Nuclear Tau and Its Potential Role in Alzheimer’s Disease
Abstract
:1. Introduction
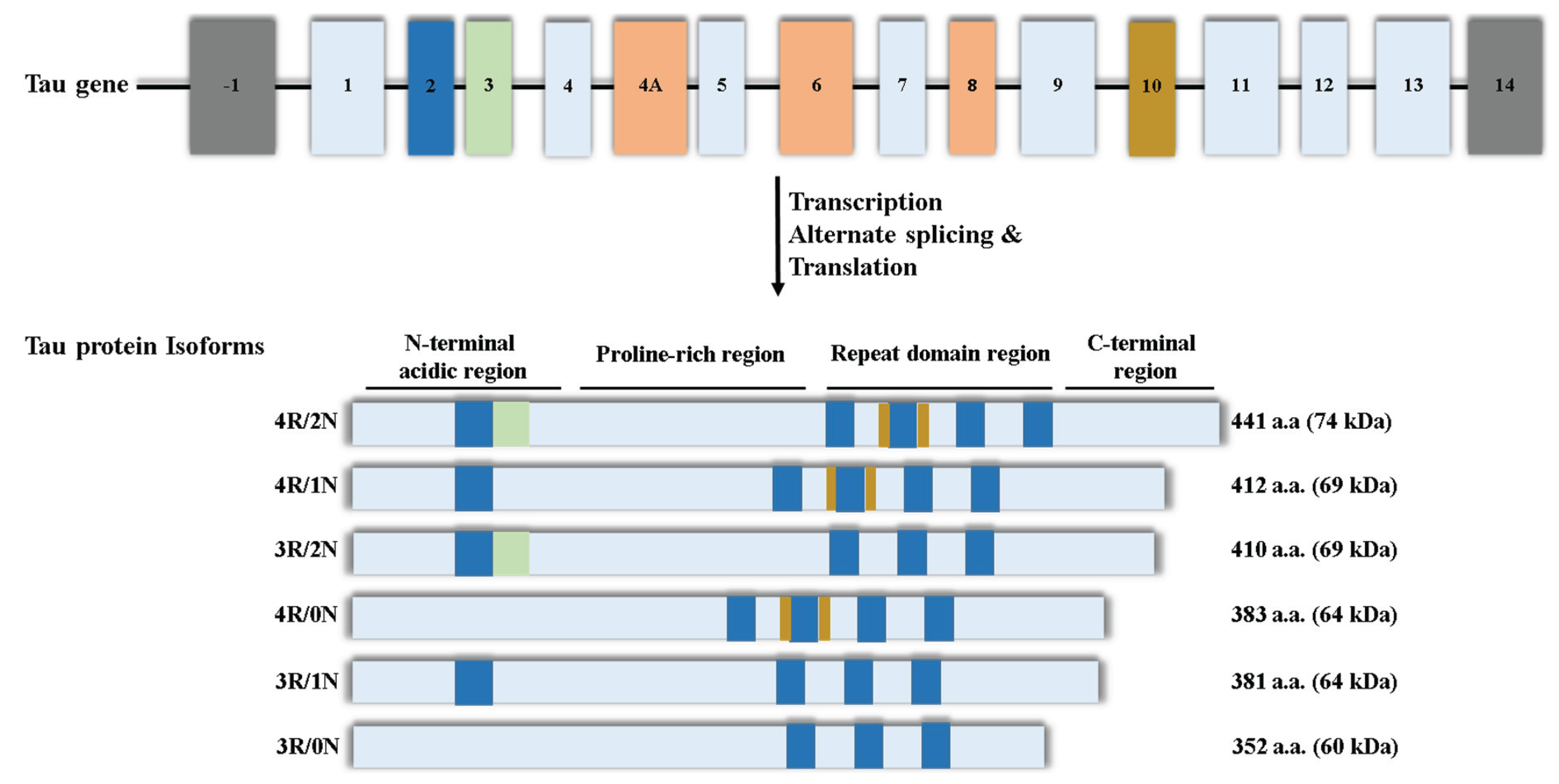

2. Nuclear Tau and Alzheimer’s Disease
2.1. Cellular Localisation of Tau
2.2. Nuclear Tau: Three Decades of Discovery
2.3. The Identity of Nuclear Tau
2.4. Capacity of Tau to Interact with the DNA
2.5. Functional Role of Nuclear Tau

2.6. Role of Nuclear Tau in Neurodegeneration
3. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Weingarten, M.D.; Lockwood, A.H.; Hwo, S.Y.; Kirschner, M.W. A protein factor essential for microtubule assembly. Proc. Natl. Acad. Sci. USA 1975, 72, 1858–1862. [Google Scholar] [CrossRef] [PubMed]
- Rossi, G.; Dalpra, L.; Crosti, F.; Lissoni, S.; Sciacca, F.L.; Catania, M.; di Fede, G.; Mangieri, M.; Giaccone, G.; Croci, D.; et al. A new function of microtubule-associated protein tau: Involvement in chromosome stability. Cell Cycle 2008, 7, 1788–1794. [Google Scholar] [CrossRef] [PubMed]
- Loomis, P.A.; Howard, T.H.; Castleberry, R.P.; Binder, L.I. Identification of nuclear tau isoforms in human neuroblastoma cells. Proc. Natl. Acad. Sci. USA 1990, 87, 8422–8426. [Google Scholar] [CrossRef] [PubMed]
- Stoothoff, W.H.; Johnson, G.V.W. Tau phosphorylation: Physiological and pathological consequences. Biochim. Biophys. Acta Mol. Basis Dis. 2005, 1739, 280–297. [Google Scholar] [CrossRef] [PubMed]
- Martin, L.; Latypova, X.; Terro, F. Post-translational modifications of tau protein: Implications for Alzheimer’s disease. Neurochem. Int. 2011, 58, 458–471. [Google Scholar] [CrossRef] [PubMed]
- Buee, L.; Bussiere, T.; Buee-Scherrer, V.; Delacourte, A.; Hof, P.R. Tau protein isoforms, phosphorylation and role in neurodegenerative disorders. Brain Res. Rev. 2000, 33, 95–130. [Google Scholar] [CrossRef]
- Dixit, R.; Ross, J.L.; Goldman, Y.E.; Holzbaur, E.L. Differential regulation of dynein and kinesin motor proteins by tau. Science 2008, 319, 1086–1089. [Google Scholar] [CrossRef] [PubMed]
- Ittner, L.M.; Ke, Y.D.; Delerue, F.; Bi, M.; Gladbach, A.; van Eersel, J.; Wolfing, H.; Chieng, B.C.; Christie, M.J.; Napier, I.A.; et al. Dendritic function of tau mediates amyloid-beta toxicity in Alzheimer’s disease mouse models. Cell 2010, 142, 387–397. [Google Scholar] [CrossRef] [PubMed]
- Neve, R.L.; Harris, P.; Kosik, K.S.; Kurnit, D.M.; Donlon, T.A. Identification of cDNA clones for the human microtubule-associated protein tau and chromosomal localization of the genes for tau and microtubule-associated protein 2. Mol. Brain Res. 1986, 1, 271–280. [Google Scholar] [CrossRef]
- Andreadis, A.; Brown, W.M.; Kosik, K.S. Structure and novel exons of the human. tau. gene. Biochemistry 1992, 31, 10626–10633. [Google Scholar] [CrossRef] [PubMed]
- Andreadis, A. Tau gene alternative splicing: Expression patterns, regulation and modulation of function in normal brain and neurodegenerative diseases. Biochim. Biophys. Acta Mol. Basis Dis. 2005, 1739, 91–103. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Loomis, P.A.; Zinkowski, R.P.; Binder, L.I. A novel tau transcript in cultured human neuroblastoma cells expressing nuclear tau. J. Cell Biol. 1993, 121, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Götz, J. Profiling murine tau with 0N, 1N and 2N isoform-specific antibodies in brain and peripheral organs reveals distinct subcellular localization, with the 1N isoform being enriched in the nucleus. PLoS ONE 2013, 8, e84849. [Google Scholar] [CrossRef] [PubMed]
- Georgieff, I.S.; Liem, R.K.; Couchie, D.; Mavilia, C.; Nunez, J.; Shelanski, M.L. Expression of high molecular weight tau in the central and peripheral nervous systems. J. Cell Sci. 1993, 10, 729–737. [Google Scholar]
- Nunez, J.; Fischer, I. Microtubule-associated proteins (maps) in the peripheral nervous system during development and regeneration. J. Mol. Neurosci. 1997, 8, 207–222. [Google Scholar] [CrossRef] [PubMed]
- Sadot, E.; Marx, R.; Barg, J.; Behar, L.; Ginzburg, I. Complete sequence of 3ʹ-untranslated region of tau from rat central nervous system: Implications for mRNA heterogeneity. J. Mol. Biol. 1994, 241, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Behar, L.; Marx, R.; Sadot, E.; Barg, J.; Ginzburg, I. cis-Acting signals and trans-acting proteins are involved in tau mRNA targeting into neurites of differentiating neuronal cells. Int. J. Dev. Neurosci. 1995, 13, 113–127. [Google Scholar] [CrossRef]
- Gupta, I.; Clauder-Münster, S.; Klaus, B.; Järvelin, A.I.; Aiyar, R.S.; Benes, V.; Wilkening, S.; Huber, W.; Pelechano, V.; Steinmetz, L.M. Alternative polyadenylation diversifies post-transcriptional regulation by selective RNA-protein interactions. Mol. Syst. Biol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Luo, M.H.; Tse, S.W.; Memmott, J.; Andreadis, A. Novel isoforms of tau that lack the microtubule-binding domain. J. Neurochem. 2004, 90, 340–351. [Google Scholar] [CrossRef] [PubMed]
- Shea, T.B.; Cressman, C.M. A 26–30 kda developmentally-regulated tau isoform localized within nuclei of mitotic human neuroblastoma cells. Int. J. Dev. Neurosci. Off. J. Int. Soc. Dev. Neurosci. 1998, 16, 41–48. [Google Scholar] [CrossRef]
- Connell, J.W.; Rodriguez-Martin, T.; Gibb, G.M.; Kahn, N.M.; Grierson, A.J.; Hanger, D.P.; Revesz, T.; Lantos, P.L.; Anderton, B.H.; Gallo, J.M. Quantitative analysis of tau isoform transcripts in sporadic tauopathies. Mol. Brain Res. 2005, 137, 104–109. [Google Scholar] [CrossRef] [PubMed]
- Cross, D.C.; Munoz, J.P.; Hernandez, P.; Maccioni, R.B. Nuclear and cytoplasmic tau proteins from human nonneuronal cells share common structural and functional features with brain tau. J. Cell. Biochem. 2000, 78, 305–317. [Google Scholar] [CrossRef]
- Uberti, D.; Rizzini, C.; Spano, P.F.; Memo, M. Characterization of tau proteins in human neuroblastoma SH-SY5Y cell line. Neurosci. Lett. 1997, 235, 149–153. [Google Scholar] [CrossRef]
- Alzheimer, A.; Stelzmann, R.A.; Schnitzlein, H.N.; Murtagh, F.R. An english translation of Alzheimer’s 1907 paper, “uber eine eigenartige erkankung der hirnrinde”. Clin. Anat. 1995, 8, 429–431. [Google Scholar] [PubMed]
- Crowther, R.A.; Wischik, C.M. Image reconstruction of the Alzheimer paired helical filament. EMBO J. 1985, 4, 3661–3665. [Google Scholar] [PubMed]
- Berriman, J.; Serpell, L.C.; Oberg, K.A.; Fink, A.L.; Goedert, M.; Crowther, R.A. Tau filaments from human brain and from in vitro assembly of recombinant protein show cross-beta structure. Proc. Natl. Acad. Sci. USA 2003, 100, 9034–9038. [Google Scholar] [CrossRef] [PubMed]
- Spillantini, M.G.; Goedert, M. Tau pathology and neurodegeneration. Lancet Neurol. 2013, 12, 609–622. [Google Scholar] [CrossRef]
- Alonso, A.; Zaidi, T.; Novak, M.; Grundke-Iqbal, I.; Iqbal, K. Hyperphosphorylation induces self-assembly of tau into tangles of paired helical filaments/straight filaments. Proc. Natl. Acad. Sci. USA 2001, 98, 6923–6928. [Google Scholar] [CrossRef] [PubMed]
- Novak, M.; Zilka, N.; Kovacech, B.; Barath, P.; Kontsekova, E. Tau truncation: The most productive post-translational modification. Alzheimer’s Dement. J. Alzheimer’s Assoc. 2012. [Google Scholar] [CrossRef]
- Kovacech, B.; Novak, M. Tau truncation is a productive posttranslational modification of neurofibrillary degeneration in Alzheimer’s disease. Curr. Alzheimer Res. 2010, 7, 708–716. [Google Scholar] [CrossRef] [PubMed]
- Wischik, C.M.; Novak, M.; Thogersen, H.C.; Edwards, P.C.; Runswick, M.J.; Jakes, R.; Walker, J.E.; Milstein, C.; Roth, M.; Klug, A. Isolation of a fragment of tau derived from the core of the paired helical filament of Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 1988, 85, 4506–4510. [Google Scholar] [CrossRef] [PubMed]
- Zilka, N.; Kovacech, B.; Barath, P.; Kontsekova, E.; Novak, M. The self-perpetuating tau truncation circle. Biochem. Soc. Trans. 2012, 40, 681–686. [Google Scholar] [CrossRef] [PubMed]
- Zilka, N.; Filipcik, P.; Koson, P.; Fialova, L.; Skrabana, R.; Zilkova, M.; Rolkova, G.; Kontsekova, E.; Novak, M. Truncated tau from sporadic Alzheimer’s disease suffices to drive neurofibrillary degeneration in vivo. FEBS Lett. 2006, 580, 3582–3588. [Google Scholar] [CrossRef] [PubMed]
- Flores-Rodriguez, P.; Ontiveros-Torres, M.A.; Cardenas-Aguayo, M.C.; Luna-Arias, J.P.; Meraz-Rios, M.A.; Viramontes-Pintos, A.; Harrington, C.R.; Wischik, C.M.; Mena, R.; Floran-Garduno, B.; et al. The relationship between truncation and phosphorylation at the C-terminus of tau protein in the paired helical filaments of Alzheimer’s disease. Front. Neurosci. 2015. [Google Scholar] [CrossRef] [PubMed]
- Papasozomenos, S.C.; Binder, L.I. Phosphorylation determines two distinct species of tau in the central nervous system. Cell Motil. Cytoskelet. 1987, 8, 210–226. [Google Scholar] [CrossRef] [PubMed]
- Binder, L.I.; Frankfurter, A.; Rebhun, L.I. The distribution of tau in the mammalian central nervous system. J. Cell Biol. 1985, 101, 1371–1378. [Google Scholar] [CrossRef] [PubMed]
- Trinczek, B.; Ebneth, A.; Mandelkow, E.M.; Mandelkow, E. Tau regulates the attachment/detachment but not the speed of motors in microtubule-dependent transport of single vesicles and organelles. J. Cell Sci. 1999, 112, 2355–2367. [Google Scholar] [PubMed]
- Di Tella, M.; Feiguin, F.; Morfini, G.; Caceres, A. Microfilament-associated growth cone component depends upon tau for its intracellular localization. Cell Motil. Cytoskelet. 1994, 29, 117–130. [Google Scholar] [CrossRef] [PubMed]
- Black, M.M.; Slaughter, T.; Moshiach, S.; Obrocka, M.; Fischer, I. Tau is enriched on dynamic microtubules in the distal region of growing axons. J. Neurosci. 1996, 16, 3601–3619. [Google Scholar] [PubMed]
- Preuss, U.; Döring, F.; Illenberger, S.; Mandelkow, E.M. Cell cycle-dependent phosphorylation and microtubule binding of tau protein stably transfected into chinese hamster ovary cells. Mol. Biol. Cell 1995, 6, 1397–1410. [Google Scholar] [CrossRef] [PubMed]
- Lee, G. Tau and src family tyrosine kinases. Biochim. Biophys. Acta Mol. Basis Dis. 2005, 1739, 323–330. [Google Scholar] [CrossRef] [PubMed]
- Arrasate, M.; Pérez, M.; Avila, J. Tau dephosphorylation at tau-1 site correlates with its association to cell membrane. Neurochem. Res. 2000, 25, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Pooler, A.M.; Usardi, A.; Evans, C.J.; Philpott, K.L.; Noble, W.; Hanger, D.P. Dynamic association of tau with neuronal membranes is regulated by phosphorylation. Neurobiol. Aging 2012, 33, e427–e438. [Google Scholar] [CrossRef] [PubMed]
- Kawarabayashi, T.; Shoji, M.; Younkin, L.H.; Wen-Lang, L.; Dickson, D.W.; Murakami, T.; Matsubara, E.; Abe, K.; Ashe, K.H.; Younkin, S.G. Dimeric amyloid beta protein rapidly accumulates in lipid rafts followed by apolipoprotein E and phosphorylated tau accumulation in the Tg2576 mouse model of Alzheimer’s disease. J. Neurosci. Off. J. Soc. Neurosci. 2004, 24, 3801–3809. [Google Scholar] [CrossRef] [PubMed]
- Williamson, R.; Usardi, A.; Hanger, D.P.; Anderton, B.H. Membrane-bound beta-amyloid oligomers are recruited into lipid rafts by a fyn-dependent mechanism. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2008, 22, 1552–1559. [Google Scholar]
- Sahara, N.; Murayama, M.; Higuchi, M.; Suhara, T.; Takashima, A. Biochemical distribution of tau protein in synaptosomal fraction of transgenic mice expressing human p301l tau. Front. Neurol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Hoover, B.R.; Reed, M.N.; Su, J.; Penrod, R.D.; Kotilinek, L.A.; Grant, M.K.; Pitstick, R.; Carlson, G.A.; Lanier, L.M.; Yuan, L.L.; et al. Tau mislocalization to dendritic spines mediates synaptic dysfunction independently of neurodegeneration. Neuron 2010, 68, 1067–1081. [Google Scholar] [CrossRef] [PubMed]
- Pooler, A.M.; Hanger, D.P. Functional implications of the association of tau with the plasma membrane. Biochem. Soc. Trans. 2010, 38, 1012–1015. [Google Scholar] [CrossRef] [PubMed]
- Papasozomenos, S.C. Tau protein immunoreactivity in dementia of the Alzheimer type: II. Electron microscopy and pathogenetic implications. Effects of fixation on the morphology of the Alzheimer’s abnormal filaments. Labor. Investig. 1989, 60, 375–389. [Google Scholar]
- Brady, R.M.; Zinkowski, R.P.; Binder, L.I. Presence of tau in isolated nuclei from human brain. Neurobiol. Aging 1995, 16, 479–486. [Google Scholar] [PubMed]
- Tang, Z.; Ioja, E.; Bereczki, E.; Hultenby, K.; Li, C.; Guan, Z.; Winblad, B.; Pei, J.J. Mtor mediates tau localization and secretion: Implication for Alzheimer’s disease. Biochim. Biophys. Acta 2015, 1853, 1646–1657. [Google Scholar] [CrossRef] [PubMed]
- Sultan, A.; Nesslany, F.; Violet, M.; Begard, S.; Loyens, A.; Talahari, S.; Mansuroglu, Z.; Marzin, D.; Sergeant, N.; Humez, S.; et al. Nuclear tau, a key player in neuronal DNA protection. J. Biol. Chem. 2011, 286, 4566–4575. [Google Scholar] [CrossRef] [PubMed]
- Violet, M.; Delattre, L.; Tardivel, M.; Sultan, A.; Chauderlier, A.; Caillierez, R.; Talahari, S.; Nesslany, F.; Lefebvre, B.; Bonnefoy, E.; et al. A major role for tau in neuronal DNA and RNA protection in vivo under physiological and hyperthermic conditions. Front. Cell. Neurosci. 2014. [Google Scholar] [CrossRef] [PubMed]
- Metuzals, J.; Robitaille, Y.; Houghton, S.; Gauthier, S.; Leblanc, R. Paired helical filaments and the cytoplasmic-nuclear interface in Alzheimer’s disease. J. Neurocytol. 1988, 17, 827–833. [Google Scholar] [CrossRef] [PubMed]
- Thurston, V.C.; Pena, P.; Pestell, R.; Binder, L.I. Nucleolar localization of the microtubule-associated protein tau in neuroblastomas using sense and anti-sense transfection strategies. Cell Motil. Cytoskelet. 1997, 38, 100–110. [Google Scholar] [CrossRef]
- Greenwood, J.A.; Johnson, G.V. Localization and in situ phosphorylation state of nuclear tau. Exp. Cell Res. 1995, 220, 332–337. [Google Scholar] [CrossRef] [PubMed]
- Thurston, V.; Zinkowski, R.; Binder, L. Tau as a nucleolar protein in human nonneural cells in vitro and in vivo. Chromosoma 1996, 105, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Lu, Q.; Wood, J.G. Characterization of fluorescently derivatized bovine tau protein and its localization and functions in cultured chinese hamster ovary cells. Cell Motil. Cytoskelet. 1993, 25, 190–200. [Google Scholar] [CrossRef] [PubMed]
- Cross, D.; Tapia, L.; Garrido, J.; Maccioni, R.B. Tau-like proteins associated with centrosomes in cultured cells. Exp. Cell Res. 1996, 229, 378–387. [Google Scholar] [CrossRef] [PubMed]
- Lambert, M.P.; Sabo, S.; Zhang, C.; Enam, S.A.; Klein, W.L. Constitutive Alzheimer’s-type tau epitopes in a neuritogenic rat cns cell line. Neurobiol. Aging 1995, 16, 583–589. [Google Scholar] [CrossRef]
- Dong, X.X.; Wang, Y.; Qin, Z.H. Molecular mechanisms of excitotoxicity and their relevance to pathogenesis of neurodegenerative diseases. Acta Pharmacol. Sin. 2009, 30, 379–387. [Google Scholar] [CrossRef] [PubMed]
- Sjoberg, M.K.; Shestakova, E.; Mansuroglu, Z.; Maccioni, R.B.; Bonnefoy, E. Tau protein binds to pericentromeric DNA: A putative role for nuclear tau in nucleolar organization. J. Cell Sci. 2006, 119, 2025–2034. [Google Scholar] [CrossRef] [PubMed]
- Jamsa, A.; Hasslund, K.; Cowburn, R.F.; Backstrom, A.; Vasange, M. The retinoic acid and brain-derived neurotrophic factor differentiated sh-sy5y cell line as a model for Alzheimer’s disease-like tau phosphorylation. Biochem. Biophys. Res. Commun. 2004, 319, 993–1000. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Li, T.; He, R.; Bartlett, P.F.; Gotz, J. Visualizing the microtubule-associated protein tau in the nucleus. Sci. China Life Sci. 2014, 57, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Frost, B.; Hemberg, M.; Lewis, J.; Feany, M.B. Tau promotes neurodegeneration through global chromatin relaxation. Nat. Neurosci. 2014, 17, 357–366. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Miao, J.; Su, T.; Liu, Y.; He, R. Formaldehyde induces hyperphosphorylation and polymerization of tau protein both in vitro and in vivo. Biochim. Biophys. Acta 2013, 1830, 4102–4116. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Nogales, M.; Cabrera, J.R.; Santos-Galindo, M.; Hoozemans, J.J.M.; Ferrer, I.; Rozemuller, A.J.M.; Hernandez, F.; Avila, J.; Lucas, J.J. Huntington’s disease is a four-repeat tauopathy with tau nuclear rods. Nat. Med. 2014, 20, 881–885. [Google Scholar] [CrossRef] [PubMed]
- Bryan, J.B.; Nagle, B.W.; Doenges, K.H. Inhibition of tubulin assembly by RNA and other polyanions: Evidence for a required protein. Proc. Natl. Acad. Sci. USA 1975, 72, 3570–3574. [Google Scholar] [CrossRef] [PubMed]
- Kampers, T.; Friedhoff, P.; Biernat, J.; Mandelkow, E.M.; Mandelkow, E. RNA stimulates aggregation of microtubule-associated protein tau into Alzheimer-like paired helical filaments. FEBS Lett. 1996, 399, 344–349. [Google Scholar] [CrossRef]
- Corces, V.G.; Salas, J.; Salas, M.L.; Avila, J. Binding of microtubule proteins to DNA: Specificity of the interaction. Eur. J. Biochem. 1978, 86, 473–479. [Google Scholar] [CrossRef] [PubMed]
- Corces, V.G.; Manso, R.; De La Torre, J.; Avila, J.; Nasr, A.; Wiche, G. Effects of DNA on microtubule assembly. Eur. J. Biochem. FEBS 1980, 105, 7–16. [Google Scholar]
- Hua, Q.; He, R. Human neuronal tau promoting the melting temperature of DNA. Chin. Sci. Bull. 2000, 45, 999–1002. [Google Scholar] [CrossRef]
- Lu, Y.; He, H.J.; Zhou, J.; Miao, J.Y.; Lu, J.; He, Y.G.; Pan, R.; Wei, Y.; Liu, Y.; He, R.Q. Hyperphosphorylation results in tau dysfunction in DNA folding and protection. J. Alzheimer’s Dis. 2013, 37, 551–563. [Google Scholar]
- Tenreiro, S.; Eckermann, K.; Outeiro, T.F. Protein phosphorylation in neurodegeneration: Friend or foe? Front. Mol. Neurosci. 2014. [Google Scholar] [CrossRef] [PubMed]
- Hua, Q.; He, R.Q. Effect of phosphorylation and aggregation on tau binding to DNA. Protein Pept. Lett. 2002, 9, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Camero, S.; Benitez, M.J.; Cuadros, R.; Hernandez, F.; Avila, J.; Jimenez, J.S. Thermodynamics of the interaction between Alzheimer’s disease related tau protein and DNA. PLoS ONE 2014, 9, e104690. [Google Scholar] [CrossRef] [PubMed]
- Qi, H.; Cantrelle, F.-X.; Benhelli-Mokrani, H.; Smet-Nocca, C.; Buée, L.; Lippens, G.; Bonnefoy, E.; Galas, M.C.; Landrieu, I. Nuclear magnetic resonance spectroscopy characterization of interaction of tau with DNA and its regulation by phosphorylation. Biochemistry 2015, 54, 1525–1533. [Google Scholar] [CrossRef] [PubMed]
- Hua, Q.; He, R.Q.; Haque, N.; Qu, M.H.; del Carmen Alonso, A.; Grundke-Iqbal, I.; Iqbal, K. Microtubule associated protein tau binds to double-stranded but not single-stranded DNA. Cell. Mol. Life Sci. 2003, 60, 413–421. [Google Scholar] [CrossRef] [PubMed]
- Krylova, S.M.; Musheev, M.; Nutiu, R.; Li, Y.; Lee, G.; Krylov, S.N. Tau protein binds single-stranded DNA sequence specifically—The proof obtained in vitro with non-equilibrium capillary electrophoresis of equilibrium mixtures. FEBS Lett. 2005, 579, 1371–1375. [Google Scholar] [CrossRef] [PubMed]
- Qu, M.H.; Li, H.; Tian, R.; Nie, C.L.; Liu, Y.; Han, B.S.; He, R.Q. Neuronal tau induces DNA conformational changes observed by atomic force microscopy. Neuroreport 2004, 15, 2723–2727. [Google Scholar] [PubMed]
- Wei, Y.; Qu, M.H.; Wang, X.S.; Chen, L.; Wang, D.L.; Liu, Y.; Hua, Q.; He, R.Q. Binding to the minor groove of the double-strand, tau protein prevents DNA from damage by peroxidation. PLoS ONE 2008, 3, e2600. [Google Scholar] [CrossRef] [PubMed]
- Olson, M.O.J.; Dundr, M. Nucleolus: Structure and function; John Wiley & Sons, Ltd: Chichester, UK, 2001. [Google Scholar]
- Pickard, A.J.; Bierbach, U. The cell’s nucleolus: An emerging target for chemotherapeutic intervention. ChemMedChem 2013, 8, 1441–1449. [Google Scholar] [CrossRef] [PubMed]
- Grummt, I.; Pikaard, C.S. Epigenetic silencing of RNA polymerase I transcription. Nat. Rev. Mol. Cell Biol. 2003, 4, 641–649. [Google Scholar] [CrossRef] [PubMed]
- Guetg, C.; Lienemann, P.; Sirri, V.; Grummt, I.; Hernandez-Verdun, D.; Hottiger, M.O.; Fussenegger, M.; Santoro, R. The NoRC complex mediates the heterochromatin formation and stability of silent rRNA genes and centromeric repeats. Embo J. 2010, 29, 2135–2146. [Google Scholar] [CrossRef] [PubMed]
- Carmo-Fonseca, M.; Mendes-Soares, L.; Campos, I. To be or not to be in the nucleolus. Nat. Cell Biol. 2000, 2, E107–E112. [Google Scholar] [CrossRef] [PubMed]
- Rossi, G.; Conconi, D.; Panzeri, E.; Redaelli, S.; Piccoli, E.; Paoletta, L.; Dalpra, L.; Tagliavini, F. Mutations in MAPT gene cause chromosome instability and introduce copy number variations widely in the genome. J. Alzheimer’s Dis. 2013, 33, 969–982. [Google Scholar]
- Camero, S.; Benitez, M.J.; Barrantes, A.; Ayuso, J.M.; Cuadros, R.; Avila, J.; Jimenez, J.S. Tau protein provides DNA with thermodynamic and structural features which are similar to those found in histone-DNA complex. J. Alzheimer’s Dis. 2014, 39, 649–660. [Google Scholar]
- Welch, W.J.; Feramisco, J.R. Nuclear and nucleolar localization of the 72,000-dalton heat shock protein in heat-shocked mammalian cells. J. Biol. Chem. 1984, 259, 4501–4513. [Google Scholar] [PubMed]
- Chu, A.; Matusiewicz, N.; Stochaj, U. Heat-induced nuclear accumulation of hsc70s is regulated by phosphorylation and inhibited in confluent cells. FASEB J. Off. Publ. Fed.Am. Soc. Exp. Biol. 2001, 15, 1478–1480. [Google Scholar] [CrossRef]
- Welch, W.J.; Mizzen, L.A. Characterization of the thermotolerant cell. II. Effects on the intracellular distribution of heat-shock protein 70, intermediate filaments, and small nuclear ribonucleoprotein complexes. J. Cell Biol. 1988, 106, 1117–1130. [Google Scholar] [CrossRef] [PubMed]
- Mandell, R.B.; Feldherr, C.M. Identification of two hsp70-related xenopus oocyte proteins that are capable of recycling across the nuclear envelope. J. Cell Biol. 1990, 111, 1775–1783. [Google Scholar] [CrossRef] [PubMed]
- Papasozomenos, S.C.; Su, Y. Altered phosphorylation of tau protein in heat-shocked rats and patients with Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 1991, 88, 4543–4547. [Google Scholar] [CrossRef] [PubMed]
- Kotoglou, P.; Kalaitzakis, A.; Vezyraki, P.; Tzavaras, T.; Michalis, L.K.; Dantzer, F.; Jung, J.U.; Angelidis, C. Hsp70 translocates to the nuclei and nucleoli, binds to XRCC1 and PARP-1, and protects hela cells from single-strand DNA breaks. Cell Stress Chaperones 2009, 14, 391–406. [Google Scholar] [CrossRef] [PubMed]
- Niu, P.; Liu, L.; Gong, Z.; Tan, H.; Wang, F.; Yuan, J.; Feng, Y.; Wei, Q.; Tanguay, R.M.; Wu, T. Overexpressed heat shock protein 70 protects cells against DNA damage caused by ultraviolet C in a dose-dependent manner. Cell Stress Chaperones 2006, 11, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Tang, D.; Kang, R.; Xiao, W.; Jiang, L.; Liu, M.; Shi, Y.; Wang, K.; Wang, H.; Xiao, X. Nuclear heat shock protein 72 as a negative regulator of oxidative stress (hydrogen peroxide)-induced HMGB1 cytoplasmic translocation and release. J. Immunol. 2007, 178, 7376–7384. [Google Scholar] [CrossRef] [PubMed]
- Ke, Y.D.; Suchowerska, A.K.; van der Hoven, J.; de Silva, D.M.; Wu, C.W.; van Eersel, J.; Ittner, A.; Ittner, L.M. Lessons from tau-deficient mice. Int. J. Alzheimer’s Dis. 2012. [Google Scholar] [CrossRef] [PubMed]
- Travers, A.A.; Ner, S.S.; Churchill, M.E. DNA chaperones: A solution to a persistence problem? Cell 1994, 77, 167–169. [Google Scholar] [CrossRef]
- Zlatanova, J.; van Holde, K. Binding to four-way junction DNA: A common property of architectural proteins? FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 1998, 12, 421–431. [Google Scholar]
- Muller, S.; Scaffidi, P.; Degryse, B.; Bonaldi, T.; Ronfani, L.; Agresti, A.; Beltrame, M.; Bianchi, M.E. New embo members’ review: The double life of HMGB1 chromatin protein: Architectural factor and extracellular signal. EMBO J. 2001, 20, 4337–4340. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.B.; Lim, C.M.; Yu, Y.M.; Lee, J.K. Induction and subcellular localization of high-mobility group box-1 (HMGB1) in the postischemic rat brain. J. Neurosci. Res. 2008, 86, 1125–1131. [Google Scholar] [CrossRef] [PubMed]
- Barucker, C.; Harmeier, A.; Weiske, J.; Fauler, B.; Albring, K.F.; Prokop, S.; Hildebrand, P.; Lurz, R.; Heppner, F.L.; Huber, O.; et al. Nuclear translocation uncovers the amyloid peptide abeta42 as a regulator of gene transcription. J. Biol. Chem. 2014, 289, 20182–20191. [Google Scholar] [CrossRef] [PubMed]
- Padmaraju, V.; Indi, S.S.; Rao, K.S. New evidences on tau-DNA interactions and relevance to neurodegeneration. Neurochem. Int. 2010, 57, 51–57. [Google Scholar] [CrossRef] [PubMed]
- Ding, Q.; Markesbery, W.R.; Chen, Q.; Li, F.; Keller, J.N. Ribosome dysfunction is an early event in Alzheimer’s disease. J. Neurosci. Off. J. Soc. Neurosci. 2005, 25, 9171–9175. [Google Scholar] [CrossRef] [PubMed]
- Carole, A.B.; Angela, M.; Gronenborn, A.; Clore, G.M. Minor groove-binding architectural proteins: Structure, function, and DNA recognition. Annu. Rev. Biophys. Biomol. Struct. 1998, 27, 105–131. [Google Scholar]
- Falvo, J.V.; Thanos, D.; Maniatis, T. Reversal of intrinsic DNA bends in the IFNβ gene enhancer by transcription factors and the architectural protein HMG I(Y). Cell 1995, 83, 1101–1111. [Google Scholar] [CrossRef]
- Von Hippel, P.H. Protein-DNA recognition: New perspectives and underlying themes. Science 1994, 263, 769–770. [Google Scholar] [CrossRef] [PubMed]
- Deaton, A.M.; Bird, A. CpG islands and the regulation of transcription. Genes Dev. 2011, 25, 1010–1022. [Google Scholar] [CrossRef] [PubMed]
- Kolell, K.J.; Crawford, D.L. Evolution of sp transcription factors. Mol. Biol. Evol. 2002, 19, 216–222. [Google Scholar] [CrossRef] [PubMed]
- Gorner, W.; Durchschlag, E.; Martinez-Pastor, M.T.; Estruch, F.; Ammerer, G.; Hamilton, B.; Ruis, H.; Schuller, C. Nuclear localization of the C2H2 zinc finger protein MSN2P is regulated by stress and protein kinase a activity. Genes Dev. 1998, 12, 586–597. [Google Scholar] [CrossRef] [PubMed]
- Hao, N.; Budnik, B.A.; Gunawardena, J.; O’Shea, E.K. Tunable signal processing through modular control of transcription factor translocation. Science 2013, 339, 460–464. [Google Scholar] [CrossRef] [PubMed]
- Gómez de Barreda, E.; Dawson, H.N.; Vitek, M.P.; Avila, J. Tau deficiency leads to the upregulation of BAF-57, a protein involved in neuron-specific gene repression. FEBS Lett. 2010, 584, 2265–2270. [Google Scholar] [CrossRef] [PubMed]
- Oyama, F.; Kotliarova, S.; Harada, A.; Ito, M.; Miyazaki, H.; Ueyama, Y.; Hirokawa, N.; Nukina, N.; Ihara, Y. Gem GTPase and tau: Morphological changes induced by gem GTPase in cho cells are antagonized by tau. J. Biol. Chem. 2004, 279, 27272–27277. [Google Scholar] [CrossRef] [PubMed]
- Payao, S.L.; Smith, M.; Kormann-Bortolotto, M.H.; Toniolo, J. Investigation of the nucleolar organizer regions in Alzheimer’s disease. Gerontology 1994, 40, 13–17. [Google Scholar] [PubMed]
- Dönmez-Altuntaş, H.; Akalın, H.; Karaman, Y.; Demirtaş, H.; İmamoğlu, N.; Özkul, Y. Evaluation of the nucleolar organizer regions in Alzheimer’s disease. Gerontology 2005, 51, 297–301. [Google Scholar] [PubMed]
- Langstrom, N.S.; Anderson, J.P.; Lindroos, H.G.; Winblad, B.; Wallace, W.C. Alzheimer’s disease-associated reduction of polysomal mRNA translation. Mol. Brain Res. 1989, 5, 259–269. [Google Scholar] [CrossRef]
- Sajdel-Sulkowska, E.M.; Marotta, C.A. Alzheimer’s disease brain: Alterations in RNA levels and in a ribonuclease-inhibitor complex. Science 1984, 225, 947–949. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, A.M.; Payao, S.L.; Borsatto, B.; Bertolucci, P.H.; Smith, M.A. Quantitative evaluation of the rRNA in Alzheimer’s disease. Mech. Ageing Dev. 2000, 120, 57–64. [Google Scholar] [CrossRef]
- Hernandez-Ortega, K.; Garcia-Esparcia, P.; Gil, L.; Lucas, J.J.; Ferrer, I. Altered machinery of protein synthesis in Alzheimer’s: From the nucleolus to the ribosome. Brain Pathol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Pietrzak, M.; Rempala, G.; Nelson, P.T.; Zheng, J.J.; Hetman, M. Epigenetic silencing of nucleolar rRNA genes in Alzheimer’s disease. PLoS ONE 2011, 6, e22585. [Google Scholar] [CrossRef] [PubMed]
- Boeras, D.I.; Granic, A.; Padmanabhan, J.; Crespo, N.C.; Rojiani, A.M.; Potter, H. Alzheimer’s presenilin 1 causes chromosome missegregation and aneuploidy. Neurobiol. Aging 2008, 29, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Granic, A.; Padmanabhan, J.; Norden, M.; Potter, H. Alzheimer Aβ peptide induces chromosome mis-segregation and aneuploidy, including trisomy 21: Requirement for tau and APP. Mol. Biol. Cell 2010, 21, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Iourov, I.Y.; Vorsanova, S.G.; Liehr, T.; Yurov, Y.B. Aneuploidy in the normal, Alzheimer’s disease and ataxia-telangiectasia brain: Differential expression and pathological meaning. Neurobiol. Dis. 2009, 34, 212–220. [Google Scholar] [CrossRef] [PubMed]
- Potter, H. Review and hypothesis: Alzheimer disease and down syndrome—Chromosome 21 nondisjunction may underlie both disorders. Am. J. Hum. Genet. 1991, 48, 1192–1200. [Google Scholar] [PubMed]
- Mondragón-Rodríguez, S.; Perry, G.; Zhu, X.; Moreira, P.I.; Acevedo-Aquino, M.C.; Williams, S. Phosphorylation of tau protein as the link between oxidative stress, mitochondrial dysfunction, and connectivity failure: Implications for Alzheimer’s disease. Oxid. Med. Cell. Longev. 2013. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, T.; Ferreira, S.; Dupont-Wallois, L.; Bussiere, T.; Dupire, M.J.; Delacourte, A.; Michalski, J.C.; Caillet-Boudin, M.L. Evidence of a balance between phosphorylation and O-glcnac glycosylation of tau proteins—A role in nuclear localization. Biochim. Biophys. Acta 2003, 1619, 167–176. [Google Scholar] [CrossRef]
- Álvarez, G.; Aldudo, J.; Alonso, M.; Santana, S.; Valdivieso, F. Herpes simplex virus type 1 induces nuclear accumulation of hyperphosphorylated tau in neuronal cells. J. Neurosci. Res. 2012, 90, 1020–1029. [Google Scholar] [CrossRef] [PubMed]
- Diniz, B.S.O.; Pinto, J.A.; Forlenza, O.V. Do CSF total tau, phosphorylated tau, and β-amyloid 42 help to predict progression of mild cognitive impairment to Alzheimer’s disease? A systematic review and meta-analysis of the literature. World J. Biol. Psychiatry 2008, 9, 172–182. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bukar Maina, M.; Al-Hilaly, Y.K.; Serpell, L.C. Nuclear Tau and Its Potential Role in Alzheimer’s Disease. Biomolecules 2016, 6, 9. https://doi.org/10.3390/biom6010009
Bukar Maina M, Al-Hilaly YK, Serpell LC. Nuclear Tau and Its Potential Role in Alzheimer’s Disease. Biomolecules. 2016; 6(1):9. https://doi.org/10.3390/biom6010009
Chicago/Turabian StyleBukar Maina, Mahmoud, Youssra K. Al-Hilaly, and Louise C. Serpell. 2016. "Nuclear Tau and Its Potential Role in Alzheimer’s Disease" Biomolecules 6, no. 1: 9. https://doi.org/10.3390/biom6010009
APA StyleBukar Maina, M., Al-Hilaly, Y. K., & Serpell, L. C. (2016). Nuclear Tau and Its Potential Role in Alzheimer’s Disease. Biomolecules, 6(1), 9. https://doi.org/10.3390/biom6010009





