Triterpenoid and Steroidal Saponins Differentially Influence Soil Bacterial Genera
Abstract
:1. Introduction
2. Materials and Methods
2.1. Chemicals and Soils
2.2. Treatment of Field Soil with Saponins
2.3. DNA Extraction and 16S rRNA Amplicon Sequencing
2.4. Sequence Data Analysis
2.5. Statistical Analysis
3. Results
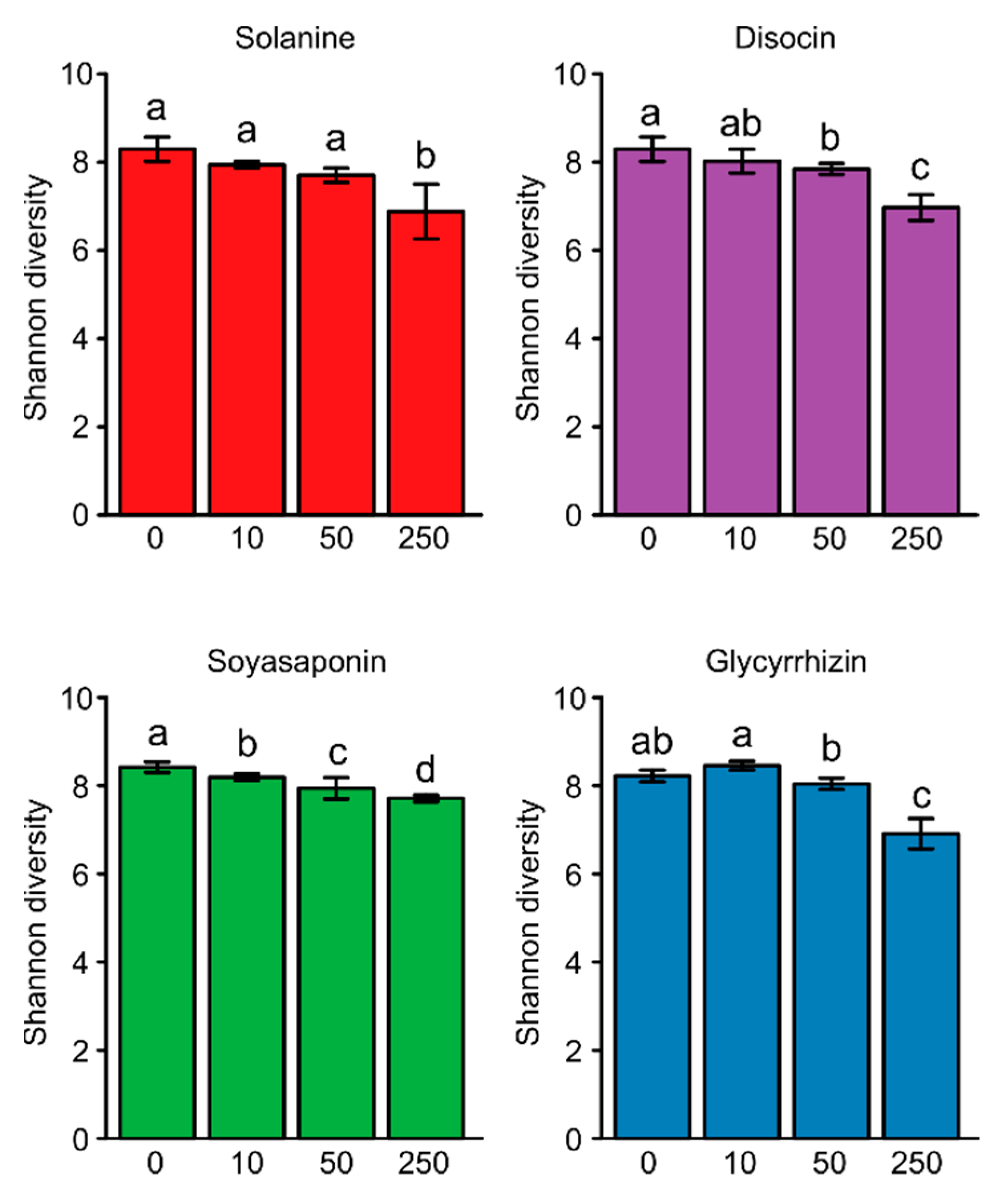
3.1. Bacterial Diversity in the Saponin-Treated Soils
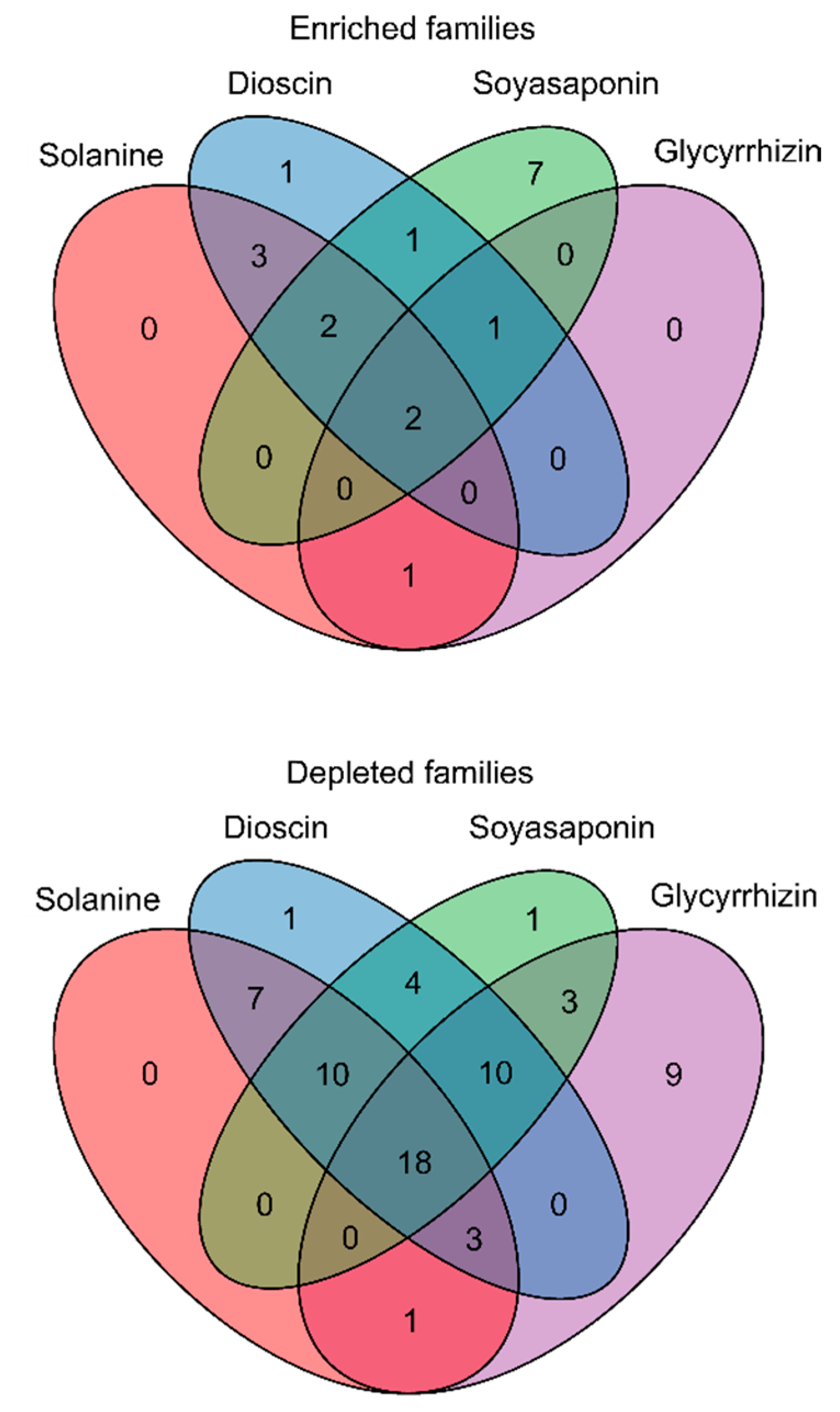
3.2. Effects of the Saponin Treatments on Bacterial Families in the Soil
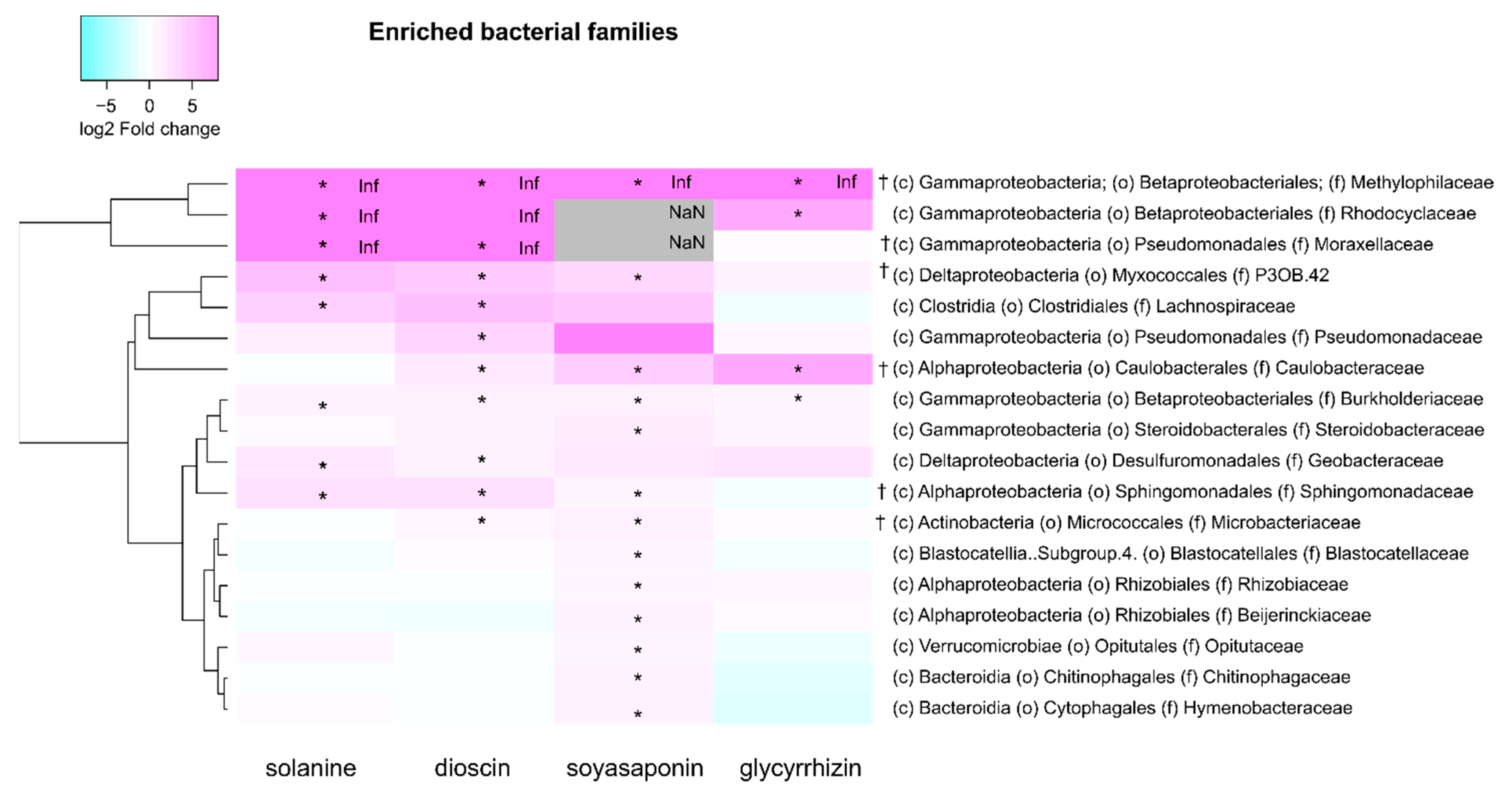
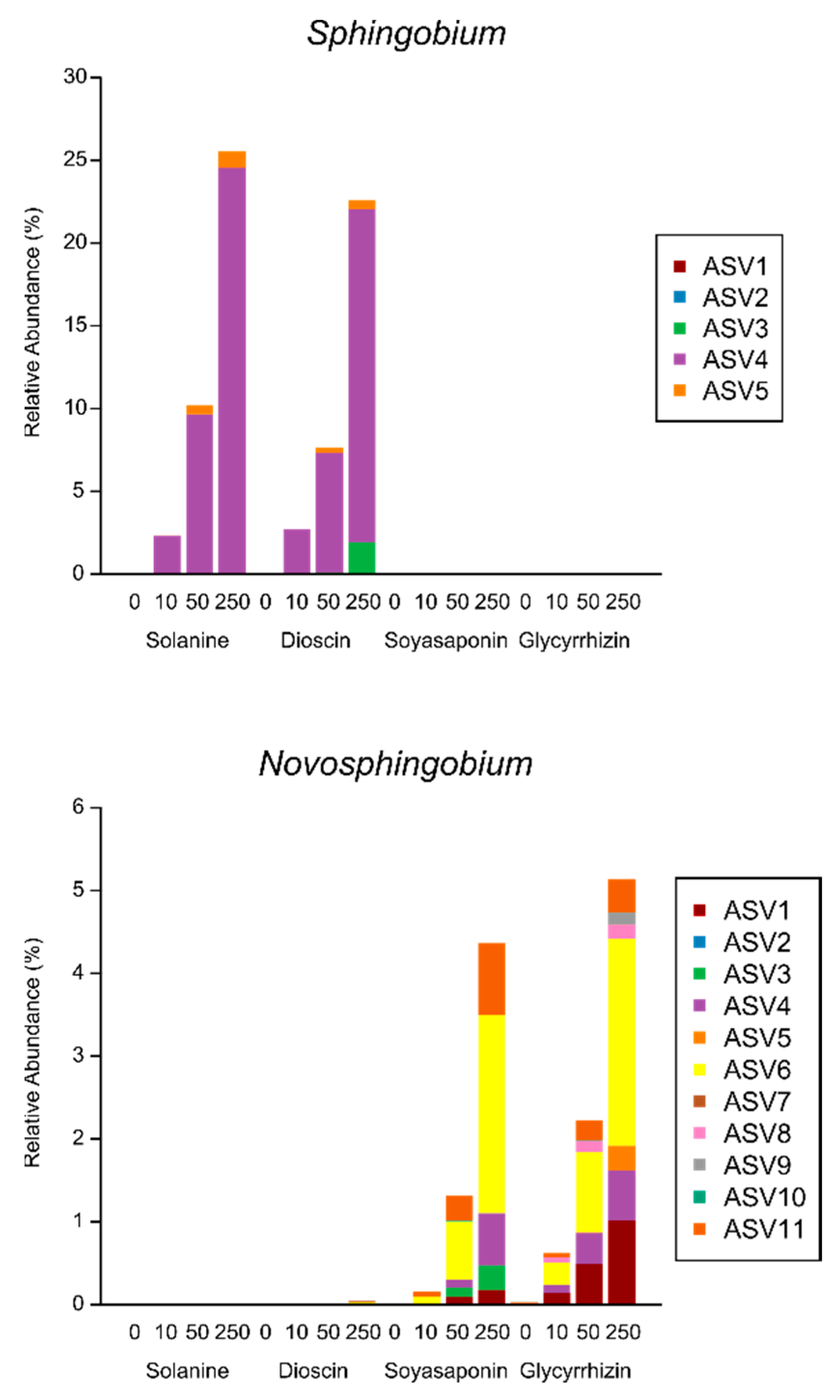
3.3. Taxonomic Composition of the Bacterial Families Enriched by the Saponin Treatments
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Afendi, F.M.; Okada, T.; Yamazaki, M.; Hirai-Morita, A.; Nakamura, Y.; Nakamura, K.; Ikeda, S.; Takahashi, H.; Altaf-Ul-Amin, M.; Darusman, L.K.; et al. KNApSAcK Family Databases: Integrated Metabolite-Plant Species Databases for Multifaceted Plant Research. Plant Cell Physiol. 2012, 53, e1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pichersky, E.; Lewinsohn, E. Convergent Evolution in Plant Specialized Metabolism. Annu. Rev. Plant Biol. 2011, 62, 549–566. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guerrieri, A.; Dong, L.M.; Bouwmeester, H.J. Role and exploitation of underground chemical signaling in plants. Pest Manag. Sci. 2019, 75, 2455–2463. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Massalha, H.; Korenblum, E.; Tholl, D.; Aharoni, A. Small molecules below-ground: The role of specialized metabolites in the rhizosphere. Plant J. 2017, 90, 788–807. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hartmann, A.; Rothballer, M.; Schmid, M. Lorenz Hiltner, a pioneer in rhizosphere microbial ecology and soil bacteriology research. Plant Soil 2008, 312, 7–14. [Google Scholar] [CrossRef]
- Hiltner, L. Uber nevere Erfahrungen und Probleme auf dem Gebiet der Boden Bakteriologie und unter besonderer Beurchsichtigung der Grundungung und Broche. Arbeit. Deut. Landw. Ges. Berl. 1904, 98, 59–78. [Google Scholar]
- Badri, D.V.; Vivanco, J.M. Regulation and function of root exudates. Plant Cell Environ. 2009, 32, 666–681. [Google Scholar] [CrossRef]
- Mendes, R.; Garbeva, P.; Raaijmakers, J.M. The rhizosphere microbiome: Significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol. Rev. 2013, 37, 634–663. [Google Scholar] [CrossRef]
- Yu, P.; He, X.; Baer, M.; Beirinckx, S.; Tian, T.; Moya, Y.A.T.; Zhang, X.; Deichmann, M.; Frey, F.P.; Bresgen, V.; et al. Plant flavones enrich rhizosphere Oxalobacteraceae to improve maize performance under nitrogen deprivation. Nat. Plants 2021, 7, 481–499. [Google Scholar] [CrossRef] [PubMed]
- Pascale, A.; Proietti, S.; Pantelides, I.S.; Stringlis, I.A. Modulation of the Root Microbiome by Plant Molecules: The Basis for Targeted Disease Suppression and Plant Growth Promotion. Front. Plant Sci. 2020, 10, 1741. [Google Scholar] [CrossRef] [PubMed]
- Jacoby, R.P.; Koprivova, A.; Kopriva, S. Pinpointing secondary metabolites that shape the composition and function of the plant microbiome. J. Exp. Bot. 2021, 72, 57–69. [Google Scholar] [CrossRef]
- Chen, Q.W.; Jiang, T.; Liu, Y.X.; Liu, H.L.; Zhao, T.; Liu, Z.X.; Gan, X.C.; Hallab, A.; Wang, X.M.; He, J.; et al. Recently duplicated sesterterpene (C25) gene clusters in Arabidopsis thaliana modulate root microbiota. Sci. China-Life Sci. 2019, 62, 947–958. [Google Scholar] [CrossRef]
- Harbort, C.J.; Hashimoto, M.; Inoue, H.; Niu, Y.; Guan, R.; Rombola, A.D.; Kopriva, S.; Voges, M.J.E.E.E.; Sattely, E.S.; Garrido-Oter, R.; et al. Root-Secreted Coumarins and the Microbiota Interact to Improve Iron Nutrition in Arabidopsis. Cell Host Microbe 2020, 28, 825–837. [Google Scholar] [CrossRef]
- Hu, L.F.; Robert, C.A.M.; Cadot, S.; Zhang, X.; Ye, M.; Li, B.B.; Manzo, D.; Chervet, N.; Steinger, T.; van der Heijden, M.G.A.; et al. Root exudate metabolites drive plant-soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat. Commun. 2018, 9, 13. [Google Scholar] [CrossRef] [Green Version]
- Huang, A.C.C.; Jiang, T.; Liu, Y.X.; Bai, Y.C.; Reed, J.; Qu, B.Y.; Goossens, A.; Nutzmann, H.W.; Bai, Y.; Osbourn, A. A specialized metabolic network selectively modulates Arabidopsis root microbiota. Science 2019, 364, eaau6389. [Google Scholar] [CrossRef] [PubMed]
- Murphy, K.M.; Edwards, J.; Louie, K.B.; Bowen, B.P.; Sundaresan, V.; Northen, T.R.; Zerbe, P. Bioactive diterpenoids impact the composition of the root-associated microbiome in maize (Zea mays). Sci. Rep. 2021, 11, 333. [Google Scholar] [CrossRef]
- Stringlis, I.A.; Yu, K.; Feussner, K.; de Jonge, R.; van Bentum, S.; Van Verk, M.C.; Berendsen, R.L.; Bakker, P.; Feussner, I.; Pieterse, C.M.J. MYB72-dependent coumarin exudation shapes root microbiome assembly to promote plant health. Proc. Natl. Acad. Sci. USA 2018, 115, E5213–E5222. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Voges, M.; Bai, Y.; Schulze-Lefert, P.; Sattely, E.S. Plant-derived coumarins shape the composition of an Arabidopsis synthetic root microbiome. Proc. Natl. Acad. Sci. USA 2019, 116, 12558–12565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maver, M.; Escudero-Martinez, C.; Abbott, J.; Morris, J.; Hedley, P.E.; Mimmo, T.; Bulgarelli, D. Applications of the indole-alkaloid gramine shape the prokaryotic microbiota thriving at the barley root-soil interface. bioRxiv 2021. [Google Scholar] [CrossRef]
- Okutani, F.; Hamamoto, S.; Aoki, Y.; Nakayasu, M.; Nihei, N.; Nishimura, T.; Yazaki, K.; Sugiyama, A. Rhizosphere modelling reveals spatiotemporal distribution of daidzein shaping soybean rhizosphere bacterial community. Plant Cell Environ. 2020, 43, 1036–1046. [Google Scholar] [CrossRef]
- Shimasaki, T.; Masuda, S.; Garrido-Oter, R.; Kawasaki, T.; Aoki, Y.; Shibata, A.; Suda, W.; Shirasu, K.; Yazaki, K.; Nakano, R.T.; et al. Tobacco Root Endophytic Arthrobacter Harbors Genomic Features Enabling the Catabolism of Host-Specific Plant Specialized Metabolites. mBio 2021, 12, e0084621. [Google Scholar] [CrossRef]
- Schütz, V.; Frindte, K.; Cui, J.; Zhang, P.; Hacquard, S.; Schulze-Lefert, P.; Knief, C.; Schulz, M.; Doermann, P. Differential Impact of Plant Secondary Metabolites on the Soil Microbiota. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Sparg, S.G.; Light, M.E.; van Staden, J. Biological activities and distribution of plant saponins. J. Ethnopharmacol. 2004, 94, 219–243. [Google Scholar] [CrossRef] [PubMed]
- Kitagawa, I. Licorice root. A-natural sweetener and an important ingredient in Chinese medicine. Pure Appl. Chem. 2002, 74, 1189–1198. [Google Scholar] [CrossRef] [Green Version]
- Sautour, M.; Mitaine-Offer, A.-C.; Lacaille-Dubois, M.-A. The Dioscorea genus: A review of bioactive steroid saponins. J. Nat. Med. 2007, 61, 91–101. [Google Scholar] [CrossRef]
- Friedman, M. Potato glycoalkaloids and metabolites: Roles in the plant and in the diet. J. Agric. Food Chem. 2006, 54, 8655–8681. [Google Scholar] [CrossRef] [PubMed]
- Cardenas, P.D.; Almeida, A.; Bak, S. Evolution of Structural Diversity of Triterpenoids. Front. Plant Sci. 2019, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Moses, T.; Papadopoulou, K.K.; Osbourn, A. Metabolic and functional diversity of saponins, biosynthetic intermediates and semi-synthetic derivatives. Crit. Rev. Biochem. Mol. Biol. 2014, 49, 439–462. [Google Scholar] [CrossRef] [Green Version]
- Sawai, S.; Saito, K. Triterpenoid biosynthesis and engineering in plants. Front. Plant Sci. 2011, 2, 25. [Google Scholar] [CrossRef] [Green Version]
- Seki, H.; Tamura, K.; Muranaka, T. P450s and UGTs: Key Players in the Structural Diversity of Triterpenoid Saponins. Plant Cell Physiol. 2015, 56, 1463–1471. [Google Scholar] [CrossRef]
- Kirwa, H.K.; Murungi, L.K.; Beck, J.J.; Torto, B. Elicitation of Differential Responses in the Root-Knot Nematode Meloidogyne incognita to Tomato Root Exudate Cytokinin, Flavonoids, and Alkaloids. J. Agric. Food Chem. 2018, 66, 11291–11300. [Google Scholar] [CrossRef]
- Rial, C.; Gomez, E.; Varela, R.M.; Molinillo, J.M.G.; Macias, F.A. Ecological Relevance of the Major Allelochemicals in Lycopersicon esculentum Roots and Exudates. J. Agric. Food Chem. 2018, 66, 4638–4644. [Google Scholar] [CrossRef]
- Korenblum, E.; Dong, Y.; Szymanski, J.; Panda, S.; Jozwiak, A.; Massalha, H.; Meir, S.; Rogachev, I.; Aharoni, A. Rhizosphere microbiome mediates systemic root metabolite exudation by root-to-root signaling. Proc. Natl. Acad. Sci. USA 2020, 117, 3874–3883. [Google Scholar] [CrossRef]
- Sugiyama, A. Flavonoids and saponins in plant rhizospheres: Roles, dynamics, and the potential for agriculture. Biosci. Biotechnol. Biochem. 2021. [Google Scholar] [CrossRef] [PubMed]
- Fujimatsu, T.; Endo, K.; Yazaki, K.; Sugiyama, A. Secretion dynamics of soyasaponins in soybean roots and effects to modify the bacterial composition. Plant Direct 2020, 4, e00259. [Google Scholar] [CrossRef]
- Tsuno, Y.; Fujimatsu, T.; Endo, K.; Sugiyama, A.; Yazaki, K. Soyasaponins: A New Class of Root Exudates in Soybean (Glycine max). Plant Cell Physiol. 2018, 59, 366–375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakayasu, M.; Ohno, K.; Takamatsu, K.; Aoki, Y.; Yamazaki, S.; Takase, H.; Shoji, T.; Yazaki, K.; Sugiyama, A. Tomato roots secrete tomatine to modulate the bacterial assemblage of the rhizosphere. Plant Physiol. 2021, 186, 270–284. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108, 4516–4522. [Google Scholar] [CrossRef] [Green Version]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [Green Version]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [Green Version]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree: Computing Large Minimum Evolution Trees with Profiles instead of a Distance Matrix. Mol. Biol. Evol. 2009, 26, 1641–1650. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Kaehler, B.D.; Rideout, J.R.; Dillon, M.; Bolyen, E.; Knight, R.; Huttley, G.A.; Caporaso, J.G. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2′s q2-feature-classifier plugin. Microbiome 2018, 6, 90. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Blanchet, F.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.; O’Hara, R.; Simpson, G.; Solymos, P.; et al. Vegan: Community Ecology Package. R Package Version 2.5-7. 2020. Available online: https://CRAN.R-project.org/package=vegan (accessed on 13 October 2021).
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Ma, B.; Chen, W.; Schlaeppi, K.; Erb, M.; Stirling, E.; Hu, L.; Wang, E.; Zhang, Y.; Zhao, K. The Symbiotic Capacity of Rhizobium Shapes Root-Associated Microbiomes. Res. Sq. 2020. Available online: https://assets.researchsquare.com/files/rs-41035/v1/de4ef2de-8b8d-4fea-bc73-2eaa09424d40.pdf?c=1631845632 (accessed on 13 October 2021).
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; van Themaat, E.V.L.; Schulze-Lefert, P. Structure and Functions of the Bacterial Microbiota of Plants. Annu. Rev. Plant Biol. 2013, 64, 807–838. [Google Scholar] [CrossRef] [Green Version]
- Duran, P.; Thiergart, T.; Garrido-Oter, R.; Agler, M.; Kemen, E.; Schulze-Lefert, P.; Hacquard, S. Microbial Interkingdom Interactions in Roots Promote Arabidopsis Survival. Cell 2018, 175, 973–983. [Google Scholar] [CrossRef] [PubMed]
- Cotton, T.E.A.; Petriacq, P.; Cameron, D.D.; Al Meselmani, M.; Schwarzenbacher, R.; Rolfe, S.A.; Ton, J. Metabolic regulation of the maize rhizobiome by benzoxazinoids. ISME J. 2019, 13, 1647–1658. [Google Scholar] [CrossRef] [Green Version]
- Kudjordjie, E.N.; Sapkota, R.; Steffensen, S.K.; Fomsgaard, I.S.; Nicolaisen, M. Maize synthesized benzoxazinoids affect the host associated microbiome. Microbiome 2019, 7, 59. [Google Scholar] [CrossRef] [PubMed]
- Tsurumaru, H.; Okubo, T.; Okazaki, K.; Hashimoto, M.; Kakizaki, K.; Hanzawa, E.; Takahashi, H.; Asanome, N.; Tanaka, F.; Sekiyama, Y.; et al. Metagenomic Analysis of the Bacterial Community Associated with the Taproot of Sugar Beet. Microbes Environ. 2015, 30, 63–69. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mikolajczyk-Bator, K.; Blaszczyk, A.; Czyzniejewski, M.; Kachlicki, P. Identification of saponins from sugar beet (Beta vulgaris) by low and high-resolution HPLC-MS/MS. J. Chromatogr. B-Anal. Technol. Biomed. Life Sci. 2016, 1029, 36–47. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Chen, Z.; Wang, B.; Wei, F.; Zhang, G.; Wang, Y.; Zhu, G.; Zhou, Y.; Zhao, Q.; He, M.; et al. Endophytes isolated from Panax notoginseng converted ginsenosides. Microb. Biotechnol. 2021, 14, 1730–1746. [Google Scholar] [CrossRef]
- Mardanova, A.; Lutfullin, M.; Hadieva, G.; Akosah, Y.; Pudova, D.; Kabanov, D.; Shagimardanova, E.; Vankov, P.; Vologin, S.; Gogoleva, N.; et al. Structure and variation of root-associated microbiomes of potato grown in alfisol. World J. Microbiol. Biotechnol. 2019, 35, 181. [Google Scholar] [CrossRef]
- Herpell, J.B.; Schindler, F.; Bejtovic, M.; Fragner, L.; Diallo, B.; Bellaire, A.; Kublik, S.; Foesel, B.U.; Gschwendtner, S.; Kerou, M.; et al. The Potato Yam Phyllosphere Ectosymbiont Paraburkholderia sp. Msb3 Is a Potent Growth Promotor in Tomato. Front. Microbiol. 2020, 11, 581. [Google Scholar] [CrossRef] [Green Version]
- Liu, H.; Chou, G.-X.; Wu, T.; Guo, Y.-L.; Wang, S.-C.; Wang, C.-H.; Wang, Z.-T. Steroidal Sapogenins and Glycosides from the Rhizomes of Dioscorea bulbifera. J. Nat. Prod. 2009, 72, 1964–1968. [Google Scholar] [CrossRef]
- Lee, S.T.; Mitchell, R.B.; Wang, Z.; Heiss, C.; Gardner, D.R.; Azadi, P. Isolation, Characterization, and Quantification of Steroidal Saponins in Switchgrass (Panicum virgatum L.). J. Agric. Food Chem. 2009, 57, 2599–2604. [Google Scholar] [CrossRef]
- Singer, E.; Bonnette, J.; Kenaley, S.C.; Woyke, T.; Juenger, T.E. Plant compartment and genetic variation drive microbiome composition in switchgrass roots. Environ. Microbiol. Rep. 2019, 11, 185–195. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Mohamad, O.A.A.; Ma, J.; Friel, A.D.; Su, Y.; Wang, Y.; Musa, Z.; Liu, Y.; Hedlund, B.P.; Li, W. Synergistic plant-microbe interactions between endophytic bacterial communities and the medicinal plant Glycyrrhiza uralensis F. Antonie Van Leeuwenhoek Int. J. Gen. Mol. Microbiol. 2018, 111, 1735–1748. [Google Scholar] [CrossRef]
- Holert, J.; Cardenas, E.; Bergstrand, L.H.; Zaikova, E.; Hahn, A.S.; Hallam, S.J.; Mohn, W.W. Metagenomes Reveal Global Distribution of Bacterial Steroid Catabolism in Natural, Engineered, and Host Environments. Mbio 2018, 9, e02345-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Belic, I.; Hirslpintaric, V.; Socic, H.; Vranjek, B. Microbial cleavage of tomatidine spiroketal sidechain. J. Steroid Biochem. Mol. Biol. 1975, 6, 1211–1212. [Google Scholar] [CrossRef]
- Belic, I.; Socic, H. Microbial dehydrogenation of tomatidine. J. Steroid Biochem. 1972, 3, 843–846. [Google Scholar] [CrossRef]
- Cheng, L.-Q.; Kim, M.K.; Lee, J.-W.; Lee, Y.-J.; Yang, D.-C. Conversion of major ginsenoside Rb-1 to ginsenoside F-2 by Caulobacter leidyia. Biotechnol. Lett. 2006, 28, 1121–1127. [Google Scholar] [CrossRef]
- Kim, J.-K.; He, D.; Liu, Q.-M.; Park, H.-Y.; Jung, M.-S.; Yoon, M.-H.; Kim, S.-C.; Im, W.-T. Novosphingobium ginsenosidimutans sp nov., with the Ability to Convert Ginsenoside. J. Microbiol. Biotechnol. 2013, 23, 444–450. [Google Scholar] [CrossRef] [Green Version]
- Fu, Y.; Yin, Z.H.; Yin, C.Y. Biotransformation of ginsenoside Rb1 to ginsenoside Rg3 by endophytic bacterium Burkholderia sp GE 17-7 isolated from Panax ginseng. J. Appl. Microbiol. 2017, 122, 1579–1585. [Google Scholar] [CrossRef]
- Zhang, S.; Shu, J.; Xue, H.; Zhang, W.; Zhang, Y.; Liu, Y.; Fang, L.; Wang, Y.; Wang, H. The Gut Microbiota in Camellia Weevils Are Influenced by Plant Secondary Metabolites and Contribute to Saponin Degradation. Msystems 2020, 5, e00692-19. [Google Scholar] [CrossRef] [Green Version]
- Carrion, V.J.; Cordovez, V.; Tyc, O.; Etalo, D.W.; de Bruijn, I.; de Jager, V.C.L.; Medema, M.H.; Eberl, L.; Raaijmakers, J.M. Involvement of Burkholderiaceae and sulfurous volatiles in disease-suppressive soils. ISME J. 2018, 12, 2307–2321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gu, S.; Wei, Z.; Shao, Z.; Friman, V.-P.; Cao, K.; Yang, T.; Kramer, J.; Wang, X.; Li, M.; Mei, X.; et al. Competition for iron drives phytopathogen control by natural rhizosphere microbiomes. Nat. Microbiol. 2020, 5, 1002–1010. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xu, J.; Riera, N.; Jin, T.; Li, J.; Wang, N. Huanglongbing impairs the rhizosphere-to-rhizoplane enrichment process of the citrus root-associated microbiome. Microbiome 2017, 5, 97. [Google Scholar] [CrossRef] [PubMed]
- Innerebner, G.; Knief, C.; Vorholt, J.A. Protection of Arabidopsis thaliana against Leaf-Pathogenic Pseudomonas syringae by Sphingomonas Strains in a Controlled Model System. Appl. Environ. Microbiol. 2011, 77, 3202–3210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vogel, C.; Innerebner, G.; Zingg, J.; Guder, J.; Vorholt, J.A. Forward Genetic In Planta Screen for Identification of Plant-Protective Traits of Sphingomonas sp Strain Fr1 against Pseudomonas syringae DC3000. Appl. Environ. Microbiol. 2012, 78, 5529–5535. [Google Scholar] [CrossRef] [Green Version]
- Khan, A.L.; Waqas, M.; Kang, S.M.; Al-Harrasi, A.; Hussain, J.; Al-Rawahi, A.; Al-Khiziri, S.; Ullah, I.; Ali, L.; Jung, H.Y.; et al. Bacterial Endophyte Sphingomonas sp LK11 Produces Gibberellins and IAA and Promotes Tomato Plant Growth. J. Microbiol. 2014, 52, 689–695. [Google Scholar] [CrossRef]
- Pan, F.S.; Meng, Q.; Wang, Q.; Luo, S.; Chen, B.; Khan, K.Y.; Yang, X.E.; Feng, Y. Endophytic bacterium Sphingomonas SaMR12 promotes cadmium accumulation by increasing glutathione biosynthesis in Sedum alfredii Hance. Chemosphere 2016, 154, 358–366. [Google Scholar] [CrossRef]
- Asaf, S.; Khan, A.L.; Khan, M.A.; Imran, Q.M.; Yun, B.W.; Lee, I.J. Osmoprotective functions conferred to soybean plants via inoculation with Sphingomonas sp LK11 and exogenous trehalose. Microbiol. Res. 2017, 205, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Santiago, C.D.; Yagi, S.; Ijima, M.; Nashimoto, T.; Sawada, M.; Ikeda, S.; Asano, K.; Orikasa, Y.; Ohwada, T. Bacterial Compatibility in Combined Inoculations Enhances the Growth of Potato Seedlings. Microbes Environ. 2017, 32, 14–23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, Y.; Wang, F.; Huang, Y.L.; Zhou, M.; Gao, J.L.; Yan, T.Z.; Sheng, H.M.; An, L.Z. Sphingomonas sp. Cra20 Increases Plant Growth Rate and Alters Rhizosphere Microbial Community Structure of Arabidopsis thaliana Under Drought Stress. Front. Microbiol. 2019, 10, 19. [Google Scholar] [CrossRef] [PubMed] [Green Version]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nakayasu, M.; Yamazaki, S.; Aoki, Y.; Yazaki, K.; Sugiyama, A. Triterpenoid and Steroidal Saponins Differentially Influence Soil Bacterial Genera. Plants 2021, 10, 2189. https://doi.org/10.3390/plants10102189
Nakayasu M, Yamazaki S, Aoki Y, Yazaki K, Sugiyama A. Triterpenoid and Steroidal Saponins Differentially Influence Soil Bacterial Genera. Plants. 2021; 10(10):2189. https://doi.org/10.3390/plants10102189
Chicago/Turabian StyleNakayasu, Masaru, Shinichi Yamazaki, Yuichi Aoki, Kazufumi Yazaki, and Akifumi Sugiyama. 2021. "Triterpenoid and Steroidal Saponins Differentially Influence Soil Bacterial Genera" Plants 10, no. 10: 2189. https://doi.org/10.3390/plants10102189
APA StyleNakayasu, M., Yamazaki, S., Aoki, Y., Yazaki, K., & Sugiyama, A. (2021). Triterpenoid and Steroidal Saponins Differentially Influence Soil Bacterial Genera. Plants, 10(10), 2189. https://doi.org/10.3390/plants10102189





