Karyotype Reorganization in Wheat–Rye Hybrids Obtained via Unreduced Gametes: Is There a Limit to the Chromosome Number in Triticale?
Abstract
:1. Introduction
2. Materials and Methods
2.1. Routine Meiosis Analysis
2.2. Giemsa C-banding
2.3. Fluorescence In Situ Hybridization (FISH)
2.4. Statistical Analysis
3. Results
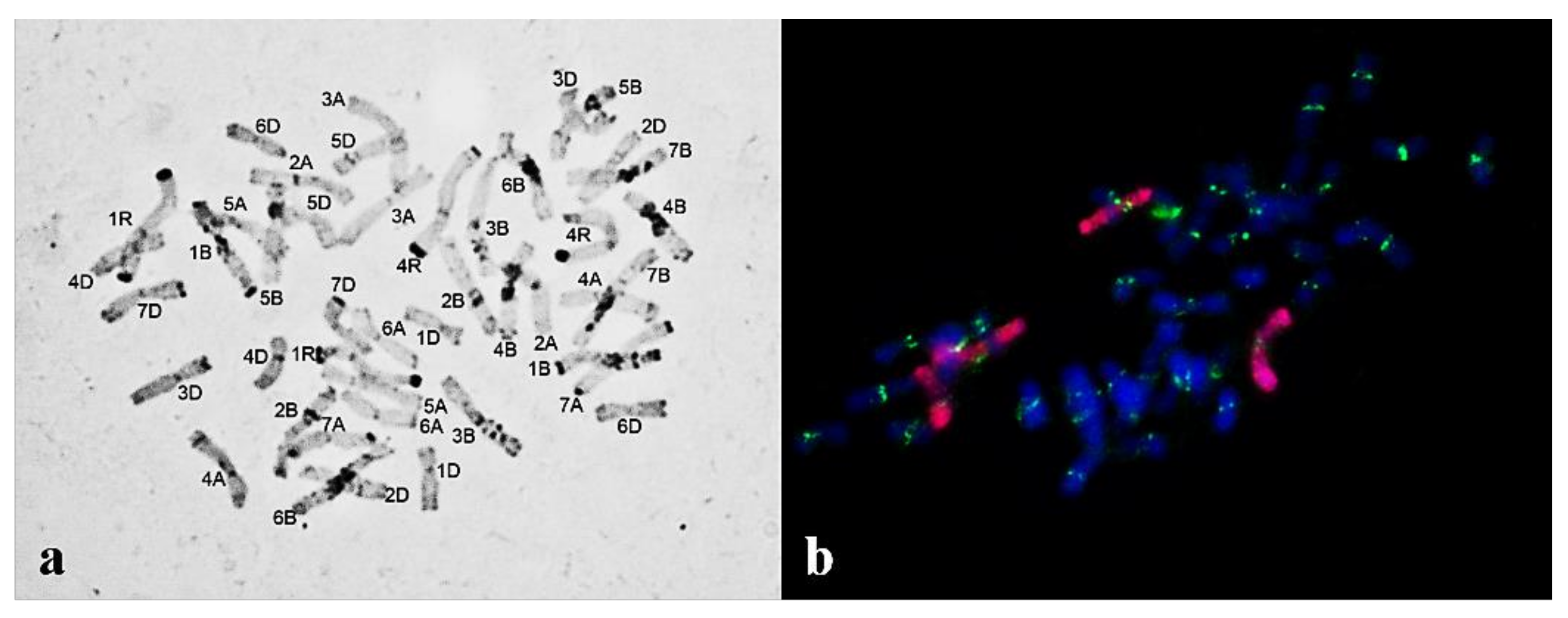
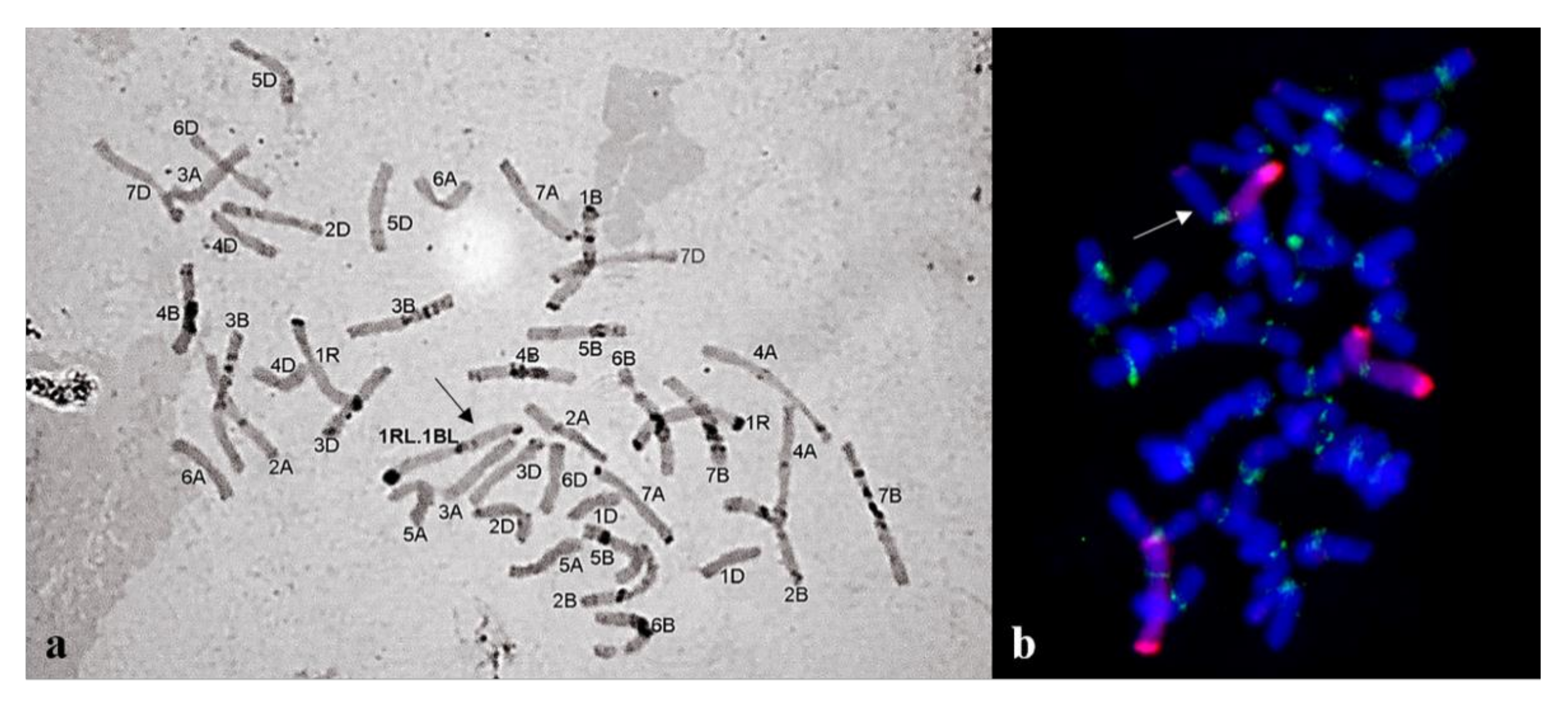
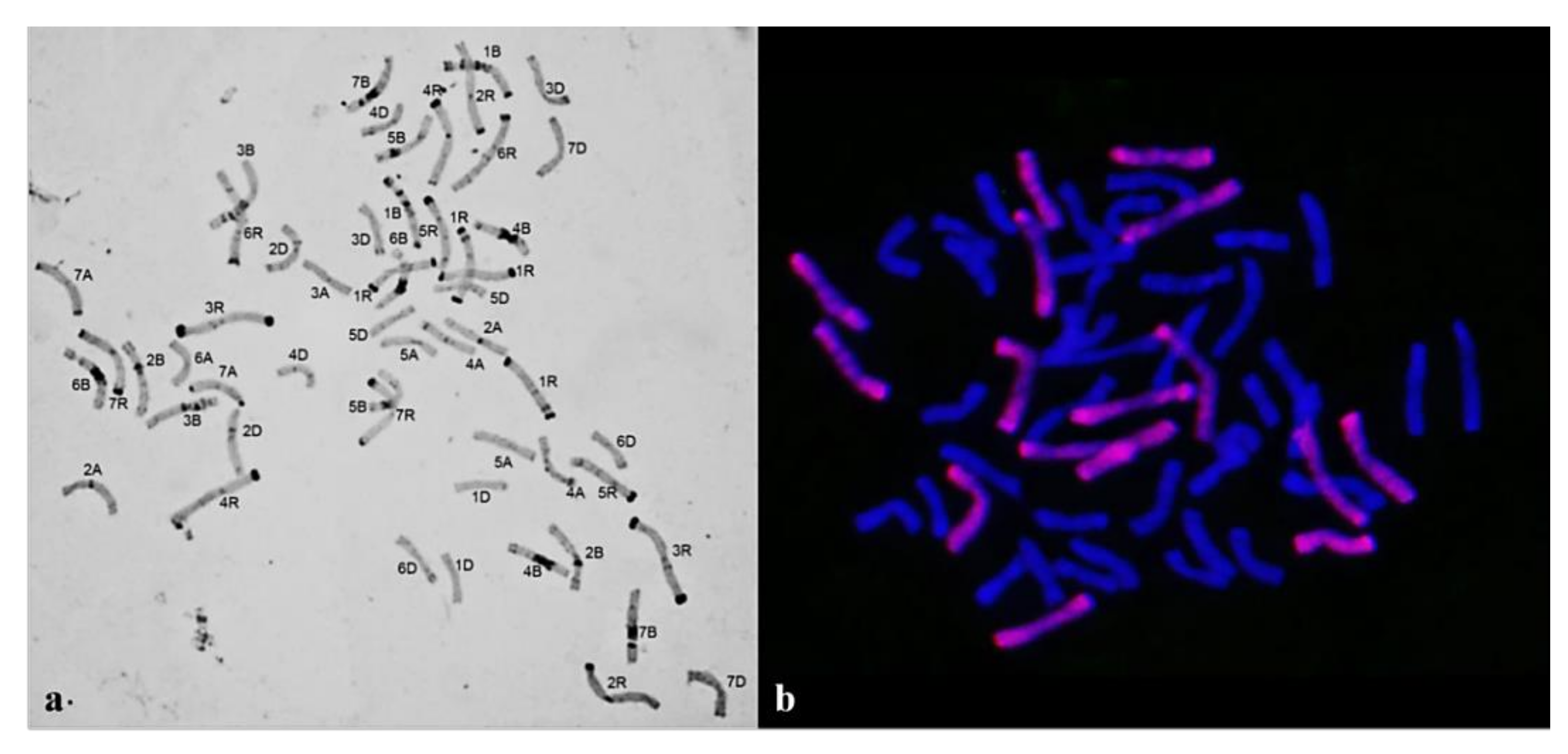
3.1. Karyotyping
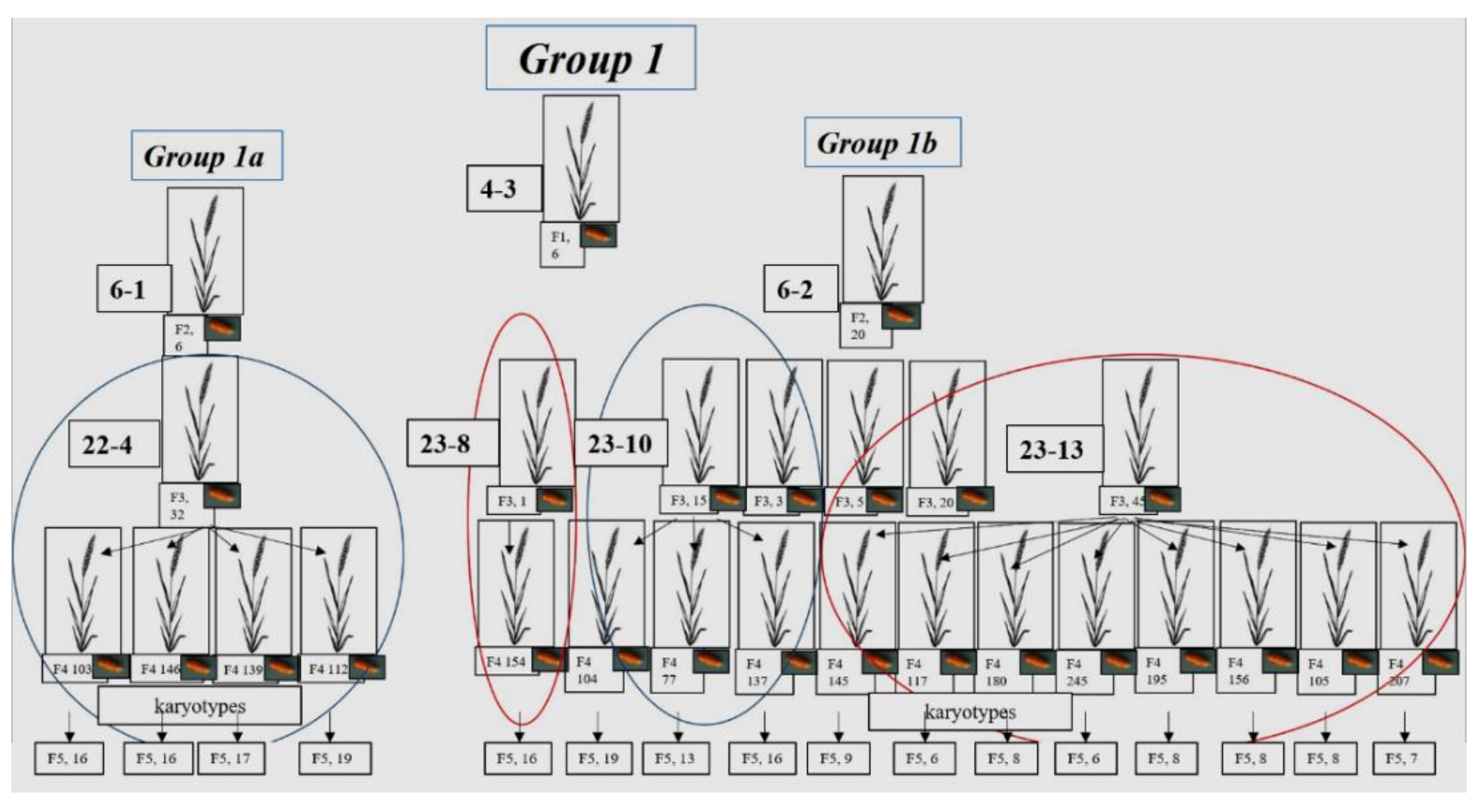
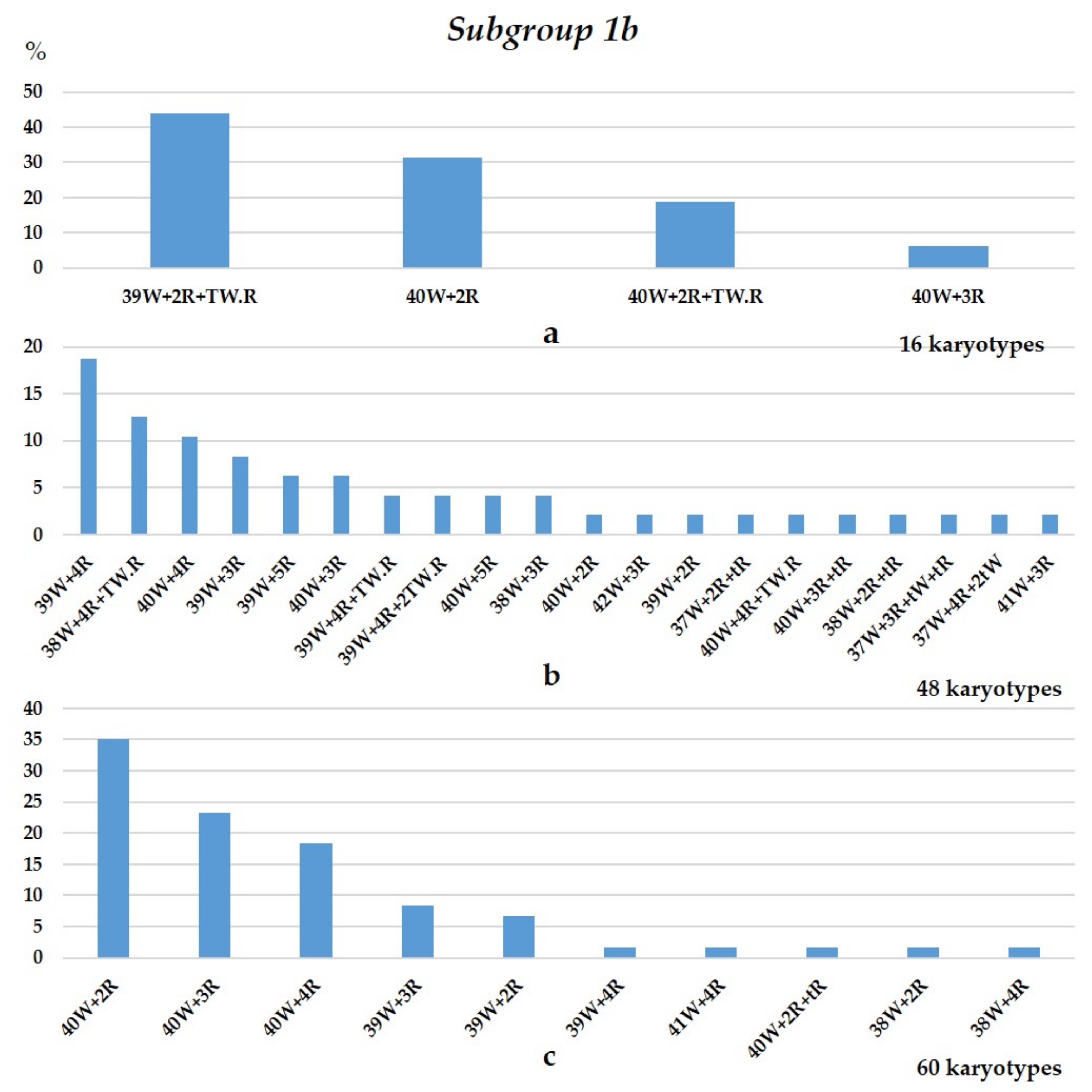
3.1.1. Group 1
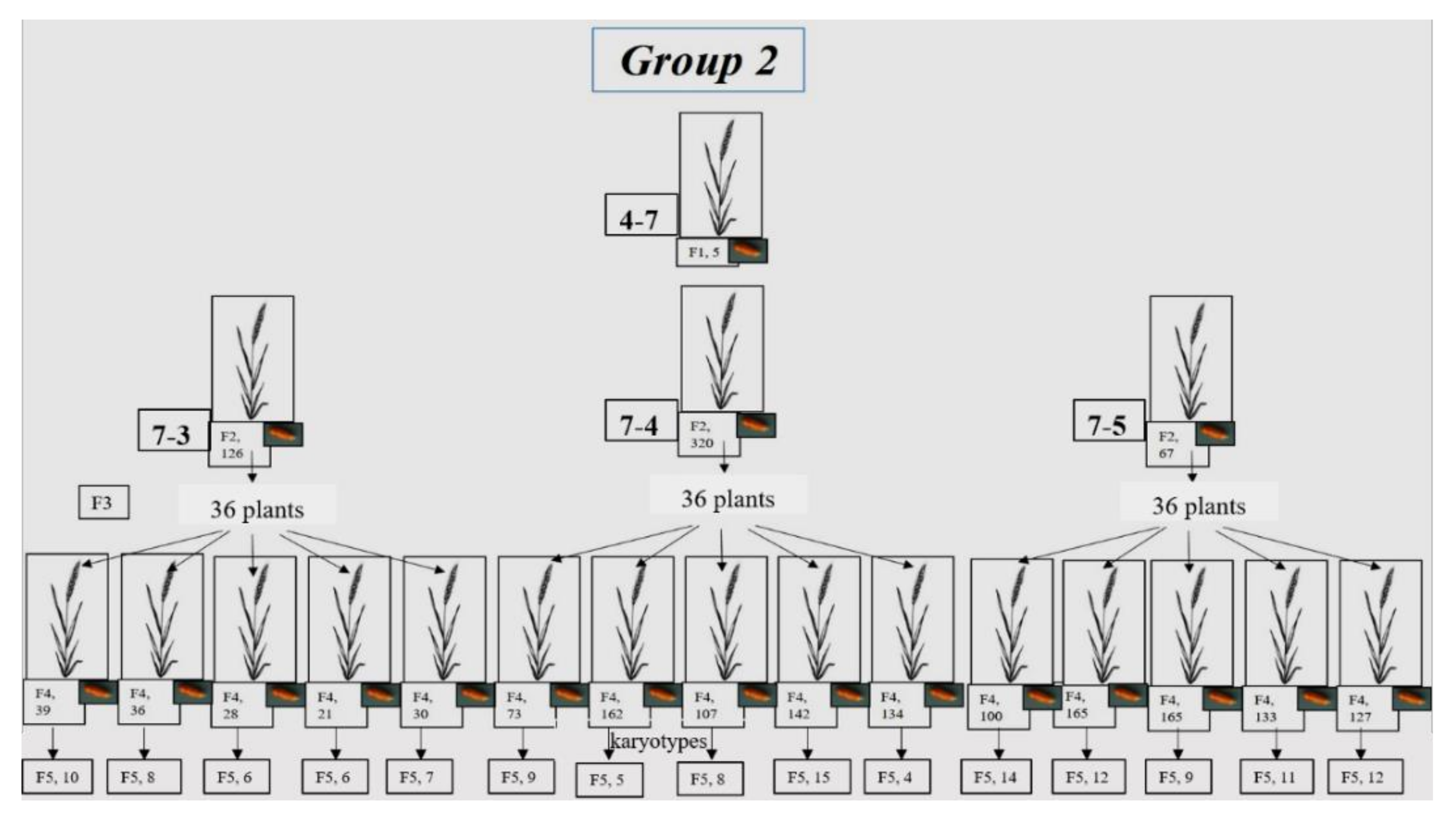
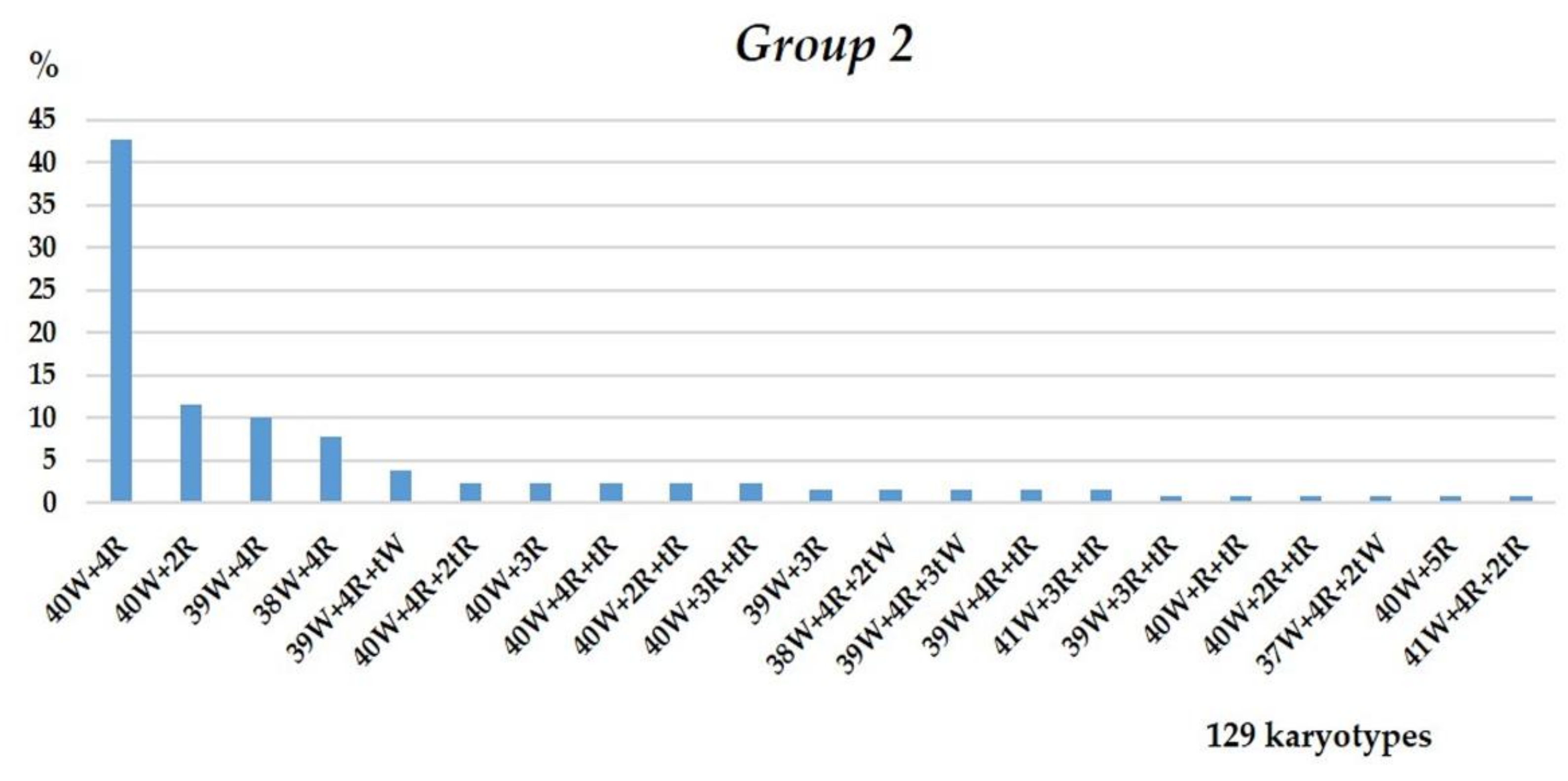
3.1.2. Group 2
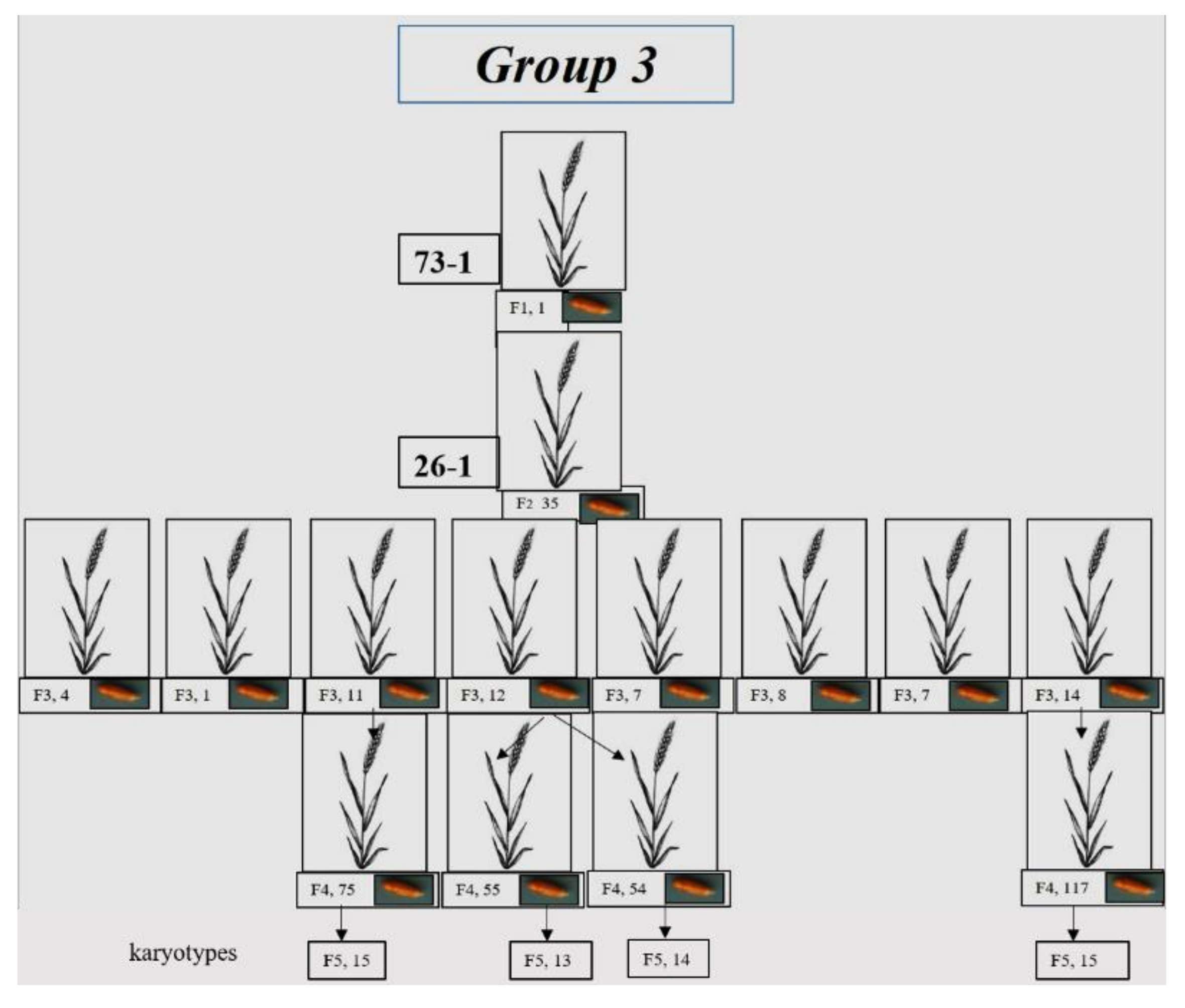
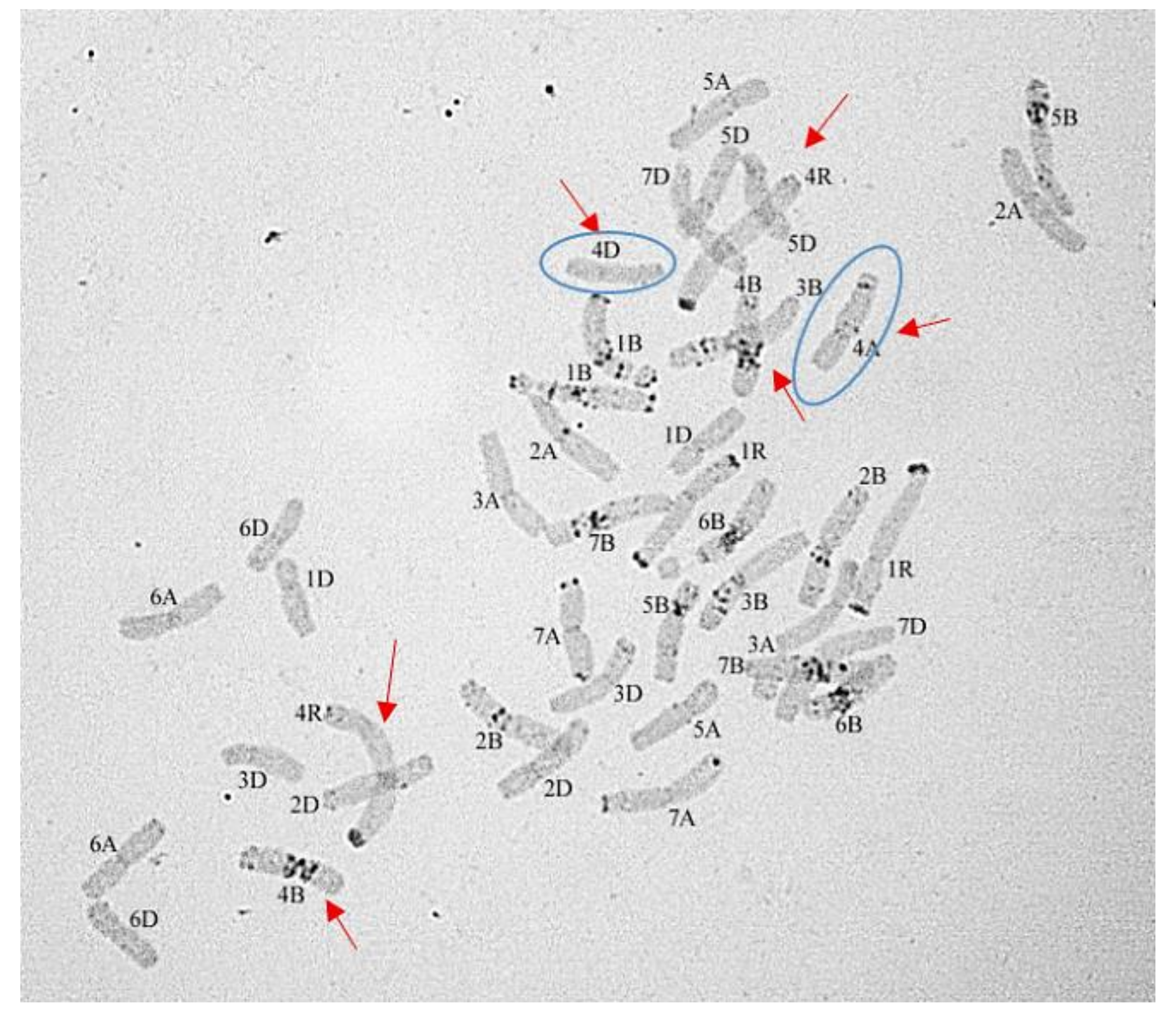
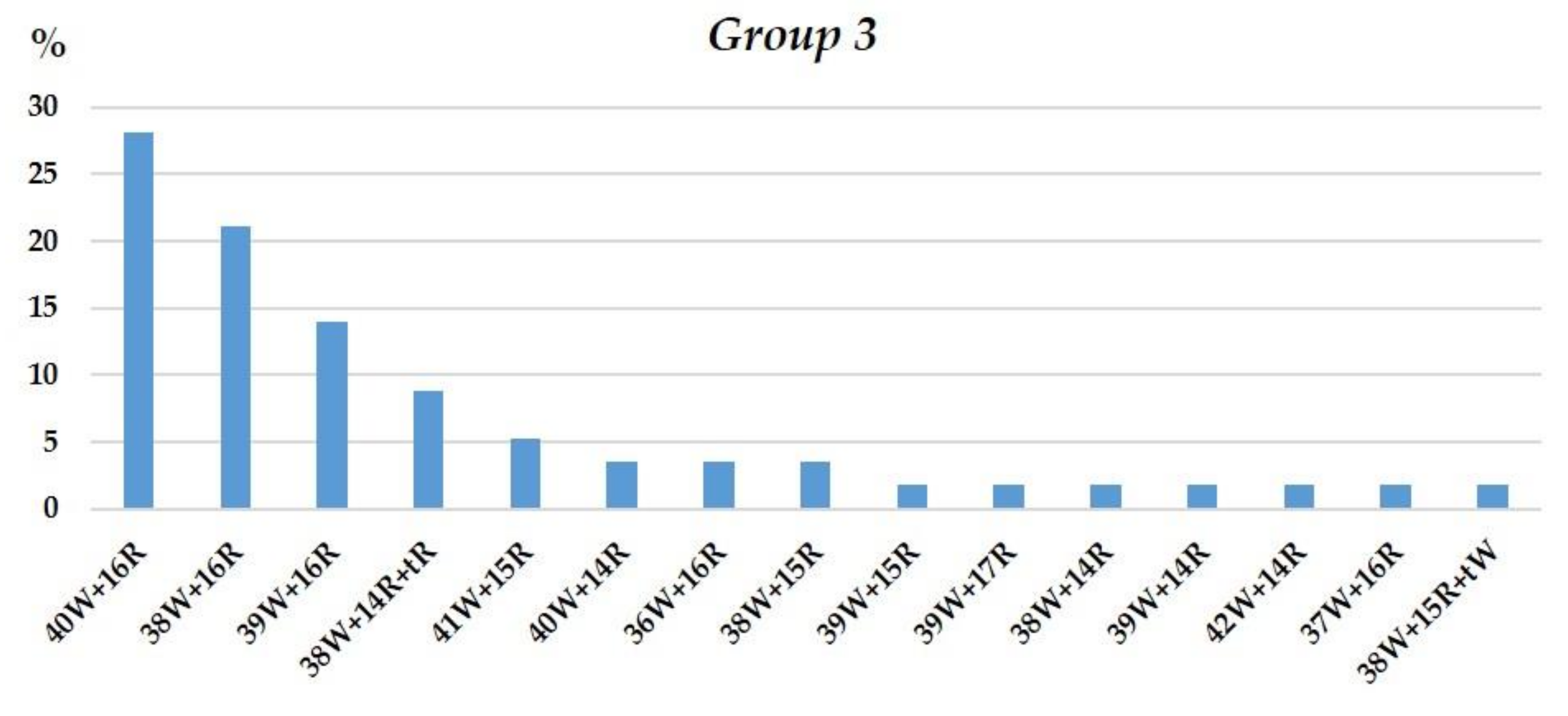
3.1.3. Group 3
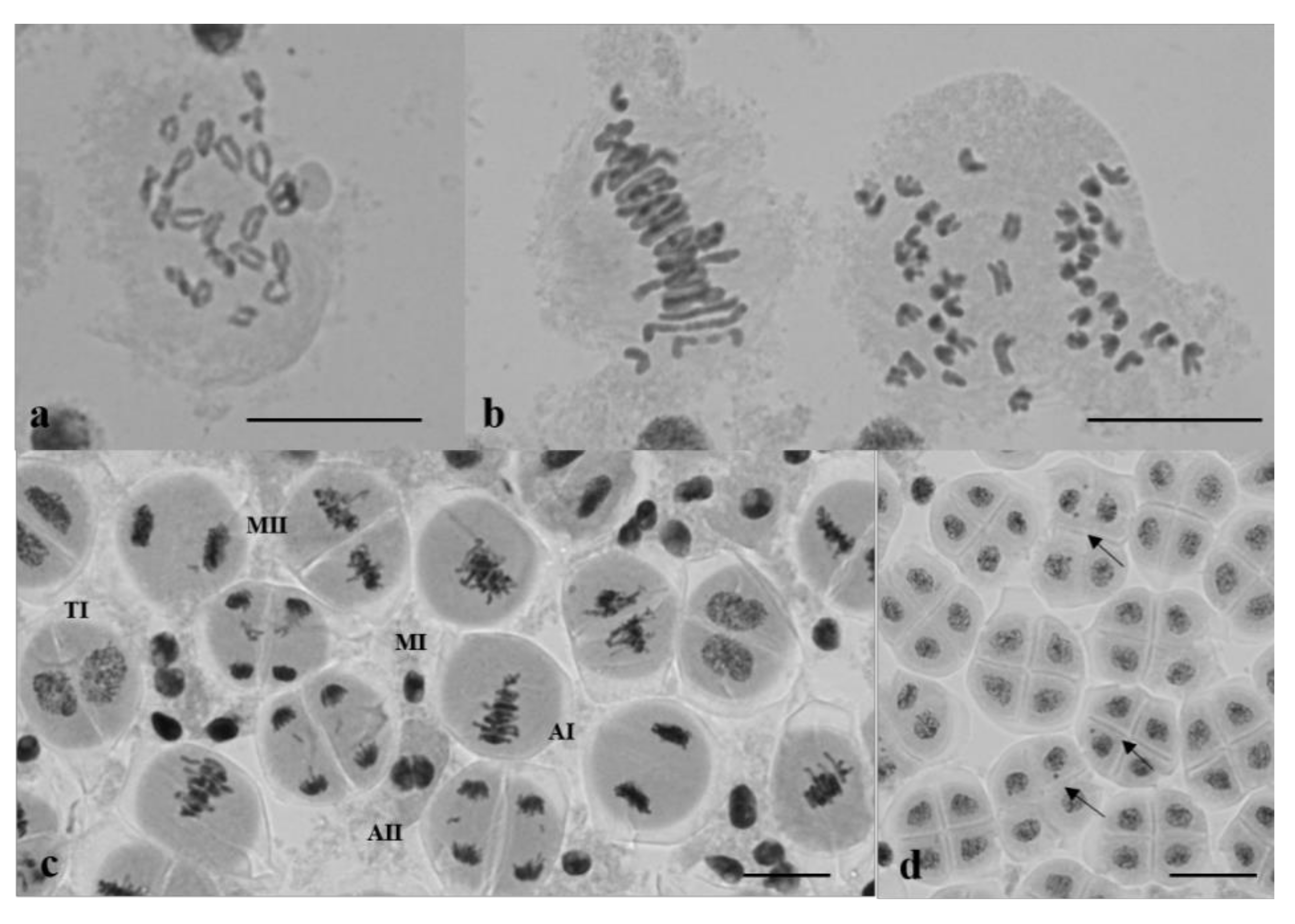
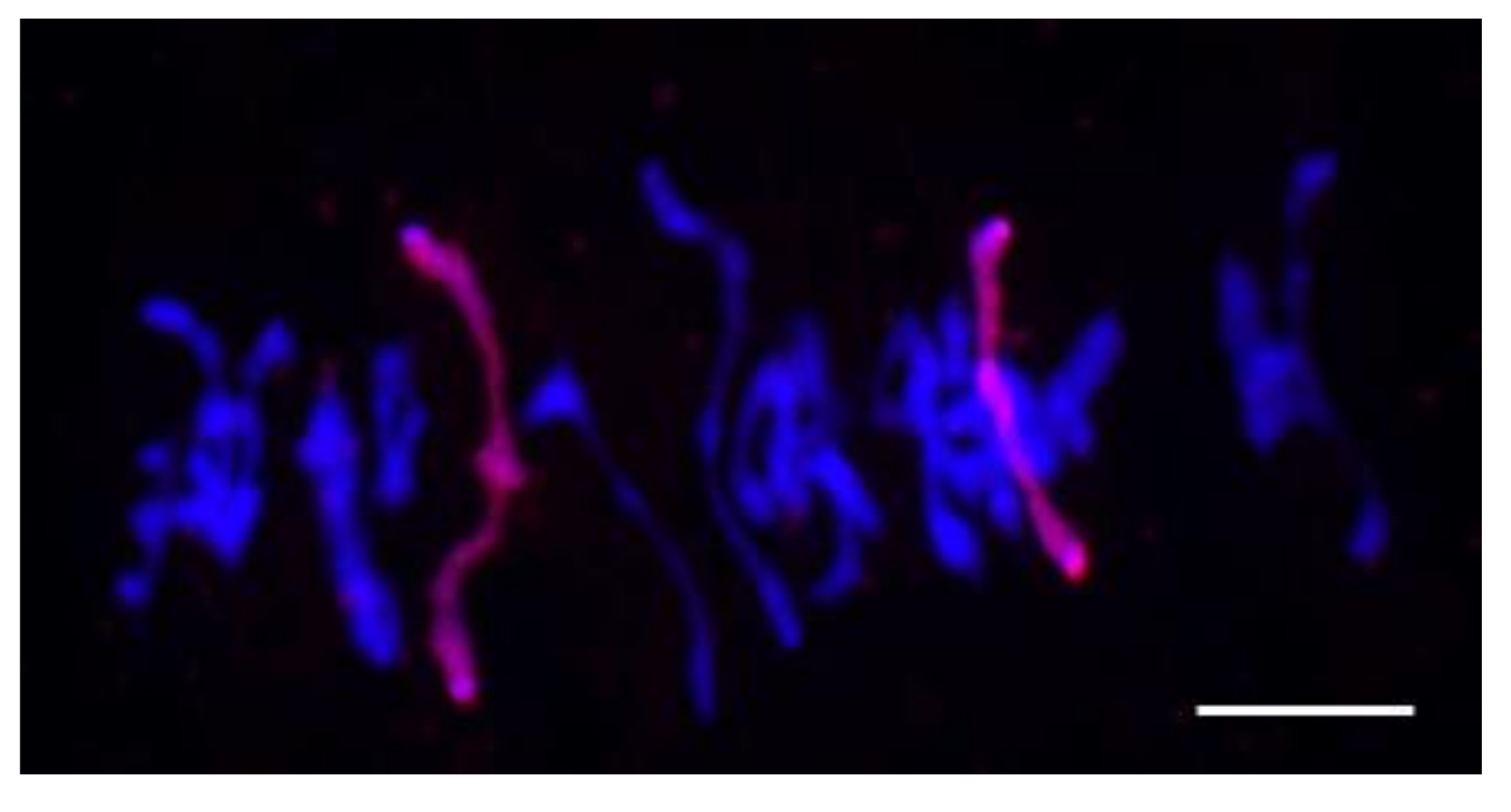
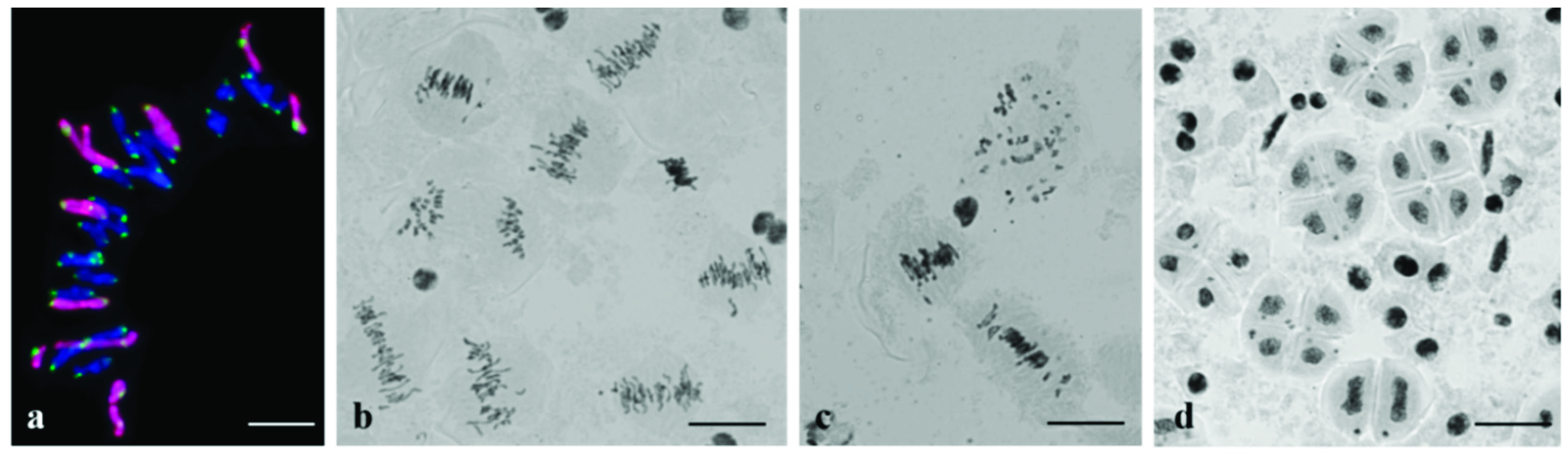
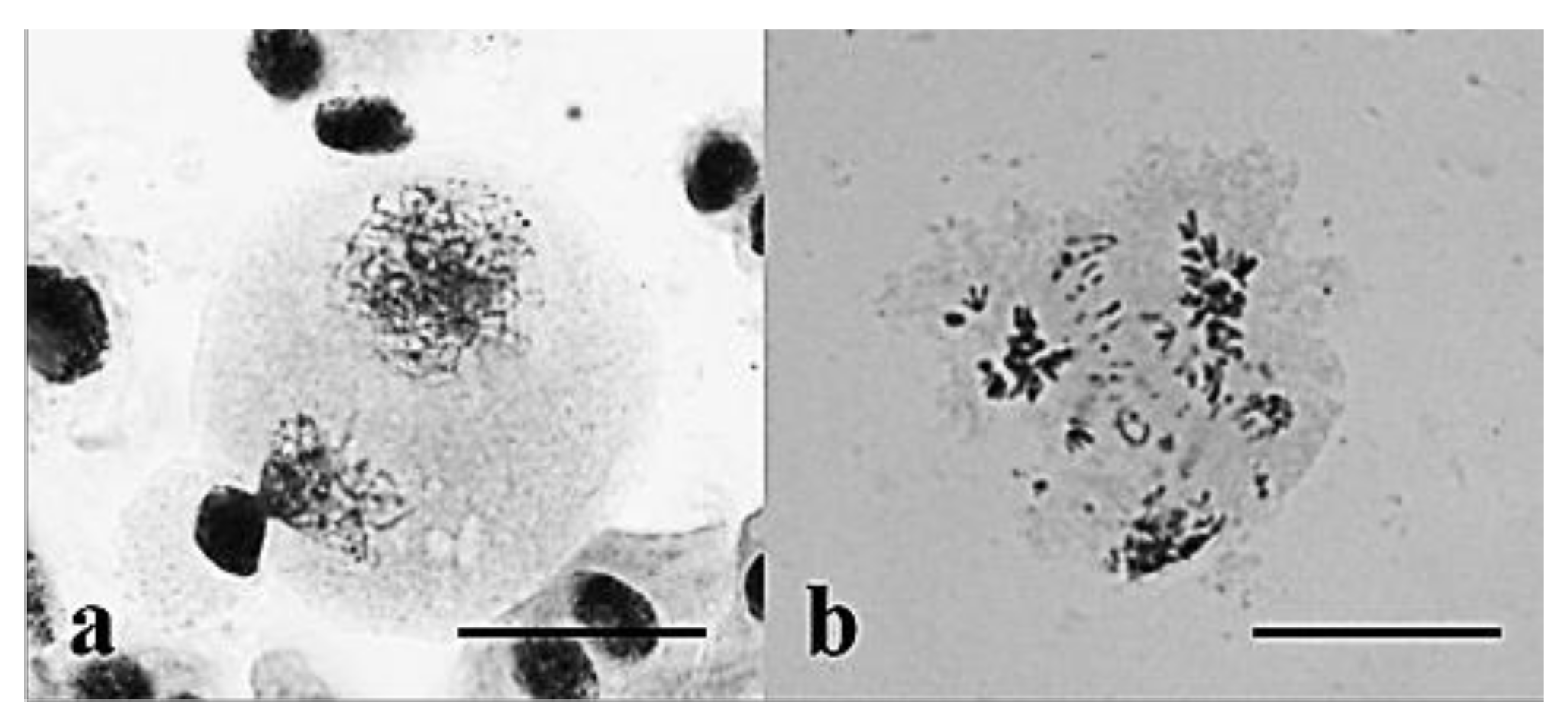
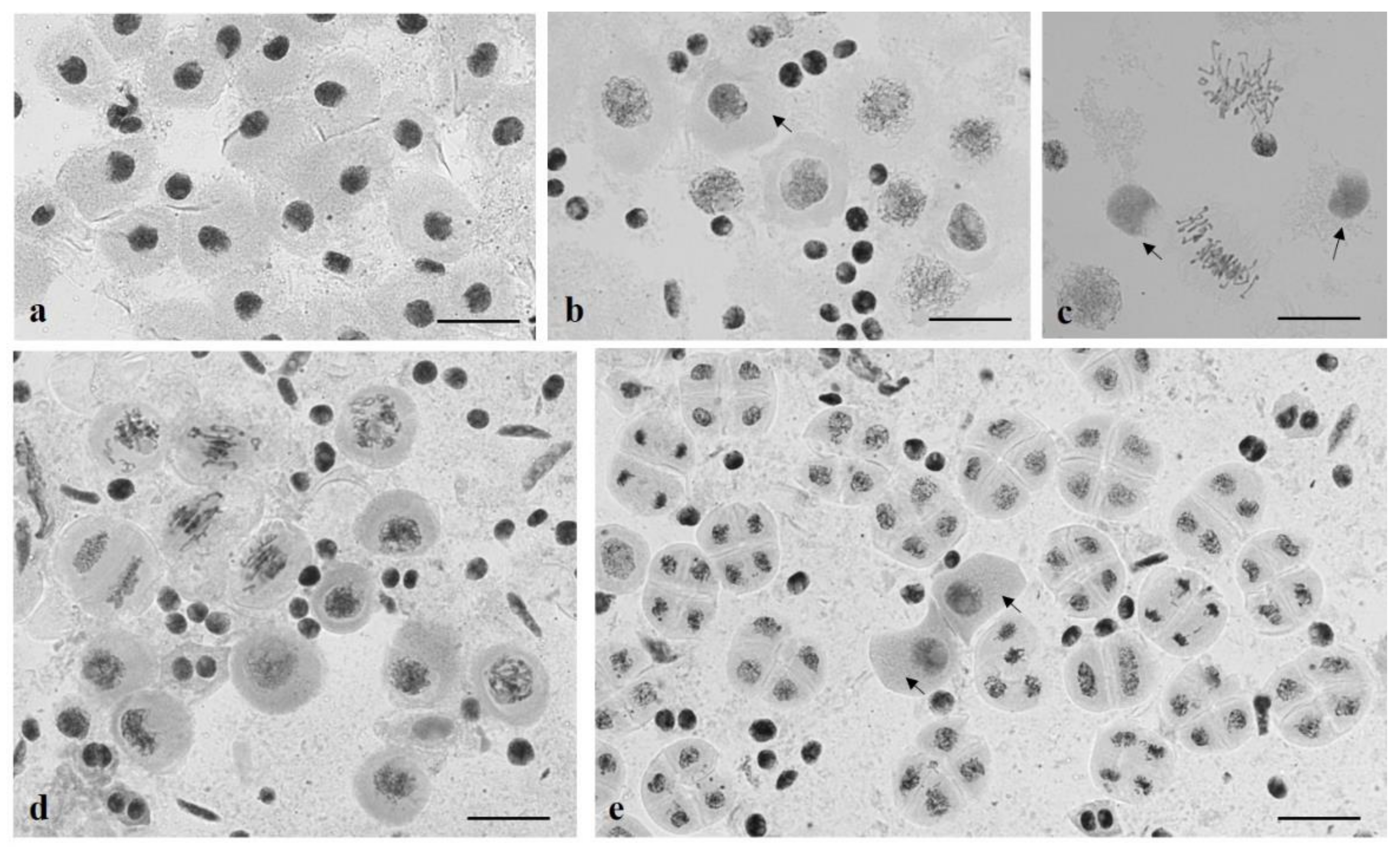
3.2. Chromosome Behavior in Meiosis
4. Discussion
4.1. Chromosome Instability in F5 1Rv(1A) × R Hybrids
4.2. Rye Chromosome 4R Is Preserved until F5 in 1Rv(1A) × R Hybrids
4.3. Alterations of Centromeric Regions
4.4. Meiotic Restitution Does Not Increase Ploidy in Progenitors of Octoploid Triticale
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Soltis, P.S.; Marchant, D.B.; Van de Peer, Y.; Soltis, D.E. Polyploidy and genome evolution in plants. Curr. Opin. Genet. Dev. 2015, 35, 119–125. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wendel, J.F.; Jackson, S.A.; Meyers, B.C.; Wing, R.A. Evolution of plant genome architecture. Genome Biol. 2016, 17, 1–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soltis, P.S.; Soltis, D.E. The role of hybridization in plant speciation. Annu. Rev. Plant Biol. 2009, 60, 561–588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramsey, J.; Schemske, D.W. Pathways, mechanisms, and rates of polyploid formation in flowering plants. Annu. Rev. Ecol. Syst. 1998, 29, 467–501. [Google Scholar] [CrossRef] [Green Version]
- Alix, K.; Gérard, P.R.; Schwarzacher, T.; Heslop-Harrison, J.S. Polyploidy and interspecific hybridisation: Partners for adaptation, speciation and evolution in plants. Ann. Bot. 2017, 120, 183–194. [Google Scholar] [CrossRef]
- Jones, R.N.; Hegarty, M. Order out of chaos in the hybrid plant nucleus. Cytogenet. Genome Res. 2009, 126, 376–389. [Google Scholar] [CrossRef]
- McClintock, B. The significance of responses of the genome to challenge. Science 1984, 226, 792–801. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.J. Genetic and epigenetic mechanisms for gene expression and phenotypic variation in plant polyploids. Annu. Rev. Plant Biol. 2007, 58, 377–406. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Doyle, J.J.; Flagel, L.E.; Paterson, A.H.; Rapp, R.A.; Soltis, D.E.; Soltis, P.S.; Wendel, J.F. Evolutionary genetics of genome merger and doubling in plants. Annu. Rev. Genet. 2008, 42, 443–461. [Google Scholar] [CrossRef] [Green Version]
- Jackson, S.; Chen, Z.J. Genomic and expression plasticity of polyploidy. Curr. Opin. Plant Biol. 2010, 13, 153–159. [Google Scholar] [CrossRef] [Green Version]
- Grover, C.; Gallagher, J.; Szadkowski, E.; Yoo, M.; Flagel, L.; Wendel, J. Homoeolog expression bias and expression level dominance in allopolyploids. New Phytol. 2012, 196, 966–971. [Google Scholar] [CrossRef]
- Hughes, T.E.; Langdale, J.A.; Kelly, S. The impact of widespread regulatory neofunctionalization on homeolog gene evolution following whole-genome duplication in maize. Genome Res. 2014, 24, 1348–1355. [Google Scholar] [CrossRef] [Green Version]
- Renny-Byfield, S.; Wendel, J.F. Doubling down on genomes: Polyplidy and crop plants. Am. J. Bot. 2014, 101, 1711–1725. [Google Scholar] [CrossRef] [Green Version]
- Panchy, N.; Lehti-Shiu, M.; Shiu, S.-H. Evolution of gene duplication in plants. Plant Physiol. 2016, 171, 2294–2316. [Google Scholar] [CrossRef] [Green Version]
- Miryeganeh, M.; Saze, H. Epigenetic inheritance and plant evolution. Popul. Ecol. 2020, 62, 17–27. [Google Scholar] [CrossRef]
- Wendel, J.F. The wondrous cycles of polyploidy in plants. Am. J. Bot. 2015, 102, 1753–1756. [Google Scholar] [CrossRef] [Green Version]
- Leitch, A.; Leitch, I. Genomic plasticity and the diversity of polyploid plants. Science 2008, 320, 481–483. [Google Scholar] [CrossRef] [PubMed]
- Heslop-Harrison, J.S.; Schwarzacher, T. Organization of the plant genome in chromosomes. Plant J. 2011, 66, 18–33. [Google Scholar] [CrossRef] [PubMed]
- International Wheat Genome Sequencing Consortium. A chromosome-based draft sequence of the hexaploid bread wheat genome. Science 2014, 345, 1251788. [Google Scholar] [CrossRef] [PubMed]
- Feldman, M.; Levy, A.A. Origin and evolution of wheat and related Triticeae species. In Alien Introgression in Wheat; Molnar-Lang, M., Ceoloni, C., Dolezel, J., Eds.; Springer International: Cham, Switzerland, 2015; pp. 21–76. [Google Scholar]
- Pont, C.; Salse, J. Wheat paleohistory created asymmetrical genomic evolution. Curr. Opin. Plant Biol. 2017, 36, 29–37. [Google Scholar] [CrossRef]
- Akhunov, E.D.; Akhunova, A.R.; Anderson, O.D.; Anderson, J.A.; Blake, N.; Clegg, M.T.; Coleman-Derr, D.; Conley, E.J.; Crossman, C.C.; Deal, K.R.; et al. Nucleotide diversity maps reveal variation in diversity among wheat genomes and chromosomes. BMC Genom. 2010, 11, 702. [Google Scholar] [CrossRef] [Green Version]
- Pont, C.; Murat, F.; Confolent, C.; Balzergue, S.; Sals, J. RNA-seq in grain unveils fate of neo- and paleopolyploidization events in bread wheat (Triticum aestivum L.). Genome Biol. 2011, 12, R119. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feldman, M.; Levy, A.A. Genome evolution due to allopolyploidization in wheat. Genetics 2012, 192, 763–774. [Google Scholar] [CrossRef] [Green Version]
- Pont, C.; Murat, F.; Guizard, S.; Flores, R.; Foucrier, S.; Bidet, Y.; Quraishi, U.M.; Alaux, M.; Doleel, J.; Fahima, T.; et al. Wheat syntenome unveils new evidences of contrasted evolutionary plasticity between paleo- and neoduplicated subgenomes. Plant. J. 2013, 76, 1030–1044. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pfeifer, M.; Kugler, K.G.; Sandve, S.R.; Zhan, B.; Rudi, H.; Hvidsten, T.R.; International Wheat Genome Sequencing Consortium; Mayer, K.F.; Olsen, O.A. Genome interplay in the grain transcriptome of hexaploid bread wheat. Science 2014, 345, 1250091. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Z.; Xin, M.; Qin, J.; Peng, H.; Ni, Z.; Yao, Y.; Sun, Q. Temporal transcriptome profiling reveals expression partitioning of homeologous genes contributing to heat and drought acclimation in wheat (Triticum aestivum L.). BMC Plant Biol. 2015, 15, 152. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Z.; Gou, X.; Xun, H.; Bian, Y.; Ma, X.; Li, J.; Lia, N.; Gong, L.; Feldman, M.; Liu, B.; et al. Homoeologous exchanges occur through intragenic recombination generating novel transcripts and proteins in wheat and other polyploids. Proc. Natl. Acad. Sci. USA 2020, 117, 14561–14571. [Google Scholar] [CrossRef]
- Murat, F.; Zhang, R.; Guizard, S.; Flores, R.; Armero, A.; Pont, C.; Steinbach, D.; Quesneville, H.; Cooke, R.; Salse, J. Shared subgenome dominance following polyploidization explains grass genome evolutionary plasticity from a seven protochromosome ancestor with 16K protogenes. Genome Biol. Evol. 2014, 6, 12–33. [Google Scholar] [CrossRef]
- Naranjo, T.; Roca, A.; Goicoechea, P.G.; Giraldez, R. Arm homoeology of wheat and rye chromosomes. Genome 1987, 29, 873–882. [Google Scholar] [CrossRef]
- Naranjo, T. Chromosome structure of durum wheat. Theor. Appl. Genet. 1990, 79, 397–400. [Google Scholar] [CrossRef]
- Liu, C.J.; Atkinson, M.D.; Chinoy, C.N.; Devos, K.M.; Gale, M.D. Nonhomoeologous translocations between group 4, 5 and 7 chromosomes within wheat and rye. Theor. Appl. Genet. 1992, 83, 305–312. [Google Scholar] [CrossRef]
- Jiang, J.; Gill, B.S. Different species-specific chromosome translocations in Triticum timopheevii and T. turgidum support the diphyletic origin of polyploid wheats. Chromosome Res. 1994, 2, 59–64. [Google Scholar] [CrossRef]
- Rodríguez, S.; Perera, E.; Maestra, B.; Díez, M.; Naranjo, T. Chromosome structure of Triticum timopheevii relative to T. turgidum. Genome 2000, 43, 923–930. [Google Scholar] [CrossRef]
- Salina, E.A.; Leonova, I.N.; Efremova, T.T.; Roder, M.S. Wheat genome structure: Translocations during the course of polyploidization. Funct. Integr. Genom. 2006, 6, 71–80. [Google Scholar] [CrossRef]
- Badaeva, E.D.; Dedkova, O.S.; Pukhalskyi, V.A.; Zelenin, A.V. Chromosomal changes over the course of polyploid wheat evolution and domestication. In Advances in Wheat Genetics: From Genome to Field; Ogihara, Y., Takumi, S., Handa, H., Eds.; Springer: Tokyo, Japan, 2015; pp. 83–89. [Google Scholar]
- Badaeva, E.D.; Chikida, N.N.; Fisenko, A.N.; Surzhikov, S.A.; Belousova, M.K.; Özkan, H.; Dragovich, A.Y.; Kochieva, E.Z. Chromosome and molecular analyses reveal significant karyotype diversity and provide new evidence on the origin of Aegilops columnaris. Plants 2021, 10, 956. [Google Scholar] [CrossRef]
- Badaeva, E.D.; Chikida, N.N.; Belousova, M.K.; Ruban, A.S.; Surzhikov, S.A.; Zoshchuk, S.A. A new insight on the evolution of polyploid Aegilops species from the complex Crassa: Molecular-cytogenetic analysis. Plant Syst. Evol. 2021, 307, 3. [Google Scholar] [CrossRef]
- Parisod, C.; Badaeva, E.D. Chromosome restructuring among hybridizing wild wheats. New Phytol. 2020, 226, 1263–1273. [Google Scholar] [CrossRef] [PubMed]
- Sattler, M.C.; Carvalho, C.R.; Clarindo, W.R. The polyploidy and its key role in plant breeding. Planta 2016, 243, 281–296. [Google Scholar] [CrossRef] [PubMed]
- Kocheshkova, A.A.; Kroupin, P.Y.; Bazhenov, M.S.; Karlov, G.I.; Pochtovyy, A.A.; Upelniek, V.P.; Belov, V.I.; Divashuk, M. GPre-harvest sprouting resistance and haplotype variation of ThVp-1 gene in the collection of wheat-wheatgrass hybrids. PLoS ONE 2017, 12, e0188049. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muntzing, A. Historical review of the development of triticale. In Triticale, Proceedings of an International Symposium, El Batan, Mexico; Development Research Centre: Ottawa, ON, Canada, 1974; pp. 13–30. [Google Scholar]
- Meister, G.K. Natural hybridization of wheat and rye in Russia. J. Hered. 1921, 12, 467–470. [Google Scholar] [CrossRef]
- Levitsky, G.A.; Benetzkaja, G.K. Cytological investigations of constant intermediate rye-wheat hybrids. In Proceedings of the Allunion Congress of Genetics Selek, Leningrad, Russia, 10–16 January 1929; pp. 345–352, (In Russian with English summary). [Google Scholar]
- Ma, X.F.; Fang, P.; Gustafson, J.P. Polyploidization-induced genome variation in triticale. Genome 2004, 47, 839–848. [Google Scholar] [CrossRef] [Green Version]
- Ma, X.F.; Gustafson, J.P. Timing and rate of genome variation in triticale following allopolyploidization. Genome 2006, 49, 950–958. [Google Scholar] [CrossRef] [Green Version]
- Ma, X.F.; Gustafson, J.P. Allopolyploidization-accommodated genomic sequence changes in Triticale. Ann. Bot. 2008, 101, 825–832. [Google Scholar] [CrossRef] [Green Version]
- Bento, M.; Pereira, H.S.; Rocheta, M.; Gustafson, P.; Viegas, W.; Silva, M. Polyploidization as a Retraction Force in Plant Genome Evolution: Sequence Rearrangements in Triticale. PLoS ONE 2008, 3, e1402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, Z.X.; Fu, S.L.; Ren, Z.L.; Zhou, J.P.; Yan, B.J.; Zhang, H.Q. Variation of tandem repeat, regulatory element, and promoter regions revealed by wheat–rye amphiploids. Genome 2008, 51, 399–408. [Google Scholar] [CrossRef]
- Martín, A.C.; Borrill, P.; Higgins, J.; Alabdullah, A.; Ramírez-González, R.H.; Swarbreck, D.; Uauy, C.; Shaw, P.; Moore, G. Genome-Wide Transcription During Early Wheat Meios Is Independent of Synapsis, Ploidy Level, and the Ph1 Locus. Front. Plant Sci. 2018, 9, 1791. [Google Scholar] [CrossRef] [PubMed]
- Badaeva, E.D.; Badaev, N.S.; Bolsheva, N.L.; Zelenin, A.V. Chromosome alterations in the karyotype of triticale in comparison with the parental forms I. Heterochromatic regions of R genome chromosomes. Theor. Appl. Genet. 1986, 72, 518–523. [Google Scholar] [CrossRef] [PubMed]
- Bolsheva, N.L.; Badaeva, E.D.; Badaev, N.S.; Zelenin, A.V. Chromosome alterations in the karyotype of triticale in comparison with the parental forms 2. Heterochromatin of the wheat chromosomes. Theor. Appl. Genet. 1986, 73, 66–71. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.; Lv, Z.; Guo, X.; Zhang, X.; Han, F. Alteration of Terminal Heterochromatin and Chromosome Rearrangements in Derivatives of Wheat–rye Hybrids. JGG 2013, 40, 413–420. [Google Scholar] [CrossRef]
- Hao, M.; Luo, J.; Zhang, L.; Yuan, Z.; Yang, Y.; Wu, M.; Chen, W.; Zheng, Y.; Zhang, H.; Liu, D. Production of hexaploid triticale by a synthetic hexaploid wheat–rye hybrid method. Euphytica 2013, 193, 347–357. [Google Scholar] [CrossRef]
- Tang, Z.; Li, M.; Chen, L.; Wang, Y.; Ren, Z.; Fu, S. New types of wheat chromosomal structural variations in derivatives of wheat–rye hybrids. PLoS ONE 2014, 9, e110282. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Guo, X.; Wang, C.; Ji, W. Spontaneous and divergent hexaploid Triticales derived from common wheat × rye by complete elimination of D-genome chromosomes. PLoS ONE 2015, 10, e0120421. [Google Scholar] [CrossRef] [Green Version]
- Lukaszewski, A.J. Introgressions between Wheat and Rye. In Alien Introgression in Wheat; Molnár-Láng, M., Ceoloni, C., Doležel, J., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 163–189. [Google Scholar]
- Dubovets, N.I.; Sycheva, Y.A. Microevolutionary differentiation of cereal tetraploid species by formation of recombinant genomes. Vavilovskii Zhurnal Genetiki i Selektsii Vavilov J. Genet. Breed. (In Russian with English summary). 2016, 20, 378–385. [Google Scholar] [CrossRef] [Green Version]
- Guo, X.; Su, H.; Shi, Q.; Fu, S.; Wang, J.; Zhang, X.; Hu, Z.; Han, F. De Novo Centromere Formation and Centromeric Sequence Expansion in Wheat and its Wide Hybrids. PLoS Genet. 2016, 12, e1005997. [Google Scholar] [CrossRef] [Green Version]
- Ramsey, J.; Schemske, D.W. Neopolyploidy in flowering plants. Annu. Rev. Ecol. Syst. 2002, 33, 589–639. [Google Scholar] [CrossRef] [Green Version]
- Otto, S.P. The evolutionary consequences of polyploidy. Cell 2007, 131, 452–462. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kreiner, J.M.; Kron, P.; Husband, B.C. Frequency and maintenance of unreduced gametes in natural plant populations: Associations with reproductive mode, life history and genome size. New Phytol. 2017, 214, 879–889. [Google Scholar] [CrossRef] [Green Version]
- Husband, B.C. The role of triploid hybrids in the evolutionary dynamics of mixed-ploidy populations. Biol. J. Linn. Soc. 2004, 82, 537–546. [Google Scholar] [CrossRef]
- Silkova, O.G.; Loginova, D.B. Sister chromatid separation and monopolar spindle organization in the first meiosis as two mechanisms of unreduced gametes formation in wheat–rye hybrids. Plant Reprod. 2016, 29, 199–213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olesczuk, S.; Lukaszewski, A.J. The origin of unusual chromosome constitutions among newly formed allopolyploids. Am. J. Bot. 2014, 101, 318–326. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, L.Q.; Yen, Y.; Zheng, Y.L.; Liu, D.C. Meiotic restriction in emmer wheat is controlled by one or more nuclear genes that continue to function in derived lines. Sex. Plant Reprod. 2007, 20, 159–166. [Google Scholar] [CrossRef]
- Oleszczuk, S.; Grzechnik, N.; Mason, A.S.; Zimny, J. Heritability of meiotic restitution and fertility restoration in haploid triticale. Plant Cell Rep. 2019, 38, 1515–1525. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.Q.; Liu, D.C.; Zheng, Y.L.; Yan, Z.H.; Dai, S.F.; Li, Y.F.; Jiang, Q.; Ye, Y.Q.; Yen, Y. Frequent occurrence of unreduced gametes in Triticum turgidum-Aegilops tauschii hybrids. Euphytica 2010, 172, 285–294. [Google Scholar] [CrossRef]
- Silkova, O.G.; Loginova, D.B.; Volodina, E.A.; Ivanova, Y.N.; Bondarevich, E.B.; Solovey, L.A.; Sycheva, E.; Dubovets, N.I. Development and Characterization of Wheat–rye Hybrids Produced by Meiotic Restitution. Russ. J. Genet. 2018, 54, 1266–1276. [Google Scholar]
- Feldman, M.; Levy, A.A. Allopolyploidy--a shaping force in the evolution of wheat genomes. Cytogenet Genome Res. 2005, 109, 250–258. [Google Scholar] [CrossRef]
- Silkova, O.G.; Dobrovolskaya, O.B.; Dubovets, N.I.; Adonina, I.G.; Kravtsova, L.A.; Shchapova, A.I.; Shumny, V.K. Production of wheat–rye substitution lines based on winter rye cultivars with karyotype identification by means of C-banding, GISH, and SSR markers. Rus. J. Genet. 2007, 43, 957–960. [Google Scholar] [CrossRef]
- Badaeva, E.D.; Badaev, N.S.; Gill, D.S.; Filatenko, A.A. Intraspecific karyotype divergence in Triticum araraticum (Poaceae). Plant Syst. Evol. 1994, 192, 117–145. [Google Scholar] [CrossRef]
- Badaeva, E.D. “Chromosomal passport” of Triticum aestivum L. em Thell. cv. Chinese Spring and standartization of chromosomal analysis of cereals. Cereal Res. Commun. 1990, 18, 273–281. [Google Scholar]
- Gill, B.S.; Kimber, G. The Giemsa C-Banded Karyotype of Rye homoeologous/constitutive heterochromatin/chromosomes. Proc. Nat. Acad. Sci. USA 1974, 71, 1247–1249. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qi, L.L.; Wu, J.J.; Friebe, B.; Qian, C.; Gu, Y.Q.; Fu, D.L.; Gill, B.S. Sequence organization and evolutionary dynamics of Brachypodium specific centromere retrotransposons. Chromosome Res. 2013, 21, 507–521. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, P.; Wanlong, L.; Fellers, J.; Friebe, B.; Gill, B.S. BAC-FISH in wheat identifies chromosome landmarks consisting of different types of transposable elements. Chromosoma 2004, 112, 288–299. [Google Scholar] [CrossRef] [PubMed]
- Francki, M.G. Identification of Bilby, a diverged centromeric T1-copia retrotransposon family from cereal rye (Secale cereale L.). Genome 2001, 44, 266–274. [Google Scholar] [CrossRef] [PubMed]
- Brownfield, L.; Köhler, C. Unreduced gamete formation in plants: Mechanisms and prospects. J. Exp. Bot. 2011, 62, 1659–1668. [Google Scholar] [CrossRef] [Green Version]
- De Storme, N.; Geelen, D. Sexual polyploidization in plants–cytological mechanisms and molecular regulation. New Phytol. 2013, 198, 670–684. [Google Scholar] [CrossRef] [Green Version]
- Mason, A.S.; Pires, J.C. Unreduced gametes: Meiotic mishap or evolutionary mechanism? Trends Genet. 2015, 31, 5–10. [Google Scholar] [CrossRef] [PubMed]
- Shchapova, A.I.; Kravtsova, L.A.; Potapova, T.A. Patterns in the transformation of the genomic structure of wheat—rye polyhaploids ABDR. Genetica 1987, 23, 295–302. [Google Scholar]
- Oleszczuk, S.; Rabiza-Swider, J.; Zimny, J.; Lukaszewski, A.J. Aneuploidy among androgenic progeny of hexaploid triticale (XTriticosecale Wittmack). Plant Cell Rep. 2011, 30, 575–586. [Google Scholar] [CrossRef] [Green Version]
- Ishii, T.; Karimi-Ashtiyani, R.; Houben, A. Haploidization via Chromosome Elimination: Means and Mechanisms. Annu. Rev. Plant Biol. 2016, 67, 421–438. [Google Scholar] [CrossRef]
- Nakajima, G. Occurrence of a haploid in Triticum turgidum. Jpn. J. Genet. 1935, 11, 246–247. [Google Scholar] [CrossRef] [Green Version]
- Gaines, E.F.; Aase, H.C. A haploid wheat plant. AJB 1926, 13, 373–385. [Google Scholar] [CrossRef]
- Barclay, I.R. High frequencies of haploid production in wheat (Triticum aestivum) by chromosome elimination. Nature 1975, 256, 410–411. [Google Scholar] [CrossRef]
- Zenkteler, M.; Straub, J. Cytoembryogical studies on the process of fertilization and the development of haploid embryos of Triticum aestivum L. (2n=42) after crossing with Hordeum bulbosum (2n=14). Z Pflanzenzüchtg 1979, 82, 36–44. [Google Scholar]
- Kasha, K.J.; Kao, K.N. High frequency haploid production in barley (Hordeum vulgare L.). Nature 1970, 225, 874–876. [Google Scholar] [CrossRef]
- Ho, K.M.; Kasha, K.J. Genetic control of chromosome elimination during haploid formation in barley. Genetics 1975, 81, 263–275. [Google Scholar] [CrossRef]
- Subrahmanyam, N.C. Haploidy from Hordeum interspecific crosses: I. Polyhaploids of H. parodii and H. procerum. Theor. Appl. Genet. 1977, 49, 209–217. [Google Scholar] [CrossRef]
- Polgári, D.; Cseh, A.; Szakács, É.; Jäger, K.; Molnár-Láng, M.; Sági, L. High-frequency generation and characterization of intergeneric hybrids and haploids from new wheat-barley crosses. Plant Cell Rep. 2014, 33, 1323–1331. [Google Scholar] [CrossRef]
- Polgári, D.; Mihók, E.; Sági, L. Composition and random elimination of paternal chromosomes in a large population of wheat × barley (Triticum aestivum L. × Hordeum vulgare L.) hybrids. Plant Cell Rep. 2019, 38, 767–775. [Google Scholar] [CrossRef] [Green Version]
- Inagaki, M.N.; Mujeeb-Kazi, A. Comparison of polyhaploid production frequencies in crosses of hexaploid wheat with maize, pearl millet and sorghum. Breed. Sci. 1995, 45, 157–161. [Google Scholar]
- Gernand, D.; Rutten, T.; Varshney, A.; Rubtsova, M.; Prodanovic, S.; Brüss, C.; Kumlehn, J.; Matzk, F.; Houben, A. Uniparental chromosome elimination at mitosis and interphase in wheat and pearl millet crosses involves micronucleus formation, progressive heterochromatinization, and DNA fragmentation. Plant Cell 2005, 17, 2431–2438. [Google Scholar] [CrossRef] [Green Version]
- Komeda, N.; Chaudhary, H.K.; Suzuki, G.; Mukai, Y. Cytological evidence for chromosome elimination in wheat×Imperata cylindrica hybrids. Genes Genet. Syst. 2007, 82, 241–248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ishii, T.; Ueda, T.; Tanaka, H.; Tsujimoto, H. Chromosome elimination by wide hybridization between Triticeae or oat plant and pearl millet: Pearl millet chromosome dynamics in hybrid embryo cells. Chromosome Res. 2010, 18, 821–831. [Google Scholar] [CrossRef]
- Zhang, J.; Yang, F.; Jiang, Y.; Guo, Y.; Wang, Y.; Zhu, X.; Li, J.; Wan, H.; Wang, Q.; Deng, Z.; et al. Preferential Subgenome Elimination and Chromosomal Structural Changes Occurring in Newly Formed Tetraploid Wheat—Aegilops ventricosa Amphiploid (AABBDvDvNvNv). Front. Genet. 2020, 11, 330. [Google Scholar] [CrossRef] [PubMed]
- Comai, L. The advantages and disadvantages of being polyploid. Nat. Rev. Genet. 2005, 6, 836–846. [Google Scholar] [CrossRef] [PubMed]
- Comai, L.; Tyagi, A.P.; Lysak, M.A. FISH analysis of meiosis in Arabidopsis allopolyploids. Chromosome Res. 2003, 11, 217–226. [Google Scholar] [CrossRef] [PubMed]
- Mestiri, I.; Chagué, V.; Tanguy, A.M.; Huneau, C.; Huteau, V.; Belcram, H.; Coriton, O.; Chalhoub, B.; Jahier, J. Newly synthesized wheat allohexaploids display progenitor-dependent meiotic stability and aneuploidy but structural genomic additivity. New Phytol. 2010, 186, 86–101. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wang, Y.J.; Guo, X.X.; Du, Y.P.; Wang, C.Y.; Ji, W.Q. Chromosomal structural changes and microsatellite variations in newly synthesized hexaploid wheat mediated by unreduced gametes. J. Genet. 2016, 95, 819–830. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Xie, D.; Fan, C.; Zhang, S.; Huang, L.; Ning, S.; Jiang, B.; Zhang, L.; Yuan, Z.; Liu, D.; et al. Chromosome Stability of Synthetic-Natural Wheat Hybrids. Front. Plant Sci. 2021, 12, 654382. [Google Scholar] [CrossRef] [PubMed]
- Chester, M.; Gallagher, J.P.; Symonds, V.V.; Cruz da Silva, A.V.; Mavrodiev, E.V.; Leitch, A.R.; Soltis, P.S.; Soltis, D.E. Extensive chromosomal variation in a recently formed natural allopolyploid species, Tragopogon miscellus (Asteraceae). Proc. Natl. Acad. Sci. USA 2012, 109, 1176–1181. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weimarck, A. Elimination of wheat and rye chromosomes in a strain of octoploid Trit icale as revealed by Giemsa banding technique. Hereditas 1974, 77, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Shkutina, F.M.; Khvostova, V.V. Cytological investigation of Triticale. Theor. Appl. Genet. 1971, 41, 109–119. [Google Scholar] [CrossRef]
- Lukaszewski, A.J.; Gustafson, J.P. Cytogenetics of triticale. Plant Breed. 1987, 5, 41–93. [Google Scholar]
- Sanei, M.; Pickering, R.; Kumke, K.; Nasuda, S.; Houben, A. Loss of centromeric histone H3 (CENH3) from centromeres precedes uniparental chromosome elimination in interspecif ic barley hybrids. Proc. Natl. Acad. Sci. USA 2011, 108, E498–E505. [Google Scholar] [CrossRef] [Green Version]
- Evtushenko, E.V.; Lipikhina, Y.A.; Stepochkin, P.I.; Vershinin, A.V. Cytogenetic and molecular characteristics of rye genome in octoploid triticale (× Triticosecale Wittmack). Comp. Cytogenet. 2019, 13, 423–434. [Google Scholar] [CrossRef]
- Devos, K.M.; Atkinson, M.D.; Chinoy, C.N.; Francis, H.A.; Harcourt, R.L.; Koebner, R.M.; Liu, C.J.; Masojć, P.; Xie, D.X.; Gale, M.D. Chromosomal rearrangements in the rye genome relative to that of wheat. Theor. Appl. Genet. 1993, 85, 673–680. [Google Scholar] [CrossRef]
- Lukaszewski, A.J.; Gustafson, J.P.; Apolinarska, B. Transmission of Chromosomes through the Eggs and Pollen of Triticale x Wheat F1 Hybrids. Theor. Appl. Genet. 1982, 63, 49–55. [Google Scholar] [CrossRef]
- Merker, A. The genome in wheat breeding. Hereditas 1984, 100, 183–191. [Google Scholar] [CrossRef]
- Kalinka, A.; Achrem, M. Reorganization of wheat and rye genomes in octoploid triticale (× Triticosecale). Planta 2018, 247, 807–829. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szakacs, E.; Molnar-Lang, M. Molecular cytogenetic evaluation of chromosome instability in Triticum aestivum–Secale cereale disomic addition lines. J. Appl. Genet. 2010, 51, 149–152. [Google Scholar] [CrossRef] [PubMed]
- Szakacs, E.; Schneider, A.; Rakszegi, M.; Molnar-Lang, M. Addition of chromosome 4R from Hungarian rye cultivar Lovászpatonai confers resistance to stripe rust and outstanding end-use quality in wheat. J. Cereal Sci. 2016, 71, 204–206. [Google Scholar] [CrossRef]
- An, D.; Ma, P.; Zheng, Q.; Fu, S.; Li, L.; Han, F.; Han, G.; Wang, J.; Xu, Y.; Jin, Y.; et al. Development and molecular cytogenetic identification of a new wheat–rye 4R chromosome disomic addition line with resistances to powdery mildew, stripe rust and sharp eyespot. Theor. Appl. Genet. 2019, 132, 257–272. [Google Scholar] [CrossRef]
- Miller, T.E. The homoeologous relationship between the chromosomes of rye and wheat. Current status. Can. J. Genet. Cytol. 1984, 26, 578–589. [Google Scholar] [CrossRef]
- Khalil, H.B.; Ehdaeivand, M.R.; Xu, Y.; Laroche, A.; Gulick, P.J. Identification and characteri zation of rye genes not expressed in allohexaploid triticale. BMC Genom. 2015, 16, 281. [Google Scholar] [CrossRef] [Green Version]
- Silkova, O.G.; Adonina, I.G.; Krivosheina, E.A.; Shchapova, A.I.; Shumny, V.K. Chromosome pairing in meiosis of partially fertile wheat/rye hybrids. Plant Reprod. 2013, 26, 33–41. [Google Scholar] [CrossRef]
- Magnard, J.-L.; Yang, M.; Chen, Y.C.S.; Leary, M.; McCormick, S. The Arabidopsis gene tardy asynchronous meiosis is required for the normal pace and synchrony of cell division during male meiosis. Plant Physiol. 2001, 127, 1157–1166. [Google Scholar] [CrossRef]
- Wang, Y.; Magnard, J.L.; McCormick, S.; Yang, M. Progression through meiosis I and meiosis II in Arabidopsis anthers is regulated by an A-type cyclin predominately expressed in prophase I. Plant Physiol. 2004, 136, 4127–4135. [Google Scholar] [CrossRef] [Green Version]
- D’Erfurth, I.; Cromer, L.; Jolivet, S.; Girard, C.; Horlow, C.; Sun, Y.; To, J.P.; Berchowitz, L.E.; Copenhaver, G.P.; Mercier, R. The cyclin-A CYCA1;2/TAM is required for the meiosis I to meiosis II transition and cooperates with OSD1 for the prophase to first meiotic division transition. PLoS Genet. 2010, 6, e1000989. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hao, M.; Luo, J.; Zeng, D.; Zhang, L.; Ning, S.; Yuan, Z.; Yan, Z.; Zhang, H.; Zheng, Y.; Feuillet, C.; et al. QTug.sau-3B is a major quantitative trait locus for wheat hexaploidization. G3: Genes Genomes Genet. 2014, 15, 1943–1953. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- D’Erfurth, I.; Jolivet, S.; Froger, N.; Catrice, O.; Novatchkova, M.; Mercier, R. Turning meiosis into mitosis. PLoS Biol. 2009, 7, e1000124. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Jha, A.K.; Chen, R.; Doonan, J.H.; Yang, M. Polyploidy-associated genomic instability in Arabidopsis thaliana. Genesis 2010, 48, 254–263. [Google Scholar] [CrossRef] [PubMed]
- Qi, F.; Zhang, F. Cell Cycle Regulation in the Plant Response to Stress. Front. Plant Sci. 2020, 10, 1765. [Google Scholar] [CrossRef] [Green Version]
- Rice, A.; Glick, L.; Abadi, S.; Einhorn, M.; Kopelman, N.M.; Salman-Minkov, A.; Mayzel, J.; Chay, O.; Mayrose, I. The Chromosome Counts Database (CCDB)—A community resource of plant chromosome numbers. New Phytol. 2015, 206, 19–26. [Google Scholar] [CrossRef]
- Hardion, L.; Verlaque, R.; Rosato, M.; Rosselló, J.A.; Vila, B. Impact of polyploidy on fertility variation of Mediterranean Arundo, L. (Poaceae). C R Biol. 2015, 338, 298–306. [Google Scholar] [CrossRef] [PubMed]
- Bowers, J.E.; Paterson, A.H. Chromosome number is key to longevity of polyploid lineages. New Phytol. 2021, 231, 19–28. [Google Scholar] [CrossRef]
- Cheng, F.; Mandakova, T.; Wu, J.; Xie, Q.; Lysak, M.A.; Wang, X.W. Deciphering the diploid ancestral genome of the mesohexaploid Brassica rapa. Plant Cell 2013, 25, 1541–1554. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.Y.; Jin, D.C.; Wang, Z.Y.; Guo, H.; Zhang, L.; Wang, L.; Li, J.P.; Paterson, A.H. Telomere-centric genome repatterning determines recurring chromosome number reductions during the evolution of eukaryotes. New Phytol. 2015, 205, 378–389. [Google Scholar] [CrossRef]
- Schubert, I.; Vu, G.T.H. Genome stability and evolution: Attempting a holistic view. Trends Plant Sci. 2016, 21, 749–757. [Google Scholar] [CrossRef]
- Hudakova, S.; Kunzel, G.; Endo, T.R.; Schubert, I. Barley chromosome arms longer than half of the spindle axis interfere with nuclear divisions. Cytogenet. Genome Res. 2002, 98, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Hidalgo, O.; Pellicer, J.; Christenhusz, M.; Schneider, H.; Leitch, A.R.; Leitch, I.J. Is There an Upper Limit to Genome Size? Trends Plant Sci. 2017, 22, 567–573. [Google Scholar] [CrossRef] [PubMed]





| Hybrid F2 | Hybrid F3 | Hybrid F4 | Hybrids F5 | |||||
|---|---|---|---|---|---|---|---|---|
| Routine Analysis | FISH | |||||||
| Plants | Anthers | Meiocytes+ Microspores | Plants | Anthers | Meiocytes | |||
| 6-1 (subgroup 1a) | 22-4 | 72-3 | 7 | 15 | 1419 | 3 | 7 | 228 |
| 72-4 | 4 | 11 | 826 | 5 | 10 | 489 | ||
| 72-11 | 4 | 17 | 1851 | 2 | 7 | 315 | ||
| 72-12 | 6 | 23 | 2934 | 6 | 10 | 684 | ||
| 6-2 (subgroup 1b) | 23-8 | 76-1 | 11 | 27 | 3304 | - | - | - |
| 23-10 | 77-1 | 4 | 11 | 1007 | - | - | - | |
| 77-4 | 6 | 16 | 1482 | - | - | - | ||
| 77-8 | 6 | 12 | 961 | - | - | - | ||
| 73-1 (group 3) | 41-7 | 87-1 | 9 | 43 | 3215 | 3 | 7 | 112 |
| 41-16 | 88-1 | 9 | 47 | 2862 | 2 | 5 | 72 | |
| 88-2 | 6 | 34 | 3117 | - | - | - | ||
| 41-22 | 89-6 | 8 | 41 | 3096 | 3 | 6 | 82 | |
| Correlation Coefficient | Interpretation |
|---|---|
| 0.90 to 1.00 (−0.90 to −1.00) | Very high positive (negative) correlations, very dependable associations |
| 0.70 to 0.90 (−0.70 to −0.90) | High positive (negative) correlations, marked associations |
| 0.50 to 0.70 (−0.50 to −0.70) | Moderate positive (negative) correlations, substantial associations |
| 0.30 to 0.50 (−0.30 to −0.50) | Low positive (negative) correlations, defined but small associations |
| 0 to 0.30 (0 to −0.30) | Negligible correlations |
| Homeologous Group 4 | Plants/% |
|---|---|
| AABBDDRR | 76/58.9% |
| A-BBDDRR | 1/0.77% |
| AA-BDDRR | 4/3.1% |
| AABB-DRR | 3/2.32% |
| --BBDDRR | 1/0.77% |
| AA--DDRR | 5/3.87% |
| A-B-DDRR | 1/0.77% |
| A-BB-DRR | 2/1.55% |
| AABBDD-R | 3/2.32% |
| AA-BDDR- | 1 /0.77% |
| AABBDD-RL | 2/1.55% |
| AABBD-RRL | 1/0.77% |
| AABBLDDRR | 1/0.77% |
| AABLBLDDRR | 7/5.43% |
| AABBDDRS- | 1/0.77% |
| AABBDDRRR | 1/0.77% |
| AABBDD-- | 19/14.73% |
| Total | 129/100% |
| Hybrid F1 | Hybrid F2 | Hybrid F3 | Hybrid F4 | Hybrids F5 | |
|---|---|---|---|---|---|
| Meiocytes with Micronuclei, % | Seed Set | ||||
| 4-7 | 6-1 (subgroup 1a) | 22-4 | 72-3 | 47 ± 3.75 | 67 ± 6.2 |
| 72-4 | 56.35 ± 8.3 | 61.83 ± 9.2 | |||
| 72-11 | 70.26 ± 6.2 | 81.88 ± 9.4 | |||
| 72-12 | 60.37 ± 6.0 | 102.47 ± 9.3 | |||
| 6-2 (subgroup 1b) | 23-8 | 76-1 | 13.93 ± 1.89 | 55.93 ± 6.7 | |
| 23-10 | 77-1 | 65.21 ± 9.4 | 37.19 ± 8.5 | ||
| 77-4 | 46.5 ± 2.45 | 40.92 ± 7.9 | |||
| 77-8 | 46.95 ± 7.95 | 40 ± 6.2 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silkova, O.G.; Ivanova, Y.N.; Loginova, D.B.; Solovey, L.A.; Sycheva, E.A.; Dubovets, N.I. Karyotype Reorganization in Wheat–Rye Hybrids Obtained via Unreduced Gametes: Is There a Limit to the Chromosome Number in Triticale? Plants 2021, 10, 2052. https://doi.org/10.3390/plants10102052
Silkova OG, Ivanova YN, Loginova DB, Solovey LA, Sycheva EA, Dubovets NI. Karyotype Reorganization in Wheat–Rye Hybrids Obtained via Unreduced Gametes: Is There a Limit to the Chromosome Number in Triticale? Plants. 2021; 10(10):2052. https://doi.org/10.3390/plants10102052
Chicago/Turabian StyleSilkova, Olga G., Yulia N. Ivanova, Dina B. Loginova, Lilia A. Solovey, Elena A. Sycheva, and Nadezhda I. Dubovets. 2021. "Karyotype Reorganization in Wheat–Rye Hybrids Obtained via Unreduced Gametes: Is There a Limit to the Chromosome Number in Triticale?" Plants 10, no. 10: 2052. https://doi.org/10.3390/plants10102052
APA StyleSilkova, O. G., Ivanova, Y. N., Loginova, D. B., Solovey, L. A., Sycheva, E. A., & Dubovets, N. I. (2021). Karyotype Reorganization in Wheat–Rye Hybrids Obtained via Unreduced Gametes: Is There a Limit to the Chromosome Number in Triticale? Plants, 10(10), 2052. https://doi.org/10.3390/plants10102052






