Chloroplast Genome of Rambutan and Comparative Analyses in Sapindaceae
Abstract
1. Introduction
2. Results
2.1. Chloroplast Genome Features of N. lappaceum
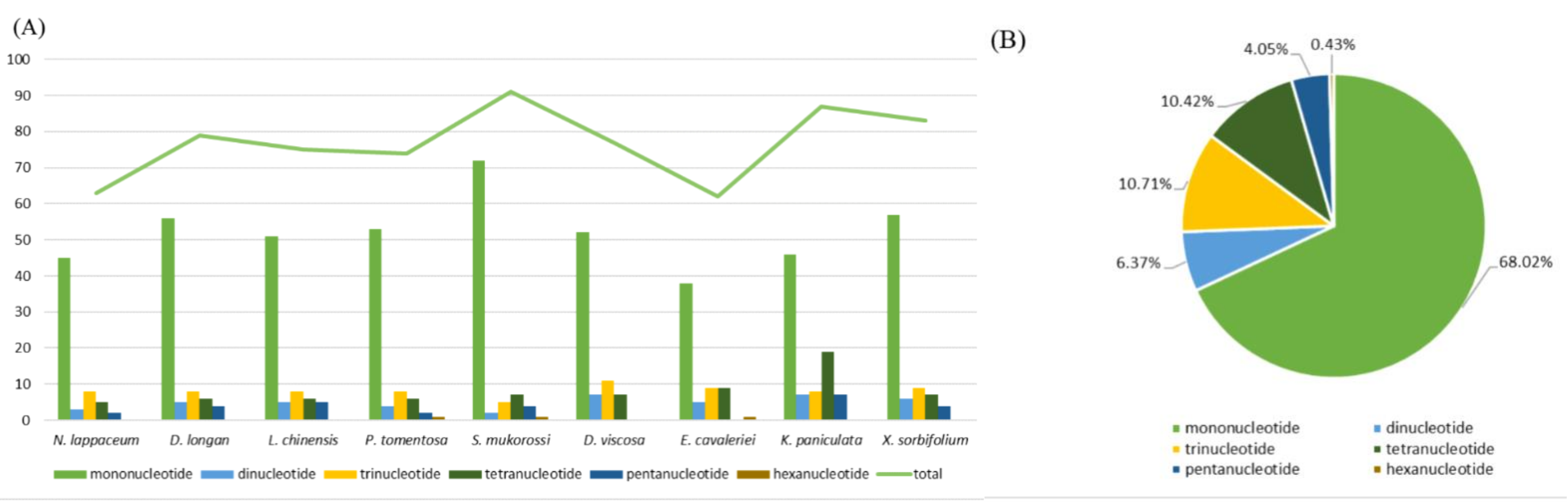
2.2. Characterization of SSRs and Repeat Sequences
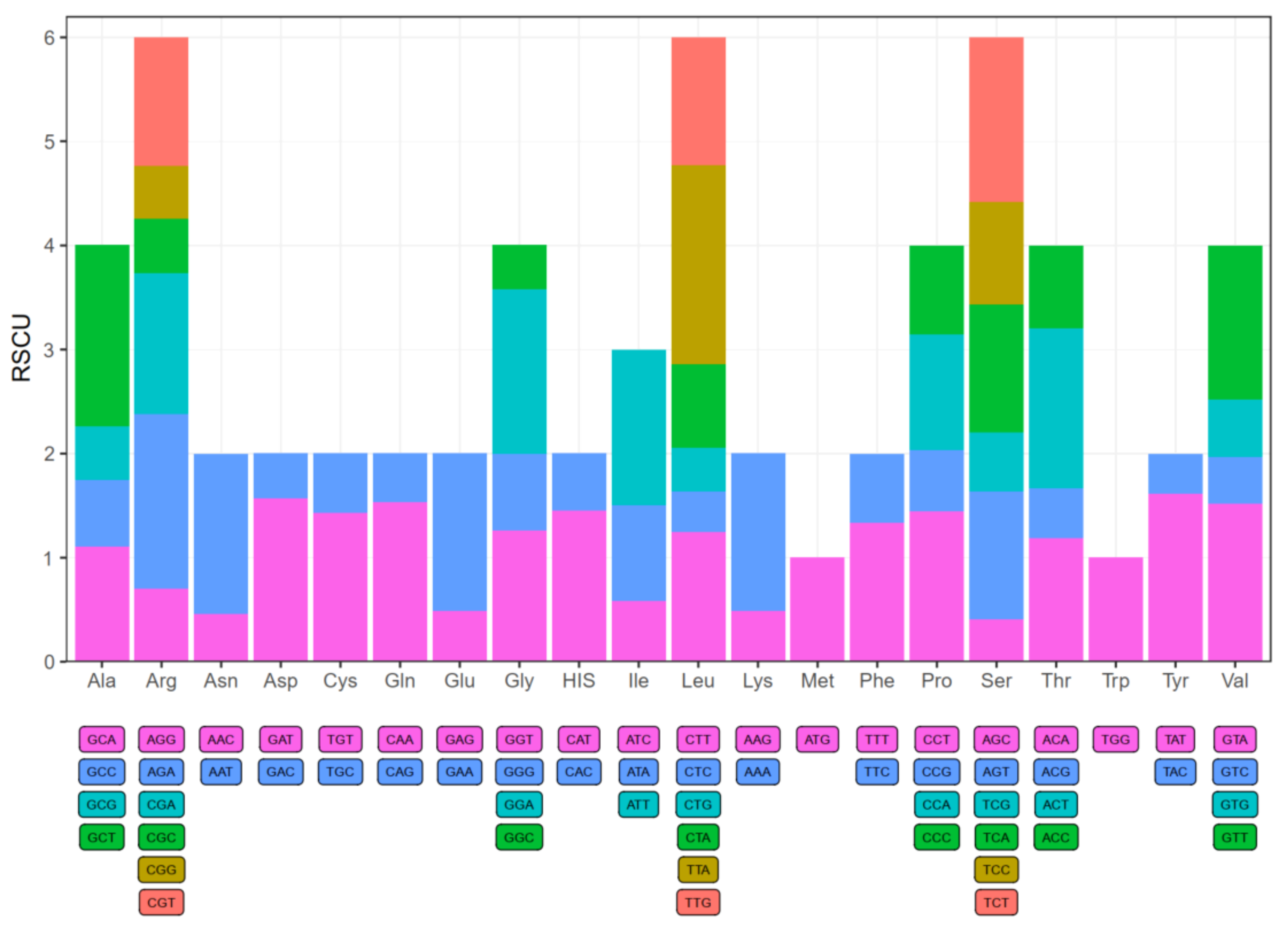
2.3. Codon Usage Analysis and RNA Editing Sites Prediction
2.4. Comparative Genomes Analysis
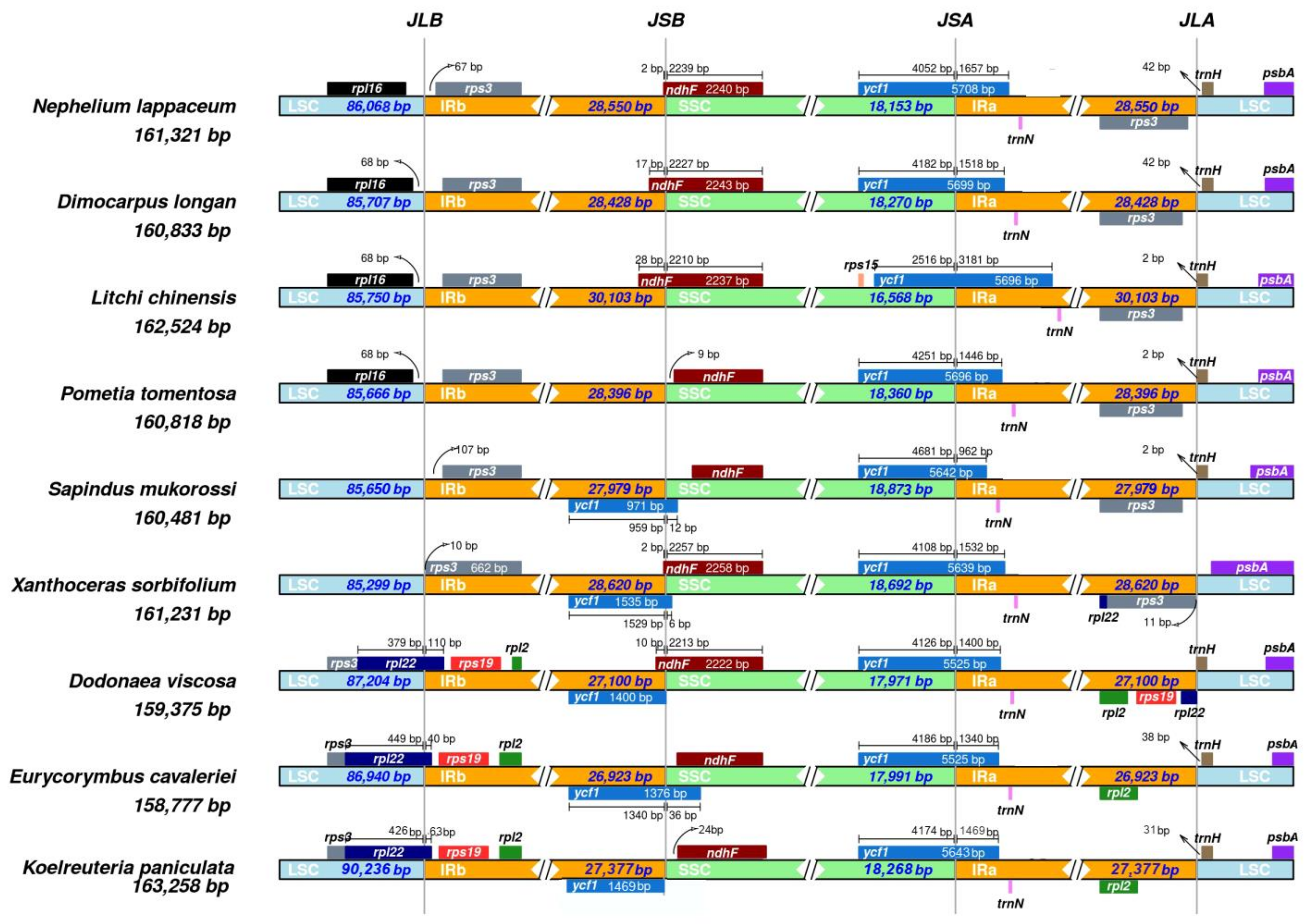
2.5. Expansion and Contraction of IR Regions
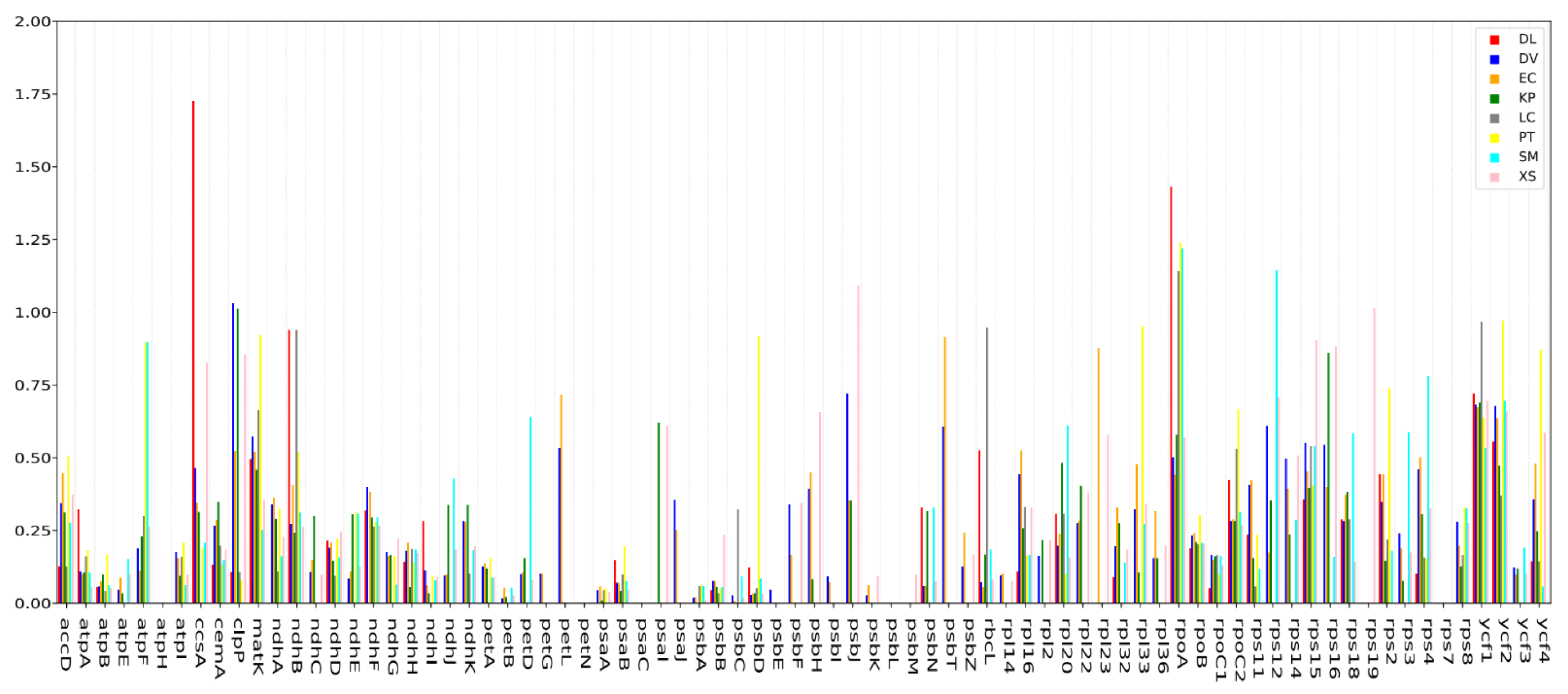
2.6. Synonymous (Ks) and Non-Synonymous (Ka) Substitution Rate Analysis
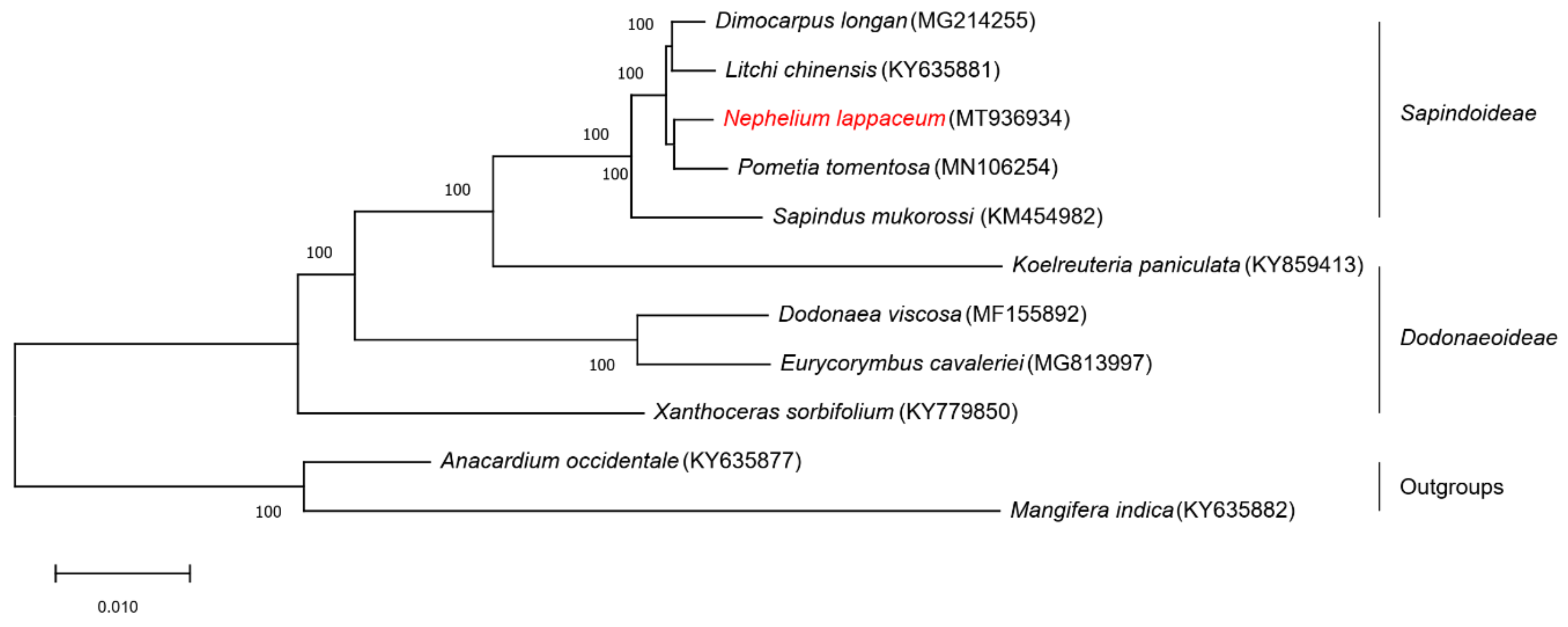
2.7. Phylogenetic Analysis
3. Discussion
4. Materials and Methods
4.1. Plant Material, DNA Extraction, and Sequencing
4.2. Chloroplast Genome Assembly and Annotation
4.3. Chloroplast Genome Analysis
4.4. Genome Comparison
4.5. Positive Selection Analysis of Protein Sequence
4.6. Phylogenetic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lim, T.K. Edible Medicinal and Non Medicinal Plants 2015; Springer: Berlin, Germany, 2012. [Google Scholar]
- Palanisamy, U.D.; Cheng, H.M.; Masilamani, T.; Subramaniam, T.; Ling, L.T.; Radhakrishnan, A.K. Rind of the rambutan, Nephelium lappaceum, a potential source of natural antioxidants. Food Chem. 2008, 109, 54–63. [Google Scholar] [CrossRef]
- Zhuang, Y.; Ma, Q.; Guo, Y.; Sun, L. Protective effects of rambutan (Nephelium lappaceum) peel phenolics on H2O2-induced oxidative damages in HepG2 cells and d-galactose-induced aging mice. Food Chem. Toxicol. 2017, 108, 554–562. [Google Scholar] [CrossRef]
- Phuong NN, M.; Le, T.T.; Van Camp, J.; Raes, K. Evaluation of antimicrobial activity of rambutan ( Nephelium lappaceum L.) peel extracts. Int. J. Food Microbiol. 2020, 321, 108539. [Google Scholar] [CrossRef]
- Harrington, M.G.; Edwards, K.J.; Johnson, S.A.; Chase, M.W.; Gadek, P.A. Phylogenetic Inference in Sapindaceae sensu lato Using Plastid matK and rbcL DNA Sequences. Syst. Bot. 2005, 30, 366–382. [Google Scholar] [CrossRef]
- Wicke, S.; Schneeweiss, G.M.; Depamphilis, C.W.; Kai, F.M.; Quandt, D. The evolution of the plastid chromosome in land plants: Gene content, gene order, gene function. Plant Mol. Biol. 2011, 76, 273–297. [Google Scholar]
- Bobik, K.; Burch-Smith, T.M. Chloroplast signaling within, between and beyond cells. Front. Plant Sci. 2015, 6, 781. [Google Scholar] [CrossRef]
- Wolfe, K.H.; Li, W.; Sharp, P.M. Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc. Natl. Acad. Sci. USA 1987, 84, 9054–9058. [Google Scholar] [CrossRef]
- Palmer, J.D. Comparative Organization of Chloroplast Genomes. Annu. Rev. Genet. 1985, 19, 325–354. [Google Scholar] [CrossRef]
- Shinozaki, K.; Ohme, M.; Tanaka, M.; Wakasugi, T.; Sugiura, M. The complete nucleotide sequence of the tobacco chloroplast genome: Its gene organization and expression. Plant Mol. Biol. Rep. 1986, 5, 2043–2049. [Google Scholar] [CrossRef]
- Li, C.; Lin, F.; An, D.; Wang, W.; Huang, R. Genome Sequencing and Assembly by Long Reads in Plants. Genes 2017, 9, 6. [Google Scholar] [CrossRef]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef]
- Marek, A.; Tomala, K. The contribution of purifying selection, linkage, and mutation bias to the negative correlation between gene expression and polymorphism density in yeast populations. Genome Biol. Evol. 2018, 10, 2986–2996. [Google Scholar] [CrossRef]
- Nguyen Dinh, S.; Sai, T.Z.T.; Nawaz, G.; Lee, K.; Kang, H. Abiotic stresses affect differently the intron splicing and expression of chloroplast genes in coffee plants (Coffea arabica) and rice (Oryza sativa). J. Plant Physiol. 2016, 201, 85–94. [Google Scholar] [CrossRef]
- Mirzaei, S.; Mansouri, M.; Mohammadi-Nejad, G.; Sablok, G. Comparative assessment of chloroplast transcriptional responses highlights conserved and unique patterns across Triticeae members under salt stress. Photosynth. Res. 2017, 136, 357–369. [Google Scholar] [CrossRef]
- Naver, H.; Boudreau, E.; Rochaix, J.D. Functional studies of YCF3: Its role in assembly of photosystem I and Interactions with some of its subunits. Plant Cell 2002, 13, 2731–2745. [Google Scholar] [CrossRef]
- Boudreau, E.; Takahashi, Y.; Lemieux, C.; Turmel, M.; Rochaix, J.D. The chloroplast ycf3 and ycf4 open reading frames of Chlamydomonas reinhardtii are required for the accumulation of the photosystem I complex. Embo. J. 1997, 16, 6095–6104. [Google Scholar] [CrossRef]
- Clarke, A.K.; Schelin, J.; Porankiewicz, J. Inactivation of the clpP1 gene for the proteolytic subunit of the ATP-dependent Clp protease in the cyanobacterium Synechococcus limits growth and light acclimation. Plant. Mol. Biol. 1998, 37, 791–801. [Google Scholar] [CrossRef]
- Bruce, C.A.; Cunningham, K.A.; Stern, D.B. The plastid clpP gene may not be essential for plant cell viability. Plant. Cell Physiol. 2003, 44, 93–95. [Google Scholar]
- Varshney, R.K.; Sigmund, R.; Borner, A.; Korzun, V.; Stein, N.; Sorrells, M.E.; Langridge, P.; Graner, A. Interspecific transferability and comparative mapping of barley EST-SSR markers in wheat, rye and rice. Plant. Sci. 2005, 168, 195–202. [Google Scholar] [CrossRef]
- Yang, A.; Zhang, J.; Tian, H.; Yao, X. Characterization of 39 novel EST-SSR markers for Liriodendron tulipifera and cross-species amplification in L. chinense (Magnoliaceae). Am. J. Bot. 2012, 99, e460–e464. [Google Scholar] [CrossRef]
- Li, B.; Lin, F.; Huang, P.; Guo, W.; Zheng, Y. Development of nuclear SSR and chloroplast genome markers in diverse Liriodendron chinense germplasm based on low-coverage whole genome sequencing. Biol. Res. 2020, 53, 21. [Google Scholar] [CrossRef]
- Gao, B.; Yuan, L.; Tang, T.; Hou, J.; Pan, K.; Wei, N. The complete chloroplast genome sequence of Alpinia oxyphylla Miq. and comparison analysis within the Zingiberaceae family. PLoS ONE 2019, 14. [Google Scholar] [CrossRef]
- Yan, C.; Du, J.; Gao, L.; Li, Y.; Hou, X. The complete chloroplast genome sequence of watercress (Nasturtium officinale R. Br.): Genome organization, adaptive evolution and phylogenetic relationships in Cardamineae. Gene 2019, 699, 24–36. [Google Scholar] [CrossRef]
- Cavalier-Smith, T. Chloroplast Evolution: Secondary Symbiogenesis and Multiple Losses. Curr. Biol. 2002, 12, R62–R64. [Google Scholar] [CrossRef]
- Hershberg, R.; Petrov, D.A. Selection on codon bias. Annu. Rev. Genet. 2008, 42, 287–299. [Google Scholar] [CrossRef]
- Raman, G.; Park, S.; Lee, E.M.; Park, S.J. Evidence of mitochondrial DNA in the chloroplast genome of Convallaria keiskei and its subsequent evolution in the Asparagales. Sci. Rep. 2019, 9, 5028. [Google Scholar] [CrossRef]
- Purabi, M.; Rofinayasmin, B.O.; Katharina, M.; Ramakrishnan, N.; Jennifer, A.H. Codon usage and codon pair patterns in non-grass monocot genomes. Ann. Bot. 2017, 120, 893–909. [Google Scholar]
- Wang, W.; Yu, H.; Wang, J.; Lei, W.; Gao, J.; Qiu, X.; Wang, J. The Complete Chloroplast Genome Sequences of the Medicinal Plant Forsythia suspensa (Oleaceae). Int. J. Mol. Sci. 2017, 18, 2288. [Google Scholar] [CrossRef]
- Smith, H.C.; Gott, J.M.; Hanson, M.R. A guide to RNA editing. RNA-Publ. RNA Soc. 1997, 3, 1105–1123. [Google Scholar]
- Hoch, B. Editing of a chloroplast mRNA by creation of an initiation codon. Nature 1991, 353, 178–180. [Google Scholar] [CrossRef] [PubMed]
- Maier, R.M.; Zeltz, P.; Kossel, H.; Bonnard, G.; Gualberto, J.M.; Grienenberger, J.M. RNA editing in plant mitochondria and chloroplasts. Plant. Mol. Biol. 1996, 32, 343–365. [Google Scholar] [CrossRef] [PubMed]
- Schmitzlinneweber, C.; Barkan, A. RNA splicing and RNA editing in chloroplasts. Top. Curr. Genet. 2007, 19, 213–248. [Google Scholar]
- Shikanai, T. RNA editing in plant organelles: Machinery, physiological function and evolution. Cell. Mol. Life Sci. 2006, 63, 698–708. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Xu, C.; Li, C.; Sun, J.; Zuo, Y.; Shi, S.; Cheng, T.; Guo, J.; Zhou, S. ycf1, the most promising plastid DNA barcode of land plants. Sci. Rep. 2015, 5, 8348. [Google Scholar] [CrossRef]
- Neubig, K.M.; Whitten, W.M.; Carlsward, B.S.; Blanco, M.A.; Endara, L.; Williams, N.H.; Moore, M. Phylogenetic utility of ycf1 in orchids: A plastid gene more variable than matK. Plant. Syst. Evol. 2009, 277, 75–84. [Google Scholar] [CrossRef]
- Dugas, D.V.; Hernandez, D.; Koenen, E.J.M.; Schwarz, E.; Straub, S.; Hughes, C.E.; Jansen, R.K.; Nageswara-Rao, M.; Staats, M.; Trujillo, J.T. Mimosoid legume plastome evolution: IR expansion, tandem repeat expansions, and accelerated rate of evolution in clpP. Sci. Rep. 2015, 5, 16958. [Google Scholar] [CrossRef]
- Yu, X.; Tan, W.; Zhang, H.; Gao, H.; Tian, X. Complete Chloroplast Genomes of Ampelopsis humulifolia and Ampelopsis japonica: Molecular Structure, Comparative Analysis, and Phylogenetic Analysis. Plants 2019, 8, 410. [Google Scholar] [CrossRef]
- Yang, Z.; Bielawski, J.P. Statistical methods for detecting molecular adaptation. Trends Ecol. Evol. 2000, 15, 496–503. [Google Scholar] [CrossRef]
- Swanson, W.J.; Wong, A.; Wolfner, M.F.; Aquadro, C.F. Evolutionary Expressed Sequence Tag Analysis of Drosophila Female Reproductive Tracts Identifies Genes Subjected to Positive Selection. Genetics 2004, 168, 1457–1465. [Google Scholar] [CrossRef]
- Gitzendanner, M.A.; Soltis, P.S.; Wong, G.K.S.; Ruhfel, B.R.; Soltis, D.E. Plastid phylogenomic analysis of green plants: A billion years of evolutionary history. Am. J. Bot. 2018, 105. [Google Scholar] [CrossRef]
- Du, Y.P.; Bi, Y.; Yang, F.P.; Zhang, M.F.; Zhang, X.H. Complete chloroplast genome sequences of Lilium: Insights into evolutionary dynamics and phylogenetic analyses. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Porebski, S.; Bailey, L.G.; Baum, B.R. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant. Mol. Biol. Rep. 1997, 15, 8–15. [Google Scholar] [CrossRef]
- Andrews, S. FastQC A Quality Control Tool for High Throughput Sequence Data. In Babraham Institute. 2015. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 1 December 2020).
- Nicolas, D.; Patrick, M.; Guillaume, S. NOVOPlasty: De novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 2017, 4, e18. [Google Scholar]
- Wang, K.; Li, L.; Zhao, M.; Li, S.; Sun, H.; Lv, Y.; Wang, Y. Characterization of the complete chloroplast genome of longan (Dimocarpus longan Lour.) using illumina paired-end sequencing. Mitochondrial DNA Part. B 2017, 2, 904–906. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.R.; Young, S. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef]
- Michael, T.; Pascal, L.; Tommaso, P.; Ulbricht-Jones, E.S.; Axel, F.; Ralph, B.; Stephan, G. GeSeq – versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 2017, 45, W6–W11. [Google Scholar]
- Shi, L.; Chen, H.; Jiang, M.; Wang, L.; Wu, X.; Huang, L.; Liu, C. CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res. 2019, 47, W65–W73. [Google Scholar] [CrossRef]
- Chan, P.P.; Lowe, T.M. tRNAscan-SE: Searching for tRNA Genes in Genomic Sequences. Methods Mol. Biol 2019, 1962, 1–14. [Google Scholar]
- Lohse, M.; Drechsel, O.; Bock, R. OrganellarGenomeDRAW (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef]
- Beier, S.; Thiel, T.; Munch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Wan, J.M. SSRHunter: Development of a local searching software for SSR sites. Yi Chuan 2005, 27, 808–810. [Google Scholar] [PubMed]
- Amiryousefi, A.; Hyvonen, J.; Poczai, P. IRscope: An online program to visualize the junction sites of chloroplast genomes. Bioinformatics 2018, 34, 3030–3031. [Google Scholar] [CrossRef] [PubMed]
- Afgan, E.; Baker, D.; Den Beek, M.V.; Blankenberg, D.; Bouvier, D.; Cech, M.; Chilton, J.; Clements, D.; Coraor, N.; Eberhard, C. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 2016, 44, W3–W10. [Google Scholar] [CrossRef]
- Wright, F. The effective number of codons used in a gene. Gene 1990, 87, 23–29. [Google Scholar] [CrossRef]
- Mower, J.P. The PREP suite: Predictive RNA editors for plant mitochondrial genes, chloroplast genes and user-defined alignments. Nucleic Acids Res. 2009, 37, 253–259. [Google Scholar] [CrossRef]
- Wang, Y.; Yuan, X.; Zhang, J. The complete chloroplast genome sequence of Pometia tomentosa. Mitochondrial DNA Part B 2019, 4, 3950–3951. [Google Scholar] [CrossRef]
- Yang, B.; Li, M.; Ma, J.; Fu, Z.; Tian, J. The complete chloroplast genome sequence of Sapindus mukorossi. Mitochondrial DNA Part A 2016, 27, 1825–1826. [Google Scholar] [CrossRef]
- Saina, J.K.; Gichira, A.W.; Li, Z.Z.; Hu, G.W.; Wang, Q.F.; Liao, K. The complete chloroplast genome sequence of Dodonaea viscosa: Comparative and phylogenetic analyses. Genetica 2017, 146, 101–113. [Google Scholar] [CrossRef]
- Du, X.; Xin, G.; Ren, X.; Liu, H.; Hao, N.; Jia, G.; Liu, W. The complete chloroplast genome of Eurycorymbus cavaleriei (Sapindaceae), a Tertiary relic species endemic to China. Conserv. Genet. Resour. 2019, 11, 283–285. [Google Scholar] [CrossRef]
- Kim, S.C.; Baek, S.H.; Hong, K.N.; Lee, J.W. Characterization of the complete chloroplast genome of Koelreuteria paniculata (Sapindaceae). Conserv. Genet. Resour. 2018, 10, 69–72. [Google Scholar] [CrossRef]
- Chen, S.Y.; Zhang, X.Z. Characterization of the complete chloroplast genome of Xanthoceras sorbifolium, an endangered oil tree. Conserv. Genet. Resour. 2017, 9, 595–598. [Google Scholar] [CrossRef]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef] [PubMed]
- Brudno, M.; Malde, S.; Poliakov, A.; Do, C.B.; Couronne, O.; Dubchak, I.; Batzoglou, S. Glocal alignment: Finding rearrangements during alignment. Bioinformatics 2003, 19, 54–62. [Google Scholar] [CrossRef]
- Zhang, Z.; Xiao, J.; Wu, J.; Zhang, H.; Liu, G.; Wang, X.; Dai, L. ParaAT: A parallel tool for constructing multiple protein-coding DNA alignments. Biochem. Biophys. Res. Commun. 2012, 419, 779–781. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A Toolkit Incorporating Gamma-Series Methods and Sliding Window Strategies. Genom. Proteom. Bioinform. 2010, 8, 77–80. [Google Scholar] [CrossRef]
- Nei, M.; Gojobori, T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol. Biol. Evol. 1986, 3, 418–426. [Google Scholar]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Cummings, M.P. PAUP* (Phylogenetic Analysis Using Parsimony (and Other Methods)). In Dictionary of Bioinformatics Computational Biology; John Wiley & Sons Inc.: Hoboken, NJ, USA, 2004. [Google Scholar]
- Azim, M.K.; Khan, I.A.; Zhang, Y. Characterization of mango (Mangifera indica L.) transcriptome and chloroplast genome. Plant. Mol. Biol. 2014, 85, 193–208. [Google Scholar] [CrossRef]


| Genome Feature | Dimocarpus longan | Litchi chinensis | Pometia tomentosa | Sapindus mukorossi | Nephelium lappaceum | Dodonaea viscosa | Eurycorymbus cavaleriei | Koelreuteria paniculata | Xanthoceras sorbifolium |
|---|---|---|---|---|---|---|---|---|---|
| GenBank | MG214255 | KY635881 | MN106254 | KM454982 | MT936934 | KM454982 | MF155892 | MG813997 | KY859413 |
| Size (bp) | 160,833 | 162,524 | 160,818 | 160,481 | 161,321 | 159,375 | 158,777 | 163,258 | 161,231 |
| LSC (bp) | 85,707 | 85,750 | 85,666 | 85,650 | 86,068 | 872,014 | 86,940 | 90,236 | 85,299 |
| SSC (bp) | 18,270 | 16,568 | 18,360 | 18,873 | 18,153 | 17,972 | 17,991 | 18,268 | 18,692 |
| IR (bp) | 28,428 | 30,103 | 28,396 | 27,979 | 28,550 | 27,099 | 26,923 | 27,377 | 28,620 |
| Total genes | 132 | 132 | 133 | 135 | 132 | 135 (2 Pseudogene) | 137 | 133 (3 Pseudogene) | 132 |
| Protein genes | 87 | 87 | 88 | 88 | 87 | 88 | 89 | 85 | 86 |
| tRNA genes | 37 | 37 | 37 | 39 | 37 | 37 | 40 | 37 | 38 |
| rRNA genes | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 |
| GC (%) | 37.79% | 37.80% | 37.87% | 37.66% | 37.77% | 37.86% | 37.92% | 37.30% | 37.69% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, F.; Lin, Z.; Lin, J.; Ming, R.; Zhang, W. Chloroplast Genome of Rambutan and Comparative Analyses in Sapindaceae. Plants 2021, 10, 283. https://doi.org/10.3390/plants10020283
Dong F, Lin Z, Lin J, Ming R, Zhang W. Chloroplast Genome of Rambutan and Comparative Analyses in Sapindaceae. Plants. 2021; 10(2):283. https://doi.org/10.3390/plants10020283
Chicago/Turabian StyleDong, Fei, Zhicong Lin, Jing Lin, Ray Ming, and Wenping Zhang. 2021. "Chloroplast Genome of Rambutan and Comparative Analyses in Sapindaceae" Plants 10, no. 2: 283. https://doi.org/10.3390/plants10020283
APA StyleDong, F., Lin, Z., Lin, J., Ming, R., & Zhang, W. (2021). Chloroplast Genome of Rambutan and Comparative Analyses in Sapindaceae. Plants, 10(2), 283. https://doi.org/10.3390/plants10020283





