Notable Aspects of Glycan-Protein Interactions
Abstract
:1. Introduction
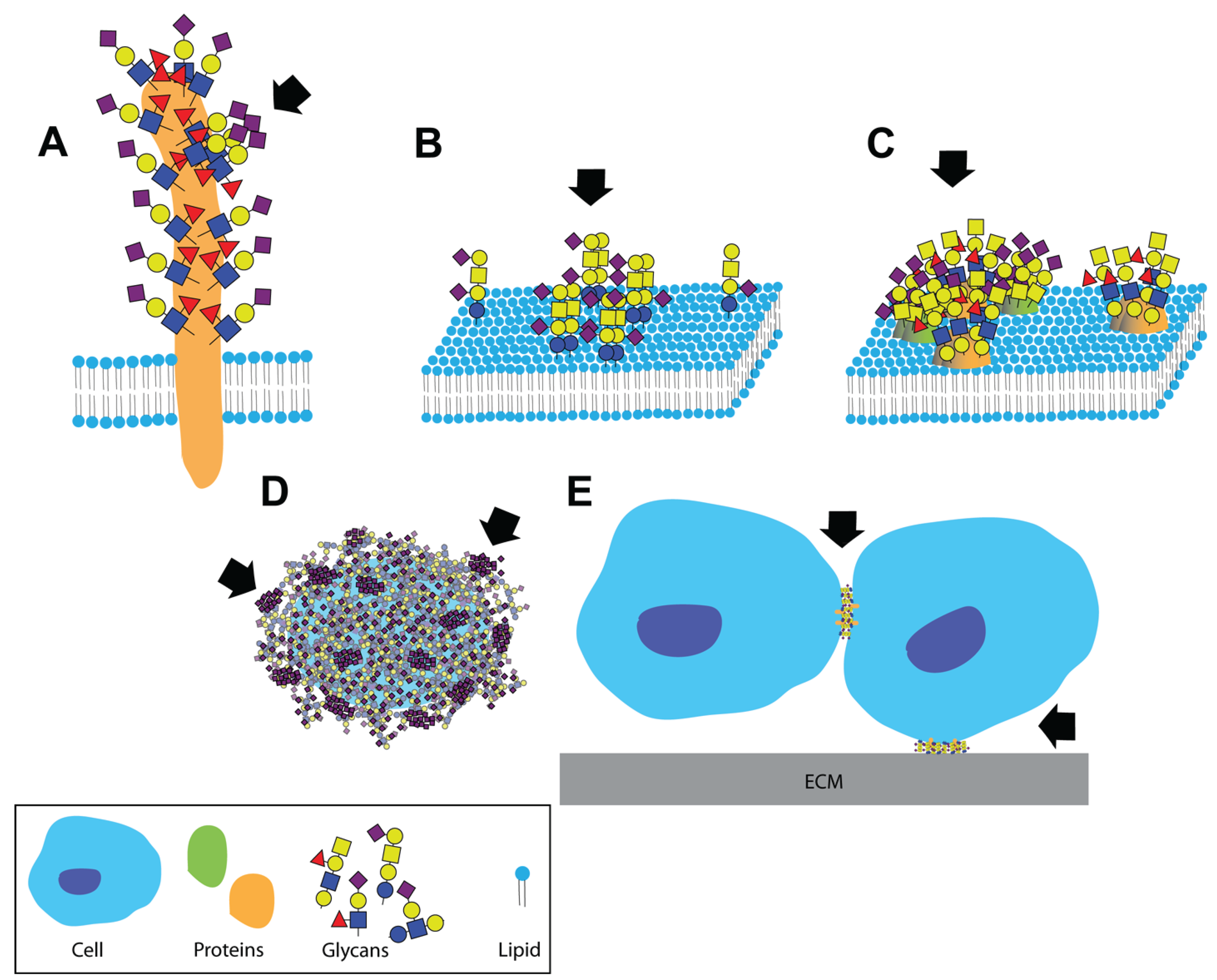
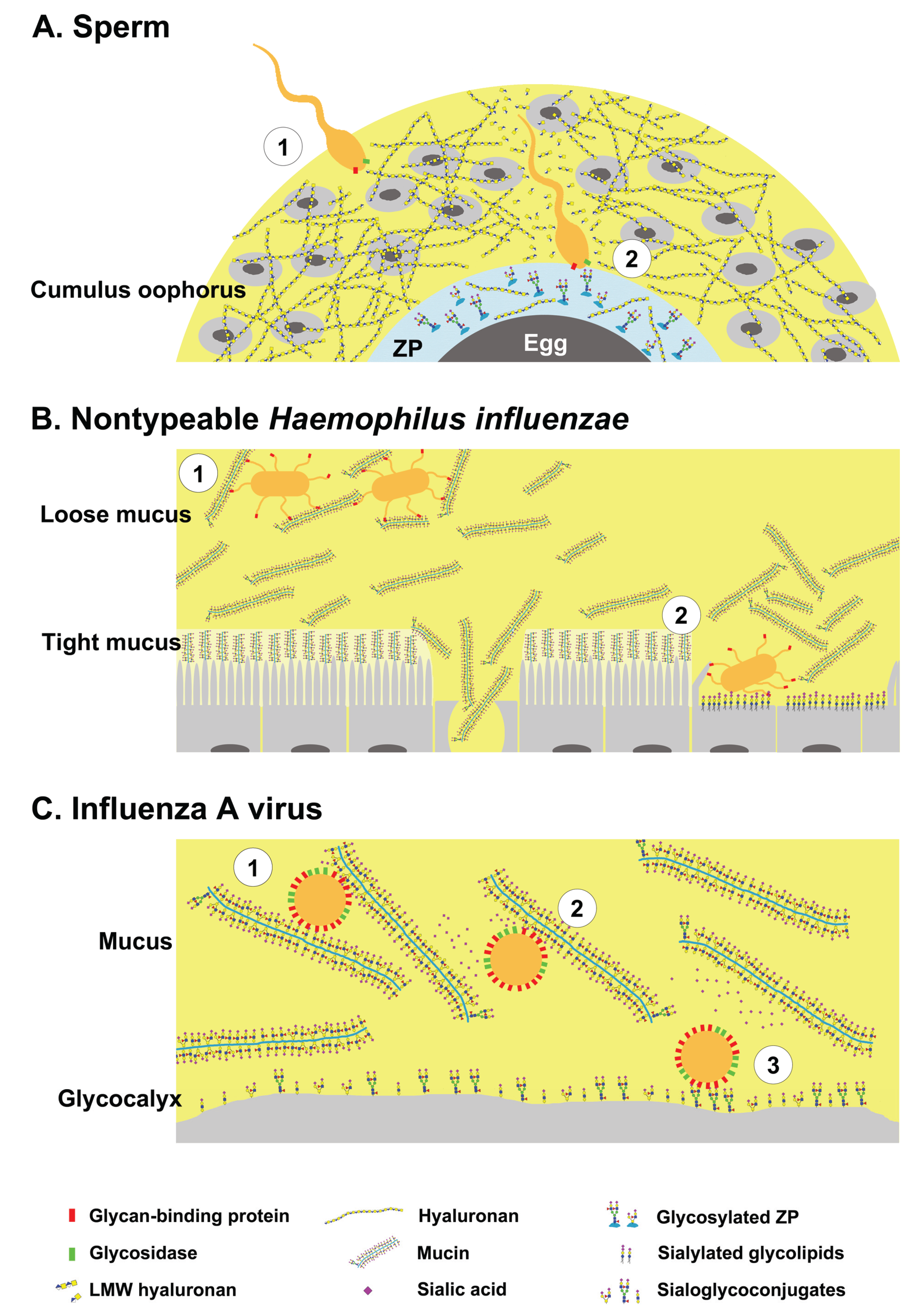

2. Dual Role of Glycans during Pathogen Invasion—Receptors and Decoys
2.1. Sperm-Egg Interactions
2.2. Nontypeable Haemophilus Influenzae and Host Mucins
2.3. Influenza A Viruses and Host Mucins
3. Glycan-Mediated Lateral Movement on Host Tissues
3.1. Selectin-Mediated Leukocyte Rolling
3.2. FimH Mediated Stick-and-Roll Adhesion
3.3. Heparan Sulfate Proteoglycans Mediated Surfacing
4. Concluding Remarks—Challenges in Investigating Glycan-Binding
Acknowledgments
Conflicts of Interest
References
- Varki, N.M.; Varki, A. Diversity in cell surface sialic acid presentations: Implications for biology and disease. Lab. Investig. 2007, 87, 851–857. [Google Scholar] [CrossRef] [PubMed]
- Varki, A. Evolutionary forces shaping the Golgi glycosylation machinery: Why cell surface glycans are universal to living cells. Cold Spring Harb. Perspect. Biol. 2011. [Google Scholar] [CrossRef] [PubMed]
- Marth, J.D. A unified vision of the building blocks of life. Nat. Cell Biol. 2008, 10, 1015–1016. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.; Varki, A. The sialome—Far more than the sum of its parts. OMICS 2010, 14, 455–464. [Google Scholar] [CrossRef] [PubMed]
- Varki, A.; Sharon, N. Historical background and overview. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2009; pp. 1–22. [Google Scholar]
- Cohen, M.; Varki, A. Modulation of glycan recognition by clustered saccharide patches. Int. Rev. Cell Mol. Biol. 2014, 308, 75–125. [Google Scholar] [PubMed]
- Varki, A. Selectin ligands. Proc. Natl. Acad. Sci. USA 1994, 91, 7390–7397. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.; Hurtado-Ziola, N.; Varki, A. ABO blood group glycans modulate sialic acid recognition on erythrocytes. Blood 2009, 114, 3668–3676. [Google Scholar] [CrossRef] [PubMed]
- Cambi, A.; Koopman, M.; Figdor, C.G. How C-type lectins detect pathogens. Cell. Microbiol. 2005, 7, 481–488. [Google Scholar] [CrossRef] [PubMed]
- McEver, R.P.; Zhu, C. Rolling cell adhesion. Annu. Rev. Cell Dev. Biol. 2010, 26, 363–396. [Google Scholar] [CrossRef] [PubMed]
- Bishop, J.R.; Gagneux, P. Evolution of carbohydrate antigens—Microbial forces shaping host glycomes? Glycobiology 2007, 17, 23R–34R. [Google Scholar] [CrossRef] [PubMed]
- Varki, A. Since there are PAMPs and DAMPs, there must be SAMPs? Glycan “self-associated molecular patterns” dampen innate immunity, but pathogens can mimic them. Glycobiology 2011, 21, 1121–1124. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, F.; Tiralongo, E.; Tiralongo, J. Sialic acid-specific lectins: Occurrence, specificity and function. Cell. Mol. Life Sci. 2006, 63, 1331–1354. [Google Scholar] [CrossRef] [PubMed]
- Angata, T.; Varki, A. Chemical diversity in the sialic acids and related alpha-keto acids: An evolutionary perspective. Chem. Rev. 2002, 102, 439–469. [Google Scholar] [CrossRef] [PubMed]
- Cone, R.A. Barrier properties of mucus. Adv. Drug Deliv. Rev. 2009, 61, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Fahy, J.V.; Dickey, B.F. Airway mucus function and dysfunction. N. Engl. J. Med. 2010, 363, 2233–2247. [Google Scholar] [CrossRef] [PubMed]
- Corfield, A.P. Mucins: A biologically relevant glycan barrier in mucosal protection. Biochim. Biophys. Acta 2015, 1850, 236–252. [Google Scholar] [CrossRef] [PubMed]
- Anifandis, G.; Messini, C.; Dafopoulos, K.; Sotiriou, S.; Messinis, I. Molecular and cellular mechanisms of sperm-oocyte interactions opinions relative to in vitro fertilization (IVF). Int. J. Mol. Sci. 2014, 15, 12972–12997. [Google Scholar] [CrossRef] [PubMed]
- Tecle, E.; Gagneux, P. Sugar-coated sperm: Unraveling the functions of the mammalian sperm glycocalyx. Mol. Reprod. Dev. 2015. [Google Scholar] [CrossRef] [PubMed]
- Suarez, S.S. Regulation of sperm storage and movement in the mammalian oviduct. Int. J. Dev. Biol. 2008, 52, 455–462. [Google Scholar] [CrossRef] [PubMed]
- Okabe, M. The cell biology of mammalian fertilization. Development 2013, 140, 4471–4479. [Google Scholar] [CrossRef] [PubMed]
- Vandevoort, C.A.; Cherr, G.N.; Overstreet, J.W. Hyaluronic acid enhances the zona pellucida-induced acrosome reaction of macaque sperm. J. Androl. 1997, 18, 1–5. [Google Scholar] [PubMed]
- Chiu, P.C.; Chung, M.K.; Koistinen, R.; Koistinen, H.; Seppala, M.; Ho, P.C.; Ng, E.H.; Lee, K.F.; Yeung, W.S. Cumulus oophorus-associated glycodelin-C displaces sperm-bound glycodelin-A and -F and stimulates spermatozoa-zona pellucida binding. J. Biol. Chem. 2007, 282, 5378–5388. [Google Scholar] [CrossRef] [PubMed]
- Sun, F.; Bahat, A.; Gakamsky, A.; Girsh, E.; Katz, N.; Giojalas, L.C.; Tur-Kaspa, I.; Eisenbach, M. Human sperm chemotaxis: Both the oocyte and its surrounding cumulus cells secrete sperm chemoattractants. Hum. Reprod. 2005, 20, 761–767. [Google Scholar] [CrossRef] [PubMed]
- Vasen, G.; Battistone, M.A.; Croci, D.O.; Brukman, N.G.; Weigel Munoz, M.; Stupirski, J.C.; Rabinovich, G.A.; Cuasnicu, P.S. The galectin-1-glycan axis controls sperm fertilizing capacity by regulating sperm motility and membrane hyperpolarization. FASEB J. 2015. [Google Scholar] [CrossRef]
- Clark, G.F. The role of carbohydrate recognition during human sperm-egg binding. Hum. Reprod. 2013, 28, 566–577. [Google Scholar] [CrossRef] [PubMed]
- Ranganathan, S.; Ganguly, A.K.; Datta, K. Evidence for presence of hyaluronan binding protein on spermatozoa and its possible involvement in sperm function. Mol. Reprod. Dev. 1994, 38, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Deb, T.B.; Datta, K. Molecular cloning of human fibroblast hyaluronic acid-binding protein confirms its identity with P-32, a protein co-purified with splicing factor SF2. Hyaluronic acid-binding protein as P-32 protein, co-purified with splicing factor SF2. J. Biol. Chem. 1996, 271, 2206–2212. [Google Scholar] [PubMed]
- Kornovski, B.S.; McCoshen, J.; Kredentser, J.; Turley, E. The regulation of sperm motility by a novel hyaluronan receptor. Fertil. Steril. 1994, 61, 935–940. [Google Scholar] [PubMed]
- Lin, Y.; Kimmel, L.H.; Myles, D.G.; Primakoff, P. Molecular cloning of the human and monkey sperm surface protein PH-20. Proc. Natl. Acad. Sci. USA 1993, 90, 10071–10075. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Mahan, K.; Lathrop, W.F.; Myles, D.G.; Primakoff, P. A hyaluronidase activity of the sperm plasma membrane protein PH-20 enables sperm to penetrate the cumulus cell layer surrounding the egg. J. Cell Biol. 1994, 125, 1157–1163. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.; Baba, D.; Kimura, M.; Yamashita, M.; Kashiwabara, S.; Baba, T. Identification of a hyaluronidase, Hyal5, involved in penetration of mouse sperm through cumulus mass. Proc. Natl. Acad. Sci. USA 2005, 102, 18028–18033. [Google Scholar] [CrossRef] [PubMed]
- Vines, C.A.; Li, M.W.; Deng, X.; Yudin, A.I.; Cherr, G.N.; Overstreet, J.W. Identification of a hyaluronic acid (HA) binding domain in the PH-20 protein that may function in cell signaling. Mol. Reprod. Dev. 2001, 60, 542–552. [Google Scholar] [CrossRef] [PubMed]
- Buffone, M.G.; Hirohashi, N.; Gerton, G.L. Unresolved questions concerning mammalian sperm acrosomal exocytosis. Biol. Reprod. 2014. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.; Yamashita, M.; Kimura, M.; Honda, A.; Kashiwabara, S.; Baba, T. Sperm penetration through cumulus mass and zona pellucida. Int. J. Dev. Biol. 2008, 52, 677–682. [Google Scholar] [CrossRef] [PubMed]
- Avella, M.A.; Baibakov, B.; Dean, J. A single domain of the ZP2 zona pellucida protein mediates gamete recognition in mice and humans. J. Cell Biol. 2014, 205, 801–809. [Google Scholar] [CrossRef] [PubMed]
- Pang, P.C.; Chiu, P.C.; Lee, C.L.; Chang, L.Y.; Panico, M.; Morris, H.R.; Haslam, S.M.; Khoo, K.H.; Clark, G.F.; Yeung, W.S.; et al. Human sperm binding is mediated by the sialyl-Lewis(x) oligosaccharide on the zona pellucida. Science 2011, 333, 1761–1764. [Google Scholar] [CrossRef] [PubMed]
- Clark, G.F. A role for carbohydrate recognition in mammalian sperm-egg binding. Biochem. Biophys. Res. Commun. 2014, 450, 1195–1203. [Google Scholar] [CrossRef] [PubMed]
- McGuckin, M.A.; Linden, S.K.; Sutton, P.; Florin, T.H. Mucin dynamics and enteric pathogens. Nat. Rev. Microbiol. 2011, 9, 265–278. [Google Scholar] [CrossRef] [PubMed]
- Moran, A.P.; Gupta, A.; Joshi, L. Sweet-talk: Role of host glycosylation in bacterial pathogenesis of the gastrointestinal tract. Gut 2011, 60, 1412–1425. [Google Scholar] [CrossRef] [PubMed]
- Nizet, V.; Esko, J.D. Bacterial and Viral Infections. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2009; pp. 537–554. [Google Scholar]
- Ouwerkerk, J.P.; de Vos, W.M.; Belzer, C. Glycobiome: Bacteria and mucus at the epithelial interface. Best Pract. Res. Clin. Gastroenterol. 2013, 27, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Voynow, J.A.; Gendler, S.J.; Rose, M.C. Regulation of mucin genes in chronic inflammatory airway diseases. Am. J. Respir. Cell Mol. Biol. 2006, 34, 661–665. [Google Scholar] [CrossRef] [PubMed]
- Geme, J.W.R. Molecular and cellular determinants of non-typeable Haemophilus influenzae adherence and invasion. Cell. Microbiol. 2002, 4, 191–200. [Google Scholar] [CrossRef]
- Foxwell, A.R.; Kyd, J.M.; Cripps, A.W. Nontypeable Haemophilus influenzae: Pathogenesis and prevention. Microbiol. Mol. Biol. Rev. 1998, 62, 294–308. [Google Scholar] [PubMed]
- Davies, J.; Carlstedt, I.; Nilsson, A.K.; Hakansson, A.; Sabharwal, H.; van Alphen, L.; van Ham, M.; Svanborg, C. Binding of Haemophilus influenzae to purified mucins from the human respiratory tract. Infect. Immun. 1995, 63, 2485–2492. [Google Scholar] [PubMed]
- Bernstein, J.M.; Reddy, M. Bacteria-mucin interaction in the upper aerodigestive tract shows striking heterogeneity: Implications in otitis media, rhinosinusitis, and pneumonia. Otolaryngol. Head Neck Surg. 2000, 122, 514–520. [Google Scholar] [PubMed]
- Severi, E.; Hood, D.W.; Thomas, G.H. Sialic acid utilization by bacterial pathogens. Microbiology 2007, 153, 2817–2822. [Google Scholar] [CrossRef] [PubMed]
- Varki, A.; Gagneux, P. Multifarious roles of sialic acids in immunity. Ann. NY Acad. Sci. 2012, 1253, 16–36. [Google Scholar] [CrossRef] [PubMed]
- Carlin, A.F.; Uchiyama, S.; Chang, Y.C.; Lewis, A.L.; Nizet, V.; Varki, A. Molecular mimicry of host sialylated glycans allows a bacterial pathogen to engage neutrophil Siglec-9 and dampen the innate immune response. Blood 2009, 113, 3333–3336. [Google Scholar] [CrossRef] [PubMed]
- Berenson, C.S.; Sayles, K.B.; Huang, J.; Reinhold, V.N.; Garlipp, M.A.; Yohe, H.C. Nontypeable Haemophilus influenzae-binding gangliosides of human respiratory (HEp-2) cells have a requisite lacto/neolacto core structure. FEMS Immunol. Med. Microbiol. 2005, 45, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Van Alphen, L.; Geelen-van den Broek, L.; Blaas, L.; van Ham, M.; Dankert, J. Blocking of fimbria-mediated adherence of Haemophilus influenzae by sialyl gangliosides. Infect. Immun. 1991, 59, 4473–4477. [Google Scholar] [PubMed]
- Noel, G.J.; Love, D.C.; Mosser, D.M. High-molecular-weight proteins of nontypeable Haemophilus influenzae mediate bacterial adhesion to cellular proteoglycans. Infect. Immun. 1994, 62, 4028–4033. [Google Scholar] [PubMed]
- Vimr, E.; Lichtensteiger, C. To sialylate, or not to sialylate: That is the question. Trends Microbiol. 2002, 10, 254–257. [Google Scholar] [CrossRef]
- Bailey, K.L.; LeVan, T.D.; Yanov, D.A.; Pavlik, J.A.; DeVasure, J.M.; Sisson, J.H.; Wyatt, T.A. Non-typeable Haemophilus influenzae decreases cilia beating via protein kinase Cepsilon. Respir. Res. 2012. [Google Scholar] [CrossRef] [PubMed]
- Shukla, D.; Spear, P.G. Herpesviruses and heparan sulfate: An intimate relationship in aid of viral entry. J. Clin. Investig. 2001, 108, 503–510. [Google Scholar] [CrossRef] [PubMed]
- Kato, K.; Ishiwa, A. The role of carbohydrates in infection strategies of enteric pathogens. Trop. Med. Health 2015, 43, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y. Sialobiology of influenza: Molecular mechanism of host range variation of influenza viruses. Biol. Pharm. Bull. 2005, 28, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Taylor, H.P.; Armstrong, S.J.; Dimmock, N.J. Quantitative relationships between an influenza virus and neutralizing antibody. Virology 1987, 159, 288–298. [Google Scholar] [CrossRef]
- Markovic, I.; Leikina, E.; Zhukovsky, M.; Zimmerberg, J.; Chernomordik, L.V. Synchronized activation and refolding of influenza hemagglutinin in multimeric fusion machines. J. Cell Biol. 2001, 155, 833–844. [Google Scholar] [CrossRef] [PubMed]
- Harris, A.; Cardone, G.; Winkler, D.C.; Heymann, J.B.; Brecher, M.; White, J.M.; Steven, A.C. Influenza virus pleiomorphy characterized by cryoelectron tomography. Proc. Natl. Acad. Sci. USA 2006, 103, 19123–19127. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.; Zhang, X.Q.; Senaati, H.P.; Chen, H.W.; Varki, N.M.; Schooley, R.T.; Gagneux, P. Influenza A penetrates host mucus by cleaving sialic acids with neuraminidase. Virol. J. 2013. [Google Scholar] [CrossRef] [PubMed]
- Tarp, M.A.; Clausen, H. Mucin-type O-glycosylation and its potential use in drug and vaccine development. Biochim. Biophys. Acta 2008, 1780, 546–563. [Google Scholar] [CrossRef] [PubMed]
- Wohlbold, T.J.; Krammer, F. In the shadow of hemagglutinin: A growing interest in influenza viral neuraminidase and its role as a vaccine antigen. Viruses 2014, 6, 2465–2494. [Google Scholar] [CrossRef] [PubMed]
- Hanaoka, K.; Pritchett, T.J.; Takasaki, S.; Kochibe, N.; Sabesan, S.; Paulson, J.C.; Kobata, A. 4-O-acetyl-N-acetylneuraminic acid in the N-linked carbohydrate structures of equine and guinea pig alpha 2-macroglobulins, potent inhibitors of influenza virus infection. J. Biol. Chem. 1989, 264, 9842–9849. [Google Scholar] [PubMed]
- De Vries, E.; de Vries, R.P.; Wienholts, M.J.; Floris, C.E.; Jacobs, M.S.; van den Heuvel, A.; Rottier, P.J.; de Haan, C.A. Influenza A virus entry into cells lacking sialylated N-glycans. Proc. Natl. Acad. Sci. USA 2012, 109, 7457–7462. [Google Scholar] [CrossRef] [PubMed]
- Byrd-Leotis, L.; Liu, R.; Bradley, K.C.; Lasanajak, Y.; Cummings, S.F.; Song, X.; Heimburg-Molinaro, J.; Galloway, S.E.; Culhane, M.R.; Smith, D.F.; et al. Shotgun glycomics of pig lung identifies natural endogenous receptors for influenza viruses. Proc. Natl. Acad. Sci. USA 2014, 111, E2241–E2250. [Google Scholar] [CrossRef] [PubMed]
- Thomas, W. Catch bonds in adhesion. Annu. Rev. Biomed. Eng. 2008, 10, 39–57. [Google Scholar] [CrossRef] [PubMed]
- Sherer, N.M.; Jin, J.; Mothes, W. Directional spread of surface-associated retroviruses regulated by differential virus-cell interactions. J. Virol. 2010, 84, 3248–3258. [Google Scholar] [CrossRef] [PubMed]
- Moore, K.L.; Patel, K.D.; Bruehl, R.E.; Li, F.; Johnson, D.A.; Lichenstein, H.S.; Cummings, R.D.; Bainton, D.F.; McEver, R.P. P-selectin glycoprotein ligand-1 mediates rolling of human neutrophils on P-selectin. J. Cell Biol. 1995, 128, 661–671. [Google Scholar] [CrossRef] [PubMed]
- Del Conde, I.; Shrimpton, C.N.; Thiagarajan, P.; Lopez, J.A. Tissue-factor-bearing microvesicles arise from lipid rafts and fuse with activated platelets to initiate coagulation. Blood 2005, 106, 1604–1611. [Google Scholar] [CrossRef] [PubMed]
- Steegmaier, M.; Borges, E.; Berger, J.; Schwarz, H.; Vestweber, D. The E-selectin-ligand ESL-1 is located in the Golgi as well as on microvilli on the cell surface. J. Cell Sci. 1997, 110, 687–694. [Google Scholar] [PubMed]
- Von Andrian, U.H.; Hasslen, S.R.; Nelson, R.D.; Erlandsen, S.L.; Butcher, E.C. A central role for microvillous receptor presentation in leukocyte adhesion under flow. Cell 1995, 82, 989–999. [Google Scholar] [CrossRef]
- Zarbock, A.; Ley, K.; McEver, R.P.; Hidalgo, A. Leukocyte ligands for endothelial selectins: Specialized glycoconjugates that mediate rolling and signaling under flow. Blood 2011, 118, 6743–6751. [Google Scholar] [CrossRef] [PubMed]
- McEver, R.P.; Cummings, R.D. Role of PSGL-1 binding to selectins in leukocyte recruitment. J. Clin. Investig. 1997, 100, S97–S103. [Google Scholar] [PubMed]
- Moore, K.L.; Stults, N.L.; Diaz, S.; Smith, D.F.; Cummings, R.D.; Varki, A.; McEver, R.P. Identification of a specific glycoprotein ligand for P-selectin (CD62) on myeloid cells. J. Cell Biol. 1992, 118, 445–456. [Google Scholar] [CrossRef] [PubMed]
- Moore, K.L. Structure and function of P-selectin glycoprotein ligand-1. Leuk. Lymphoma 1998, 29, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, P.P.; McEver, R.P.; Cummings, R.D. Structures of the O-glycans on P-selectin glycoprotein ligand-1 from HL-60 cells. J. Biol. Chem. 1996, 271, 18732–18742. [Google Scholar] [CrossRef] [PubMed]
- Pouyani, T.; Seed, B. PSGL-1 recognition of P-selectin is controlled by a tyrosine sulfation consensus at the PSGL-1 amino terminus. Cell 1995, 83, 333–343. [Google Scholar] [CrossRef]
- Von Andrian, U.H.; Mempel, T.R. Homing and cellular traffic in lymph nodes. Nat. Rev. Immunol. 2003, 3, 867–878. [Google Scholar] [CrossRef] [PubMed]
- McEver, R.P. A sulfated address for lymphocyte homing. Nat. Immunol. 2005, 6, 1067–1069. [Google Scholar] [CrossRef] [PubMed]
- Park, E.Y.; Smith, M.J.; Stropp, E.S.; Snapp, K.R.; di Vietro, J.A.; Walker, W.F.; Schmidtke, D.W.; Diamond, S.L.; Lawrence, M.B. Comparison of PSGL-1 microbead and neutrophil rolling: Microvillus elongation stabilizes P-selectin bond clusters. Biophys. J. 2002, 82, 1835–1847. [Google Scholar] [CrossRef]
- Mehta, P.; Cummings, R.D.; McEver, R.P. Affinity and kinetic analysis of P-selectin binding to P-selectin glycoprotein ligand-1. J. Biol. Chem. 1998, 273, 32506–32513. [Google Scholar] [CrossRef] [PubMed]
- Beachey, E.H. Bacterial adherence: Adhesin-receptor interactions mediating the attachment of bacteria to mucosal surface. J. Infect. Dis. 1981, 143, 325–345. [Google Scholar] [CrossRef] [PubMed]
- Hahn, E.; Wild, P.; Hermanns, U.; Sebbel, P.; Glockshuber, R.; Haner, M.; Taschner, N.; Burkhard, P.; Aebi, U.; Muller, S.A. Exploring the 3D molecular architecture of Escherichia coli type 1 pili. J. Mol. Biol. 2002, 323, 845–857. [Google Scholar] [CrossRef]
- Air, G.M.; Laver, W.G. The neuraminidase of influenza virus. Proteins 1989, 6, 341–356. [Google Scholar] [CrossRef] [PubMed]
- Taganna, J.; de Boer, A.R.; Wuhrer, M.; Bouckaert, J. Glycosylation changes as important factors for the susceptibility to urinary tract infection. Biochem. Soc. Trans. 2011, 39, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Serafini-Cessi, F.; Monti, A.; Cavallone, D. N-Glycans carried by Tamm-Horsfall glycoprotein have a crucial role in the defense against urinary tract diseases. Glycoconj. J. 2005, 22, 383–394. [Google Scholar] [CrossRef] [PubMed]
- Thomas, W.E.; Nilsson, L.M.; Forero, M.; Sokurenko, E.V.; Vogel, V. Shear-dependent “stick-and-roll” adhesion of type 1 fimbriated Escherichia coli. Mol. Microbiol. 2004, 53, 1545–1557. [Google Scholar] [CrossRef] [PubMed]
- Sokurenko, E.V.; Vogel, V.; Thomas, W.E. Catch-bond mechanism of force-enhanced adhesion: Counterintuitive, elusive, but… widespread? Cell Host Microbe 2008, 4, 314–323. [Google Scholar] [CrossRef] [PubMed]
- Aprikian, P.; Interlandi, G.; Kidd, B.A.; le Trong, I.; Tchesnokova, V.; Yakovenko, O.; Whitfield, M.J.; Bullitt, E.; Stenkamp, R.E.; Thomas, W.E.; et al. The bacterial fimbrial tip acts as a mechanical force sensor. PLoS Biol. 2011, 9, e1000617. [Google Scholar] [CrossRef] [PubMed]
- Esko, J.D.; Kimata, K.; Lindahl, U. Proteoglycans and Sulfated Glycosaminoglycans. In Essentials of Glycobiology, 2nd Ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2009; pp. 229–248. [Google Scholar]
- Ugolini, S.; Mondor, I.; Sattentau, Q.J. HIV-1 attachment: Another look. Trends Microbiol. 1999, 7, 144–149. [Google Scholar] [CrossRef]
- Walker, S.J.; Pizzato, M.; Takeuchi, Y.; Devereux, S. Heparin binds to murine leukemia virus and inhibits Env-independent attachment and infection. J. Virol. 2002, 76, 6909–6918. [Google Scholar] [CrossRef] [PubMed]
- Connell, B.J.; Lortat-Jacob, H. Human immunodeficiency virus and heparan sulfate: From attachment to entry inhibition. Front. Immunol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Schowalter, R.M.; Pastrana, D.V.; Buck, C.B. Glycosaminoglycans and sialylated glycans sequentially facilitate Merkel cell polyomavirus infectious entry. PLoS Pathog. 2011, 7, e1002161. [Google Scholar] [CrossRef] [PubMed]
- Tsai, B.; Gilbert, J.M.; Stehle, T.; Lencer, W.; Benjamin, T.L.; Rapoport, T.A. Gangliosides are receptors for murine polyoma virus and SV40. EMBO J. 2003, 22, 4346–4355. [Google Scholar] [CrossRef] [PubMed]
- Sapp, M.; Day, P.M. Structure, attachment and entry of polyoma- and papillomaviruses. Virology 2009, 384, 400–409. [Google Scholar] [CrossRef] [PubMed]
- Schelhaas, M.; Ewers, H.; Rajamaki, M.L.; Day, P.M.; Schiller, J.T.; Helenius, A. Human papillomavirus type 16 entry: Retrograde cell surface transport along actin-rich protrusions. PLoS Pathog. 2008, 4, e1000148. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, M.J.; Sherer, N.M.; Marks, C.B.; Pypaert, M.; Mothes, W. Actin- and myosin-driven movement of viruses along filopodia precedes their entry into cells. J. Cell Biol. 2005, 170, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Lakadamyali, M.; Rust, M.J.; Babcock, H.P.; Zhuang, X. Visualizing infection of individual influenza viruses. Proc. Natl. Acad. Sci. USA 2003, 100, 9280–9285. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.L.; Lidke, D.S.; Ozbun, M.A. Virus activated filopodia promote human papillomavirus type 31 uptake from the extracellular matrix. Virology 2008, 381, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Ewers, H.; Smith, A.E.; Sbalzarini, I.F.; Lilie, H.; Koumoutsakos, P.; Helenius, A. Single-particle tracking of murine polyoma virus-like particles on live cells and artificial membranes. Proc. Natl. Acad. Sci. USA 2005, 102, 15110–15115. [Google Scholar] [CrossRef] [PubMed]
- Laughlin, S.T.; Bertozzi, C.R. Imaging the glycome. Proc. Natl. Acad. Sci. USA 2009, 106, 12–17. [Google Scholar] [CrossRef] [PubMed]
- Chuh, K.N.; Pratt, M.R. Chemistry-enabled methods for the visualization of cell-surface glycoproteins in Metazoans. Glycoconj. J. 2015. [Google Scholar] [CrossRef] [PubMed]
- Narla, S.N.; Nie, H.; Li, Y.; Sun, X.L. Multi-dimensional glycan microarrays with glyco-macroligands. Glycoconj. J. 2015. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.L.; Cohen, M.; Fisher, C.J.; Schooley, R.T.; Gagneux, P.; Godula, K. Determination of receptor specificities for whole influenza viruses using multivalent glycan arrays. Chem. Commun. 2015, 51, 5326–5329. [Google Scholar] [CrossRef] [PubMed]
- Arthur, C.M.; Cummings, R.D.; Stowell, S.R. Using glycan microarrays to understand immunity. Curr. Opin. Chem. Biol. 2014, 18, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Lasanajak, Y.; Xia, B.; Heimburg-Molinaro, J.; Rhea, J.M.; Ju, H.; Zhao, C.; Molinaro, R.J.; Cummings, R.D.; Smith, D.F. Shotgun glycomics: A microarray strategy for functional glycomics. Nat. Methods 2011, 8, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.Y.; Holtz, B.; Wang, Y.; Wang, L.X.; Orndorff, P.E.; Guo, A. Quantitative glycomics from fluidic glycan microarrays. J. Am. Chem. Soc. 2009, 131, 13646–13650. [Google Scholar] [CrossRef] [PubMed]
- Palma, A.S.; Feizi, T.; Childs, R.A.; Chai, W.; Liu, Y. The neoglycolipid (NGL)-based oligosaccharide microarray system poised to decipher the meta-glycome. Curr. Opin. Chem. Biol. 2014, 18, 87–94. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cohen, M. Notable Aspects of Glycan-Protein Interactions. Biomolecules 2015, 5, 2056-2072. https://doi.org/10.3390/biom5032056
Cohen M. Notable Aspects of Glycan-Protein Interactions. Biomolecules. 2015; 5(3):2056-2072. https://doi.org/10.3390/biom5032056
Chicago/Turabian StyleCohen, Miriam. 2015. "Notable Aspects of Glycan-Protein Interactions" Biomolecules 5, no. 3: 2056-2072. https://doi.org/10.3390/biom5032056
APA StyleCohen, M. (2015). Notable Aspects of Glycan-Protein Interactions. Biomolecules, 5(3), 2056-2072. https://doi.org/10.3390/biom5032056




