Molecular Characterization of a Novel Family VIII Esterase with β-Lactamase Activity (PsEstA) from Paenibacillus sp.
Abstract
1. Introduction
2. Materials and Methods
2.1. Sequence Analysis and Model Building
2.2. Protein Purification
2.3. Characterization of PsEstA
2.4. pH Indicator-Based Hydrolysis Assay
2.5. β-Lactamase Assay
2.6. Immobilization of PsEstA
3. Results and Discussion
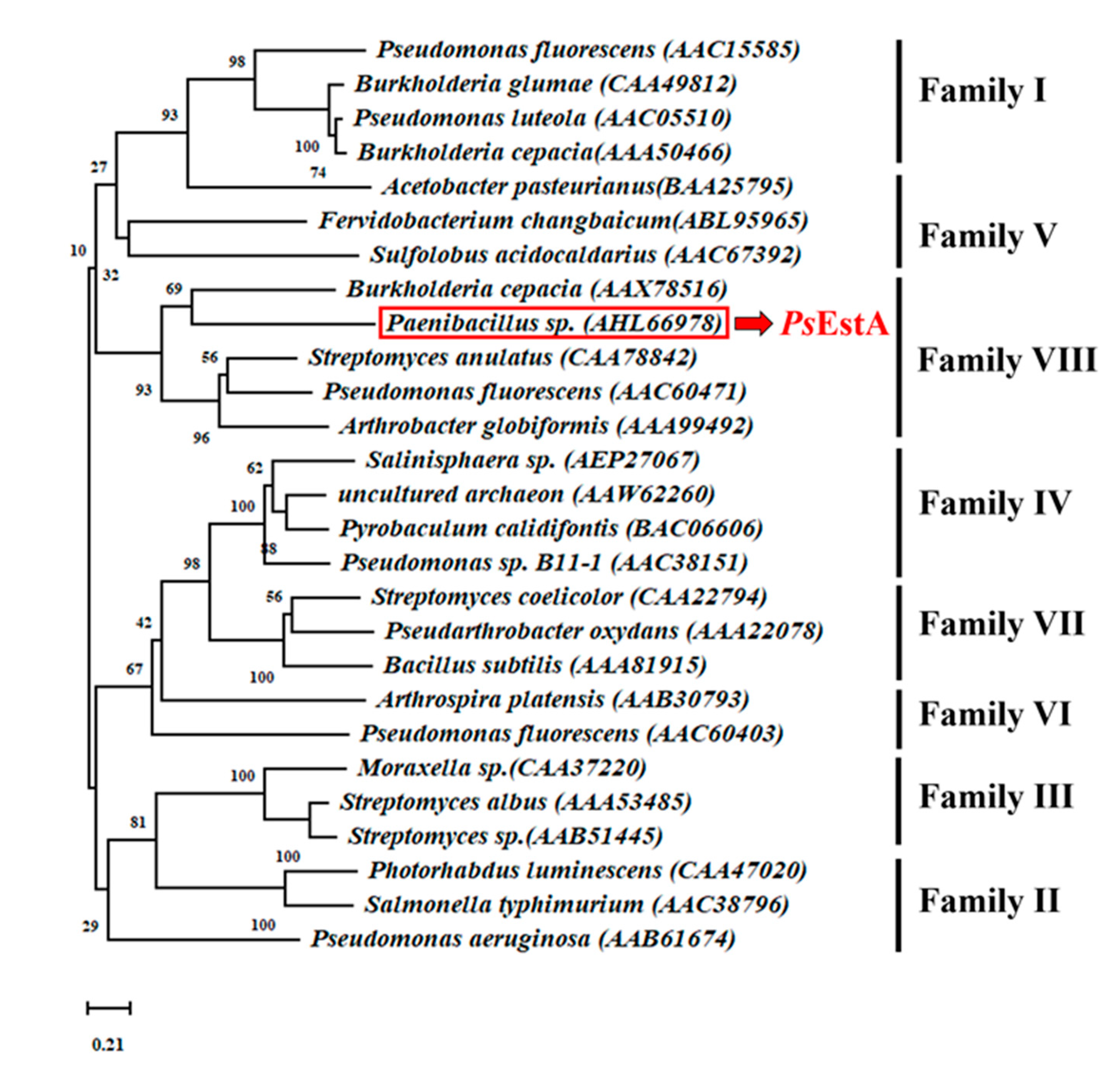
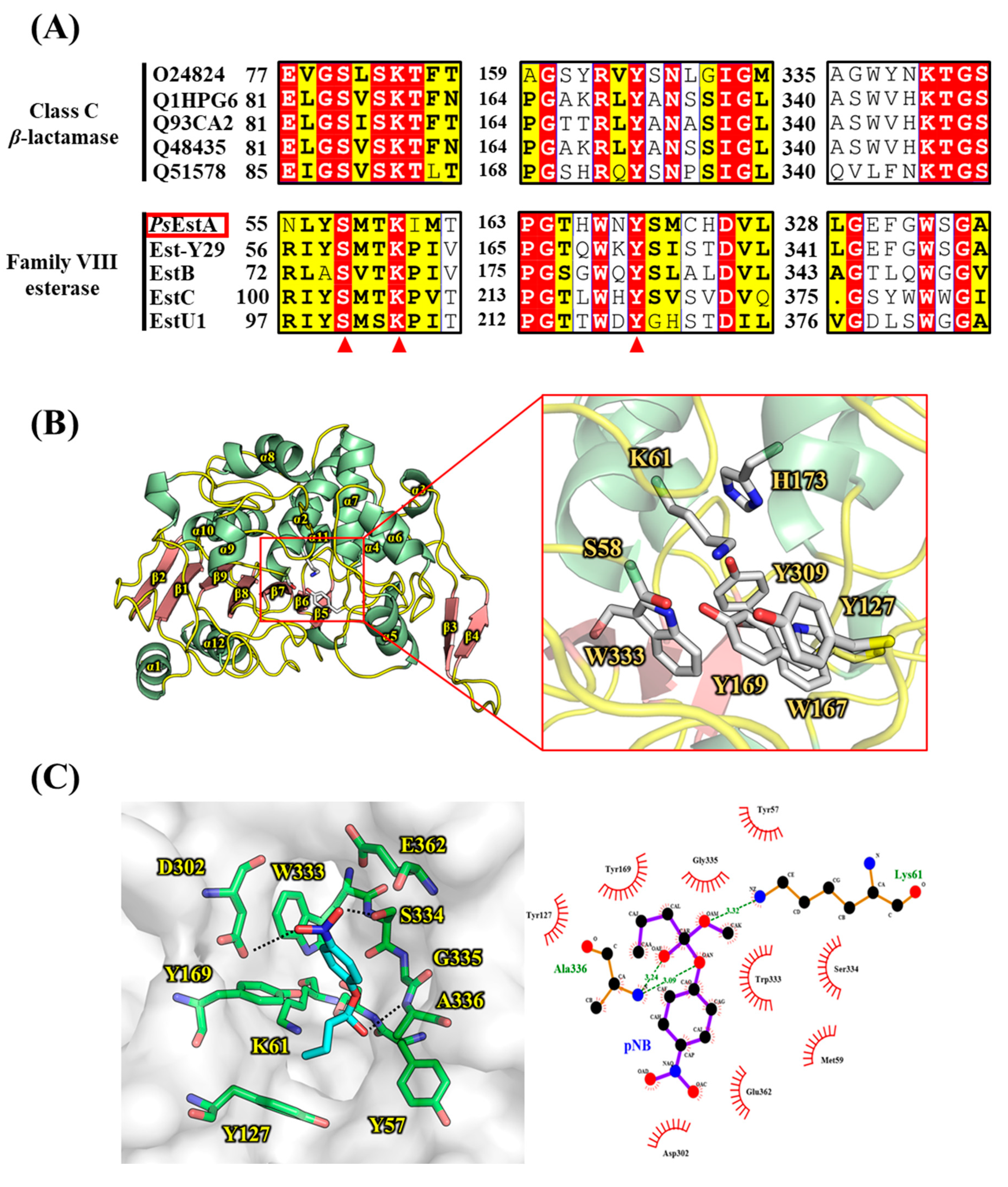
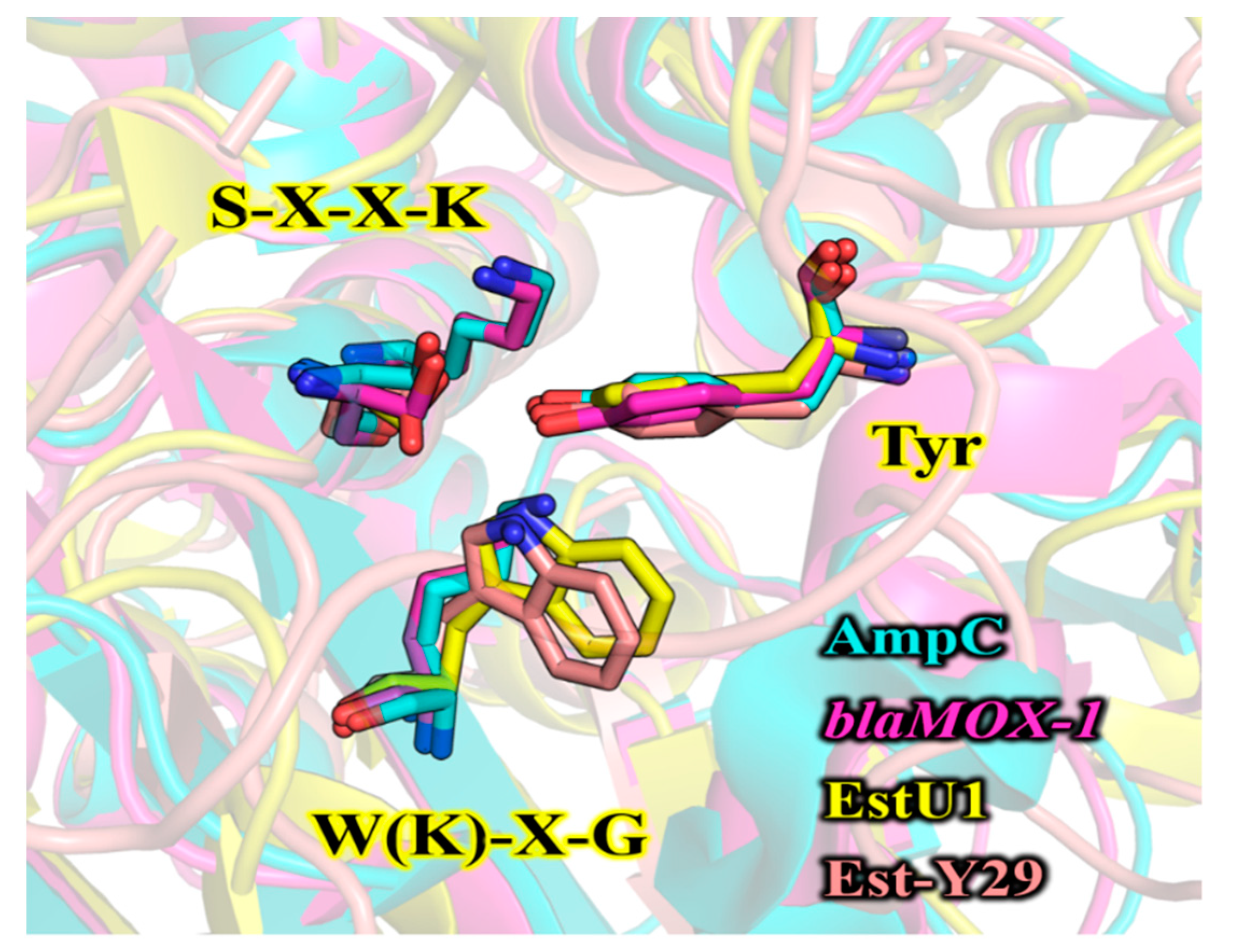
3.1. Bioinformatic Analysis
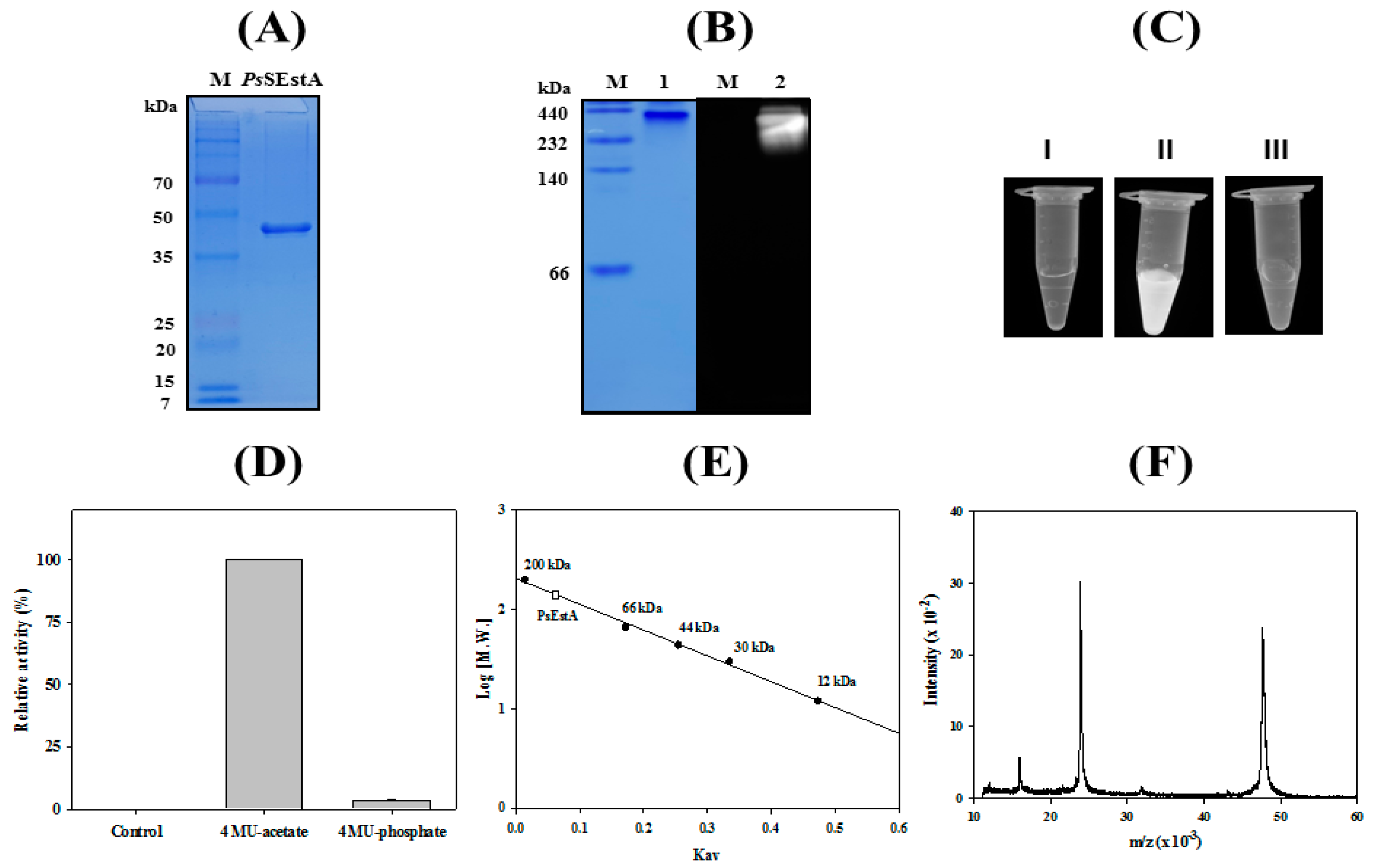
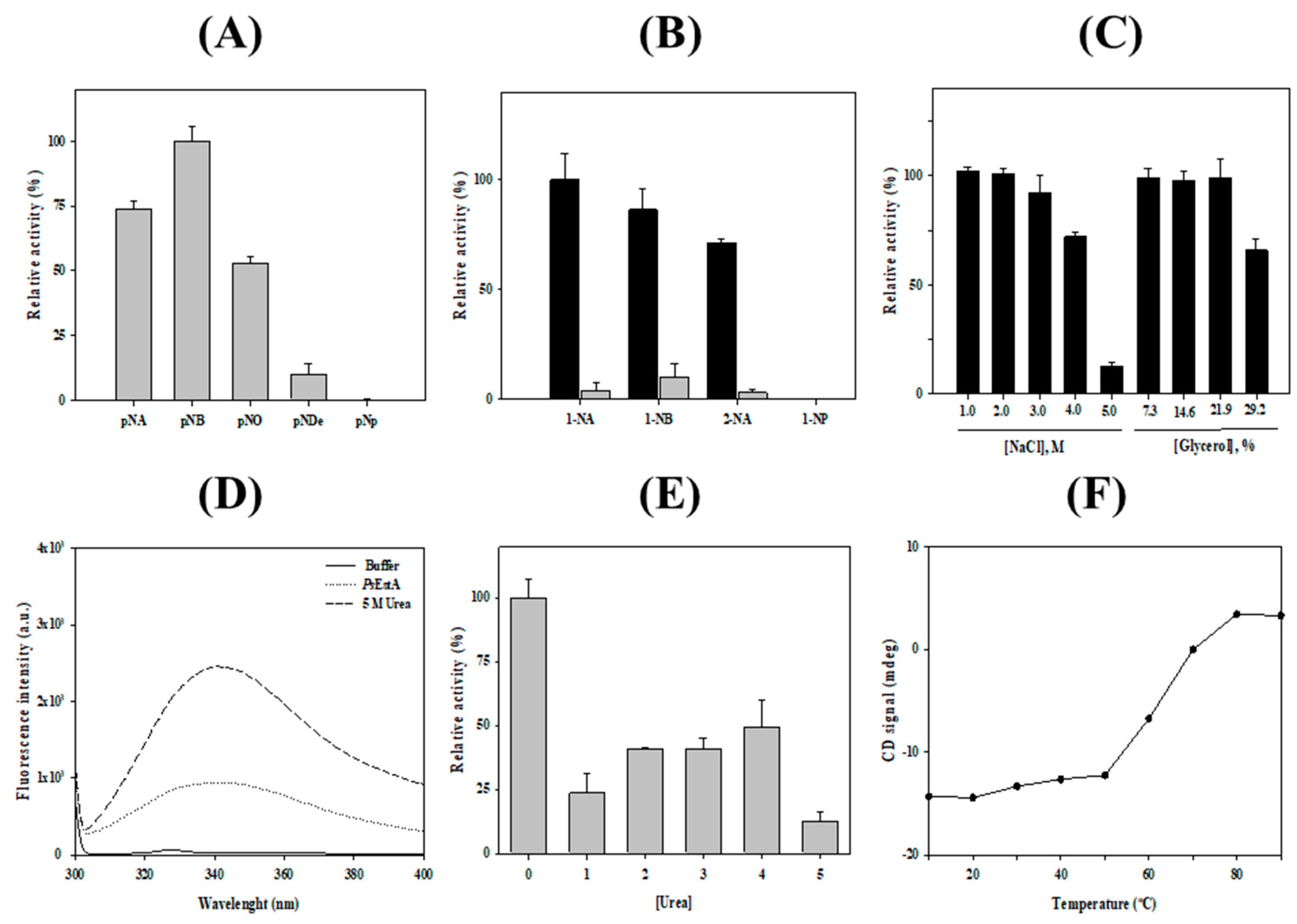
3.2. Characterization of PsEstA
3.3. Biochemical Assay
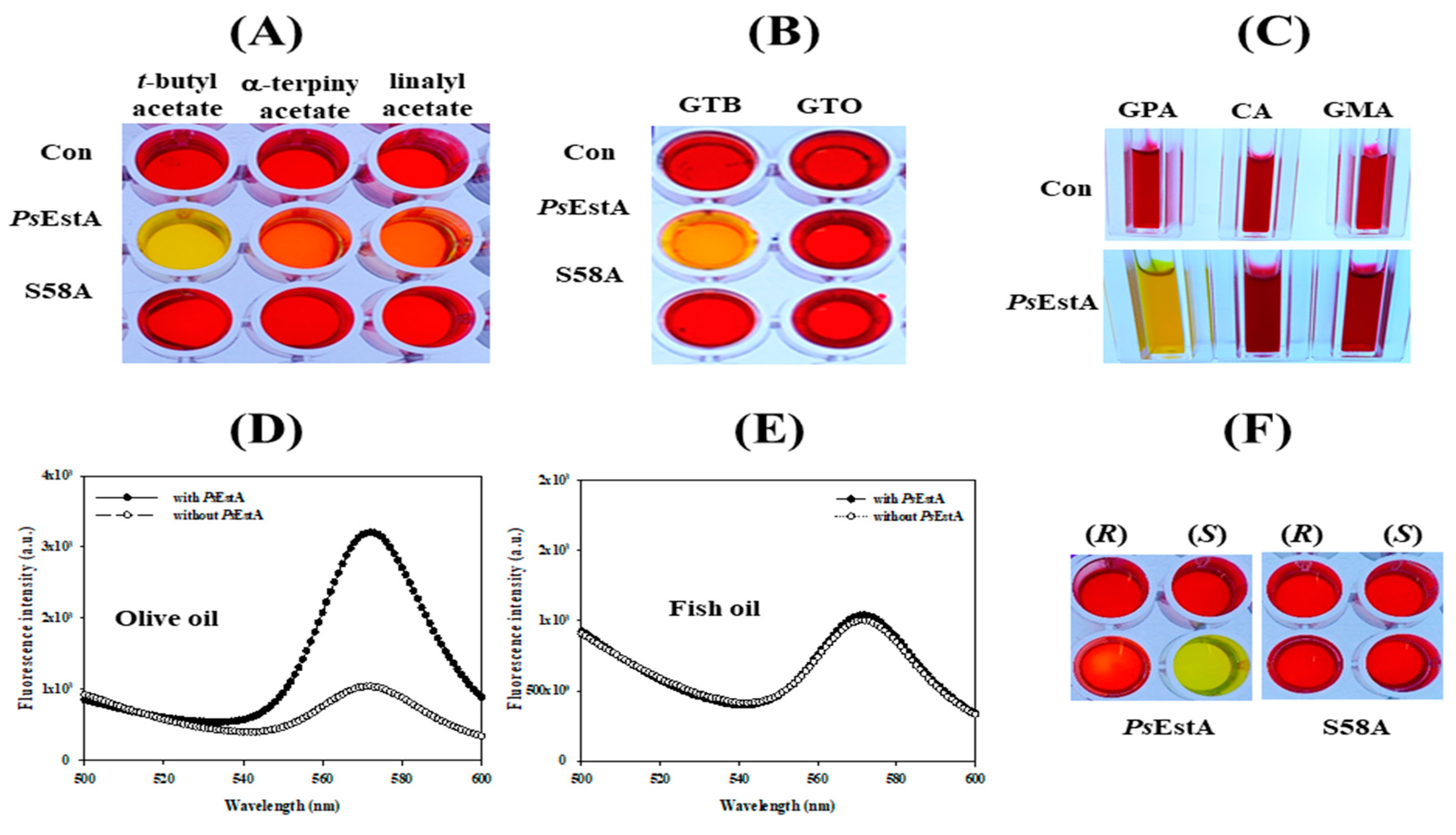
3.4. Substrate Analysis
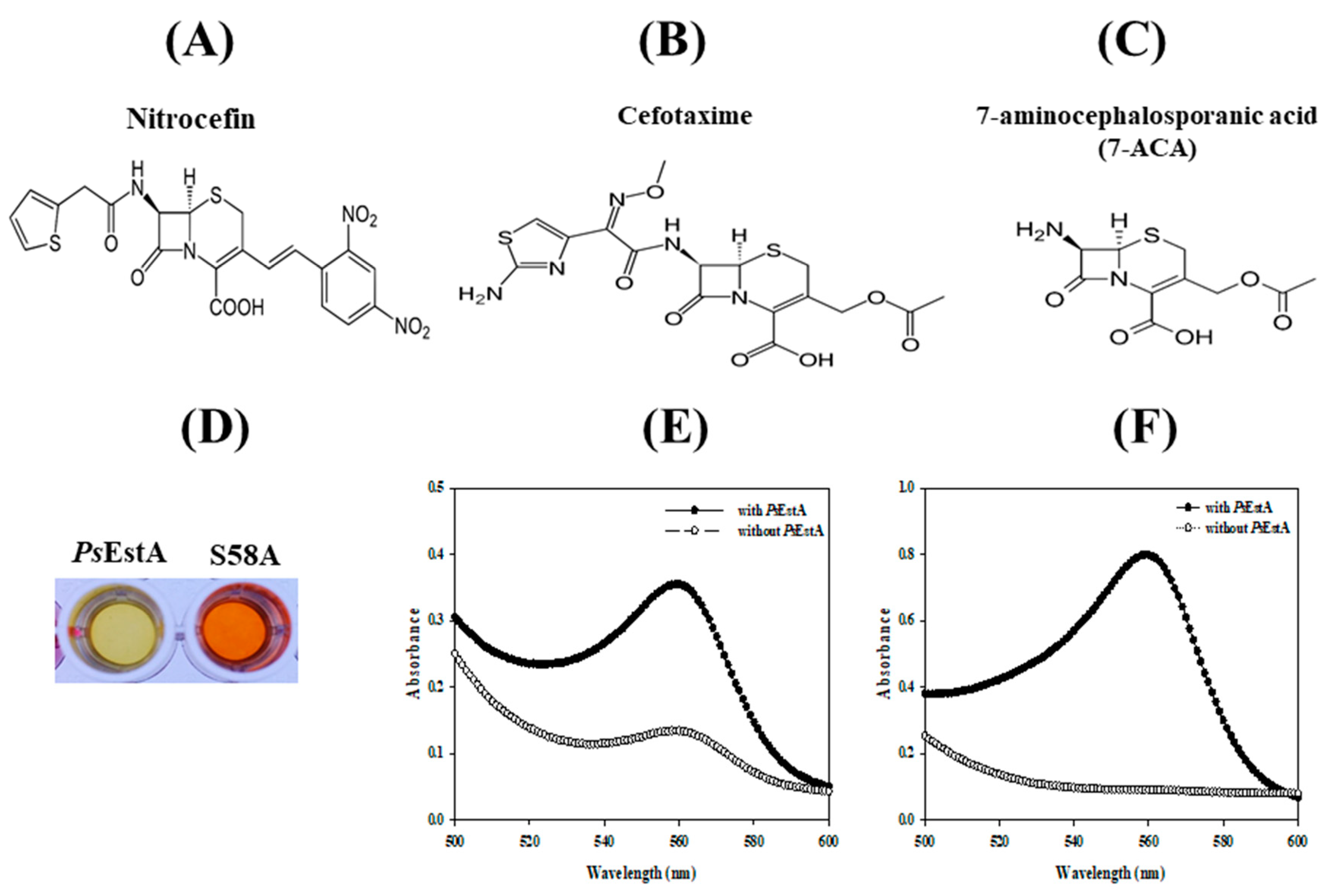
3.5. β-Lactamase Activity of PsEstA
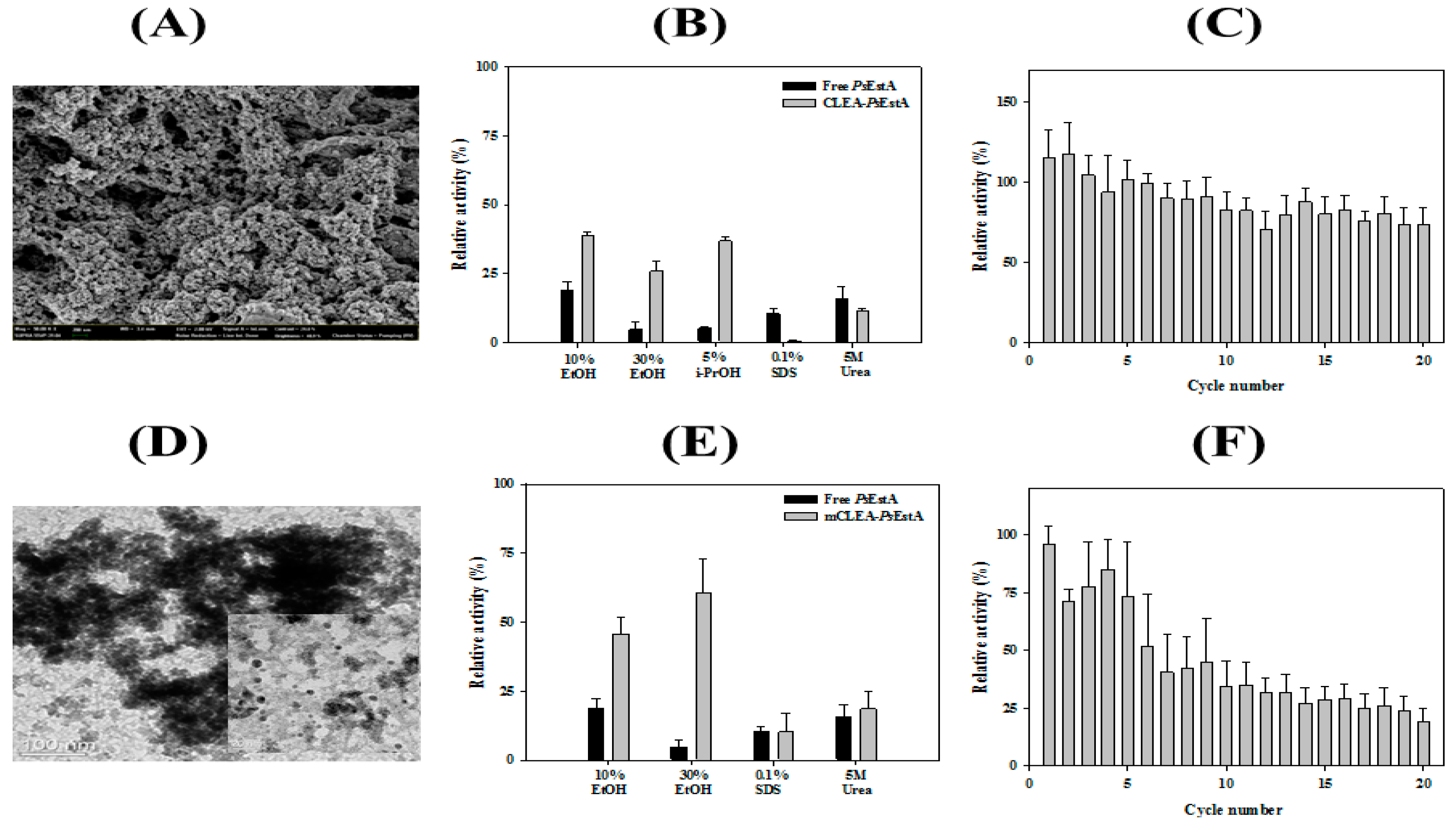
3.6. Immobilization of PsEstA
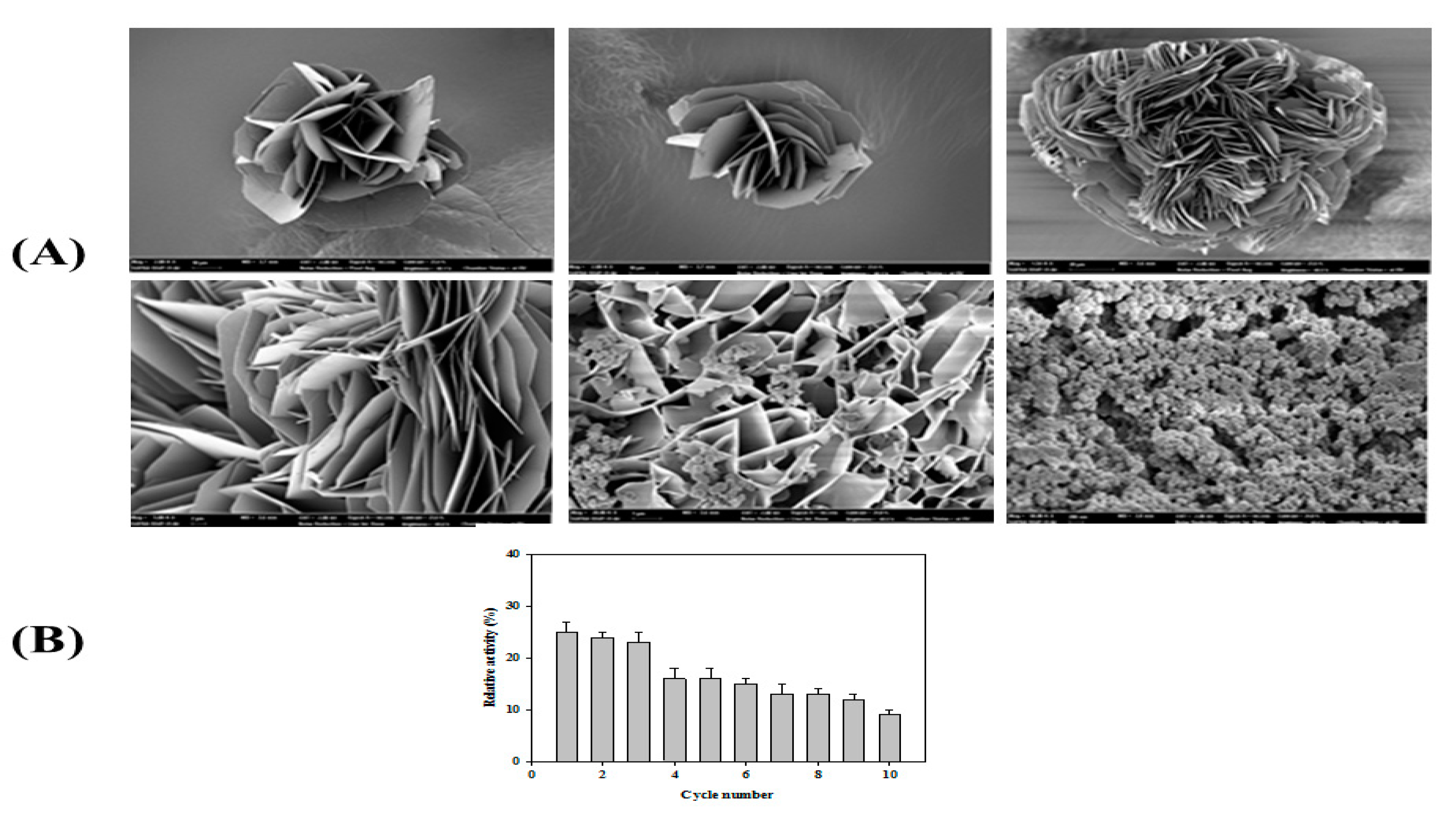
3.7. Formation of Organic–Inorganic Hybrid PsEstA Nanoflower
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Fisher, J.F.; Meroueh, S.O.; Mobashery, S. Bacterial resistance to beta-lactam antibiotics: Compelling opportunism, compelling opportunity. Chem. Rev. 2005, 105, 395–424. [Google Scholar] [CrossRef] [PubMed]
- Salahuddin, P.; Kumar, A.; Khan, A.U. Structure, function of serine and metallo-β-lactamases and their inhibitors. Curr. Protein Pept. Sci. 2018, 19, 130–144. [Google Scholar] [CrossRef] [PubMed]
- Bush, K. Proliferation and significance of clinically relevant β-lactamases. Ann. N. Y. Acad. Sci. 2013, 1277, 84–90. [Google Scholar] [CrossRef] [PubMed]
- Bush, K.; Jacoby, G.A. Updated functional classification of beta-lactamases. Antimicrob. Agents Chemother. 2010, 54, 969–976. [Google Scholar] [CrossRef] [PubMed]
- Bush, K. Past and present perspectives on β-lactamases. Antimicrob. Agents Chemother. 2018, 62, e01076-18. [Google Scholar] [CrossRef] [PubMed]
- Powers, R.A. Structural and functional aspects of extended spectrum AmpC cephalosporinases. Curr. Drug Targets. 2016, 17, 1051–1060. [Google Scholar] [CrossRef]
- Pozzi, C.; Di Pisa, F.; De Luca, F.; Benvenuti, M.; Docquier, J.D.; Mangani, S. Atomic-resolution structure of a class C β-lactamase and its complex with Avibactam. ChemMedChem 2018, 13, 1437–1446. [Google Scholar] [CrossRef]
- Tooke, C.L.; Hinchliffe, P.; Bragginton, E.C.; Colenso, C.K.; Hirvonen, V.H.A.; Takebayashi, Y.; Spencer, J. β-Lactamases and β-Lactamase Inhibitors in the 21st Century. J. Mol. Biol. 2019, 431, 3472–3500. [Google Scholar] [CrossRef]
- Na, J.H.; Cha, S.S. Structural basis for the extended substrate spectrum of AmpC BER and structure-guided discovery of the inhibition activity of citrate against the class C β-lactamases AmpC BER and CMY-10. Acta Crystallogr. D Struct. Biol. 2016, 72, 976–985. [Google Scholar] [CrossRef]
- Oguri, T.; Ishii, Y.; Shimizu-Ibuka, A. Conformational change observed in the active site of class C β-lactamase MOX-1 upon binding to aztreonam. Antimicrob. Agents Chemother. 2015, 59, 5069–5072. [Google Scholar] [CrossRef]
- Bhattacharya, M.; Toth, M.; Antunes, N.T.; Smith, C.A.; Vakulenko, S.B. Structure of the extended-spectrum class C β-lactamase ADC-1 from Acinetobacter baumannii. Acta Crystallogr. D Biol. Crystallogr. 2014, 70, 760–771. [Google Scholar] [CrossRef] [PubMed]
- Rashamuse, K.; Magomani, V.; Ronneburg, T.; Brady, D. A novel family VIII carboxylesterase derived from a leachate metagenome library exhibits promiscuous beta-lactamase activity on nitrocefin. Appl. Microbiol. Biotechnol. 2009, 83, 491–500. [Google Scholar] [CrossRef] [PubMed]
- Mokoena, N.; Mathiba, K.; Tsekoa, T.; Steenkamp, P.; Rashamuse, K. Functional characterisation of a metagenome derived family VIII esterase with a deacetylation activity on β-lactam antibiotics. Biochem. Biophys. Res. Commun. 2013, 437, 342–348. [Google Scholar] [CrossRef] [PubMed]
- Cha, S.S.; An, Y.J. Crystal structure of EstSRT1, a family VIII carboxylesterase displaying hydrolytic activity toward oxyimino cephalosporins. Biochem. Biophys. Res. Commun. 2016, 478, 818–824. [Google Scholar] [PubMed]
- Lee, H.W.; Jung, W.K.; Kim, Y.H.; Ryu, B.H.; Kim, T.D.; Kim, J.; Kim, H. Characterization of a novel alkaline family VIII esterase with S-enantiomer preference from a compost metagenomic library. J. Microbiol. Biotechnol. 2016, 26, 315–325. [Google Scholar] [CrossRef] [PubMed]
- Ngo, T.D.; Ryu, B.H.; Ju, H.; Jang, E.J.; Kim, K.K.; Kim, T.D. Crystallographic analysis and biochemical applications of a novel penicillin-binding protein/β-lactamase homologue from a metagenomic library. Acta Crystallogr. D Biol. Crystallogr. 2014, 70, 2455–2466. [Google Scholar] [CrossRef]
- Ryu, B.H.; Ngo, T.D.; Yoo, W.; Lee, S.; Kim, B.Y.; Lee, E.; Kim, K.K.; Kim, T.D. Biochemical and structural analysis of a novel esterase from Caulobacter crescentus related to penicillin-binding protein (PBP). Sci. Rep. 2016, 6, 37978. [Google Scholar] [CrossRef]
- Kim, Y.O.; Park, I.S.; Nam, B.H.; Kim, D.G.; Jee, Y.J.; Lee, S.J.; An, C.M. A novel esterase from Paenibacillus sp. PBS-2 is a new member of the β-lactamase belonging to the family VIII lipases/esterases. J. Microbiol. Biotechnol. 2014, 24, 1260–1268. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA 7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. 2018, 27, 135–145. [Google Scholar] [CrossRef]
- Gouet, P.; Robert, X.; Courcelle, E. ESPript/ENDscript: Extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 2003, 31, 3320–3323. [Google Scholar] [CrossRef] [PubMed]
- Bianco, G.; Forli, S.; Goodsell, D.S.; Olson, A.J. Covalent docking using autodock: Two-point attractor and flexible side chain methods. Protein Sci. 2016, 25, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Schüttelkopf, A.W.; Van Aalten, D.M. PRODRG: A tool for high-throughput crystallography of protein–ligand complexes. Acta Crystallogr. D Struct. Biol. 2004, 60, 1355–1363. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Seeliger, D.; de Groot, B.L. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Aided Mol. Des. 2010, 24, 417–422. [Google Scholar] [CrossRef]
- Wang, Y.; Ryu, B.H.; Yoo, W.; Lee, C.W.; Kim, K.K.; Lee, J.H.; Kim, T.D. Identification, characterization, immobilization, and mutational analysis of a novel acetylesterase with industrial potential (LaAcE) from Lactobacillus acidophilus. Biochim. Biophys. Acta Gen. Subj. 2018, 1862, 197–210. [Google Scholar] [CrossRef]
- Lee, C.W.; Kwon, S.; Park, S.H.; Kim, B.Y.; Yoo, W.; Ryu, B.H.; Kim, H.W.; Shin, S.C.; Kim, S.; Park, H.; et al. Crystal structure and functional characterization of an esterase (EaEST) from Exiguobacterium antarcticum. PLoS ONE 2017, 12, e0169540. [Google Scholar] [CrossRef]
- Oh, C.; Ryu, B.H.; Yoo, W.; Nguyen, D.D.; Kim, T.; Ha, S.C.; Kim, T.D.; Kim, K.K. Identification and Crystallization of Penicillin-Binding Protein/β-Lactamase Homolog (Rp46) from Ruegeria Pomeroyi. Crystals 2017, 7, 6. [Google Scholar]
- Zottig, X.; Meddeb-Mouelhi, F.; Beauregard, M. Development of a high-throughput liquid state assay for lipase activity using natural substrates and rhodamine B. Anal. Biochem. 2016, 496, 25–29. [Google Scholar] [CrossRef]
- Cui, J.; Cui, L.; Jia, S.; Su, Z.; Zhang, S. Hybrid cross-linked lipase aggregates with magnetic nanoparticles: A robust and recyclable biocatalysis for the epoxidation of oleic acid. J. Agric. Food Chem. 2016, 64, 7179–7187. [Google Scholar] [CrossRef] [PubMed]
- Yoo, W.; Le, L.T.H.L.; Lee, J.H.; Kim, K.K.; Kim, T.D. A novel enantioselective SGNH family esterase (NmSGNH1) from Neisseria meningitides: Characterization, mutational analysis, and ester synthesis. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids. 2019, 1864, 1438–1448. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.; Kim, S.; Ryu, Y.; Kim, T.D. Identification and characterization of a novel (S)-ketoprofen-specific esterase. Int. J. Biol. Macromol. 2007, 41, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Wagner, U.G.; Petersen, E.I.; Schwab, H.; Kratky, C. EstB from Burkholderia gladioli: A novel esterase with a beta-lactamase fold reveals steric factors to discriminate between esterolytic and beta-lactam cleaving activity. Protein Sci. 2002, 11, 467–478. [Google Scholar] [CrossRef]
- Cha, S.S.; An, Y.J.; Jeong, C.S.; Kim, M.K.; Jeon, J.H.; Lee, C.M.; Lee, H.S.; Kang, S.G.; Lee, J.H. Structural basis for the β-lactamase activity of EstU1, a family VIII carboxylesterase. Proteins 2013, 81, 2045–2051. [Google Scholar] [CrossRef]
- Wouters, J.; Fonze, E.; Vermeire, M.; Frere, J.M.; Charlier, P. Crystal structure of Enterobacter cloacae 908R class C beta-lactamase bound to iodoacetamido phenyl boronic acid, a transition-state analogue. Cell. Mol. Life Sci. 2003, 60, 1764–1773. [Google Scholar] [CrossRef]
- Oguri, T.; Furuyama, T.; Okuno, T.; Ishii, Y.; Tateda, K.; Bonomo, R.A.; Shimizu-Ibuka, A. Crystal structure of Mox-1, a unique plasmid-mediated class C β-lactamase with hydrolytic activity towards moxalactam. Antimicrob. Agents. Chemother. 2014, 58, 3914–3920. [Google Scholar] [CrossRef]
- Schutte, M.; Fetzner, S. EstA from Arthrobacter nitroguajacolicus Rü61a, a thermo- and solvent-tolerant carboxylesterase related to class C beta-lactamases. Curr. Microbiol. 2007, 54, 230–236. [Google Scholar] [CrossRef]
- Kim, S.; Joo, S.; Yoon, S.; Kim, S.; Moon, J.; Ryu, Y.; Kim, K.K.; Kim, T.D. Purification, crystallization and preliminary crystallographic analysis of Est-Y29: A novel oligomeric beta-lactamase. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2009, 65, 310–312. [Google Scholar] [CrossRef]
- Rashamuse, K.J.; Burton, S.G.; Stafford, W.H.; Cowan, D.A. Molecular characterization of a novel family VIII esterase from Burkholderia multivorans UWC10. J. Mol. Microbiol. Biotechnol. 2007, 13, 181–188. [Google Scholar] [CrossRef]
- Oh, C.; Ryu, B.H.; Yoo, W.; Nguyen, D.D.; Kim, T.; Ha, S.-C.; Kim, T.D.; Kim, K.K. Identification and Crystallographic Analysis of a New Carbohydrate Acetylesterase (SmAcE1) from Sinorhizobium meliloti. Crystals 2018, 8, 12. [Google Scholar] [CrossRef]
- Jeon, J.H.; Kim, S.J.; Lee, H.S.; Cha, S.S.; Lee, J.H.; Yoon, S.H.; Koo, B.S.; Lee, C.M.; Choi, S.H.; Lee, S.H.; et al. Novel metagenome-derived carboxylesterase that hydrolyzes β-lactam antibiotics. Appl. Environ. Microbiol. 2011, 77, 7830–7836. [Google Scholar] [CrossRef] [PubMed]
- Yu, E.Y.; Kwon, M.A.; Lee, M.; Oh, J.Y.; Choi, J.E.; Lee, J.Y.; Song, B.K.; Hahm, D.H.; Song, J.K. Isolation and characterization of cold-active family VIII esterases from an arctic soil metagenome. Appl. Microbiol. Biotechnol. 2011, 90, 573–581. [Google Scholar] [CrossRef] [PubMed]
- Sirisha, V.L.; Jain, A.; Jain, A. Enzyme immobilization: An overview on methods, support material, and applications of immobilized enzymes. Adv. Food. Nutr. Res. 2016, 79, 179–211. [Google Scholar] [PubMed]
- Di Cosimo, R.; McAuliffe, J.; Poulose, A.J.; Bohlmann, G. Industrial use of immobilized enzymes. Chem. Soc. Rev. 2013, 42, 6437–6474. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, R.A. Characteristic features and biotechnological applications of cross-linked enzyme aggregates (CLEAs). Appl. Microbiol. Biotechnol. 2011, 92, 467–477. [Google Scholar] [CrossRef]
- Altinkaynak, C.; Tavlasoglu, S.; Özdemir, N.; Ocsoy, I. A new generation approach in enzyme immobilization: Organic-inorganic hybrid nanoflowers with enhanced catalytic activity and stability. Enzyme Microb. Technol. 2016, 93, 105–112. [Google Scholar] [CrossRef]
- Lee, S.W.; Cheon, S.A.; Kim, M.I.; Park, T.J. Organic-inorganic hybrid nanoflowers: Types, characteristics, and future prospects. J. Nanobiotechnology 2015, 13, 54. [Google Scholar] [CrossRef]
- Wagner, U.G.; DiMaio, F.; Kolkenbrock, S.; Fetzner, S. Crystal structure analysis of EstA from Arthrobacter sp. Rue61a—An insight into catalytic promiscuity. FEBS Lett. 2014, 588, 1154–1160. [Google Scholar] [CrossRef]
- Littlechild, J.A. Enzymes from extreme environments and their industrial applications. Front. Bioeng. Biotechnol. 2015, 3, 161. [Google Scholar] [CrossRef]
- Mitusińska, K.; Magdziarz, T.; Bzówka, M.; Stańczak, A.; Góra, A. Exploring Solanum tuberosum Epoxide Hydrolase Internal Architecture by Water Molecules Tracking. Biomolecules 2018, 8, 143. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kwon, S.; Yoo, W.; Kim, Y.-O.; Kim, K.K.; Kim, T.D. Molecular Characterization of a Novel Family VIII Esterase with β-Lactamase Activity (PsEstA) from Paenibacillus sp. Biomolecules 2019, 9, 786. https://doi.org/10.3390/biom9120786
Kwon S, Yoo W, Kim Y-O, Kim KK, Kim TD. Molecular Characterization of a Novel Family VIII Esterase with β-Lactamase Activity (PsEstA) from Paenibacillus sp. Biomolecules. 2019; 9(12):786. https://doi.org/10.3390/biom9120786
Chicago/Turabian StyleKwon, Sena, Wanki Yoo, Young-Ok Kim, Kyeong Kyu Kim, and T. Doohun Kim. 2019. "Molecular Characterization of a Novel Family VIII Esterase with β-Lactamase Activity (PsEstA) from Paenibacillus sp." Biomolecules 9, no. 12: 786. https://doi.org/10.3390/biom9120786
APA StyleKwon, S., Yoo, W., Kim, Y.-O., Kim, K. K., & Kim, T. D. (2019). Molecular Characterization of a Novel Family VIII Esterase with β-Lactamase Activity (PsEstA) from Paenibacillus sp. Biomolecules, 9(12), 786. https://doi.org/10.3390/biom9120786






