.; Hou, J.; Hao, C.; Wang, L.; Ge, H.; Dong, Y.; Zhang, X. The wheat (T. aestivum) sucrose synthase 2 gene (TaSus2) active in endosperm development is associated with yield traits. Funct. Integr. Genom. 2011, 11, 49–61. [Google Scholar] [CrossRef]
- Hou, J.; Jiang, Q.; Hao, C.; Wang, Y.; Zhang, H.; Zhang, X. Global selection on sucrose synthase haplotypes during a century of wheat breeding. Plant Physiol. 2014, 164, 1918–1929. [Google Scholar] [CrossRef]
- Su, Z.; Hao, C.; Wang, L.; Dong, Y.; Zhang, X. Identification and development of a functional marker of TaGW2 associated with grain weight in bread wheat (Triticum aestivum L.). Theor. Appl. Genet. 2011, 122, 211–223. [Google Scholar] [CrossRef]
- Yang, Z.; Bai, Z.; Li, X.; Wang, P.; Wu, Q.; Yang, L.; Li, L.; Li, X. SNP identification and allelic-specific PCR markers development for TaGW2, a gene linked to wheat kernel weight. Theor. Appl. Genet. 2012, 125, 1057–1068. [Google Scholar] [CrossRef]
- Ma, L.; Li, T.; Hao, C.; Wang, Y.; Chen, X.; Zhang, X. TaGS5–3A, a grain size gene selected during wheat improvement for larger kernel and yield. Plant Biotechnol. J. 2016, 14, 1269–1280. [Google Scholar] [CrossRef]
- Ma, D.; Yan, J.; He, Z.; Wu, L.; Xia, X. Characterization of a cell wall invertase gene TaCwi-A1 on common wheat chromosome 2A and development of functional markers. Mol. Breed. 2010, 29, 43–52. [Google Scholar] [CrossRef]
- Zhang, L.; Zhao, Y.L.; Gao, L.F.; Zhao, G.Y.; Zhou, R.H.; Zhang, B.S.; Jia, J.Z. TaCKX6-D1, the ortholog of rice OsCKX2, is associated with grain weight in hexaploid wheat. New Phytol. 2012, 195, 574–584. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.K.; Rustgi, S.; Kumar, N. Genetic and molecular basis of grain size and grain number and its relevance to grain productivity in higher plants. Genome 2006, 49, 565–571. [Google Scholar] [CrossRef]
- Varshney, R.K.; Prasad, M.; Roy, J.K.; Kumar, N.; Harjit, S.; Dhaliwal, H.S.; Balyan, H.S.; Gupta, P.K. Identification of eight chromosomes and a microsatellite marker on 1AS associated with QTL for grain weight in bread wheat. Theor. Appl. Genet. 2000, 100, 1290–1294. [Google Scholar] [CrossRef]
- Cao, P.; Liang, X.; Zhao, H.; Feng, B.; Xu, E.; Wang, L.; Hu, Y. Identification of the quantitative trait loci controlling spike-related traits in hexaploid wheat (Triticum aestivum L.). Planta 2019, 250, 1967–1981. [Google Scholar] [CrossRef]
- Campbell, B.T.; Baenziger, P.S.; Gill, K.S.; Eskridge, K.M.; Budak, H.; Erayman, M.; Dweikat, I.; Yen, Y. Identification of QTLs and Environmental Interactions Associated with Agronomic Traits on Chromosome 3A of Wheat. Crop Sci. 2003, 43, 1493–1505. [Google Scholar] [CrossRef]
- Borner, A.; Schumann, E.; Furste, A.; Coster, H.; Leithold, B.; Roder, S.; Weber, E. Mapping of quantitative trait loci determining agronomic important characters in hexaploid wheat (Triticum aestivum L.). Theor. Appl. Genet. 2002, 105, 921–936. [Google Scholar] [CrossRef]
- Huang, X.Q.; Coster, H.; Ganal, M.W.; Roder, M.S. Advanced backcross QTL analysis for the identification of quantitative trait loci alleles from wild relatives of wheat (Triticum aestivum L.). Theor. Appl. Genet. 2003, 106, 1379–1389. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.Q.; Cloutier, S.; Lycar, L.; Radovanovic, N.; Humphreys, D.G.; Noll, J.S.; Somers, D.J.; Brown, P.D. Molecular detection of QTLs for agronomic and quality traits in a doubled haploid population derived from two Canadian wheats (Triticum aestivum L.). Theor. Appl. Genet. 2006, 113, 753–766. [Google Scholar] [CrossRef] [PubMed]
- Narasimhamoorthy, B.; Gill, B.S.; Fritz, A.K.; Nelson, J.C.; Brown-Guedira, G.L. Advanced backcross QTL analysis of a hard winter wheat × synthetic wheat population. Theor. Appl. Genet. 2006, 112, 787–796. [Google Scholar] [CrossRef] [PubMed]
- Breseghello, F.; Sorrells, M.E. Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 2006, 172, 1165–1177. [Google Scholar] [CrossRef] [PubMed]
- Breseghello, F.; Sorrells, M.E. QTL analysis of kernel size and shape in two hexaploid wheat mapping populations. Field Crops Res. 2007, 101, 172–179. [Google Scholar] [CrossRef]
- Ramya, P.; Chaubal, A.; Kulkarni, K.; Gupta, L.; Kadoo, N.; Dhaliwal, H.S.; Chhuneja, P.; Lagu, M.; Gupta, V. QTL mapping of 1000-kernel weight, kernel length, and kernel width in bread wheat (Triticum aestivum L.). J. Appl. Genet. 2010, 51, 421–429. [Google Scholar] [CrossRef] [PubMed]
- Mir, R.R.; Kumar, N.; Jaiswal, V.; Girdharwal, N.; Prasad, M.; Balyan, H.S.; Gupta, P.K. Genetic dissection of grain weight in bread wheat through quantitative trait locus interval and association mapping. Mol. Breed. 2012, 29, 963–972. [Google Scholar] [CrossRef]
- Rasheed, A.; Xia, X.; Ogbonnaya, F.; Mahmood, T.; Zhang, Z.; Mujeeb-Kazi, A.; He, Z. Genome-wide association for grain morphology in synthetic hexaploid wheats using digital imaging analysis. BMC Plant Biol. 2014, 14, 128. [Google Scholar] [CrossRef] [PubMed]
- Williams, K.; Munkvold, J.; Sorrells, M. Comparison of digital image analysis using elliptic Fourier descriptors and major dimensions to traits seed shape in hexaploid wheat (Triticum aestivum L.). Euphytica 2012, 190, 99–116. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, J.; Xia, X.; He, Z. TaGS-D1, an ortholog of rice OsGS3, is associated with grain weight and grain length in common wheat. Mol. Breed. 2014, 34, 1097–1107. [Google Scholar] [CrossRef]
- Jia, H.; Wan, H.; Yang, S.; Zhang, Z.; Kong, Z.; Xue, S.; Zhang, L.; Ma, Z. Genetic dissection of yield-related traits in a recombinant inbred line population created using a key breeding parent in China’s wheat breeding. Theor. Appl. Genet. 2013, 126, 2123–2139. [Google Scholar] [CrossRef] [PubMed]
- Guan, P.; Lu, L.; Jia, L.; Kabir, M.R.; Zhang, J.; Lan, T.; Zhao, Y.; Xin, M.; Hu, Z.; Yao, Y.; et al. Global QTL Analysis Identifies Genomic Regions on Chromosomes 4A and 4B Harboring Stable Loci for Yield-Related Traits Across Different Environments in Wheat (Triticum aestivum L.). Front. Plant Sci. 2018, 9, 529. [Google Scholar] [CrossRef]
- Cuthbert, J.L.; Somers, D.J.; Brule-Babel, A.L.; Brown, P.D.; Crow, G.H. Molecular mapping of quantitative trait loci for yield and yield components in spring wheat (Triticum aestivum L.). Theor. Appl. Genet. 2008, 117, 595–608. [Google Scholar] [CrossRef] [PubMed]
- Mohler, V.; Albrecht, T.; Castell, A.; Diethelm, M.; Schweizer, G.; Hartl, L. Considering causal genes in the genetic dissection of kernel traits in common wheat. J. Appl. Genet. 2016, 57, 467–476. [Google Scholar] [CrossRef]
- Brinton, J.; Simmonds, J.; Minter, F.; Leverington-Waite, M.; Snape, J.; Uauy, C. Increased pericarp cell length underlies a major quantitative trait locus for grain weight in hexaploid wheat. New Phytol. 2017, 215, 1026–1038. [Google Scholar] [CrossRef]
- Onyemaobi, I.; Ayalew, H.; Liu, H.; Siddique, K.H.M.; Yan, G. Identification and validation of a major chromosome region for high grain number per spike under meiotic stage water stress in wheat (Triticum aestivum L.). PLoS ONE 2018, 13, e0194075. [Google Scholar] [CrossRef]
- Li, F.; Wen, W.; He, Z.; Liu, J.; Jin, H.; Cao, S.; Geng, H.; Yan, J.; Zhang, P.; Wan, Y.; et al. Genome-wide linkage mapping of yield-related traits in three Chinese bread wheat populations using high-density SNP markers. Theor. Appl. Genet. 2018, 131, 1903–1924. [Google Scholar] [CrossRef]
- Kumar, A.; Mantovani, E.E.; Seetan, R.; Soltani, A.; Echeverry-Solarte, M.; Jain, S.; Simsek, S.; Doehlert, D.; Alamri, M.S.; Elias, E.M.; et al. Dissection of Genetic Factors underlying Wheat Kernel Shape and Size in an Elite x Nonadapted Cross using a High Density SNP Linkage Map. Plant. Genome 2016, 9, 1. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.H.; Chen, Y.X.; Zhou, S.H.; Fu, L.; Chen, J.J.; Xiao, Y.; Zhang, D.; Ouyang, S.H.; Zhao, X.J.; Cui, Y.; et al. High-density genetic linkage map construction and QTL mapping of grain shape and size in the wheat population Yanda1817 x Beinong6. PLoS ONE 2015, 10, e0118144. [Google Scholar] [CrossRef] [PubMed]
- Cui, F.; Zhao, C.; Ding, A.; Li, J.; Wang, L.; Li, X.; Bao, Y.; Li, J.; Wang, H. Construction of an integrative linkage map and QTL mapping of grain yield-related traits using three related wheat RIL populations. Theor. Appl. Genet. 2014, 127, 659–675. [Google Scholar] [CrossRef]
- Fan, X.; Cui, F.; Ji, J.; Zhang, W.; Zhao, X.; Liu, J.; Meng, D.; Tong, Y.; Wang, T.; Li, J. Dissection of Pleiotropic QTL Regions Controlling Wheat Spike Characteristics Under Different Nitrogen Treatments Using Traditional and Conditional QTL Mapping. Front. Plant Sci. 2019, 10, 187. [Google Scholar] [CrossRef]
- Liu, G.; Jia, L.; Lu, L.; Qin, D.; Zhang, J.; Guan, P.; Ni, Z.; Yao, Y.; Sun, Q.; Peng, H. Mapping QTLs of yield-related traits using RIL population derived from common wheat and Tibetan semi-wild wheat. Theor. Appl. Genet. 2014, 127, 2415–2432. [Google Scholar] [CrossRef]
- Liu, J.; Wu, B.; Singh, R.P.; Velu, G. QTL mapping for micronutrients concentration and yield component traits in a hexaploid wheat mapping population. J. Cereal Sci. 2019, 88, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Quarrie, S.A.; Steed, A.; Calestani, C.; Semikhodskii, A.; Lebreton, C.; Chinoy, C.; Steele, N.; Pljevljakusic, D.; Waterman, E.; Weyen, J.; et al. A high-density genetic map of hexaploid wheat (Triticum aestivum L.) from the cross Chinese Spring x SQ1 and its use to compare QTLs for grain yield across a range of environments. Theor. Appl. Genet. 2005, 110, 865–880. [Google Scholar] [CrossRef]
- Hai, L.; Guo, H.; Wagner, C.; Xiao, S.; Friedt, W. Genomic regions for yield and yield parameters in Chinese winter wheat (Triticum aestivum L.) genotypes tested under varying environments correspond to QTL in widely different wheat materials. Plant Sci. 2008, 175, 226–232. [Google Scholar] [CrossRef]
- Wang, J.; Liu, W.; Wang, H.; Li, L.; Wu, J.; Yang, X.; Li, X.; Gao, A. QTL mapping of yield-related traits in the wheat germplasm 3228. Euphytica 2010, 177, 277–292. [Google Scholar] [CrossRef]
- Lee, H.S.; Jung, J.-U.; Kang, C.-S.; Heo, H.-Y.; Park, C.S. Mapping of QTL for yield and its related traits in a doubled haploid population of Korean wheat. Plant Biotechnol. Rep. 2014, 8, 443–454. [Google Scholar] [CrossRef]
- Roder, M.S.; Huang, X.Q.; Borner, A. Fine mapping of the region on wheat chromosome 7D controlling grain weight. Funct. Integr. Genom. 2008, 8, 79–86. [Google Scholar] [CrossRef]
- Tian, X.; Wen, W.; Xie, L.; Fu, L.; Xu, D.; Fu, C.; Wang, D.; Chen, X.; Xia, X.; Chen, Q.; et al. Molecular Mapping of Reduced Plant Height Gene Rht24 in Bread Wheat. Front. Plant Sci. 2017, 8, 1379. [Google Scholar] [CrossRef]
- Mo, Y.; Vanzetti, L.S.; Hale, I.; Spagnolo, E.J.; Guidobaldi, F.; Al-Oboudi, J.; Odle, N.; Pearce, S.; Helguera, M.; Dubcovsky, J. Identification and characterization of Rht25, a locus on chromosome arm 6AS affecting wheat plant height, heading time, and spike development. Theor. Appl. Genet. 2018, 131, 2021–2035. [Google Scholar] [CrossRef]
- Agarwal, P.; Balyan, H.S.; Gupta, P.K. Identification of modifiers of the plant height in wheat using an induced dwarf mutant controlled by RhtB4c allele. Physiol. Mol. Biol. Plants 2020, 26, 2283–2289. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Dell, B.; Biddulph, B.; Drake-Brockman, F.; Walker, E.; Khan, N.; Wong, D.; Hayden, M.; Appels, R. Wild-type alleles of Rht-B1 and Rht-D1 as independent determinants of thousand-grain weight and kernel number per spike in wheat. Mol. Breed. 2013, 32, 771–783. [Google Scholar] [CrossRef]
- Chai, L.; Chen, Z.; Bian, R.; Zhai, H.; Cheng, X.; Peng, H.; Yao, Y.; Hu, Z.; Xin, M.; Guo, W.; et al. Dissection of two quantitative trait loci with pleiotropic effects on plant height and spike length linked in coupling phase on the short arm of chromosome 2D of common wheat (Triticum aestivum L.). Theor. Appl. Genet. 2019, 132, 1815–1831. [Google Scholar] [CrossRef]
- Chen, Z.; Cheng, X.; Chai, L.; Wang, Z.; Bian, R.; Li, J.; Zhao, A.; Xin, M.; Guo, W.; Hu, Z.; et al. Dissection of genetic factors underlying grain size and fine mapping of QTgw.cau-7D in common wheat (Triticum aestivum L.). Theor. Appl. Genet. 2020, 133, 149–162. [Google Scholar] [CrossRef]
- Sun, C.; Dong, Z.; Zhao, L.; Ren, Y.; Zhang, N.; Chen, F. The Wheat 660K SNP array demonstrates great potential for marker-assisted selection in polyploid wheat. Plant Biotechnol. J. 2020, 18, 1354–1360. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Chen, J.; Zhang, W.; Hu, Y.; Chang, L.; Fang, L.; Wang, Q.; Lv, F.; Wu, H.; Si, Z.; et al. Sequence-based ultra-dense genetic and physical maps reveal structural variations of allopolyploid cotton genomes. Genome Biol. 2015, 16, 108. [Google Scholar] [CrossRef]
- Schwarzkopf, E.J.; Motamayor, J.C.; Cornejo, O.E. Genetic differentiation and intrinsic genomic features explain variation in recombination hotspots among cocoa tree populations. BMC Genom. 2020, 21, 332. [Google Scholar] [CrossRef]
- Cavanagh, C.R.; Chao, S.; Wang, S.; Huang, B.E.; Stephen, S.; Kiani, S.; Forrest, K.; Saintenac, C.; Brown-Guedira, G.L.; Akhunova, A.; et al. Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc. Natl. Acad. Sci. USA 2013, 110, 8057–8062. [Google Scholar] [CrossRef]
- Wang, S.; Wong, D.; Forrest, K.; Allen, A.; Chao, S.; Huang, B.E.; Maccaferri, M.; Salvi, S.; Milner, S.G.; Cattivelli, L.; et al. Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol. J. 2014, 12, 787–796. [Google Scholar] [CrossRef]
- Sun, C.; Zhang, F.; Yan, X.; Zhang, X.; Dong, Z.; Cui, D.; Chen, F. Genome-wide association study for 13 agronomic traits reveals distribution of superior alleles in bread wheat from the Yellow and Huai Valley of China. Plant Biotechnol. J. 2017, 15, 953–969. [Google Scholar] [CrossRef] [PubMed]
- Winfield, M.O.; Allen, A.M.; Burridge, A.J.; Barker, G.L.; Benbow, H.R.; Wilkinson, P.A.; Coghill, J.; Waterfall, C.; Davassi, A.; Scopes, G.; et al. High-density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol. J. 2016, 14, 1195–1206. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Huang, S.; Zeng, Q.; Wang, X.; Yu, R.; Wang, Q.; Singh, R.P.; Bhavani, S.; Kang, Z.; Wu, J.; et al. Refined mapping of stripe rust resistance gene YrP10090 within a desirable haplotype for wheat improvement on chromosome 6A. Theor. Appl. Genet. 2021, 3, 1–17. [Google Scholar]
- Liu, J.; Luo, W.; Qin, N.; Ding, P.; Zhang, H.; Yang, C.; Mu, Y.; Tang, H.; Liu, Y.; Li, W.; et al. A 55 K SNP array-based genetic map and its utilization in QTL mapping for productive tiller number in common wheat. Theor. Appl. Genet. 2018, 131, 2439–2450. [Google Scholar] [CrossRef]
- Ma, J.; Zhang, H.; Li, S.; Zou, Y.; Li, T.; Liu, J.; Ding, P.; Mu, Y.; Tang, H.; Deng, M.; et al. Identification of quantitative trait loci for kernel traits in a wheat cultivar Chuannong16. BMC Genet. 2019, 20, 77. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Ding, P.; Liu, J.; Li, T.; Zou, Y.; Habib, A.; Mu, Y.; Tang, H.; Jiang, Q.; Liu, Y.; et al. Identification and validation of a major and stably expressed QTL for spikelet number per spike in bread wheat. Theor. Appl. Genet. 2019, 132, 3155–3167. [Google Scholar] [CrossRef]
- Ren, T.; Hu, Y.; Tang, Y.; Li, C.; Yan, B.; Ren, Z.; Tan, F.; Tang, Z.; Fu, S.; Li, Z. Utilization of a Wheat55K SNP Array for Mapping of Major QTL for Temporal Expression of the Tiller Number. Front. Plant Sci. 2018, 9, 333. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Wu, J.; Wang, X.; Mu, J.; Xu, Z.; Zeng, Q.; Liu, S.; Wang, Q.; Kang, Z.; Han, D. Utilization of the Genomewide Wheat 55K SNP Array for Genetic Analysis of Stripe Rust Resistance in Common Wheat Line P9936. Phytopathology 2019, 109, 819–827. [Google Scholar] [CrossRef] [PubMed]
- Rasheed, A.; Xia, X. From markers to genome-based breeding in wheat. Theor. Appl. Genet. 2019, 132, 767–784. [Google Scholar] [CrossRef]
- Cui, F.; Fan, X.; Chen, M.; Zhang, N.; Zhao, C.; Zhang, W.; Han, J.; Ji, J.; Zhao, X.; Yang, L.; et al. QTL detection for wheat kernel size and quality and the responses of these traits to low nitrogen stress. Theor. Appl. Genet. 2016, 129, 469–484. [Google Scholar] [CrossRef]
- Hao, C.; Jiao, C.; Hou, J.; Li, T.; Liu, H.; Wang, Y.; Zheng, J.; Liu, H.; Bi, Z.; Xu, F.; et al. Resequencing of 145 Landmark Cultivars Reveals Asymmetric Sub-genome Selection and Strong Founder Genotype Effects on Wheat Breeding in China. Mol. Plant 2020, 12, 1733–1751. [Google Scholar] [CrossRef]
- Brinton, J.; Ramirez-Gonzalez, R.H.; Simmonds, J.; Wingen, L.; Orford, S.; Griffiths, S.; Wheat Genome, P.; Haberer, G.; Spannagl, M.; Walkowiak, S.; et al. A haplotype-led approach to increase the precision of wheat breeding. Commun. Biol. 2020, 3, 712. [Google Scholar] [CrossRef]
- Meng, L.; Li, H.; Zhang, L.; Wang, J. QTL IciMapping: Integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J. 2015, 3, 269–283. [Google Scholar] [CrossRef]
- Cui, F.; Zhang, N.; Fan, X.L.; Zhang, W.; Zhao, C.H.; Yang, L.J.; Pan, R.Q.; Chen, M.; Han, J.; Zhao, X.Q.; et al. Utilization of a Wheat660K SNP array-derived high-density genetic map for high-resolution mapping of a major QTL for kernel number. Sci. Rep. 2017, 7, 3788. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Yu, R.; Wang, H.; Zhou, C.; Huang, S.; Jiao, H.; Yu, S.; Nie, X.; Wang, Q.; Liu, S.; et al. A large-scale genomic association analysis identifies the candidate causal genes conferring stripe rust resistance under multiple field environments. Plant Biotechnol. J. 2021, 19, 177–191. [Google Scholar] [CrossRef] [PubMed]
- Banta, J.A.; Stevens, M.H.H.; Pigliucci, M. A comprehensive test of the ‘limiting resources’ framework applied to plant tolerance to apical meristem damage. Oikos 2010, 119, 359–369. [Google Scholar] [CrossRef]
- Wu, Y.; Bhat, P.R.; Close, T.J.; Lonardi, S. Efficient and accurate construction of genetic linkage maps from the minimum spanning tree of a graph. PLoS Genet. 2008, 4, e1000212. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; He, F.; Chen, G.L.; Cui, F.; Li, X.F. Identification of 1BL·1RS Wheat-Rye Chromosome Translocations via 1RS Specific Molecular Markers and Genomic in situ Hybridization. Acta Agron. Sin. 2011, 37, 563–569. [Google Scholar] [CrossRef]
| Traits | Environment | Xinong1376 | Xiaoyan81 | Mean ± SD | Minimum | Maximum | p-Value | Heritability |
|---|---|---|---|---|---|---|---|---|
| Plant height | 19NY | 65.25 | 77.75 ** | 67.08 ± 13.78 | 32.2 | 96.8 | 2.19 × 10−3 | 0.73 |
| 20NY | 68.24 | 81.22 ** | 80.03 ± 14.43 | 40.2 | 109.8 | 5.66 × 10−6 | ||
| 19YL | 68.36 | 78.23 ** | 65.78 ± 12.78 | 34.6 | 90.9 | 6.14 × 10−4 | ||
| 20YL | 72.33 | 83.25 ** | 72.24 ± 15.08 | 38.3 | 109.2 | 3.82 × 10−2 | ||
| Thousand Grain Weight | 19NY | 41.35 ** | 36.23 | 40.72 ± 4.37 | 27.81 | 52.19 | 1.12 × 10−1 | 0.62 |
| 20NY | 42.13 ** | 39.48 | 42.62 ± 4.51 | 26.28 | 51.76 | 1.24 × 10−3 | ||
| 19YL | 44.32 ** | 41.75 | 45.32 ± 4.41 | 34.22 | 55.05 | 2.80 × 10−2 | ||
| 20YL | 46.23 ** | 42.32 | 45.21 ± 4.40 | 29.5 | 54.83 | 3.68 × 10−1 | ||
| Grain length | 19NY | 7.12 ** | 6.87 | 7.23 ± 0.37 | 6.27 | 8.04 | 3.93 × 10−2 | 0.61 |
| 20NY | 7.32 ** | 6.75 | 7.14 ± 0.35 | 6.34 | 8.03 | 6.79 × 10−2 | ||
| 19YL | 7.51 ** | 7.24 | 7.44 ± 0.34 | 6.68 | 8.23 | 1.81 × 10−2 | ||
| 20YL | 7.36 ** | 7.14 | 7.51 ± 0.38 | 6.67 | 8.51 | 1.96 × 10−1 | ||
| Grain width | 19NY | 3.31 | 3.21 | 3.37 ± 0.15 | 2.88 | 3.69 | 1.45 × 10−3 | 0.72 |
| 20NY | 3.88 ** | 3.62 | 3.45 ± 0.18 | 2.81 | 3.83 | 2.18 × 10−3 | ||
| 19YL | 3.51 ** | 3.28 | 3.60 ± 0.16 | 3.11 | 3.9 | 1.38 × 10−2 | ||
| 20YL | 3.66 ** | 3.42 | 3.60 ± 0.16 | 3.16 | 3.95 | 3.02 × 10−3 |
| Chromosome | Linkage Group | Length(cM) | Maker Numbers | Bin Number | Insinuation Markers | Maximum Clearance | Average Bin | Bin Density |
|---|---|---|---|---|---|---|---|---|
| 1A | LG1A | 192.66 | 1064 | 112 | 1045 | 25.68 | 1.72 | 0.58 |
| 1B | LG1B | 144.61 | 558 | 118 | 447 | 26.86 | 1.23 | 0.82 |
| 1D | LG1D | 130.85 | 394 | 42 | 336 | 18.01 | 3.12 | 0.32 |
| 2A | LG2A | 215.97 | 951 | 140 | 940 | 23.46 | 1.54 | 0.65 |
| 2B | LG2B | 244.43 | 676 | 173 | 597 | 25.44 | 1.41 | 0.71 |
| 2D | LG2D.1 | 132.89 | 161 | 48 | 154 | 25.42 | 2.77 | 0.36 |
| LG2D.2 | 38.69 | 77 | 11 | 75 | 10.06 | 3.52 | 0.28 | |
| 3A | LG3A | 311.23 | 1322 | 285 | 1301 | 16.8 | 1.09 | 0.92 |
| 3B | LG3B | 160.61 | 487 | 144 | 458 | 12.59 | 1.12 | 0.9 |
| 3D | LG3D.1 | 17.46 | 38 | 8 | 36 | 13.71 | 2.18 | 0.46 |
| LG3D.2 | 214.66 | 999 | 87 | 1026 | 22.84 | 2.47 | 0.41 | |
| 4A | LG4A | 228.42 | 614 | 123 | 592 | 24.85 | 1.86 | 0.54 |
| 4B | LG4B | 169.56 | 1185 | 193 | 1156 | 8.57 | 0.88 | 1.14 |
| 4D | LG4D | 166.01 | 203 | 77 | 199 | 16.52 | 2.16 | 0.46 |
| 5A | LG5A.1 | 234.18 | 969 | 169 | 963 | 16.26 | 1.39 | 0.72 |
| LG5A.2 | 52.94 | 139 | 39 | 134 | 9.87 | 1.36 | 0.74 | |
| 5B | LG5B.1 | 68.44 | 682 | 88 | 675 | 8.09 | 0.78 | 1.29 |
| LG5B.2 | 172.4 | 538 | 164 | 529 | 15.43 | 1.05 | 0.95 | |
| 5D | LG5D.1 | 223.58 | 192 | 119 | 171 | 13.69 | 1.88 | 0.53 |
| LG5D.2 | 61.6 | 427 | 42 | 415 | 8.03 | 1.47 | 0.68 | |
| 6A | LG6A.1 | 112.71 | 154 | 50 | 137 | 20.91 | 2.25 | 0.44 |
| LG6A.2 | 54.95 | 161 | 36 | 151 | 17.46 | 1.53 | 0.66 | |
| 6B | LG6B | 167.65 | 852 | 188 | 783 | 7.7 | 0.89 | 1.12 |
| 6D | LG6D.1 | 31.08 | 34 | 7 | 34 | 10.4 | 4.44 | 0.23 |
| LG6D.2 | 170.88 | 506 | 124 | 497 | 12.41 | 1.38 | 0.73 | |
| 7A | LG7A.1 | 75.25 | 194 | 76 | 176 | 14.91 | 0.99 | 1.01 |
| LG7A.2 | 225.38 | 647 | 201 | 633 | 18.9 | 1.12 | 0.89 | |
| 7B | LG7B | 170.54 | 882 | 129 | 845 | 18.19 | 1.32 | 0.76 |
| 7D | LG7D.1 | 237.8 | 453 | 130 | 446 | 15.51 | 1.83 | 0.55 |
| LG7D.2 | 85.38 | 17 | 13 | 16 | 24.96 | 6.57 | 0.15 | |
| 1st homologous | 3 | 468.12 | 2016 | 272 | 1828 | 26.86 | 1.72 | 0.58 |
| 2nd homologous | 4 | 631.98 | 1865 | 372 | 1766 | 25.44 | 1.7 | 0.59 |
| 3rd homologous | 4 | 703.96 | 2846 | 524 | 2821 | 22.84 | 1.34 | 0.74 |
| 4th homologous | 3 | 563.99 | 2002 | 393 | 1947 | 24.85 | 1.44 | 0.7 |
| 5th homologous | 6 | 813.14 | 2947 | 621 | 2887 | 15.43 | 1.31 | 0.76 |
| 6th homologous | 4 | 537.27 | 1707 | 405 | 1602 | 20.91 | 1.33 | 0.75 |
| 7th homologous | 5 | 794.35 | 2193 | 549 | 2116 | 24.96 | 1.45 | 0.69 |
| A genome | 10 | 1703.69 | 6215 | 1231 | 6072 | 25.68 | 1.38 | 0.72 |
| B genome | 8 | 1298.23 | 5860 | 1197 | 5490 | 26.86 | 1.08 | 0.92 |
| D genome | 12 | 1510.87 | 3501 | 708 | 3405 | 25.44 | 2.13 | 0.47 |
| TOTAL | 30 | 4512.79 | 15576 | 3136 | 14967 | 26.44 | 1.44 | 0.69 |
| Trait | QTLs Name | Environment | Position | LOD | PVE (%) | Add | Left and Right Marker | Interval | Physical Interval | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| PH | Qph.nwafu-2D.1 | 19YL | 17 | 10.7 | 3.73 | −3.77 | AX-111561744/AX-179557748 | 16.5–20.5 | 23.416254/28.417456 | (Zhai et al., 2016) |
| PH | 20YL | 17 | 8.82 | 3.3 | −3.75 | AX-111561744/AX-179557748 | 16.5–20.5 | 23.416254/28.417456 | ||
| PH | Qph.nwafu-2D.2 | 19YL | 103 | 53.34 | 33.08 | 11.29 | AX-94570302/AX-109998182 | 102.5–103.5 | 413.778968/425.474614 | |
| PH | Qph.nwafu-4B | 19NY | 59 | 7.39 | 10.23 | 4.93 | AX-179477460/AX-110984065 | 58.5–59.5 | 30.805339/32.961929 | (Mohler et al., 2016) |
| PH | 20NY | 59 | 9.36 | 13.76 | 5.9 | AX-179477460/AX-110984065 | 58.5–59.5 | 30.805339/32.961929 | ||
| PH | 19YL | 59 | 23.33 | 9.32 | 6.28 | AX-179477460/AX-110984065 | 58.5–59.5 | 30.805339/32.961929 | ||
| PH | 20YL | 59 | 26.85 | 12.26 | 7.62 | AX-179477460/AX-110984065 | 58.5–59.5 | 30.805339/32.961929 | ||
| PH | Qph.nwafu-4D | 19NY | 62 | 17.27 | 27.08 | −7.69 | AX-86170701/AX-89445201 | 61.5–62.5 | 18.781207/19.459614 | (Zhang et al., 2013) |
| PH | 20NY | 62 | 16.78 | 27.09 | −7.93 | AX-86170701/AX-89445201 | 61.5–62.5 | 18.781207/19.459614 | ||
| PH | 19YL | 62 | 40.17 | 20.11 | −8.84 | AX-86170701/AX-89445201 | 61.5–62.5 | 18.781207/19.459614 | ||
| PH | 20YL | 62 | 42.21 | 23.71 | −10.16 | AX-86170701/AX-89445201 | 61.5–62.5 | 18.781207/19.459614 | ||
| PH | Qph.nwafu-5B | 20YL | 55 | 4.32 | 1.49 | 2.52 | AX-109908739/AX-86174612 | 54.5–55.5 | 422.122099/425.671678 | |
| PH | Qph.nwafu-5D | 20YL | 190 | 46.92 | 29.23 | −11.18 | AX-94390434/AX-110033637 | 189.5–190.5 | 466.230408/469.357817 | (Quarrie et al., 2005; Hai et al., 2008) |
| PH | Qph.nwafu-6B.1 | 20YL | 139 | 6.19 | 2.19 | 3.07 | AX-109987590/AX-86162252 | 137.5–139.5 | 687.177084/688.20385 | |
| PH | Qph.nwafu-6B.2 | 19YL | 160 | 4.15 | 1.36 | 2.27 | AX-110632551/AX-109509377 | 159.5–160.5 | 712.125253/711.370298 | |
| TGW | Qtgw.nwafu-2A | 20YL | 186 | 2.6 | 5.08 | 0.95 | AX-95103231/AX-94508212 | 185.5–186.5 | 733.854404/734.347961 | (Cui et al., 2014) |
| TGW | Qtgw.nwafu-2B | 20YL | 101 | 4.24 | 10.14 | 1.34 | AX-108905289/AX-95235626 | 95.5–106.5 | 153.585606/568.468886 | (Li et al., 2018) |
| TGW | Qtgw.nwafu-3A | 19YL | 133 | 2.7 | 4.14 | −0.89 | AX-179477407/AX-94457296 | 132.5–134.5 | 457.796943/431.074614 | |
| TGW | Qtgw.nwafu-4B | 20NY | 51 | 4.18 | 3.43 | 1.23 | AX-111494900/AX-94438527 | 48.5–53.5 | 25.847125/26.491497 | [67] |
| TGW | 19YL | 51 | 4.37 | 6.85 | 1.18 | AX-111494900/AX-94438527 | 49.5–53.5 | 25.847125/26.491497 | ||
| TGW | 19NY | 52 | 2.85 | 5.02 | 1.06 | AX-94438527/AX-110383634 | 48.5–55.5 | 26.491497/28.71668 | ||
| TGW | Qtgw.nwafu-4D.1 | 19NY | 60 | 5.87 | 9.73 | −1.44 | AX-89703298/AX-86170701 | 56.5–60.5 | 16.926631/18.781207 | (Mohler et al., 2016) |
| TGW | 20NY | 60 | 6.2 | 5.25 | −1.48 | AX-89703298/AX-86170701 | 56.5–60.5 | 16.926631/18.781207 | ||
| TGW | Qtgw.nwafu-4D.2 | 19YL | 111 | 3.54 | 5.55 | −1.03 | AX-111926032/AX-94818797 | 107.5–112.5 | 476.884228/477.371597 | |
| TGW | Qtgw.nwafu-5A | 19YL | 44 | 7.18 | 11.94 | 1.51 | AX-95510385/AX-95117188 | 43.5–45.5 | 698.508129/702.466804 | |
| TGW | 20YL | 44 | 3.51 | 6.99 | 1.11 | AX-95510385/AX-95117188 | 43.5–45.5 | 698.508129/702.466804 | ||
| TGW | Qtgw.nwafu-5D.1 | 19NY | 37 | 5.51 | 9.32 | 1.4 | AX-111543112/AX-110576074 | 34.5–38.5 | 38.070293/41.294446 | |
| TGW | 20NY | 37 | 14.79 | 14.24 | 2.43 | AX-111543112/AX-110576074 | 35.5–38.5 | 38.070293/41.294446 | ||
| TGW | Qtgw.nwafu-5D.2 | 20NY | 46 | 6.46 | 5.56 | −1.51 | AX-111019963/AX-110085499 | 44.5–49.5 | 42.928674/44.192407 | |
| TGW | Qtgw.nwafu-5D.3 | 19YL | 81 | 3.66 | 5.83 | 1.05 | AX-110867187/AX-108827297 | 79.5–81.5 | 369.202139/370.064947 | |
| TGW | Qtgw.nwafu-6A | 20YL | 29 | 3.06 | 6.26 | −1.06 | AX-109431286/AX-109358667 | 27.5–30.5 | 606.979733/608.046298 | (Cui et al., 2014) |
| GL | Qgl.nwafu-1A | 19YL | 150 | 3.3 | 3.11 | −0.06 | AX-95682344/AX-108726119 | 148.5–150.5 | 572.350803/572.658176 | (Mir et al., 2012) |
| GL | Qgl.nwafu-1B.1 | 20YL | 0 | 3.62 | 4.51 | 0.09 | AX-94835306/AX-179476279 | 0–0.5 | 59.471177/94.978091 | |
| GL | Qgl.nwafu-1B.2 | 19NY | 65 | 3.51 | 4.49 | 0.08 | AX-94650293/AX-112288501 | 64.5–66.5 | 640.845515/641.632325 | |
| GL | 19YL | 65 | 5.23 | 5.14 | 0.08 | AX-94650293/AX-112288501 | 64.5–65.5 | 640.845515/641.632325 | ||
| GL | Qgl.nwafu-3A | 20YL | 135 | 5.71 | 7.61 | −0.1 | AX-94426283/AX-110122062 | 134.5–136.5 | 511.755031/510.853056 | |
| GL | 20NY | 137 | 3.78 | 6.88 | −0.09 | AX-179557644/AX-94387510 | 136.5–137.5 | 541.482465/540.048345 | ||
| GL | Qgl.nwafu-4A | 20NY | 49 | 4.41 | 9.11 | 0.1 | AX-111251110/AX-179476673 | 46.5–53.5 | 407.389107/129.089816 | |
| GL | 19NY | 50 | 4.57 | 6.07 | 0.1 | AX-111251110/AX-179476673 | 47.5–52.5 | 407.389107/129.089816 | ||
| GL | Qgl.nwafu-4B.1 | 19YL | 51 | 4.28 | 4.17 | 0.07 | AX-179476673/AX-110173140 | 47.5–52.5 | 129.089816/140.310606 | |
| GL | Qgl.nwafu-4B.2 | 19YL | 68 | 5 | 4.94 | −0.08 | AX-109507847/AX-109427900 | 67.5–69.5 | 114.952789/161.548436 | (Wang et al., 2010) |
| GL | 20YL | 68 | 4.03 | 5.11 | −0.08 | AX-109507847/AX-109427900 | 67.5–69.5 | 114.952789/161.548436 | ||
| GL | Qgl.nwafu-4D | 19YL | 16 | 2.67 | 2.55 | 0.05 | AX-108892806/AX-109447997 | 15.5–18.5 | 6.598631/7.048661 | |
| GL | Qgl.nwafu-5A | 19NY | 44 | 6.93 | 9.59 | 0.12 | AX-95510385/AX-95117188 | 43.5–44.5 | 698.508129/698.508129 | |
| GL | 19YL | 44 | 6.47 | 6.72 | 0.09 | AX-95510385/AX-95117188 | 43.5–44.5 | 698.508129/700.34701 | ||
| GL | 20YL | 44 | 5.8 | 7.73 | 0.1 | AX-95510385/AX-95117188 | 43.5–45.5 | 698.508129/700.34701 | ||
| GL | Qgl.nwafu-5B | 19YL | 6 | 2.75 | 2.56 | −0.05 | AX-112288130/AX-95631525 | 5.5–6.5 | 6.654131/8.917454 | |
| GL | Qgl.nwafu-5D.1 | 20NY | 37 | 3.13 | 5.76 | 0.08 | AX-111543112/AX-110576074 | 33.5–39.5 | 38.070293/41.294446 | |
| GL | Qgl.nwafu-5D.2 | 19YL | 82 | 6.13 | 5.99 | 0.08 | AX-111496494/AX-109707913 | 81.5–84.5 | 370.135626/379.028214 | |
| GL | 20YL | 82 | 6 | 7.75 | 0.1 | AX-111496494/AX-109707913 | 81.5–84.5 | 370.135626/379.028214 | ||
| GL | 19NY | 89 | 4.36 | 5.62 | 0.09 | AX-110558491/AX-111903917 | 88.5–91.5 | 385.893875/386.126855 | ||
| GL | Qgl.nwafu-5D.3 | 20YL | 191 | 7.45 | 9.83 | 0.11 | AX-110033637/AX-110830424 | 190.5–191.5 | 469.357817/469.523881 | |
| GL | Qgl.nwafu-5D.4 | 19YL | 218 | 6.85 | 7.84 | 0.09 | AX-110777538/AX-111512534 | 215.5–221.5 | 485.909071/491.01105 | |
| GL | Qgl.nwafu-6B | 19YL | 162 | 8.79 | 8.95 | 0.1 | AX-110287286/AX-111572797 | 161.5–162.5 | 712.125253/712.245125 | (Li et al., 2018) |
| GL | 19NY | 167 | 7.88 | 10.73 | 0.13 | AX-89379712/AX-94499484 | 166.5–167 | 704.884934/718.376276 | ||
| GL | 20NY | 167 | 3.55 | 6.68 | 0.09 | AX-89379712/AX-94499484 | 166.5–167 | 704.884934/718.376276 | ||
| GL | 20YL | 167 | 7.73 | 10.33 | 0.12 | AX-89379712/AX-94499484 | 166.5–167 | 704.884934/718.376276 | ||
| GW | Qgw.nwafu-2B | 20YL | 94 | 3.86 | 6.13 | 0.04 | AX-109423066/AX-108990832 | 93.5–94.5 | 152.611396/153.128588 | |
| GW | Qgw.nwafu-2D | 19NY | 12 | 2.55 | 4.2 | 0.04 | AX-179477408/AX-111367738 | 11.5–12.5 | 20.768547/21.405473 | (Huang et al., 2006; Guan et al., 2018; Wu et al., 2015) |
| GW | Qgw.nwafu-3A | 19YL | 311 | 3.18 | 4.66 | 0.04 | AX-110915909/AX-110475339 | 308.5–311 | 746.360221/749.849798 | (Lee et al., 2014) |
| GW | Qgw.nwafu-4B.1 | 19NY | 51 | 4.15 | 6.95 | 0.05 | AX-111494900/AX-94438527 | 49.5–54.5 | 25.847125/26.491497 | |
| GW | 20NY | 51 | 3.5 | 6.85 | 0.05 | AX-111494900/AX-94438527 | 48.5–54.5 | 25.847125/26.491497 | ||
| GW | Qgw.nwafu-4B.2 | 20YL | 68 | 3.91 | 6.23 | 0.04 | AX-109507847/AX-109427900 | 67.5–68.5 | 114.952789/161.548436 | (Wang et al., 2010) |
| GW | Qgw.nwafu-4B.3 | 19YL | 77 | 9.23 | 15.24 | 0.07 | AX-179559104/AX-95658798 | 76.5–77.5 | 520.214474/523.447693 | |
| GW | Qgw.nwafu-4D | 20NY | 60 | 3.2 | 6.32 | −0.05 | AX-89703298/AX-86170701 | 59.5–61.5 | 16.926631/18.781207 | |
| GW | 19NY | 61 | 4.15 | 7.22 | −0.05 | AX-86170701/AX-110572006 | 59.5–61.5 | 18.781207/19.179341 | ||
| GW | 19YL | 63 | 7.93 | 12.12 | −0.06 | AX-86170701/AX-89445201 | 61.5–64.5 | 18.781207/19.459614 | ||
| GW | 20YL | 63 | 6.37 | 10.37 | −0.05 | AX-86170701/AX-89445201 | 61.5–63.5 | 18.781207/19.459614 | ||
| GW | Qgw.nwafu-5D | 19NY | 163 | 3.5 | 6.01 | 0.04 | AX-109317498/AX-109855976 | 159.5–166.5 | 448.686533/449.292436 | |
| GW | Qgw.nwafu-6D | 20NY | 4 | 2.51 | 5.8 | 0.05 | AX-111594857/AX-109406081 | 0–12.5 | 12.650045/8.255713 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lv, D.; Zhang, C.; Yv, R.; Yao, J.; Wu, J.; Song, X.; Jian, J.; Song, P.; Zhang, Z.; Han, D.; et al. Utilization of a Wheat50K SNP Microarray-Derived High-Density Genetic Map for QTL Mapping of Plant Height and Grain Traits in Wheat. Plants 2021, 10, 1167. https://doi.org/10.3390/plants10061167
Lv D, Zhang C, Yv R, Yao J, Wu J, Song X, Jian J, Song P, Zhang Z, Han D, et al. Utilization of a Wheat50K SNP Microarray-Derived High-Density Genetic Map for QTL Mapping of Plant Height and Grain Traits in Wheat. Plants. 2021; 10(6):1167. https://doi.org/10.3390/plants10061167
Chicago/Turabian StyleLv, Dongyun, Chuanliang Zhang, Rui Yv, Jianxin Yao, Jianhui Wu, Xiaopeng Song, Juntao Jian, Pengbo Song, Zeyuan Zhang, Dejun Han, and et al. 2021. "Utilization of a Wheat50K SNP Microarray-Derived High-Density Genetic Map for QTL Mapping of Plant Height and Grain Traits in Wheat" Plants 10, no. 6: 1167. https://doi.org/10.3390/plants10061167
APA StyleLv, D., Zhang, C., Yv, R., Yao, J., Wu, J., Song, X., Jian, J., Song, P., Zhang, Z., Han, D., & Sun, D. (2021). Utilization of a Wheat50K SNP Microarray-Derived High-Density Genetic Map for QTL Mapping of Plant Height and Grain Traits in Wheat. Plants, 10(6), 1167. https://doi.org/10.3390/plants10061167





