m1A Post‐Transcriptional Modification in tRNAs
Abstract
:1. Introduction
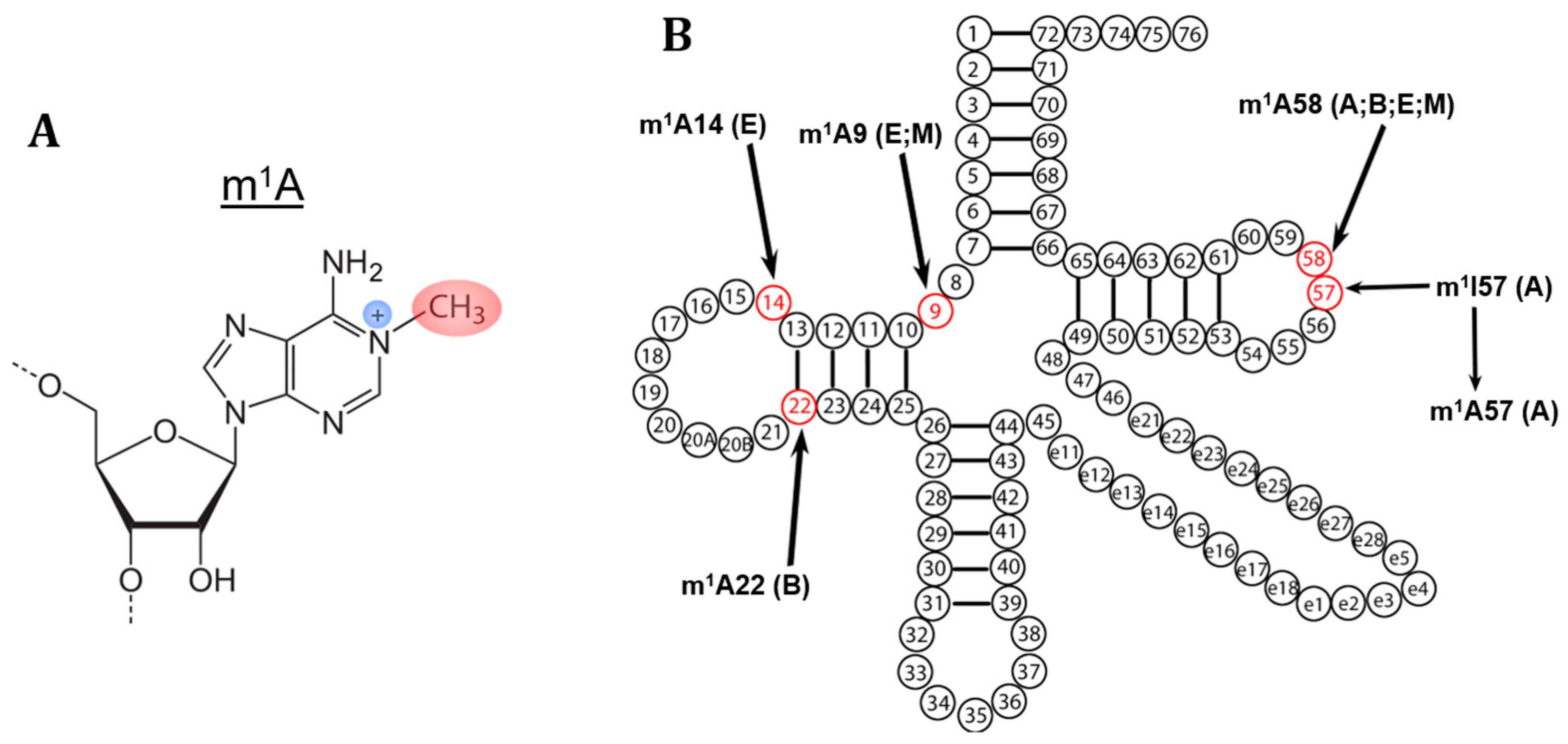
2. m1A Modifications in tRNA
3. The Biological Role of m1A Modifications in tRNA
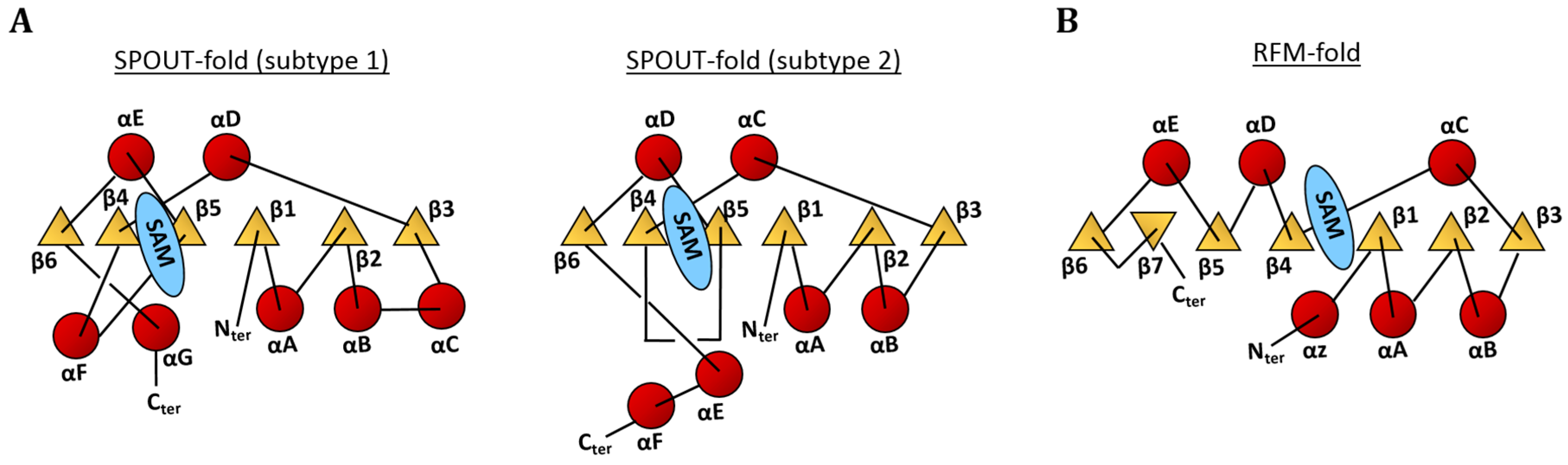
4. Enzymes Responsible for the m1A Modifications in tRNA
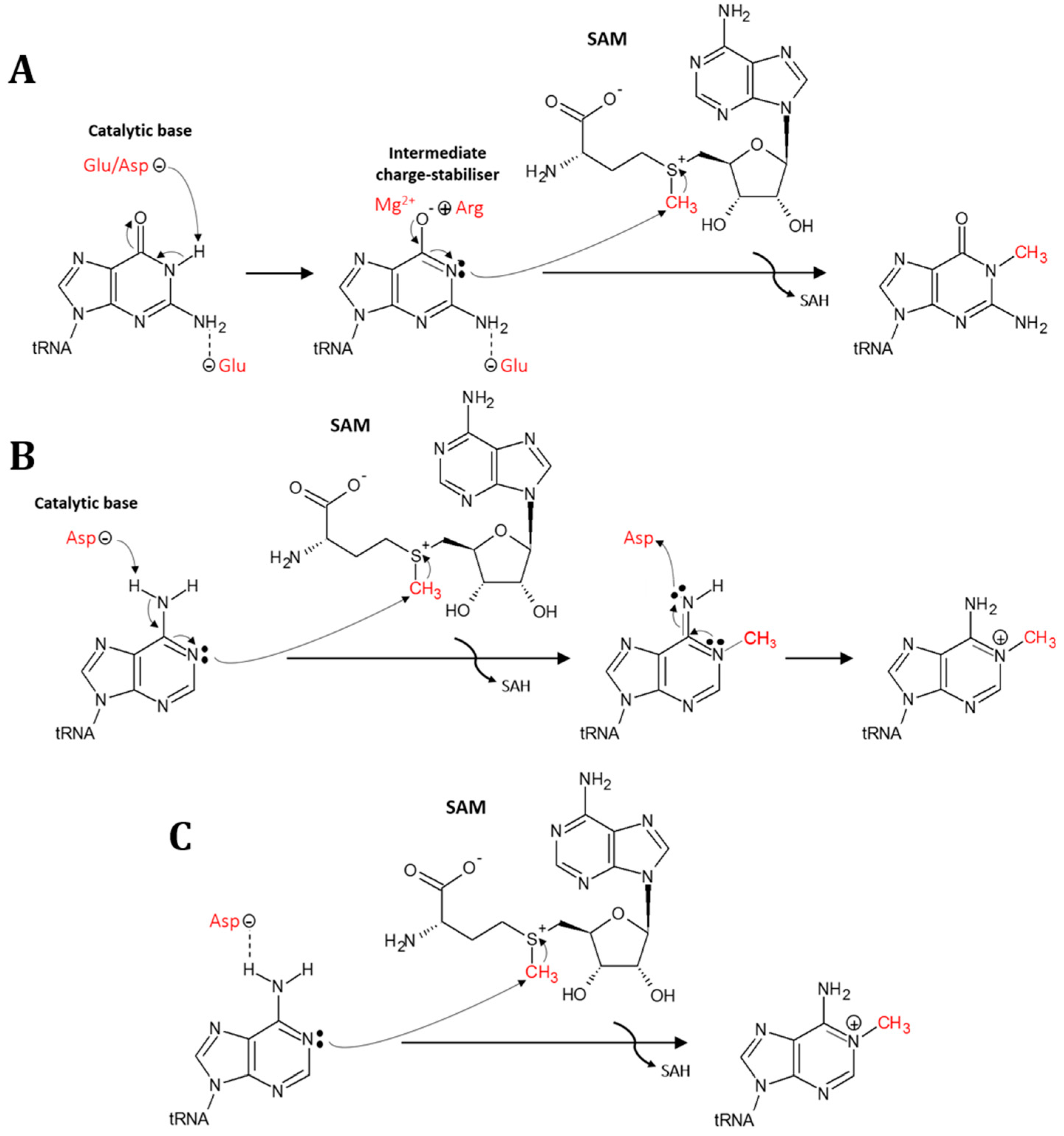
5. Mechanism for Formation of the N1-Methylation in tRNA
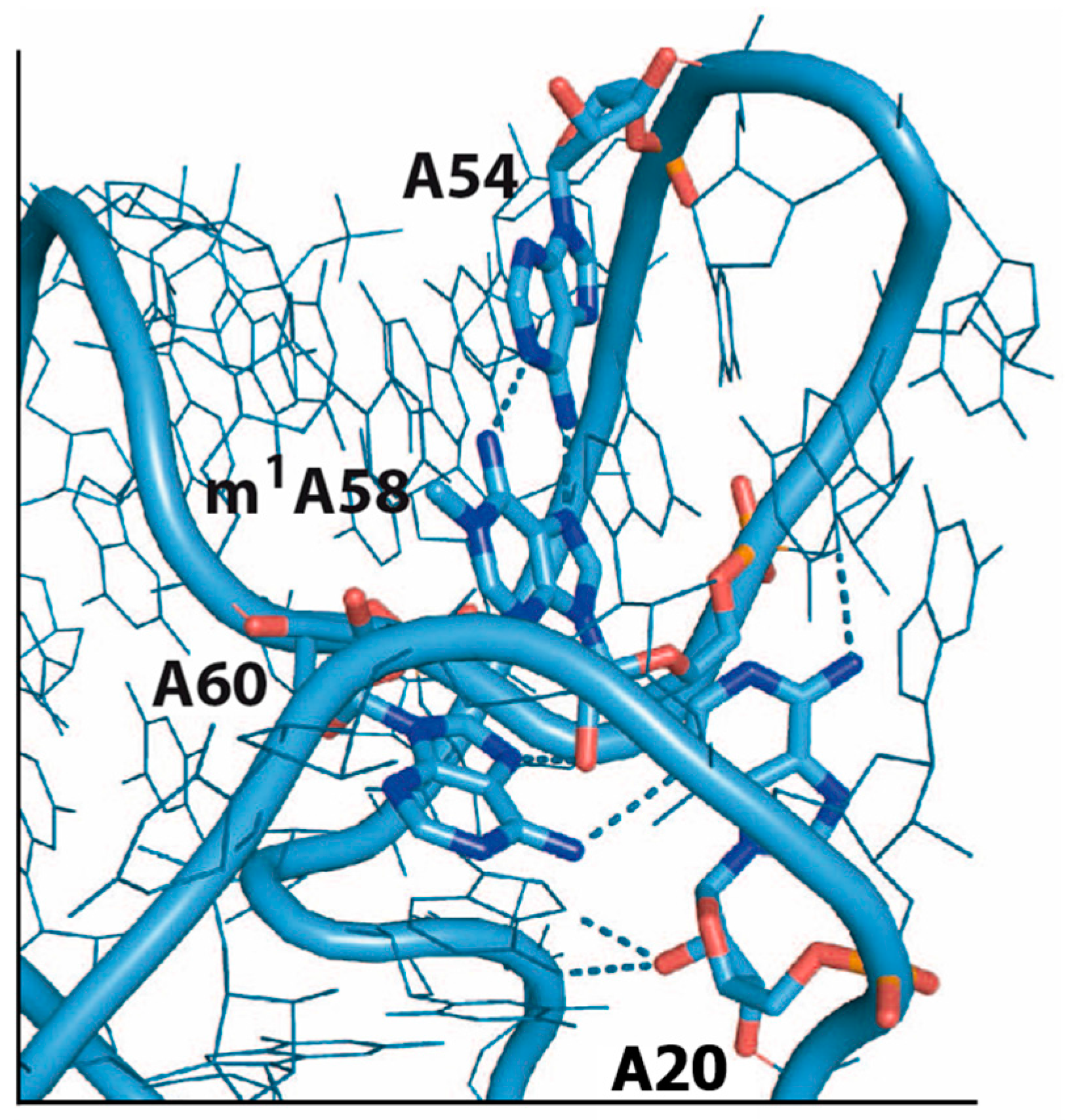
6. m1A58
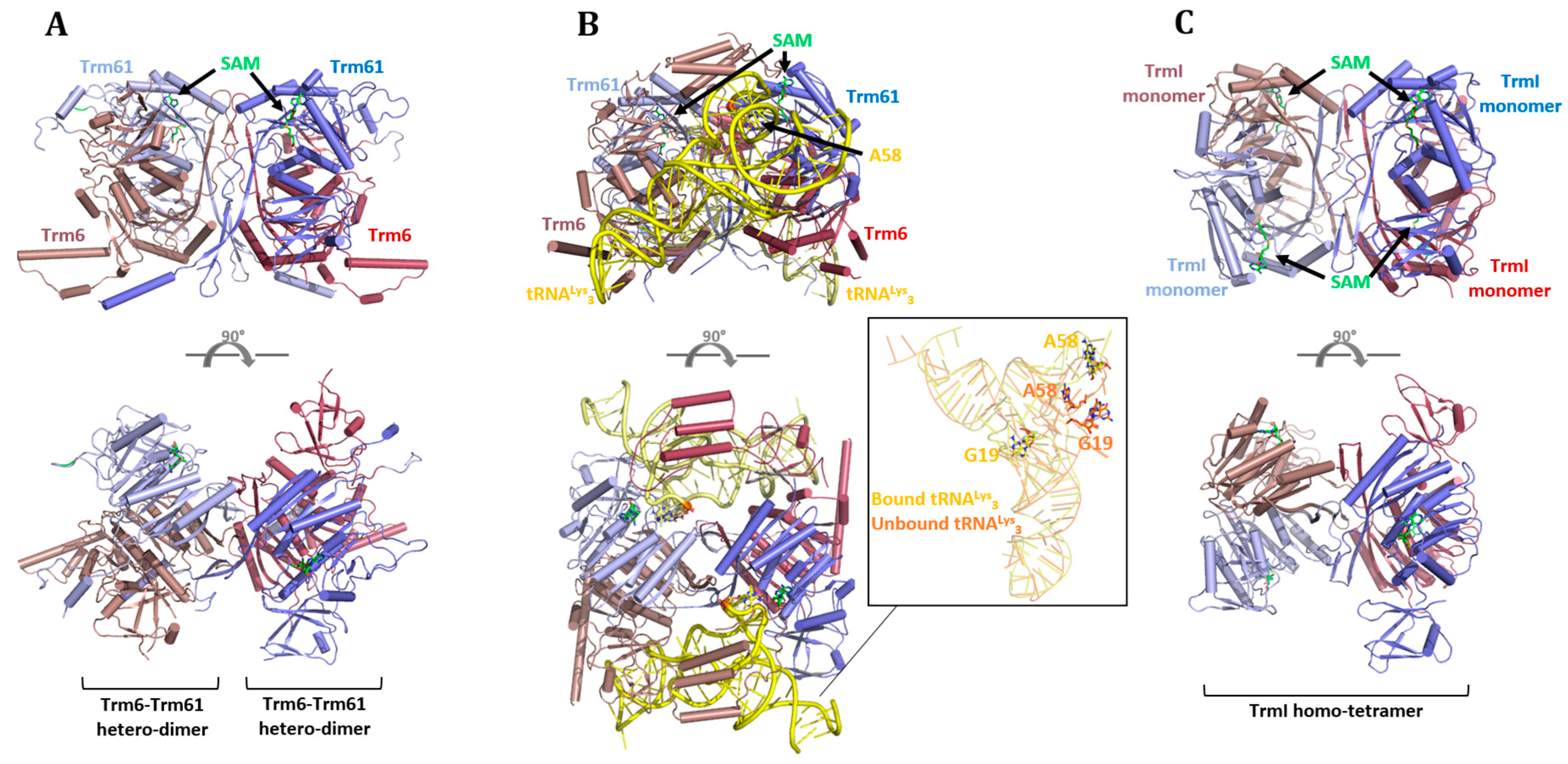
6.1. Eukaryotes
6.2. Archaea and Bacteria
6.3. Mitochondria
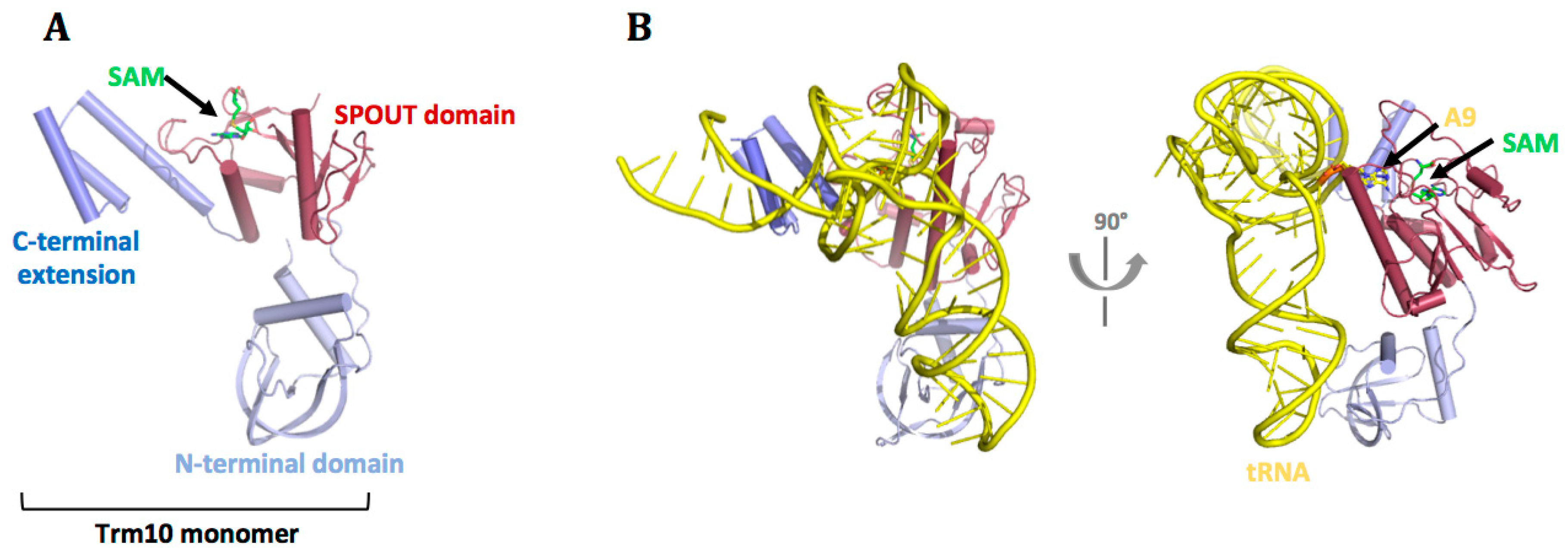
7. m1A9
7.1. Eukaryotes
7.2. Archaea
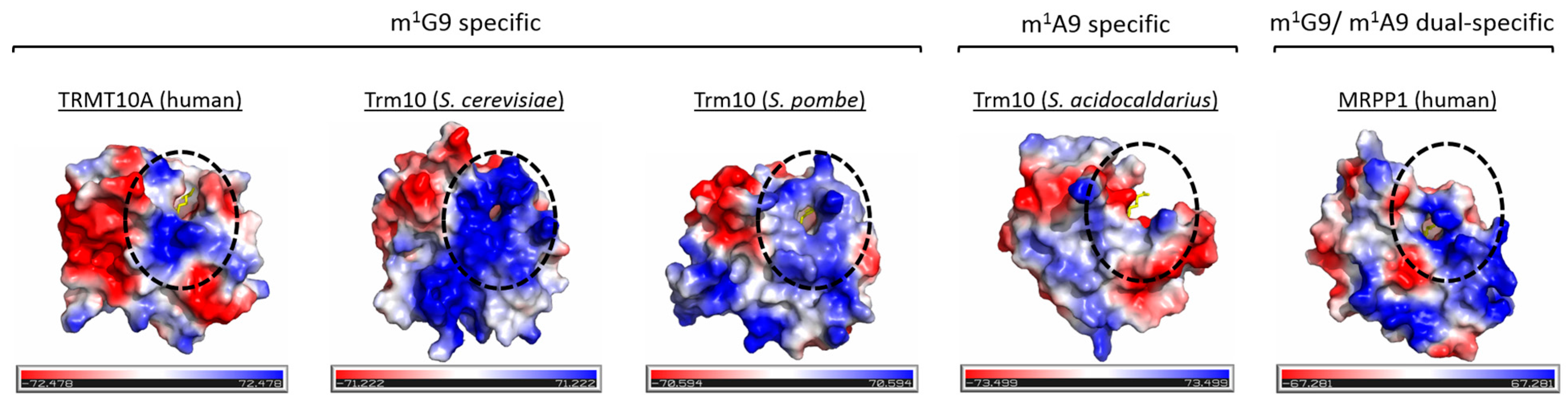
7.3. tRNA Recognition by Trm10 Proteins
8. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Phizicky, E.M.; Hopper, A.K. tRNA biology charges to the front. Genes Dev. 2010, 24, 1832–1860. [Google Scholar] [CrossRef] [PubMed]
- Hopper, A.K.; Phizicky, E.M. tRNA transfers to the limelight. Genes Dev. 2003, 17, 162–180. [Google Scholar] [CrossRef] [PubMed]
- Motorin, Y.; Helm, M. tRNA stabilization by modified nucleotides. Biochemistry 2010, 49, 4934–4944. [Google Scholar] [CrossRef]
- Li, H. Complexes of tRNA and maturation enzymes: Shaping up for translation. Curr. Opin. Struct. Biol. 2007, 17, 293–301. [Google Scholar] [CrossRef]
- Hopper, A.K. Transfer RNA post-transcriptional processing, turnover, and subcellular dynamics in the yeast Saccharomyces cerevisiae. Genetics 2013, 194, 43–67. [Google Scholar] [CrossRef] [PubMed]
- Machnicka, M.A.; Olchowik, A.; Grosjean, H.; Bujnicki, J.M. Distribution and frequencies of post- transcriptional modifications in tRNAs. RNA Biol. 2014, 11, 1619–1629. [Google Scholar] [CrossRef] [PubMed]
- tRNAmodviz. Available online: http://genesilico.pl/trnamodviz (accessed on 9 February 2017).
- Helm, M.; Brulé, H.; Degoul, F.; Cepanec, C.; Leroux, J.P.; Giegé, R.; Florentz, C. The presence of modified nucleotides is required for cloverleaf folding of a human mitochondrial tRNA. Nucleic Acids Res. 1998, 26, 1636–1643. [Google Scholar] [CrossRef] [PubMed]
- Voigts-Hoffmann, F.; Hengesbach, M.; Kobitski, A.Y.; Van Aerschot, A.; Herdewijn, P.; Ulrich Nienhaus, G.; Helm, M. A methyl group controls conformational equilibrium in human mitochondrial tRNALys. J. Am. Chem. Soc. 2007, 129, 13382–13383. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K.; Shinma, M.; Oshima, T.; Nishimura, S. Heat-induced stability of tRNA from an extreme thermophile, Thermus thermophilus. Biochem. Biophys. Res. Commun. 1976, 72, 1137–1144. [Google Scholar] [CrossRef]
- Motorin, Y.; Helm, M. RNA nucleotide methylation. Wiley Interdiscip. Rev. RNA 2011, 2, 611–631. [Google Scholar] [CrossRef] [PubMed]
- Hori, H. Methylated nucleosides in tRNA and tRNA methyltransferases. Front. Genet. 2014, 5, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Czerwoniec, A.; Kasprzak, J.M.; Kaminska, K.H.; Rother, K.; Purta, E.; Bujnicki, J.M. Folds and Functions of Domains in RNA Modification Enzymes. In DNA and RNA Modification Enzymes: Structure, Mechanism, Function and Evolution; Grosjean, H., Ed.; Landes Bioscience: Austin, TX, USA, 2009; pp. 289–302. [Google Scholar]
- Schubert, H.L.; Blumenthal, R.M.; Cheng, X. Many paths to methyltransfer: A chronicle of convergence. Trends Biochem. Sci. 2003, 28, 329–335. [Google Scholar] [CrossRef]
- Anantharaman, V.; Koonin, E.V.; Aravind, L. SPOUT: A Class of Methyltransferases that Includes spoU and trmD RNA Methylase Superfamilies, and Novel Superfamilies of Predicted Prokaryotic RNA Methylases. J. Mol. Microbiol. Biotechnol. 2002, 4, 71–75. [Google Scholar] [PubMed]
- Tkaczuk, K.; Dunin-Horkawicz, S.; Purta, E.; Bujnicki, J.M. Structural and evolutionary bioinformatics of the SPOUT superfamily of methyltransferases. BMC Bioinform. 2007, 8, 73. [Google Scholar] [CrossRef] [PubMed]
- Roovers, M.; Kaminska, K.H.; Tkaczuk, K.L.; Gigot, D.; Droogmans, L.; Bujnicki, J.M. The YqfN protein of Bacillus subtilis is the tRNA: m1A22 methyltransferase (TrmK). Nucleic Acids Res. 2008, 36, 3252–3262. [Google Scholar] [CrossRef] [PubMed]
- Sprinzl, M.; Horn, C.; Brown, M.; Ioudovitch, A.; Steinberg, S. Compilation of tRNA sequences and sequences of tRNA genes. Nucleic Acids Res. 1998, 26, 148–153. [Google Scholar]
- Menichi, B.; Arnold, H.H.; Heyman, T.; Dirheimer, G.; Keith, G. Primary Structure of Bacillus subtilis tRNAsTyr. Biochem. Biophys. Res. Commun. 1980, 95, 461–467. [Google Scholar] [CrossRef]
- Matsugi, J.; Jia, H.T.; Murao, K.; Ishikura, H. Nucleotide sequences of serine tRNAs from Bacillus subtilis. Biochem. Biophys. Acta. 1992, 1130, 333–335. [Google Scholar] [CrossRef]
- Andachi, Y.; Yamao, F.; Muto, A.; Osawa, S. Codon recognition patterns as deduced from sequences of the complete set of transfer RNA species in Mycoplasma capricolum. Resemblance to mitochondria. J. Mol. Biol. 1989, 209, 37–54. [Google Scholar] [CrossRef]
- Raettig, R.; Kersten, H.; Weissenbach, J.; Dirheimer, G. Methylation of an adenosine in the D-loop of specific transfer RNAs from yeast by a procaryotic tRNA (adenine-1) methyltransferase. Nucleic Acids Res. 1977, 4, 1769–1782. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Grosjean, H.; Constantinesco, F.; Foiret, D.; Benachenhou, N. A novel enzymatic pathway leading to 1-methylinosine modification in Haloferax volcanii tRNA. Nucleic Acids Res. 1995, 23, 4312–4319. [Google Scholar] [CrossRef]
- Grosjean, H.; Auxilien, S.; Constantinesco, F.; Simon, C.; Corda, Y.; Becker, H.F.; Foiret, D.; Morin, A.; Jin, Y.X.; Fournier, M.; et al. Enzymatic conversion of adenosine to inosine and to N1-methylinosine in transfer RNAs: A review. Biochimie 1996, 78, 488–501. [Google Scholar] [CrossRef]
- Helm, M.; Attardi, G. Nuclear Control of Cloverleaf Structure of Human Mitochondrial tRNALys. J. Mol. Biol. 2004, 337, 545–560. [Google Scholar] [CrossRef]
- Jühling, F.; Mörl, M.; Hartmann, R.K.; Sprinzl, M.; Stadler, P.F.; Pütz, J. tRNAdb 2009: Compilation of tRNA sequences and tRNA genes. Nucleic Acids Res. 2009, 37, D159–D162. [Google Scholar] [CrossRef] [PubMed]
- Kobitski, A.Y.; Hengesbach, M.; Helm, M.; Nienhaus, G.U. Sculpting an RNA conformational energy landscape by a methyl group modification—a single-molecule FRET study. Angew. Chem. Int. Ed. 2008, 47, 4326–4330. [Google Scholar] [CrossRef] [PubMed]
- Helm, M.; Giegé, R.; Florentz, C. A Watson−Crick Base-Pair-Disrupting Methyl Group (m1A9) Is Sufficient for Cloverleaf Folding of Human Mitochondrial tRNALys. Biochemistry 1999, 38, 13338–13346. [Google Scholar] [CrossRef] [PubMed]
- Wittenhagen, L.M.; Kelley, S.O. Impact of disease-related mitochondrial mutations on tRNA structure and function. Trends Biochem. Sci. 2003, 28, 605–611. [Google Scholar] [CrossRef] [PubMed]
- Sohm, B.; Frugier, M.; Brulé, H.; Olszak, K.; Przykorska, A.; Florentz, C. Towards understanding human mitochondrial leucine aminoacylation identity. J. Mol. Biol. 2003, 328, 995–1010. [Google Scholar] [PubMed]
- Sohm, B.; Sissler, M.; Park, H.; King, M.P.; Florentz, C. Recognition of human mitochondrial tRNALeu(UUR) by its cognate leucyl-tRNA synthetase. J. Mol. Biol. 2004, 339, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Sen, A.; Karasik, A.; Shanmuganathan, A.; Mirkovic, E.; Koutmos, M.; Cox, R.T. Loss of the mitochondrial protein-only ribonuclease P complex causes aberrant tRNA processing and lethality in Drosophila. Nucleic Acids Res. 2016, 2, gkw338. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, M.I.; Mercer, T.R.; Davies, S.M.; Shearwood, A.M.; Nygård, K.K.; Richman, T.R.; Mattick, J.S.; Rackham, O.; Filipovska, A. RNA processing in human mitochondria. Cell Cycle 2011, 10, 2904–2916. [Google Scholar] [CrossRef] [PubMed]
- Gillis, D.; Krishnamohan, A.; Yaacov, B.; Shaag, A.; Jackman, J.E.; Elpeleg, O. TRMT10A dysfunction is associated with abnormalities in glucose homeostasis, short stature and microcephaly. J. Med. Genet. 2014, 51, 581–586. [Google Scholar] [CrossRef] [PubMed]
- Igoillo-Esteve, M.; Genin, A.; Lambert, N.; Désir, J.; Pirson, I.; Abdulkarim, B.; Simonis, N.; Drielsma, A.; Marselli, L.; Marchetti, P.; et al. tRNA methyltransferase homolog gene TRMT10A mutation in young onset diabetes and primary microcephaly in humans. PLoS Genet. 2013, 9, e1003888. [Google Scholar] [CrossRef] [PubMed]
- Metodiev, M.D.; Thompson, K.; Alston, C.L.; Morris, A.A.M.; He, L.; Assouline, Z.; Rio, M.; Bahi-Buisson, N.; Pyle, A.; Griffin, H.; et al. Recessive Mutations in TRMT10C Cause Defects in Mitochondrial RNA Processing and Multiple Respiratory Chain Deficiencies. Am. J. Hum. Genet. 2016, 98, 993–1000. [Google Scholar] [CrossRef] [PubMed]
- Vilardo, E.; Nachbagauer, C.; Buzet, A.; Taschner, A.; Holzmann, J.; Rossmanith, W. A subcomplex of human mitochondrial RNase P is a bifunctional methyltransferase—extensive moonlighting in mitochondrial tRNA biogenesis. Nucleic Acids Res. 2012, 40, 11583–11593. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Sun, H.; Liang, X.; Lima, W.F.; Crooke, S.T. Human RNase H1 Is Associated with Protein P32 and Is Involved in Mitochondrial Pre-rRNA Processing. PLoS ONE 2013, 8, e71006. [Google Scholar] [CrossRef] [PubMed]
- Holzmann, J.; Frank, P.; Löffler, E.; Bennett, K.L.; Gerner, C.; Rossmanith, W. RNase P without RNA: Identification and functional reconstitution of the human mitochondrial tRNA processing enzyme. Cell 2008, 135, 462–474. [Google Scholar] [CrossRef] [PubMed]
- Horie, N.; Hara-Yokoyama, M.; Yokoyama, S.; Watanabe, K.; Kuchino, Y.; Nishimura, S.; Miyazawa, T. Two tRNAIle1 species from an extreme thermophile, Thermus thermophilus HB8: effect of 2-thiolation of ribothymidine on the thermostability of tRNA. Biochemistry 1985, 24, 5711–5715. [Google Scholar] [CrossRef] [PubMed]
- Shigi, N.; Sakaguchi, Y.; Suzuki, T.; Watanabe, K. Identification of two tRNA thiolation genes required for cell growth at extremely high temperatures. J. Biol. Chem. 2006, 281, 14296–14306. [Google Scholar] [CrossRef] [PubMed]
- Tomikawa, C.; Yokogawa, T.; Kanai, T.; Hori, H. N7-Methylguanine at position 46 (m7G46) in tRNA from Thermus thermophilus is required for cell viability at high temperatures through a tRNA modification network. Nucleic Acids Res. 2009, 38, 942–957. [Google Scholar] [CrossRef] [PubMed]
- Droogmans, L.; Roovers, M.; Bujnicki, J.M.; Tricot, C.; Hartsch, T.; Stalon, V.; Grosjean, H. Cloning and characterization of tRNA (m1A58) methyltransferase (TrmI) from Thermus thermophilus HB27, a protein required for cell growth at extreme temperatures. Nucleic Acids Res. 2003, 31, 2148–2156. [Google Scholar] [CrossRef] [PubMed]
- Auxilien, S.; Keith, G.; Le Grice, S.F.J.; Darlix, J.L. Role of Post-transcriptional Modifications of Primer tRNALys,3 in the Fidelity and Efficacy of Plus Strand DNA Transfer during HIV-1 Reverse Transcription. J. Biol. Chem. 1999, 274, 4412–4420. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.; Phan, L.; Cuesta, R.; Carlson, B.A.; Pak, M.; Asano, K.; Bjo, G.R.; Tamame, M.; Hinnebusch, A.G. The essential Gcd10p–Gcd14p nuclear complex is required for 1-methyladenosine modification and maturation of initiator methionyl-tRNA. Genes Dev. 1998, 3, 3650–3662. [Google Scholar] [CrossRef]
- Basavappa, R.; Sigler, P.B. The 3 Å crystal structure of yeast initiator tRNA: Functional implications in initiator/elongator discrimination. EMBO J. 1991, 10, 3105–3111. [Google Scholar] [PubMed]
- Anderson, J.; Phan, L.; Hinnebusch, A.G. The Gcd10p/Gcd14p complex is the essential two-subunit tRNA(1-methyladenosine) methyltransferase of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 2000, 97, 5173–5178. [Google Scholar] [CrossRef] [PubMed]
- Saikia, M.; Fu, Y.E.; Pavon-Eternod, M.; He, C.; Pan, T. Genome-wide analysis of N1-methyl-adenosine modification in human tRNAs. RNA 2010, 16, 1317–1327. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Clark, W.; Luo, G.; Wang, X.; Fu, Y.; Wei, J.; Wang, X.; Hao, Z.; Dai, Q.; Zheng, G.; et al. ALKBH1-Mediated tRNA Demethylation Regulates Translation. Cell 2016, 167, 816–828. [Google Scholar] [CrossRef] [PubMed]
- Thanassi, J.A.; Hartman-Neumann, S.L.; Dougherty, T.J.; Dougherty, B.A.; Pucci, M.J. Identification of 113 conserved essential genes using a high-throughput gene disruption system in Streptococcus pneumoniae. Nucleic Acids Res. 2002, 30, 3152–3162. [Google Scholar] [CrossRef] [PubMed]
- Hoernes, T.P.; Hüttenhofer, A.; Erlacher, M.D. mRNA modifications: Dynamic regulators of gene expression? RNA Biol. 2016, 13, 760–765. [Google Scholar] [CrossRef] [PubMed]
- Dominissini, D.; Nachtergaele, S.; Moshitch-Moshkovitz, S.; Peer, E.; Kol, N.; Ben-Haim, M.S.; Dai, Q.; Di Segni, A.; Salmon-Divon, M.; Clark, W.C.; et al. The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA. Nature 2016, 530, 441–446. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xiong, X.; Wang, K.; Wang, L.; Shu, X.; Ma, S.; Yi, C. Transcriptome-wide mapping reveals reversible and dynamic N1-methyladenosine methylome. Nat. Chem. Biol. 2016, 12, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Duncan, T.; Trewick, S.C.; Koivisto, P.; Bates, P.A.; Lindahl, T.; Sedgwick, B. Reversal of DNA alkylation damage by two human dioxygenases. Proc. Natl. Acad. Sci. USA 2002, 99, 16660–16665. [Google Scholar] [CrossRef] [PubMed]
- Aas, P.A.; Otterlei, M.; Falnes, P.O.; Vågbø, C.B.; Skorpen, F.; Akbari, M.; Sundheim, O.; Bjørås, M.; Slupphaug, G.; Seeberg, E.; et al. Human and bacterial oxidative demethylases repair alkylation damage in both RNA and DNA. Nature 2003, 421, 859–863. [Google Scholar] [CrossRef] [PubMed]
- Ito, T.; Masuda, I.; Yoshida, K.; Goto-Ito, S.; Sekine, S.; Suh, S.W.; Hou, Y.M.; Yokoyama, S. Structural basis for methyl-donor-dependent and sequence-specific binding to tRNA substrates by knotted methyltransferase TrmD. Proc. Natl. Acad. Sci. USA 2015, 112, E4197–E4205. [Google Scholar] [CrossRef] [PubMed]
- Aravind, L.; Koonin, E.V. THUMP—a predicted RNA-binding domain shared by 4-thiouridine, pseudouridine synthases and RNA methylases. Trends Biochem. Sci. 2001, 26, 215–217. [Google Scholar] [CrossRef]
- Aravind, L.; Koonin, E.V. Novel predicted RNA-binding domains associated with the translation machinery. J. Mol. Evol. 1999, 48, 291–302. [Google Scholar] [CrossRef] [PubMed]
- Salas, C.E.; Dirheimer, G. In vitro methylation of yeast tRNAAsp by rat brain cortical tRNA-(adenine-1) methyltransferase. Nucleic Acids Res. 1979, 6, 1123–1133. [Google Scholar] [CrossRef]
- Ahn, H.J.; Kim, H.W.; Yoon, H.J.; Lee, B.I.; Suh, S.W.; Yang, J.K. Crystal structure of tRNA(m1G37)methyltransferase: Insights into tRNA recognition. EMBO J. 2003, 22, 2593–2603. [Google Scholar] [CrossRef] [PubMed]
- Christian, T.; Lahoud, G.; Liu, C.; Hoffmann, K.; Perona, J.J.; Hou, Y.M. Mechanism of N-methylation by the tRNA m1G37 methyltransferase Trm5. RNA 2010, 16, 2484–2492. [Google Scholar] [CrossRef] [PubMed]
- Goto-Ito, S.; Ito, T.; Kuratani, M.; Bessho, Y.; Yokoyama, S. Tertiary structure checkpoint at anticodon loop modification in tRNA functional maturation. Nat. Struct. Mol. Biol. 2009, 16, 1109–1115. [Google Scholar] [CrossRef] [PubMed]
- Elkins, P.A.; Watts, J.M.; Zalacain, M.; van Thiel, A.; Vitazka, P.R.; Redlak, M.; Andraos-Selim, C.; Rastinejad, F.; Holmes, W.M. Insights into Catalysis by a Knotted TrmD tRNA Methyltransferase. J. Mol. Biol. 2003, 333, 931–949. [Google Scholar] [CrossRef] [PubMed]
- Sakaguchi, R.; Lahoud, G.; Christian, T.; Gamper, H.; Hou, Y.M. A Divalent Metal Ion-Dependent N1-Methyl Transfer to G37-tRNA. Chem. Biol. 2014, 21, 1351–1360. [Google Scholar] [CrossRef] [PubMed]
- Barraud, P.; Golinelli-Pimpaneau, B.; Atmanene, C.; Sanglier, S.; Van Dorsselaer, A.; Droogmans, L.; Dardel, F.; Tisné, C. Crystal Structure of Thermus thermophilus tRNA m1A58 Methyltransferase and Biophysical Characterization of Its Interaction with tRNA. J. Mol. Biol. 2008, 377, 535–550. [Google Scholar] [CrossRef] [PubMed]
- Dégut, C.; Ponchon, L.; Folly-Klan, M.; Barraud, P.; Tisné, C. The m1A58 modification in eubacterial tRNA: An overview of tRNA recognition and mechanism of catalysis by TrmI. Biophys. Chem. 2016, 210, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Finer-Moore, J.; Czudnochowski, N.; O’Connell, J.D.; Wang, A.L.; Stroud, R.M. Crystal Structure of the Human tRNA m1A58 Methyltransferase–tRNA3Lys Complex: Refolding of Substrate tRNA Allows Access to the Methylation Target. J. Mol. Biol. 2015, 427, 3862–3876. [Google Scholar] [CrossRef] [PubMed]
- Bujnicki, J.M.; Rychlewski, L. Grouping together highly diverged PD-(D/E)XK nucleases and identification of novel superfamily members using structure-guided alignment of sequence profiles. J. Mol. Microbiol. Biotechnol. 2001, 3, 69–72. [Google Scholar] [PubMed]
- Bujnicki, J.M. In silico analysis of the tRNA:m1A58 methyltransferase family: Homology-based fold prediction and identification of new members from Eubacteria and Archaea. FEBS Lett. 2001, 507, 123–127. [Google Scholar] [CrossRef]
- Chujo, T.; Suzuki, T. Trmt61B is a methyltransferase responsible for 1-methyladenosine at position 58 of human mitochondrial tRNAs. RNA 2012, 18, 2269–2276. [Google Scholar] [CrossRef] [PubMed]
- Roovers, M.; Wouters, J.; Bujnicki, J.M.; Tricot, C.; Stalon, V.; Grosjean, H.; Droogmans, L. A primordial RNA modification enzyme: The case of tRNA (m1A) methyltransferase. Nucleic Acids Res. 2004, 32, 465–476. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Zhu, Y.; Wang, C.; Fan, X.; Jiang, X.; Ebrahimi, M.; Qiao, Z.; Niu, L.; Teng, M.; Li, X. Crystal structure of the two-subunit tRNA m1A58 methyltransferase TRM6–TRM61 from Saccharomyces cerevisiae. Sci. Rep. 2016, 6, 32562. [Google Scholar] [CrossRef] [PubMed]
- Guelorget, A.; Roovers, M.; Guérineau, V.; Barbey, C.; Li, X.; Golinelli-Pimpaneau, B. Insights into the hyperthermostability and unusual region-specificity of archaeal Pyrococcus abyssi tRNA m1A57/58 methyltransferase. Nucleic Acids Res. 2010, 38, 6206–6218. [Google Scholar] [CrossRef] [PubMed]
- Kuratani, M.; Yanagisawa, T.; Ishii, R.; Matsuno, M.; Si, S.Y.; Katsura, K.; Ushikoshi-Nakayama, R.; Shibata, R.; Shirouzu, M.; Bessho, Y.; et al. Crystal structure of tRNA m1A58 methyltransferase TrmI from Aquifex aeolicus in complex with S-adenosyl-l-methionine. J. Struct. Funct. Genomics. 2014, 15, 173–180. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Kumar, P.H.; Dineshkumar, T.K.; Varshney, U.; Subramanya, H.S. Crystal structure of Rv2118c: an AdoMet-dependent methyltransferase from Mycobacterium tuberculosis H37Rv. J. Mol. Biol. 2001, 312, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Ozanick, S.; Krecic, A.; Andersland, J.; Anderson, J.T. The bipartite structure of the tRNA m1A58 methyltransferase from S. cerevisiae is conserved in humans. RNA 2005, 11, 1281–1290. [Google Scholar] [CrossRef] [PubMed]
- Kadaba, S.; Krueger, A.; Trice, T.; Krecic, A.M.; Hinnebusch, A.G.; Anderson, J. Nuclear surveillance and degradation of hypomodified initiator tRNAMet in S. cerevisiae. Genes Dev. 2004, 18, 1227–1240. [Google Scholar] [CrossRef] [PubMed]
- Guelorget, A.; Barraud, P.; Tisné, C.; Golinelli-Pimpaneau, B. Structural comparison of tRNA m1A58 methyltransferases revealed different molecular strategies to maintain their oligomeric architecture under extreme conditions. BMC Struct. Biol. 2011, 11, 48. [Google Scholar] [CrossRef] [PubMed]
- Yasukawa, T.; Suzuki, T.; Suzuki, T.; Ueda, T.; Ohta, S.; Watanabe, K. Modification defect at anticodon wobble nucleotide of mitochondrial tRNAsLeu(UUR) with pathogenic mutations of mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes. J. Biol. Chem. 2000, 275, 4251–4257. [Google Scholar] [CrossRef] [PubMed]
- Yokogawa, T.; Watanabe, Y.; Kumazawa, Y.; Ueda, T.; Hirao, I.; Miura, K.I.; Watanabe, K. A novel cloverleaf strcuture found in mammalian mitochondrial tRNASer(UCN). Nucleic Acids Res. 1991, 19, 6101–6105. [Google Scholar] [CrossRef] [PubMed]
- Kempenaers, M.; Roovers, M.; Oudjama, Y.; Tkaczuk, K.L.; Bujnicki, J.M.; Droogmans, L. New archaeal methyltransferases forming 1-methyladenosine or 1-methyladenosine and 1-methylguanosine at position 9 of tRNA. Nucleic Acids Res. 2010, 38, 6533–6543. [Google Scholar] [CrossRef] [PubMed]
- Jackman, J.E. Identification of the yeast gene encoding the tRNA m1G methyltransferase responsible for modification at position 9. RNA 2003, 9, 574–585. [Google Scholar] [CrossRef] [PubMed]
- Shao, Z.; Yan, W.; Peng, J.; Zuo, X.; Zou, Y.; Li, F.; Gong, D.; Ma, R.; Wu, J.; Shi, Y.; et al. Crystal structure of tRNA m1G9 methyltransferase Trm10: Insight into the catalytic mechanism and recognition of tRNA substrate. Nucleic Acids Res. 2014, 42, 509–525. [Google Scholar] [CrossRef] [PubMed]
- Van Laer, B.; Roovers, M.; Wauters, L.; Kasprzak, J.M.; Dyzma, M.; Deyaert, E.; Kumar Singh, R.; Feller, A.; Bujnicki, J.M.; Droogmans, L.; et al. Structural and functional insights into tRNA binding and adenosine N1-methylation by an archaeal Trm10 homologue. Nucleic Acids Res. 2015, 44, 940–953. [Google Scholar] [CrossRef] [PubMed]
- Pütz, J.; Dupuis, B.; Sissler, M.; Florentz, C. Mamit-tRNA, a database of mammalian mitochondrial tRNA primary and secondary structures. RNA 2007, 13, 1184–1190. [Google Scholar] [CrossRef] [PubMed]
- Jomvall, H.; Persson, B.; Krook, M.; Atrian, S.; Gonzàlez-Duarte, R.; Jeffery, J.; Ghosh, D. Short-Chain Dehydrogenases/Reductases (SDR). Biochemistry 1995, 34, 6003–6013. [Google Scholar]
- Kallberg, Y.; Oppermann, U.; Jörnvall, H.; Persson, B. Short-chain dehydrogenase/reductase (SDR) relationships: A large family with eight clusters common to human, animal, and plant genomes. Protein Sci. 2002, 11, 636–641. [Google Scholar] [CrossRef] [PubMed]
- Kavanagh, K.L.; Jörnvall, H.; Persson, B.; Oppermann, U. Medium- and short-chain dehydrogenase/reductase gene and protein families: the SDR superfamily: functional and structural diversity within a family of metabolic and regulatory enzymes. Cell. Mol. Life Sci. 2008, 65, 3895–3906. [Google Scholar] [CrossRef] [PubMed]
- Lopez Sanchez, M.I.; Mercer, T.R.; Davies, S.M.; Shearwood, A.M.J.; Nygård, K.K.; Richman, T.R.; Mattick, J.S.; Rackham, O.; Filipovska, A. RNA processing in human mitochondria. Cell Cycle 2011, 10, 2904–2916. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.J.; Zhou, M.; Fang, Z.P.; Wang, M.; Zhou, X.L.; Wang, E.D. The tRNA recognition mechanism of the minimalist SPOUT methyltransferase, TrmL. Nucleic Acids Res. 2013, 41, 7828–7842. [Google Scholar] [CrossRef] [PubMed]
- Christian, T.; Lahoud, G.; Liu, C.; Hou, Y.M. Control of Catalytic Cycle by a Pair of Analogous tRNA Modification Enzymes. J. Mol. Biol. 2010, 400, 204–217. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K.; Nureki, O.; Fukai, S.; Ishii, R.; Okamoto, H.; Yokoyama, S.; Endo, Y.; Hori, H. Roles of conserved amino acid sequence motifs in the SpoU (TrmH) RNA methyltransferase family. J. Biol. Chem. 2005, 280, 10368–10377. [Google Scholar] [CrossRef] [PubMed]
- Schilling, O.; Späth, B.; Kostelecky, B.; Marchfelder, A.; Meyer-Klaucke, W.; Vogel, A. Exosite modules guide substrate recognition in the ZiPD/ElaC protein family. J. Biol. Chem. 2005, 280, 17857–17862. [Google Scholar] [CrossRef] [PubMed]
- Kostelecky, B.; Pohl, E.; Vogel, A.; Schilling, O.; Meyer-Klaucke, W. The crystal structure of the zinc phosphodiesterase from Escherichia coli provides insight into function and cooperativity of tRNase Z-family proteins. J. Bacteriol. 2006, 188, 1607–1614. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.J.; Long, T.; Zhou, M.; Zhou, X.L.; Wang, E.D. tRNA recognition by a bacterial tRNA Xm32 modification enzyme from the SPOUT methyltransferase superfamily. Nucleic Acids Res. 2015, 43, 7489–7503. [Google Scholar] [CrossRef] [PubMed]
| tRNA nt-position | Domain (tRNA cellular location) | MTase Superfamily | MTase Subfamily |
|---|---|---|---|
| 9 | E (mt), A | SPOUT/class IV | Trm10 * |
| 14 | E (cyt) | Unknown | Unknown |
| 22 | B | RFM/class I | TrmK |
| 58 | E (cyt) | RFM/class I | Trm6/Trm61 |
| 58 | E (mt) | RFM/class I | Trm61 * |
| 58 | A, B | RFM/class I | TrmI |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oerum, S.; Dégut, C.; Barraud, P.; Tisné, C. m1A Post‐Transcriptional Modification in tRNAs. Biomolecules 2017, 7, 20. https://doi.org/10.3390/biom7010020
Oerum S, Dégut C, Barraud P, Tisné C. m1A Post‐Transcriptional Modification in tRNAs. Biomolecules. 2017; 7(1):20. https://doi.org/10.3390/biom7010020
Chicago/Turabian StyleOerum, Stephanie, Clément Dégut, Pierre Barraud, and Carine Tisné. 2017. "m1A Post‐Transcriptional Modification in tRNAs" Biomolecules 7, no. 1: 20. https://doi.org/10.3390/biom7010020
APA StyleOerum, S., Dégut, C., Barraud, P., & Tisné, C. (2017). m1A Post‐Transcriptional Modification in tRNAs. Biomolecules, 7(1), 20. https://doi.org/10.3390/biom7010020





