Local Order in the Unfolded State: Conformational Biases and Nearest Neighbor Interactions
Abstract
:1. Introduction
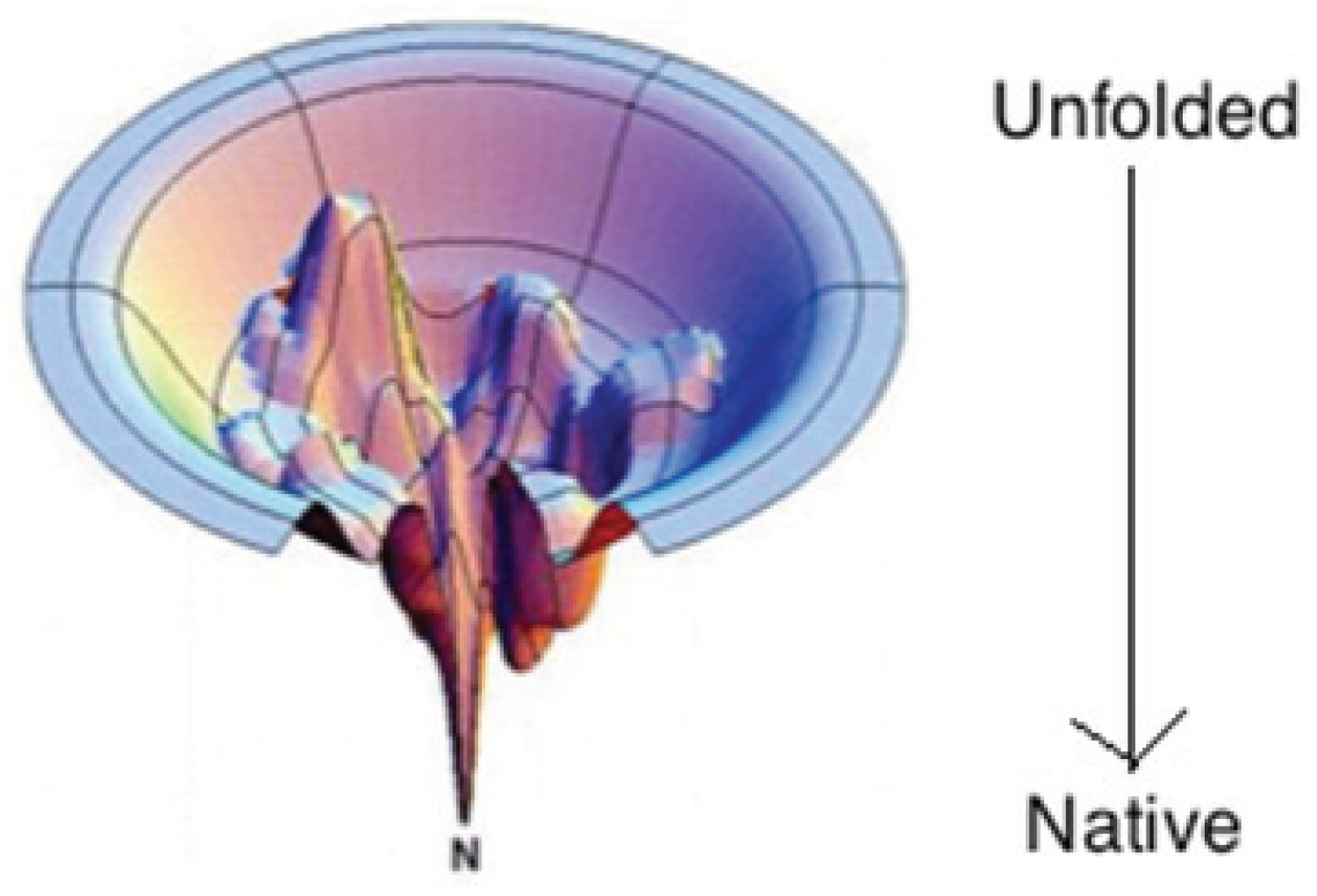
2. Conformational Propensities in the Unfolded State and the Alanine Debate
2.1. Unfolded Does Not Mean Random
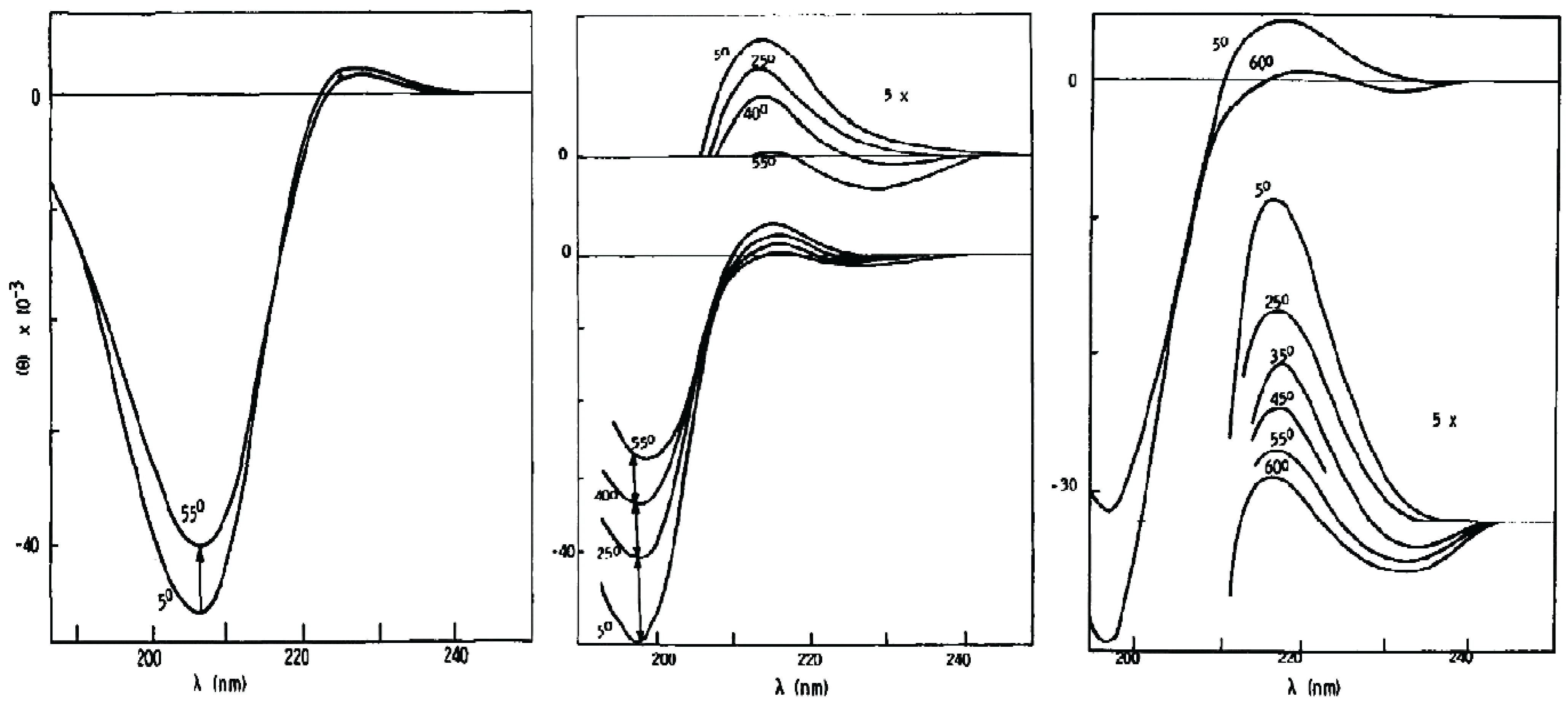
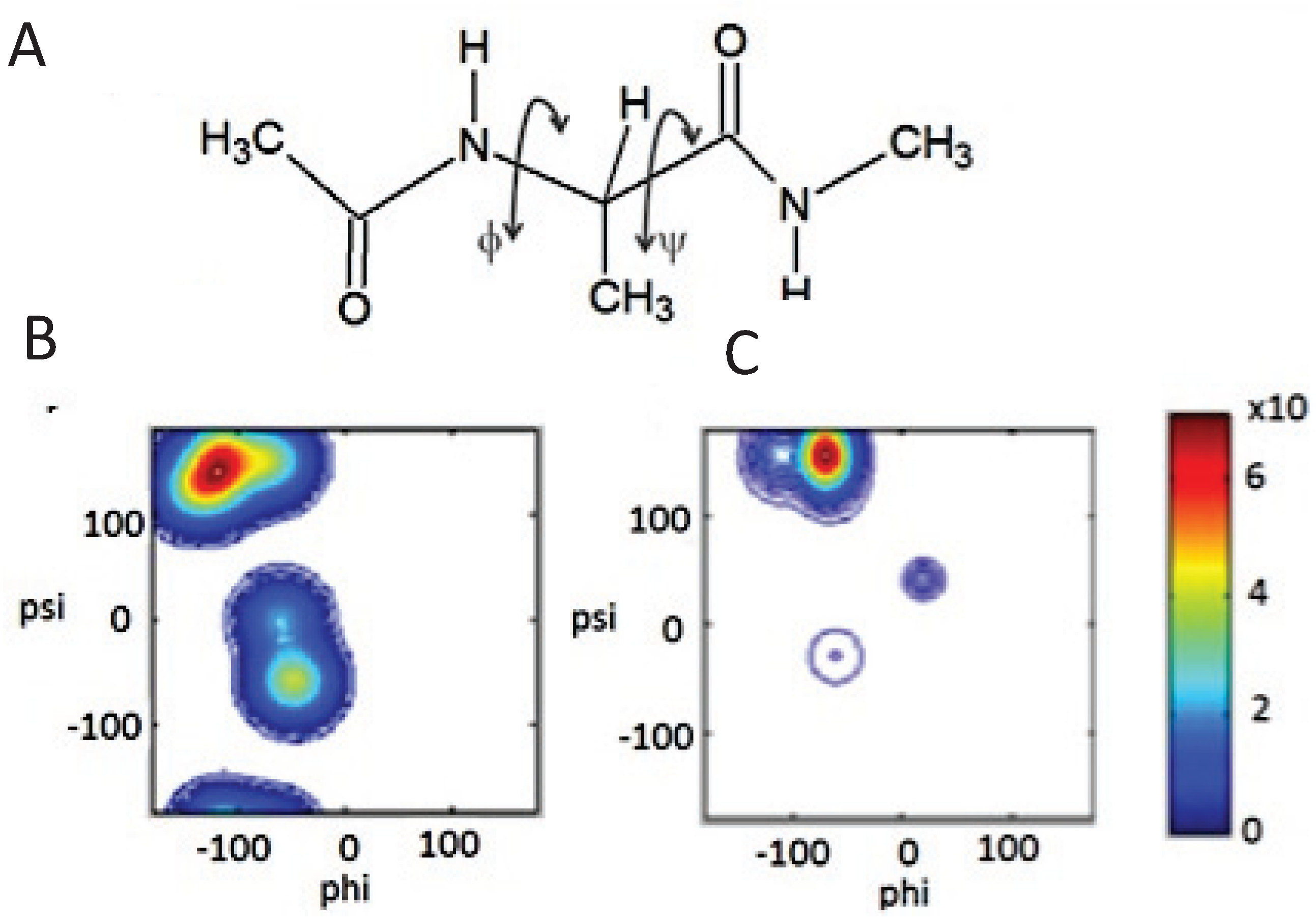
2.2. Conformational Preference of Alanine in Water
2.2.1. Experimental Studies
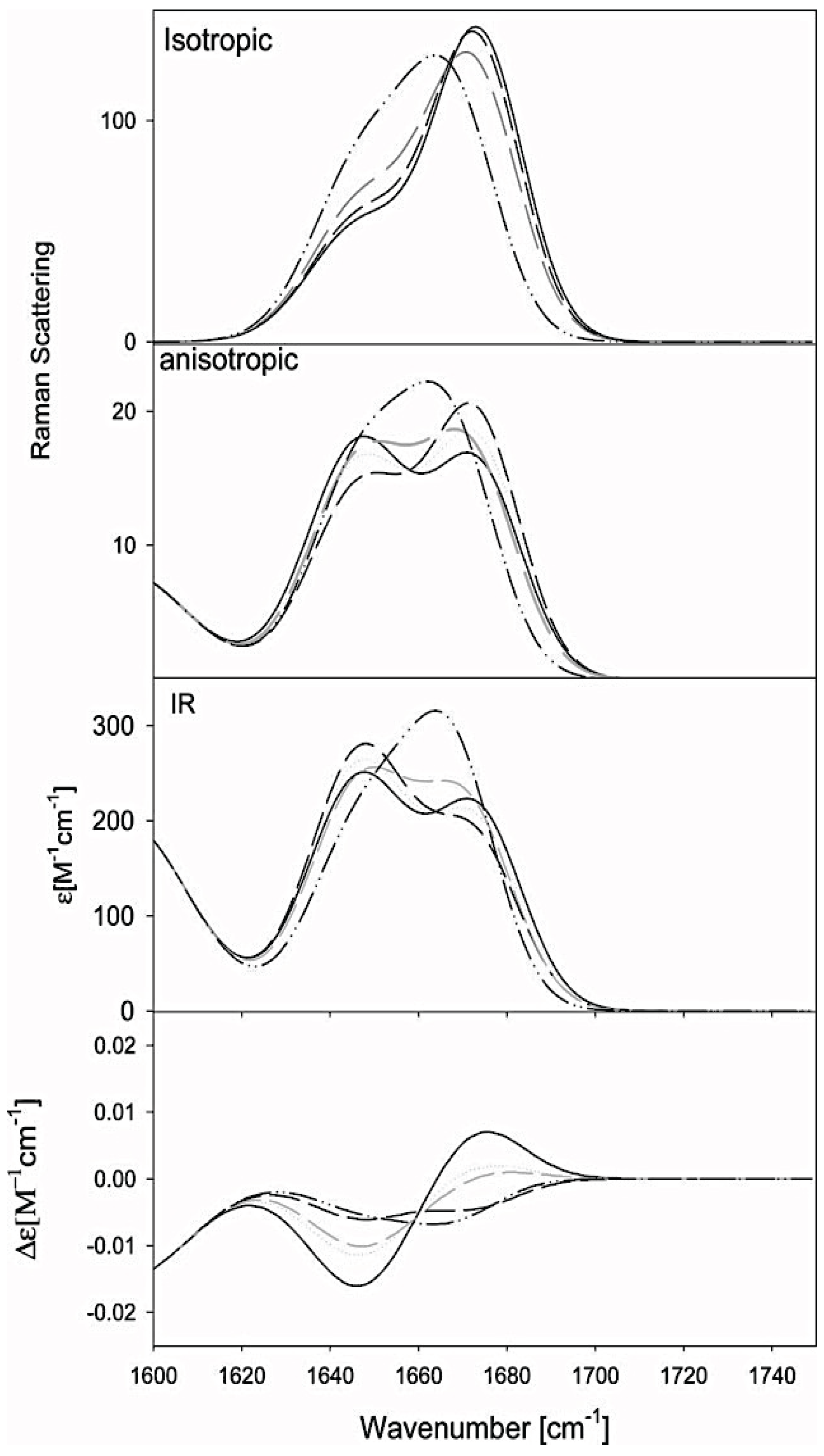
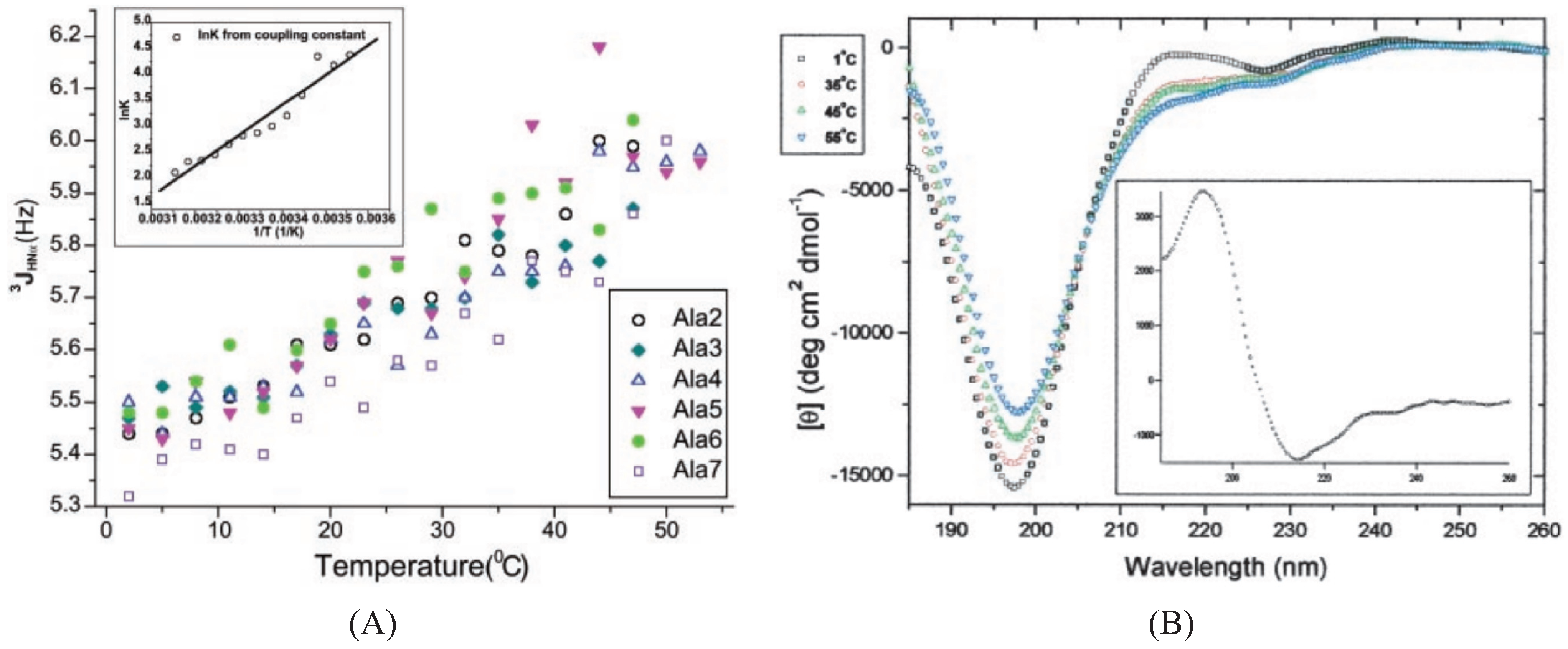
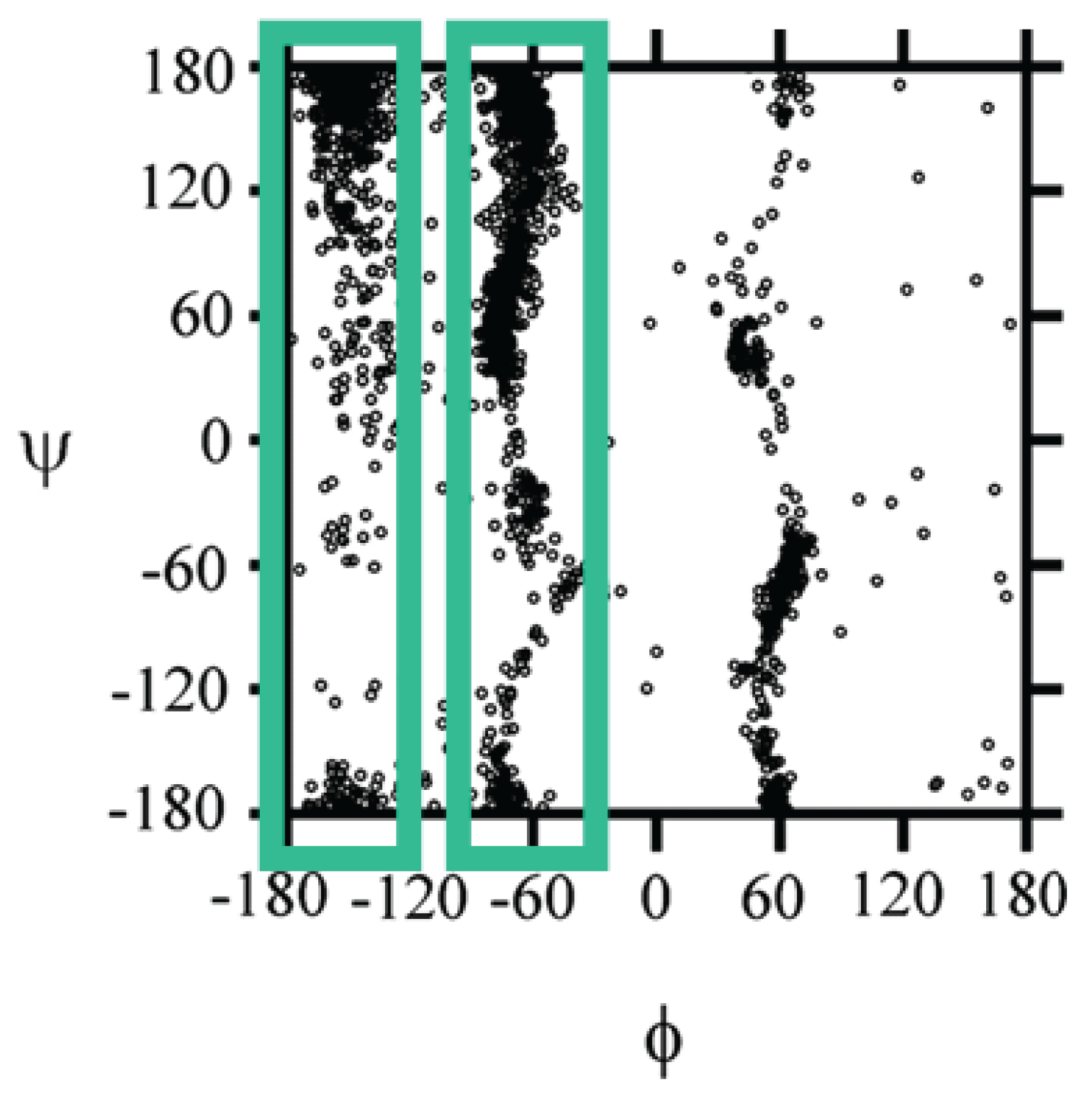
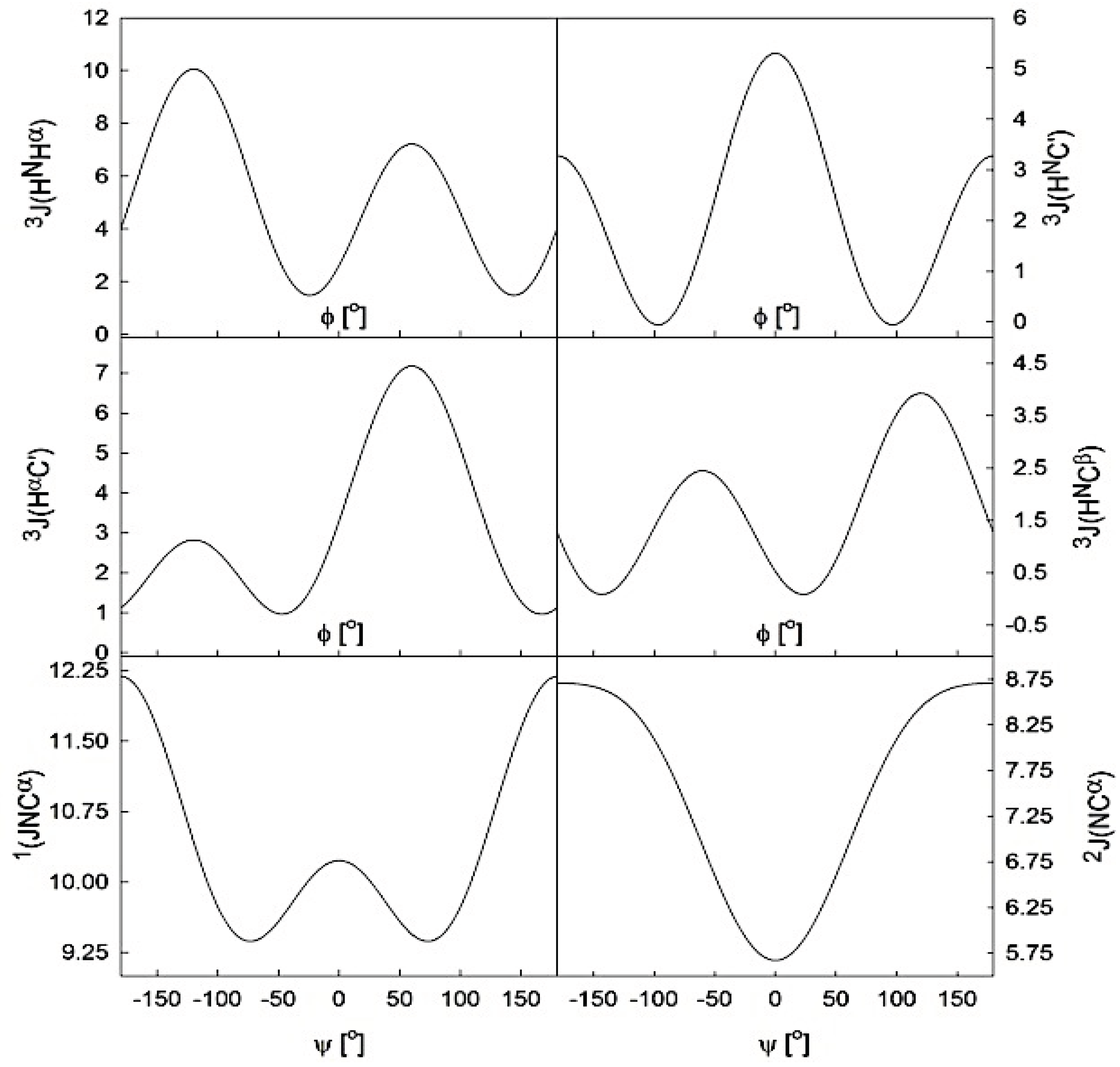
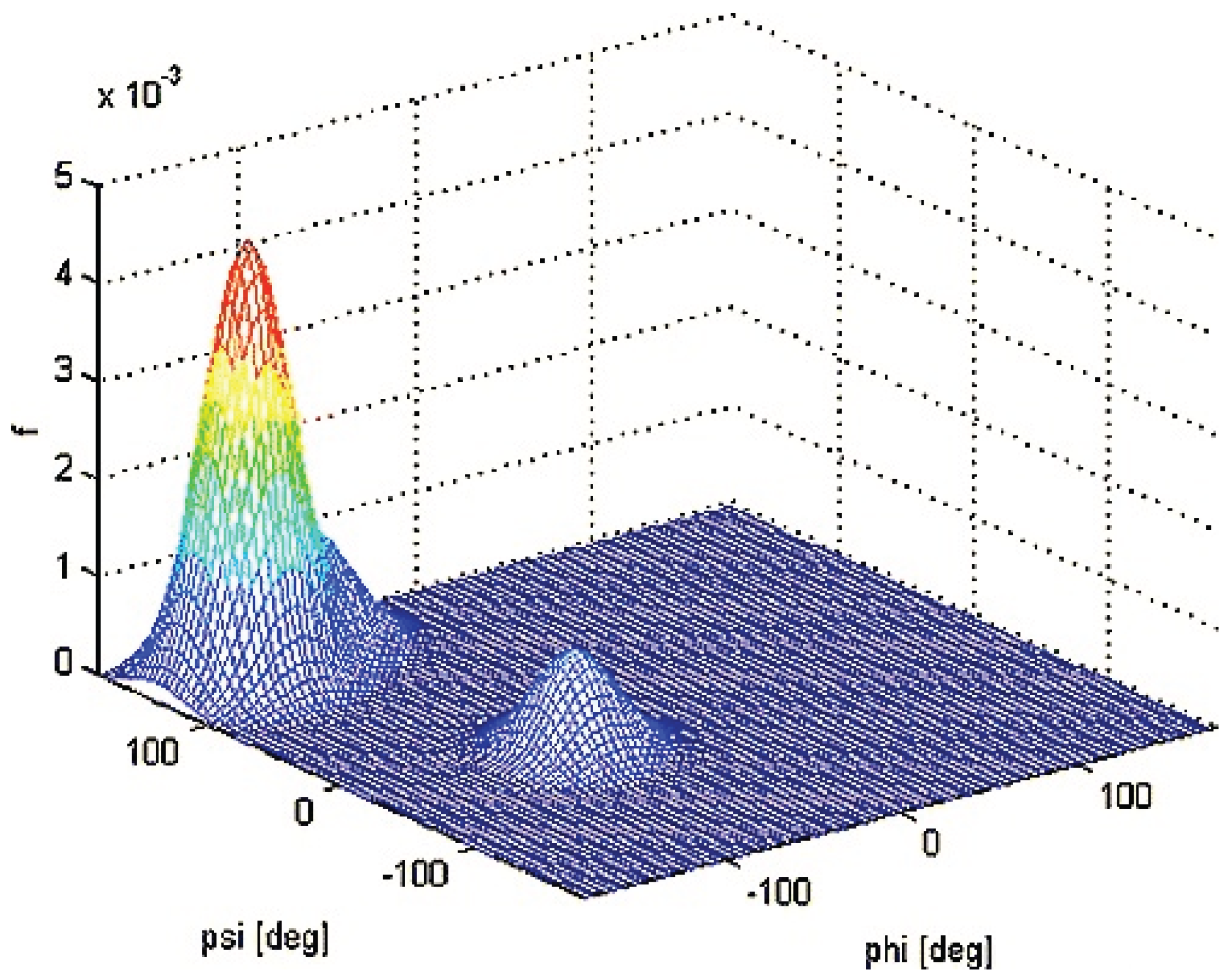
2.2.2. Theoretical Studies on Alanine
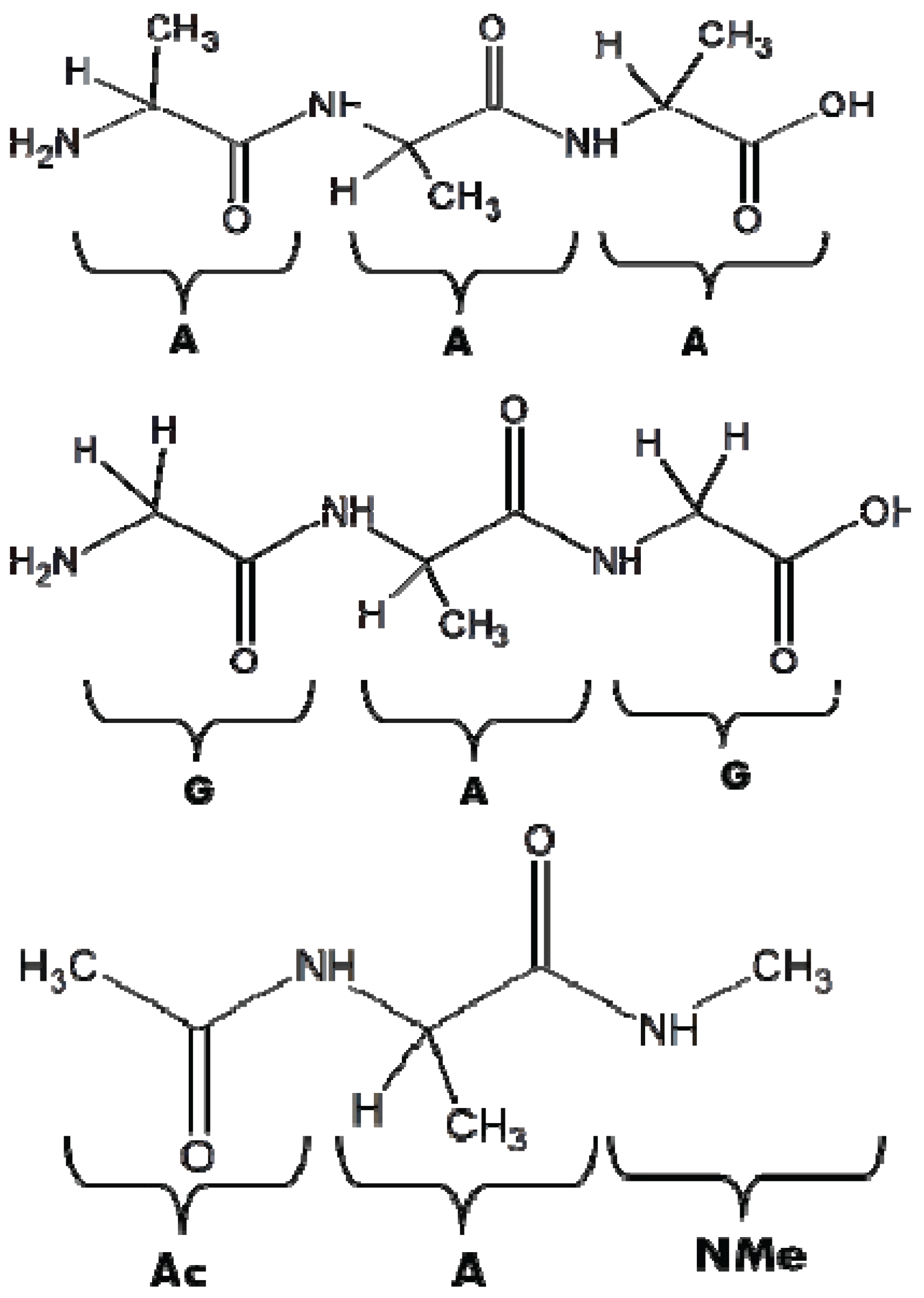
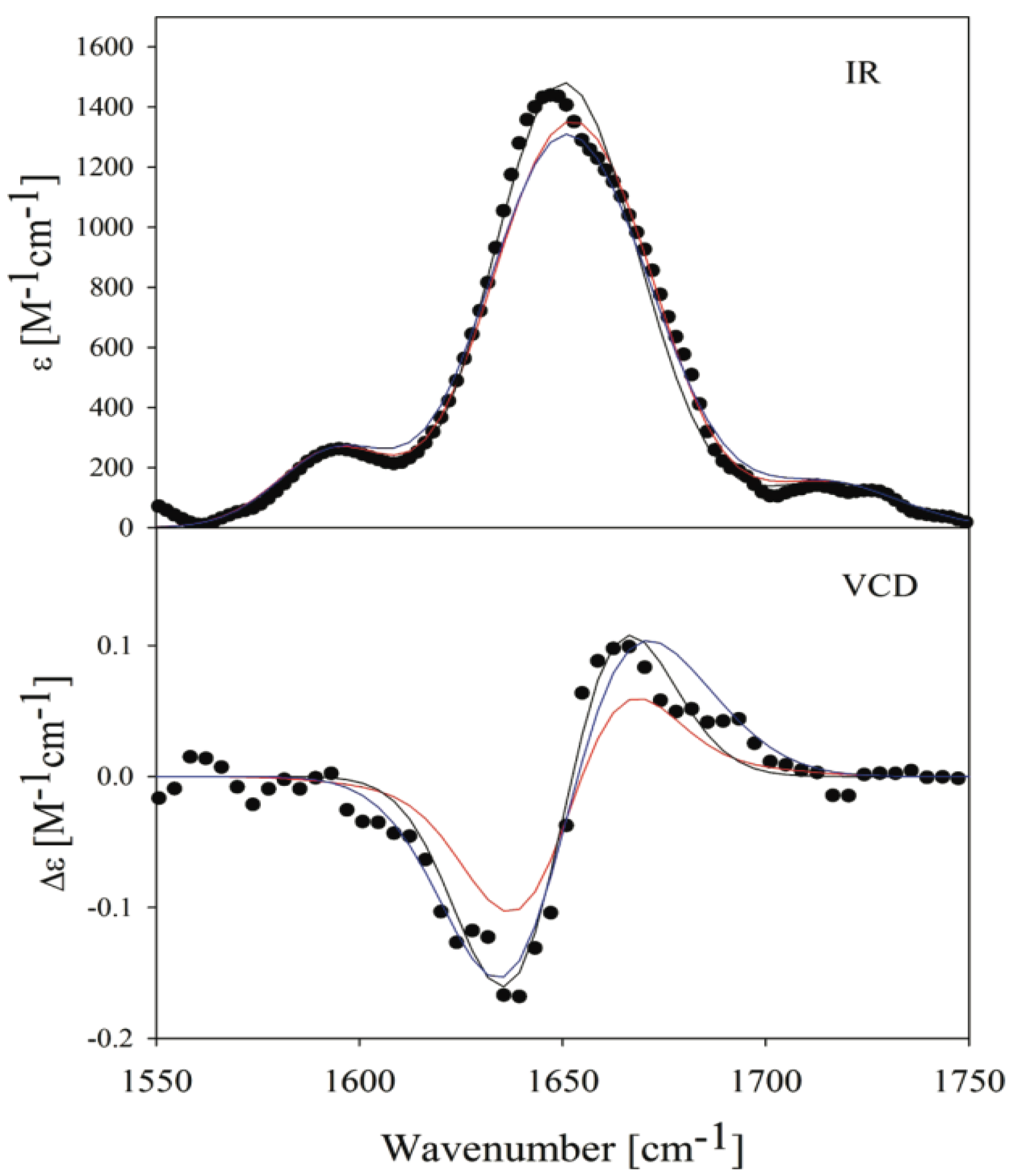
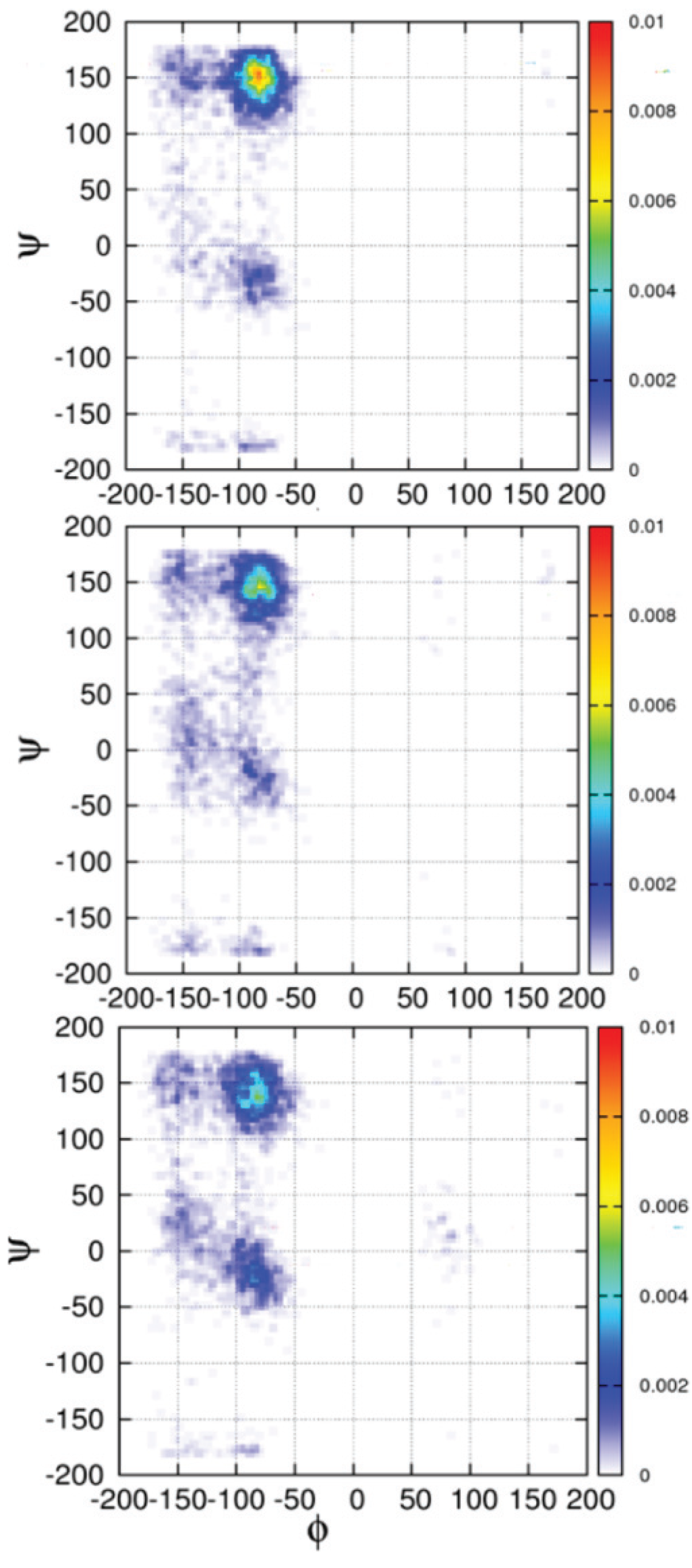
3. Conformational Biases of Other Amino Acid Residues
3.1. Experimental Studies

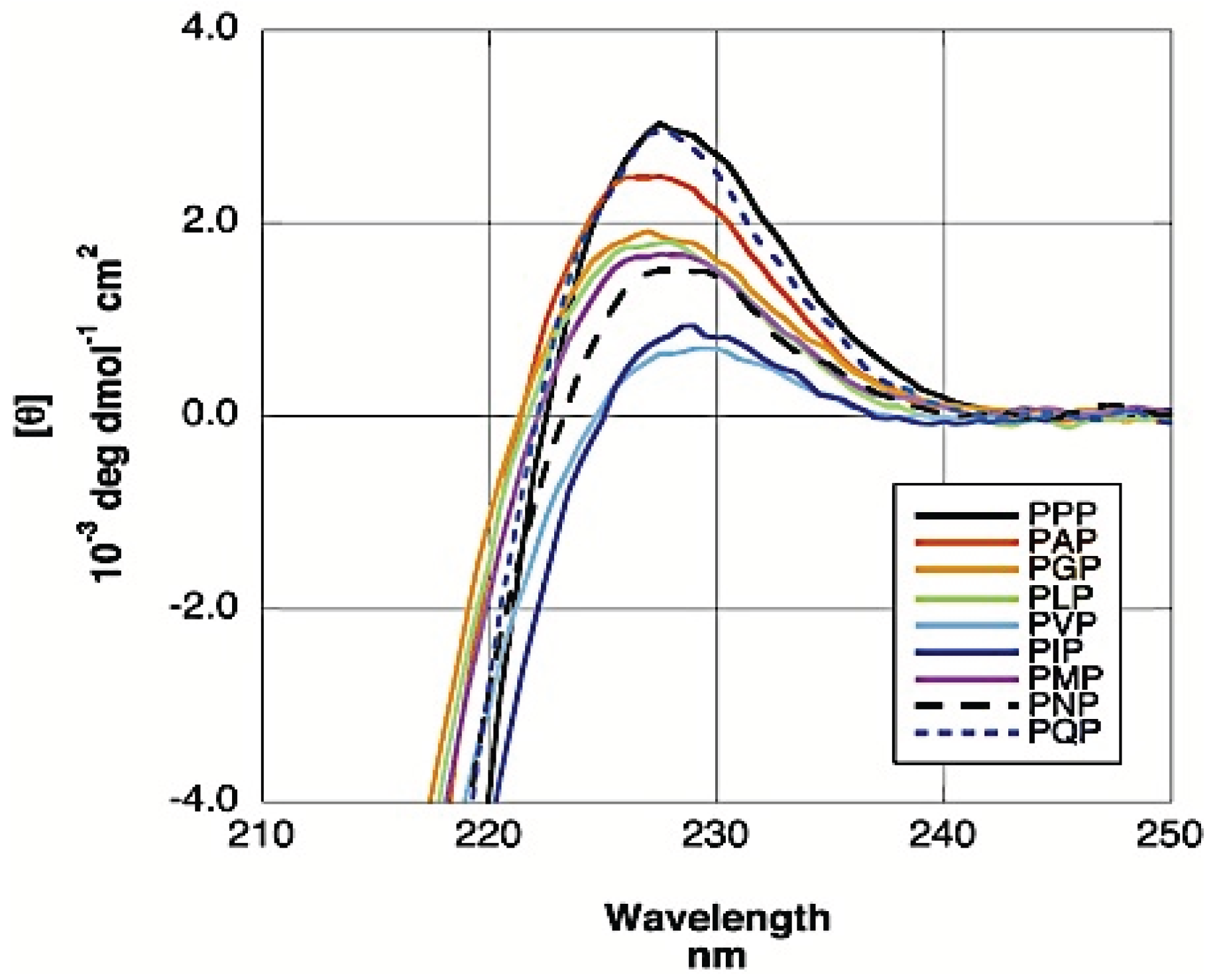

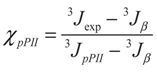
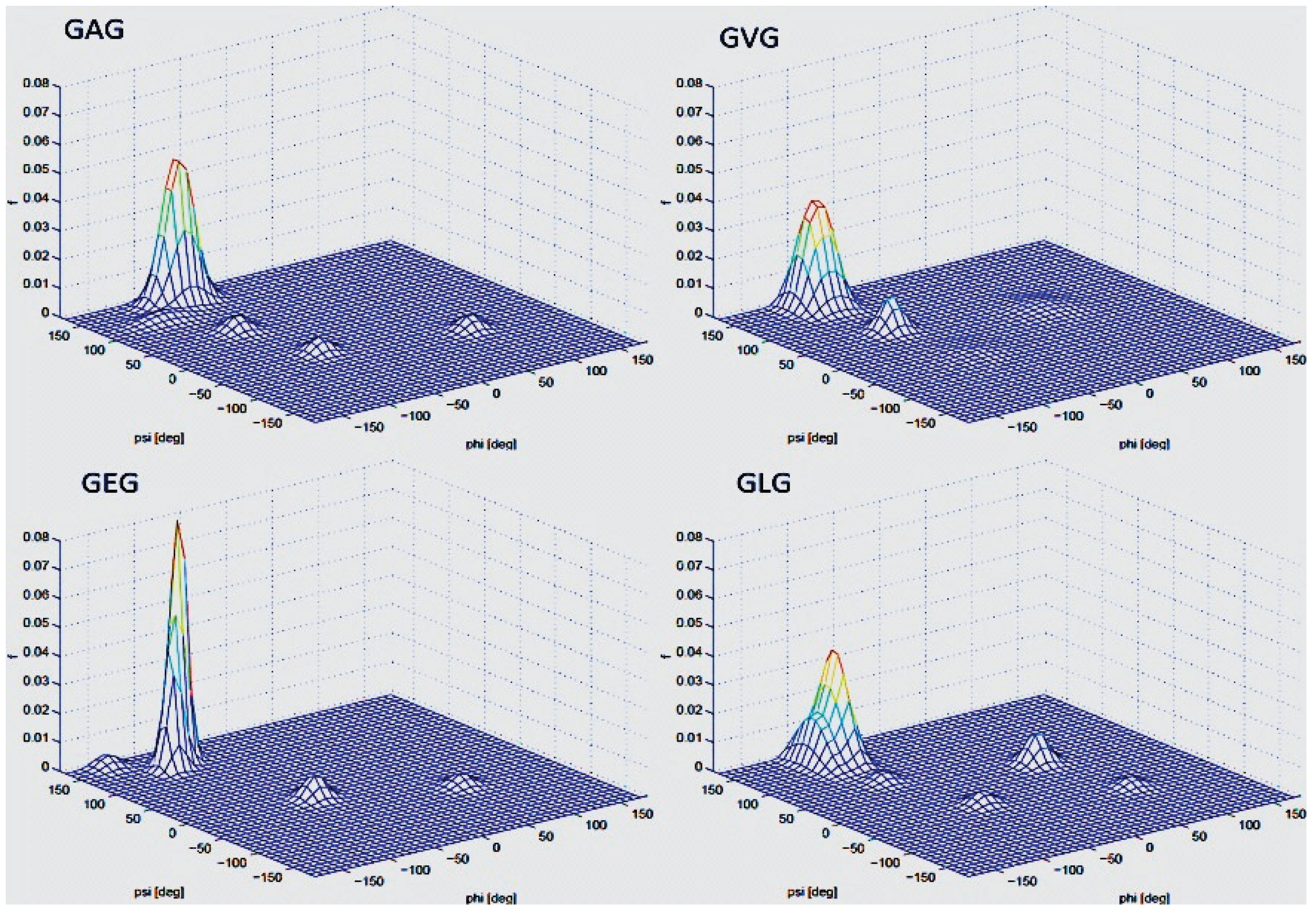
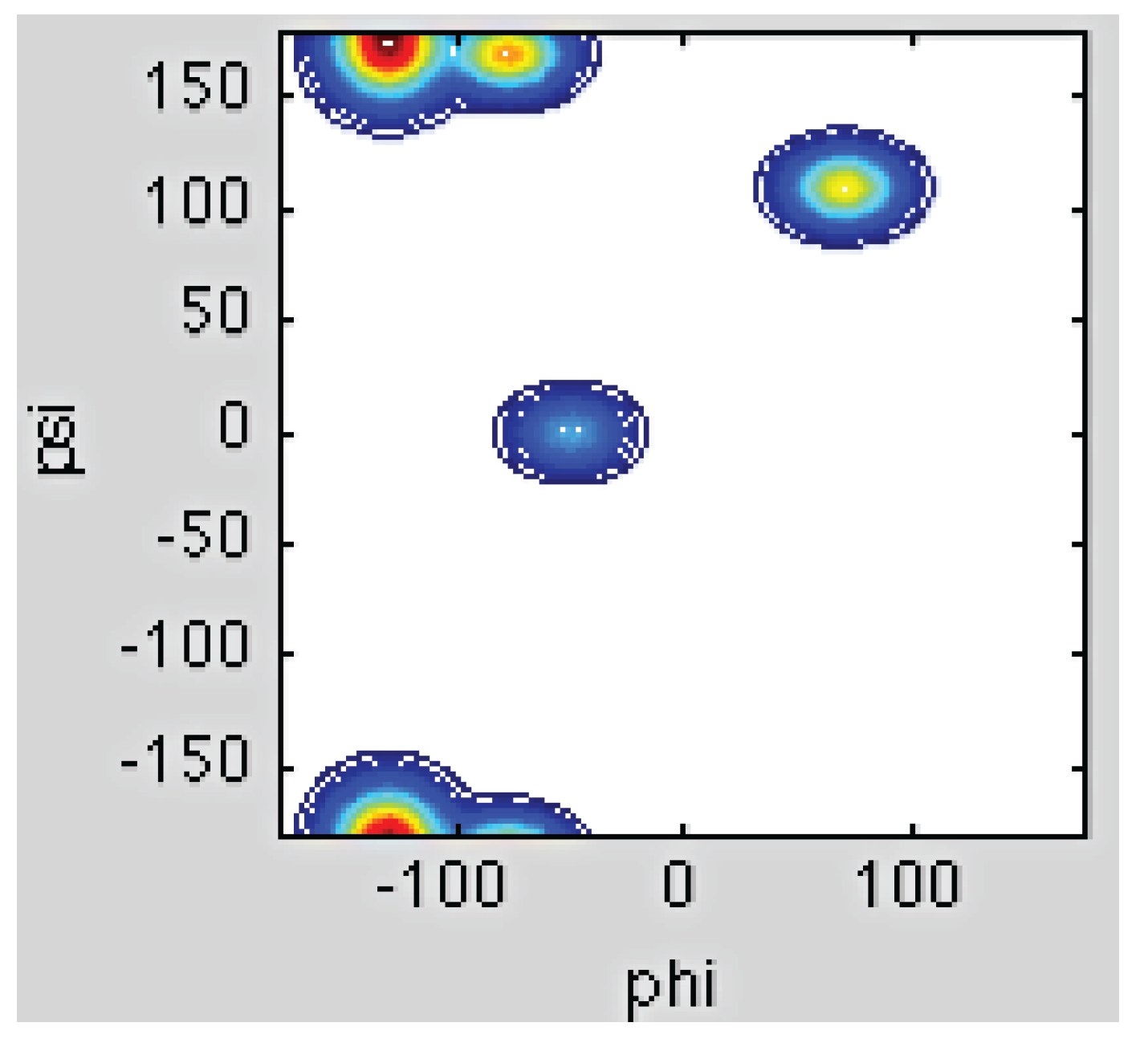
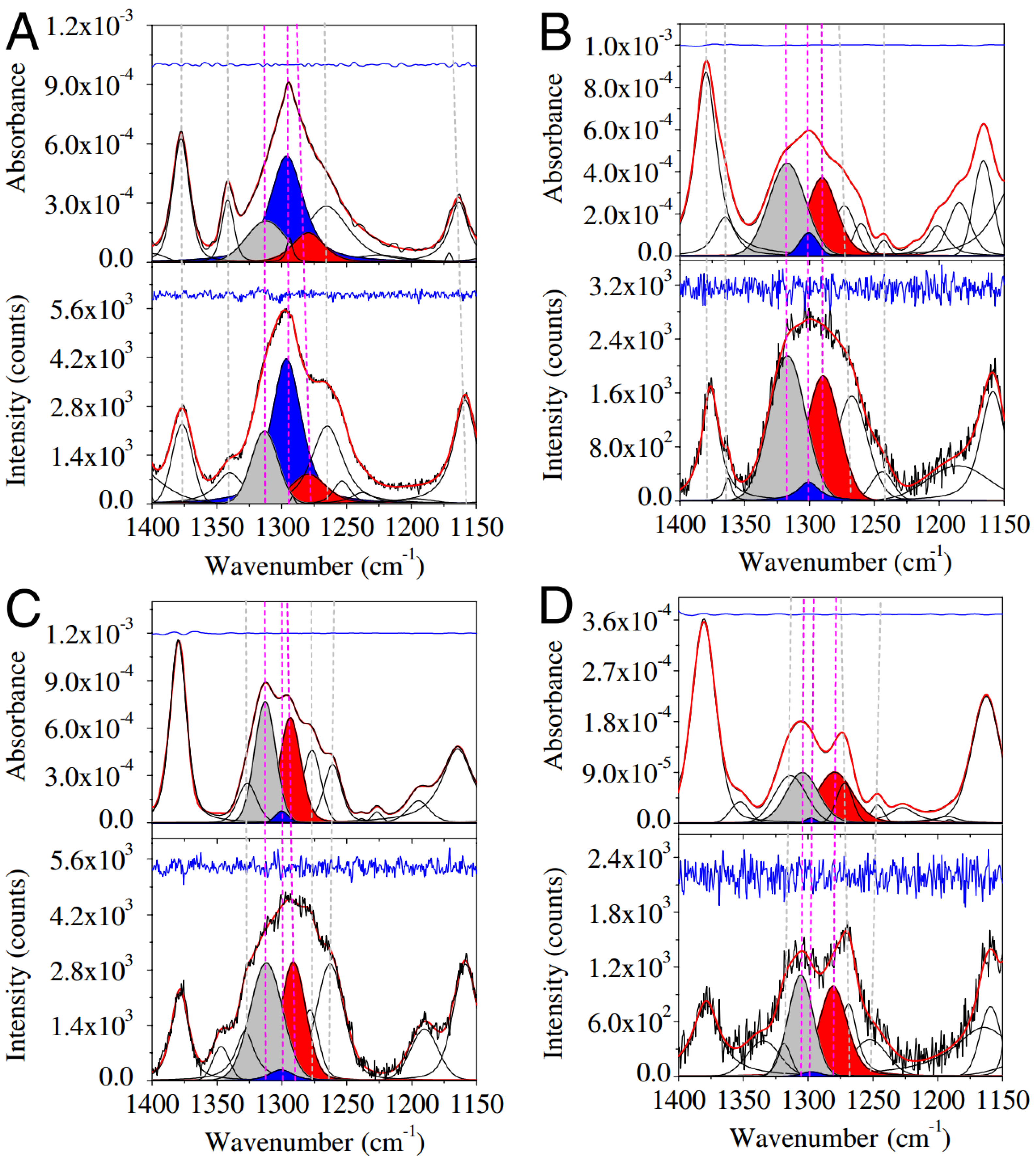
3.2. Analysis of Coil Libraries
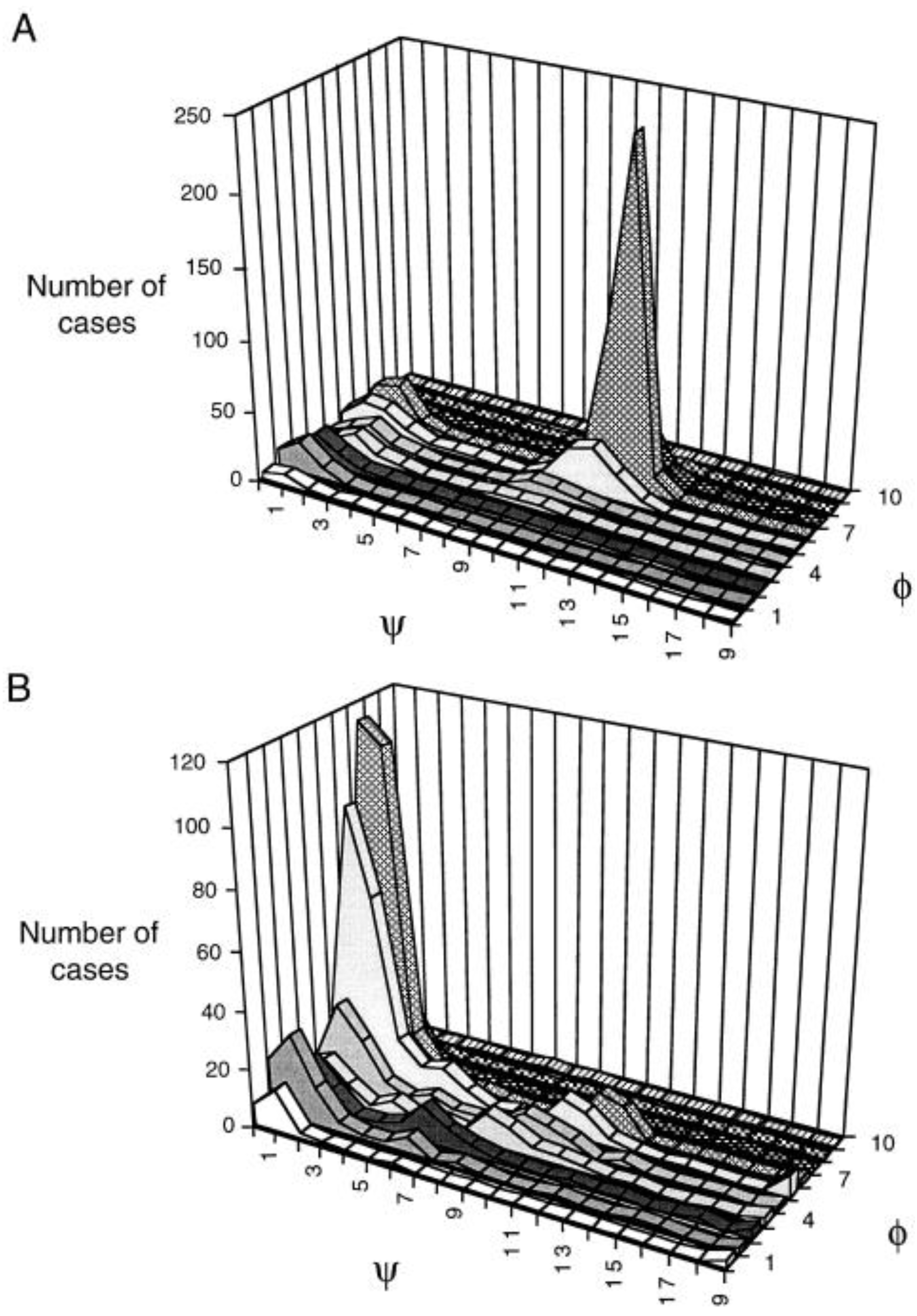
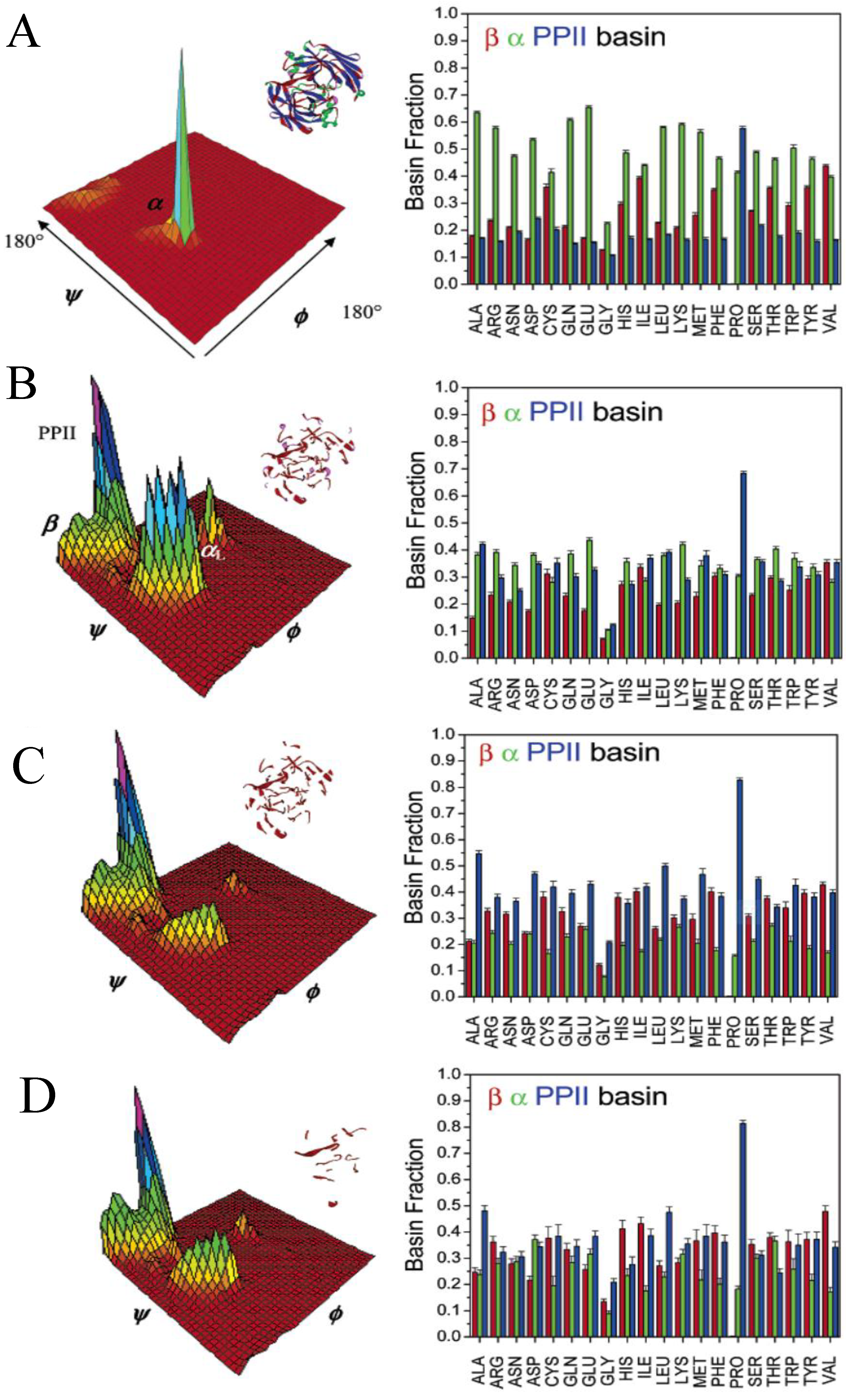
| pPII | β | II’/III turn | γ-turn | III’ turn | I’/γ turn | |
|---|---|---|---|---|---|---|
| GYGc | 0.38 | 0.19 | 0.31 | 0.05 | ||
| GYGe | 0.45 | 0.34 | 0.16 | 0.12 | ||
| GFGc | 0.29 | 0.21 | 0.39 | 0.1 | ||
| GFGe | 0.45 | 0.45 | 0.05 | 0.05 | ||
| GIGc | 0.26 | 0.36 | 0.38 | |||
| GIGe | 0.42 | 0.37 | 0.18 | 0.08 | ||
| GVGc | 0.33 | 0.28 | 0.39 | |||
| GVGe | 0.32 | 0.46 | 0.04 | 0.11 | 0.07 | |
| GRGc | 0.31 | 0.15 | 0.48 | 0.06 | ||
| GRGe | 0.58 | 0.2 | 0.08 | 0.03 | 0.03 | 0.08 |
| GEGc | 0.39 | 0.17 | 0.41 | 0.03 | ||
| GEGe | 0.54 | 0.32 | 0.08 | 0.04 | 0.04 |
4. Nearest Neighbor Influence of Conformational Propensities
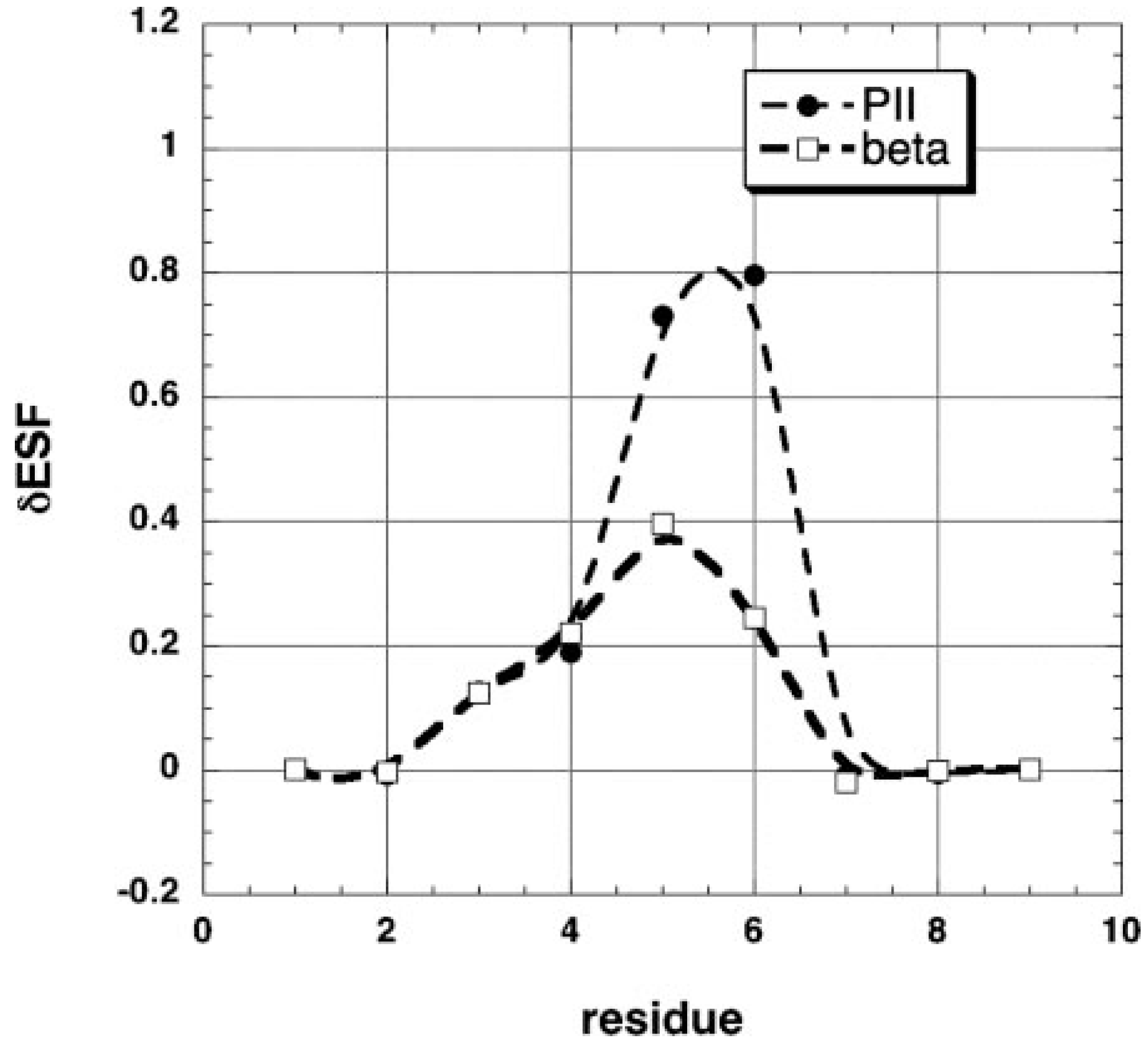
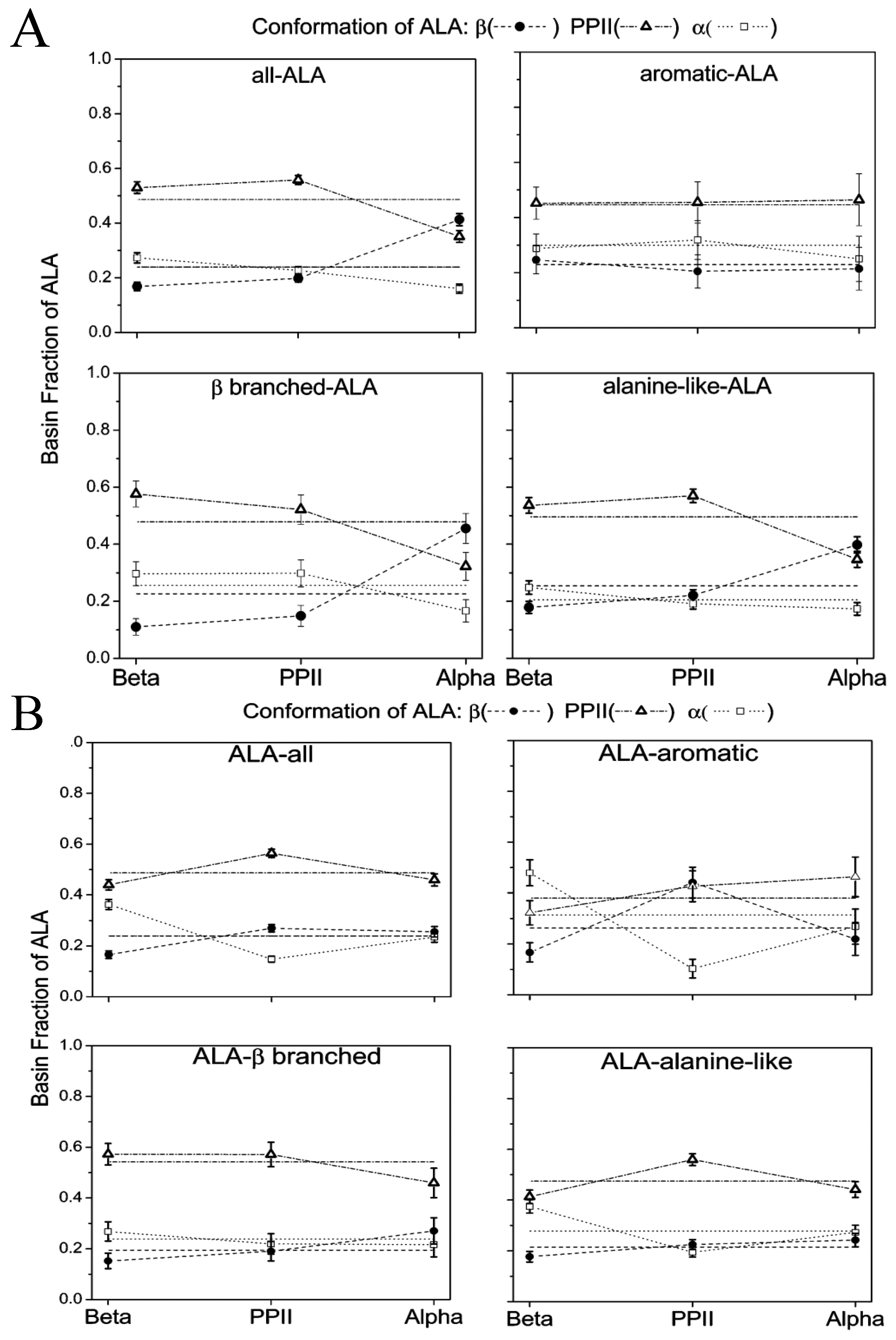

5. Conclusions and Outlook
Acknowledgments
Authors Contributions
Conflicts of Interest
References
- Uversky, V.; Dunker, A.K. Why are we interested in the unfolded peptides and proteins? In Protein and Peptide Folding, Misfolding, and Non-folding; Schweitzer-Stenner, R., Ed.; Wiley & Sons: Hoboken, NJ, USA, 2012; pp. 3–54. [Google Scholar]
- Onuchic, J.N.; Luthey-Schulten, Z.; Wolynes, P.G. Theory of protein folding; the energy landscape persepective. Annu. Rev. Phys. Chem. 1997, 48, 545–600. [Google Scholar] [CrossRef]
- Dill, K.A. Dominant forces in protein folding. Biochemistry 1990, 29, 7133–7155. [Google Scholar] [CrossRef]
- Baldwin, R.L.; Rose, G.D. Is protein folidng hierarchic. I. Local structure and peptide folding. Trends Biochem. Sci. 1999, 24, 26–33. [Google Scholar] [CrossRef]
- Ben Naim, A. The role of hydrogen bonds in protein folding and protein association. J. Phys. Chem. 1991, 95, 1437–1444. [Google Scholar] [CrossRef]
- Chan, H.S.; Dill, K.A. Polymer principles in protein structure and stability. Annu. Rev. Biophys. Biophys. Chem. 1991, 20, 447–490. [Google Scholar] [CrossRef]
- Anfinsen, C.B.; Haber, E.; Sela, M.; White, F.H., Jr. The kinetics of formation of native ribonuclease during oxidation of the reduced polypeptide chain. Proc. Natl. Acad. Sci. USA 1961, 47, 1309–1314. [Google Scholar] [CrossRef]
- Levinthal, C. Are there pathways for protein folding? J. Chim. Phys. Phys. 1968, 65, 44–45. [Google Scholar]
- Dill, K.A.; Chan, H.S. From levinthal to pathways to funnels. Nat. Struct. Biol. 1997, 4, 10–19. [Google Scholar] [CrossRef]
- Bryngelson, J.D.; Wolynes, P.G. Spin glasses and the statistical mechanics of protein folding. Proc. Natl. Acad. Sci. USA 1987, 84, 7524–7528. [Google Scholar] [CrossRef]
- Ramachandran, G.N.; Ramachandran, C.; Sasisekharan, V. Stereochemistry of polypeptide chain configurations. J. Mol. Biol. 1963, 7, 95–99. [Google Scholar] [CrossRef]
- Flory, P.J. Statistical Mechanics of Chain Molecules; Wiley & Sons: New York, NY, USA, 1969; pp. 30–31. [Google Scholar]
- Brant, D.A.; Flory, P.J.J. The configuration of random polypeptide chains. II. Theory. J. Am. Chem. Soc. 1965, 87, 2791–2800. [Google Scholar] [CrossRef]
- Tanford, C. Protein denaturation. Adv. Protein Chem. 1968, 23, 121–282. [Google Scholar] [CrossRef]
- Toal, S.; Meral, D.; Verbaro, D.; Urbanc, B.; Schweitzer-Stenner, R. pH-independence of trialanine and the effects of termini blocking in short peptides: A combined vibrational, NMR, UVCD, and molecular dynamics study. J. Phys. Chem. B 2013, 117, 3689–3706. [Google Scholar]
- Bernado, P.; Mylonas, E.; Petoukhov, M.V.; Blackledge, M.; Svergun, D.I. Structural characterization of flexible proteins using small-angle X-ray scattering. J. Am. Chem. Soc. 2007, 129, 5656–5664. [Google Scholar]
- Bernado, P.; Bertoncini, C.W.; Griesinger, C.; Zweckstetter, M.; Blackledge, M. Defining long-range order and local disorder in native a-synuclein using residua dipolar couplings. J. Am. Chem. Soc. 2005, 127, 17968–17969. [Google Scholar] [CrossRef]
- Bernado, P.; Blanchard, L.; Timmins, P.; Marion, D.; Ruigrok, R.W.; Blackledge, M. A structural model for unfolded proteins from residual dipolar couplings and small-angle X-ray scattering. Proc. Natl. Acad. Sci. USA 2005, 102, 17002–17007. [Google Scholar] [CrossRef]
- Mukrasch, M.D.; Markwick, P.; Biernat, J.; von Bergen, M.; Bernado, P.; Greisinger, C.; Mandelkow, E.; Zweckstetter, M.; Blackledge, M. Highly populated turn conformations in natively unfolded tau protein identified from residual dipolar couplings and molecular simulation. J. Am. Chem. Soc. 2007, 129, 5235–5243. [Google Scholar] [CrossRef]
- Shortle, D. The expanded denaturated state: An ensemble of conformations trapped in a locally encoded topological space. Adv. Protein Chem. 2002, 62, 1–23. [Google Scholar] [CrossRef]
- Ohnishi, S.; Lee, A.L.; Egdell, M.H.; Shortle, D. Direct demonstration of structural similarity between native and denatured eglin C. Biochemistry 2004, 43, 4064–4070. [Google Scholar]
- Alexandrescu, A.T.; Abeygunawardana, C.; Shortle, D. Structure and dynamics of a denaturared 131-residue fragment of staphylococcal nuclease: A heteronuclear nmr study. Biochemistry 1994, 33, 1063–1072. [Google Scholar] [CrossRef]
- Uversky, V.N. What does it mean to be natively unfolded? Eur. J. Biochem. 2002, 269, 2–12. [Google Scholar] [CrossRef]
- Dunker, A.K.; Cortese, M.S.; Romero, P.; Iakoucheva, L.M.; Uversky, V.N. Flexible nets. The roles of intrinsic disorder in protein interaction networks. FEBS J. 2005, 272, 5129–5148. [Google Scholar] [CrossRef]
- Uversky, V.N. Natively unfolded proteins. In Unfolded Proteins. From Denaturated to Intrinsically Disordered; Creamer, T.P., Ed.; Nova: Nauppauge, NY, USA, 2008; pp. 237–294. [Google Scholar]
- Wright, P.E.; Dyson, H.J.; Lerner, R.A. Intrinsically unstructured proteins: Re-assessing the protein structure-function paradigm. Biochemistry 1988, 27, 7167–7175. [Google Scholar] [CrossRef]
- Dobson, C.M. Protein misfolding, evolution and disease. Trends Biochem. Sci. 1999, 24, 329–332. [Google Scholar] [CrossRef]
- Calamai, M.; Canale, C.; Relini, A.; Stefani, M.; Chiti, F.; Dobson, C.M. Reversal of protein aggregation provides evidence for multiple aggregated states. J. Mol. Biol. 2005, 346, 603–616. [Google Scholar] [CrossRef]
- Cho, M.K.; Nodet, G.; Kim, H.Y.; Jensen, M.R.; Bernado, P.; Fernandez, C.O.; Becker, S.; Blackledge, M.; Zweckstetter, M. Structural characterization of alpha-synuclein in an aggregation prone state. Protein Sci. 2009, 18, 1840–1846. [Google Scholar] [CrossRef]
- Syme, C.D.; Blanch, E.W.; Holt, C.; Ross, J.; Goedert, M.; Hecht, L.; Barron, L.D. A Raman optical activity study of rheomorphism in caseins, synucleins and tau new insight into the structure and behaviour of natively unfolded proteins. Eur. J. Biochem. 2002, 269, 148–156. [Google Scholar] [CrossRef]
- Toriumi, K.; Oma, Y.; Kino, Y.; Futai, E.; Sasagawa, N.; Ishiura, S. Expression of polyalanine stretches induces mitochondrial dysfunction. J. Neurosci. Res. 2008, 86, 1529–1537. [Google Scholar] [CrossRef]
- Gerum, C.; Silvers, R.; Wirmer-Bartoschek, J.; Schwalbe, H. Unfolded-state structure and dynamic influence the fibril formation in human prion protein. Angew. Chem. Int. Ed. Engl. 2009, 48, 9452–9456. [Google Scholar] [CrossRef]
- Gsponer, J.; Haberthür, U.; Calfisch, A. The role of side-chain interactions in the early steps of aggregation: Molecular dynamics simulations of an amyloid-forming peptide from yeat prion sup35. Proc. Natl. Acad. Sci. USA 2003, 100, 5154–5159. [Google Scholar] [CrossRef]
- Kelly, J.W. The alternative conformations of amyloidogenic proteins and their multi-step assembly pathways. Curr. Opin. Struct. Biol. 1998, 8, 101–106. [Google Scholar] [CrossRef]
- McColl, I.H.; Blanch, E.W.; Gill, A.C.; Rhie, A.G.O.; Ritchie, M.A.; Hecht, L.; Nielsen, K.; Barron, L.D. A new perspective on β-sheet structures using vibrational raman optical activity: From poly(L-lysine) to the prion protein. J. Am. Chem. Soc. 2003, 125, 10019–10026. [Google Scholar] [CrossRef]
- Dill, K.A.; Shortle, D. Denatured state of proteins. Annu. Rev. Biochem. 1991, 60, 795–825. [Google Scholar] [CrossRef]
- Fiebig, K.M.; Schwalbe, H.; Buck, M.; Smith, L.J.; Dobson, C.M. Toward a description of the conformations of denatured states of proteins. Comparison of a random coil model with NMR measurements. J. Phys. Chem. 1996, 100, 2661–2666. [Google Scholar] [CrossRef]
- Schwalbe, H.; Fiebig, K.M.; Buck, M.; Jones, J.A.; Grimshaw, S.B.; Spencer, A.; Glaser, S.J.; Smith, L.J.; Dobson, C.M. Structural and dynamical properties of a denatured protein. Heteronuclear 3D NMR experiments and theoretical simulations of lysozyme in 8 M urea. Biochemistry 1997, 36, 8977–8991. [Google Scholar] [CrossRef]
- Smith, L.J.; Bolin, K.A.; Schwalbe, H.; MacArthur, M.W.; Thornton, J.M.; Dobson, C.M. Analysis of main chain torsion angles in proteins: Predicition of nmr coupling constants for native and random coil conformations. J. Mol. Biol. 1996, 255, 494–506. [Google Scholar] [CrossRef]
- Bai, Y.; Chung, J.; Dyson, H.J.; Wright, P.E. Structural and dynamic characterization of an unfolded state of polar apo-plastocyanin formed under denaturing conditions. Protein Sci. 2001, 10, 1056–1066. [Google Scholar] [CrossRef]
- Chandrasekar, K.; Profy, A.T.; Dyson, H.J. Solution conformational preferences of immunogenic peptides derived from the principal neutralizing determinant of the HIV-1 envelope glycoprotein gp 120. Biochemistry 1991, 30, 9187–9194. [Google Scholar]
- Dyson, H.J.; Wright, P.E. Peptide conformation and protein folding. Curr. Opin. Struct. Biol. 1993, 3, 60–65. [Google Scholar]
- Dyson, H.J.; Wright, P.E. Unfolded proteins and protein folding studied by NMR. Chem. Rev. 2004, 104, 3607–3622. [Google Scholar]
- Jensen, M.R.; Salmon, L.; Nodet, G.; Blackledge, M. Defining conformational ensembles of intrinsically disordered and partially folded proteins directly from chemical shifts. J. Am. Chem. Soc. 2010, 132, 1270–1272. [Google Scholar]
- Meier, S.; Grzesiek, S.; Blackledge, M. Mapping the conformational landscape of urea-denaturated ubiquitin using residual dipolar couplings. J. Am. Chem. Soc. 2007, 129, 9799–9807. [Google Scholar]
- Bruschweiler, R.; Blackledge, M.; Ernst, R.R. Multi-conformational peptide dynamics derived from nmr data: A new search algorithm and its application to antamanide. J. Biomol. NMR 1991, 1, 3–11. [Google Scholar]
- Silvers, R.; Sziegat, F.; Tachibana, H.; Segawa, S.-I.; Whittaker, S.; Günther, U.; Gabel, F.; Huang, J.; Blackledge, M.; Wirmer-Bartoschek, J. Modulation of structure and dynamics by disulfide bond formation in unfolded states. J. Am. Chem. Soc. 2012, 134, 6846–6854. [Google Scholar] [CrossRef]
- Sziegat, F.; Silvers, R.; Hähnke, M.J.; Jensen, M.R.; Blackledge, M.; Wirmer-Bartoschek, J.; Schwalbe, H. Disentangling the coil: Modulation of conformationa and dynamic properties by site-directed mutation in the non-native state of hen egg white lysozyme. Biochemistry 2012, 51, 3361–3372. [Google Scholar] [CrossRef]
- Shi, Z.; Shen, K.; Liu, Z.; Kallenbach, N.R. Conformation in the backbone in unfolded proteins. Chem. Rev. 2006, 106, 1877–1897. [Google Scholar] [CrossRef]
- Shi, Z.; Olson, C.A.; Rose, G.D.; Baldwin, R.L.; Kallenbach, N.R. Polyproline II structure in a sequence of seven alanine residues. Proc. Natl. Acad. Sci. USA 2002, 99, 9190–9195. [Google Scholar]
- Shi, Z.; Chen, K.; Liu, Z.; Ng, A.; Bracken, W.C.; Kallenbach, N.R. Polyproline II propensities from GGXGG peptides reveal an anticorrelation with β-sheet scales. Proc. Natl. Acad. Sci. USA 2005, 102, 17964–17968. [Google Scholar] [CrossRef]
- Hagarman, A.; Measey, T.J.; Mathieu, D.; Schwalbe, H.; Schweitzer-Stenner, R. Intrinsic propensities of amino acid residues in GxG peptides inferred from amide I’ band profiles and NMR scalar coupling constants. J. Am. Chem. Soc. 2010, 132, 540–551. [Google Scholar]
- Cowan, P.M.; Mc Gavin, S. Structure of poly-L-proline. Nature 1955, 176, 501–503. [Google Scholar] [CrossRef]
- Shoulders, M.D.; Raines, R.T. Collagen structure and stability. Annu. Rev. Biochem. 2009, 78, 929–958. [Google Scholar] [CrossRef]
- Tiffany, M.L. Optical activity related to ordered aggregation in some biological molecules. 1. Poly(L-proline) and collagen. Int. J. Polym. Mater. 1976, 4, 293–302. [Google Scholar] [CrossRef]
- Tumanian, V.G. Conformation of a triple collagen spiral as a function of primary structure. BIOFIZIKA 1992, 37, 5–8. [Google Scholar]
- Tumanyan, V.G.; Abagyan, R.A.; Esipova, N.G. Conformational analysis of polytripeptides (Gly-Pro-Ala)n, (Gly-Ala-Hyp)n, and (Gly-Ala-Ala)n in connection with the problem of collagen structure. Biopolymers 1984, 23, 1499–1512. [Google Scholar] [CrossRef]
- Esipova, N.G.; Tumanyan, V.G.; Vlasova, A.V.; Vlasov, P.K. A tetrapeptide-based method for polyproline II-type secondary structure prediction. Proteins 2005, 61, 763–768. [Google Scholar] [CrossRef]
- Esipova, N.G.; Raqulina, L.E.; Davydova, L.I.; Lobachev, V.M.; Makeev, V.; Bogush, V.G.; Tumanyan, V.G.; Debabov, V.G. Left helix of polyproline II type and genesis of β-structures in spidroins 1 and 2 and their recombinant analogs. Biophysics 2009, 54, 271–274. [Google Scholar] [CrossRef]
- Chellgren, B.W.; Creamer, T.P. Short sequences of non-proline residues can adopt the polyproline II helical conformaiton. Biochemistry 2004, 43, 5864–5869. [Google Scholar] [CrossRef]
- Whittington, S.J.; Chellgren, B.W.; Herman, V.M.; Creamer, T.P. Urea promotes polyproline II helix formation: Implications for protein denatured states. Biochemistry 2005, 44, 6269–6275. [Google Scholar] [CrossRef]
- Rath, A.; Davidson, A.R.; Deber, C.M. The structure of “unstructured” regions in peptides and proteins: Role of the polyproline II helix in protein folding and recognition. Biopolymers 2005, 80, 179–185. [Google Scholar] [CrossRef]
- Bochiccio, B.; Tamburro, A.M. Polyproline II structure in proteins: Identification by chirotical spectroscopies, stability and function. Chirality 2002, 14, 782–792. [Google Scholar] [CrossRef]
- Blanch, E.W.; Gill, A.C.; Rhie, A.G.O.; Hope, J.; Hecht, L.; Nielsen, K.; Barron, L.D. Raman optical activity demonstrates poly(L-proline II helix in the N-terminal region of the ovine prion protein: Implications for function and misfunction. J. Mol. Biol. 2004, 343, 467–476. [Google Scholar] [CrossRef]
- Adzhubei, A.A.; Sternberg, M.J. Conservation of polyproline II helices in homologous proteins: Implications for structure prediction by model building. Protein Sci. 1994, 3, 2395–2410. [Google Scholar] [CrossRef]
- Zagrovic, B.; Lipfert, J.; Sorin, E.J.; Millett, I.S.; van Gunsteren, W.F.; Doniach, S.; Pande, V.S. Unusual compactness of a polyproline type II structure. Proc. Natl. Acad. Sci. USA 2005, 102, 11698–11703. [Google Scholar] [CrossRef]
- Kwac, K.; Cho, M. Molecular dynamics simulation study of N-methylacetamide in water I. Amide mode frequency fluctuations. J. Chem. Phys. 2003, 119, 2247–2255. [Google Scholar] [CrossRef]
- Ishizuka, R.; Huber, G.A.; McCammon, J.A. Solvation effect on the conformation of alanine dipeptide: Integral equation approach. J. Phys. Chem. Lett. 2011, 1, 2279–2283. [Google Scholar] [CrossRef]
- Makowska, J.; Rodziewicz-Motowidlo, S.; Baginska, K.; Vila, J.A.; Liwo, A.; Chmurzynski, L.; Scheraga, H.A. Polyproline II conformation is one of many local conformational states and is not an overall conformation of unfolded peptides and proteins. Proc. Natl. Acad. Sci. USA 2006, 103, 1744–1749. [Google Scholar] [CrossRef]
- Makowska, J.; Rodziewicz, S.; Baginska, K.; Makowski, M.; Vila, J.A.; Liwo, A.; Chmurzyňski, L.; Scheraga, H.A. Further evidence for the absence of polyrpoline II stretch in the XAO peptide. Biophys. J. 2007, 92, 2904–2917. [Google Scholar] [CrossRef]
- Best, R.B.; Hummer, G. Optimized molecular dynamics force fields applied to the helix-coil transition of polypeptides. J. Phys. Chem. B 2009, 113, 9004–9015. [Google Scholar] [CrossRef]
- Nerenberg, P.; Head-Gordon, T. Optimizing protein-solvent force fields to reproduce conformational preferences of model peptides. J. Chem. Theory Comp. 2011, 7, 1220–1230. [Google Scholar] [CrossRef]
- Avbelj, F.; Grdadolnik, S.G.; Grdadolnik, J.; Baldwin, R.L. Intrinsic backbone preferences are fully present in blocked amino acids. Proc. Natl. Acad. Sci. USA 2006, 103, 1272–1277. [Google Scholar] [CrossRef]
- Avbelj, F. Solvation and electrostatics as determined of local structural order in unfolded peptides and proteins. In Protein and Peptide Folding, Misfolding and Non-folding; Schweitzer-Stenner, R., Ed.; John Wiley & Sons: Hobocken, NJ, USA, 2012; pp. 131–158. [Google Scholar]
- Serrano, L. Comparison between the ϕ-distribution of the amino acids in the protein data base and NMR data indicates that amino acids have various ϕ propensities in the random coil conformation. J. Mol. Biol. 1995, 254, 322–333. [Google Scholar] [CrossRef]
- Grdadolnik, J.; Mohacek-Grosev, V.; Baldwin, R.L.; Avbelj, F. Populations of the three makor backbone conformations in 19 amino acid dipeptides. Proc. Natl. Acad. Sci. USA 2011, 108, 1794–1798. [Google Scholar] [CrossRef]
- Hagarman, A.; Mathieu, D.; Toal, S.; Measey, T.J.; Schwalbe, H.; Schweitzer-Stenner, R. Amino acids with hydrogen-bonding side chains have an intrinsic tendency to sample various turn conformations in aqueous solution. Chemistry 2011, 17, 6789–6797. [Google Scholar] [CrossRef]
- Rybka, K.; Toal, S.E.; Verbaro, D.J.; Mathieu, D.; Schwalbe, H.; Schweitzer-Stenner, R. Disorder and order in unfolded and disordered peptides and proteins: A view derived from tripeptide conformational analysis. II. Tripeptides with short side chains populating asx and beta-type like turn conformations. Proteins 2013, 81, 968–983. [Google Scholar] [CrossRef]
- Avbelj, F.; Baldwin, R.L. Role of backbone solvation and electrostatics in generating preferred peptide backbone conformations: Distributions of phi. Proc. Natl. Acad. Sci. USA 2003, 100, 5742–5747. [Google Scholar] [CrossRef]
- Jha, A.K.; Colubri, A.; Freed, K.F.; Sosnick, T.R. Statistical coil model of the unfolded state: Resolving the reconciliation problem. Proc. Natl. Acad. Sci. USA 2005, 102, 13099–13104. [Google Scholar]
- Jha, A.K.; Colubri, A.; Zaman, M.H.; Koide, S.; Sosnick, T.R.; Freed, K.F. Helix, sheet, and polyproline II frequencies and strong nearest neighbor effects in a restricted coil library. Biochemistry 2005, 44, 9691–9702. [Google Scholar] [CrossRef]
- Pappu, R.V.; Srinivasan, R.; Rose, G.D. The flory isolated-pair hypothesis is not valid for polypeptide chains: Implications for protein folding. Proc. Natl. Acad. Sci. USA 2000, 97, 12565–12570. [Google Scholar] [CrossRef]
- DeBartolo, J.; Jha, A.; Freed, K.F.; Sosnick, T.R. Local backbone preferences and nearest neighbor effects in the unfolded and native states. In Proteins and Peptides. Folding, Misfolding and Unfolding; Schweitzer-Stenner, R., Ed.; Wiley & Sons: Chichester, UK, 2012; pp. 79–98. [Google Scholar]
- Penkett, C.J.; Redfield, C.; Dodd, I.; Hubbard, J.; McBay, D.L.; Mossakowska, D.E.; Smith, R.A.; Dobson, C.M.; Smith, L.J. NMR analysis of main-chain conformational preferences in an unfolded fibronectin-binding protein. J. Mol. Biol. 1997, 274, 152–159. [Google Scholar] [CrossRef]
- Avbelj, F.; Baldwin, R.L. Origin of the neighboring residue effect on peptide backbone conformation. Proc. Natl. Acad. Sci. USA 2004, 101, 10967–10972. [Google Scholar] [CrossRef]
- Tiffany, M.L.; Krimm, S. New chain conformations of poly(glutamicacid) and polylysine. Biopolymers 1968, 6, 1379–1382. [Google Scholar] [CrossRef]
- Tiffany, M.L.; Krimm, S. Effect of temperature on the circular dichroism spectra of polypeptides in the extended state. Biopolymers 1972, 11, 2309–2316. [Google Scholar] [CrossRef]
- Woody, R.W. Circular dichroism of unordered polypeptides. Adv. Biophys. Chem. 1992, 2, 37–79. [Google Scholar]
- Woody, R.W. Circular dichroism spectrum of peptides in the poly(pro)II conformation. J. Am. Chem. Soc. 2009, 131, 8234–8245. [Google Scholar] [CrossRef]
- Sreerama, N.; Woody, R.W. Structural composition of βi and β2 proteins. Protein Sci. 2003, 12, 384–388. [Google Scholar] [CrossRef]
- Woody, R. The exciton model and the circular dichroism of polypeptides. Monatshefte Chem. 2005, 136, 347–366. [Google Scholar] [CrossRef]
- Gokce, I.; Woody, R.W.; Anderluh, G.; Lakey, J.H. Single peptide bonds exhibit poly(pro)II (“random coil”) circular dichroism spectra. J. Am. Chem. Soc. 2005, 127, 9700–9701. [Google Scholar]
- Mattice, W.L. The effect of temperature and salt concentration on the circular dichroism exhibited by unionized derivatives of L-alanine in aqueous solution. Biopolymers 1974, 13, 169–183. [Google Scholar] [CrossRef]
- Dukor, R.K.; Keiderling, T.A. Reassessment of the random coil conformation: Vibrational cd study of proline oligopeptides and related polypeptides. Biopolymers 1991, 31, 1747–1761. [Google Scholar] [CrossRef]
- Keiderling, T.A. Vibrational circular dichroism: Application to conformational analysis of biomolecules. In Circular Dichroism and the Conformational Analysis of Biomolecules; Fasman, G.D., Ed.; Plenumn Press: New York, NY, USA, 1996; p. 555. [Google Scholar]
- Keiderling, T.A.; Xu, Q. Unfolded proteins studied with IR and VCD spectra. Adv. Protein Chem. 2002, 62, 111–161. [Google Scholar] [CrossRef]
- Kubelka, J.; Huang, R.; Keiderling, T.A. Solvent effects on IR and VCD spectra of helical peptides: DFT-based static spectral simulations with explicit water. J. Phys. Chem. B 2005, 109, 8231–8243. [Google Scholar] [CrossRef]
- Kubelka, J.; Keiderling, T.A. The anomalous infrared amide I intensity distribution in 13C isotopically labeled peptide β-sheets comes from extended, multiple-stranded structures. An ab initio study. J. Am. Chem. Soc. 2001, 123, 6142–6150. [Google Scholar] [CrossRef]
- Kubelka, J.; Keiderling, T.A. Differentiation of β-sheet-forming structures: Ab initio-based simulations of IR absorption and vibrational CD For model peptide and protein β-sheets. J. Am. Chem. Soc. 2001, 123, 12048–12058. [Google Scholar] [CrossRef]
- Schweitzer-Stenner, R. Secondary structure analysis of polypeptides based on an excitonic coupling model to describe the band profile of amide I of IR, raman and vibrational circular dichroism spectra. J. Phys. Chem. B 2004, 108, 16965–16975. [Google Scholar] [CrossRef]
- Schweitzer-Stenner, R. Distribution of conformations sampled by the central amino acid residue in tripeptides inferred from amide I band profiles and nmr scalar coupling constants. J. Am. Chem. Soc. B 2009, 113, 2922–2932. [Google Scholar]
- Krimm, S.; Bandekar, J. Polypeptides, and proteins. Adv. Protein Chem. 1986, 38, 181–364. [Google Scholar] [CrossRef]
- Mirkin, N.G.; Krimm, S. Ab initio vibrational analysis of hydrogen-bonded trans- and cis-N-methylacetamide. J. Am. Chem. Soc. 1991, 113, 9742–9747. [Google Scholar] [CrossRef]
- Torii, H.; Tasumi, M. Ab initio molecular orbital study of the amide I vibrational interactions between the peptide groups in di- and tripeptides and considerations on the conformation of the extended helix. J. Raman Spectrosc. 1998, 29, 81–86. [Google Scholar] [CrossRef]
- Torii, H.; Tasumi, M. Model calculations on the amide-I infrared bands of globular proteins. J. Chem. Phys. 1992, 96, 3379–3387. [Google Scholar] [CrossRef]
- Torii, H.; Tasumi, M. Application of the three-dimensional doorway-state theory to analyses of the amide-I band of globular proteins. J. Chem. Phys. 1992, 97, 92–98. [Google Scholar] [CrossRef]
- Torii, H.; Tatsumi, T.; Kanazawa, T.; Tasumi, M. Effects of intramolecular hydrogen-bonding interactions on the amide I mode of N-methylacetamide: Matrix isolation infrared studies and Ab initio molecular orbital calculations. J. Phys. Chem. B 1998, 102, 309–314. [Google Scholar]
- Gorbunov, R.D.; Kosov, D.S.; Stock, G. Ab initio-based exciton model of amide I vibrations in peptides: Definition, conformational dependence and transferrability. J. Chem. Phys. 2005, 122, 224904–224915. [Google Scholar] [CrossRef]
- Scholz, J.M.; Baldwin, R.L. The mechanism of alpha helix formation of peptides. Annu. Rev. Biophys. Biomol. Struct. 1992, 21, 95–118. [Google Scholar] [CrossRef]
- Scholtz, J.M.; Marqusee, S.; Baldwin, R.L.; York, E.J.; Stewart, J.M.; Santoro, M.; Bolen, D.W. Calorimetric determination of the enthalpy change for the alpha-helix to coil transition of an alanine peptide in water. Proc. Natl. Acad. Sci. USA 1991, 88, 2854–2858. [Google Scholar] [CrossRef]
- Karplus, M. Theoretical calculation links NMR coupling constant to molecular geometry. J. Chem. Phys. 1959, 30, 11–15. [Google Scholar] [CrossRef]
- Wang, A.C.; Bax, A. Determination of the backbone dihedral angles ϕ in human ubiquitin from reparametrized empirical Karplus equations. J. Am. Chem. Soc. 1996, 118, 2483–2494. [Google Scholar] [CrossRef]
- Brahms, S.; Brahms, J.; Spach, G.; Brack, A. Identification of β,β-turns and unordered conformations in polypeptide chains by vacuum ultraviolet circular dichroism. Proc. Natl. Acad. Sci. USA 1977, 74, 3208–3212. [Google Scholar] [CrossRef]
- Woutersen, S.; Hamm, P. Structure determination of trialanine in water using polarized sensitive two-dimensional vibrational spectroscopy. J. Phys. Chem. B 2000, 104, 11316–11320. [Google Scholar] [CrossRef]
- Woutersen, S.; Hamm, P. Isotope-edited two-dimensional vibrational spectroscopy of trialanine in aqueous solution. J. Chem. Phys. 2001, 114, 2727–2737. [Google Scholar] [CrossRef]
- Woutersen, S.; Pfister, R.; Hamm, P.; Mu, Y.; Kosov, D.S.; Stock, G. Peptide conformational heterogeneity revealed from nonlinear vibrational spectroscopy and molecular-dynamics simulations. J. Chem. Phys. 2002, 117, 6833–6840. [Google Scholar] [CrossRef]
- Eker, F.; Cao, X.; Nafie, L.; Huang, Q.; Griebenow, K.; Schweitzer-Stenner, R. The structure of alanine based tripeptides in water and dimethyl sulfoxide probed by vibrational spectroscopy. J. Phys. Chem. B 2003, 107, 358–365. [Google Scholar]
- Eker, F.; Cao, X.; Nafie, L.; Schweitzer-Stenner, R. Tripeptides adopt stable structures in water. A combined polarized visible Raman, FTIR, and VCD spectroscopy study. J. Am. Chem. Soc. 2002, 124, 14330–14341. [Google Scholar] [CrossRef]
- Eker, F.; Griebenow, K.; Cao, X.; Nafie, L.A.; Schweitzer-Stenner, R. Preferred peptide backbone conformations in the unfolded state revealed by the structure analysis of alanine-based (AXA) tripeptides in aqueous solution. Proc. Natl. Acad. Sci. USA 2004, 101, 10054–10059. [Google Scholar] [CrossRef]
- Graf, J.; Nguyen, P.H.; Stock, G.; Schwalbe, H. Structure and dynamics of the homologous series of alanine peptides: A joint molecular dynamics/nmr study. J. Am. Chem. Soc. 2007, 129, 1179–1189. [Google Scholar] [CrossRef]
- Han, W.-G.; Jakanen, K.J.; Elstner, M.; Suhai, S. Theoretical study of aqueous N-acetyl-L-alanine N-methylamide: Structures and Raman, VCD, and ROA spectra. J. Phys. Chem. B 1998, 102, 2587–2602. [Google Scholar] [CrossRef]
- Poon, C.D.; Samulsi, E.T.; Weise, C.F.; Weisshaar, J.C. Do bridging water molecules dictate the structure of a model dipeptide in aqueous solution? J. Am. Chem. Soc. 2000, 122, 5612–5613. [Google Scholar]
- Weise, C.F.; Weisshaar, J.C. Conformational analysis of alanine dipeptide from dipolar couplings in a water-based liquid crystal. J. Phys. Chem. B 2003, 107, 3265–3277. [Google Scholar] [CrossRef]
- Kim, Y.S.; Wang, J.; Hochstrasser, R.M. Two-dimensional infrared spectroscopy of the alanine dipeptide in aqueous solution. J. Phys. Chem. B 2005, 109, 7511–7521. [Google Scholar] [CrossRef]
- Feig, M. Is alanine dipeptide a good model for representing the torsional preferences of protein backbones. J. Chem. Theory Comput. 2008, 4, 1555–1564. [Google Scholar] [CrossRef]
- Garcia-Pietro, F.F.; Galván, I.F.; Aguliar, M.A.; Martin, M.E. Study on the conformational equilibrium of the alanine dipeptide in water solution by using the averaged solvent electrostatic potential from molecular dynamics methodology. J. Chem. Phys. 2011. [Google Scholar] [CrossRef]
- Hu, H.; Elstner, M.; Hermans, J. Comparison of a QM/MM force field and molecular mechanics force fields in simulations of alanine and glycine “dipeptides” (Ace-Ala-Nme and Ace-Gly-Nme) in water in relation to the problem of modeling the unfolded peptide backbone in solution. Proteins 2003, 50, 451–463. [Google Scholar] [CrossRef]
- Ioannou, F.; Archontis, G.; Leontidis, E. Specific interactions of sodium salts with alanine dipeptide and tetrapeptide in water: Insights from molecular dynamics. J. Phys. Chem. B 2011, 115, 13389–13400. [Google Scholar] [CrossRef]
- Kaminski, G.A.; Friesner, R.A.; Tirado-Rives, J.; Jorgensen, W.L. Evaluation and reparametrization of the OPLS-AA force field for proteins via comparison with accurate quantum chemical calculations on peptides. J. Phys. Chem. B 2001, 105, 6474–6487. [Google Scholar] [CrossRef]
- McColl, I.H.; Blanch, E.W.; Hecht, L.; Kallenbach, N.R.; Barron, L.D. Vibrational Raman optical activity characterization of poly(L-proline) II helix in alanine oligopeptides. J. Am. Chem. Soc. 2004, 126, 5076–5077. [Google Scholar]
- Mikhonin, A.V.; Ahmed, Z.; Ianoul, A.; Asher, S.A. Assignments and conformational dependencies of the amide iii peptide backbone UV resonance Raman bands. J. Phys. Chem. B 2004, 108, 19020–19028. [Google Scholar] [CrossRef]
- Mikhonin, A.V.; Asher, S.A. Uncoupled peptide bond vibrations in a α-helical and polyproline II conformation of polyalanine peptides. J. Phys. Chem. B 2005, 109, 3047–3052. [Google Scholar] [CrossRef]
- Schweitzer-Stenner, R.; Eker, F.; Griebenow, K.; Cao, X.; Nafie, L. The conformation of tetraalanine in water determined by polarized raman, ftir and VCD spectroscopy. J. Am. Chem. Soc. 2004, 126, 2768–2776. [Google Scholar] [CrossRef]
- Tanaka, S.; Scheraga, H.A. Statistical mechanical treatment of protein conformation. I. Conformational properties of amino acids in proteins. Macromolecules 1976, 9, 142–157. [Google Scholar] [CrossRef]
- Tanaka, S.; Scheraga, H.A. Statistical mechanical treatment of protein conformation. II. A three-state model for specific-sequence copolymers of amino acids. Macromolecules 1976, 9, 150–167. [Google Scholar]
- Schweitzer-Stenner, R.; Measey, T. The alanine-rich XAO peptide adopts a heterogeneous population, including turn-like and PpII conformations. Proc. Natl. Acad. Sci. USA 2007, 104, 6649–6654. [Google Scholar] [CrossRef]
- Ding, L.; Chen, K.; Santini, P.A.; Shi, Z.; Kallenbach, N.R. The pentapeptide GGAGG has pII conformation. J. Am. Chem. Soc. 2003, 125, 8092–8093. [Google Scholar] [CrossRef]
- Schweitzer-Stenner, R.; Hagarman, A.; Toal, S.; Mathieu, D.; Schwalbe, H. Disorder and order in unfolded and disordered peptides and proteins: A view derived from tripeptide conformational analysis. I. Tripeptides with long and predominantly hydrophobic side chains. Proteins 2013, 81, 955–967. [Google Scholar] [CrossRef]
- He, L.; Navarro, A.E.; Shi, Z.; Kallenbach, N.R. End effects influence short model peptide conformation. J. Am. Chem. Soc. 2011, 134, 1571–1576. [Google Scholar]
- Schweitzer-Stenner, R.; Eker, F.; Huang, Q.; Griebenow, K. Dihedral angles of trialanine in D2O determined by combining ftir and polarized visible raman spectroscopy. J. Am. Chem. Soc. 2001, 123, 9628–9633. [Google Scholar] [CrossRef]
- Oh, K.-I.; Lee, K.-K.; Park, E.-K.; Jung, Y.-S.; Hwang, G.-S.; Cho, M. A comprehensive library of blocked dipeptides reveals intrinsic backbone conformational propensities of unfolded proteins. Proteins 2012, 80, 977–990. [Google Scholar] [CrossRef]
- Cruz, V.; Ramos, J.; Martinez-Salazar, J. Water-mediated conformations of the alanine dipeptide as revealed by distributed umbrella sampling simulations, quantum mechanics based calculations, and experimental data. J. Phys. Chem. B 2011, 115, 4880–4886. [Google Scholar] [CrossRef]
- Kwac, K.; Lee, K.K.; Han, J.B.; Oh, K.I.; Cho, M. Classical and quantum mechanical/molecular mechanical molecular dynamics simulations of alanine dipeptide in water: Comparisons with IR and vibrational circular dichroism spectra. J. Chem. Phys. 2008. [Google Scholar] [CrossRef]
- Beck, D.A.C.; Alonso, D.O.V.; Inoyama, D.; Dagget, V. The intrinsic conformational propensities of the 20 naturally occuring amino acids and reflection of these propensities in proteins. Proc. Natl. Acad. Sci. USA 2008, 105, 12259–12264. [Google Scholar] [CrossRef]
- Drozdov, A.N.; Grossfield, A.; Pappu, R.V. Role of solvent in determining conformational preferences of alanine dipeptide in water. J. Am. Chem. Soc. 2004, 126, 2574–2581. [Google Scholar] [CrossRef]
- Gnanakaran, S.; Garcia, A.E. Helix-coil transition of alanine peptides in water: Force field dependence on the folded and unfolded structures. Proteins 2005, 59, 773–782. [Google Scholar] [CrossRef]
- Best, R.B.; Buchete, N.V.; Hummer, G. Are current molecular dynamics force fields too helical? Biophys. J. 2008, 95, L07–L09. [Google Scholar]
- Mu, Y.; Kosov, D.S.; Stock, G. Conformational dynamics of trialanine in water. 2. Comparison of amber, charmm, gromos, and opls force fields to NMR and infrared experiments. J. Phys. Chem. B 2003, 107, 5064–5073. [Google Scholar] [CrossRef]
- Gnanakaran, S.; Garcia, A.E. Validation of an all-atom protein force field: From dipeptides to larger peptides. J. Phys. Chem. B 2003, 107, 12555–12557. [Google Scholar] [CrossRef]
- Garcia, A.E. Characterization of non-alpha conformations in Ala peptides. Polymer 2004, 120, 885–890. [Google Scholar]
- Kentsis, A.; Mezei, M.; Gindin, T.; Osman, R. Unfolded state of polyalanine is a segmented polyproline II helix. Proteins 2004, 55, 493–501. [Google Scholar] [CrossRef]
- Zaman, M.H.; Shen, M.-Y.; Berry, R.S.; Freed, K.F.; Sosnick, T.R. Investigations into sequence and conformational dependence of backbone entropy, inter-basin dynamics and the flory isolated-pair hypothesis for peptides. J. Mol. Biol. 2003, 331, 693–711. [Google Scholar]
- Verbaro, D.; Ghosh, I.; Nau, W.M.; Schweitzer-Stenner, R. Discrepancies between conformational distributions of a polyalanine peptide in solution obtained from molecular dynamics force fields and amide I’ band profiles. J. Phys. Chem. B 2010, 114, 17201–17208. [Google Scholar] [CrossRef]
- Duan, Y.; Wu, C.; Chowdury, S.; Lee, M.C.; Xiong, G.; Zhang, W.; Yang, R.; Cieplak, P.; Luo, R.; Lee, T.; et al. A point-charge force field for molecular mechanics simulations of protein based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 2003, 24, 1999–2012. [Google Scholar] [CrossRef]
- Lanza, G.; Chiacchio, M.A. Comprehensive and accurate ab initio energy surface of simple alanine peptides. Chem. Phys. Chem. 2013, 14, 3284–3293. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Berendsen, H.J.C.; Grigera, J.R.; Straatsma, T.P. The missing term in effective pair potentials. J. Phys. Chem. 1987, 91, 6269–6271. [Google Scholar] [CrossRef]
- Mahoney, M.W.; Jorgensen, W.L. A five-site model for liquid water and the reproduction of the density anomaly by rigid, nonpolarizable potential functions. J. Chem. Phys. 2000, 112, 8910–8922. [Google Scholar] [CrossRef]
- Horn, H.W.; Swope, W.C.; Pitera, J.W.; Madura, J.D.; Dick, T.J.; Hura, G.L.; Head-Gordon, T. Development of an improved four-site water model for biomolecular simulations: TIP4P-EW. J. Chem. Phys. 2004, 120, 9665–9678. [Google Scholar] [CrossRef]
- Wickstrom, L.; Okur, A.; Simmerling, C. Evaluating the performance of the FF99SB force field based on nmr scalar coupling data. Biophys. J. 2009, 97, 853–856. [Google Scholar] [CrossRef]
- Ren, P.; Ponder, J.W. Polarizable atomic multipole water model for molecular mechanics simulation. J. Phys. Chem. B 2003, 107, 5933–5947. [Google Scholar]
- Dixon, R.W.; Kollman, P.A. Advancing beyond the atom-centered model in additive and nonadditive molecular mechanics. J. Comput. Chem. 1997, 18, 1632–1646. [Google Scholar] [CrossRef]
- Rucker, A.L.; Pager, C.T.; Campbell, M.N.; Qualls, J.E.; Creamer, T.P. Host-guest scale of left-handed polyproline II helix formation. Proteins 2003, 53, 68–75. [Google Scholar] [CrossRef]
- Kelly, M.A.; Chellgren, B.W.; Rucker, A.L.; Troutman, J.M.; Fried, M.G.; Miller, A.F.; Creamer, T.P. Host-guest study of left-handed polyproline II helix formation. Biochemistry 2001, 40, 14376–14383. [Google Scholar] [CrossRef]
- Moradi, M.; Babin, V.; Sagui, C.; Roland, C. A statistical analysis of the PPII propensity of amino acid guests in proline-rich peptides. Biophys. J. 2011, 100, 1083–1093. [Google Scholar] [CrossRef]
- Moradi, M.; Babin, V.; Celeste, S.; Roland, C. PPII propensity of multiple-guest amino acids in a proline-rich environment. J. Phys. Chem. B 2011, 115, 8645–8656. [Google Scholar] [CrossRef]
- Toal, S.E.; Verbaro, D.J.; Schweitzer-Stenner, R. Role of enthalpy-entropy compensation interactions in determining the conformational propensities of amino acid residues in unfolded peptides. J. Phys. Chem. B 2014, 118, 1309–1318. [Google Scholar]
- Mertuka, G.; Dyson, H.J.; Wright, P.E. “Random coil” 1H chemical shifts obtained as a function of temperature and trifluoroethanol concentration for the peptide series GGXGG. J. Biomol. NMR 1995, 5, 14–24. [Google Scholar] [CrossRef]
- Plaxco, K.W.; Morton, C.J.; Grimshaw, S.B.; Jones, J.A.; Pitkeathly, M.; Campbell, I.D.; Dobson, C.M. The effects of guanidine hydrochloride on the “random coil” conformations and nmr chemical shifts of the peptide series GGXGG. J. Biomol. NMR 1997, 10, 221–230. [Google Scholar] [CrossRef]
- Schwarzinger, S.; Kroon, G.J.A.; Foss, T.R.; Chung, J.; Wright, P.E.; Dyson, H.J. Sequence-dependent correction of random coil NMR chemical shifts. J. Am. Chem. Soc. 2001, 123, 2970–2978. [Google Scholar]
- Van der Velde, F.; Pereira, L.; Rollema, H.S. The revised NMR chemical shift data of carrageenans. Carbohydr. Res. 2004, 29, 813–822. [Google Scholar]
- Wishart, D.S.; Sykes, B.D. Chemical shifts as a tool for structure determination. Methods Enzymol. 1994, 239, 363–392. [Google Scholar] [CrossRef]
- Swindells, M.B.; MacArthur, M.W.; Thornton, J.M. Intrinsic ϕ,ψ propensities of amino acids, derived from the coil regions of known structures. Nat. Struct. Biol. 1995, 2, 596–603. [Google Scholar] [CrossRef]
- Sosnick, T.R. Sampling Library. Available online: http://godzilla.uchicago.edu/cgi-bin/rama.cgi/ (accessed on 1 February 2014).
- Xu, C.; Wang, J.; Liu, H. A hamiltonian replica exchange approach and its application to the study of side-chain type and neighbor effects on peptide backbone conformations. J. Chem. Theory Comput. 2008, 4, 1348–1359. [Google Scholar] [CrossRef]
- Ohkubo, Y.Z.; Brooks, C.L. Exploring flory’s isolated-pair hypothesis: Statistical mechanics of helix-coil transitions in polyalanine and the C-peptide from RNase A. Proc. Natl. Acad. Sci. 2003, 100, 13916–13921. [Google Scholar] [CrossRef]
- Zimm, B.H.; Bragg, J.K. Theory of the phase transition between helix and random coil. J. Chem. Phys. 1959, 31, 526–535. [Google Scholar] [CrossRef]
- Baxa, M.C.; Haddadain, E.J.; Jha, A.K.; Fred, K.F.; Sosnick, T.R. Context and firce field dependence of the loss of protein backbone entropy upon folding using realistic denatured and native state ensembles. J. Am. Chem. Soc. 2012, 134, 15929–15936. [Google Scholar] [CrossRef]
- Verbaro, D.; Mathieu, D.; Toal, S.E.; Schwalbe, H.; Schweitzer-Stenner, R. Ionized trilysine: A model system for understanding the nonrandom structure of poly-L-lysine and lysine-containing motifs in proteins. J. Phys. Chem. B 2012, 116, 8084–8094. [Google Scholar] [CrossRef]
- Duitch, L.; Toal, S.; Measey, T.J.; Schweitzer-Stenner, R. Triaspartate: A model system for conformationally flexible DDD motifs in proteins. J. Phys. Chem. B 2012, 116, 5160–5171. [Google Scholar] [CrossRef]
- Pizzanelli, S.; Forte, C.; Monti, S.; Zandomeneghi, G.; Hagarman, A.; Measey, T.J.; Schweitzer-Stenner, R. Conformation of phenylalanine in the trpeptides AFA and GFG probed by combining MD simulations with NMR, FTIR, polarized raman and VCD spectroscopy. J. Phys. Chem. B 2010, 114, 3965–3978. [Google Scholar] [CrossRef]
- Oh, K.-I.; Jung, Y.-S.; Hwang, G.-S.; Cho, M. Conformational distributions of denaturated and unstructured proteins are similar to those of 20 × 20 blocked dipeptides. J. Biomol. NMR 2012, 53, 25–41. [Google Scholar] [CrossRef]
- Jung, Y.-S.; Oh, K.-I.; Hwang, G.-S.; Cho, M. Neighboring residues effects in terminally blocked dipeptides: Implications for residual secondary structures in intrinsically unfolded/disordered proteins. Chirality 2014. [Google Scholar] [CrossRef]
- Chen, K.; Liu, Z.; Zhou, C.; Shi, Z.; Kallenbach, N.R. Neighbor effect on PPII conformation in alanine peptides. J. Am. Chem. Soc. 2005, 127, 10146–10147. [Google Scholar]
- Schweitzer-Stenner, R. Different degrees of disorder in long disordered peptides can be discriminated by vibrational spectroscopy. J. Phys. Chem. B 2013, 117, 6927–6936. [Google Scholar] [CrossRef]
- Toal, S.; Omidi, A.; Schweitzer-Stenner, R. Conformational changes of trialanine induced by direct interactions between alanine residues and alcohols in binary mixtures of water with glycerol and ethanol. J. Am. Chem. Soc. 2011, 133, 12728–12739. [Google Scholar] [CrossRef]
- Schwalbe, M.; Ozenne, V.; Bibow, S.; Jaremko, M.; Jaremko, L.; Gajda, M.; Jensen, M.R.; Biernat, J.; Mandelkow, E.; Zweckstetter, M.; et al. Predictive atomic resolution descriptions of intrinsically disordered hTau40 and α-synuclein in solution from NMR and small angle scattering. Structure 2014, 22, 1–12. [Google Scholar]
- Barron, L.D.; Blanch, E.W.; Hecht, L. Unfolded proteins studied by Raman optical activity. Adv. Protein Chem. 2002, 62, 51–90. [Google Scholar] [CrossRef]
- Barron, L.D.; Hecht, L.; Blanch, E.W.; Bell, A.F. Solution structure and dynamics of biomolecules from Raman optical activity. Prog. Biophys. Mol. Biol. 2000, 73, 1–49. [Google Scholar] [CrossRef]
- Blanch, E.W.; Morozowa-Roche, L.A.; Cochran, D.A.E.; Doig, A.J.; Hecht, L.; Barron, L.D. Is polyproline II helix the killer conformation? A raman optical activity study of the amyloidogenic prefibrillar intermediate of human lysozyme. J. Mol. Biol. 2000, 301, 553–563. [Google Scholar] [CrossRef]
- Danielsson, J.; Jarvet, J.; Damberg, P.; Gräslund, A. The alzheimer β-peptide shows temperature-dependent transitions between left-handed 31-helix, β-strand and random coil secondary structure. FEBS J. 2005, 272, 3938–3949. [Google Scholar] [CrossRef]
- Schweitzer-Stenner, R.; Measey, T.; Hagarman, A.; Eker, F.; Griebenow, K. Salmon calcitonin and amyloid β: Two peptides with amyloidogenic capacity adopt different conformational manifolds in their unfolded states. Biochemistry 2006, 45, 2810–2819. [Google Scholar] [CrossRef]
- Fitzkee, N.C.; Rose, G.D. Reassessing random-coil statistics in unfolded proteins. Proc. Natl. Acad. Sci. USA 2004, 101, 12497–12502. [Google Scholar] [CrossRef]
- Kohn, J.E.; Millett, I.S.; Jacob, J.; Zagrovic, B.; Dillon, T.M.; Cingel, N.; Dothager, R.S.; Seifer, S.; Thiyagarajan, R.; Sosnick, T.R.; et al. Random-coil behavior and the dimensions of chemically unfolded proteins. Proc. Natl. Acad. Sci. USA 2004, 101, 12491–12496. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Toal, S.; Schweitzer-Stenner, R. Local Order in the Unfolded State: Conformational Biases and Nearest Neighbor Interactions. Biomolecules 2014, 4, 725-773. https://doi.org/10.3390/biom4030725
Toal S, Schweitzer-Stenner R. Local Order in the Unfolded State: Conformational Biases and Nearest Neighbor Interactions. Biomolecules. 2014; 4(3):725-773. https://doi.org/10.3390/biom4030725
Chicago/Turabian StyleToal, Siobhan, and Reinhard Schweitzer-Stenner. 2014. "Local Order in the Unfolded State: Conformational Biases and Nearest Neighbor Interactions" Biomolecules 4, no. 3: 725-773. https://doi.org/10.3390/biom4030725
APA StyleToal, S., & Schweitzer-Stenner, R. (2014). Local Order in the Unfolded State: Conformational Biases and Nearest Neighbor Interactions. Biomolecules, 4(3), 725-773. https://doi.org/10.3390/biom4030725





