Small “Nested” Introgressions from Wild Thinopyrum Species, Conferring Effective Resistance to Fusarium Diseases, Positively Impact Durum Wheat Yield Potential
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Field Trials and Trait Measurements
2.3. Statistical Analysis
3. Results and Discussion
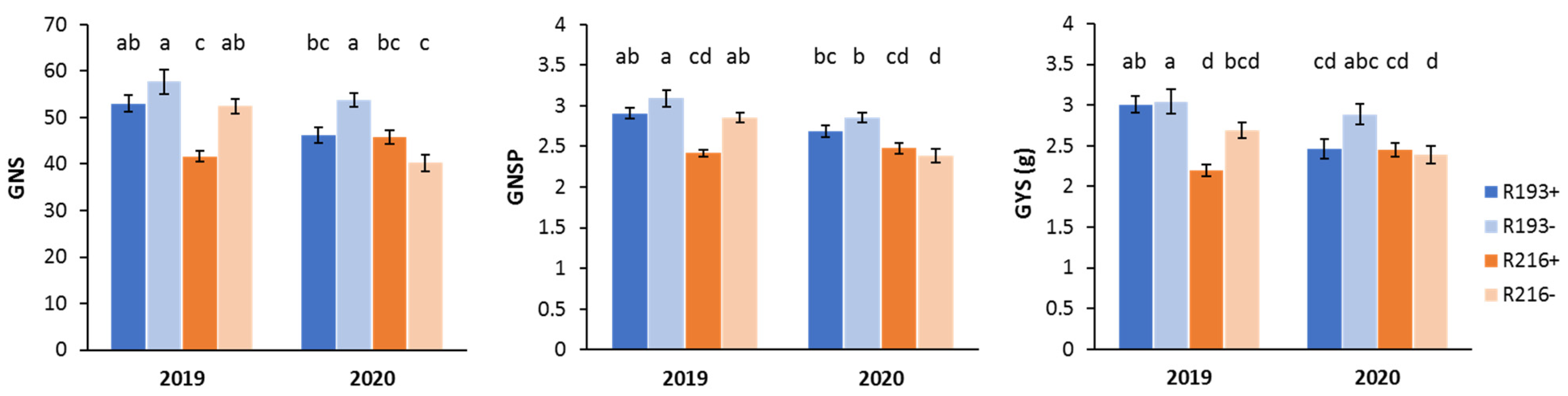
3.1. 7el2 Effect on Yield-Related Traits under Rainfed Conditions
3.2. 7E Effect on Yield-Related Traits under Rainfed Condition
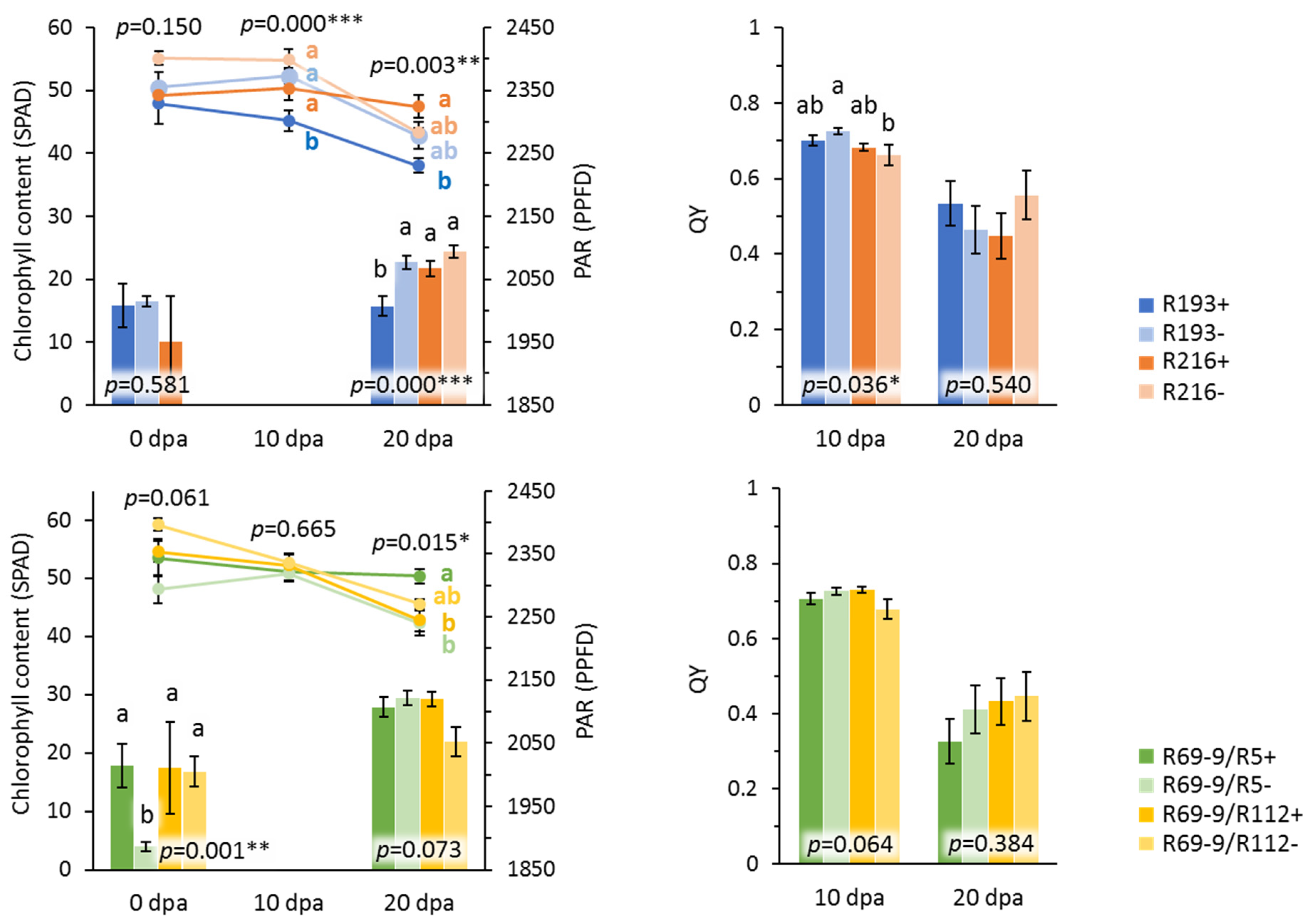
3.3. Novel Alien Segment Effects on Physiological Traits
3.4. Yield Performance of Durum Wheat-Thinopyrum spp. Recombinants vs. Top-Yielding Cultivars under Rainfed and Irrigated Conditions
4. Conclusions and Perspectives
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Xynias, I.N.; Mylonas, I.; Korpetis, E.G.; Ninou, E. Durum Wheat Breeding in the Mediterranean Region: Current Status and Future Prospects. Agronomy 2020, 10, 432. [Google Scholar] [CrossRef]
- FAOSTAT. Available online: http://www.fao.org/faostat/en/#data/QC (accessed on 8 January 2021).
- Sall, A.T.; Chiari, T.; Legesse, W.; Seid-Ahmed, K.; Ortiz, R.; van Ginkel, M.; Bassi, F.M. Durum Wheat (Triticum durum Desf.): Origin, Cultivation and Potential Expansion in Sub-Saharan Africa. Agronomy 2019, 9, 263. [Google Scholar] [CrossRef]
- Martínez-Moreno, F.; Solís, I.; Noguero, D.; Blanco, A.; Özberk, İ.; Nsarellah, N.; Elias, E.; Mylonas, I.; Soriano, J. Durum wheat in the Mediterranean rim: Historical evolution and genetic resources. Genet. Resour. Crop Evol. 2020, 67, 1415–1436. [Google Scholar] [CrossRef]
- Maccaferri, M.; Harris, N.S.; Twardziok, S.O.; Pasam, R.K.; Gundlach, H.; Spannagl, M.; Ormanbekova, D.; Lux, T.; Prade, V.M.; Milner, S.G.; et al. Durum wheat genome highlights past domestication signatures and future improvement targets. Nat. Genet. 2019, 51, 885–895. [Google Scholar] [CrossRef] [PubMed]
- Tester, M.; Langridge, P. Breeding technologies to increase crop production in a changing world. Science 2010, 327, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Ray, D.K.; Ramankutty, N.; Mueller, N.D.; West, P.C.; Foley, J.A. Recent patterns of crop yield growth and stagnation. Nat. Commun. 2012, 3, 1293. [Google Scholar] [CrossRef] [PubMed]
- Ray, D.K.; Mueller, N.D.; West, P.C.; Foley, J.A. Yield Trends Are Insufficient to Double Global Crop Production by 2050. PLoS ONE 2013, 8, e66428. [Google Scholar] [CrossRef] [PubMed]
- Senapati, N.; Brown, H.E.; Semenov, M.A. Raising genetic yield potential in high productive countries: Designing wheat ideotypes under climate change. Agric. For. Meteorol. 2019, 271, 33–45. [Google Scholar] [CrossRef]
- Xiong, W.; Reynolds, M.P.; Crossa, J.; Payne, T.S.; Schulthess, U.; Sonder, K.; Addimando, N.; Singh, R.P.; Ammar, K.; Gerard, B. Climate change has increased genotype-environment interactions in wheat breeding. Res. Sq. 2020. [Google Scholar] [CrossRef]
- Ceoloni, C.; Kuzmanović, L.; Forte, P.; Gennaro, A.; Bitti, A. Targeted exploitation of gene pools of alien Triticeae species for sustainable and multi-faceted improvement of the durum wheat crop. Crop Pasture Sci. 2014, 65, 96–111. [Google Scholar] [CrossRef]
- Gregory, P.J.; Johnson, S.N.; Newton, A.C.; Ingram, J.S.I. Integrating pests and pathogens into the climate change/food security debate. J. Exp. Bot. 2009, 60, 2827–2838. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, S.; Newton, A.C. Climate change, plant diseases and food security: An overview. Plant Pathol. 2011, 60, 2–14. [Google Scholar] [CrossRef]
- Zhang, X.; Halder, J.; White, R.P.; Hughes, D.J.; Ye, Z.; Wang, C.; Xu, R.; Gan, B.; Fitt, B.D.L. Climate change increases risk of fusarium ear blight on wheat in central China. Ann. Appl. Biol. 2014, 164, 384–395. [Google Scholar] [CrossRef]
- Alahmad, S.; Kang, Y.; Dinglasan, E.; Mazzucotelli, E.; Voss-Fels, K.P.; Able, J.A.; Christopher, J.; Bassi, F.M.; Hickey, L.T. Adaptive traits to improve durum wheat yield in drought and crown rot environments. Int. J. Mol. Sci. 2020, 21, 5260. [Google Scholar] [CrossRef]
- Buerstmayr, M.; Steiner, B.; Buerstmayr, H. Breeding for Fusarium head blight resistance in wheat—Progress and challenges. Plant Breed. 2020, 139, 429–454. [Google Scholar] [CrossRef]
- Maresca, M. From the gut to the brain: Journey and pathophysiological effects of the food-associated trichothecene mycotoxin deoxynivalenol. Toxins 2013, 5, 784–820. [Google Scholar] [CrossRef] [PubMed]
- Walkowiak, S.; Gao, L.; Monat, C.; Haberer, G.; Kassa, M.T.; Brinton, J.; Ramirez-Gonzalez, R.H.; Kolodziej, M.C.; Delorean, E.; Thambugala, D.; et al. Multiple wheat genomes reveal global variation in modern breeding. Nature 2020, 588, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Wang, S.; Liang, Z.; Shi, W.; Gao, C.; Xia, G. From Genetic Stock to Genome Editing: Gene Exploitation in Wheat. Trends Biotechnol. 2018, 36, 160–172. [Google Scholar] [CrossRef] [PubMed]
- Brinton, J.; Ramirez-Gonzalez, R.H.; Simmonds, J.; Wingen, L.; Orford, S.; Griffiths, S.; Haberer, G.; Spannagl, M.; Walkowiak, S.; Pozniak, C.; et al. A haplotype-led approach to increase the precision of wheat breeding. Commun. Biol. 2020, 3, 712. [Google Scholar] [CrossRef] [PubMed]
- Ceoloni, C.; Kuzmanović, L.; Forte, P.; Virili, M.E.; Bitti, A. Wheat-Perennial Triticeae Introgressions: Major Achievements and Prospects. In Alien Introgression in Wheat—Cytogenetics, Molecular Biology, and Genomics; Molnár-Láng, M., Ceoloni, C., Doležel, J., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 273–313. [Google Scholar]
- Feuillet, C.; Langridge, P.; Waugh, R. Cereal breeding takes a walk on the wild side. Trends Genet. 2008, 24, 24–32. [Google Scholar] [CrossRef] [PubMed]
- Mondal, S.; Rutkoski, J.E.; Velu, G.; Singh, P.K.; Crespo-Herrera, L.A.; Guzman, C.; Bhavani, S.; Lan, C.; He, X.; Singh, R.P. Harnessing diversity in wheat to enhance grain yield, climate resilience, disease and insect pest resistance and nutrition through conventional and modern breeding approaches. Front. Plant Sci. 2016, 7, 91. [Google Scholar] [CrossRef] [PubMed]
- Prohens, J.; Gramazio, P.; Plazas, M.; Dempewolf, H.; Kilian, B.; Díez, M.J.; Fita, A.; Herraiz, F.J.; Rodríguez-Burruezo, A.; Soler, S.; et al. Introgressiomics: A new approach for using crop wild relatives in breeding for adaptation to climate change. Euphytica 2017, 213, 158. [Google Scholar] [CrossRef]
- Sears, E.R. Chromosome engineering in wheat. In Stadler Genetics Symposia, 4th ed.; Kimber, G., Rédei, G.R., Eds.; University of Missouri: Columbia, MO, USA, 1972; pp. 23–38. [Google Scholar]
- Sears, E.R. Transfer of alien genetic material to wheat. In Wheat Science: Today and Tomorrow; Evans, L.T., Peacock, W.J., Eds.; Cambridge University Press: Cambridge, UK, 1981; pp. 75–89. [Google Scholar]
- Qi, L.; Friebe, B.; Zhang, P.; Gill, B.S. Homoeologous recombination, chromosome engineering and crop improvement. Chromosom. Res. 2007, 15, 3–19. [Google Scholar] [CrossRef] [PubMed]
- Kuzmanović, L.; Rossini, F.; Ruggeri, R.; Pagnotta, M.; Ceoloni, C. Engineered Durum Wheat Germplasm with Multiple Alien Introgressions: Agronomic and Quality Performance. Agronomy 2020, 10, 486. [Google Scholar] [CrossRef]
- Kuzmanović, L.; Gennaro, A.; Benedettelli, S.; Dodd, I.C.; Quarrie, S.A.; Ceoloni, C. Structural-functional dissection and characterization of yield-contributing traits originating from a group 7 chromosome of the wheatgrass species Thinopyrum ponticum after transfer into durum wheat. J. Exp. Bot. 2014, 65, 509–525. [Google Scholar] [CrossRef]
- Sharma, S.; Xu, S.; Ehdaie, B.; Hoops, A.; Close, T.J.; Lukaszewski, A.J.; Waines, J.G. Dissection of QTL effects for root traits using a chromosome arm-specific mapping population in bread wheat. Theor. Appl. Genet. 2011, 122, 759–769. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, M.; Langridge, P. Physiological breeding. Curr. Opin. Plant Biol. 2016, 31, 162–171. [Google Scholar] [CrossRef]
- Forte, P.; Virili, M.E.; Kuzmanović, L.; Moscetti, I.; Gennaro, A.; D’Ovidio, R.; Ceoloni, C. A novel assembly of Thinopyrum ponticum genes into the durum wheat genome: Pyramiding Fusarium head blight resistance onto recombinant lines previously engineered for other beneficial traits from the same alien species. Mol. Breed. 2014, 34, 1701–1716. [Google Scholar] [CrossRef]
- Kuzmanović, L.; Mandalà, G.; Tundo, S.; Ciorba, R.; Frangella, M.; Ruggeri, R.; Rossini, F.; Gevi, F.; Rinalducci, S.; Ceoloni, C. Equipping Durum Wheat—Thinopyrum ponticum Recombinant Lines with a Thinopyrum elongatum Major QTL for Resistance to Fusarium Diseases Through a Cytogenetic Strategy. Front. Plant Sci. 2019, 10, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Ceoloni, C.; Forte, P.; Kuzmanović, L.; Tundo, S.; Moscetti, I.; De Vita, P.; Virili, M.E.; D’Ovidio, R. Cytogenetic mapping of a major locus for resistance to Fusarium head blight and crown rot of wheat on Thinopyrum elongatum 7EL and its pyramiding with valuable genes from a Th. ponticum homoeologous arm onto bread wheat 7DL. Theor. Appl. Genet. 2017, 130, 2005–2024. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sun, S.; Ge, W.; Zhao, L.; Hou, B.; Wang, K.; Lyu, Z.; Chen, L.; Xu, S.; Guo, J.; et al. Horizontal gene transfer of Fhb7 from fungus underlies Fusarium head blight resistance in wheat. Science 2020, 368, 844. [Google Scholar] [CrossRef] [PubMed]
- Ceoloni, C.; Forte, P.; Gennaro, A.; Micali, S.; Carozza, R.; Bitti, A. Recent developments in durum wheat chromosome engineering. Cytogenet. Genome Res. 2005, 109, 328–334. [Google Scholar] [CrossRef] [PubMed]
- Kuzmanović, L.; Ruggeri, R.; Virili, M.E.; Rossini, F.; Ceoloni, C. Effects of Thinopyrum ponticum chromosome segments transferred into durum wheat on yield components and related morpho-physiological traits in Mediterranean rain-fed conditions. Field Crop. Res. 2016, 186, 86–98. [Google Scholar] [CrossRef]
- Kuzmanović, L.; Ruggeri, R.; Able, J.A.; Bassi, F.M.; Maccaferri, M.; Tuberosa, R.; De Vita, P.; Rossini, F.; Ceoloni, C. Yield of chromosomally engineered durum wheat-Thinopyrum ponticum recombinant lines in a range of contrasting rain-fed environments. Field Crop. Res. 2018, 228, 147–157. [Google Scholar] [CrossRef]
- Ceoloni, C.; Kuzmanović, L.; Ruggeri, R.; Rossini, F.; Forte, P.; Cuccurullo, A.; Bitti, A. Harnessing genetic diversity of wild gene pools to enhance wheat crop production and sustainability: Challenges and opportunities. Diversity 2017, 9, 55. [Google Scholar] [CrossRef]
- Virili, M.; Kuzmanović, L.; Bitti, A.; Salvi, S.; Tuberosa, R.; Ceoloni, C. Analysis of seminal root architecture in durum wheat-Thinopyrum ponticum recombinant lines. In Proceedings of the Joint Meeting SIBV-SIGA, Milano, Italy, 8–11 September 2015. [Google Scholar]
- Rossini, F.; Provenzano, M.E.; Kuzmanović, L.; Ceoloni, C.; Ruggeri, R. Assessing the Ability of Durum Wheat-Thinopyrum ponticum Recombinant Lines to Suppress Naturally Occurring Weeds under Different Sowing Densities. Agronomy 2020, 10, 709. [Google Scholar] [CrossRef]
- Quaranta, F.; Arcangeli, A.; Basili, O.; Belocchi, A.; Bottazzi, P.; Fabbrini, L.; Malagesi, F.; Mariotti, R.; Mazzon, V.; Cacciatori, P.; et al. Speciale Grano Duro: Dettaglio regionale dei risultati 2020—Centro Italia versante tirrenico. L’informatore Agrar. 2020, 29, 52–56. [Google Scholar]
- Carboni, G.; Dettori, M.; Mameli, L.; Rinaldi, M.; Colecchia, S.A.; Belocchi, A.; Quaranta, F. Prove di grano duro su sodo in Sardegna e Puglia. L’informatore Agrar. 2018, 39, 52–59. [Google Scholar]
- Giorgi, B. Origin, behaviour and utilization of a Ph1 mutant of durum wheat, Triticum turgidum (L.) var. durum. In Proceedings of the 6th International Wheat Genetics Symposium, Kyoto, Japan, 28 November–3 December 1983; pp. 1033–1040. [Google Scholar]
- Allen, R.G.; Pereira, L.S.; Raes, D.; Smith, M. Crop Evapotranspiration—Guidelines for Computing Crop Water Requirements; FAO Irrigation and Drainage Paper 56; Food and Agriculture Organization (FAO): Rome, Italy, 1998. [Google Scholar]
- Slafer, G.A.; Elia, M.; Savin, R.; García, G.A.; Terrile, I.I.; Ferrante, A.; Miralles, D.J.; González, F.G. Fruiting efficiency: An alternative trait to further rise wheat yield. Food Energy Secur. 2015, 4, 92–109. [Google Scholar] [CrossRef]
- Gennaro, A.; Forte, P.; Carozza, R.; Savo Sardaro, M.L.; Ferri, D.; Bitti, A.; Borrelli, G.M.; D’Egidio, M.G.; Ceoloni, C. Pyramiding different alien chromosome segments in durum wheat: Feasibility and breeding potential. Isr. J. Plant Sci. 2007, 55, 267–276. [Google Scholar] [CrossRef]
- Valluru, R.; Reynolds, M.P.; Lafarge, T. Food security through translational biology between wheat and rice. Food Energy Secur. 2015, 4, 203–218. [Google Scholar] [CrossRef]
- Royo, C.; Álvaro, F.; Martos, V.; Ramdani, A.; Isidro, J.; Villegas, D.; García Del Moral, L.F. Genetic changes in durum wheat yield components and associated traits in Italian and Spanish varieties during the 20th century. Euphytica 2007, 155, 259–270. [Google Scholar] [CrossRef]
- Reynolds, M.; Chapman, S.; Crespo-Herrera, L.; Molero, G.; Mondal, S.; Pequeno, D.N.L.; Pinto, F.; Pinera-Chavez, F.J.; Poland, J.; Rivera-Amado, C.; et al. Breeder friendly phenotyping. Plant Sci. 2020, 295, 110396. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, M.; Bonnett, D.; Chapman, S.C.; Furbank, R.T.; Manés, Y.; Mather, D.E.; Parry, M.A.J. Raising yield potential of wheat. I. Overview of a consortium approach and breeding strategies. J. Exp. Bot. 2011, 62, 439–452. [Google Scholar] [CrossRef] [PubMed]
- Furbank, R.T.; Jimenez-Berni, J.A.; George-Jaeggli, B.; Potgieter, A.B.; Deery, D.M. Field crop phenomics: Enabling breeding for radiation use efficiency and biomass in cereal crops. New Phytol. 2019, 223, 1714–1727. [Google Scholar] [CrossRef] [PubMed]
- Furbank, R.T.; Sharwood, R.; Estavillo, G.M.; Silva-Perez, V.; Condon, A.G. Photons to food: Genetic improvement of cereal crop photosynthesis. J. Exp. Bot. 2020, 71, 2226–2238. [Google Scholar] [CrossRef] [PubMed]
- Asseng, S.; Kassie, B.T.; Labra, M.H.; Amador, C.; Calderini, D.F. Simulating the impact of source-sink manipulations in wheat. Field Crop. Res. 2017, 202, 47–56. [Google Scholar] [CrossRef]
- Monsen, S.B.; Stevens, R.; Shaw, N.L. Grasses. In Restoring Western Ranges and Wildlands; U.S. Department of Agriculture Forest Service, Rocky Mountain Research Station: Fort Collins, CO, USA, 2004; Volume 2, pp. 295–424. [Google Scholar]
- Shu, Y.; Zhang, J.; Ao, Y.; Song, L.; Guo, C. Analysis of the Thinopyrum elongatum transcriptome under water deficit stress. Int. J. Genomics 2015, 265791. [Google Scholar] [CrossRef]
- Ceoloni, C.; Kuzmanović, L.; Gennaro, A.; Forte, P.; Giorgi, D.; Grossi, M.R.; Bitti, A. Genomes, chromosomes and genes of perennial triticeae of the genus Thinopyrum: The value of their transfer into wheat for gains in cytogenomic knowledge and ‘precision’ breeding. In Advances in Genomics of Plant Genetic Resources; Tuberosa, R., Graner, A., Frison, E., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 333–358. ISBN 9789400775756. [Google Scholar]
- Li, Z.; Li, B.; Tong, Y. The contribution of distant hybridization with decaploid Agropyron elongatum to wheat improvement in China. J. Genet. Genomics 2008, 35, 451–456. [Google Scholar] [CrossRef]
- Placido, D.F.; Campbell, M.T.; Folsom, J.J.; Cui, X.; Kruger, G.R.; Baenziger, P.S.; Walia, H. Introgression of novel traits from a wild wheat relative improves drought adaptation in wheat. Plant Physiol. 2013, 161, 1806–1819. [Google Scholar] [CrossRef] [PubMed]
- Placido, D.F.; Sandhu, J.; Sato, S.J.; Nersesian, N.; Quach, T.; Clemente, T.E.; Staswick, P.E.; Walia, H. The LATERAL ROOT DENSITY gene regulates root growth during water stress in wheat. Plant Biotechnol. J. 2020, 18, 1955–1968. [Google Scholar] [CrossRef] [PubMed]
- El Haddad, N.; Kabbaj, H.; Zaïm, M.; El Hassouni, K.; Tidiane Sall, A.; Azouz, M.; Ortiz, R.; Baum, M.; Amri, A.; Gamba, F.; et al. Crop wild relatives in durum wheat breeding: Drift or thrift? Crop Sci. 2021, 61, 37–54. [Google Scholar] [CrossRef]
- Savary, S.; Willocquet, L.; Pethybridge, S.J.; Esker, P.; McRoberts, N.; Nelson, A. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 2019, 3, 430–439. [Google Scholar] [CrossRef] [PubMed]
- Alahmad, S.; Dinglasan, E.; Leung, K.M.; Riaz, A.; Derbal, N.; Voss-Fels, K.P.; Able, J.A.; Bassi, F.M.; Christopher, J.; Hickey, L.T. Speed breeding for multiple quantitative traits in durum wheat. Plant Methods 2018, 14, 36. [Google Scholar] [CrossRef]



| Recombinant | Pedigree | Progeny | Alien Segment | Alien Genes | Field Trial | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Presence | %7AL | Lr19 | Yp | Fhb-7el2 | Fhb-7EL | RF | RF + IRR | |||
| R112+ | (Tr#12 a × Creso ph1c b)/5*Simeto | BC5F11 | hom+ | 28 | + | + | - | - | 2020 (sp) | |
| R5+ | (Tr#12 × Creso ph1c)/5*Simeto | BC5F11 | hom+ | 23 | + | + | - | - | 2020 (sp) | |
| R193+ | (KS24-1 c × R112+)/Simeto/IDYT22 | F4-5 | hom+ | 28 | + | + | + | - | 2019 (sp) | 2020 (sp) |
| R193- | (KS24-1 × R112+)Simeto/IDYT22 | F4-5 | hom− | - | - | - | - | - | 2019 (sp) | 2020 (sp) |
| R216+ | (KS24-1 × R5+)/Simeto/IDYT22 | F4-5 | hom+ | 23 | + | + | + | - | 2019 (sp) | 2020 (sp) |
| R216- | (KS24-1 × R5+)/Simeto/IDYT22 | F4-5 | hom− | - | - | - | - | - | 2019 (sp) | 2020 (sp) |
| R74-10/R112+ | (R74-10 d × R112+)/2*Simeto | BC2F4 | hom+ | 28 | + | - | - | + | 2019 (sr) | |
| R74-10/R112- | (R74-10 × R112+)/2*Simeto | BC2F4 | hom− | - | - | - | - | - | 2019 (sr) | |
| R74-10/R5+ | (R74-10 × R5+)/2*Simeto | BC2F4 | hom+ | 23 | + | - | - | + | 2019 (sr) | |
| R74-10/R5- | (R74-10 × R5+)/2*Simeto | BC2F4 | hom− | - | - | - | - | - | 2019 (sr) | |
| R69-9/R112+ | (R69-9 d × R112+)/2*Simeto | BC2F4-5 | hom+ | 28 | + | + | - | + | 2019 (sr) | 2020 (sp) |
| R69-9/R112- | (R69-9 × R112+)/2*Simeto | BC2F4-5 | hom− | - | - | - | - | - | 2019 (sr) | 2020 (sp) |
| R69-9/R5+ | (R69-9 × R5+)/2*Simeto | BC2F4 | hom+ | 23 | + | + | - | + | 2020 (sp) | |
| R69-9/R5- | (R69-9 × R5+)/2*Simeto | BC2F4 | hom− | - | - | - | - | - | 2020 (sp) | |
| Genotype | Plant Traits | Spike Traits | ||||||
|---|---|---|---|---|---|---|---|---|
| GY (g) | TILN | GN | TGW (g) | GYS (g) | SPN | GNS | GNSP | |
| R193+ | 13.3 | 7.5 | 265.5 | 50.2 | 2.7 ab | 17.5 b | 48.9 b | 2.8 ab |
| R193- | 13.3 | 6.9 | 278.0 | 49.8 | 2.9 a | 18.7 a | 55.3 a | 2.9 a |
| R216+ | 11.1 | 7.0 | 226.3 | 50.3 | 2.4 c | 18.0 b | 44.2 c | 2.5 c |
| R216- | 11.1 | 7.4 | 223.1 | 52.6 | 2.5 bc | 17.3 b | 44.8 bc | 2.6 bc |
| ANOVA p | ||||||||
| Genotype (G) | 0.090 | 0.980 | 0.040 * | 0.681 | 0.000 *** | 0.001 ** | 0.000 *** | 0.000 *** |
| Year (Y) | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.025 * | 0.280 | 0.000 *** | 0.000 *** |
| G × Y | 0.147 | 0.241 | 0.107 | 0.040 * | 0.007 ** | 0.000 *** | 0.000 *** | 0.007 ** |
| Genotype | Plant Traits | Spike Traits | |||||||
|---|---|---|---|---|---|---|---|---|---|
| GY (g) | TILN | GN | TGW (g) | GYS (g) | SPN | GNS | GNSP | SFI | |
| 2019 (sr) | |||||||||
| R69-9/R112+ | - | - | - | 49.6 ab | 3.0 a | 19.2 ab | 61.2 a | 3.2 a | 55.3 a |
| R69-9/R112- | - | - | - | 51.8 a | 2.8 ab | 20.0 a | 54.9 b | 2.7 bc | 52.7 ab |
| R74-10/R112+ | - | - | - | 53.9 a | 2.7 b | 17.9 cd | 49.5 bc | 2.8 b | 46.5 c |
| R74-10/R112- | - | - | - | 52.2 a | 2.3 c | 17.3 d | 44.2 d | 2.5 cd | 50.7 ac |
| R74-10/R5+ | - | - | - | 45.1 b | 2.1 c | 18.8 bc | 47.3 cd | 2.5 d | 51.8 ab |
| R74-10/R5- | 47.8 ab | 2.2 c | 18.3 bd | 47.3 cd | 2.6 bd | 49.7 bc | |||
| ANOVA p | - | - | - | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.001 ** |
| 2020 (sp) | |||||||||
| R69-9/R112+ | 11.3 | 4.2 | 191.2 | 59.1 ab | 3.0 ab | 18.4 b | 50.7 ab | 2.7 a | 58.7 b |
| R69-9/R112- | 11.2 | 4.0 | 199.6 | 56.6 bc | 3.3 a | 20.1 a | 55.9 a | 2.8 a | 68.1 a |
| R69-9/R5+ | 13.4 | 5.0 | 213.5 | 63.4 a | 2.9 bc | 19.6 a | 45.6 b | 2.3 b | 54.7 b |
| R69-9/R5- | 8.6 | 4.0 | 159.9 | 53.7 c | 2.6 c | 19.7 a | 46.7 b | 2.4 b | 64.0 a |
| ANOVA p | 0.178 | 0.325 | 0.560 | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** |
| Genotype | Plant Traits | Spike Traits | |||||||
|---|---|---|---|---|---|---|---|---|---|
| GY (g) | TILN | GN | TGW (g) | GYS (g) | SPN | GNS | GNSP | SFI | |
| R216+ | 13.5 | 5.9 a | 239.4 a | 56.4 de | 2.8 bd | 18.3 ce | 48.9 ab | 2.7 ad | 62.0 a |
| R69-9/R5+ | 13.4 | 5.0 ac | 208.2 ac | 64.8 ab | 3.0 ab | 19.1 bc | 45.9 bd | 2.4 e | 54.0 ab |
| R69-9/R112+ | 13.3 | 4.8 ac | 222.0 ab | 59.8 bd | 3.2 a | 18.4 cd | 52.6 a | 2.9 a | 61.8 a |
| Monastir | 12.6 | 5.2 ac | 231.7 ab | 54.3 e | 2.7 be | 19.6 b | 49.3 ab | 2.5 ce | 59.1 ab |
| R193+ | 12.0 | 5.0 ab | 206.2 ac | 58.3 de | 2.7 be | 17.1 f | 46.1 bd | 2.7 ac | 52.4 ab |
| Ettore | 11.6 | 5.3 ab | 222.3 ab | 52.5 e | 2.5 e | 18.7 c | 47.8 ac | 2.6 be | 59.8 e |
| R5+ | 11.0 | 4.5 ac | 172.1 ac | 63.1 ac | 2.7 be | 16.0 g | 42.6 de | 2.6 ad | 49.5 bc |
| R112+ | 10.7 | 4.4 bc | 185.9 ac | 58.3 ce | 2.7 ce | 15.8 g | 45.2 bd | 2.8 ab | 61.8 a |
| Kanakis | 10.6 | 4.7 ac | 185.5 ac | 56.3 de | 2.5 de | 17.4 ef | 43.3 cd | 2.5 ce | 54.9 ab |
| IDYT22 | 10.2 | 4.6 ac | 157.8 bc | 64.3 ab | 2.5 e | 16.1 g | 37.9 e | 2.4 e | 42.0 c |
| Antalis | 9.8 | 4.0 bc | 180.3 ac | 55.5 de | 2.9 ac | 20.6 a | 50.2 ab | 2.4 e | 58.4 ab |
| Simeto | 9.2 | 3.3 c | 134.8 c | 66.7 a | 2.8 ac | 17.5 df | 41.4 ce | 2.4 de | 51.4 ac |
| Mean | 11.5 | 4.7 | 195.5 | 59.2 | 2.7 | 17.9 | 45.9 | 2.6 | 55.6 |
| ANCOVA p | |||||||||
| Genotype (G) | 0.031 * | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** |
| Irrigation (I) | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.000 *** | 0.322 | 0.000 *** | 0.000 *** | 0.717 |
| G × I | 0.019 * | 0.078 | 0.019 * | 0.014 * | 0.000 *** | 0.005 ** | 0.000 *** | 0.000 *** | 0.174 |
| Replica | 0.000 *** | 0.109 | 0.002 ** | 0.015 * | 0.000 *** | 0.000 *** | 0.000 *** | 0.016 * | 0.695 |
| Genotype | Flag Leaf Traits | ||||||
|---|---|---|---|---|---|---|---|
| SPAD1 | SPAD2 | SPAD3 | PAR1 | PAR3 | QY2 | QY3 | |
| R216+ | 49.2 ab | 50.7 ab | 50.9 ab | 1950.0 | 2055.0 ac | 0.69 ac | 0.46 ab |
| R69-9/R5+ | 53.5 ab | 51.8 ab | 52.1 a | 2014.4 | 2098.3 a | 0.71 a | 0.42 ab |
| R69-9/R112+ | 54.5 ab | 52.7 ab | 46.3 ad | 2011.1 | 2093.3 ab | 0.71 a | 0.46 ab |
| Monastir | 49.6 ab | 50.3 ab | 46.1 ad | 1845.6 | 2011.7 cf | 0.72 a | 0.51 ab |
| R193+ | 47.9 ab | 48.8 b | 44.1 cd | 2007.8 | 2023.3 ce | 0.70 ab | 0.54 ab |
| Ettore | 45.7 b | 50.5 ab | 44.5 bd | 1796.7 | 2016.7 cf | 0.66 bc | 0.47 ab |
| R5+ | 52.0 ab | 51.6 ab | 48.1 ad | 2025.6 | 1975.8 df | 0.68 ac | 0.45 ab |
| R112+ | 51.5 ab | 52.7 ab | 52.4 a | 1893.3 | 2103.0 ab | 0.70 ab | 0.46 ab |
| Kanakis | 45.5 b | 49.3 ab | 41.9 d | 1841.1 | 1912.5 g | 0.69 ac | 0.44 ab |
| IDYT22 | 56.7 ab | 51.6 ab | 44.9 bd | 2020.0 | 1951.7 fg | 0.69 ac | 0.49 ab |
| Antalis | 52.9 ab | 52.1 ab | 42.2 cd | N/A | 2031.7 bd | 0.67 ac | 0.59 a |
| Simeto | 60.6 a | 54.3 a | 48.9 ac | 1923.3 | 1971.1 ef | 0.64 c | 0.41 b |
| Mean | 51.6 | 51.4 | 46.9 | 1939.0 | 2020.3 | 0.69 | 0.47 |
| ANCOVA p | |||||||
| Genotype (G) | 0.006 ** | 0.041 * | 0.000 *** | 0.055 | 0.000 *** | 0.000 *** | 0.038 * |
| Irrigation (I) | - | 0.008 ** | 0.000 *** | - | 0.063 | 0.245 | 0.000 *** |
| G × I | - | 0.629 | 0.009 ** | - | 0.000 *** | 0.065 | 0.360 |
| Replica | 0.647 | 0.788 | 0.049 * | 0.014 * | 0.681 | 0.268 | 0.486 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kuzmanović, L.; Giovenali, G.; Ruggeri, R.; Rossini, F.; Ceoloni, C. Small “Nested” Introgressions from Wild Thinopyrum Species, Conferring Effective Resistance to Fusarium Diseases, Positively Impact Durum Wheat Yield Potential. Plants 2021, 10, 579. https://doi.org/10.3390/plants10030579
Kuzmanović L, Giovenali G, Ruggeri R, Rossini F, Ceoloni C. Small “Nested” Introgressions from Wild Thinopyrum Species, Conferring Effective Resistance to Fusarium Diseases, Positively Impact Durum Wheat Yield Potential. Plants. 2021; 10(3):579. https://doi.org/10.3390/plants10030579
Chicago/Turabian StyleKuzmanović, Ljiljana, Gloria Giovenali, Roberto Ruggeri, Francesco Rossini, and Carla Ceoloni. 2021. "Small “Nested” Introgressions from Wild Thinopyrum Species, Conferring Effective Resistance to Fusarium Diseases, Positively Impact Durum Wheat Yield Potential" Plants 10, no. 3: 579. https://doi.org/10.3390/plants10030579
APA StyleKuzmanović, L., Giovenali, G., Ruggeri, R., Rossini, F., & Ceoloni, C. (2021). Small “Nested” Introgressions from Wild Thinopyrum Species, Conferring Effective Resistance to Fusarium Diseases, Positively Impact Durum Wheat Yield Potential. Plants, 10(3), 579. https://doi.org/10.3390/plants10030579







