From Traditional Breeding to Genome Editing for Boosting Productivity of the Ancient Grain Tef [Eragrostis tef (Zucc.) Trotter]
Abstract
1. Introduction
2. Mechanisms of Tolerance to Lodging and Environmental Constraints in Tef
2.1. Lodging Tolerance
2.2. Drought Tolerance
2.3. Weed Competition and Herbicide Tolerance
2.4. Panicle Architecture
3. Status of Tef Improvement
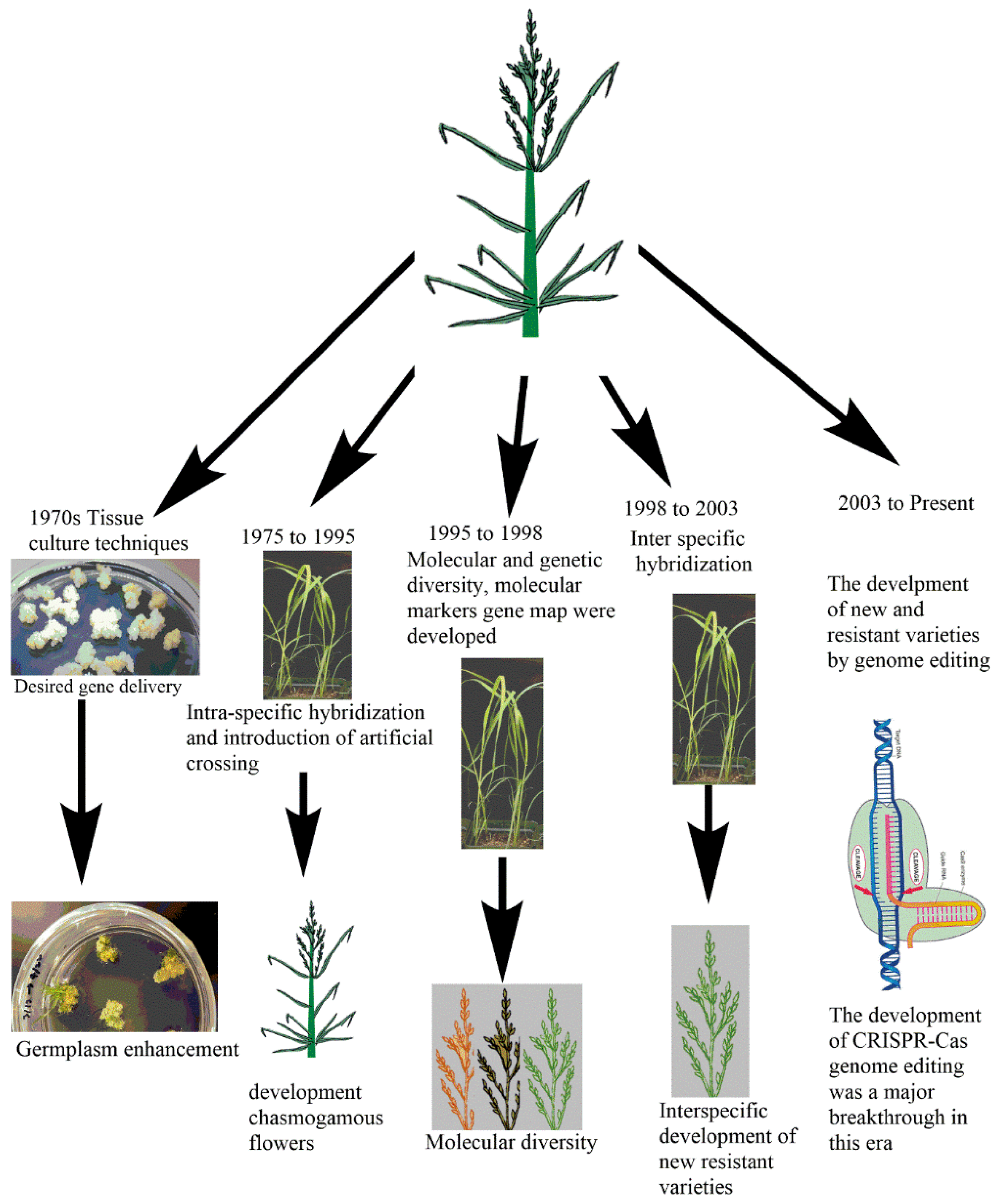
3.1. Traditional Breeding: Past and Current Status of Tef Improvement
3.2. Molecular Marker Development
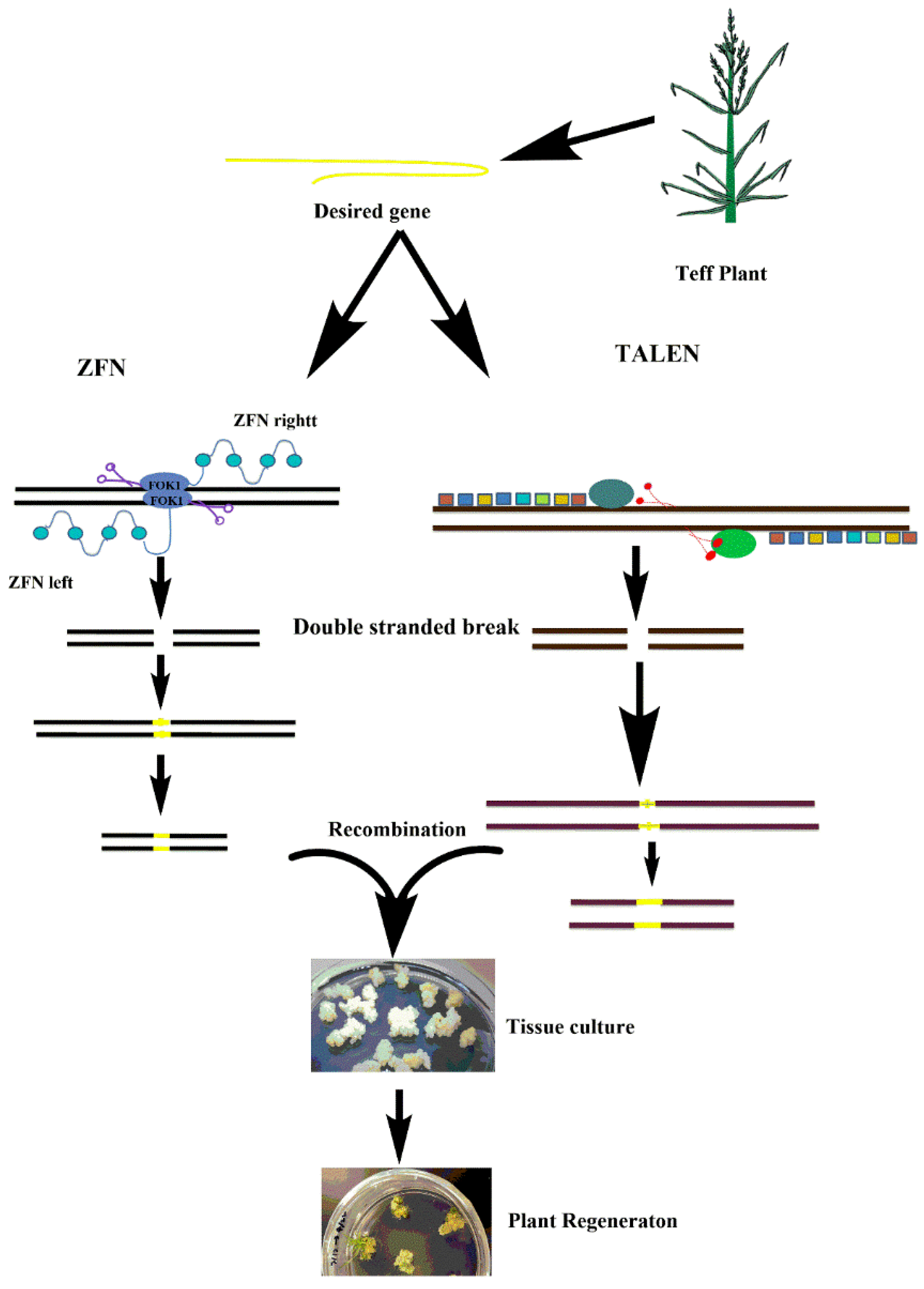
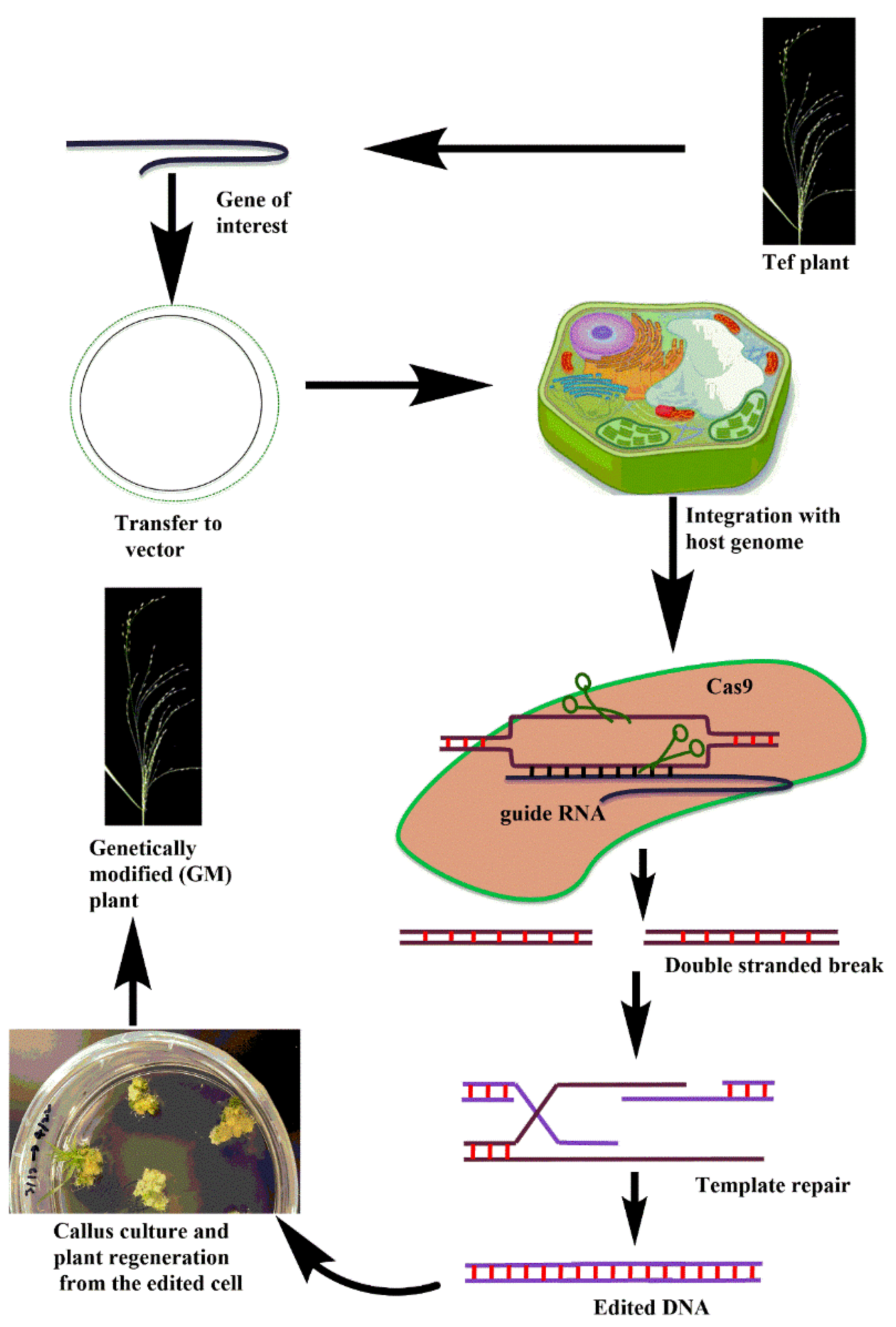
4. Potential of Genome Editing Technologies for Tef Improvement
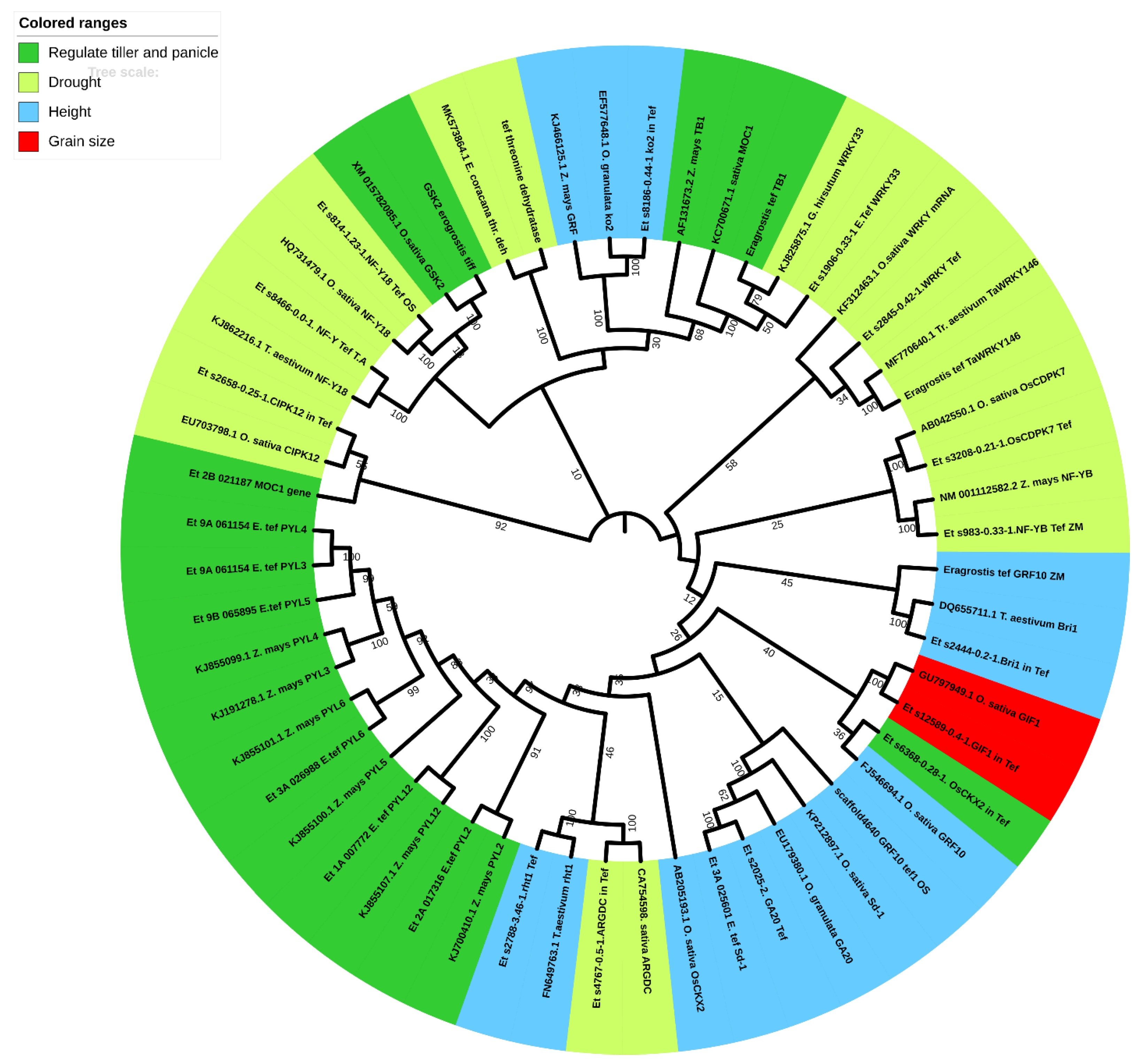
Candidate Tef Genes for CRISPR-Cas Technology
5. Constraints and Solutions Related to CRISPR-Cas Genome Editing
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cheng, A. Shaping a sustainable food future by rediscovering long-forgotten ancient grains. Plant Sci. 2018, 269, 136–142. [Google Scholar] [CrossRef]
- Ray, D.K.; Gerber, J.S.; MacDonald, G.K.; West, P.C. Climate variation explains a third of global crop yield variability. Nat. Commun. 2015, 6, 1–9. [Google Scholar] [CrossRef]
- Cochrane, L.; Bekele, Y.W. Average crop yield (2001–2017) in Ethiopia: Trends at national, regional and zonal levels. Data Brief 2018, 16, 1025. [Google Scholar] [CrossRef] [PubMed]
- CSA. Agricultural Sample Survey 2015/2016; CSA: Addis Ababa, Ethiopia, 2016. [Google Scholar]
- Tadele, Z.; Assefa, K. Increasing food production in Africa by boosting the productivity of understudied crops. Agronomy 2012, 2, 240–283. [Google Scholar] [CrossRef]
- El-Alfy, T.S.; Ezzat, S.M.; Sleem, A.A. Chemical and biological study of the seeds of Eragrostis tef (Zucc.) Trotter. Nat. Prod. Res. 2012, 26, 619–629. [Google Scholar] [CrossRef]
- Spaenij-Dekking, L.; Kooy-Winkelaar, Y.; Koning, F. The Ethiopian cereal tef in celiac disease. N. Engl. J. Med. 2005, 353, 1748–1749. [Google Scholar] [CrossRef]
- Saturni, L.; Ferretti, G.; Bacchetti, T. The gluten-free diet: Safety and nutritional quality. Nutrients 2010, 2, 16–34. [Google Scholar] [CrossRef]
- Gujral, N.; Freeman, H.J.; Thomson, A.B. Celiac disease: Prevalence, diagnosis, pathogenesis and treatment. World J. Gastroenterol. WJG 2012, 18, 6036. [Google Scholar] [CrossRef] [PubMed]
- Yami, A. 17. Tef Straw: A Valuable Feed Resource to Improve Animal Production and Productivity; Tef Improvement: Bern, Switzerland, 2013; Volume 233. [Google Scholar]
- Assefa, K.; Yu, J.K.; Zeid, M.; Belay, G.; Tefera, H.; Sorrells, M. Breeding tef [Eragrostis tef (Zucc.) Trotter]: Conventional and molecular approaches. Plant Breed. 2011, 130, 1–9. [Google Scholar] [CrossRef]
- Fang, X.; Li, Y.; Nie, J.; Wang, C.; Huang, K.; Zhang, Y.; Zhang, Y.; She, H.; Liu, X.; Ruan, R. Effects of nitrogen fertilizer and planting density on the leaf photosynthetic characteristics, agronomic traits and grain yield in common buckwheat (Fagopyrum esculentum M.). Field Crops Res. 2018, 219, 160–168. [Google Scholar] [CrossRef]
- Zhao, H.; Mo, Z.; Lin, Q.; Pan, S.; Duan, M.; Tian, H.; Wang, S.; Tang, X. Relationships between grain yield and agronomic traits of rice in southern China. Chil. J. Agric. Res. 2020, 80, 72–79. [Google Scholar] [CrossRef]
- Belete, T.; Mathewos, T.; Daba, G. Correlation of yield and yield related traits of Tef (Eragrostis tef (Zucc.) Trotter) varieties in Ethiopia. J. Genet. Environ. Resour. Conserv. 2020, 8, 35–39. [Google Scholar]
- Tilahun, Z.M. Effect of row spacing and nitrogen fertilizer levels on yield and yield components of rice varieties. World Sci. News 2019, 116, 180–193. [Google Scholar]
- Arefaine, A.; Adhanom, D.; Tekeste, N. Response of Teff (Eragrostis tef (Zucc.) Trotter) to Seeding Rate and Methods of Sowing on Yield and Yield Attributes in a Subhumid Environment, Northern Ethiopia. Int. J. Agron. 2020, 2020, 1–7. [Google Scholar] [CrossRef]
- Haftamu, G.; Mitiku, H.; Yamoah, C. Tillage frequency, soil compaction and N-fertilizer rate effects on yield of teff (Eragrostis tef (Zucc.) Trotter) in central zone of Tigray, Northern Ethiopia. Momona Ethiop. J. Sci. 2009, 1, 82–94. [Google Scholar]
- Menalled, F.D. Sustainable Agriculture and Integrated Weed Management. In Weed Control: Sustainability, Hazards, and Risks in Cropping Systems Worldwide; CRC Press: Boca Raton, FL, USA, 2018; Volume 3. [Google Scholar]
- Abrha, B.; Tsegay, A.; Gebrehiwot, K. Economic analysis of tef (Eragrostis tef (zucc.) trotter) yield in response to soil water, weed and fertilizer managements in the northern highlands of Ethiopia. J. Drylands 2017, 2, 675–682. [Google Scholar]
- Rockström, J.; Falkenmark, M. Agriculture: Increase water harvesting in Africa. Nat. News 2015, 519, 283. [Google Scholar] [CrossRef]
- Matsuura, A.; Tsuji, W.; An, P.; Inanaga, S.; Murata, K. Effect of pre-and post-heading water deficit on growth and grain yield of four millets. Plant Prod. Sci. 2012, 15, 323–331. [Google Scholar] [CrossRef]
- Winkel, T.; Renno, J.-F.; Payne, W. Effect of the timing of water deficit on growth, phenology and yield of pearl millet (Pennisetum glaucum (L.) R. Br.) grown in Sahelian conditions. J. Exp. Bot. 1997, 48, 1001–1009. [Google Scholar] [CrossRef]
- Abraha, M.T.; Hussein, S.; Laing, M.; Assefa, K. Genetic management of drought in tef: Current status and future research directions. Glob. J. Crop Soil Sci. Plant Breed. 2015, 3, 156–161. [Google Scholar]
- Assefa, K.; Chanyalew, S.; Tadele, Z. Tef, Eragrostis tef (Zucc.) Trotter. Millets and Sorghum: Biology and Genetic Improvement. 2017, pp. 226–266. Available online: https://www.researchgate.net/publication/312353091_Tef_Eragrostis_tef_Zucc_Trotter_Biology_and_Genetic_Improvement (accessed on 19 March 2021).
- Blösch, R.; Plaza-Wüthrich, S.; de Reuille, P.B.; Weichert, A.; Routier-Kierzkowska, A.-L.; Cannarozzi, G.; Robinson, S.; Tadele, Z. Panicle angle is an important factor in tef lodging tolerance. Front. Plant Sci. 2020, 11, 1–12. [Google Scholar] [CrossRef]
- Van Delden, S.; Vos, J.; Ennos, A.; Stomph, T. Analysing lodging of the panicle bearing cereal teff (Eragrostis tef). New Phytol. 2010, 186, 696–707. [Google Scholar] [CrossRef] [PubMed]
- Ketema, S. Tef (Eragrostis Tef) Breeding, Genetic Resources, Agronomy, Utilization and Role in Ethiopian Agriculture; Institute of Agricultural Research: Zaria, Nigeria, 1993. [Google Scholar]
- Jifar, H.; Tesfaye, K.; Assefa, K.; Chanyalew, S.; Tadele, Z. Semi-dwarf tef lines for high seed yield and lodging tolerance in Central Ethiopia. Afr. Crop Sci. J. 2017, 25, 419–439. [Google Scholar] [CrossRef]
- Paff, K.; Asseng, S. A review of tef physiology for developing a tef crop model. Eur. J. Agron. 2018, 94, 54–66. [Google Scholar] [CrossRef]
- Debbarma, J.; Sarki, Y.N.; Saikia, B.; Boruah, H.P.D.; Singha, D.L.; Chikkaputtaiah, C. Ethylene response factor (ERF) family proteins in abiotic stresses and CRISPR–Cas9 genome editing of ERFs for multiple abiotic stress tolerance in crop plants: A review. Mol. Biotechnol. 2019, 61, 153–172. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, Y.K.; Yadav, S.K. High temperature stress tolerance in maize (Zea mays L.): Physiological and molecular mechanisms. J. Plant Biol. 2019, 62, 93–102. [Google Scholar] [CrossRef]
- Zhou, H.; Liu, B.; Weeks, D.P.; Spalding, M.H.; Yang, B. Large chromosomal deletions and heritable small genetic changes induced by CRISPR/Cas9 in rice. Nucleic Acids Res. 2014, 42, 10903–10914. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Zhou, H.; Bi, H.; Fromm, M.; Yang, B.; Weeks, D.P. Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res. 2013, 41, e188. [Google Scholar] [CrossRef]
- Saika, H.; Mori, A.; Endo, M.; Toki, S. Targeted deletion of rice retrotransposon Tos17 via CRISPR/Cas9. Plant Cell Rep. 2019, 38, 455–458. [Google Scholar] [CrossRef]
- Lee, K.; Eggenberger, A.L.; Banakar, R.; McCaw, M.E.; Zhu, H.; Main, M.; Kang, M.; Gelvin, S.B.; Wang, K. CRISPR/Cas9-mediated targeted T-DNA integration in rice. Plant Mol. Biol. 2019, 99, 317–328. [Google Scholar] [CrossRef]
- Dayani, S.; Sabzalian, M.R.; Mazaheri-Tirani, M. CRISPR/Cas9 Genome Editing in Bread Wheat (Triticum aestivum L.) Genetic Improvement. In Advances in Plant Breeding Strategies: Cereals; Springer: Berlin/Heidelberg, Germany, 2019; pp. 453–469. [Google Scholar]
- Jaganathan, D.; Ramasamy, K.; Sellamuthu, G.; Jayabalan, S.; Venkataraman, G. CRISPR for crop improvement: An update review. Front. Plant Sci. 2018, 9, 985. [Google Scholar] [CrossRef] [PubMed]
- Basso, M.F.; Ferreira, P.C.G.; Kobayashi, A.K.; Harmon, F.G.; Nepomuceno, A.L.; Molinari, H.B.C.; Grossi-de-Sa, M.F. Micro RNA s and new biotechnological tools for its modulation and improving stress tolerance in plants. Plant Biotechnol. J. 2019, 17, 1482–1500. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Gao, H.; Wang, H.; Lafitte, H.R.; Archibald, R.L.; Yang, M.; Hakimi, S.M.; Mo, H.; Habben, J.E. ARGOS 8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol. J. 2017, 15, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.; Que, Z.; Xia, Y.; Tang, N.; Li, D.; He, R.; Cao, M. Knock out of the annexin gene OsAnn3 via CRISPR/Cas9-mediated genome editing decreased cold tolerance in rice. J. Plant Biol. 2017, 60, 539–547. [Google Scholar] [CrossRef]
- Zhang, A.; Liu, Y.; Wang, F.; Li, T.; Chen, Z.; Kong, D.; Bi, J.; Zhang, F.; Luo, X.; Wang, J. Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol. Breed. 2019, 39, 47. [Google Scholar] [CrossRef] [PubMed]
- Sadanandom, A.; Srivastava, A.K.; Zhang, C. Targeted mutagenesis of the SUMO protease, Overly Tolerant to Salt1 in rice through CRISPR/Cas9-mediated genome editing reveals a major role of this SUMO protease in salt tolerance. BioRxiv 2019. [Google Scholar] [CrossRef]
- Zhang, Y.; Liang, Z.; Zong, Y.; Wang, Y.; Liu, J.; Chen, K.; Qiu, J.-L.; Gao, C. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 2016, 7, 1–8. [Google Scholar] [CrossRef]
- Wang, W.; Akhunova, A.; Chao, S.; Akhunov, E. Optimizing multiplex CRISPR/Cas9-based genome editing for wheat. BioRxiv 2016. [Google Scholar] [CrossRef]
- Howells, R.M.; Craze, M.; Bowden, S.; Wallington, E.J. Efficient generation of stable, heritable gene edits in wheat using CRISPR/Cas9. BMC Plant Biol. 2018, 18, 215. [Google Scholar] [CrossRef] [PubMed]
- Berry, P.; Sterling, M.; Spink, J.; Baker, C.; Sylvester-Bradley, R.; Mooney, S.; Tams, A.; Ennos, A. Understanding and reducing lodging in cereals. Adv. Agron. 2004, 84, 215–269. [Google Scholar]
- Dagnaw, H. Influence of Nitrogen Fertilizer Rates and Varieties on Grain Yield, Grain Nutrition and Injera Sensory Quality of Tef [Eragrostis tef (Zucc.) Trotter] Varieties. Master’s Thesis, Adis Ababa University, Addis Ababa, Ethiopia, 2018. [Google Scholar]
- Sterling, M.; Baker, C.; Berry, P.; Wade, A. An experimental investigation of the lodging of wheat. Agric. For. Meteorol. 2003, 119, 149–165. [Google Scholar] [CrossRef]
- Merchuk-Ovnat, L.; Bimro, J.; Yaakov, N.; Kutsher, Y.; Amir-Segev, O.; Reuveni, M. In-Depth Field Characterization of Teff [Eragrostis tef (Zucc.) Trotter] Variation: From Agronomic to Sensory Traits. Agronomy 2020, 10, 1107. [Google Scholar] [CrossRef]
- Assefa, K.; Cannarozzi, G.; Girma, D.; Kamies, R.; Chanyalew, S.; Plaza-Wüthrich, S.; Blösch, R.; Rindisbacher, A.; Rafudeen, S.; Tadele, Z. Genetic diversity in tef [Eragrostis tef (Zucc.) Trotter]. Front. Plant Sci. 2015, 6, 177. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Ma, B.L. A new method for assessing plant lodging and the impact of management options on lodging in canola crop production. Sci. Rep. 2016, 6, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Mariani, L.; Ferrante, A. Agronomic management for enhancing plant tolerance to abiotic stresses—Drought, salinity, hypoxia, and lodging. Horticulturae 2017, 3, 52. [Google Scholar] [CrossRef]
- Epstein, E. The anomaly of silicon in plant biology. Proc. Natl. Acad. Sci. USA 1994, 91, 11–17. [Google Scholar] [CrossRef]
- Ma, J.F. Role of silicon in enhancing the resistance of plants to biotic and abiotic stresses. Soil Sci. Plant Nutr. 2004, 50, 11–18. [Google Scholar] [CrossRef]
- De Carvalho, D.D.; Costa, F.T.; Duran, N.; Haun, M. Cytotoxic activity of violacein in human colon cancer cells. Toxicol. In Vitro 2006, 20, 1514–1521. [Google Scholar] [CrossRef]
- Liang, Y.; Nikolic, M.; Bélanger, R.; Gong, H.; Song, A. Silicon in Agriculture; Springer: Dordrecht, The Netherlands, 2015; Volume 10, pp. 978–994. [Google Scholar]
- Ligaba-Osena, A.; Guo, W.; Choi, S.C.; Limmer, M.A.; Seyfferth, A.L.; Hankoua, B.B. Silicon enhances biomass and grain yield in an ancient crop tef [Eragrostis tef (Zucc.) Trotter]. Front. Plant Sci. 2020, 11, 608503. [Google Scholar] [CrossRef]
- Deshmukh, R.K.; Ma, J.F.; Bélanger, R.R. Role of silicon in plants. Front. Plant Sci. 2017, 8, 1858. [Google Scholar] [CrossRef]
- Jöst, M.; Esfeld, K.; Burian, A.; Cannarozzi, G.; Chanyalew, S.; Kuhlemeier, C.; Assefa, K.; Tadele, Z. Semi-dwarfism and lodging tolerance in tef (Eragrostis tef) is linked to a mutation in the α-Tubulin 1 gene. J. Exp. Bot. 2015, 66, 933–944. [Google Scholar] [CrossRef]
- Cannarozzi, G.; Chanyalew, S.; Assefa, K.; Bekele, A.; Blösch, R.; Weichert, A.; Klauser, D.; Plaza-Wüthrich, S.; Esfeld, K.; Jöst, M. Technology generation to dissemination: Lessons learned from the tef improvement project. Euphytica 2018, 214, 1–20. [Google Scholar] [CrossRef]
- Rademacher, W. Growth retardants: Effects on gibberellin biosynthesis and other metabolic pathways. Annu. Rev. Plant Biol. 2000, 51, 501–531. [Google Scholar] [CrossRef] [PubMed]
- Sanvicente, P.; Lazarevitch, S.; Blouet, A.; Guckert, A. Morphological and anatomical modifications in winter barley culm after late plant growth regulator treatment. Eur. J. Agron. 1999, 11, 45–51. [Google Scholar] [CrossRef]
- Hedden, P. The genes of the Green Revolution. Trends Genet. 2003, 19, 5–9. [Google Scholar] [CrossRef]
- Itoh, H.; Ueguchi-Tanaka, M.; Sentoku, N.; Kitano, H.; Matsuoka, M.; Kobayashi, M. Cloning and functional analysis of two gibberellin 3β-hydroxylase genes that are differently expressed during the growth of rice. Proc. Natl. Acad. Sci. USA 2001, 98, 8909–8914. [Google Scholar] [CrossRef] [PubMed]
- Monna, L.; Kitazawa, N.; Yoshino, R.; Suzuki, J.; Masuda, H.; Maehara, Y.; Tanji, M.; Sato, M.; Nasu, S.; Minobe, Y. Positional cloning of rice semidwarfing gene, sd-1: Rice “green revolution gene” encodes a mutant enzyme involved in gibberellin synthesis. DNA Res. 2002, 9, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, A.; Ashikari, M.; Ueguchi-Tanaka, M.; Itoh, H.; Nishimura, A.; Swapan, D.; Ishiyama, K.; Saito, T.; Kobayashi, M.; Khush, G.S. Green revolution: A mutant gibberellin-synthesis gene in rice. Nature 2002, 416, 701. [Google Scholar] [CrossRef]
- Hong, Z.; Ueguchi-Tanaka, M.; Umemura, K.; Uozu, S.; Fujioka, S.; Takatsuto, S.; Yoshida, S.; Ashikari, M.; Kitano, H.; Matsuoka, M. A rice brassinosteroid-deficient mutant, ebisu dwarf (d2), is caused by a loss of function of a new member of cytochrome P450. Plant Cell 2003, 15, 2900–2910. [Google Scholar] [CrossRef] [PubMed]
- Multani, D.S.; Briggs, S.P.; Chamberlin, M.A.; Blakeslee, J.J.; Murphy, A.S.; Johal, G.S. Loss of an MDR transporter in compact stalks of maize br2 and sorghum dw3 mutants. Science 2003, 302, 81–84. [Google Scholar] [CrossRef]
- Asano, K.; Hirano, K.; Ueguchi-Tanaka, M.; Angeles-Shim, R.B.; Komura, T.; Satoh, H.; Kitano, H.; Matsuoka, M.; Ashikari, M. Isolation and characterization of dominant dwarf mutants, Slr1-d, in rice. Mol. Genet. Genom. 2009, 281, 223–231. [Google Scholar] [CrossRef]
- Spielmeyer, W.; Ellis, M.H.; Chandler, P.M. Semidwarf (sd-1),“green revolution” rice, contains a defective gibberellin 20-oxidase gene. Proc. Natl. Acad. Sci. USA 2002, 99, 9043–9048. [Google Scholar] [CrossRef] [PubMed]
- Muangprom, A.; Thomas, S.G.; Sun, T.-p.; Osborn, T.C. A novel dwarfing mutation in a green revolution gene from Brassica rapa. Plant Physiol. 2005, 137, 931–938. [Google Scholar] [CrossRef]
- Peng, J.; Richards, D.E.; Hartley, N.M.; Murphy, G.P.; Devos, K.M.; Flintham, J.E.; Beales, J.; Fish, L.J.; Worland, A.J.; Pelica, F. ‘Green revolution’genes encode mutant gibberellin response modulators. Nature 1999, 400, 256. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, L.; Du, Y.; Yang, Z.; Condon, A.G.; Hu, Y.-G. Genetic effect of dwarfing gene Rht13 compared with Rht-D1b on plant height and some agronomic traits in common wheat (Triticum aestivum L.). Field Crops Res. 2014, 162, 39–47. [Google Scholar] [CrossRef]
- Itoh, H.; Tatsumi, T.; Sakamoto, T.; Otomo, K.; Toyomasu, T.; Kitano, H.; Ashikari, M.; Ichihara, S.; Matsuoka, M. A rice semi-dwarf gene, Tan-Ginbozu (D35), encodes the gibberellin biosynthesis enzyme, ent-kaurene oxidase. Plant Mol. Biol. 2004, 54, 533–547. [Google Scholar] [CrossRef]
- Hilley, J.L.; Weers, B.D.; Truong, S.K.; McCormick, R.F.; Mattison, A.J.; McKinley, B.A.; Morishige, D.T.; Mullet, J.E. Sorghum Dw2 encodes a protein kinase regulator of stem internode length. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, M.; Fujimoto, H.; Hirano, K.; Araki-Nakamura, S.; Ohmae-Shinohara, K.; Fujii, A.; Tsunashima, M.; Song, X.J.; Ito, Y.; Nagae, R. Sorghum Dw1, an agronomically important gene for lodging resistance, encodes a novel protein involved in cell proliferation. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Mickelbart, M.V.; Hasegawa, P.M.; Bailey-Serres, J. Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat. Rev. Genet. 2015, 16, 237–251. [Google Scholar] [CrossRef] [PubMed]
- Duque, A.S.; de Almeida, A.M.; da Silva, A.B.; da Silva, J.M.; Farinha, A.P.; Santos, D.; Fevereiro, P.; de Sousa Araújo, S. Abiotic stress responses in plants: Unraveling the complexity of genes and networks to survive. In Abiotic Stress-Plant Responses and Applications in Agriculture; IntechOpen: London, UK, 2013; pp. 49–101. [Google Scholar] [CrossRef]
- Ayele, M.; Blum, A.; Nguyen, H.T. Diversity for osmotic adjustment and root depth in tef [Eragrostis tef (Zucc.) Trotter]. Euphytica 2001, 121, 237–249. [Google Scholar] [CrossRef]
- Araya, A.; Stroosnijder, L.; Girmay, G.; Keesstra, S. Crop coefficient, yield response to water stress and water productivity of teff (Eragrostis tef (Zucc.). Agric. Water Manag. 2011, 98, 775–783. [Google Scholar] [CrossRef]
- Plaza-Wüthrich, S.; Blösch, R.; Rindisbacher, A.; Cannarozzi, G.; Tadele, Z. Gibberellin deficiency confers both lodging and drought tolerance in small cereals. Front. Plant Sci. 2016, 7, 643. [Google Scholar] [CrossRef]
- Kusaka, M.; Lalusin, A.G.; Fujimura, T. The maintenance of growth and turgor in pearl millet (Pennisetum glaucum [L.] Leeke) cultivars with different root structures and osmo-regulation under drought stress. Plant Sci. 2005, 168, 1–14. [Google Scholar] [CrossRef]
- Farooq, M.; Wahid, A.; Kobayashi, N.; Fujita, D.; Basra, S. Plant drought stress: Effects, mechanisms and management. In Sustainable Agriculture; Springer: Berlin/Heidelberg, Germany, 2009; pp. 153–188. [Google Scholar]
- Merrill, S.D.; Tanaka, D.L.; Hanson, J.D. Root length growth of eight crop species in Haplustoll soils. Soil Sci. Soc. Am. J. 2002, 66, 913–923. [Google Scholar] [CrossRef]
- Renton, M.; Chauhan, B.S. Modelling crop-weed competition: Why, what, how and what lies ahead? Crop Prot. 2017, 95, 101–108. [Google Scholar] [CrossRef]
- Andrew, I.; Storkey, J.; Sparkes, D. A review of the potential for competitive cereal cultivars as a tool in integrated weed management. Weed Res. 2015, 55, 239–248. [Google Scholar] [CrossRef]
- Hansen, P.K.; Kristensen, K.; Willas, J. A weed suppressive index for spring barley (Hordeum vulgare) varieties. Weed Res. 2008, 48, 225–236. [Google Scholar] [CrossRef]
- Gebrehiwot, H.G.; Aune, J.B.; Netland, J.; Eklo, O.M.; Torp, T.; Brandsæter, L.O. Weed-Competitive Ability of Teff (Eragrostis tef (Zucc.) Trotter) Varieties. Agronomy 2020, 10, 108. [Google Scholar] [CrossRef]
- Laizer, H.C.; Chacha, M.N.; Ndakidemi, P.A. Farmers’ knowledge, perceptions and practices in managing weeds and insect pests of common bean in Northern Tanzania. Sustainability 2019, 11, 4076. [Google Scholar] [CrossRef]
- Dezfulian, M.H.; Foreman, C.; Jalili, E.; Pal, M.; Dhaliwal, R.K.; Roberto, D.K.A.; Imre, K.M.; Kohalmi, S.E.; Crosby, W.L. Acetolactate synthase regulatory subunits play divergent and overlapping roles in branched-chain amino acid synthesis and Arabidopsis development. BMC Plant Biol. 2017, 17, 71. [Google Scholar] [CrossRef]
- Singh, S.; Kumar, V.; Dhanjal, D.S.; Singh, J. Herbicides and Plant Growth Regulators: Current Developments and Future Challenges. In Natural Bioactive Products in Sustainable Agriculture; Singh, J., Yadav, A.N., Eds.; Springer: Singapore, 2020. [Google Scholar] [CrossRef]
- Küpper, A.; Borgato, E.A.; Patterson, E.L.; Netto, A.G.; Nicolai, M.; Carvalho, S.J.P.d.; Nissen, S.J.; Gaines, T.A.; Christoffoleti, P.J. Multiple Resistance to Glyphosate and Acetolactate Synthase Inhibitors in Palmer Amaranth (Amaranthus palmeri) Identified in Brazil. Weed Sci. 2017, 65, 317–326. [Google Scholar] [CrossRef]
- Xue, W.; Xing, Y.; Weng, X.; Zhao, Y.; Tang, W.; Wang, L.; Zhou, H.; Yu, S.; Xu, C.; Li, X. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 2008, 40, 761–767. [Google Scholar] [CrossRef]
- Guillen-Portal, F.R.; Stougaard, R.N.; Xue, Q.; Eskridge, K.M. Compensatory mechanisms associated with the effect of spring wheat seed size on wild oat competition. Crop Sci. 2006, 46, 935–945. [Google Scholar] [CrossRef]
- Gupta, P.K.; Rustgi, S.; Kumar, N. Genetic and molecular basis of grain size and grain number and its relevance to grain productivity in higher plants. Genome 2006, 49, 565–571. [Google Scholar] [CrossRef]
- Huang, M.; Zou, Y.-B.; Jiang, P.; Bing, X.; Md, I.; Ao, H.-J. Relationship between grain yield and yield components in super hybrid rice. Agric. Sci. China 2011, 10, 1537–1544. [Google Scholar] [CrossRef]
- Yao, F.; Huang, J.; Cui, K.; Nie, L.; Xiang, J.; Liu, X.; Wu, W.; Chen, M.; Peng, S. Agronomic performance of high-yielding rice variety grown under alternate wetting and drying irrigation. Field Crops Res. 2012, 126, 16–22. [Google Scholar] [CrossRef]
- Terao, T.; Nagata, K.; Morino, K.; Hirose, T. A gene controlling the number of primary rachis branches also controls the vascular bundle formation and hence is responsible to increase the harvest index and grain yield in rice. Theor. Appl. Genet. 2010, 120, 875–893. [Google Scholar] [CrossRef]
- Zhang, Y.-C.; Yu, Y.; Wang, C.-Y.; Li, Z.-Y.; Liu, Q.; Xu, J.; Liao, J.-Y.; Wang, X.-J.; Qu, L.-H.; Chen, F. Overexpression of microRNA OsmiR397 improves rice yield by increasing grain size and promoting panicle branching. Nat. Biotechnol. 2013, 31, 848–852. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Qian, Q.; Liu, Z.; Sun, H.; He, S.; Luo, D.; Xia, G.; Chu, C.; Li, J.; Fu, X. Natural variation at the DEP1 locus enhances grain yield in rice. Nat. Genet. 2009, 41, 494–497. [Google Scholar] [CrossRef]
- Miura, K.; Ikeda, M.; Matsubara, A.; Song, X.-J.; Ito, M.; Asano, K.; Matsuoka, M.; Kitano, H.; Ashikari, M. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 2010, 42, 545–549. [Google Scholar] [CrossRef]
- Assefa, K.; Chanyalew, S.; Metaferia, G. Conventional and Molecular Tef Breeding; Tef Improvement: Bern, Switzerland, 2011; Volume 33. [Google Scholar]
- Tadele, Z.; Ferede Haile, B.; Abreha, E.; Assefa, K.; Chanyalew, S.; Mekbib, F. Morpho-Physiologic, Genotype X Environment Interaction and In Vitro Evaluation for Drought Tolerance in Tef Eragrostis tef (Zucc.) Trotter, Ethiopia; Haramaya University: Haromaya, Ethiopia, 2018. [Google Scholar]
- Berehe, T. Breakthrough in tef breeding technique. FAO Inf. Bull. Cerealimprovement Prod. Near East Proj. 1975, 3, 11–23. [Google Scholar]
- Belay, G.; Tefera, H.; Getachew, A.; Assefa, K.; Metaferia, G. Highly client-oriented breeding with farmer participation in the Ethiopian cereal tef [Eragrostis tef (Zucc.) Trotter]. Afr. J. Agric. Res. 2008, 3, 22–28. [Google Scholar]
- Belay, G.; Tefera, H.; Tadesse, B.; Metaferia, G.; Jarra, D.; Tadesse, T. Participatory variety selection in the Ethiopian cereal tef (Eragrostis tef). Exp. Agric. 2006, 42, 91–101. [Google Scholar] [CrossRef]
- Chanyalew, S.; Assefa, K.; Tadele, Z. Tef [Eragrostis tef (Zucc.) Trotter] Breeding. In Advances in Plant Breeding Strategies: Cereals; Springer: Berlin/Heidelberg, Germany, 2019; pp. 373–403. [Google Scholar]
- Ibitoye, D.; Akin-Idowu, P. Marker-assisted-selection (MAS): A fast track to increase genetic gain in horticultural crop breeding. Afr. J. Biotechnol. 2010, 9, 8889–8895. [Google Scholar]
- Jiang, G.-L. Molecular markers and marker-assisted breeding in plants. Plant Breed. Lab. Fields 2013, 45–83. [Google Scholar] [CrossRef]
- Yu, J.-K.; Sun, Q.; Rota, M.L.; Edwards, H.; Tefera, H.; Sorrells, M.E. Expressed sequence tag analysis in tef (Eragrostis tef (Zucc.) Trotter). Genome 2006, 49, 365–372. [Google Scholar] [CrossRef]
- Bai, G.; Ayele, M.; Tefera, H.; Nguyen, H.T. Amplified fragment length polymorphism analysis of tef [Eragrostis tef (Zucc.) Trotter]. Crop Sci. 1999, 39, 819–824. [Google Scholar] [CrossRef]
- Abraha, M.T.; Shimelis, H.; Laing, M.; Assefa, K.; Amelework, B. Assessment of the genetic relationship of tef (Eragrostis tef) genotypes using SSR markers. S. Afr. J. Bot. 2016, 105, 106–110. [Google Scholar] [CrossRef]
- Ayele, M.; Nguyen, H. Evaluation of amplified fragment length polymorphism markers in tef, Eragrostis tef (Zucc.) Trotter, and related species. Plant Breed. 2000, 119, 403–409. [Google Scholar] [CrossRef]
- Kashtwari, M.; Wani, A.A.; Rather, R.N. TILLING: An alternative path for crop improvement. J. Crop Improv. 2019, 33, 83–109. [Google Scholar] [CrossRef]
- Tadele, Z. Orphan crops: Their importance and the urgency of improvement. Planta 2019, 250, 677–694. [Google Scholar] [CrossRef]
- Voytas, D.F. Plant Genome Engineering with Sequence-Specific Nucleases. Annu. Rev. Plant Biol. 2013, 64, 327–350. [Google Scholar] [CrossRef]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A. Multiplex genome engineering using CRISPR/Cas systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef]
- Feng, Z.; Zhang, B.; Ding, W.; Liu, X.; Yang, D.-L.; Wei, P.; Cao, F.; Zhu, S.; Zhang, F.; Mao, Y. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 2013, 23, 1229. [Google Scholar] [CrossRef]
- Shan, Q.; Wang, Y.; Li, J.; Zhang, Y.; Chen, K.; Liang, Z.; Zhang, K.; Liu, J.; Xi, J.J.; Qiu, J.-L. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 2013, 31, 686. [Google Scholar] [CrossRef] [PubMed]
- Christian, M.; Cermak, T.; Doyle, E.L.; Schmidt, C.; Zhang, F.; Hummel, A.; Bogdanove, A.J.; Voytas, D.F. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 2010, 186, 757–761. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, F.; Li, X.; Baller, J.A.; Qi, Y.; Starker, C.G.; Bogdanove, A.J.; Voytas, D.F. Transcription activator-like effector nucleases enable efficient plant genome engineering. Plant Physiol. 2013, 161, 20–27. [Google Scholar] [CrossRef] [PubMed]
- Shan, Q.; Wang, Y.; Chen, K.; Liang, Z.; Li, J.; Zhang, Y.; Zhang, K.; Liu, J.; Voytas, D.F.; Zheng, X. Rapid and efficient gene modification in rice and Brachypodium using TALENs. Mol. Plant 2013, 6, 1365–1368. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Maeder, M.L.; Unger-Wallace, E.; Hoshaw, J.P.; Reyon, D.; Christian, M.; Li, X.; Pierick, C.J.; Dobbs, D.; Peterson, T. High frequency targeted mutagenesis in Arabidopsis thaliana using zinc finger nucleases. Proc. Natl. Acad. Sci. USA 2010, 107, 12028–12033. [Google Scholar] [CrossRef]
- Sander, J.D.; Dahlborg, E.J.; Goodwin, M.J.; Cade, L.; Zhang, F.; Cifuentes, D.; Curtin, S.J.; Blackburn, J.S.; Thibodeau-Beganny, S.; Qi, Y. Selection-free zinc-finger-nuclease engineering by context-dependent assembly (CoDA). Nat. Methods 2011, 8, 67. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Hong, X.; Zhang, S.; Yao, R.; Xiao, Y. CRISPR base editing and prime editing: DSB and template-free editing systems for bacteria and plants. Synth. Syst. Biotechnol. 2020, 5, 277–292. [Google Scholar]
- Alok, A.; Sandhya, D.; Jogam, P.; Rodrigues, V.; Bhati, K.K.; Sharma, H.; Kumar, J. The rise of the CRISPR/Cpf1 system for efficient genome editing in plants. Front. Plant Sci. 2020, 11, 264. [Google Scholar] [CrossRef] [PubMed]
- Lin, Q.; Zong, Y.; Xue, C.; Wang, S.; Jin, S.; Zhu, Z.; Wang, Y.; Anzalone, A.V.; Raguram, A.; Doman, J.L. Prime genome editing in rice and wheat. Nat. Biotechnol. 2020, 38, 582–585. [Google Scholar] [CrossRef]
- Li, S.; Zhang, X.; Wang, W.; Guo, X.; Wu, Z.; Du, W.; Zhao, Y.; Xia, L. Expanding the scope of CRISPR/Cpf1-mediated genome editing in rice. Mol. Plant 2018, 11, 995–998. [Google Scholar] [CrossRef]
- Mohanta, T.K.; Bashir, T.; Hashem, A.; Abd_Allah, E.F.; Bae, H. Genome editing tools in plants. Genes 2017, 8, 399. [Google Scholar] [CrossRef] [PubMed]
- Lowe, K.; Wu, E.; Wang, N.; Hoerster, G.; Hastings, C.; Cho, M.-J.; Scelonge, C.; Lenderts, B.; Chamberlin, M.; Cushatt, J. Morphogenic regulators Baby boom and Wuschel improve monocot transformation. Plant Cell 2016, 28, 1998–2015. [Google Scholar] [CrossRef] [PubMed]
- Debernardi, J.M.; Tricoli, D.M.; Ercoli, M.F.; Hayta, S.; Ronald, P.; Palatnik, J.F.; Dubcovsky, J. A GRF–GIF chimeric protein improves the regeneration efficiency of transgenic plants. Nat. Biotechnol. 2020, 38, 1274–1279. [Google Scholar] [CrossRef]
- Li, J.-F.; Norville, J.E.; Aach, J.; McCormack, M.; Zhang, D.; Bush, J.; Church, G.M.; Sheen, J. Multiplex and homologous recombination–mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat. Biotechnol. 2013, 31, 688. [Google Scholar] [CrossRef]
- Nekrasov, V.; Staskawicz, B.; Weigel, D.; Jones, J.D.; Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotechnol. 2013, 31, 691. [Google Scholar] [CrossRef] [PubMed]
- Paul, J.W.; Qi, Y. CRISPR/Cas9 for plant genome editing: Accomplishments, problems and prospects. Plant Cell Rep. 2016, 35, 1417–1427. [Google Scholar] [CrossRef]
- Demirci, Y.; Zhang, B.; Unver, T. CRISPR/Cas9: An RNA-guided highly precise synthetic tool for plant genome editing. J. Cell. Physiol. 2018, 233, 1844–1859. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Zhang, H.; Lou, D.; Yu, D. Selection of highly efficient sgRNAs for CRISPR/Cas9-based plant genome editing. Sci. Rep. 2016, 6, 1–8. [Google Scholar] [CrossRef]
- Ran, F.A.; Hsu, P.D.; Wright, J.; Agarwala, V.; Scott, D.A.; Zhang, F. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 2013, 8, 2281. [Google Scholar] [CrossRef]
- Samanta, M.K.; Dey, A.; Gayen, S. CRISPR/Cas9: An advanced tool for editing plant genomes. Transgenic Res. 2016, 25, 561–573. [Google Scholar] [CrossRef]
- Fu, Y.; Sander, J.D.; Reyon, D.; Cascio, V.M.; Joung, J.K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 2014, 32, 279. [Google Scholar] [CrossRef] [PubMed]
- VanBuren, R.; Wai, C.M.; Wang, X.; Pardo, J.; Yocca, A.E.; Wang, H.; Chaluvadi, S.R.; Han, G.; Bryant, D.; Edger, P.P. Exceptional subgenome stability and functional divergence in the allotetraploid Ethiopian cereal teff. Nat. Commun. 2020, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Joyce, B.; Baltzell, A.; Bomhoff, M.; Lyons, E. Comparative Genomics Using CoGe, Hook, Line, and Sinker. In Bioinformatics in Aquaculture: Principles and Methods; John Wiley & Sons: Hoboken, NJ, USA, 2017. [Google Scholar] [CrossRef]
- Kumar, S.; Tamura, K.; Nei, M. MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief. Bioinform. 2004, 5, 150–163. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Ketema, S. Tef-Eragrostis tef (Zucc.); Bioversity International: Rome, Italy, 1997; Volume 12. [Google Scholar]
- Zhu, Q.; Smith, S.M.; Ayele, M.; Yang, L.; Jogi, A.; Chaluvadi, S.R.; Bennetzen, J.L. High-throughput discovery of mutations in tef semi-dwarfing genes by next-generation sequencing analysis. Genetics 2012, 192, 819–829. [Google Scholar] [CrossRef]
- Miller, D. Teff Grass: A new alternative. In Proceedings of the 2009 California Alfalfa & Forage Symposium and Western Seed Conference, Reno, NV, USA, 2–4 December 2009; UC Cooperative Extension, Plant Sciences Department, University of California: Davis, CA, USA, 2009. [Google Scholar]
- Shi, L.; Wei, X.; Adedze, Y.; Sheng, Z.; Tang, S.; Hu, P.; Wang, J. Characterization and gene cloning of the rice (Oryza sativa L.) dwarf and narrow-leaf mutant dnl3. Genet. Mol. Res. 2016, 15, 1–12. [Google Scholar] [CrossRef]
- Zhao, Z.; Xue, Y.; Yang, H.; Li, H.; Sun, G.; Zhao, X.; Ding, D.; Tang, J. Genome-Wide Identification of miRNAs and Their Targets Involved in the Developing Internodes under Maize Ears by Responding to Hormone Signaling. PLoS ONE 2016, 11, e0164026. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Gu, L.; Li, P.; Song, X.; Wei, L.; CHen, Z.; Cao, X. Degradome sequencing reveals endogenous small RNA targets in rice (Oryza sativa L. ssp. indica). Front. Biol. 2010, 5, 67–90. [Google Scholar] [CrossRef]
- Yang, Y.-H.; Zhang, F.-M.; Ge, S. Evolutionary rate patterns of the Gibberellin pathway genes. BMC Evol. Biol. 2009, 9, 206. [Google Scholar] [CrossRef] [PubMed]
- Singla, B.; Khurana, J.P.; Khurana, P. Characterization of three somatic embryogenesis receptor kinase genes from wheat, Triticum aestivum. Plant Cell Rep. 2008, 27, 833–843. [Google Scholar] [CrossRef]
- Pinthus, M.; Levy, A. The relationship between the Rht 1 and Rht 2 dwarfing genes and grain weight in Triticum aestivum L. spring wheat. Theor. Appl. Genet. 1983, 66, 153–157. [Google Scholar] [CrossRef]
- Gouda, G.; Gupta, M.K.; Donde, R.; Mohapatra, T.; Vadde, R.; Behera, L. Marker-assisted selection for grain number and yield-related traits of rice (Oryza sativa L.). Physiol. Mol. Biol. Plants Int. J. Funct. Plant Biol. 2020, 26, 885. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, W.; Yang, B.; Currey, C.; Fei, S.-Z. Functional analysis of the teosinte branched 1 gene in the tetraploid switchgrass (Panicum virgatum L.) by CRISPR/Cas9-directed mutagenesis. BioRxiv 2020, 1–33. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Zhang, G.; Liu, D.; Niu, M.; Tong, H.; Chu, C. GSK2 stabilizes OFP3 to suppress brassinosteroid responses in rice. Plant J. 2020, 102, 1187–1201. [Google Scholar] [CrossRef]
- Miao, C.; Xiao, L.; Hua, K.; Zou, C.; Zhao, Y.; Bressan, R.A.; Zhu, J.-K. Mutations in a subfamily of abscisic acid receptor genes promote rice growth and productivity. Proc. Natl. Acad. Sci. USA 2018, 115, 6058–6063. [Google Scholar] [CrossRef]
- Xu, C.; Wang, Y.; Yu, Y.; Duan, J.; Liao, Z.; Xiong, G.; Meng, X.; Liu, G.; Qian, Q.; Li, J. Degradation of MONOCULM 1 by APC/C TAD1 regulates rice tillering. Nat. Commun. 2012, 3, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Bhatia, D.; Joshi, S.; Das, A.; Vikal, Y.; Sahi, G.K.; Neelam, K.; Kaur, K.; Singh, K. Introgression of yield component traits in rice (Oryza sativa ssp. indica) through interspecific hybridization. Crop Sci. 2017, 57, 1557–1573. [Google Scholar] [CrossRef]
- Wang, N.-N.; Xu, S.-W.; Sun, Y.-L.; Liu, D.; Zhou, L.; Li, Y.; Li, X.-B. The cotton WRKY transcription factor (GhWRKY33) reduces transgenic Arabidopsis resistance to drought stress. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Satapathy, L.; Kumar, D.; Kumar, M.; Mukhopadhyay, K. Functional and DNA–protein binding studies of WRKY transcription factors and their expression analysis in response to biotic and abiotic stress in wheat (Triticum aestivum L.). 3 Biotech 2018, 8, 40. [Google Scholar] [CrossRef]
- Hittalmani, S.; Mahesh, H.; Shirke, M.D.; Biradar, H.; Uday, G.; Aruna, Y.; Lohithaswa, H.; Mohanrao, A. Genome and transcriptome sequence of finger millet (Eleusine coracana (L.) Gaertn.) provides insights into drought tolerance and nutraceutical properties. BMC Genom. 2017, 18, 465. [Google Scholar] [CrossRef]
- Wang, Y.; Tong, X.; Qiu, J.; Li, Z.; Zhao, J.; Hou, Y.; Tang, L.; Zhang, J. A phosphoproteomic landscape of rice (Oryza sativa) tissues. Physiol. Plant. 2017, 160, 458–475. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Gao, X.; Liu, Q.; Shao, Y.; Zhang, D.; Jiang, L.; Li, C. Overexpression of TaWRKY146 increases drought tolerance through inducing stomatal closure in Arabidopsis thaliana. Front. Plant Sci. 2017, 8, 2036. [Google Scholar] [CrossRef] [PubMed]
- Soltani Najafabadi, M. Improving rice (Oryza sativa L.) drought tolerance by suppressing a NF-YA transcription factor. Iran. J. Biotechnol. 2012, 10, 40–48. [Google Scholar]
- Capell, T.; Escobar, C.; Liu, H.; Burtin, D.; Lepri, O.; Christou, P. Over-expression of the oat arginine decarboxylase cDNA in transgenic rice (Oryza sativa L.) affects normal development patterns in vitro and results in putrescine accumulation in transgenic plants. Theor. Appl. Genet. 1998, 97, 246–254. [Google Scholar] [CrossRef]
- Li, X.-Y.; Mantovani, R.; Hooft van Huijsduijnen, R.; Andre, I.; Benoist, C.; Mathis, D. Evolutionary variation of the CCAAT-binding transcription factor NF-Y. Nucleic Acids Res. 1992, 20, 1087–1091. [Google Scholar] [CrossRef] [PubMed]
- Sprink, T.; Eriksson, D.; Schiemann, J.; Hartung, F. Regulatory hurdles for genome editing: Process-vs. product-based approaches in different regulatory contexts. Plant Cell Rep. 2016, 35, 1493–1506. [Google Scholar] [CrossRef]
- Liang, Z.; Chen, K.; Li, T.; Zhang, Y.; Wang, Y.; Zhao, Q.; Liu, J.; Zhang, H.; Liu, C.; Ran, Y. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat. Commun. 2017, 8, 14261. [Google Scholar] [CrossRef] [PubMed]
- Woo, J.W.; Kim, J.; Kwon, S.I.; Corvalán, C.; Cho, S.W.; Kim, H.; Kim, S.-G.; Kim, S.-T.; Choe, S.; Kim, J.-S. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat. Biotechnol. 2015, 33, 1162. [Google Scholar] [CrossRef] [PubMed]
- Latorre, A.; Latorre, A.; Somoza, Á. Modified RNAs in CRISPR/Cas9: An old trick works again. Angew. Chem. Int. Ed. 2016, 55, 3548–3550. [Google Scholar] [CrossRef] [PubMed]
- Gerashchenkov, G.; Rozhnova, N.; Kuluev, B.; Kiryanova, O.Y.; Gumerova, G.; Knyazev, A.; Vershinina, Z.; Mikhailova, E.; Chemeris, D.; Matniyazov, R. Design of Guide RNA for CRISPR/Cas Plant Genome Editing. Mol. Biol. 2020, 54, 24–42. [Google Scholar] [CrossRef]
- Salvi, S.; Druka, A.; Milner, S.G.; Gruszka, D. Induced genetic variation, TILLING and NGS-based cloning. In Biotechnological Approaches to Barley Improvement; Springer: Berlin/Heidelberg, Germany, 2014; pp. 287–310. [Google Scholar]
- Zhang, D.; Li, Z.; Li, J.-F. Targeted gene manipulation in plants using the CRISPR/Cas technology. J. Genet. Genom. 2016, 43, 251–262. [Google Scholar] [CrossRef]
- Crosetto, N.; Mitra, A.; Silva, M.J.; Bienko, M.; Dojer, N.; Wang, Q.; Karaca, E.; Chiarle, R.; Skrzypczak, M.; Ginalski, K. Nucleotide-resolution DNA double-strand break mapping by next-generation sequencing. Nat. Methods 2013, 10, 361. [Google Scholar] [CrossRef]
- Frock, R.L.; Hu, J.; Meyers, R.M.; Ho, Y.-J.; Kii, E.; Alt, F.W. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nat. Biotechnol. 2015, 33, 179. [Google Scholar] [CrossRef]
- Kim, D.; Bae, S.; Park, J.; Kim, E.; Kim, S.; Yu, H.R.; Hwang, J.; Kim, J.-I.; Kim, J.-S. Digenome-seq: Genome-wide profiling of CRISPR-Cas9 off-target effects in human cells. Nat. Methods 2015, 12, 237. [Google Scholar] [CrossRef]
- Tsai, S.Q.; Zheng, Z.; Nguyen, N.T.; Liebers, M.; Topkar, V.V.; Thapar, V.; Wyvekens, N.; Khayter, C.; Iafrate, A.J.; Le, L.P. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 2015, 33, 187. [Google Scholar] [CrossRef]
- Hsu, P.D.; Lander, E.S.; Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 2014, 157, 1262–1278. [Google Scholar] [CrossRef] [PubMed]
- Fauser, F.; Schiml, S.; Puchta, H. Both CRISPR/C as-based nucleases and nickases can be used efficiently for genome engineering in A rabidopsis thaliana. Plant J. 2014, 79, 348–359. [Google Scholar] [CrossRef] [PubMed]
- Slaymaker, I.M.; Gao, L.; Zetsche, B.; Scott, D.A.; Yan, W.X.; Zhang, F. Rationally engineered Cas9 nucleases with improved specificity. Science 2016, 351, 84–88. [Google Scholar] [CrossRef] [PubMed]
- Kleinstiver, B.P.; Pattanayak, V.; Prew, M.S.; Tsai, S.Q.; Nguyen, N.T.; Zheng, Z.; Joung, J.K. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 2016, 529, 490. [Google Scholar] [CrossRef]
| Gene | Plant Name | Accession Number | Reference |
|---|---|---|---|
| Plant Height | |||
| KO2 | Oryza sativa Japonica | AY660664 | [148] |
| GA regulatory factor-like (GRF) mRNA | Zea mays | KJ466125 | [149] |
| growth-regulating factor 10 (GRF10) | Oryza sativa Indica | FJ546694 | [150] |
| GA20-oxidase (GA20ox2) | Oryza granulata | EU179380 | [151] |
| BRI1 | Triticum aestivum | DQ655711 | [152] |
| Sd-1 (used in green revl) | Oryza sativa | KP212897.1 | [70] |
| RHT1 | Triticum aestivum | FN649763 | [153] |
| Number of Tillers and Panicle Branches | |||
| OsCKX2 | Oryza sativa | AB205193.1 | [154] |
| teosinte branched1 (tb1) | switchgrass | AF131673.2 | [155] |
| GSK2 | Oryza sativa | XM_015782085 | [156] |
| PYL2 | Oryza sativa | KJ700410.1 | [157] |
| PYL3, | Oryza sativa | KJ191278.1 | |
| PYL4, | Oryza sativa | KJ855099.1 | |
| PYL5, | Oryza sativa | KJ855100.1 | |
| PYL6 | Oryza sativa | KJ855101.1 | |
| PYL12 | Oryza sativa | KJ855107.1 | |
| monoculm1 MOC1 | Oryza sativa Japonica | KC700671.1 | [158] |
| Grain Size | |||
| G1F1A | Oryza sativa | GU797949 | [159] |
| Drought Tolerance | |||
| GhWRKY33 | Gossypium hirsutum | KJ825875.1 | [160] |
| WRKY mRNA | Triticum aestivum | KT865879 | [161] |
| threonine dehydratase mRNA | Eleusine coracana | MK573864 | [162] |
| OsCDPK7 | Oryza sativa Japonica | AB042550 | [163] |
| TaWRKY146 | Triticum aestivum | MF770640.1 | [164] |
| NF-Y18 | Oryza sativa Japonica | HQ731479 | [165] |
| Arginine decarboxylase (ADC) | Oryza sativa Japonica | CA754598.1 | [166] |
| CIPK12 | Oryza sativa Japonica | EU703798 | [166] |
| NF-YB | Zea mays | NM_001112582 | [167] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Numan, M.; Khan, A.L.; Asaf, S.; Salehin, M.; Beyene, G.; Tadele, Z.; Ligaba-Osena, A. From Traditional Breeding to Genome Editing for Boosting Productivity of the Ancient Grain Tef [Eragrostis tef (Zucc.) Trotter]. Plants 2021, 10, 628. https://doi.org/10.3390/plants10040628
Numan M, Khan AL, Asaf S, Salehin M, Beyene G, Tadele Z, Ligaba-Osena A. From Traditional Breeding to Genome Editing for Boosting Productivity of the Ancient Grain Tef [Eragrostis tef (Zucc.) Trotter]. Plants. 2021; 10(4):628. https://doi.org/10.3390/plants10040628
Chicago/Turabian StyleNuman, Muhammad, Abdul Latif Khan, Sajjad Asaf, Mohammad Salehin, Getu Beyene, Zerihun Tadele, and Ayalew Ligaba-Osena. 2021. "From Traditional Breeding to Genome Editing for Boosting Productivity of the Ancient Grain Tef [Eragrostis tef (Zucc.) Trotter]" Plants 10, no. 4: 628. https://doi.org/10.3390/plants10040628
APA StyleNuman, M., Khan, A. L., Asaf, S., Salehin, M., Beyene, G., Tadele, Z., & Ligaba-Osena, A. (2021). From Traditional Breeding to Genome Editing for Boosting Productivity of the Ancient Grain Tef [Eragrostis tef (Zucc.) Trotter]. Plants, 10(4), 628. https://doi.org/10.3390/plants10040628







