Spatio-Temporal Prediction of the Epidemic Spread of Dangerous Pathogens Using Machine Learning Methods
Abstract
1. Introduction
2. Materials and Methods
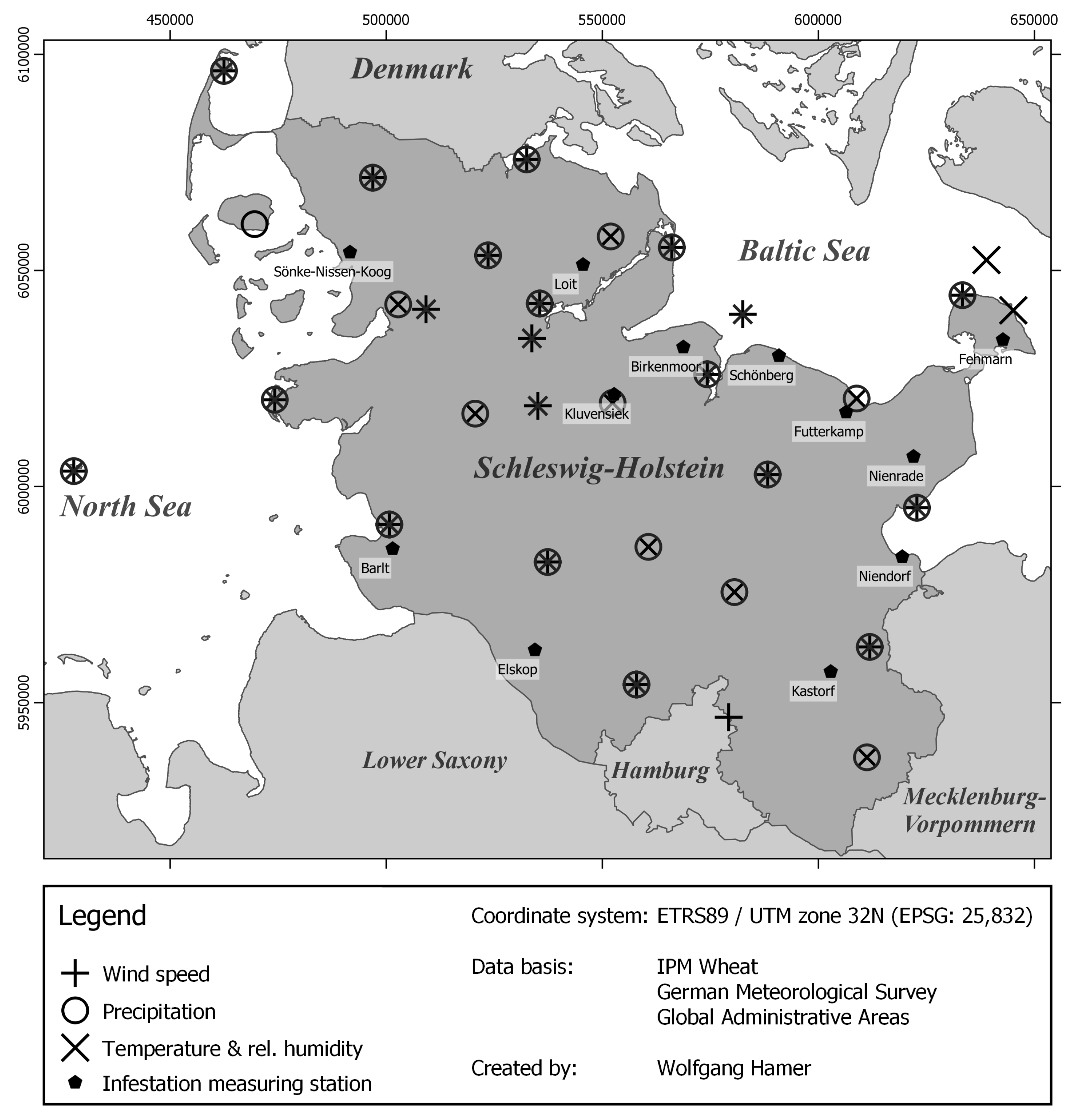
2.1. Study Area
2.2. Sources of Data Used
2.3. Methods Applied
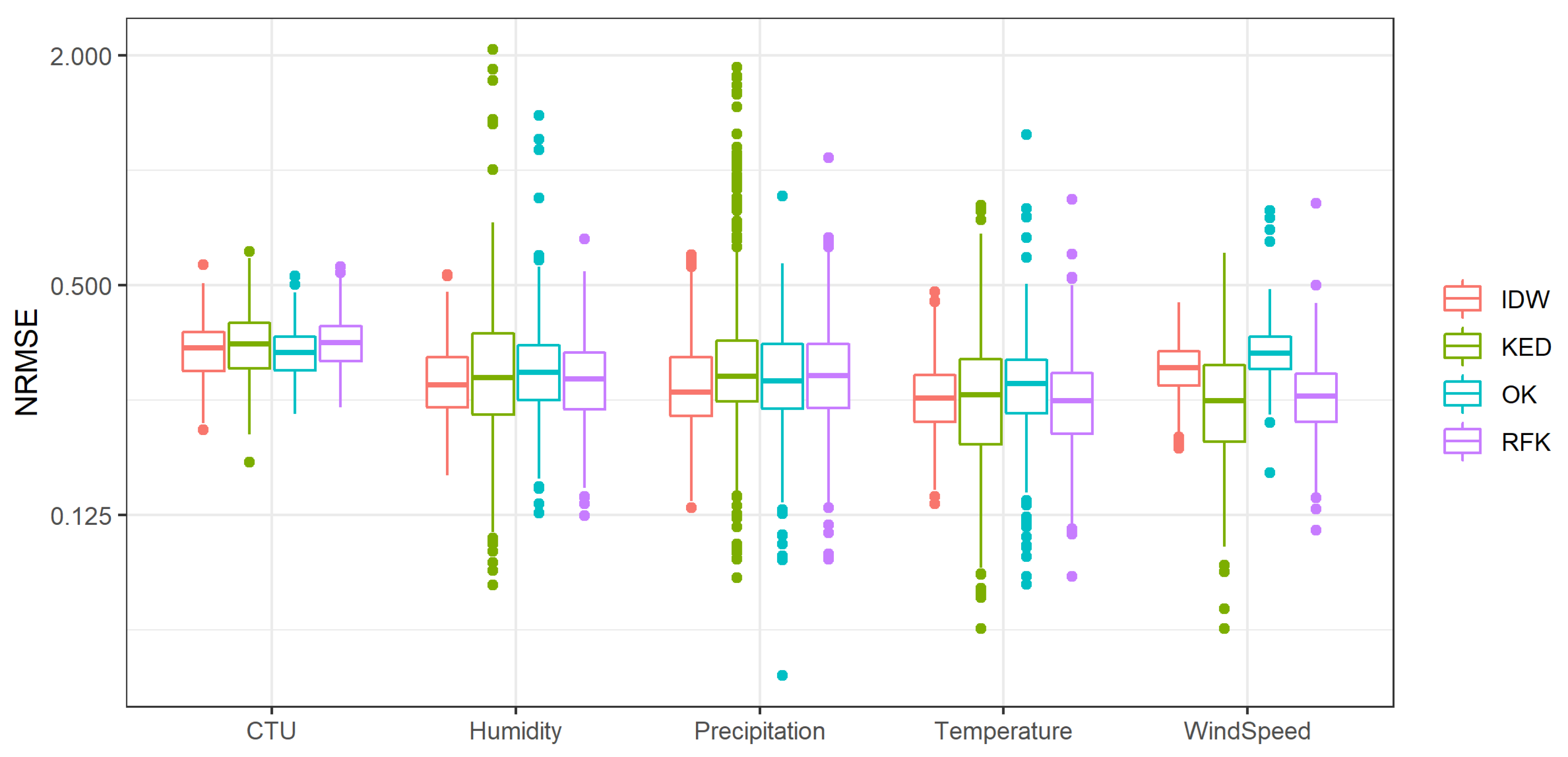
- Inverse distance weighting (IDW) is a common deterministic spatial interpolation method, which is defined as a weighted average of the data point values [27]. The weighting itself is a function of the distance:where is the value to be estimated at the location , is the known value at a specific location , is the distance between the estimated and the known data points, p is the inverse distance power, and n is the number of known data points closest to the estimated location. Although IDW does not model the spatial autocorrelation of the target variable (as the methods described below do), the procedure can still achieve good results. Wagner et al. [28] and Borges et al. [29], for example, found that the IDW interpolation achieved the best results for interpolating precipitation.
- Ordinary kriging (OK) was first established by Krige [30] and mathematically derived by Matheron [31]. Using this statistical interpolation method, the spatial dependence of the variable is not just assumed (as it would be with the IDW method), but analysed and integrated in the regionalisation. The analysis of the spatial dependence is based on variography: the similarity of point pairs (semivariance) is compared to their distance, and a function is adapted to the semivariance, which decreases with increasing distance Matheron [31]. By means of this variogram model, it is possible to calculate the weights () by which the values of the surrounding points () are multiplied, according to the krige Equation (2) [31,32]:
- Kriging with external drift (KED) uses the integration of independent variables in the process of Kriging interpolation. The principle of the procedure was established by Matheron [33] as a combination of OK and multiple linear regression of the dependent variable (the parameter of interest) with the independent variable (e.g., the coordinates or an elevation model). As described by Equation (3), a linear regression is fitted to the variables to predict the dependent variable by the (already regionalised) independent variables. The spatial prediction of the dependent variable starts with the application of the regression model () on the regionalised independent variables. In order to increase the accuracy of the prediction, the Kriging procedure (, as in Equation (2), is used to interpolate the residuals of the regression model. These two spatial data sets are then summed to calculate the aim variable ():
- Random forest kriging (RFK) is similar to the KED procedure. However, instead of multiple linear regression, the relationship between the dependent and independent variables is represented by a random forest. The functionality of such a random forest is explained in more detail in the description of the machine learning methods.
- Decision trees (DT) are based on the idea of recursive partitioning—splitting the data repeatedly into smaller subsets until they are homogeneous regarding the target variable. Of the different algorithmic methods created to find the ideal split, the C5.0 DT algorithm (developed by Ross Quinlan as an improvement to his C4.5 algorithm [36]), which uses the concept of entropy to create subsets with maximum purity, is the most common. The entropy for a specific subset is calculated as:where S is the subset, c is the number of class levels, and is the proportion of values in the specific class [37]. The quality of the splits, thus, is based on the summed entropy values of all subsets:with the total entropy (T) and the weighting () obtained by the proportion of examples in the subset [37]. The entropy values of the potential splits are weighted against each other, resulting in the information gain of potential splits, of which the one with the highest information gain is chosen.
- Random forests (RF), developed by Breiman [38], combine the aforementioned DT with the bagging procedure, which uses bootstrap sampling—a random sampling with replacement method—to generate multiple predictions for a data set using the same prediction method [39]. The individual results are combined into one final prediction, using a plurality vote for classification and the average value for numerical outputs. The RF algorithm creates random subsets of the data set at each node of the developing tree [38]. The splits are evaluated using estimates, which test the ongoing grown trees at the cases not yet integrated into the model, in order to give an error estimate [38]. As a result, a forest of randomly grown DTs emerges. The combination of these trees, as an average for regression tasks and as the most frequent value for classification tasks, results in the final prediction of the RF model. RFs are assumed to be less prone to overfitting, as only parts of the data set are used to generate the individual trees [40]. Furthermore, they are assumed to be better at learning from larger data sets with a large number of features [37].
- Boosted decision trees (BDT) utilise the boosting procedure. Similar to the bagging method explained above, the boosting procedure creates a number of DTs [41]. In contrast to the decision trees of the RFs, however, the trees in BDT do not consist of random sub-data sets. At first, only one DT is generated, based on the entire data set. The weight of misclassified instances in this first tree is, then, increased when the next tree is generated [42]. This process is repeated until the requested number of trees is reached. The final prognosis of this method is given by the majority decision of the generated DTs.
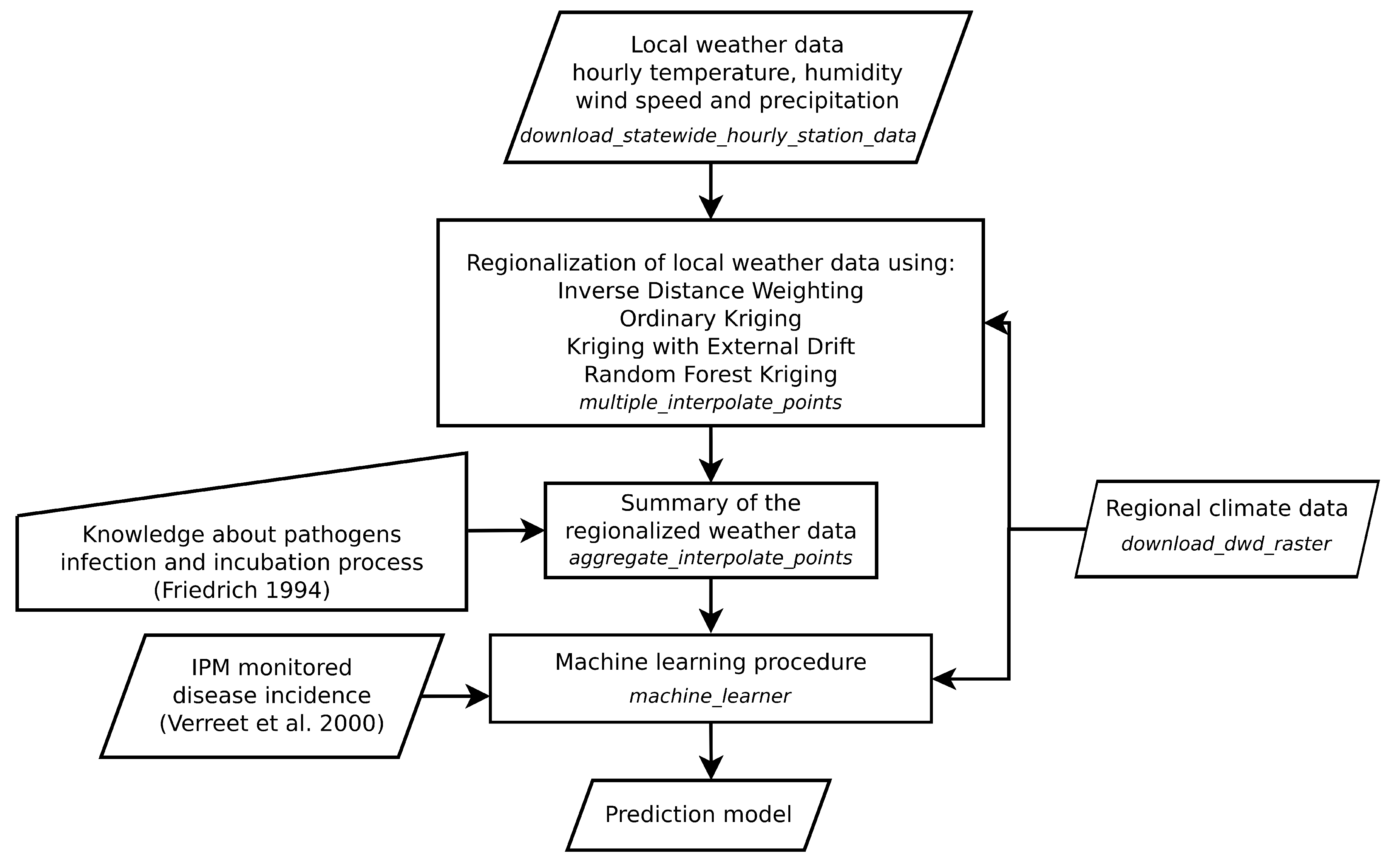
2.4. Modelling Approach
2.5. Use Case: Predict Powdery Mildew Infestation Events
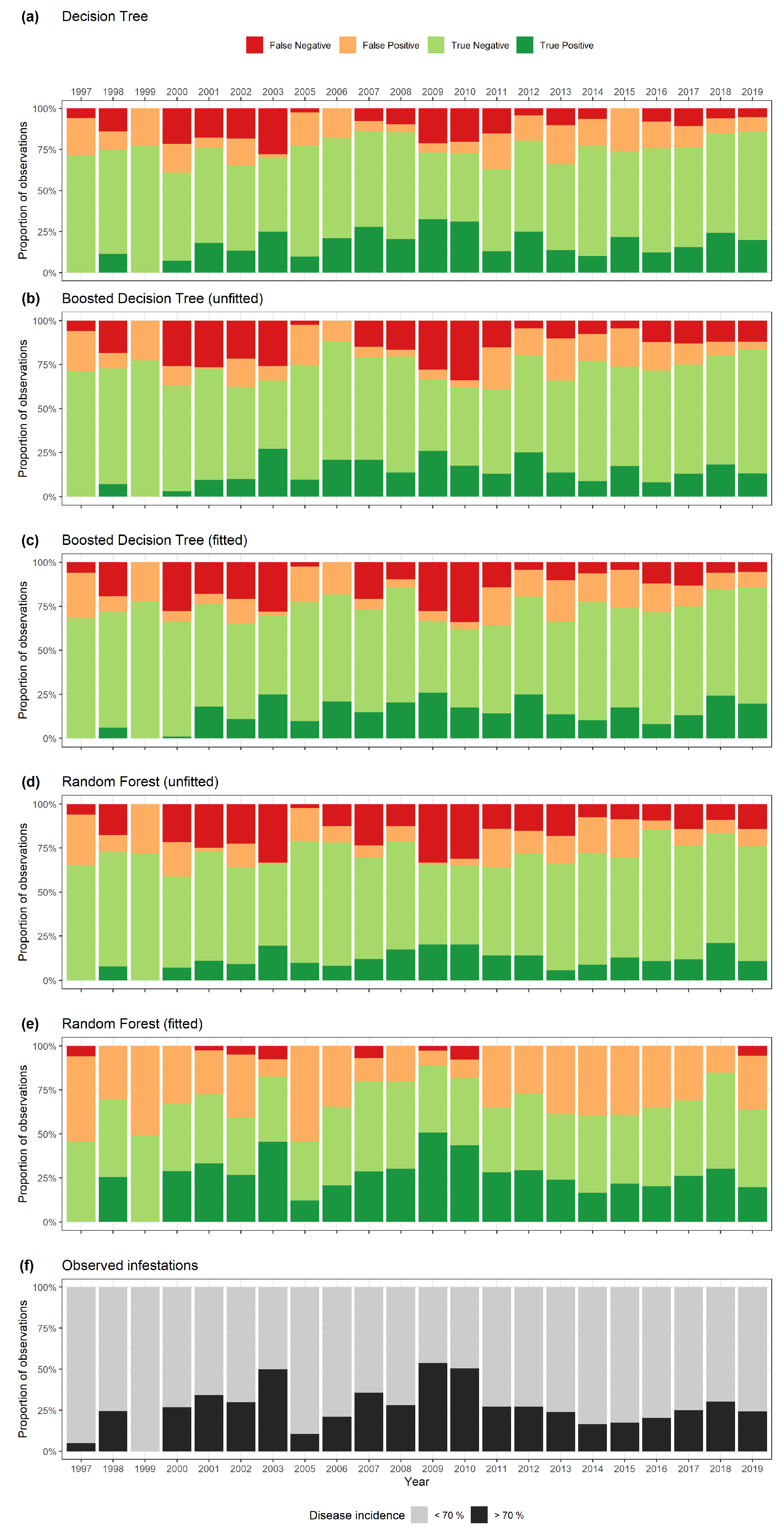
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| BDT | boosted decision tree |
| DT | decision tree |
| f | fitted (by ROC AUC) |
| IDW | inverse distance weighting |
| IPM | integrated pest management |
| KED | kriging with external drift |
| OK | ordinary kriging |
| RF | random forest |
| RFK | random forest kriging |
| ROC | receive operating characteristic |
| ROC AUC | receive operating characteristic area under the curve |
| uf | unfitted (by ROC AUC) |
References
- FAOSTAT Pesticides Use. Available online: fenixservices.fao.org/faostat/static/bulkdownloads/Inputs_Pesticides_Use_E_All_Data.zip (accessed on 15 October 2019).
- European Parliament. DIRECTIVE 2009/128/EC OF THE EUROPEAN PARLIAMENT AND OF THE COUNCIL of 21 October 2009 Establishing a Framework for Community Action to Achieve the Sustainable Use of Pesticides. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=CELEX:02009L0128-20190726 (accessed on 15 October 2019).
- Hau, B. Epidemiologische Simulatoren als Instrumente der Systemanalyse mit Besonderer Berücksichtigung eines Modells des Gerstenmehltaus; Acta Phytomedica; P. Parey: Hamburg, Germany, 1985. [Google Scholar]
- Friedrich, S. Prognose der Infektionswahrscheinlichkeit durch Echten Mehltau an Winterweizen (Erysiphe graminis DC. f. sp. tritici) anhand Meteorologischer Eingangsparameter; Wissenschaftsverlag Mainz Aachen: Mainz, Germany, 1994. [Google Scholar]
- Jensen, A.L.; Jensen, F.V. MIDAS—An Influence Diagram for Management of Mildew in Winter Wheat. In UAI’96: Proceedings of the Twelfth International Conference on Uncertainty in Artificial Intelligence; Morgan Kaufmann Publishers Inc.: San Francisco, CA, USA, 1996; pp. 349–356. [Google Scholar]
- Bruns, J.B. Untersuchungen zur wetterbasierten Befallssimulation und Verlustprognose von Echtem Mehltau (Erysiphe graminis D.C. f. sp. tritici Marchal) an Winterweizen. Ph.D. Thesis, Georg-August-Universität Göttingen, Göttingen, Germany, 1996. [Google Scholar]
- Rossi, V.; Giosuè, S. A dynamic simulation model for powdery mildew epidemics on winterwheat. EPPO Bull. 2003, 33, 389–396. [Google Scholar] [CrossRef]
- Willocquet, L.; Aubertot, J.; Lebard, S.; Robert, C.; Lannou, C.; Savary, S. Simulating multiple pest damage in varying winter wheat production situations. Field Crops Res. 2008, 107, 12–28. [Google Scholar] [CrossRef]
- Zhang, S.; Shang, Y.; Wang, L. Plant disease recognition based on plant leaf image. J. Anim. Plant Sci. 2015, 25, 42–45. [Google Scholar]
- Delwiche, S.R.; Yang, I.C.; Graybosch, R.A. Multiple view image analysis of freefalling U.S. wheat grains for damage assessment. Comput. Electron. Agric. 2013, 98, 62–73. [Google Scholar] [CrossRef]
- Lu, J.; Ehsani, R.; Shi, Y.; Abdulridha, J.; de Castro, A.I.; Xu, Y. Field detection of anthracnose crown rot in strawberry using spectroscopy technology. Comput. Electron. Agric. 2017, 135, 289–299. [Google Scholar] [CrossRef]
- Mwebaze, E.; Biehl, M. Prototype-Based Classification for Image Analysis and Its Application to Crop Disease Diagnosis; Springer International Publishing: Cham, Switzerland, 2016; pp. 329–339. [Google Scholar] [CrossRef]
- Fränzle, O. Streifzug durch 6000 Jahre Landnutzungs- und Landschaftswandel in Schleswig-Holstein; Chapter Reliefentwicklung und Bodenbildung in Schleswig-Holstein, EcoSys; Verein zur Förderung der Ökosystemforschung zu Kiel e.V.: Kiel, Germany, 2004; Volume 41, pp. 11–35. [Google Scholar]
- Meynen, E.; Schmithüsen, J. Handbuch der Naturräumlichen Gliederung Deutschlands: 1953–1962; Bundesanst. für Landeskunde u. Raumforschung: Bad Godesberg, Germany, 1962; Number Bd. 2. [Google Scholar]
- Schlenger, H.; Paffen, K.; Stewig, R. Schleswig-Holstein: Ein Geographisch-Landeskundlicher Exkursionführer; Schriften des Geographischen Instituts der Universität Kiel, Hirt: Kiel, Germany, 1969. [Google Scholar]
- DWD 2019 Open Data Server. Available online: https://www.dwd.de/EN/ourservices/opendata/opendata.html (accessed on 7 May 2019).
- Verreet, J.; Klink, H.; Hoffmann, G. Regional monitoring for disease prediction and optimization of plant protection measuares: The IPM wheat model. Plant Dis. 2000, 84, 816–826. [Google Scholar] [CrossRef]
- Zadoks, J.C.; Chang, T.T.; Konzak, C.F. A decimal code for the growth stages of cereals. Weed Res. 1974, 14, 415–421. [Google Scholar] [CrossRef]
- Cao, X.; Duan, X.; Zhou, Y.; Luo, Y. Dynamics in concentrations of Blumeria graminis f. sp tritici conidia and its relationship to local weather conditions and disease index in wheat. Eur. J. Plant Pathol. 2012, 132, 525–535. [Google Scholar] [CrossRef]
- Eckhardt, H.; Steubing, L.; Kranz, J. Untersuchungen zur Infektionseffizienz, Inkubations- und Latenzzeit beim Gerstenmehltau Erysiphe graminis f. sp. hordei. J. Plant Dis. Prot. 1984, 91, 590–600. [Google Scholar]
- Beest, D.E.T.; Paveley, N.D.; Shaw, M.W.; van den Bosch, F. Disease-weather relationships for powdery mildew and yellow rust on winter wheat. Phytopathology 2008, 98, 609–617. [Google Scholar] [CrossRef]
- Hau, B.; de Vallavieille-Pope, C. Wind-dispersed diseases. In The Epidemiology of Plant Diseases; Cooke, B., Jones, D.G., Kaye, B., Eds.; Springer: Dordrecht, The Netherlands, 2006; pp. 387–416. [Google Scholar] [CrossRef]
- Merchán, V.; Kranz, J. Studies on the effect of rain on the infection of wheat by Erysiphe graminis DC. f. sp. tritici Marchal. J. Plant Dis. Prot. 1986, 93, 255–261. [Google Scholar]
- Soltani, A.; Sinclair, T. Modeling Physiology of Crop Development, Growth and Yield; CAB Books; CABI: Wallingford, UK, 2012. [Google Scholar]
- Klink, H. Geoepidemiologische Erhebungen von Weizenpathogenen in Schleswig-Holstein unter Anwendung und Entwicklung des Integrierten Pflanzenschutzsystems (IPS-Modell Weizen) Für Einen Minimierten, Bedarfsgerechten Fungizideinsatz (1993–1996). Ph.D. Thesis, Christian-Albrechts-Universität zu Kiel, Kiel, Germany, 1997. [Google Scholar]
- Hengl, T. A Practical Guide to Geostatistical Mapping; Amsterdam University Press: Amsterdam, The Netherlands, 2011. [Google Scholar]
- Shepard, D. A Two-dimensional Interpolation Function for Irregularly-spaced Data. In ACM’68: Proceedings of the 1968 23rd ACM National Conference; ACM: New York, NY, USA, 1968; pp. 517–524. [Google Scholar] [CrossRef]
- Wagner, P.D.; Fiener, P.; Wilken, F.; Kumar, S.; Schneider, K. Comparison and evaluation of spatial interpolation schemes for daily rainfall in data scarce regions. J. Hydrol. 2012, 464–465, 388–400. [Google Scholar] [CrossRef]
- Borges, P.d.A.; Franke, J.; da Anunciação, Y.M.T.; Weiss, H.; Bernhofer, C. Comparison of spatial interpolation methods for the estimation of precipitation distribution in Distrito Federal, Brazil. Theor. Appl. Climatol. 2016, 123, 335–348. [Google Scholar] [CrossRef]
- Krige, D.G. A Statistical Approach to Some Mine Valuation and Allied Problems on the Witwatersrand; University of the Witwatersrand: Johannesburg, South Africa, 1951. [Google Scholar]
- Matheron, G. Principles of geostatistics. Econ. Geol. 1963, 58, 1246–1266. [Google Scholar] [CrossRef]
- Cressie, N. The origins of kriging. Math. Geol. 1990, 22, 239–252. [Google Scholar] [CrossRef]
- Matheron, G. Le Krigeage Universel; Cahiers du Centre de Morphologie Mathėmatique; Ecole des Mines de Paris: Fontainebleau, France, 1969. [Google Scholar]
- Benavides, R.; Montes, F.; Rubio, A.; Osoro, K. Geostatistical modelling of air temperature in a mountainous region of Northern Spain. Agric. Forest Meteorol. 2007, 146, 173–188. [Google Scholar] [CrossRef]
- Eguía, P.; Granada, E.; Alonso, J.; Arce, E.; Saavedra, A. Weather datasets generated using kriging techniques to calibrate building thermal simulations with TRNSYS. J. Build. Eng. 2016, 7, 78–91. [Google Scholar] [CrossRef]
- Quinlan, J.R. C4.5: Programs for Machine Learning; Morgan Kaufmann Publishers Inc.: San Francisco, CA, USA, 1993. [Google Scholar]
- Lantz, B. Machine Learning with R, 2nd ed.; Packt Publishing Ltd.: Birmingham, UK, 2015. [Google Scholar]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Breiman, L. Bagging predictors. Mach. Learn. 1996, 24, 123–140. [Google Scholar] [CrossRef]
- Wyner, A.J.; Olson, M.; Bleich, J.; Mease, D. Explaining the success of adaboost and random forests as interpolating classifiers. J. Mach. Learn. Res. 2017, 18, 1–33. [Google Scholar]
- Quinlan, J.R. Bagging, Boosting, and C4. 5; AAAI/IAAI: Palo Alto, CA, USA, 1996; Volume 1, pp. 725–730. [Google Scholar]
- Schapire, R.; Freund, Y. Boosting: Foundations and Algorithms; Adaptive Computation and Machine Learning; MIT Press: Cambridge, MA, USA, 2012. [Google Scholar]
- Hamer, W. Papros–PAthogen PROgnosis System; 2019. Available online: https://zenodo.org/record/2574046 (accessed on 7 May 2019). [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- DWD 2016 Multi-Annual Means of Grids of Air Temperature (2m) over Germany 1981–2010, Version v1.0. Available online: ftp://opendata.dwd.de/climate_environment/CDC/grids_germany/multi_annual/air_temperature_mean/DESCRIPTION_gridsgermany_multi_annual_air_temperature_mean_8110_en.pdf (accessed on 7 May 2019).
- Pietzsch, S.; Bissolli, P. A modified drought index for WMO RA VI. Adv. Sci. Res. 2011, 6, 275–279. [Google Scholar] [CrossRef]
- DWD 2014 Gridded Mean of Annual Wind Speeds from 10 m to 100 m (in 10 m Steps) above Ground and Weibull Parameters, for Germany, Version V0.1. Available online: ftp://opendata.dwd.de/climate_environment/CDC/grids_germany/multi_annual/wind_parameters/resol_1000x1000/DESCRIPTION_gridsgermany_resol_1000x1000_en.pdf (accessed on 7 May 2019).
- EU Copernicus Programme. European Digital Elevation Model (EU-DEM), Version 1.1; European Environment Agency: 2016. Available online: https://land.copernicus.eu/imagery-in-situ/eu-dem/eu-dem-v1.1/view (accessed on 7 May 2019).
- Nicodemus, K.K. Letter to the Editor: On the stability and ranking of predictors from random forest variable importance measures. Brief. Bioinform. 2011, 12, 369. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, M.; Quinlan, R. C50: C5.0 Decision Trees and Rule-Based Models; R Package Version 0.1.2. 2018. Available online: https://cran.r-project.org/web/packages/C50/index.html (accessed on 7 May 2019).
- Liaw, A.; Wiener, M. Classification and Regression by randomForest. R News 2002, 2, 18–22. [Google Scholar]
- Fawcett, T. An introduction to ROC analysis. Pattern Recognit. Lett. 2006, 27, 861–874. [Google Scholar] [CrossRef]
- Altman, D.G.; Bland, J.M. Diagnostic tests. 1: Sensitivity and specificity. BMJ Br. Med J. 1994, 308, 1552. [Google Scholar] [CrossRef] [PubMed]
- Wilcoxon, F. Individual Comparisons by Ranking Methods. Biom. Bull. 1945, 1, 80–83. [Google Scholar] [CrossRef]
- Powers, D.M. Evaluation: From precision, recall and F-measure to ROC, informedness, markedness & correlation. J. Mach. Learn. Technol. 2011, 2, 2229–3981. [Google Scholar]
- Aggarwal, P.; Kalra, N.; Chander, S.; Pathak, H. InfoCrop: A dynamic simulation model for the assessment of crop yields, losses due to pests, and environmental impact of agro-ecosystems in tropical environments. I. Model description. Agric. Syst. 2006, 89, 1–25. [Google Scholar] [CrossRef]
- Wen, L.; Bowen, C.R.; Hartman, G.L. Prediction of Short-Distance Aerial Movement of Phakopsora pachyrhizi Urediniospores Using Machine Learning. Phytopathology 2017, 107, 1187–1198. [Google Scholar] [CrossRef]
- FAOSTAT Crops Production. Available online: fenixservices.fao.org/faostat/static/bulkdownloads/Production_Crops_E_All_Data.zip (accessed on 15 October 2019).
| Modell Feature | Selected Parameter |
|---|---|
| Air temperature interpolation method | Random Forest Kriging |
| Air humidity interpolation method | Inverse Distance Weighting |
| Precipitation interpolation method | Inverse Distance Weighting |
| Wind speed interpolation method | Kriging with External Drift |
| CTU interpolation method | Ordinary Kriging |
| Target variable | exceedance of 70 % disease incidence |
| minimum air temperature | |
| mean air temperature | |
| maximum air temperature | |
| minimum air humidity | |
| mean air humidity | |
| maximum air humidity | |
| minimum precipitation | |
| mean precipitation | |
| Independent variables | maximum precipitation |
| minimum wind speed | |
| mean wind speed | |
| maximum wind speed | |
| CTU (based upon air temperature) | |
| elevation information [48] | |
| air temperature (climatic) [16] | |
| precipitation (climatic) [16] | |
| drought index (climatic) [16] | |
| wind speed (climatic) [16] |
| Decision Tree | Boosted DT (uf) | Random Forest (uf) | Boosted DT (f) | Random Forest (f) | |
|---|---|---|---|---|---|
| Accuracy | 0.751 | 0.727 | 0.711 | 0.733 | 0.679 |
| Sensitivity | 0.596 | 0.475 | 0.404 | 0.503 | 0.919 |
| Specificity | 0.812 | 0.826 | 0.832 | 0.824 | 0.584 |
| Precision | 0.556 | 0.520 | 0.488 | 0.531 | 0.466 |
| ROC AUC | 0.704 | 0.651 | 0.618 | 0.664 | 0.752 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hamer, W.B.; Birr, T.; Verreet, J.-A.; Duttmann, R.; Klink, H. Spatio-Temporal Prediction of the Epidemic Spread of Dangerous Pathogens Using Machine Learning Methods. ISPRS Int. J. Geo-Inf. 2020, 9, 44. https://doi.org/10.3390/ijgi9010044
Hamer WB, Birr T, Verreet J-A, Duttmann R, Klink H. Spatio-Temporal Prediction of the Epidemic Spread of Dangerous Pathogens Using Machine Learning Methods. ISPRS International Journal of Geo-Information. 2020; 9(1):44. https://doi.org/10.3390/ijgi9010044
Chicago/Turabian StyleHamer, Wolfgang B., Tim Birr, Joseph-Alexander Verreet, Rainer Duttmann, and Holger Klink. 2020. "Spatio-Temporal Prediction of the Epidemic Spread of Dangerous Pathogens Using Machine Learning Methods" ISPRS International Journal of Geo-Information 9, no. 1: 44. https://doi.org/10.3390/ijgi9010044
APA StyleHamer, W. B., Birr, T., Verreet, J.-A., Duttmann, R., & Klink, H. (2020). Spatio-Temporal Prediction of the Epidemic Spread of Dangerous Pathogens Using Machine Learning Methods. ISPRS International Journal of Geo-Information, 9(1), 44. https://doi.org/10.3390/ijgi9010044





