Initial Events in Bacterial Transcription Initiation
Abstract
:1. Introduction
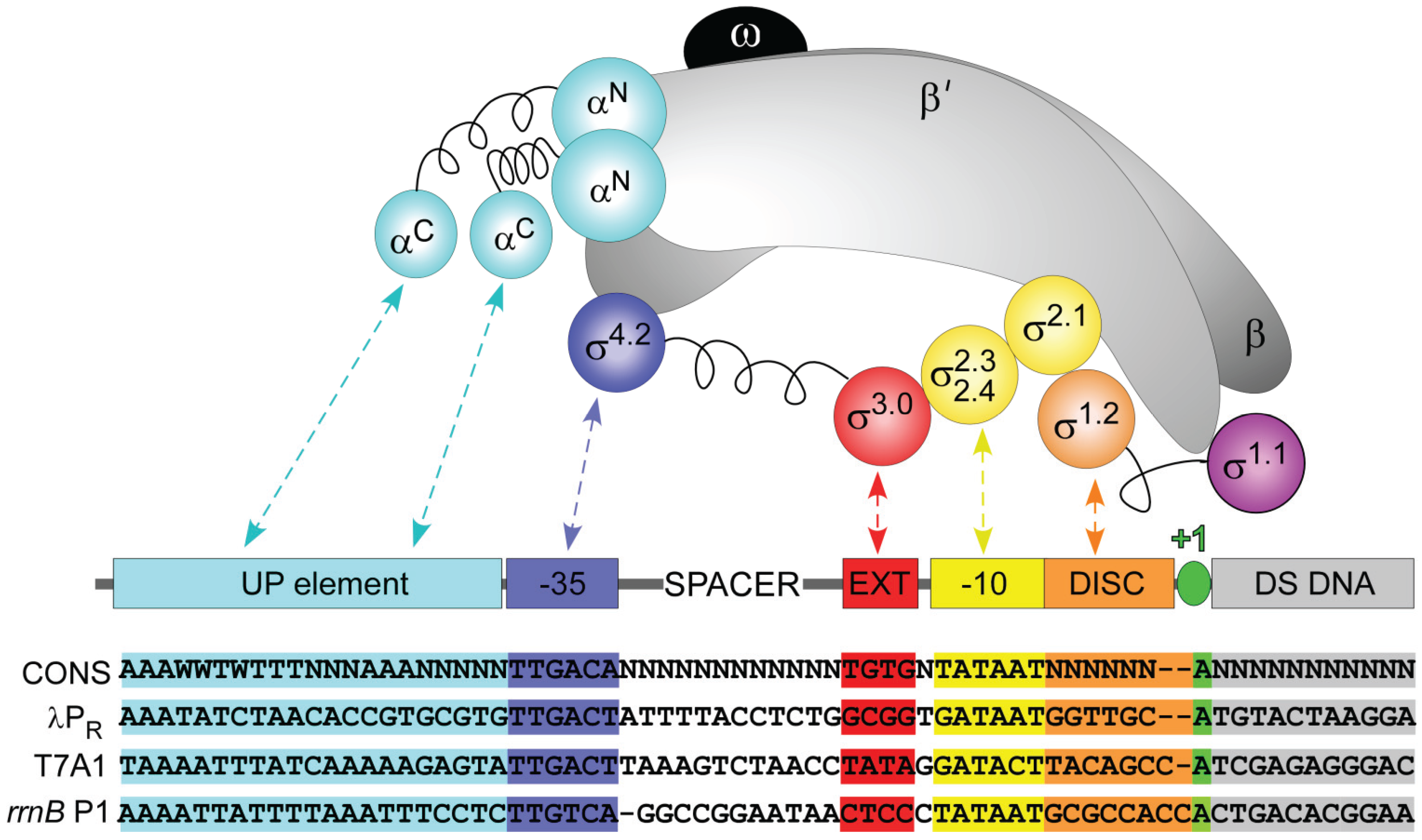
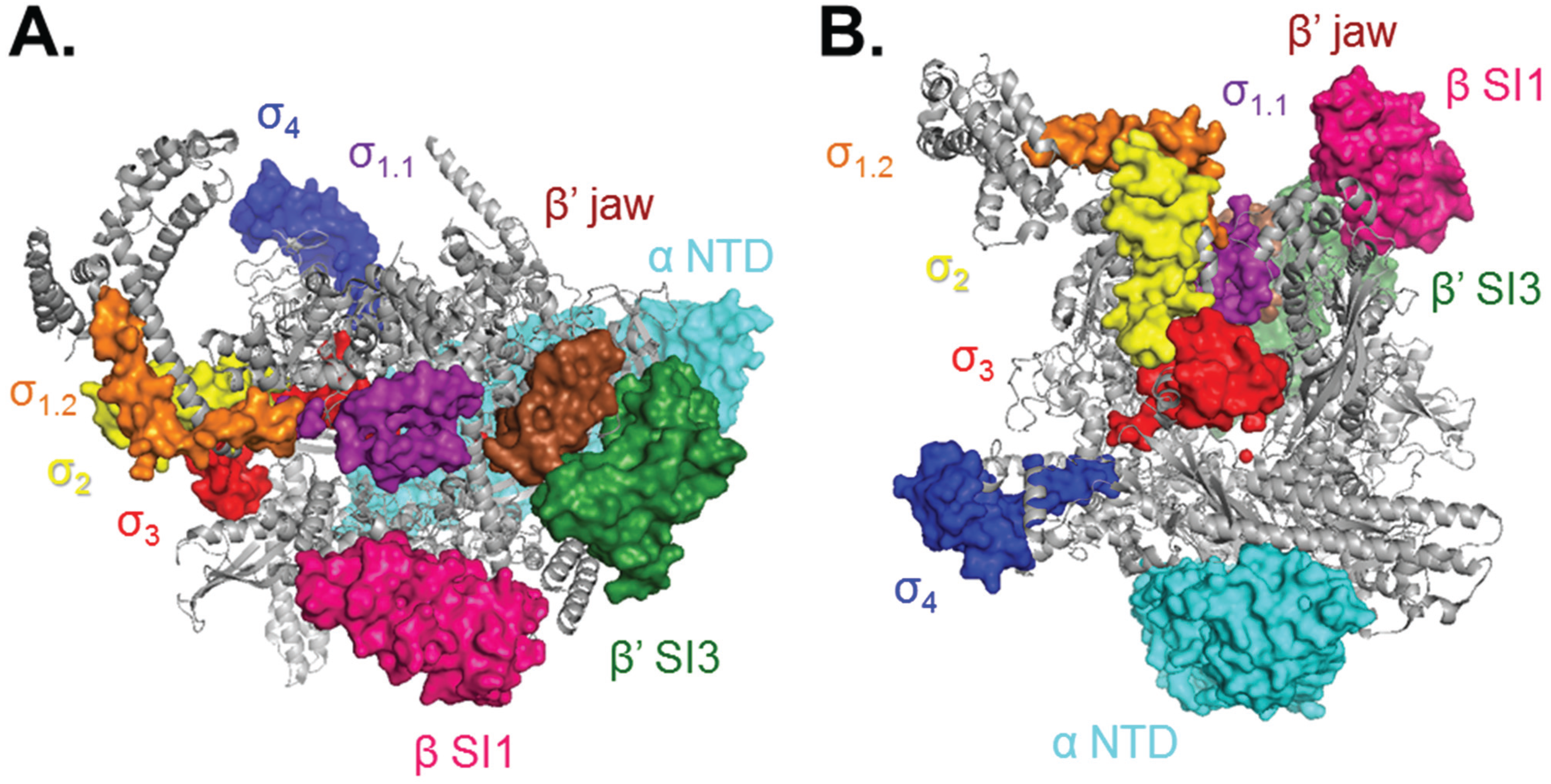
2. Bacterial RNAP σ70 Holoenzyme
3. E. coli σ70 Promoter Recognition
4. Initial Binding and Subsequent Conformational Changes to Form and Stabilize the Open Complex at Eσ70 Promoters
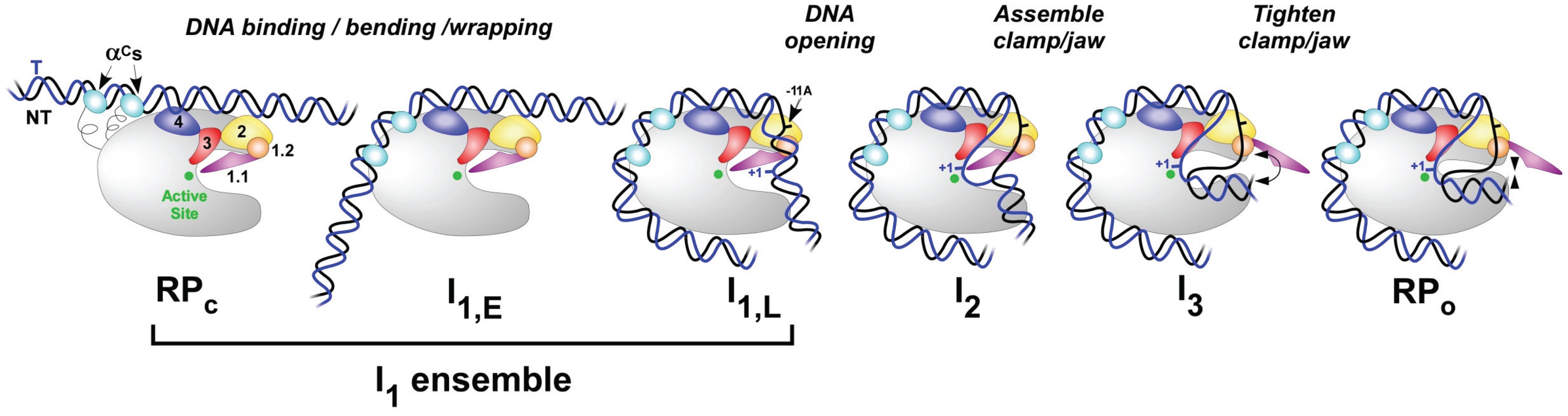
4.1. The Promoter Search and Initial Specific Binding to Form RPC
4.2. Upstream Bending and Wrapping to Form More Advanced Closed Complexes
4.3. Bending the Downstream Duplex into the Cleft
4.4. DNA Opening and Closing in the Cleft
4.5. Open Complex Stabilization
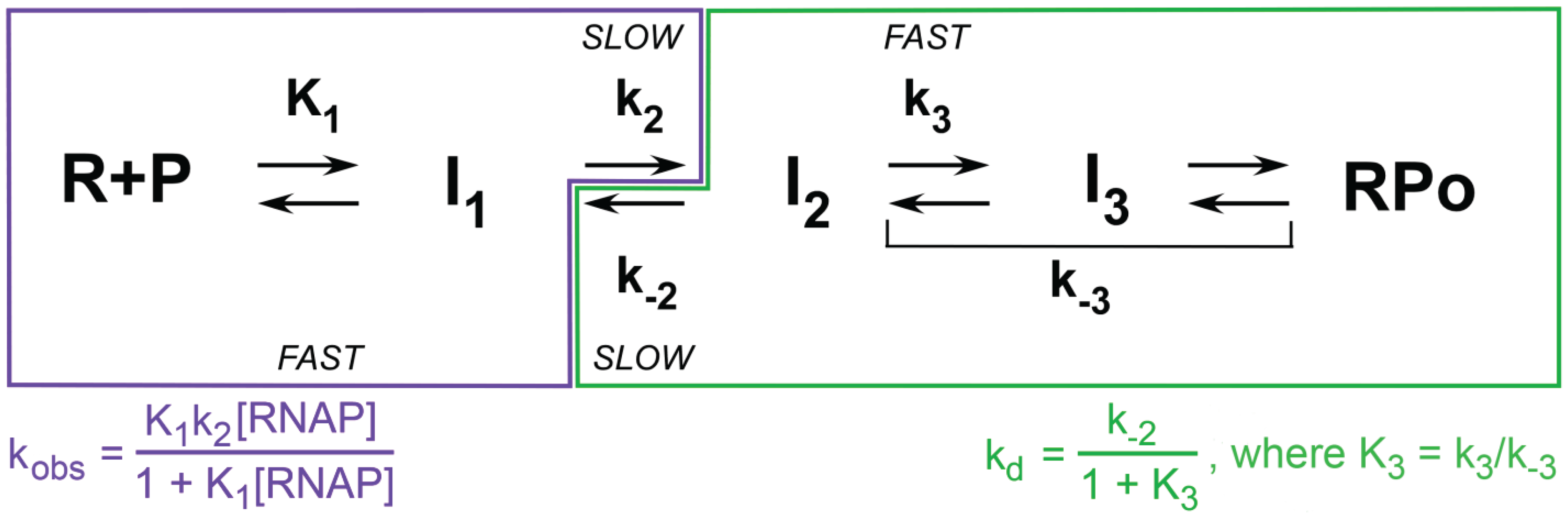
5. Forming and Stabilizing the Open Complex; Regulation of the Kinetics by Promoter Sequence, Length and Other Variables
5.1. Kinetics and Mechanism of Open Complex Formation
5.2. Kinetics of Open Complex Formation for λPR and λPR Variants: Interpretation of kobs
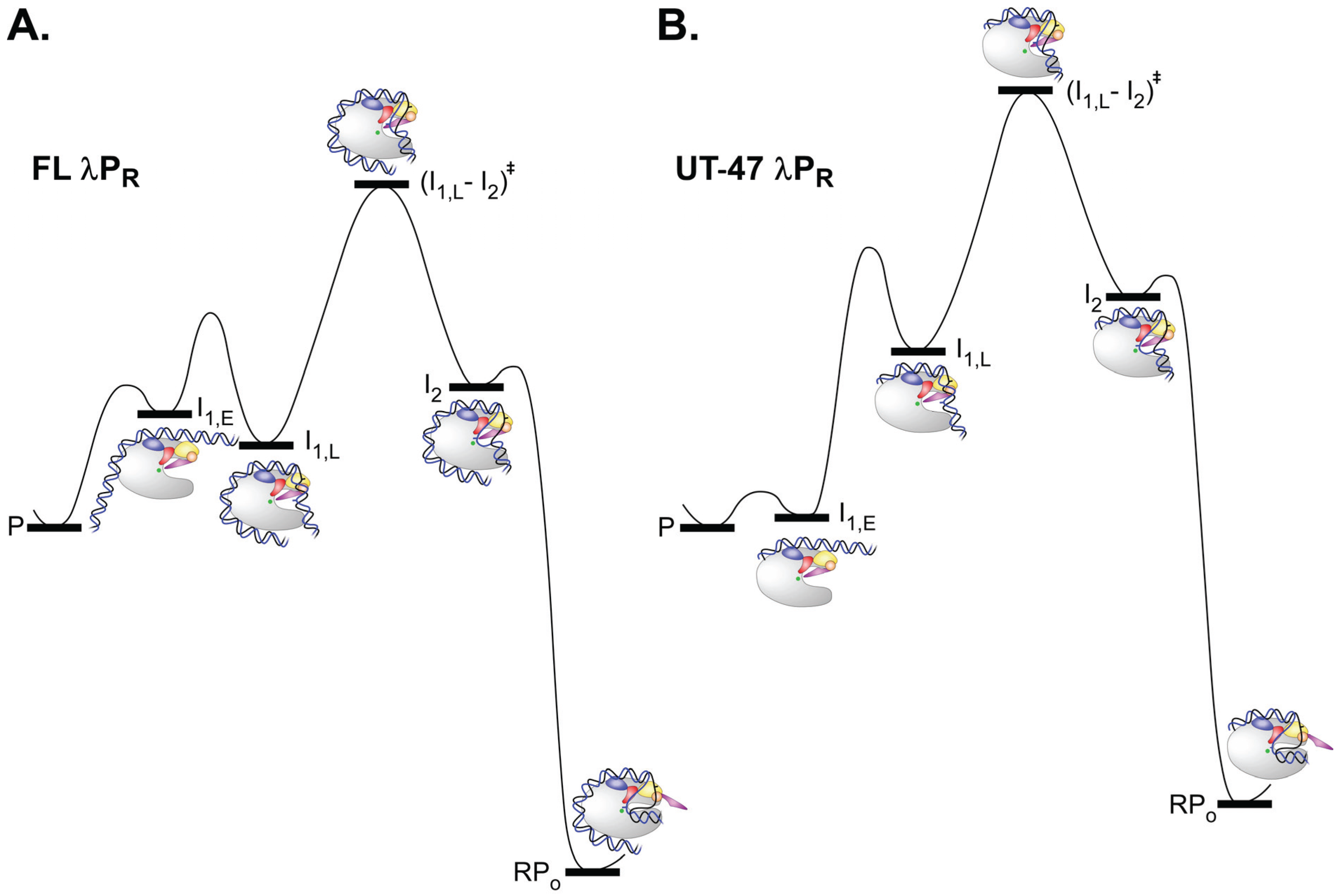
5.3. Kinetics and Mechanism of Open Complex Dissociation
5.4. Determining the DNA Closing Rate Constant (k-2) and the Stabilization (K3) of the Initial Open Complex
5.5. Fundamental Similarities of the RNAP-Promoter Mechanism to a Mechanism of Enzyme Catalysis; Implications for Regulation of Open Complex Formation and Lifetime
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Decker, K.B.; Hinton, D.M. Transcription regulation at the core: Similarities among bacterial, archaeal, and eukaryotic RNA polymerases. Annu. Rev. Microbiol. 2013, 67, 113–139. [Google Scholar] [CrossRef] [PubMed]
- Ebright, R.H. RNA polymerase: Structural similarities between bacterial RNA polymerase and eukaryotic RNA polymerase II. J. Mol. Biol. 2000, 304, 687–698. [Google Scholar] [CrossRef] [PubMed]
- Saecker, R.M.; Record, M.T., Jr.; Dehaseth, P.L. Mechanism of bacterial transcription initiation: RNA polymerase—Promoter binding, isomerization to initiation-competent open complexes, and initiation of RNA synthesis. J. Mol. Biol. 2011, 412, 754–771. [Google Scholar] [CrossRef] [PubMed]
- Dorman, C.J. Co-operative roles for DNA supercoiling and nucleoid-associated proteins in the regulation of bacterial transcription. Biochem. Soc. Trans. 2013, 41, 542–547. [Google Scholar] [CrossRef] [PubMed]
- Feklistov, A.; Sharon, B.D.; Darst, S.A.; Gross, C.A. Bacterial sigma factors: A historical, structural, and genomic perspective. Annu. Rev. Microbiol. 2014, 68, 357–376. [Google Scholar] [CrossRef] [PubMed]
- Haugen, S.P.; Ross, W.; Gourse, R.L. Advances in bacterial promoter recognition and its control by factors that do not bind DNA. Nat. Rev. Microbiol. 2008, 6, 507–519. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.J.; Minchin, S.D.; Busby, S.J. Activating transcription in bacteria. Annu. Rev. Microbiol. 2012, 66, 125–152. [Google Scholar] [CrossRef] [PubMed]
- Bae, B.; Davis, E.; Brown, D.; Campbell, E.A.; Wigneshweraraj, S.; Darst, S.A. Phage T7 Gp2 inhibition of Escherichia coli RNA polymerase involves misappropriation of σ70 domain 1.1. Proc. Natl. Acad. Sci. USA 2013, 110, 19772–19777. [Google Scholar] [CrossRef] [PubMed]
- Basu, R.S.; Warner, B.A.; Molodtsov, V.; Pupov, D.; Esyunina, D.; Fernandez-Tornero, C.; Kulbachinskiy, A.; Murakami, K.S. Structural basis of transcription initiation by bacterial RNA polymerase holoenzyme. J. Biol. Chem. 2014, 289, 24549–24559. [Google Scholar] [CrossRef] [PubMed]
- Feklistov, A.; Darst, S.A. Structural basis for promoter-10 element recognition by the bacterial RNA polymerase sigma subunit. Cell 2011, 147, 1257–1269. [Google Scholar] [CrossRef] [PubMed]
- Lawson, C.L.; Swigon, D.; Murakami, K.S.; Darst, S.A.; Berman, H.M.; Ebright, R.H. Catabolite activator protein: DNA binding and transcription activation. Curr. Opin. Struct. Biol. 2004, 14, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Murakami, K.S. X-ray crystal structure of Escherichia coli RNA polymerase σ70 holoenzyme. J. Biol. Chem. 2013, 288, 9126–9134. [Google Scholar] [CrossRef] [PubMed]
- Murakami, K.S.; Masuda, S.; Campbell, E.A.; Muzzin, O.; Darst, S.A. Structural basis of transcription initiation: An RNA polymerase holoenzyme-DNA complex. Science 2002, 296, 1285–1290. [Google Scholar] [CrossRef] [PubMed]
- Murakami, K.S.; Masuda, S.; Darst, S.A. Structural basis of transcription initiation: RNA polymerase holoenzyme at 4 Å resolution. Science 2002, 296, 1280–1284. [Google Scholar] [CrossRef] [PubMed]
- Murakami, K.S.; Masuda, S.; Darst, S.A. Crystallographic analysis of Thermus aquaticus RNA polymerase holoenzyme and a holoenzyme/promoter DNA complex. Methods Enzymol. 2003, 370, 42–53. [Google Scholar] [PubMed]
- Vassylyev, D.G.; Sekine, S.; Laptenko, O.; Lee, J.; Vassylyeva, M.N.; Borukhov, S.; Yokoyama, S. Crystal structure of a bacterial RNA polymerase holoenzyme at 2.6 Å resolution. Nature 2002, 417, 712–719. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Feng, Y.; Chatterjee, S.; Tuske, S.; Ho, M.X.; Arnold, E.; Ebright, R.H. Structural basis of transcription initiation. Science 2012, 338, 1076–1080. [Google Scholar] [CrossRef] [PubMed]
- Ruff, E.F.; Drennan, A.; Capp, M.W.; Poulos, M.A.; Artsimovitch, I.; Record, M.T., Jr. Escherichia coli RNA polymerase determinants of open complex lifetime and structure. J. Mol. Biol. In press. 2015. [Google Scholar]
- Rutherford, S.T. Insights into the Mechanism of DksA Action at Ribosomal RNA Promoters in Escherichia coli; University of Wisconsin: Madison, WI, USA, 2008. [Google Scholar]
- Rammohan, J.; Ruiz Manzano, A.; Garner, A.L.; Stallings, C.L.; Galburt, E.A. CarD stabilizes mycobacterial open complexes via a two-tiered kinetic mechanism. Nucleic Acids Res. 2015, 43, 3272–3285. [Google Scholar] [CrossRef] [PubMed]
- Friedman, L.J.; Gelles, J. Mechanism of transcription initiation at an activator-dependent promoter defined by single-molecule observation. Cell 2012, 148, 679–689. [Google Scholar] [CrossRef] [PubMed]
- Heyduk, E.; Heyduk, T. Next generation sequencing-based parallel analysis of melting kinetics of 4096 variants of a bacterial promoter. Biochemistry 2014, 53, 282–292. [Google Scholar] [CrossRef] [PubMed]
- Gries, T.J.; Kontur, W.S.; Capp, M.W.; Saecker, R.M.; Record, M.T., Jr. One-step DNA melting in the RNA polymerase cleft opens the initiation bubble to form an unstable open complex. Proc. Natl. Acad. Sci. USA 2010, 107, 10418–10423. [Google Scholar] [CrossRef] [PubMed]
- Kontur, W.S.; Saecker, R.M.; Capp, M.W.; Record, M.T., Jr. Late steps in the formation of E. coli RNA polymerase-λPR promoter open complexes: Characterization of conformational changes by rapid [perturbant] upshift experiments. J. Mol. Biol. 2008, 376, 1034–1047. [Google Scholar] [CrossRef] [PubMed]
- Rogozina, A.; Zaychikov, E.; Buckle, M.; Heumann, H.; Sclavi, B. DNA melting by RNA polymerase at the T7A1 promoter precedes the rate-limiting step at 37 degrees C and results in the accumulation of an off-pathway intermediate. Nucleic Acids Res. 2009, 37, 5390–5404. [Google Scholar] [CrossRef] [PubMed]
- Sclavi, B.; Zaychikov, E.; Rogozina, A.; Walther, F.; Buckle, M.; Heumann, H. Real-time characterization of intermediates in the pathway to open complex formation by Escherichia coli RNA polymerase at the T7A1 promoter. Proc. Natl. Acad. Sci. USA 2005, 102, 4706–4711. [Google Scholar] [CrossRef] [PubMed]
- Davis, C.A.; Bingman, C.A.; Landick, R.; Record, M.T., Jr.; Saecker, R.M. Real-time footprinting of DNA in the first kinetically significant intermediate in open complex formation by Escherichia coli RNA polymerase. Proc. Natl. Acad. Sci. USA 2007, 104, 7833–7838. [Google Scholar] [CrossRef] [PubMed]
- Ko, J.; Heyduk, T. Kinetics of promoter escape by bacterial RNA polymerase: Effects of promoter contacts and transcription bubble collapse. Biochem. J. 2014, 463, 135–144. [Google Scholar] [CrossRef] [PubMed]
- Mekler, V.; Minakhin, L.; Borukhov, S.; Mustaev, A.; Severinov, K. Coupling of downstream RNA polymerase-promoter interactions with formation of catalytically competent transcription initiation complex. J. Mol. Biol. 2014, 426, 3973–3984. [Google Scholar] [CrossRef] [PubMed]
- Mekler, V.; Minakhin, L.; Severinov, K. A critical role of downstream RNA polymerase-promoter interactions in the formation of initiation complex. J. Biol. Chem. 2011, 286, 22600–22608. [Google Scholar] [CrossRef] [PubMed]
- Mekler, V.; Pavlova, O.; Severinov, K. Interaction of Escherichia coli RNA polymerase σ70 subunit with promoter elements in the context of free σ70, RNA polymerase holoenzyme, and the β'-σ70 complex. J. Biol. Chem. 2011, 286, 270–279. [Google Scholar] [CrossRef] [PubMed]
- Revyakin, A.; Ebright, R.H.; Strick, T.R. Promoter unwinding and promoter clearance by RNA polymerase: Detection by single-molecule DNA nanomanipulation. Proc. Natl. Acad. Sci. USA 2004, 101, 4776–4780. [Google Scholar] [CrossRef] [PubMed]
- Revyakin, A.; Liu, C.; Ebright, R.H.; Strick, T.R. Abortive initiation and productive initiation by RNA polymerase involve DNA scrunching. Science 2006, 314, 1139–1143. [Google Scholar] [CrossRef] [PubMed]
- Robb, N.C.; Cordes, T.; Hwang, L.C.; Gryte, K.; Duchi, D.; Craggs, T.D.; Santoso, Y.; Weiss, S.; Ebright, R.H.; Kapanidis, A.N. The transcription bubble of the RNA polymerase-promoter open complex exhibits conformational heterogeneity and millisecond-scale dynamics: Implications for transcription start-site selection. J. Mol. Biol. 2013, 425, 875–885. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, L.A.; Gries, T.J.; Saecker, R.M.; Record, M.T., Jr.; Harris, M.E.; DeHaseth, P.L. Evidence for a tyrosine-adenine stacking interaction and for a short-lived open intermediate subsequent to initial binding of Escherichia coli RNA polymerase to promoter DNA. J. Mol. Biol. 2009, 385, 339–349. [Google Scholar] [CrossRef] [PubMed]
- Burgess, R.R. Separation and characterization of the subunits of ribonucleic acid polymerase. J. Biol. Chem. 1969, 244, 6168–6176. [Google Scholar] [PubMed]
- Murakami, K.S.; Darst, S.A. Bacterial RNA polymerases: The wholo story. Curr. Opin. Struct. Biol. 2003, 13, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Campbell, E.A.; Minakhin, L.; Richter, C.; Severinov, K.; Darst, S.A. Crystal structure of Thermus aquaticus core RNA polymerase at 3.3 Å resolution. Cell 1999, 98, 811–824. [Google Scholar] [CrossRef] [PubMed]
- Cramer, P.; Bushnell, D.A.; Kornberg, R.D. Structural basis of transcription: RNA polymerase II at 2.8 angstrom resolution. Science 2001, 292, 1863–1876. [Google Scholar] [CrossRef] [PubMed]
- Gnatt, A.L.; Cramer, P.; Fu, J.; Bushnell, D.A.; Kornberg, R.D. Structural basis of transcription: An RNA polymerase II elongation complex at 3.3 Å resolution. Science 2001, 292, 1876–1882. [Google Scholar] [CrossRef] [PubMed]
- Lane, W.J.; Darst, S.A. Molecular evolution of multisubunit RNA polymerases: Sequence analysis. J. Mol. Biol. 2010, 395, 671–685. [Google Scholar] [CrossRef] [PubMed]
- Gruber, T.M.; Gross, C.A. Multiple sigma subunits and the partitioning of bacterial transcription space. Annu. Rev. Microbiol. 2003, 57, 441–466. [Google Scholar] [CrossRef] [PubMed]
- Campbell, E.A.; Muzzin, O.; Chlenov, M.; Sun, J.L.; Olson, C.A.; Weinman, O.; Trester-Zedlitz, M.L.; Darst, S.A. Structure of the bacterial RNA polymerase promoter specificity sigma subunit. Mol. Cell 2002, 9, 527–539. [Google Scholar] [CrossRef] [PubMed]
- Keilty, S.; Rosenberg, M. Constitutive function of a positively regulated promoter reveals new sequences essential for activity. J. Biol. Chem. 1987, 262, 6389–6395. [Google Scholar] [PubMed]
- Schwartz, E.C.; Shekhtman, A.; Dutta, K.; Pratt, M.R.; Cowburn, D.; Darst, S.; Muir, T.W. A full-length group 1 bacterial sigma factor adopts a compact structure incompatible with DNA binding. Chem. Biol. 2008, 15, 1091–1103. [Google Scholar] [CrossRef] [PubMed]
- Dombroski, A.J.; Walter, W.A.; Gross, C.A. Amino-terminal amino acids modulate sigma-factor DNA-binding activity. Genes Dev. 1993, 7, 2446–2455. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.; Dombroski, A.J. Region 1 of σ70 is required for efficient isomerization and initiation of transcription by Escherichia coli RNA polymerase. J. Mol. Biol. 1997, 267, 60–74. [Google Scholar] [CrossRef] [PubMed]
- Darst, S.A. Bacterial RNA polymerase. Curr. Opin. Struct. Biol. 2001, 11, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, A.; Wang, D.; Ebright, Y.W.; Korlann, Y.; Kortkhonjia, E.; Kim, T.; Chowdhury, S.; Wigneshweraraj, S.; Irschik, H.; Jansen, R.; et al. Opening and closing of the bacterial RNA polymerase clamp. Science 2012, 337, 591–595. [Google Scholar] [CrossRef] [PubMed]
- Weixlbaumer, A.; Leon, K.; Landick, R.; Darst, S.A. Structural basis of transcriptional pausing in bacteria. Cell 2013, 152, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Tagami, S.; Sekine, S.; Kumarevel, T.; Hino, N.; Murayama, Y.; Kamegamori, S.; Yamamoto, M.; Sakamoto, K.; Yokoyama, S. Crystal structure of bacterial RNA polymerase bound with a transcription inhibitor protein. Nature 2010, 468, 978–982. [Google Scholar] [CrossRef] [PubMed]
- Ross, W.; Vrentas, C.E.; Sanchez-Vazquez, P.; Gaal, T.; Gourse, R.L. The magic spot: A ppGpp binding site on E. coli RNA polymerase responsible for regulation of transcription initiation. Mol. Cell 2013, 50, 420–429. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Y.; Wang, Y.; Steitz, T.A. The mechanism of E. coli RNA polymerase regulation by ppGpp is suggested by the structure of their complex. Mol. Cell 2013, 50, 430–436. [Google Scholar] [CrossRef] [PubMed]
- Belogurov, G.A.; Vassylyeva, M.N.; Sevostyanova, A.; Appleman, J.R.; Xiang, A.X.; Lira, R.; Webber, S.E.; Klyuyev, S.; Nudler, E.; Artsimovitch, I.; et al. Transcription inactivation through local refolding of the RNA polymerase structure. Nature 2009, 457, 332–335. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.X.; Hudson, B.P.; Das, K.; Arnold, E.; Ebright, R.H. Structures of RNA polymerase-antibiotic complexes. Curr. Opin. Struct. Biol. 2009, 19, 715–723. [Google Scholar] [CrossRef] [PubMed]
- Tupin, A.; Gualtieri, M.; Leonetti, J.P.; Brodolin, K. The transcription inhibitor lipiarmycin blocks DNA fitting into the RNA polymerase catalytic site. EMBO J. 2010, 29, 2527–2537. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Darst, S.A.; Thirumalai, D. Promoter melting triggered by bacterial RNA polymerase occurs in three steps. Proc. Natl. Acad. Sci. USA 2010, 107, 12523–12528. [Google Scholar] [CrossRef] [PubMed]
- Cowing, D.W.; Mecsas, J.; Record, M.T., Jr.; Gross, C.A. Intermediates in the formation of the open complex by RNA polymerase holoenzyme containing the sigma factor σ32 at the groE promoter. J. Mol. Biol. 1989, 210, 521–530. [Google Scholar] [CrossRef] [PubMed]
- Craig, M.L.; Tsodikov, O.V.; McQuade, K.L.; Schlax, P.E., Jr.; Capp, M.W.; Saecker, R.M.; Record, M.T., Jr. DNA footprints of the two kinetically significant intermediates in formation of an RNA polymerase-promoter open complex: Evidence that interactions with start site and downstream DNA induce sequential conformational changes in polymerase and DNA. J. Mol. Biol. 1998, 283, 741–756. [Google Scholar] [CrossRef] [PubMed]
- Saecker, R.M.; Tsodikov, O.V.; McQuade, K.L.; Schlax, P.E., Jr.; Capp, M.W.; Record, M.T., Jr. Kinetic studies and structural models of the association of E. coli σ70 RNA polymerase with the λPR promoter: Large scale conformational changes in forming the kinetically significant intermediates. J. Mol. Biol. 2002, 319, 649–671. [Google Scholar] [CrossRef] [PubMed]
- McClure, W.R. Mechanism and control of transcription initiation in prokaryotes. Annu. Rev. Biochem. 1985, 54, 171–204. [Google Scholar] [CrossRef] [PubMed]
- Record, M.T., Jr.; Reznikoff, W.S.; Craig, M.L.; McQuade, K.L.; Schlax, P.J. Escherichia coli RNA polymerase (Eσ70), promoters, and the kinetics of the steps of transcription initiation. In Escherichia coli and Salmonella Cellular and Molecular Biology; Neidhardt, F.C., Ed.; ASM Press: Washington, DC, USA, 1996; pp. 792–821. [Google Scholar]
- Kontur, W.S.; Capp, M.W.; Gries, T.J.; Saecker, R.M.; Record, M.T., Jr. Probing DNA binding, DNA opening, and assembly of a downstream clamp/jaw in Escherichia coli RNA polymerase-λPR promoter complexes using salt and the physiological anion glutamate. Biochemistry 2010, 49, 4361–4373. [Google Scholar] [CrossRef] [PubMed]
- Kontur, W.S.; Saecker, R.M.; Davis, C.A.; Capp, M.W.; Record, M.T., Jr. Solute probes of conformational changes in open complex (RPo) formation by Escherichia coli RNA polymerase at the λPR promoter: Evidence for unmasking of the active site in the isomerization step and for large-scale coupled folding in the subsequent conversion to RPo. Biochemistry 2006, 45, 2161–2177. [Google Scholar] [CrossRef] [PubMed]
- Roe, J.H.; Burgess, R.R.; Record, M.T., Jr. Kinetics and mechanism of the interaction of Escherichia coli RNA polymerase with the λPR promoter. J. Mol. Biol. 1984, 176, 495–522. [Google Scholar] [CrossRef] [PubMed]
- Roe, J.H.; Burgess, R.R.; Record, M.T., Jr. Temperature dependence of the rate constants of the Escherichia coli RNA polymerase-λPR promoter interaction. Assignment of the kinetic steps corresponding to protein conformational change and DNA opening. J. Mol. Biol. 1985, 184, 441–453. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, W. Starting and stopping sequences for the RNA polymerase. In RNA Polymerase; Losick, R., Chamberlin, M.J., Eds.; Cold Spring Harbor Laboratory: Cold Spring Harbor, NY, USA, 1976; pp. 193–205. [Google Scholar]
- Gourse, R.L.; Ross, W.; Gaal, T. UPs and downs in bacterial transcription initiation: The role of the alpha subunit of RNA polymerase in promoter recognition. Mol. Microbiol. 2000, 37, 687–695. [Google Scholar] [CrossRef] [PubMed]
- Ross, W.; Gosink, K.K.; Salomon, J.; Igarashi, K.; Zou, C.; Ishihama, A.; Severinov, K.; Gourse, R.L. A third recognition element in bacterial promoters: DNA binding by the alpha subunit of RNA polymerase. Science 1993, 262, 1407–1413. [Google Scholar] [CrossRef] [PubMed]
- Estrem, S.T.; Gaal, T.; Ross, W.; Gourse, R.L. Identification of an UP element consensus sequence for bacterial promoters. Proc. Natl. Acad. Sci. USA 1998, 95, 9761–9766. [Google Scholar] [CrossRef] [PubMed]
- Rao, L.; Ross, W.; Appleman, J.A.; Gaal, T.; Leirmo, S.; Schlax, P.J.; Record, M.T., Jr.; Gourse, R.L. Factor independent activation of rrnB P1. An “extended” promoter with an upstream element that dramatically increases promoter strength. J. Mol. Biol. 1994, 235, 1421–1435. [Google Scholar] [CrossRef] [PubMed]
- Estrem, S.T.; Ross, W.; Gaal, T.; Chen, Z.W.; Niu, W.; Ebright, R.H.; Gourse, R.L. Bacterial promoter architecture: Subsite structure of UP elements and interactions with the carboxy-terminal domain of the RNA polymerase alpha subunit. Genes Dev. 1999, 13, 2134–2147. [Google Scholar] [CrossRef] [PubMed]
- Gaal, T.; Rao, L.; Estrem, S.T.; Yang, J.; Wartell, R.M.; Gourse, R.L. Localization of the intrinsically bent DNA region upstream of the E.coli rrnB P1 promoter. Nucleic Acids Res. 1994, 22, 2344–2350. [Google Scholar] [CrossRef] [PubMed]
- Koo, H.S.; Wu, H.M.; Crothers, D.M. DNA bending at adenine · thymine tracts. Nature 1986, 320, 501–506. [Google Scholar] [CrossRef] [PubMed]
- Davis, C.A.; Capp, M.W.; Record, M.T., Jr.; Saecker, R.M. The effects of upstream DNA on open complex formation by Escherichia coli RNA polymerase. Proc. Natl. Acad. Sci. USA 2005, 102, 285–290. [Google Scholar] [CrossRef] [PubMed]
- Mangiarotti, L.; Cellai, S.; Ross, W.; Bustamante, C.; Rivetti, C. Sequence-dependent upstream DNA-RNA polymerase interactions in the open complex with λPR and λPRM promoters and implications for the mechanism of promoter interference. J. Mol. Biol. 2009, 385, 748–760. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Y.; Steitz, T.A. Crystal structures of the E. coli transcription initiation complexes with a complete bubble. Mol. Cell 2015, 58, 534–540. [Google Scholar] [CrossRef] [PubMed]
- Ross, W.; Gourse, R.L. Sequence-independent upstream DNA-alphaCTD interactions strongly stimulate Escherichia coli RNA polymerase-lacUV5 promoter association. Proc. Natl. Acad. Sci. USA 2005, 102, 291–296. [Google Scholar] [CrossRef] [PubMed]
- Hawley, D.K.; McClure, W.R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983, 11, 2237–2255. [Google Scholar] [CrossRef] [PubMed]
- Lisser, S.; Margalit, H. Compilation of E. coli mRNA promoter sequences. Nucleic Acids Res. 1993, 21, 1507–1516. [Google Scholar] [CrossRef] [PubMed]
- Ross, W.; Schneider, D.A.; Paul, B.J.; Mertens, A.; Gourse, R.L. An intersubunit contact stimulating transcription initiation by E coli RNA polymerase: Interaction of the alpha C-terminal domain and sigma region 4. Genes Dev. 2003, 17, 1293–1307. [Google Scholar] [CrossRef] [PubMed]
- Dove, S.L.; Huang, F.W.; Hochschild, A. Mechanism for a transcriptional activator that works at the isomerization step. Proc. Natl. Acad. Sci. USA 2000, 97, 13215–13220. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Tang, H.; Ebright, R.H. Functional interaction between RNA polymerase alpha subunit C-terminal domain and σ70 in UP-element- and activator-dependent transcription. Mol. Cell 2003, 11, 1621–1633. [Google Scholar] [CrossRef] [PubMed]
- Dove, S.L.; Darst, S.A.; Hochschild, A. Region 4 of sigma as a target for transcription regulation. Mol. Microbiol. 2003, 48, 863–874. [Google Scholar] [CrossRef] [PubMed]
- Niu, W.; Kim, Y.; Tau, G.; Heyduk, T.; Ebright, R.H. Transcription activation at class II CAP-dependent promoters: Two interactions between CAP and RNA polymerase. Cell 1996, 87, 1123–1134. [Google Scholar] [CrossRef] [PubMed]
- Nickels, B.E.; Roberts, C.W.; Roberts, J.W.; Hochschild, A. RNA-mediated destabilization of the σ70 region 4/β flap interaction facilitates engagement of RNA polymerase by the Q antiterminator. Mol. Cell 2006, 24, 457–468. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, J.E.; Zheng, D.; Busby, S.J.; Minchin, S.D. Identification and analysis of ‘extended-10’ promoters in Escherichia coli. Nucleic Acids Res. 2003, 31, 4689–4695. [Google Scholar] [CrossRef] [PubMed]
- Shimada, T.; Yamazaki, Y.; Tanaka, K.; Ishihama, A. The whole set of constitutive promoters recognized by RNA polymerase RpoD holoenzyme of Escherichia coli. PLoS ONE 2014, 9, e90447. [Google Scholar] [CrossRef] [PubMed]
- Aoyama, T.; Takanami, M.; Ohtsuka, E.; Taniyama, Y.; Marumoto, R.; Sato, H.; Ikehara, M. Essential structure of E. coli promoter: Effect of spacer length between the two consensus sequences on promoter function. Nucleic Acids Res. 1983, 11, 5855–5864. [Google Scholar] [CrossRef] [PubMed]
- Mulligan, M.E.; Brosius, J.; McClure, W.R. Characterization in vitro of the effect of spacer length on the activity of Escherichia coli RNA polymerase at the TAC promoter. J. Biol. Chem. 1985, 260, 3529–3538. [Google Scholar] [PubMed]
- Stefano, J.E.; Gralla, J.D. Spacer mutations in the lac ps promoter. Proc. Natl. Acad. Sci. USA 1982, 79, 1069–1072. [Google Scholar] [CrossRef] [PubMed]
- Hook-Barnard, I.G.; Hinton, D.M. The promoter spacer influences transcription initiation via σ70 region 1.1 of Escherichia coli RNA polymerase. Proc. Natl. Acad. Sci. USA 2009, 106, 737–742. [Google Scholar] [CrossRef] [PubMed]
- Yuzenkova, Y.; Tadigotla, V.R.; Severinov, K.; Zenkin, N. A new basal promoter element recognized by RNA polymerase core enzyme. EMBO J. 2011, 30, 3766–3775. [Google Scholar] [CrossRef] [PubMed]
- Barne, K.A.; Bown, J.A.; Busby, S.J.; Minchin, S.D. Region 2.5 of the Escherichia coli RNA polymerase σ70 subunit is responsible for the recognition of the ‘extended-10’ motif at promoters. EMBO J. 1997, 16, 4034–4040. [Google Scholar] [CrossRef] [PubMed]
- Haugen, S.P.; Berkmen, M.B.; Ross, W.; Gaal, T.; Ward, C.; Gourse, R.L. rRNA promoter regulation by nonoptimal binding of sigma region 1.2: An additional recognition element for RNA polymerase. Cell 2006, 125, 1069–1082. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Lew, C.M.; Gralla, J.D. Promoter opening by σ54 and σ70 RNA polymerases: σ factor-directed alterations in the mechanism and tightness of control. Genes Dev. 2000, 14, 2242–2255. [Google Scholar] [CrossRef] [PubMed]
- Waldburger, C.; Gardella, T.; Wong, R.; Susskind, M.M. Changes in conserved region 2 of Escherichia coli σ70 affecting promoter recognition. J. Mol. Biol. 1990, 215, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Marr, M.T.; Roberts, J.W. Promoter recognition as measured by binding of polymerase to nontemplate strand oligonucleotide. Science 1997, 276, 1258–1260. [Google Scholar] [CrossRef] [PubMed]
- Roberts, C.W.; Roberts, J.W. Base-specific recognition of the nontemplate strand of promoter DNA by E. coli RNA polymerase. Cell 1996, 86, 495–501. [Google Scholar] [CrossRef] [PubMed]
- Cook, V.M.; Dehaseth, P.L. Strand opening-deficient Escherichia coli RNA polymerase facilitates investigation of closed complexes with promoter DNA: Effects of DNA sequence and temperature. J. Biol. Chem. 2007, 282, 21319–21326. [Google Scholar] [CrossRef] [PubMed]
- Shultzaberger, R.K.; Chen, Z.; Lewis, K.A.; Schneider, T.D. Anatomy of Escherichia coli σ70 promoters. Nucleic Acids Res. 2007, 35, 771–788. [Google Scholar] [CrossRef] [PubMed]
- Campagne, S.; Marsh, M.E.; Capitani, G.; Vorholt, J.A.; Allain, F.H. Structural basis for –10 promoter element melting by environmentally induced sigma factors. Nat. Struct. Mol. Biol. 2014, 21, 269–276. [Google Scholar] [CrossRef] [PubMed]
- Haugen, S.P.; Ross, W.; Manrique, M.; Gourse, R.L. Fine structure of the promoter-sigma region 1.2 interaction. Proc. Natl. Acad. Sci. USA 2008, 105, 3292–3297. [Google Scholar] [CrossRef] [PubMed]
- Jeong, W.; Kang, C. Start site selection at lacUV5 promoter affected by the sequence context around the initiation sites. Nucleic Acids Res. 1994, 22, 4667–4672. [Google Scholar] [CrossRef] [PubMed]
- Lewis, D.E.; Adhya, S. Axiom of determining transcription start points by RNA polymerase in Escherichia coli. Mol. Microbiol. 2004, 54, 692–701. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Turnbough, C.L., Jr. Effects of transcriptional start site sequence and position on nucleotide-sensitive selection of alternative start sites at the pyrC promoter in Escherichia coli. J. Bacteriol. 1994, 176, 2938–2945. [Google Scholar] [PubMed]
- O’Neill, M.C. Escherichia coli promoters. I. Consensus as it relates to spacing class, specificity, repeat substructure, and three-dimensional organization. J. Biol. Chem. 1989, 264, 5522–5530. [Google Scholar] [PubMed]
- Thomason, M.K.; Bischler, T.; Eisenbart, S.K.; Forstner, K.U.; Zhang, A.; Herbig, A.; Nieselt, K.; Sharma, C.M.; Storz, G. Global transcriptional start site mapping using differential RNA sequencing reveals novel antisense RNAs in Escherichia coli. J. Bacteriol. 2015, 197, 18–28. [Google Scholar] [CrossRef] [PubMed]
- Feklistov, A.; Barinova, N.; Sevostyanova, A.; Heyduk, E.; Bass, I.; Vvedenskaya, I.; Kuznedelov, K.; Merkiene, E.; Stavrovskaya, E.; Klimasauskas, S.; et al. A basal promoter element recognized by free RNA polymerase sigma subunit determines promoter recognition by RNA polymerase holoenzyme. Mol. Cell 2006, 23, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Travers, A.A. Promoter sequence for stringent control of bacterial ribonucleic acid synthesis. J. Bacteriol. 1980, 141, 973–976. [Google Scholar] [PubMed]
- Guerin, M.; Leng, M.; Rahmouni, A.R. High resolution mapping of E.coli transcription elongation complex in situ reveals protein interactions with the non-transcribed strand. EMBO J. 1996, 15, 5397–5407. [Google Scholar] [PubMed]
- Vvedenskaya, I.O.; Vahedian-Movahed, H.; Bird, J.G.; Knoblauch, J.G.; Goldman, S.R.; Zhang, Y.; Ebright, R.H.; Nickels, B.E. Transcription. Interactions between RNA polymerase and the “core recognition element” counteract pausing. Science 2014, 344, 1285–1289. [Google Scholar] [CrossRef] [PubMed]
- Gaal, T.; Bartlett, M.S.; Ross, W.; Turnbough, C.L., Jr.; Gourse, R.L. Transcription regulation by initiating NTP concentration: RRNA synthesis in bacteria. Science 1997, 278, 2092–2097. [Google Scholar] [CrossRef] [PubMed]
- Guthold, M.; Zhu, X.; Rivetti, C.; Yang, G.; Thomson, N.H.; Kasas, S.; Hansma, H.G.; Smith, B.; Hansma, P.K.; Bustamante, C. Direct observation of one-dimensional diffusion and transcription by Escherichia coli RNA polymerase. Biophys. J. 1999, 77, 2284–2294. [Google Scholar] [CrossRef] [PubMed]
- Harada, Y.; Funatsu, T.; Murakami, K.; Nonoyama, Y.; Ishihama, A.; Yanagida, T. Single-molecule imaging of RNA polymerase-DNA interactions in real time. Biophys. J. 1999, 76, 709–715. [Google Scholar] [CrossRef] [PubMed]
- Kabata, H.; Kurosawa, O.; Arai, I.; Washizu, M.; Margarson, S.A.; Glass, R.E.; Shimamoto, N. Visualization of single molecules of RNA polymerase sliding along DNA. Science 1993, 262, 1561–1563. [Google Scholar] [CrossRef] [PubMed]
- Ricchetti, M.; Metzger, W.; Heumann, H. One-dimensional diffusion of Escherichia coli DNA-dependent RNA polymerase: A mechanism to facilitate promoter location. Proc. Natl. Acad. Sci. USA 1988, 85, 4610–4614. [Google Scholar] [CrossRef] [PubMed]
- Singer, P.; Wu, C.W. Promoter search by Escherichia coli RNA polymerase on a circular DNA template. J. Biol. Chem. 1987, 262, 14178–14189. [Google Scholar] [PubMed]
- Friedman, L.J.; Mumm, J.P.; Gelles, J. RNA polymerase approaches its promoter without long-range sliding along DNA. Proc. Natl. Acad. Sci. USA 2013, 110, 9740–9745. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Redding, S.; Finkelstein, I.J.; Gorman, J.; Reichman, D.R.; Greene, E.C. The promoter-search mechanism of Escherichia coli RNA polymerase is dominated by three-dimensional diffusion. Nat. Struct. Mol. Biol. 2013, 20, 174–181. [Google Scholar] [CrossRef] [PubMed]
- Bakshi, S.; Dalrymple, R.M.; Li, W.; Choi, H.; Weisshaar, J.C. Partitioning of RNA polymerase activity in live Escherichia coli from analysis of single-molecule diffusive trajectories. Biophys. J. 2013, 105, 2676–2686. [Google Scholar] [CrossRef] [PubMed]
- deHaseth, P.L.; Lohman, T.M.; Burgess, R.R.; Record, M.T., Jr. Nonspecific interactions of Escherichia coli RNA polymerase with native and denatured DNA: Differences in the binding behavior of core and holoenzyme. Biochemistry 1978, 17, 1612–1622. [Google Scholar] [CrossRef] [PubMed]
- Hofer, B.; Muller, D.; Koster, H. The pathway of E. coli RNA polymerase-promoter complex formation as visualized by footprinting. Nucleic Acids Res. 1985, 13, 5995–6013. [Google Scholar] [CrossRef] [PubMed]
- Kovacic, R.T. The 0 degree C closed complexes between Escherichia coli RNA polymerase and two promoters, T7-A3 and lacUV5. J. Biol. Chem. 1987, 262, 13654–13661. [Google Scholar] [PubMed]
- Mecsas, J.; Cowing, D.W.; Gross, C.A. Development of RNA polymerase-promoter contacts during open complex formation. J. Mol. Biol. 1991, 220, 585–597. [Google Scholar] [CrossRef] [PubMed]
- Schickor, P.; Metzger, W.; Werel, W.; Lederer, H.; Heumann, H. Topography of intermediates in transcription initiation of E. coli. EMBO J. 1990, 9, 2215–2220. [Google Scholar] [PubMed]
- Spassky, A.; Kirkegaard, K.; Buc, H. Changes in the DNA structure of the lac UV5 promoter during formation of an open complex with Escherichia coli RNA polymerase. Biochemistry 1985, 24, 2723–2731. [Google Scholar] [CrossRef] [PubMed]
- Ruff, E.F.; Zorn, K.; Capp, M.W.; Artsimovitch, I.; Record, M.T., Jr. Effects of upstream Escherichia coli RNA polymerase-DNA interactions on open complex formation. Unpublished work. 2015. [Google Scholar]
- Naryshkin, N.; Revyakin, A.; Kim, Y.; Mekler, V.; Ebright, R.H. Structural organization of the RNA polymerase-promoter open complex. Cell 2000, 101, 601–611. [Google Scholar] [CrossRef] [PubMed]
- Bartlett, M.S.; Gaal, T.; Ross, W.; Gourse, R.L. RNA polymerase mutants that destabilize RNA polymerase-promoter complexes alter NTP-sensing by rrn P1 promoters. J. Mol. Biol. 1998, 279, 331–345. [Google Scholar] [CrossRef] [PubMed]
- Rutherford, S.T.; Villers, C.L.; Lee, J.H.; Ross, W.; Gourse, R.L. Allosteric control of Escherichia coli rRNA promoter complexes by DksA. Genes Dev. 2009, 23, 236–248. [Google Scholar] [CrossRef] [PubMed]
- Li, X.Y.; McClure, W.R. Characterization of the closed complex intermediate formed during transcription initiation by Escherichia coli RNA polymerase. J. Biol. Chem. 1998, 273, 23549–23557. [Google Scholar] [CrossRef] [PubMed]
- Browning, D.F.; Busby, S.J. The regulation of bacterial transcription initiation. Nat. Rev. Microbiol. 2004, 2, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Drennan, A.C.; Kraemer, M.; Capp, M.; Gries, T.; Ruff, E.; Sheppard, C.; Wigneshweraraj, S.; Artsimovitch, I.; Record, M.T., Jr. Key roles of the downstream mobile jaw of Escherichia coli RNA polymerase in transcription initiation. Biochemistry 2012, 51, 9447–9459. [Google Scholar] [CrossRef] [PubMed]
- Mekler, V.; Minakhin, L.; Sheppard, C.; Wigneshweraraj, S.; Severinov, K. Molecular mechanism of transcription inhibition by phage T7 Gp2 protein. J. Mol. Biol. 2011, 413, 1016–1027. [Google Scholar] [CrossRef] [PubMed]
- Artsimovitch, I.; Svetlov, V.; Murakami, K.S.; Landick, R. Co-overexpression of Escherichia coli RNA polymerase subunits allows isolation and analysis of mutant enzymes lacking lineage-specific sequence insertions. J. Biol. Chem. 2003, 278, 12344–12355. [Google Scholar] [CrossRef] [PubMed]
- Hawley, D.K.; Malan, T.P.; Mulligan, M.E.; McClure, W.R. Intermediates on the pathway to open complex formation. In Promoters: Structure and Function; Praeger: New York, NY, USA, 1982; pp. 54–68. [Google Scholar]
- McClure, W.R. Rate-limiting steps in RNA chain initiation. Proc. Natl. Acad. Sci. USA 1980, 77, 5634–5638. [Google Scholar] [CrossRef] [PubMed]
- Tsodikov, O.V.; Record, M.T., Jr. General method of analysis of kinetic equations for multistep reversible mechanisms in the single-exponential regime: Application to kinetics of open complex formation between Eσ70 RNA polymerase and λPR promoter DNA. Biophys. J. 1999, 76, 1320–1329. [Google Scholar] [CrossRef] [PubMed]
- Drennan, A.C. Key conformational changes of Escherichia coli RNA polymerase and promoter DNA in transcription initiation. Ph.D. Thesis, University of Wisconsin, Madison, WI, USA, 2012. [Google Scholar]
- Heitkamp, S.E. Real-Time characterization of transcription initiation intermediates for E. coli RNA polymerase using fast footprinting and equilibrium and stopped-flow fluorescence. Ph.D. Thesis, University of Wisconsin, Madison, WI, USA, 2012. [Google Scholar]
- Hook-Barnard, I.G.; Hinton, D.M. Transcription initiation by mix and match elements: Flexibility for polymerase binding to bacterial promoters. Gene Regul. Syst. Biol. 2007, 1, 275–293. [Google Scholar]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruff, E.F.; Record, M.T., Jr.; Artsimovitch, I. Initial Events in Bacterial Transcription Initiation. Biomolecules 2015, 5, 1035-1062. https://doi.org/10.3390/biom5021035
Ruff EF, Record MT Jr., Artsimovitch I. Initial Events in Bacterial Transcription Initiation. Biomolecules. 2015; 5(2):1035-1062. https://doi.org/10.3390/biom5021035
Chicago/Turabian StyleRuff, Emily F., M. Thomas Record, Jr., and Irina Artsimovitch. 2015. "Initial Events in Bacterial Transcription Initiation" Biomolecules 5, no. 2: 1035-1062. https://doi.org/10.3390/biom5021035
APA StyleRuff, E. F., Record, M. T., Jr., & Artsimovitch, I. (2015). Initial Events in Bacterial Transcription Initiation. Biomolecules, 5(2), 1035-1062. https://doi.org/10.3390/biom5021035




