Mitochondrial Aging in the CNS: Unravelling Implications for Neurological Health and Disease
Abstract
1. Introduction
2. Mitochondrial Quality Control During Aging in CNS
2.1. Decline in Mitophagy During Aging
2.2. Dysregulation of Mitochondrial Dynamics During Aging in CNS
3. Energy Demands of the CNS During Aging
4. Mitochondrial ROS Production and Oxidative Stress in the CNS
5. Mitochondrial DNA Mutations During Aging
6. Mitochondrial-Nuclear Communication and How This Affects Health and Neuronal Function During Aging
6.1. Mitochondria–Nuclear Communication
6.2. Mitochondrial–ER, Mitochondrial–Peroxisome, and Mitochondrial–Lysosomes Communication in Aging
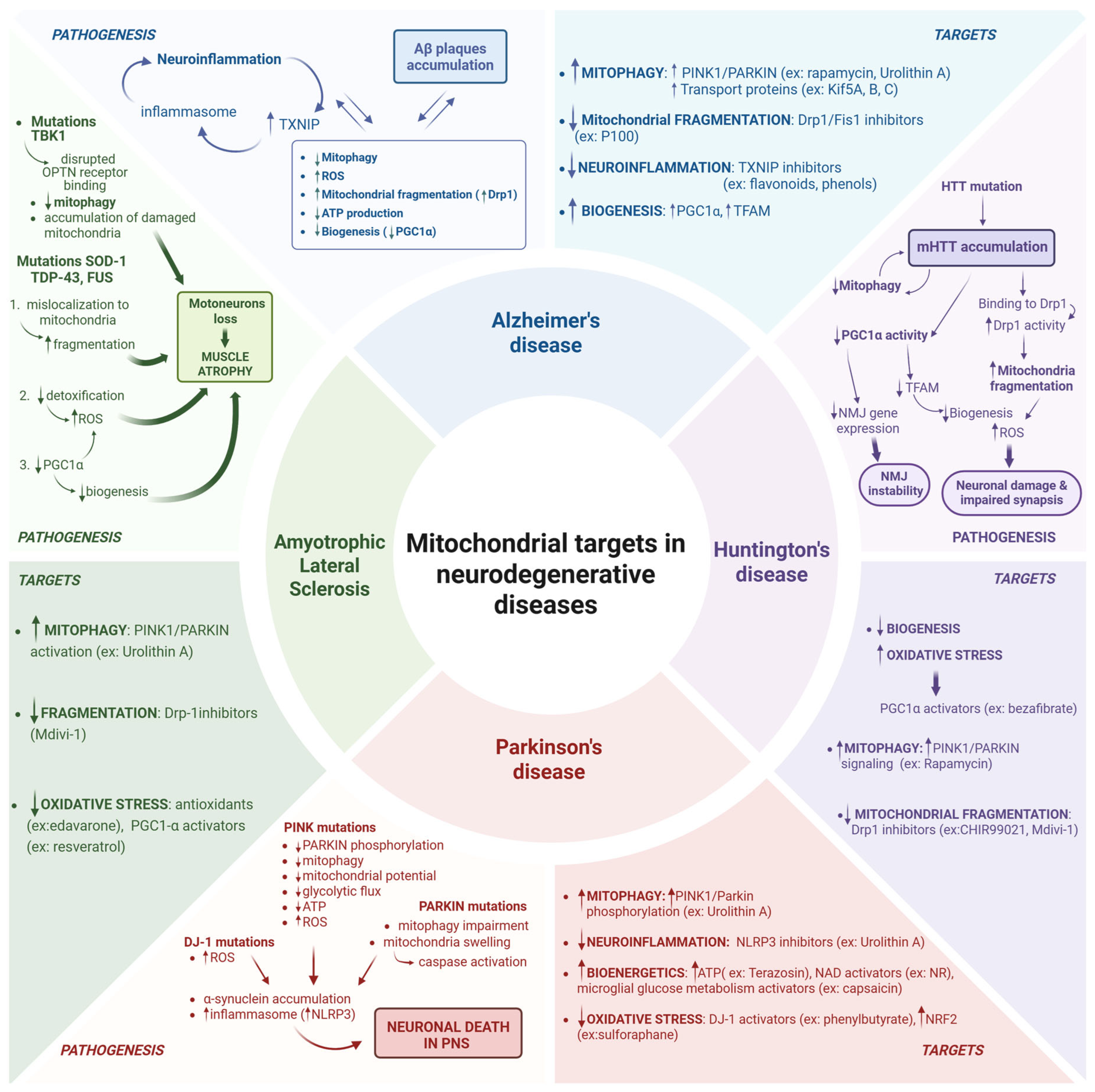
7. Neurodegenerative Diseases and Mitochondrial Molecular Targets
7.1. Alzheimer’s Disease (AD)
7.1.1. Enhancing Mitophagy
- A.
- Activation of the PINK1/PARKIN pathway: Upregulation of PINK1 or PARKIN content has shown neuroprotective effects and therapeutic potential in the treatment of AD [163]. In microglia, it was demonstrated to significantly reduce neuroinflammation and the levels of insoluble Aβ content, ameliorating the cognitive deficits in the AD mouse model [24]. Importantly, restoration of mitophagy by oral administration of drugs such as Urolithin A and Rapamycin, which promote mitophagy, were effective in improving the cognitive deficits in both the nematode and rodent models of AD, decreasing Aβ plaques, and promoting the elimination of tau hyperphosphorylation [25,164,165].
- B.
- Enhancing mitochondrial transport proteins: A recent study found that in AD, the presence of Aβ oligomers (Aβo) and amyloid precursor protein-C terminal fragments (APP-CTFs) alters the expression of several mitochondrial transport proteins, including SNPH and Miro1; adapter proteins such as TRANK1 and TRAK2; and components of the dynein and kinesin motor complexes, including Kif5A, Kif5B, and Kif5C. These proteins are crucial for axonal mitochondrial transport supporting mitophagy. Disruptions in their function contribute to impaired mitophagy and disease progression and restoring those genes showed improvement in AD pathogenesis [166,167].
7.1.2. Decreasing Mitochondria Fragmentation
- A.
- Inhibition of Fis1 and Drp1: Biochemical assessments of peripheral blood from AD patients showed significantly high levels of the mitochondrial fission factor Fis1, with post-mortem brain specimens further revealing upregulated Drp1 expression in AD cases [170,171]. P110 treatment disrupts Drp1/Fis1 interaction without affecting the interaction of Drp1 with its other adaptors, which results in reduced Aβ deposition and improved behavioral deficits [172].
- B.
- Promotion of mitochondria fusion via OPA1 and Mfn2: Han et al. recently revealed a marked downregulation of mitochondrial fusion regulators OPA1 and Mfn2 in AD groups compared to non-AD controls [173].
- C.
- Reducing neuroinflammation: The higher expression of Drp1 and TXNIP during AD are also involved in early inflammatory responses in oligodendrocytes and microglia of AD animal models, triggering NLRP3 inflammasome activation and caspase-3 cleavage, which, in turn, amplify neuroinflammatory cascades [174,175]. These pathological processes exacerbate Aβ deposition and tau-mediated neurodegeneration, ultimately leading to neuronal death or functional deficits. Some flavonoids and phenols (e.g., rosmarinic acid, rutic, puerarin) have been proposed as potent TXNIP inhibitors [176].
7.1.3. Enhancing Biogenesis
- A.
- Activation of PGC-1α: PGC-1α protein levels are inversely proportional to the concentration of Aβ in the CNS in AD making it a promising therapeutic target for AD [178]. Consistently, the overexpression of PGC-1α via adeno-associated virus in the brain of the AD murine transgenic model APP23 is sufficient to enhance the transcription of growth factors and to blunt Aβ-mediated neuroinflammation and neuronal death, leading to reduced-amyloid production and neuronal loss [179], together with decreased mitochondrial damage, thus restoring AD cognitive deficits [180].
- B.
- Enhancing TFAM: Human mitochondrial transcription factor A (hTFAM) activation was shown to protect mtDNA, reduce oxidative damage and intracellular Aβ, and improve cognitive function in both mouse and human AD models, making it a promising therapeutic target [181].
7.2. Parkinson’s Disease (PD)
7.2.1. Boosting Mitophagy
- A.
- Activation of PINK/Parkin signaling pathway for preventing α-syn accumulation: The pathological accumulation of phosphorylated α-syn is a key contributor to age-dependent neurodegeneration in PD [183]. In aging primates’ brains, reduced expression of Parkin and a decline in its PINK1-dependent phosphorylation contribute to substantia nigra neurodegeneration. Phosphorylated Parkin plays a critical role in neuroprotection by promoting the clearance of α-syn. Indeed, reintroducing wild-type (WT) Parkin expression, which is capable of being phosphorylated by PINK1, reduced the accumulation of pα-syn [16]. This research highlights the crucial role of Parkin phosphorylation in the development of PD, pointing to it as a potential avenue for new treatment strategies.
- B.
- Promoting mitophagy in microglia and reducing neuroinflammation through Urolithin A (UA): Microglial-mediated neuroinflammation and mitophagy are closely interconnected. In PD murine models, neuroinflammasome pathways, particularly NLRP3, are highly activated, a response amplified by ROS generated from dysfunctional mitochondria. Urolithin A (UA), a naturally occurring compound, has been shown to promote mitophagy, restore mitochondrial function, and suppress NLRP3 inflammasome activation in PD mouse models [192]. UA administration in aged humans also effectively reduced neuroinflammation markers, ameliorated mitochondrial function, and improved muscle strength and performance, suggesting its potential as a therapeutic strategy [193,194].
7.2.2. Boosting Bioenergetics
- A.
- Rescue microglial energy metabolism: Considering the major role of microglial glucose metabolism in neuroinflammation, manipulating microglial glucose metabolism has been proposed as a potential therapeutic approach. Compounds such as capsaicin preserve dopamine neurons degeneration, blocking the activity of glial cells in inducing oxidative stress and neuroinflammation [196]. Particularly, capsaicin rescues microglial energy metabolism by increasing TRPV1 and suppressing NADPH oxidase-driven ROS production. This protects nigrostriatal dopaminergic neurons via inhibition of glial activation-mediated oxidative stress.
- B.
- Increasing NAD+ levels: A significant decline in the NAD+/NADH ratio supports the reduction in ETC enzyme activity in mitochondria during aging and in pathologies such as PD. A Phase I trial (NCT03816020) with nicotinamide riboside (NR) showed elevated NAD+ levels in PD patients, enhancing mitochondrial and lysosomal function, and reducing inflammatory cytokines in both serum and CSF [197]. This study highlighted that 1000 mg/day for 30 days is safe, increases brain NAD+, and mildly improves clinical symptoms and cerebral metabolism in early-stage PD.
- C.
- Increasing ATP levels: Impaired glycolytic flux in PD triggers a reduction in ATP levels. Terazosin enhanced ATP via phosphoglycerate kinase 1 (PGK1) which stimulates glycolysis and increases brain ATP production, slowing neurodegeneration in preclinical and clinical studies [198].
7.2.3. Reducing Oxidative Stress
- A.
- Increasing DJ-1 levels: Among the endogenous cytoprotective pathways, the DJ-1 protein (encoded by the PARK7 gene) has emerged as a crucial regulator of cellular redox homeostasis. DJ-1 mutations are linked to autosomal recessive-early onset PD and have been shown to compromise mitochondrial function and antioxidant defense mechanisms. Under mitochondrial stress, DJ-1 translocates into mitochondria, where it assists protein trafficking and regulates mitochondrial metabolism. Importantly, DJ-1 also acts as a sensor and scavenger of oxidative stress, participating in the detoxification of ROS and maintaining neuronal viability [199]. Loss of DJ-1 function increases susceptibility to neurotoxins, impairs motor coordination, and shortens lifespan in animal models, emphasizing its critical role in protecting neurons from oxidative damage [200]. Phenylbutyrate is a histone deacetylase inhibitor that increases DJ-1 expression ~2–3-fold in neurons and mouse brains. In cell and mouse models of PD (including MPTP-toxicity and α-synuclein aggregation models), phenylbutyrate reduces oxidative stress, decreases α-synuclein aggregates, and preserves motor/cognitive function [201].
- B.
- Nrf2 Pathway Activation: Nrf2 is an important regulator of antioxidant expression, such as HO-1, NQO1, SOD. A novel multitarget compound that exhibits NRF2 induction activity and MAO-B selective inhibition, combined with anti-inflammatory, antioxidant, and blood–brain barrier permeation properties, has been recently developed and has benefits in PD [202]. Moreover, sulforaphane is an interesting component derived from glucoraphanin and found in many vegetables such as broccoli and cauliflower that activates the Nrf2 pathway. Oral consumption of broccoli leads to a rapid increase in plasma sulforaphane levels within hours. Once absorbed, sulforaphane efficiently crosses the blood–brain barrier, providing neuroprotective benefits against both acute brain injury and chronic neurodegenerative diseases, such as PD [203].
7.3. Huntington’s Disease (HD)
- A.
- Enhancing mitochondrial biogenesis by increasing PGC-1α: Research evidence indicates that mutant huntingtin (mHTT) directly interacts with the promoter region of PGC-1α and interferes with the CREB/TAF-dependent transcriptional pathway, leading to reduced transcriptional activity. This downregulation impairs the expression of key downstream targets, including mitochondrial transcription factor A (TFAM), thereby decreasing biogenesis and contributing to mitochondrial dysfunction, increased oxidative stress susceptibility, and progressive neuronal degeneration [212]. Additionally, mHTT-mediated suppression of PGC-1α affects genes critical for the maintenance of neuromuscular junction stability [213]. Oral administration of bezafibrate restored levels of PGC-1α and downstream targets such as NRF-1 and TFAM, showing improvements in an HD mouse model by increasing mitochondrial biogenesis and reducing lipid accumulation [209,214].
- B.
- Enhancing mitophagy through PINK activation: Research studies have demonstrated that mHTT is also involved in the process of mitophagy. mHTT inhibits the commitment of mitophagy receptors, such as p62 and OPTN, to damaged mitochondria and impairs their interaction with LC3, thereby blocking autophagosome formation and disrupting mitophagy initiation [207]. mHTT also impairs Parkin recruitment to mitochondria and inhibits PINK1 accumulation on the outer mitochondrial membrane [207,215]. This results in defects in mitophagy, where a large number of dysfunctional mitochondria are not removed in time and excessively accumulate in neuronal cells, inducing a vicious cycle [216]. It was observed that the overexpression of PINK1 in a Drosophila model of HD resulted in the amelioration of Parkin-mediated mitophagy defects and the attenuation of mHTT-induced neurotoxicity, promoting neuronal activity and protecting neuronal integrity [215]. Consequently, mitophagy serves to protect neurons from damage in HD, and a deficiency in this process contributes to further malignant deterioration of HD pathology [157].
- C.
- Decreasing mitochondria fragmentation through Drp1 inhibition: The accumulation of mHTT in neuronal cells in the brains of HD patients has been shown to induce mitochondrial hyper-fragmentation and to affect the mitochondrial respiratory chain complex by disrupting the balance of mitochondrial dynamics, which, in turn, induces impairment of mitochondrial function [217]. Moreover, in the brains of HD patients, the expression levels of the mitochondrial fission/fusion proteins Drp1, Fis1, and Mfn are significantly altered [218]. Sawant et al. demonstrated that mHTT is able to bind to the proteins Mfn and Drp1. This binding enhances the activity of Drp1 and, together with a downregulation of fusion proteins such as Mfn1/2 and OPA1, results in mitochondrial fusion and fission imbalances. Fragmented mitochondria are less efficient at ATP production and produce more ROS. Moreover, this fragmentation triggers mitochondrial distribution abnormalities and mitochondrial axonal transport defects, which ultimately lead to impaired synaptic plasticity [219]. Coherently, inhibition of Drp1 has been demonstrated to restore mitochondrial and neuronal dysfunction in mHTT-induced HD animal models. In particular, treatment with CHIR99021 stabilizes calpastatin (CAST), thereby inhibiting calpain activation and preventing Drp1-mediated mitochondrial fragmentation, reducing HD-associated neuropathology and behavioral defects [157,220].
7.4. Amyotrophic Lateral Sclerosis (ALS)
- A.
- Promoting mitophagy: In ALS, ninety distinct mutations in TBK1 are linked to ALS and frontotemporal dementia, including missense mutations that disrupt TBK1’s ability to dimerize and associate with the mitophagy receptor optineurin (OPTN), thereby impairing autophagy/mitophagy initiation and mitochondrial clearance. Additionally, mutations in C9ORF72 disrupt endosomal and autophagic trafficking, leading to defective clearance of damaged mitochondria and accumulation of dysfunctional mitochondria [224]. Moreover, TDP-43 and FUS aggregates sequester essential mitochondrial proteins and impair mitochondrial transport along axons, further compounding mitophagy defects [225,226]. While urolithin A has shown promise as treatment for improving motor dysfunction by activating mitophagy [227], rapamycin, which also increases mitophagy by inhibiting mTOR, only showed positive effects in decreasing neuroinflammation in clinical studies, even though preclinical studies have shown reduced TDP-43 aggregation and improved motor neuron function in ALS [228]. The same has been observed with trehalose, another autophagy activator, for which preclinical studies showed promising results, but in large-scale trials did not demonstrate efficacy in patients [229]. However, trehalose ameliorates the pathogenesis of some subgroups, suggesting that more personalized research is needed.
- B.
- Reducing fragmentation though downregulation of Drp-1: Mutant SOD1, TDP-43, and FUS mislocalize to mitochondria and promote excessive fission via Drp1 upregulation and OPA1/Mfn2 downregulation [230,231]. On the other hand, CHCHD10 mutations affect cristae structure and fusion [232]. Excessive Drp1-mediated mitochondrial fission contributes to mitochondrial fragmentation, which impairs ATP synthesis and facilitates cytochrome c release, triggering apoptotic pathways and motor neuron death. Suppression of the Drp-1 cascade prevents ALS-related symptoms [233].
- C.
- Reducing oxidative stress and ROS production: Mitochondria in ALS show increased ROS production, partly due to SOD1 mutations, which impair detoxification of superoxide radicals, and TDP-43, which inhibits mitochondrial antioxidant responses. Increased mitochondrial ROS also trigger activation of the mitochondrial permeability transition pore (mPTP), promoting mitochondrial swelling and release of pro-apoptotic factors [234,235]. Targeting ROS with antioxidants such as edaravone, which is FDA-approved for ALS [236], helps scavenge ROS, thereby reducing oxidative stress and protecting mtDNA and mitochondrial function.
- D.
- PGC-1α activation: ALS models show reduced PGC-1α levels and downstream regulators such as NRF1 and TFAM, limiting mitochondrial renewal and repair [213], which further exacerbates energy failure and oxidative stress. PGC-1α also regulates the expression of key antioxidant enzymes and mitochondrial fusion/fission proteins; thus, its reduction amplifies mitochondrial vulnerability in ALS [237]. Activation of PGC-1α by resveratrol ameliorates mitochondrial function in ALS animal models [238].
8. Challenges and Future Directions
9. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Liu, X.; Ye, M.; Ma, L. The Emerging Role of Autophagy and Mitophagy in Tauopathies: From Pathogenesis to Translational Implications in Alzheimer’s Disease. Front. Aging Neurosci. 2022, 14, 1022821. [Google Scholar] [CrossRef]
- Nabi, S.U.; Khan, A.; Siddiqui, E.M.; Rehman, M.U.; Alshahrani, S.; Arafah, A.; Mehan, S.; Alsaffar, R.M.; Alexiou, A.; Shen, B. Mechanisms of Mitochondrial Malfunction in Alzheimer’s Disease: New Therapeutic Hope. Oxidative Med. Cell. Longev. 2022, 2022, 4759963. [Google Scholar] [CrossRef]
- Lemasters, J.J. Selective Mitochondrial Autophagy, or Mitophagy, as a Targeted Defense Against Oxidative Stress, Mitochondrial Dysfunction, and Aging. Rejuvenation Res. 2005, 8, 3–5. [Google Scholar] [CrossRef] [PubMed]
- Palikaras, K.; Lionaki, E.; Tavernarakis, N. Mechanisms of Mitophagy in Cellular Homeostasis, Physiology and Pathology. Nat. Cell Biol. 2018, 20, 1013–1022. [Google Scholar] [CrossRef]
- Killackey, S.A.; Philpott, D.J.; Girardin, S.E. Mitophagy Pathways in Health and Disease. J. Cell Biol. 2020, 219, e202004029. [Google Scholar] [CrossRef]
- Siwach, A.; Patel, H.; Khairnar, A.; Parekh, P. Molecular Symphony of Mitophagy: Ubiquitin-Specific Protease-30 as a Maestro for Precision Management of Neurodegenerative Diseases. CNS Neurosci. Ther. 2025, 31, e70192. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, M.; Hou, X.O.; Hu, L.F. Roles of Microglial Mitophagy in Neurological Disorders. Front. Aging Neurosci. 2022, 14, 979869. [Google Scholar] [CrossRef] [PubMed]
- Sarraf, S.A.; Raman, M.; Guarani-Pereira, V.; Sowa, M.E.; Huttlin, E.L.; Gygi, S.P.; Harper, J.W. Landscape of the PARKIN-Dependent Ubiquitylome in Response to Mitochondrial Depolarization. Nature 2013, 496, 372–376. [Google Scholar] [CrossRef]
- Geisler, S.; Holmström, K.M.; Skujat, D.; Fiesel, F.C.; Rothfuss, O.C.; Kahle, P.J.; Springer, W. PINK1/Parkin-Mediated Mitophagy Is Dependent on VDAC1 and P62/SQSTM1. Nat. Cell Biol. 2010, 12, 119–131. [Google Scholar] [CrossRef]
- Heo, J.M.; Ordureau, A.; Paulo, J.A.; Rinehart, J.; Harper, J.W. The PINK1-PARKIN Mitochondrial Ubiquitylation Pathway Drives a Program of OPTN/NDP52 Recruitment and TBK1 Activation to Promote Mitophagy. Mol. Cell 2015, 60, 7–20. [Google Scholar] [CrossRef]
- Caponio, D.; Veverová, K.; Zhang, S.Q.; Shi, L.; Wong, G.; Vyhnalek, M.; Fang, E.F. Compromised Autophagy and Mitophagy in Brain Ageing and Alzheimer’s Diseases. Aging Brain 2022, 2, 100056. [Google Scholar] [CrossRef]
- Swerdlow, N.S.; Wilkins, H.M. Mitophagy and the Brain. Int. J. Mol. Sci. 2020, 21, 9661. [Google Scholar] [CrossRef]
- Castellazzi, M.; Patergnani, S.; Donadio, M.; Giorgi, C.; Bonora, M.; Bosi, C.; Brombo, G.; Pugliatti, M.; Seripa, D.; Zuliani, G.; et al. Autophagy and Mitophagy Biomarkers Are Reduced in Sera of Patients with Alzheimer’s Disease and Mild Cognitive Impairment. Sci. Rep. 2019, 9, 20009. [Google Scholar] [CrossRef] [PubMed]
- McWilliams, T.G.; Prescott, A.R.; Montava-Garriga, L.; Ball, G.; Singh, F.; Barini, E.; Muqit, M.M.K.; Brooks, S.P.; Ganley, I.G. Basal Mitophagy Occurs Independently of PINK1 in Mouse Tissues of High Metabolic Demand. Cell Metab. 2018, 27, 439–449.e5. [Google Scholar] [CrossRef]
- Perez, F.A.; Palmiter, R.D. Parkin-Deficient Mice Are Not a Robust Model of Parkinsonism. Proc. Natl. Acad. Sci. USA 2005, 102, 2174–2179. [Google Scholar] [CrossRef]
- Han, R.; Wang, Q.; Xiong, X.; Chen, X.; Tu, Z.; Li, B.; Zhang, F.; Chen, C.; Pan, M.; Xu, T.; et al. Deficiency of Parkin Causes Neurodegeneration and Accumulation of Pathological α-Synuclein in Monkey Models. J. Clin. Investig. 2024, 134, e179633. [Google Scholar] [CrossRef] [PubMed]
- Qian, S.; He, H.; Xiong, X.; Ai, R.; Wang, W.; Zhu, H.; Ye, Q.; Zhou, S.; Nilsen, H.; Xie, C. Identification of Mitophagy-Associated Proteins Profile as Potential Plasma Biomarkers of Idiopathic Parkinson’s Disease. CNS Neurosci. Ther. 2024, 30, e14532. [Google Scholar] [CrossRef]
- Lee, J.J.; Sanchez-Martinez, A.; Martinez Zarate, A.; Benincá, C.; Mayor, U.; Clague, M.J.; Whitworth, A.J. Basal Mitophagy Is Widespread in Drosophila but Minimally Affected by Loss of Pink1 or Parkin. J. Cell Biol. 2018, 217, 1613–1622. [Google Scholar] [CrossRef] [PubMed]
- Wrighton, P.J.; Shwartz, A.; Heo, J.-M.; Quenzer, E.D.; LaBella, K.A.; Harper, J.W.; Goessling, W. Quantitative Intravital Imaging in Zebrafish Reveals in Vivo Dynamics of Physiological-Stress-Induced Mitophagy. J. Cell Sci. 2021, 134, jcs256255. [Google Scholar] [CrossRef]
- Rappe, A.; Vihinen, H.A.; Suomi, F.; Hassinen, A.J.; Ehsan, H.; Jokitalo, E.S.; McWilliams, T.G. Longitudinal Autophagy Profiling of the Mammalian Brain Reveals Sustained Mitophagy throughout Healthy Aging. EMBO J. 2024, 43, 6199–6231. [Google Scholar] [CrossRef]
- Schmid, E.T.; Pyo, J.H.; Walker, D.W. Neuronal Induction of BNIP3-Mediated Mitophagy Slows Systemic Aging in Drosophila. Nat. Aging 2022, 2, 494–507. [Google Scholar] [CrossRef]
- Baker, D.J.; Petersen, R.C. Cellular Senescence in Brain Aging and Neurodegenerative Diseases: Evidence and Perspectives. J. Clin. Investig. 2018, 128, 1208–1216. [Google Scholar] [CrossRef]
- Di Micco, R.; Krizhanovsky, V.; Baker, D.; d’Adda di Fagagna, F. Cellular Senescence in Ageing: From Mechanisms to Therapeutic Opportunities. Nat. Rev. Mol. Cell Biol. 2021, 22, 75–95. [Google Scholar] [CrossRef]
- Fang, E.F.; Hou, Y.; Palikaras, K.; Adriaanse, B.A.; Kerr, J.S.; Yang, B.; Lautrup, S.; Hasan-Olive, M.M.; Caponio, D.; Dan, X.; et al. Mitophagy Inhibits Amyloid-β and Tau Pathology and Reverses Cognitive Deficits in Models of Alzheimer’s Disease. Nat. Neurosci. 2019, 22, 401–412. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Zhuang, X.-X.; Niu, Z.; Ai, R.; Lautrup, S.; Zheng, S.; Jiang, Y.; Han, R.; Gupta, T.S.; Cao, S.; et al. Amelioration of Alzheimer’s Disease Pathology by Mitophagy Inducers Identified via Machine Learning and a Cross-Species Workflow. Nat. Biomed. Eng. 2022, 6, 76–93. [Google Scholar] [CrossRef] [PubMed]
- Sadovnikova, I.S.; Gureev, A.P.; Ignatyeva, D.A.; Gryaznova, M.V.; Chernyshova, E.V.; Krutskikh, E.P.; Novikova, A.G.; Popov, V.N. Nrf2/ARE Activators Improve Memory in Aged Mice via Maintaining of Mitochondrial Quality Control of Brain and the Modulation of Gut Microbiome. Pharmaceuticals 2021, 14, 607. [Google Scholar] [CrossRef]
- Singh, P.; Gollapalli, K.; Mangiola, S.; Schranner, D.; Yusuf, M.A.; Chamoli, M.; Shi, S.L.; Lopes Bastos, B.; Nair, T.; Riermeier, A.; et al. Taurine Deficiency as a Driver of Aging. Science 2023, 380, eabn9257. [Google Scholar] [CrossRef] [PubMed]
- Gruendler, R.; Hippe, B.; Sendula Jengic, V.; Peterlin, B.; Haslberger, A.G. Nutraceutical Approaches of Autophagy and Neuroinflammation in Alzheimer’s Disease: A Systematic Review. Molecules 2020, 25, 6018. [Google Scholar] [CrossRef]
- Makarov, M.; Korkotian, E. Differential Role of Active Compounds in Mitophagy and Related Neurodegenerative Diseases. Toxins 2023, 15, 202. [Google Scholar] [CrossRef]
- Park, H.; Chung, K.M.; An, H.K.; Gim, J.-E.; Hong, J.; Woo, H.; Cho, B.; Moon, C.; Yu, S.W. Parkin Promotes Mitophagic Cell Death in Adult Hippocampal Neural Stem Cells Following Insulin Withdrawal. Front. Mol. Neurosci. 2019, 12, 46. [Google Scholar] [CrossRef]
- Zaninello, M.; Palikaras, K.; Naon, D.; Iwata, K.; Herkenne, S.; Quintana-Cabrera, R.; Semenzato, M.; Grespi, F.; Ross-Cisneros, F.N.; Carelli, V.; et al. Inhibition of Autophagy Curtails Visual Loss in a Model of Autosomal Dominant Optic Atrophy. Nat. Commun. 2020, 11, 4029. [Google Scholar] [CrossRef]
- Zaninello, M.; Palikaras, K.; Sotiriou, A.; Tavernarakis, N.; Scorrano, L. Sustained Intracellular Calcium Rise Mediates Neuronal Mitophagy in Models of Autosomal Dominant Optic Atrophy. Cell Death Differ. 2022, 29, 167–177. [Google Scholar] [CrossRef]
- Patergnani, S.; Bonora, M.; Ingusci, S.; Previati, M.; Marchi, S.; Zucchini, S.; Perrone, M.; Wieckowski, M.R.; Castellazzi, M.; Pugliatti, M.; et al. Antipsychotic Drugs Counteract Autophagy and Mitophagy in Multiple Sclerosis. Proc. Natl. Acad. Sci. USA 2021, 118, e2020078118. [Google Scholar] [CrossRef]
- Sun, N.; Yun, J.; Liu, J.; Malide, D.; Liu, C.; Rovira, I.I.; Holmström, K.M.; Fergusson, M.M.; Yoo, Y.H.; Combs, C.A.; et al. Measuring In Vivo Mitophagy. Mol. Cell 2015, 60, 685–696. [Google Scholar] [CrossRef]
- Blüml, S.; Moreno-Torres, A.; Shic, F.; Nguy, C.H.; Ross, B.D. Tricarboxylic Acid Cycle of Glia in the in Vivo Human Brain. NMR Biomed. 2002, 15, 1–5. [Google Scholar] [CrossRef]
- Mulica, P.; Grünewald, A.; Pereira, S.L. Astrocyte-Neuron Metabolic Crosstalk in Neurodegeneration: A Mitochondrial Perspective. Front. Endocrinol. 2021, 12, 668517. [Google Scholar] [CrossRef] [PubMed]
- Boisvert, M.M.; Erikson, G.A.; Shokhirev, M.N.; Allen, N.J. The Aging Astrocyte Transcriptome from Multiple Regions of the Mouse Brain. Cell Rep. 2018, 22, 269–285. [Google Scholar] [CrossRef] [PubMed]
- Matias, I.; Diniz, L.P.; Damico, I.V.; Araujo, A.P.B.; Neves, L.d.S.; Vargas, G.; Leite, R.E.P.; Suemoto, C.K.; Nitrini, R.; Jacob-Filho, W.; et al. Loss of Lamin-B1 and Defective Nuclear Morphology Are Hallmarks of Astrocyte Senescence in Vitro and in the Aging Human Hippocampus. Aging Cell 2022, 21, e13521. [Google Scholar] [CrossRef] [PubMed]
- Chinta, S.J.; Woods, G.; Demaria, M.; Rane, A.; Zou, Y.; McQuade, A.; Rajagopalan, S.; Limbad, C.; Madden, D.T.; Campisi, J.; et al. Cellular Senescence Is Induced by the Environmental Neurotoxin Paraquat and Contributes to Neuropathology Linked to Parkinson’s Disease. Cell Rep. 2018, 22, 930–940. [Google Scholar] [CrossRef]
- Scott, K.M.; Williams-Gray, C.H. Targeting Aged Astrocytes May Be a New Therapeutic Strategy in Parkinson’s Disease. Mov. Disord. 2018, 33, 758–759. [Google Scholar] [CrossRef]
- Diniz, L.P.; Araujo, A.P.B.; Carvalho, C.F.; Matias, I.; de Sá Hayashide, L.; Marques, M.; Pessoa, B.; Andrade, C.B.V.; Vargas, G.; Queiroz, D.D.; et al. Accumulation of Damaged Mitochondria in Aging Astrocytes Due to Mitophagy Dysfunction: Implications for Susceptibility to Mitochondrial Stress. Biochim. Et Biophys. Acta (BBA) Mol. Basis Dis. 2024, 1870, 167470. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhu, Z.; Yan, Y.; Sun, H.; Wang, G.; Du, X.; Li, F.; Yuan, S.; Wang, W.; Wang, M.; et al. P7C3 Suppresses Astrocytic Senescence to Protect Dopaminergic Neurons: Implication in the Mouse Model of Parkinson’s Disease. CNS Neurosci. Ther. 2024, 30, e14819. [Google Scholar] [CrossRef]
- Cohen, J.; Torres, C. Astrocyte Senescence: Evidence and Significance. Aging Cell 2019, 18, e12937. [Google Scholar] [CrossRef]
- Traa, A.; Tamez González, A.A.; Van Raamsdonk, J.M. Developmental Disruption of the Mitochondrial Fission Gene Drp-1 Extends the Longevity of Daf-2 Insulin/IGF-1 Receptor Mutant. Geroscience 2025, 47, 877–902. [Google Scholar] [CrossRef]
- Friedman, J.R.; Nunnari, J. Mitochondrial Form and Function. Nature 2014, 505, 335–343. [Google Scholar] [CrossRef]
- Sukhorukov, V.S.; Baranich, T.I.; Egorova, A.V.; Akateva, A.V.; Okulova, K.M.; Ryabova, M.S.; Skvortsova, K.A.; Dmitriev, O.V.; Mudzhiri, N.M.; Voronkov, D.N.; et al. Mitochondrial Dynamics in Brain Cells During Normal and Pathological Aging. Int. J. Mol. Sci. 2024, 25, 12855. [Google Scholar] [CrossRef]
- Bondy, S.C.; Wu, M. The Critical Role of Autophagy and Phagocytosis in the Aging Brain. Int. J. Mol. Sci. 2024, 26, 57. [Google Scholar] [CrossRef]
- Yao, C.H.; Wang, R.; Wang, Y.; Kung, C.P.; Weber, J.D.; Patti, G.J. Mitochondrial Fusion Supports Increased Oxidative Phosphorylation during Cell Proliferation. eLife 2019, 8, e41351. [Google Scholar] [CrossRef] [PubMed]
- Ng, M.Y.W.; Wai, T.; Simonsen, A. Quality Control of the Mitochondrion. Dev. Cell 2021, 56, 881–905. [Google Scholar] [CrossRef] [PubMed]
- Machiela, E.; Liontis, T.; Dues, D.J.; Rudich, P.D.; Traa, A.; Wyman, L.; Kaufman, C.; Cooper, J.F.; Lew, L.; Nadarajan, S.; et al. Disruption of Mitochondrial Dynamics Increases Stress Resistance through Activation of Multiple Stress Response Pathways. FASEB J. 2020, 34, 8475–8492. [Google Scholar] [CrossRef]
- Wilson, P.D.; Franks, L.M. The Effect of Age on Mitochondrial Ultrastructure. Gerontologia 1975, 21, 81–94. [Google Scholar] [CrossRef]
- Yasuda, K.; Ishii, T.; Suda, H.; Akatsuka, A.; Hartman, P.S.; Goto, S.; Miyazawa, M.; Ishii, N. Age-Related Changes of Mitochondrial Structure and Function in Caenorhabditis Elegans. Mech. Ageing Dev. 2006, 127, 763–770. [Google Scholar] [CrossRef]
- Sharma, A.; Smith, H.J.; Yao, P.; Mair, W.B. Causal Roles of Mitochondrial Dynamics in Longevity and Healthy Aging. EMBO Rep. 2019, 20, e48395. [Google Scholar] [CrossRef]
- Knott, A.B.; Perkins, G.; Schwarzenbacher, R.; Bossy-Wetzel, E. Mitochondrial Fragmentation in Neurodegeneration. Nat. Rev. Neurosci. 2008, 9, 505–518. [Google Scholar] [CrossRef]
- Liu, Y.J.; McIntyre, R.L.; Janssens, G.E.; Houtkooper, R.H. Mitochondrial Fission and Fusion: A Dynamic Role in Aging and Potential Target for Age-Related Disease. Mech. Ageing Dev. 2020, 186, 111212. [Google Scholar] [CrossRef] [PubMed]
- Son, J.M.; Sarsour, E.H.; Kakkerla Balaraju, A.; Fussell, J.; Kalen, A.L.; Wagner, B.A.; Buettner, G.R.; Goswami, P.C. Mitofusin 1 and Optic Atrophy 1 Shift Metabolism to Mitochondrial Respiration during Aging. Aging Cell 2017, 16, 1136–1145. [Google Scholar] [CrossRef]
- Sgarbi, G.; Matarrese, P.; Pinti, M.; Lanzarini, C.; Ascione, B.; Gibellini, L.; Dika, E.; Patrizi, A.; Tommasino, C.; Capri, M.; et al. Mitochondria Hyperfusion and Elevated Autophagic Activity Are Key Mechanisms for Cellular Bioenergetic Preservation in Centenarians. Aging (Albany NY) 2014, 6, 296–310. [Google Scholar] [CrossRef]
- Chen, H.; Chomyn, A.; Chan, D.C. Disruption of Fusion Results in Mitochondrial Heterogeneity and Dysfunction*. J. Biol. Chem. 2005, 280, 26185–26192. [Google Scholar] [CrossRef] [PubMed]
- Oh, J.; Lee, Y.D.; Wagers, A.J. Stem Cell Aging: Mechanisms, Regulators and Therapeutic Opportunities. Nat. Med. 2014, 20, 870–880. [Google Scholar] [CrossRef] [PubMed]
- Signer, R.A.J.; Morrison, S.J. Mechanisms That Regulate Stem Cell Aging and Life Span. Cell Stem Cell 2013, 12, 152–165. [Google Scholar] [CrossRef]
- Scheckhuber, C.Q.; Erjavec, N.; Tinazli, A.; Hamann, A.; Nyström, T.; Osiewacz, H.D. Reducing Mitochondrial Fission Results in Increased Life Span and Fitness of Two Fungal Ageing Models. Nat. Cell Biol. 2007, 9, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Braun, R.J.; Westermann, B. Mitochondrial Dynamics in Yeast Cell Death and Aging. Biochem. Soc. Trans. 2011, 39, 1520–1526. [Google Scholar] [CrossRef] [PubMed]
- Traa, A.; Machiela, E.; Rudich, P.D.; Soo, S.K.; Senchuk, M.M.; Van Raamsdonk, J.M. Identification of Novel Therapeutic Targets for Polyglutamine Diseases That Target Mitochondrial Fragmentation. Int. J. Mol. Sci. 2021, 22, 13447. [Google Scholar] [CrossRef]
- Yang, C.C.; Chen, D.; Lee, S.S.; Walter, L. The Dynamin-Related Protein DRP-1 and the Insulin Signaling Pathway Cooperate to Modulate Caenorhabditis Elegans Longevity. Aging Cell 2011, 10, 724–728. [Google Scholar] [CrossRef]
- Weir, H.J.; Yao, P.; Huynh, F.K.; Escoubas, C.C.; Goncalves, R.L.; Burkewitz, K.; Laboy, R.; Hirschey, M.D.; Mair, W.B. Dietary Restriction and AMPK Increase Lifespan via Mitochondrial Network and Peroxisome Remodeling. Cell Metab. 2017, 26, 884–896.e5. [Google Scholar] [CrossRef]
- Espino de la Fuente-Muñoz, C.; Rosas-Lemus, M.; Moreno-Castilla, P.; Bermúdez-Rattoni, F.; Uribe-Carvajal, S.; Arias, C. Age-Dependent Decline in Synaptic Mitochondrial Function Is Exacerbated in Vulnerable Brain Regions of Female 3xTg-AD Mice. Int. J. Mol. Sci. 2020, 21, 8727. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Zhang, Q.; Zhang, P.; Sun, L.; Peng, C.; Yuan, Z.; Cheng, J. C-Abl-Mediated Drp1 Phosphorylation Promotes Oxidative Stress-Induced Mitochondrial Fragmentation and Neuronal Cell Death. Cell Death Dis. 2017, 8, e3117. [Google Scholar] [CrossRef]
- Li, Y.; Yu, H.; Chen, C.; Li, S.; Zhang, Z.; Xu, H.; Zhu, F.; Liu, J.; Spencer, P.S.; Dai, Z.; et al. Proteomic Profile of Mouse Brain Aging Contributions to Mitochondrial Dysfunction, DNA Oxidative Damage, Loss of Neurotrophic Factor, and Synaptic and Ribosomal Proteins. Oxid. Med. Cell Longev. 2020, 2020, 5408452. [Google Scholar] [CrossRef]
- Park, J.; Choi, H.; Kim, B.; Chae, U.; Lee, D.G.; Lee, S.R.; Lee, S.; Lee, H.S.; Lee, D.S. Peroxiredoxin 5 (Prx5) Decreases LPS-Induced Microglial Activation through Regulation of Ca2+/Calcineurin-Drp1-Dependent Mitochondrial Fission. Free Radic. Biol. Med. 2016, 99, 392–404. [Google Scholar] [CrossRef]
- Katoh, M.; Wu, B.; Nguyen, H.B.; Thai, T.Q.; Yamasaki, R.; Lu, H.; Rietsch, A.M.; Zorlu, M.M.; Shinozaki, Y.; Saitoh, Y.; et al. Polymorphic Regulation of Mitochondrial Fission and Fusion Modifies Phenotypes of Microglia in Neuroinflammation. Sci. Rep. 2017, 7, 4942. [Google Scholar] [CrossRef]
- Park, J.; Choi, H.; Min, J.S.; Park, S.J.; Kim, J.H.; Park, H.J.; Kim, B.; Chae, J.I.; Yim, M.; Lee, D.S. Mitochondrial Dynamics Modulate the Expression of Pro-Inflammatory Mediators in Microglial Cells. J. Neurochem. 2013, 127, 221–232. [Google Scholar] [CrossRef]
- Zhou, K.; Chen, J.; Wu, J.; Wu, Q.; Jia, C.; Xu, Y.X.Z.; Chen, L.; Tu, W.; Yang, G.; Kong, J.; et al. Atractylenolide III Ameliorates Cerebral Ischemic Injury and Neuroinflammation Associated with Inhibiting JAK2/STAT3/Drp1-Dependent Mitochondrial Fission in Microglia. Phytomedicine 2019, 59, 152922. [Google Scholar] [CrossRef]
- Traa, A.; Keil, A.; AlOkda, A.; Jacob-Tomas, S.; Tamez González, A.A.; Zhu, S.; Rudich, Z.; Van Raamsdonk, J.M. Overexpression of Mitochondrial Fission or Mitochondrial Fusion Genes Enhances Resilience and Extends Longevity. Aging Cell 2024, 23, e14262. [Google Scholar] [CrossRef]
- Pulido, C.; Ryan, T.A. Synaptic Vesicle Pools Are a Major Hidden Resting Metabolic Burden of Nerve Terminals. Sci. Adv. 2021, 7, eabi9027. [Google Scholar] [CrossRef] [PubMed]
- Du, F.; Zhu, X.H.; Zhang, Y.; Friedman, M.; Zhang, N.; Ugurbil, K.; Chen, W. Tightly Coupled Brain Activity and Cerebral ATP Metabolic Rate. Proc. Natl. Acad. Sci. USA 2008, 105, 6409–6414. [Google Scholar] [CrossRef] [PubMed]
- Pan, R.Y.; He, L.; Zhang, J.; Liu, X.; Liao, Y.; Gao, J.; Liao, Y.; Yan, Y.; Li, Q.; Zhou, X.; et al. Positive Feedback Regulation of Microglial Glucose Metabolism by Histone H4 Lysine 12 Lactylation in Alzheimer’s Disease. Cell Metab. 2022, 34, 634–648.e6. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Cao, K.; Wang, F.; Wu, W.; Mai, W.; Qiu, L.; Luo, Y.; Ge, W.P.; Sun, B.; Shi, L.; et al. Dual Roles of Hexokinase 2 in Shaping Microglial Function by Gating Glycolytic Flux and Mitochondrial Activity. Nat. Metab. 2022, 4, 1756–1774. [Google Scholar] [CrossRef]
- Saab, A.S.; Tzvetavona, I.D.; Trevisiol, A.; Baltan, S.; Dibaj, P.; Kusch, K.; Möbius, W.; Goetze, B.; Jahn, H.M.; Huang, W.; et al. Oligodendroglial NMDA Receptors Regulate Glucose Import and Axonal Energy Metabolism. Neuron 2016, 91, 119–132. [Google Scholar] [CrossRef]
- Pellerin, L.; Magistretti, P.J. Glutamate Uptake into Astrocytes Stimulates Aerobic Glycolysis: A Mechanism Coupling Neuronal Activity to Glucose Utilization. Proc. Natl. Acad. Sci. USA 1994, 91, 10625–10629. [Google Scholar] [CrossRef]
- Kasischke, K.A.; Vishwasrao, H.D.; Fisher, P.J.; Zipfel, W.R.; Webb, W.W. Neural Activity Triggers Neuronal Oxidative Metabolism Followed by Astrocytic Glycolysis. Science 2004, 305, 99–103. [Google Scholar] [CrossRef]
- Grinblat, L.; Pacheco Bolaños, L.F.; Stoppani, A.O. Decreased Rate of Ketone-Body Oxidation and Decreased Activity of D-3-Hydroxybutyrate Dehydrogenase and Succinyl-CoA:3-Oxo-Acid CoA-Transferase in Heart Mitochondria of Diabetic Rats. Biochem. J. 1986, 240, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Davis, M.L.; Sheng, Z.H. Regulation of Axonal Mitochondrial Transport and Its Impact on Synaptic Transmission. Neurosci. Res. 2011, 70, 9–15. [Google Scholar] [CrossRef]
- Bartman, S.; Coppotelli, G.; Ross, J.M. Mitochondrial Dysfunction: A Key Player in Brain Aging and Diseases. Curr. Issues Mol. Biol. 2024, 46, 1987–2026. [Google Scholar] [CrossRef]
- Sutherland, T.C.; Sefiani, A.; Horvat, D.; Huntington, T.E.; Lei, Y.; West, A.P.; Geoffroy, C.G. Age-Dependent Decline in Neuron Growth Potential and Mitochondria Functions in Cortical Neurons. Cells 2021, 10, 1625. [Google Scholar] [CrossRef] [PubMed]
- Bowling, A.C.; Mutisya, E.M.; Walker, L.C.; Price, D.L.; Cork, L.C.; Beal, M.F. Age-Dependent Impairment of Mitochondrial Function in Primate Brain. J. Neurochem. 1993, 60, 1964–1967. [Google Scholar] [CrossRef]
- Sandhu, S.K.; Kaur, G. Mitochondrial Electron Transport Chain Complexes in Aging Rat Brain and Lymphocytes. Biogerontology 2003, 4, 19–29. [Google Scholar] [CrossRef]
- Davie, K.; Janssens, J.; Koldere, D.; De Waegeneer, M.; Pech, U.; Kreft, Ł.; Aibar, S.; Makhzami, S.; Christiaens, V.; Bravo González-Blas, C.; et al. A Single-Cell Transcriptome Atlas of the Aging Drosophila Brain. Cell 2018, 174, 982–998.e20. [Google Scholar] [CrossRef]
- Zhu, X.H.; Lu, M.; Lee, B.Y.; Ugurbil, K.; Chen, W. In Vivo NAD Assay Reveals the Intracellular NAD Contents and Redox State in Healthy Human Brain and Their Age Dependences. Proc. Natl. Acad. Sci. USA 2015, 112, 2876–2881. [Google Scholar] [CrossRef]
- Yoshino, J.; Baur, J.A.; Imai, S.I. NAD+ Intermediates: The Biology and Therapeutic Potential of NMN and NR. Cell Metab. 2018, 27, 513–528. [Google Scholar] [CrossRef]
- Shichkova, P.; Coggan, J.S.; Kanari, L.; Boci, E.; Favreau, C.; Antonel, S.M.; Keller, D.; Markram, H. Breakdown and Repair of Metabolism in the Aging Brain. Front. Sci. 2025, 3, 1441297. [Google Scholar] [CrossRef]
- Zhao, X.; Yang, X.; Du, C.; Hao, H.; Liu, S.; Liu, G.; Zhang, G.; Fan, K.; Ma, J. Up-Regulated Succinylation Modifications Induce a Senescence Phenotype in Microglia by Altering Mitochondrial Energy Metabolism. J. Neuroinflammation 2024, 21, 296. [Google Scholar] [CrossRef]
- Na, D.; Zhang, Z.; Meng, M.; Li, M.; Gao, J.; Kong, J.; Zhang, G.; Guo, Y. Energy Metabolism and Brain Aging: Strategies to Delay Neuronal Degeneration. Cell Mol. Neurobiol. 2025, 45, 38. [Google Scholar] [CrossRef]
- Fakouri, N.B.; Hansen, T.L.; Desler, C.; Anugula, S.; Rasmussen, L.J. From Powerhouse to Perpetrator-Mitochondria in Health and Disease. Biology 2019, 8, 35. [Google Scholar] [CrossRef]
- Ionescu-Tucker, A.; Cotman, C.W. Emerging Roles of Oxidative Stress in Brain Aging and Alzheimer’s Disease. Neurobiol. Aging 2021, 107, 86–95. [Google Scholar] [CrossRef]
- Gómez, J.; Mota-Martorell, N.; Jové, M.; Pamplona, R.; Barja, G. Mitochondrial ROS Production, Oxidative Stress and Aging within and between Species: Evidences and Recent Advances on This Aging Effector. Exp. Gerontol. 2023, 174, 112134. [Google Scholar] [CrossRef] [PubMed]
- Maynard, S.; Fang, E.F.; Scheibye-Knudsen, M.; Croteau, D.L.; Bohr, V.A. DNA Damage, DNA Repair, Aging, and Neurodegeneration. Cold Spring Harb. Perspect. Med. 2015, 5, a025130. [Google Scholar] [CrossRef] [PubMed]
- Chung, K.K.K.; Thomas, B.; Li, X.; Pletnikova, O.; Troncoso, J.C.; Marsh, L.; Dawson, V.L.; Dawson, T.M. S-Nitrosylation of Parkin Regulates Ubiquitination and Compromises Parkin’s Protective Function. Science 2004, 304, 1328–1331. [Google Scholar] [CrossRef] [PubMed]
- Xiao, B.; Kuruvilla, J.; Tan, E.K. Mitophagy and Reactive Oxygen Species Interplay in Parkinson’s Disease. npj Parkinsons Dis. 2022, 8, 135. [Google Scholar] [CrossRef]
- Xiao, Q.; Yang, Y.; Zhao, X.Y.; He, L.S.; Qin, Y.; He, Y.H.; Zhang, G.P.; Luo, J.D. Oxidative Stress Contributes to the Impaired Sonic Hedgehog Pathway in Type 1 Diabetic Mice with Myocardial Infarction. Exp. Ther. Med. 2015, 10, 1750–1758. [Google Scholar] [CrossRef]
- Yang, C.; Qi, Y.; Sun, Z. The Role of Sonic Hedgehog Pathway in the Development of the Central Nervous System and Aging-Related Neurodegenerative Diseases. Front. Mol. Biosci. 2021, 8, 711710. [Google Scholar] [CrossRef]
- Snezhkina, A.V.; Kudryavtseva, A.V.; Kardymon, O.L.; Savvateeva, M.V.; Melnikova, N.V.; Krasnov, G.S.; Dmitriev, A.A. ROS Generation and Antioxidant Defense Systems in Normal and Malignant Cells. Oxid. Med. Cell Longev. 2019, 2019, 6175804. [Google Scholar] [CrossRef] [PubMed]
- Venkateshappa, C.; Harish, G.; Mythri, R.B.; Mahadevan, A.; Bharath, M.M.S.; Shankar, S.K. Increased Oxidative Damage and Decreased Antioxidant Function in Aging Human Substantia Nigra Compared to Striatum: Implications for Parkinson’s Disease. Neurochem. Res. 2012, 37, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Lana, J.V.; Rios, A.; Takeyama, R.; Santos, N.; Pires, L.; Santos, G.S.; Rodrigues, I.J.; Jeyaraman, M.; Purita, J.; Lana, J.F. Nebulized Glutathione as a Key Antioxidant for the Treatment of Oxidative Stress in Neurodegenerative Conditions. Nutrients 2024, 16, 2476. [Google Scholar] [CrossRef]
- Luchsinger, J.A.; Tang, M.X.; Shea, S.; Mayeux, R. Caloric Intake and the Risk of Alzheimer Disease. Arch. Neurol. 2002, 59, 1258–1263. [Google Scholar] [CrossRef] [PubMed]
- Spina Purrello, V.; Cormaci, G.; Denaro, L.; Reale, S.; Costa, A.; Lalicata, C.; Sabbatini, M.; Marchetti, B.; Avola, R. Effect of Growth Factors on Nuclear and Mitochondrial ADP-Ribosylation Processes during Astroglial Cell Development and Aging in Culture. Mech. Ageing Dev. 2002, 123, 511–520. [Google Scholar] [CrossRef]
- Muftuoglu, M.; Mori, M.P.; de Souza-Pinto, N.C. Formation and Repair of Oxidative Damage in the Mitochondrial DNA. Mitochondrion 2014, 17, 164–181. [Google Scholar] [CrossRef]
- Lawless, C.; Greaves, L.; Reeve, A.K.; Turnbull, D.M.; Vincent, A.E. The Rise and Rise of Mitochondrial DNA Mutations. Open Biol. 2020, 10, 200061. [Google Scholar] [CrossRef]
- Yakes, F.M.; Van Houten, B. Mitochondrial DNA Damage Is More Extensive and Persists Longer than Nuclear DNA Damage in Human Cells Following Oxidative Stress. Proc. Natl. Acad. Sci. USA 1997, 94, 514–519. [Google Scholar] [CrossRef]
- Rong, Z.; Tu, P.; Xu, P.; Sun, Y.; Yu, F.; Tu, N.; Guo, L.; Yang, Y. The Mitochondrial Response to DNA Damage. Front. Cell Dev. Biol. 2021, 9, 669379. [Google Scholar] [CrossRef]
- Wallace, D.C. A Mitochondrial Paradigm of Metabolic and Degenerative Diseases, Aging, and Cancer: A Dawn for Evolutionary Medicine. Annu. Rev. Genet. 2005, 39, 359–407. [Google Scholar] [CrossRef]
- Oldfors, A.; Moslemi, A.R.; Fyhr, I.M.; Holme, E.; Larsson, N.G.; Lindberg, C. Mitochondrial DNA Deletions in Muscle Fibers in Inclusion Body Myositis. J. Neuropathol. Exp. Neurol. 1995, 54, 581–587. [Google Scholar] [CrossRef] [PubMed]
- Nido, G.S.; Dölle, C.; Flønes, I.; Tuppen, H.A.; Alves, G.; Tysnes, O.B.; Haugarvoll, K.; Tzoulis, C. Ultradeep Mapping of Neuronal Mitochondrial Deletions in Parkinson’s Disease. Neurobiol. Aging 2018, 63, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Bender, A.; Krishnan, K.J.; Morris, C.M.; Taylor, G.A.; Reeve, A.K.; Perry, R.H.; Jaros, E.; Hersheson, J.S.; Betts, J.; Klopstock, T.; et al. High Levels of Mitochondrial DNA Deletions in Substantia Nigra Neurons in Aging and Parkinson Disease. Nat. Genet. 2006, 38, 515–517. [Google Scholar] [CrossRef]
- Bender, A.; Schwarzkopf, R.M.; McMillan, A.; Krishnan, K.J.; Rieder, G.; Neumann, M.; Elstner, M.; Turnbull, D.M.; Klopstock, T. Dopaminergic Midbrain Neurons Are the Prime Target for Mitochondrial DNA Deletions. J. Neurol. 2008, 255, 1231–1235. [Google Scholar] [CrossRef]
- Wei Soong, N.; Hinton, D.R.; Cortopassi, G.; Arnheim, N. Mosaicism for a Specific Somatic Mitochondrial DNA Mutation in Adult Human Brain. Nat. Genet. 1992, 2, 318–323. [Google Scholar] [CrossRef]
- Krokan, H.E.; Bjørås, M. Base Excision Repair. Cold Spring Harb. Perspect. Biol. 2013, 5, a012583. [Google Scholar] [CrossRef]
- Kazak, L.; Reyes, A.; Holt, I.J. Minimizing the Damage: Repair Pathways Keep Mitochondrial DNA Intact. Nat. Rev. Mol. Cell Biol. 2012, 13, 659–671. [Google Scholar] [CrossRef] [PubMed]
- Youle, R.J.; van der Bliek, A.M. Mitochondrial Fission, Fusion, and Stress. Science 2012, 337, 1062–1065. [Google Scholar] [CrossRef]
- Yamazaki, T.; Kirchmair, A.; Sato, A.; Buqué, A.; Rybstein, M.; Petroni, G.; Bloy, N.; Finotello, F.; Stafford, L.; Navarro Manzano, E.; et al. Mitochondrial DNA Drives Abscopal Responses to Radiation That Are Inhibited by Autophagy. Nat. Immunol. 2020, 21, 1160–1171. [Google Scholar] [CrossRef]
- Elson, J.L.; Samuels, D.C.; Turnbull, D.M.; Chinnery, P.F. Random Intracellular Drift Explains the Clonal Expansion of Mitochondrial DNA Mutations with Age. Am. J. Hum. Genet. 2001, 68, 802–806. [Google Scholar] [CrossRef]
- Kraytsberg, Y.; Kudryavtseva, E.; McKee, A.C.; Geula, C.; Kowall, N.W.; Khrapko, K. Mitochondrial DNA Deletions Are Abundant and Cause Functional Impairment in Aged Human Substantia Nigra Neurons. Nat. Genet. 2006, 38, 518–520. [Google Scholar] [CrossRef] [PubMed]
- Itoh, K.; Weis, S.; Mehraein, P.; Müller-Höcker, J. Defects of Cytochrome c Oxidase in the Substantia Nigra of Parkinson’s Disease: An Immunohistochemical and Morphometric Study. Mov. Disord. 1997, 12, 9–16. [Google Scholar] [CrossRef]
- Diaz, F.; Bayona-Bafaluy, M.P.; Rana, M.; Mora, M.; Hao, H.; Moraes, C.T. Human Mitochondrial DNA with Large Deletions Repopulates Organelles Faster than Full-length Genomes under Relaxed Copy Number Control. Nucleic Acids Res. 2002, 30, 4626–4633. [Google Scholar] [CrossRef]
- Moraes, C.T.; Kenyon, L.; Hao, H. Mechanisms of Human Mitochondrial DNA Maintenance: The Determining Role of Primary Sequence and Length over Function. MBoC 1999, 10, 3345–3356. [Google Scholar] [CrossRef]
- Kowald, A.; Kirkwood, T.B.L. Resolving the Enigma of the Clonal Expansion of mtDNA Deletions. Genes. 2018, 9, 126. [Google Scholar] [CrossRef]
- Vincent, A.E.; Rosa, H.S.; Pabis, K.; Lawless, C.; Chen, C.; Grünewald, A.; Rygiel, K.A.; Rocha, M.C.; Reeve, A.K.; Falkous, G.; et al. Subcellular Origin of Mitochondrial DNA Deletions in Human Skeletal Muscle. Ann. Neurol. 2018, 84, 289–301. [Google Scholar] [CrossRef]
- Kang, J.S.; Tian, J.H.; Pan, P.Y.; Zald, P.; Li, C.; Deng, C.; Sheng, Z.H. Docking of Axonal Mitochondria by Syntaphilin Controls Their Mobility and Affects Short-Term Facilitation. Cell 2008, 132, 137–148. [Google Scholar] [CrossRef]
- Miller, K.E.; Sheetz, M.P. Axonal Mitochondrial Transport and Potential Are Correlated. J. Cell Sci. 2004, 117, 2791–2804. [Google Scholar] [CrossRef]
- McWilliams, T.G.; Prescott, A.R.; Allen, G.F.G.; Tamjar, J.; Munson, M.J.; Thomson, C.; Muqit, M.M.K.; Ganley, I.G. Mito-QC Illuminates Mitophagy and Mitochondrial Architecture in Vivo. J. Cell Biol. 2016, 214, 333–345. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.F.; Lee, H.J.; Chi, C.S.; Tsai, C.R.; Chang, T.K.; Wang, C.J. The Neurological Evolution of Pearson Syndrome: Case Report and Literature Review. Eur. J. Paediatr. Neurol. 2007, 11, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Rocha, M.C.; Rosa, H.S.; Grady, J.P.; Blakely, E.L.; He, L.; Romain, N.; Haller, R.G.; Newman, J.; McFarland, R.; Ng, Y.S.; et al. Pathological Mechanisms Underlying Single Large-Scale Mitochondrial DNA Deletions. Ann. Neurol. 2018, 83, 115–130. [Google Scholar] [CrossRef]
- Pistilli, D.; di Gioia, C.R.T.; D’Amati, G.; Sciacchitano, S.; Quaglione, R.; Quitadamo, R.; Casali, C.; Gallo, P.; Santorelli, F.M. Detection of Deleted Mitochondrial DNA in Kearns-Sayre Syndrome Using Laser Capture Microdissection. Human. Pathol. 2003, 34, 1058–1061. [Google Scholar] [CrossRef] [PubMed]
- Savu, D.I.; Moisoi, N. Mitochondria—Nucleus Communication in Neurodegenerative Disease. Who Talks First, Who Talks Louder? Biochim. Et Biophys. Acta (BBA) Bioenerg. 2022, 1863, 148588. [Google Scholar] [CrossRef]
- Morgenstern, M.; Peikert, C.D.; Lübbert, P.; Suppanz, I.; Klemm, C.; Alka, O.; Steiert, C.; Naumenko, N.; Schendzielorz, A.; Melchionda, L.; et al. Quantitative High-Confidence Human Mitochondrial Proteome and Its Dynamics in Cellular Context. Cell Metab. 2021, 33, 2464–2483.e18. [Google Scholar] [CrossRef] [PubMed]
- Fairbrother-Browne, A.; Ali, A.T.; Reynolds, R.H.; Garcia-Ruiz, S.; Zhang, D.; Chen, Z.; Ryten, M.; Hodgkinson, A. Mitochondrial-Nuclear Cross-Talk in the Human Brain Is Modulated by Cell Type and Perturbed in Neurodegenerative Disease. Commun. Biol. 2021, 4, 1262. [Google Scholar] [CrossRef]
- Mottis, A.; Herzig, S.; Auwerx, J. Mitocellular Communication: Shaping Health and Disease. Science 2019, 366, 827–832. [Google Scholar] [CrossRef] [PubMed]
- Herzig, S.; Shaw, R.J. AMPK: Guardian of Metabolism and Mitochondrial Homeostasis. Nat. Rev. Mol. Cell Biol. 2018, 19, 121–135. [Google Scholar] [CrossRef]
- Verdin, E. NAD+ in Aging, Metabolism, and Neurodegeneration. Science 2015, 350, 1208–1213. [Google Scholar] [CrossRef]
- Jain, I.H.; Zazzeron, L.; Goli, R.; Alexa, K.; Schatzman-Bone, S.; Dhillon, H.; Goldberger, O.; Peng, J.; Shalem, O.; Sanjana, N.E.; et al. Hypoxia as a Therapy for Mitochondrial Disease. Science 2016, 352, 54–61. [Google Scholar] [CrossRef]
- Chandel, N.S.; Maltepe, E.; Goldwasser, E.; Mathieu, C.E.; Simon, M.C.; Schumacker, P.T. Mitochondrial Reactive Oxygen Species Trigger Hypoxia-Induced Transcription. Proc. Natl. Acad. Sci. USA 1998, 95, 11715–11720. [Google Scholar] [CrossRef]
- Diebold, L.; Chandel, N.S. Mitochondrial ROS Regulation of Proliferating Cells. Free Radic. Biol. Med. 2016, 100, 86–93. [Google Scholar] [CrossRef]
- Campbell, S.L.; Wellen, K.E. Metabolic Signaling to the Nucleus in Cancer. Mol. Cell 2018, 71, 398–408. [Google Scholar] [CrossRef]
- Ovciarikova, J.; Shikha, S.; Lacombe, A.; Courjol, F.; McCrone, R.; Hussain, W.; Maclean, A.; Lemgruber, L.; Martins-Duarte, E.S.; Gissot, M.; et al. Two Ancient Membrane Pores Mediate Mitochondrial-Nucleus Membrane Contact Sites. J. Cell Biol. 2024, 223, e202304075. [Google Scholar] [CrossRef]
- Desai, R.; East, D.A.; Hardy, L.; Faccenda, D.; Rigon, M.; Crosby, J.; Alvarez, M.S.; Singh, A.; Mainenti, M.; Hussey, L.K.; et al. Mitochondria Form Contact Sites with the Nucleus to Couple Prosurvival Retrograde Response. Sci. Adv. 2020, 6, eabc9955. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Li, C.; Yang, S.; Xiao, Y.; Xiong, X.; Chen, W.; Zhao, H.; Zhang, Q.; Han, Y.; Sun, L. Mitochondria-Associated ER Membranes—The Origin Site of Autophagy. Front. Cell Dev. Biol. 2020, 8, 595. [Google Scholar] [CrossRef] [PubMed]
- Cieri, D.; Vicario, M.; Giacomello, M.; Vallese, F.; Filadi, R.; Wagner, T.; Pozzan, T.; Pizzo, P.; Scorrano, L.; Brini, M.; et al. SPLICS: A Split Green Fluorescent Protein-Based Contact Site Sensor for Narrow and Wide Heterotypic Organelle Juxtaposition. Cell Death Differ. 2018, 25, 1131–1145. [Google Scholar] [CrossRef]
- Cisneros, J.; Belton, T.B.; Shum, G.C.; Molakal, C.G.; Wong, Y.C. Mitochondria-Lysosome Contact Site Dynamics and Misregulation in Neurodegenerative Diseases. Trends Neurosci. 2022, 45, 312–322. [Google Scholar] [CrossRef]
- Fransen, M.; Lismont, C.; Walton, P. The Peroxisome-Mitochondria Connection: How and Why? Int. J. Mol. Sci. 2017, 18, 1126. [Google Scholar] [CrossRef] [PubMed]
- Paillusson, S.; Stoica, R.; Gomez-Suaga, P.; Lau, D.H.W.; Mueller, S.; Miller, T.; Miller, C.C.J. There’s Something Wrong with My MAM; the ER–Mitochondria Axis and Neurodegenerative Diseases. Trends Neurosci. 2016, 39, 146–157. [Google Scholar] [CrossRef]
- Wong, Y.C.; Ysselstein, D.; Krainc, D. Mitochondria–Lysosome Contacts Regulate Mitochondrial Fission via RAB7 GTP Hydrolysis. Nature 2018, 554, 382–386. [Google Scholar] [CrossRef]
- Shai, N.; Yifrach, E.; van Roermund, C.W.T.; Cohen, N.; Bibi, C.; IJlst, L.; Cavellini, L.; Meurisse, J.; Schuster, R.; Zada, L.; et al. Systematic Mapping of Contact Sites Reveals Tethers and a Function for the Peroxisome-Mitochondria Contact. Nat. Commun. 2018, 9, 1761. [Google Scholar] [CrossRef]
- English, A.M.; Schuler, M.H.; Xiao, T.; Kornmann, B.; Shaw, J.M.; Hughes, A.L. ER-Mitochondria Contacts Promote Mitochondrial-Derived Compartment Biogenesis. J. Cell Biol. 2020, 219, e202002144. [Google Scholar] [CrossRef]
- Sugiura, A.; McLelland, G.; Fon, E.A.; McBride, H.M. A New Pathway for Mitochondrial Quality Control: Mitochondrial-derived Vesicles. EMBO J. 2014, 33, 2142–2156. [Google Scholar] [CrossRef] [PubMed]
- König, T.; McBride, H.M. Mitochondrial-Derived Vesicles in Metabolism, Disease, and Aging. Cell Metab. 2024, 36, 21–35. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liao, Z.; Zhang, Q.; Han, X.; Liu, C.; Wang, J. Mitochondrial Dysfunction in Alzheimer’s Disease: A Key Frontier for Future Targeted Therapies. Front. Immunol. 2025, 15, 1484373. [Google Scholar] [CrossRef] [PubMed]
- Zvěřová, M. Clinical Aspects of Alzheimer’s Disease. Clin. Biochem. 2019, 72, 3–6. [Google Scholar] [CrossRef]
- Liu, T.; Sun, W.; Guo, S.; Yuan, Z.; Zhu, M.; Lu, J.; Chen, T.; Qu, Y.; Feng, C.; Yang, T. Role of Mitochondrial Quality Control in Neurodegenerative Disease Progression. Front. Cell Neurosci. 2025, 19, 1588645. [Google Scholar] [CrossRef]
- Swerdlow, R.H. Mitochondria and Mitochondrial Cascades in Alzheimer’s Disease. J. Alzheimer’s Dis. 2018, 62, 1403–1416. [Google Scholar] [CrossRef]
- Guha, S.; Johnson, G.V.W.; Nehrke, K. The Crosstalk Between Pathological Tau Phosphorylation and Mitochondrial Dysfunction as a Key to Understanding and Treating Alzheimer’s Disease. Mol. Neurobiol. 2020, 57, 5103–5120. [Google Scholar] [CrossRef]
- Perluigi, M.; Di Domenico, F.; Butterfield, D.A. Oxidative Damage in Neurodegeneration: Roles in the Pathogenesis and Progression of Alzheimer Disease. Physiol. Rev. 2024, 104, 103–197. [Google Scholar] [CrossRef]
- Mary, A.; Eysert, F.; Checler, F.; Chami, M. Mitophagy in Alzheimer’s Disease: Molecular Defects and Therapeutic Approaches. Mol. Psychiatry 2023, 28, 202–216. [Google Scholar] [CrossRef]
- Colacurcio, D.J.; Nixon, R.A. Disorders of Lysosomal Acidification-The Emerging Role of v-ATPase in Aging and Neurodegenerative Disease. Ageing Res. Rev. 2016, 32, 75–88. [Google Scholar] [CrossRef]
- Jiang, X.J.; Wu, Y.Q.; Ma, R.; Chang, Y.M.; Li, L.L.; Zhu, J.H.; Liu, G.P.; Li, G. PINK1 Alleviates Cognitive Impairments via Attenuating Pathological Tau Aggregation in a Mouse Model of Tauopathy. Front. Cell Dev. Biol. 2022, 9, 736267. [Google Scholar] [CrossRef]
- Hou, S.J.; Zhang, S.X.; Li, Y.; Xu, S.Y. Rapamycin Responds to Alzheimer’s Disease: A Potential Translational Therapy. Clin. Interv. Aging 2023, 18, 1629–1639. [Google Scholar] [CrossRef]
- Hou, Y.; Chu, X.; Park, J.H.; Zhu, Q.; Hussain, M.; Li, Z.; Madsen, H.B.; Yang, B.; Wei, Y.; Wang, Y.; et al. Urolithin A Improves Alzheimer’s Disease Cognition and Restores Mitophagy and Lysosomal Functions. Alzheimer’s Dement. 2024, 20, 4212–4233. [Google Scholar] [CrossRef]
- Vaillant-Beuchot, L.; Eysert, F.; Duval, B.; Kinoshita, P.F.; Pardossi-Piquard, R.; Bauer, C.; Eddarkaoui, S.; Buée, L.; Checler, F.; Chami, M. The Amyloid Precursor Protein and Its Derived Fragments Concomitantly Contribute to the Alterations of Mitochondrial Transport Machinery in Alzheimer’s Disease. Cell Death Dis. 2024, 15, 367. [Google Scholar] [CrossRef]
- Wang, Q.; Tian, J.; Chen, H.; Du, H.; Guo, L. Amyloid Beta-Mediated KIF5A Deficiency Disrupts Anterograde Axonal Mitochondrial Movement. Neurobiol. Dis. 2019, 127, 410–418. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Yin, J.; Ma, X.; Zhao, F.; Siedlak, S.L.; Wang, Z.; Torres, S.; Fujioka, H.; Xu, Y.; Perry, G.; et al. Inhibition of Mitochondrial Fragmentation Protects against Alzheimer’s Disease in Rodent Model. Human. Mol. Genet. 2017, 26, 4118–4131. [Google Scholar] [CrossRef] [PubMed]
- Bhatti, J.S.; Kaur, S.; Mishra, J.; Dibbanti, H.; Singh, A.; Reddy, A.P.; Bhatti, G.K.; Reddy, P.H. Targeting Dynamin-Related Protein-1 as a Potential Therapeutic Approach for Mitochondrial Dysfunction in Alzheimer’s Disease. Biochim. Biophys. Acta Mol. Basis Dis. 2023, 1869, 166798. [Google Scholar] [CrossRef] [PubMed]
- Bera, A.; Lavanya, G.; Reshmi, R.; Dev, K.; Kumar, R. Mechanistic and Therapeutic Role of Drp1 in the Pathogenesis of Alzheimer’s Disease. Eur. J. Neurosci. 2022, 56, 5516–5531. [Google Scholar] [CrossRef]
- Pakpian, N.; Phopin, K.; Kitidee, K.; Govitrapong, P.; Wongchitrat, P. Alterations in Mitochondrial Dynamic-Related Genes in the Peripheral Blood of Alzheimer’s Disease Patients. Curr. Alzheimer Res. 2020, 17, 616–625. [Google Scholar] [CrossRef]
- Joshi, A.U.; Saw, N.L.; Shamloo, M.; Mochly-Rosen, D. Drp1/Fis1 Interaction Mediates Mitochondrial Dysfunction, Bioenergetic Failure and Cognitive Decline in Alzheimer’s Disease. Oncotarget 2017, 9, 6128–6143. [Google Scholar] [CrossRef]
- Han, A.R.; Moon, T.K.; Kang, I.K.; Yu, D.B.; Kim, Y.; Byon, C.; Park, S.; Kim, H.L.; Kim, H.L.; Lee, K.J.; et al. Integrative Analysis of microRNA-Mediated Mitochondrial Dysfunction in Hippocampal Neural Progenitor Cell Death in Relation with Alzheimer’s Disease. BMB Rep. 2024, 57, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Elsherbini, A.; Kirov, A.S.; Dinkins, M.B.; Wang, G.; Qin, H.; Zhu, Z.; Tripathi, P.; Crivelli, S.M.; Bieberich, E. Association of Aβ with Ceramide-Enriched Astrosomes Mediates Aβ Neurotoxicity. Acta Neuropathol. Commun. 2020, 8, 60. [Google Scholar] [CrossRef]
- Sbai, O.; Bazzani, V.; Tapaswi, S.; McHale, J.; Vascotto, C.; Perrone, L. Is Drp1 a Link between Mitochondrial Dysfunction and Inflammation in Alzheimer’s Disease? Front. Mol. Neurosci. 2023, 16, 1166879. [Google Scholar] [CrossRef]
- Zhang, M.; Hu, G.; Shao, N.; Qin, Y.; Chen, Q.; Wang, Y.; Zhou, P.; Cai, B. Thioredoxin-Interacting Protein (TXNIP) as a Target for Alzheimer’s Disease: Flavonoids and Phenols. Inflammopharmacology 2021, 29, 1317–1329. [Google Scholar] [CrossRef] [PubMed]
- Kerr, J.S.; Adriaanse, B.A.; Greig, N.H.; Mattson, M.P.; Cader, M.Z.; Bohr, V.A.; Fang, E.F. Mitophagy and Alzheimer’s Disease: Cellular and Molecular Mechanisms. Trends Neurosci. 2017, 40, 151–166. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.N.; Zhang, M.Q.; Yu, F.L.; Han, B.; Bao, M.Y.; Li, X.; Zhang, Y. Peroxisom Proliferator-Activated Receptor-γ Coactivator-1α in Neurodegenerative Disorders: A Promising Therapeutic Target. Biochem. Pharmacol. 2023, 215, 115717. [Google Scholar] [CrossRef]
- Katsouri, L.; Lim, Y.M.; Blondrath, K.; Eleftheriadou, I.; Lombardero, L.; Birch, A.M.; Mirzaei, N.; Irvine, E.E.; Mazarakis, N.D.; Sastre, M. PPARγ-Coactivator-1α Gene Transfer Reduces Neuronal Loss and Amyloid-β Generation by Reducing β-Secretase in an Alzheimer’s Disease Model. Proc. Natl. Acad. Sci. USA 2016, 113, 12292–12297. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, C.; Jiang, Y.; Wang, S.; Wu, X.; Wang, K. PPARγ Coactivator-1α (PGC-1α) Protects Neuroblastoma Cells against Amyloid-Beta (Aβ) Induced Cell Death and Neuroinflammation via NF-κB Pathway. BMC Neurosci. 2017, 18, 69. [Google Scholar] [CrossRef]
- Oka, S.; Leon, J.; Sakumi, K.; Ide, T.; Kang, D.; LaFerla, F.M.; Nakabeppu, Y. Human Mitochondrial Transcriptional Factor A Breaks the Mitochondria-Mediated Vicious Cycle in Alzheimer’s Disease. Sci. Rep. 2016, 6, 37889. [Google Scholar] [CrossRef] [PubMed]
- Mhyre, T.R.; Boyd, J.T.; Hamill, R.W.; Maguire-Zeiss, K.A. Parkinson’s Disease. Subcell. Biochem. 2012, 65, 389–455. [Google Scholar] [CrossRef]
- Fujiwara, H.; Hasegawa, M.; Dohmae, N.; Kawashima, A.; Masliah, E.; Goldberg, M.S.; Shen, J.; Takio, K.; Iwatsubo, T. α-Synuclein Is Phosphorylated in Synucleinopathy Lesions. Nat. Cell Biol. 2002, 4, 160–164. [Google Scholar] [CrossRef]
- Dimant, H.; Ebrahimi-Fakhari, D.; McLean, P.J. Molecular Chaperones and Co-Chaperones in Parkinson Disease. Neuroscientist 2012, 18, 589–601. [Google Scholar] [CrossRef]
- Langston, J.W.; Ballard, P.; Tetrud, J.W.; Irwin, I. Chronic Parkinsonism in Humans Due to a Product of Meperidine-Analog Synthesis. Science 1983, 219, 979–980. [Google Scholar] [CrossRef]
- Labbé, C.; Ross, O.A. Association Studies of Sporadic Parkinson’s Disease in the Genomic Era. Curr. Genom. 2014, 15, 2–10. [Google Scholar] [CrossRef]
- Valente, E.M.; Abou-Sleiman, P.M.; Caputo, V.; Muqit, M.M.K.; Harvey, K.; Gispert, S.; Ali, Z.; Del Turco, D.; Bentivoglio, A.R.; Healy, D.G.; et al. Hereditary Early-Onset Parkinson’s Disease Caused by Mutations in PINK1. Science 2004, 304, 1158–1160. [Google Scholar] [CrossRef]
- Darios, F.; Corti, O.; Lücking, C.B.; Hampe, C.; Muriel, M.-P.; Abbas, N.; Gu, W.J.; Hirsch, E.C.; Rooney, T.; Ruberg, M.; et al. Parkin Prevents Mitochondrial Swelling and Cytochrome c Release in Mitochondria-Dependent Cell Death. Hum. Mol. Genet. 2003, 12, 517–526. [Google Scholar] [CrossRef] [PubMed]
- Morais, V.A.; Verstreken, P.; Roethig, A.; Smet, J.; Snellinx, A.; Vanbrabant, M.; Haddad, D.; Frezza, C.; Mandemakers, W.; Vogt-Weisenhorn, D.; et al. Parkinson’s Disease Mutations in PINK1 Result in Decreased Complex I Activity and Deficient Synaptic Function. EMBO Mol. Med. 2009, 1, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.H.; Song, K.; Yoon, S.H.; Shehzad, O.; Kim, Y.S.; Son, J.H. Rescue of PINK1 Protein Null-Specific Mitochondrial Complex IV Deficits by Ginsenoside Re Activation of Nitric Oxide Signaling. J. Biol. Chem. 2012, 287, 44109–44120. [Google Scholar] [CrossRef]
- Clark, E.H.; Vázquez de la Torre, A.; Hoshikawa, T.; Briston, T. Targeting Mitophagy in Parkinson’s Disease. J. Biol. Chem. 2021, 296, 100209. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Chen, Y.; Zhuo, J.; Zhang, L.; Liu, J.; Wang, B.; Sun, D.; Yu, S.; Lou, H. Urolithin A Promotes Mitophagy and Suppresses NLRP3 Inflammasome Activation in Lipopolysaccharide-Induced BV2 Microglial Cells and MPTP-Induced Parkinson’s Disease Model. Neuropharmacology 2022, 207, 108963. [Google Scholar] [CrossRef] [PubMed]
- Kuerec, A.H.; Lim, X.K.; Khoo, A.L.Y.; Sandalova, E.; Guan, L.; Feng, L.; Maier, A.B. Targeting Aging with Urolithin A in Humans: A Systematic Review. Ageing Res. Rev. 2024, 100, 102406. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; D’Amico, D.; Andreux, P.A.; Fouassier, A.M.; Blanco-Bose, W.; Evans, M.; Aebischer, P.; Auwerx, J.; Rinsch, C. Urolithin A Improves Muscle Strength, Exercise Performance, and Biomarkers of Mitochondrial Health in a Randomized Trial in Middle-Aged Adults. Cell Rep. Med. 2022, 3, 100633. [Google Scholar] [CrossRef]
- Castagnet, P.I.; Golovko, M.Y.; Barceló-Coblijn, G.C.; Nussbaum, R.L.; Murphy, E.J. Fatty Acid Incorporation Is Decreased in Astrocytes Cultured from Alpha-Synuclein Gene-Ablated Mice. J. Neurochem. 2005, 94, 839–849. [Google Scholar] [CrossRef]
- Chung, Y.C.; Baek, J.Y.; Kim, S.R.; Ko, H.W.; Bok, E.; Shin, W.H.; Won, S.Y.; Jin, B.K. Capsaicin Prevents Degeneration of Dopamine Neurons by Inhibiting Glial Activation and Oxidative Stress in the MPTP Model of Parkinson’s Disease. Exp. Mol. Med. 2017, 49, e298. [Google Scholar] [CrossRef]
- Brakedal, B.; Dölle, C.; Riemer, F.; Ma, Y.; Nido, G.S.; Skeie, G.O.; Craven, A.R.; Schwarzlmüller, T.; Brekke, N.; Diab, J.; et al. The NADPARK Study: A Randomized Phase I Trial of Nicotinamide Riboside Supplementation in Parkinson’s Disease. Cell Metab. 2022, 34, 396–407.e6. [Google Scholar] [CrossRef]
- Cai, R.; Zhang, Y.; Simmering, J.E.; Schultz, J.L.; Li, Y.; Fernandez-Carasa, I.; Consiglio, A.; Raya, A.; Polgreen, P.M.; Narayanan, N.S.; et al. Enhancing Glycolysis Attenuates Parkinson’s Disease Progression in Models and Clinical Databases. J. Clin. Investig. 2019, 129, 4539–4549. [Google Scholar] [CrossRef]
- Junn, E.; Jang, W.H.; Zhao, X.; Jeong, B.S.; Mouradian, M.M. Mitochondrial Localization of DJ-1 Leads to Enhanced Neuroprotection. J. Neurosci. Res. 2009, 87, 123–129. [Google Scholar] [CrossRef]
- Skou, L.D.; Johansen, S.K.; Okarmus, J.; Meyer, M. Pathogenesis of DJ-1/PARK7-Mediated Parkinson’s Disease. Cells 2024, 13, 296. [Google Scholar] [CrossRef]
- Zhou, W.; Bercury, K.; Cummiskey, J.; Luong, N.; Lebin, J.; Freed, C.R. Phenylbutyrate Up-Regulates the DJ-1 Protein and Protects Neurons in Cell Culture and in Animal Models of Parkinson Disease. J. Biol. Chem. 2011, 286, 14941–14951. [Google Scholar] [CrossRef]
- Duarte, P.; Michalska, P.; Crisman, E.; Cuadrado, A.; León, R. Novel Series of Dual NRF2 Inducers and Selective MAO-B Inhibitors for the Treatment of Parkinson’s Disease. Antioxidants 2022, 11, 247. [Google Scholar] [CrossRef]
- Wu, N.; Luo, Z.; Deng, R.; Zhang, Z.; Zhang, J.; Liu, S.; Luo, Z.; Qi, Q. Sulforaphane: An Emerging Star in Neuroprotection and Neurological Disease Prevention. Biochem. Pharmacol. 2025, 233, 116797. [Google Scholar] [CrossRef]
- Rawlins, M.D.; Wexler, N.S.; Wexler, A.R.; Tabrizi, S.J.; Douglas, I.; Evans, S.J.W.; Smeeth, L. The Prevalence of Huntington’s Disease. Neuroepidemiology 2016, 46, 144–153. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, M.E.; Ambrose, C.M.; Duyao, M.P.; Myers, R.H.; Lin, C.; Srinidhi, L.; Barnes, G.; Taylor, S.A.; James, M.; Groot, N.; et al. A Novel Gene Containing a Trinucleotide Repeat That Is Expanded and Unstable on Huntington’s Disease Chromosomes. Cell 1993, 72, 971–983. [Google Scholar] [CrossRef]
- Jodeiri Farshbaf, M.; Ghaedi, K. Huntington’s Disease and Mitochondria. Neurotox. Res. 2017, 32, 518–529. [Google Scholar] [CrossRef] [PubMed]
- Franco-Iborra, S.; Plaza-Zabala, A.; Montpeyo, M.; Sebastian, D.; Vila, M.; Martinez-Vicente, M. Mutant HTT (Huntingtin) Impairs Mitophagy in a Cellular Model of Huntington Disease. Autophagy 2021, 17, 672–689. [Google Scholar] [CrossRef]
- Funayama, M.; Nishioka, K.; Li, Y.; Hattori, N. Molecular Genetics of Parkinson’s Disease: Contributions and Global Trends. J. Hum. Genet. 2023, 68, 125–130. [Google Scholar] [CrossRef] [PubMed]
- Joshi, D.C.; Chavan, M.B.; Gurow, K.; Gupta, M.; Dhaliwal, J.S.; Ming, L.C. The Role of Mitochondrial Dysfunction in Huntington’s Disease: Implications for Therapeutic Targeting. Biomed. Pharmacother. 2025, 183, 117827. [Google Scholar] [CrossRef]
- Guedes-Dias, P.; Pinho, B.R.; Soares, T.R.; de Proença, J.; Duchen, M.R.; Oliveira, J.M.A. Mitochondrial Dynamics and Quality Control in Huntington’s Disease. Neurobiol. Dis. 2016, 90, 51–57. [Google Scholar] [CrossRef]
- McColgan, P.; Tabrizi, S.J. Huntington’s Disease: A Clinical Review. Eur. J. Neurol. 2018, 25, 24–34. [Google Scholar] [CrossRef] [PubMed]
- Cui, L.; Jeong, H.; Borovecki, F.; Parkhurst, C.N.; Tanese, N.; Krainc, D. Transcriptional Repression of PGC-1α by Mutant Huntingtin Leads to Mitochondrial Dysfunction and Neurodegeneration. Cell 2006, 127, 59–69. [Google Scholar] [CrossRef]
- Thau, N.; Knippenberg, S.; Körner, S.; Rath, K.J.; Dengler, R.; Petri, S. Decreased mRNA Expression of PGC-1α and PGC-1α-Regulated Factors in the SOD1G93A ALS Mouse Model and in Human Sporadic ALS. J. Neuropathol. Exp. Neurol. 2012, 71, 1064–1074. [Google Scholar] [CrossRef] [PubMed]
- Johri, A.; Calingasan, N.Y.; Hennessey, T.M.; Sharma, A.; Yang, L.; Wille, E.; Chandra, A.; Beal, M.F. Pharmacologic Activation of Mitochondrial Biogenesis Exerts Widespread Beneficial Effects in a Transgenic Mouse Model of Huntington’s Disease. Hum. Mol. Genet. 2012, 21, 1124–1137. [Google Scholar] [CrossRef] [PubMed]
- Khalil, B.; El Fissi, N.; Aouane, A.; Cabirol-Pol, M.J.; Rival, T.; Liévens, J.C. PINK1-Induced Mitophagy Promotes Neuroprotection in Huntington’s Disease. Cell Death Dis. 2015, 6, e1617. [Google Scholar] [CrossRef]
- Wong, Y.C.; Holzbaur, E.L.F. The Regulation of Autophagosome Dynamics by Huntingtin and HAP1 Is Disrupted by Expression of Mutant Huntingtin, Leading to Defective Cargo Degradation. J. Neurosci. 2014, 34, 1293–1305. [Google Scholar] [CrossRef]
- Cherubini, M.; Lopez-Molina, L.; Gines, S. Mitochondrial Fission in Huntington’s Disease Mouse Striatum Disrupts ER-Mitochondria Contacts Leading to Disturbances in Ca2+ Efflux and Reactive Oxygen Species (ROS) Homeostasis. Neurobiol. Dis. 2020, 136, 104741. [Google Scholar] [CrossRef]
- Davies, S.W.; Turmaine, M.; Cozens, B.A.; DiFiglia, M.; Sharp, A.H.; Ross, C.A.; Scherzinger, E.; Wanker, E.E.; Mangiarini, L.; Bates, G.P. Formation of Neuronal Intranuclear Inclusions Underlies the Neurological Dysfunction in Mice Transgenic for the HD Mutation. Cell 1997, 90, 537–548. [Google Scholar] [CrossRef]
- Sawant, N.; Morton, H.; Kshirsagar, S.; Reddy, A.P.; Reddy, P.H. Mitochondrial Abnormalities and Synaptic Damage in Huntington’s Disease: A Focus on Defective Mitophagy and Mitochondria-Targeted Therapeutics. Mol. Neurobiol. 2021, 58, 6350–6377. [Google Scholar] [CrossRef]
- Hu, D.; Sun, X.; Magpusao, A.; Fedorov, Y.; Thompson, M.; Wang, B.; Lundberg, K.; Adams, D.J.; Qi, X. Small-Molecule Suppression of Calpastatin Degradation Reduces Neuropathology in Models of Huntington’s Disease. Nat. Commun. 2021, 12, 5305. [Google Scholar] [CrossRef]
- Brotman, R.G.; Moreno-Escobar, M.C.; Joseph, J.; Munakomi, S.; Pawar, G. Amyotrophic Lateral Sclerosis. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2025. [Google Scholar]
- Cozzolino, M.; Rossi, S.; Mirra, A.; Carrì, M.T. Mitochondrial Dynamism and the Pathogenesis of Amyotrophic Lateral Sclerosis. Front. Cell. Neurosci. 2015, 9, 31. [Google Scholar] [CrossRef]
- Genin, E.C.; Abou-Ali, M.; Paquis-Flucklinger, V. Mitochondria, a Key Target in Amyotrophic Lateral Sclerosis Pathogenesis. Genes. 2023, 14, 1981. [Google Scholar] [CrossRef]
- Shi, Y.; Lin, S.; Staats, K.A.; Li, Y.; Chang, W.-H.; Hung, S.T.; Hendricks, E.; Linares, G.R.; Wang, Y.; Son, E.Y.; et al. Haploinsufficiency Leads to Neurodegeneration in C9ORF72 ALS/FTD Human Induced Motor Neurons. Nat. Med. 2018, 24, 313–325. [Google Scholar] [CrossRef]
- Wang, W.; Wang, L.; Lu, J.; Siedlak, S.L.; Fujioka, H.; Liang, J.; Jiang, S.; Ma, X.; Jiang, Z.; da Rocha, E.L.; et al. The Inhibition of TDP-43 Mitochondrial Localization Blocks Its Neuronal Toxicity. Nat. Med. 2016, 22, 869–878. [Google Scholar] [CrossRef]
- Deng, J.; Yang, M.; Chen, Y.; Chen, X.; Liu, J.; Sun, S.; Cheng, H.; Li, Y.; Bigio, E.H.; Mesulam, M.; et al. FUS Interacts with HSP60 to Promote Mitochondrial Damage. PLOS Genet. 2015, 11, e1005357. [Google Scholar] [CrossRef]
- Luan, P.; D’Amico, D.; Andreux, P.A.; Laurila, P.P.; Wohlwend, M.; Li, H.; Lima, T.I.d.; Place, N.; Rinsch, C.; Zanou, N.; et al. Urolithin A Improves Muscle Function by Inducing Mitophagy in Muscular Dystrophy. Sci. Transl. Med. 2021, 13, eabb0319. [Google Scholar] [CrossRef]
- Osaki, T.; Uzel, S.G.M.; Kamm, R.D. Microphysiological 3D Model of Amyotrophic Lateral Sclerosis (ALS) from Human iPS-Derived Muscle Cells and Optogenetic Motor Neurons. Sci. Adv. 2018, 4, eaat5847. [Google Scholar] [CrossRef] [PubMed]
- Safety and Efficacy of Trehalose in Amyotrophic Lateral Sclerosis (HEALEY ALS Platform Trial): An Adaptive, Phase 2/3, Double-Blind, Randomised, Placebo-Controlled Trial. Lancet Neurol. 2025, 24, 500–511. [CrossRef] [PubMed]
- Pham, J.; Keon, M.; Brennan, S.; Saksena, N. Connecting RNA-Modifying Similarities of TDP-43, FUS, and SOD1 with MicroRNA Dysregulation Amidst A Renewed Network Perspective of Amyotrophic Lateral Sclerosis Proteinopathy. Int. J. Mol. Sci. 2020, 21, 3464. [Google Scholar] [CrossRef]
- Xu, Y.F.; Gendron, T.F.; Zhang, Y.J.; Lin, W.L.; D’Alton, S.; Sheng, H.; Casey, M.C.; Tong, J.; Knight, J.; Yu, X.; et al. Wild-Type Human TDP-43 Expression Causes TDP-43 Phosphorylation, Mitochondrial Aggregation, Motor Deficits, and Early Mortality in Transgenic Mice. J. Neurosci. 2010, 30, 10851–10859. [Google Scholar] [CrossRef] [PubMed]
- Akkum, F.I.; Ozbas, C.E.; Damar, M.; Uversky, V.N.; Fayetorbay, R.; Kang, D.E.; Woo, J.A.; Coskuner-Weber, O. Impacts of Pathogenic Mutations on the Structures of the CHCHD10 Monomer: An AlphaFold3 Study Linked to the Generation of Conformational Ensembles. Int. J. Biol. Macromol. 2025, 318, 144970. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.Y.; Lee, J.H.; Chung, A.-Y.; Jo, Y.; Shin, J.H.; Park, H.C.; Kim, H.; Lopez-Gonzalez, R.; Ryu, J.R.; Sun, W. Prevention of Mitochondrial Impairment by Inhibition of Protein Phosphatase 1 Activity in Amyotrophic Lateral Sclerosis. Cell Death Dis. 2020, 11, 888. [Google Scholar] [CrossRef]
- Yu, C.H.; Davidson, S.; Harapas, C.R.; Hilton, J.B.; Mlodzianoski, M.J.; Laohamonthonkul, P.; Louis, C.; Low, R.R.J.; Moecking, J.; De Nardo, D.; et al. TDP-43 Triggers Mitochondrial DNA Release via mPTP to Activate cGAS/STING in ALS. Cell 2020, 183, 636–649.e18. [Google Scholar] [CrossRef]
- Martin, L.J. The Mitochondrial Permeability Transition Pore: A Molecular Target for Amyotrophic Lateral Sclerosis Therapy. Biochim. Biophys. Acta 2010, 1802, 186–197. [Google Scholar] [CrossRef]
- Witzel, S.; Maier, A.; Steinbach, R.; Grosskreutz, J.; Koch, J.C.; Sarikidi, A.; Petri, S.; Günther, R.; Wolf, J.; Hermann, A.; et al. Safety and Effectiveness of Long-Term Intravenous Administration of Edaravone for Treatment of Patients with Amyotrophic Lateral Sclerosis. JAMA Neurol. 2022, 79, 121–130. [Google Scholar] [CrossRef]
- Varghese, M.; Zhao, W.; Trageser, K.J.; Pasinetti, G.M. Peroxisome Proliferator Activator Receptor Gamma Coactivator-1α Overexpression in Amyotrophic Lateral Sclerosis: A Tale of Two Transgenics. Biomolecules 2020, 10, 760. [Google Scholar] [CrossRef] [PubMed]
- Mancuso, R.; del Valle, J.; Modol, L.; Martinez, A.; Granado-Serrano, A.B.; Ramirez-Núñez, O.; Pallás, M.; Portero-Otin, M.; Osta, R.; Navarro, X. Resveratrol Improves Motoneuron Function and Extends Survival in SOD1G93A ALS Mice. Neurotherapeutics 2014, 11, 419–432. [Google Scholar] [CrossRef]
- Ashwani; Sharma, A.; Choudhary, M.K.; Gugulothu, D.; Pandita, D.; Verma, S.; Vora, L.K.; Khatri, D.K.; Garabadu, D. Epigenetic and Mitochondrial Metabolic Dysfunction in Multiple Sclerosis: A Review of Herbal Drug Approaches and Current Clinical Trials. Mol. Neurobiol. 2025, 62, 10045–10090. [Google Scholar] [CrossRef]
- Pegoretti, V.; Swanson, K.A.; Bethea, J.R.; Probert, L.; Eisel, U.L.M.; Fischer, R. Inflammation and Oxidative Stress in Multiple Sclerosis: Consequences for Therapy Development. Oxid. Med. Cell Longev. 2020, 2020, 7191080. [Google Scholar] [CrossRef] [PubMed]
- Karuppagounder, S.S.; Brahmachari, S.; Lee, Y.; Dawson, V.L.; Dawson, T.M.; Ko, H.S. The C-Abl Inhibitor, Nilotinib, Protects Dopaminergic Neurons in a Preclinical Animal Model of Parkinson’s Disease. Sci. Rep. 2014, 4, 4874. [Google Scholar] [CrossRef]
- Manczak, M.; Reddy, P.H. Mitochondrial Division Inhibitor 1 Protects against Mutant Huntingtin-Induced Abnormal Mitochondrial Dynamics and Neuronal Damage in Huntington’s Disease. Hum. Mol. Genet. 2015, 24, 7308–7325. [Google Scholar] [CrossRef]
- Ciuro, M.; Sangiorgio, M.; Cacciato, V.; Cantone, G.; Fichera, C.; Salvatorelli, L.; Magro, G.; Leanza, G.; Vecchio, M.; Valle, M.S.; et al. Mitigating the Functional Deficit after Neurotoxic Motoneuronal Loss by an Inhibitor of Mitochondrial Fission. Int. J. Mol. Sci. 2024, 25, 7059. [Google Scholar] [CrossRef] [PubMed]
- Marx, N.; Ritter, N.; Disse, P.; Seebohm, G.; Busch, K.B. Detailed Analysis of Mdivi-1 Effects on Complex I and Respiratory Supercomplex Assembly. Sci. Rep. 2024, 14, 19673. [Google Scholar] [CrossRef] [PubMed]
- Kaufmann, P.; Thompson, J.L.P.; Levy, G.; Buchsbaum, R.; Shefner, J.; Krivickas, L.S.; Katz, J.; Rollins, Y.; Barohn, R.J.; Jackson, C.E.; et al. Phase II Trial of CoQ10 for ALS Finds Insufficient Evidence to Justify Phase III. Ann. Neurol. 2009, 66, 235–244. [Google Scholar] [CrossRef] [PubMed]
- McGarry, A.; McDermott, M.; Kieburtz, K.; de Blieck, E.A.; Beal, F.; Marder, K.; Ross, C.; Shoulson, I.; Gilbert, P.; Mallonee, W.M.; et al. A Randomized, Double-Blind, Placebo-Controlled Trial of Coenzyme Q10 in Huntington Disease. Neurology 2017, 88, 152–159. [Google Scholar] [CrossRef]
- Gonzales, M.M.; Garbarino, V.R.; Kautz, T.F.; Song, X.; Lopez-Cruzan, M.; Linehan, L.; Van Skike, C.E.; De Erausquin, G.A.; Galvan, V.; Orr, M.E.; et al. Rapamycin Treatment for Alzheimer’s Disease and Related Dementias: A Pilot Phase 1 Clinical Trial. Commun. Med. 2025, 5, 189. [Google Scholar] [CrossRef]
- Bennett, C.F.; Krainer, A.R.; Cleveland, D.W. Antisense Oligonucleotide Therapies for Neurodegenerative Diseases. Annu. Rev. Neurosci. 2019, 42, 385–406. [Google Scholar] [CrossRef]
- Ansori, A.N.M.; Antonius, Y.; Susilo, R.J.K.; Hayaza, S.; Kharisma, V.D.; Parikesit, A.A.; Zainul, R.; Jakhmola, V.; Saklani, T.; Rebezov, M.; et al. Application of CRISPR-Cas9 Genome Editing Technology in Various Fields: A Review. Narra J. 2023, 3, e184. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Steffan, D.; Pezzini, C.; Esposito, M.; Franco-Romero, A. Mitochondrial Aging in the CNS: Unravelling Implications for Neurological Health and Disease. Biomolecules 2025, 15, 1252. https://doi.org/10.3390/biom15091252
Steffan D, Pezzini C, Esposito M, Franco-Romero A. Mitochondrial Aging in the CNS: Unravelling Implications for Neurological Health and Disease. Biomolecules. 2025; 15(9):1252. https://doi.org/10.3390/biom15091252
Chicago/Turabian StyleSteffan, Davide, Camilla Pezzini, Martina Esposito, and Anais Franco-Romero. 2025. "Mitochondrial Aging in the CNS: Unravelling Implications for Neurological Health and Disease" Biomolecules 15, no. 9: 1252. https://doi.org/10.3390/biom15091252
APA StyleSteffan, D., Pezzini, C., Esposito, M., & Franco-Romero, A. (2025). Mitochondrial Aging in the CNS: Unravelling Implications for Neurological Health and Disease. Biomolecules, 15(9), 1252. https://doi.org/10.3390/biom15091252








