Nicotinamide N-Methyltransferase as Promising Tool for Management of Gastrointestinal Neoplasms
Abstract
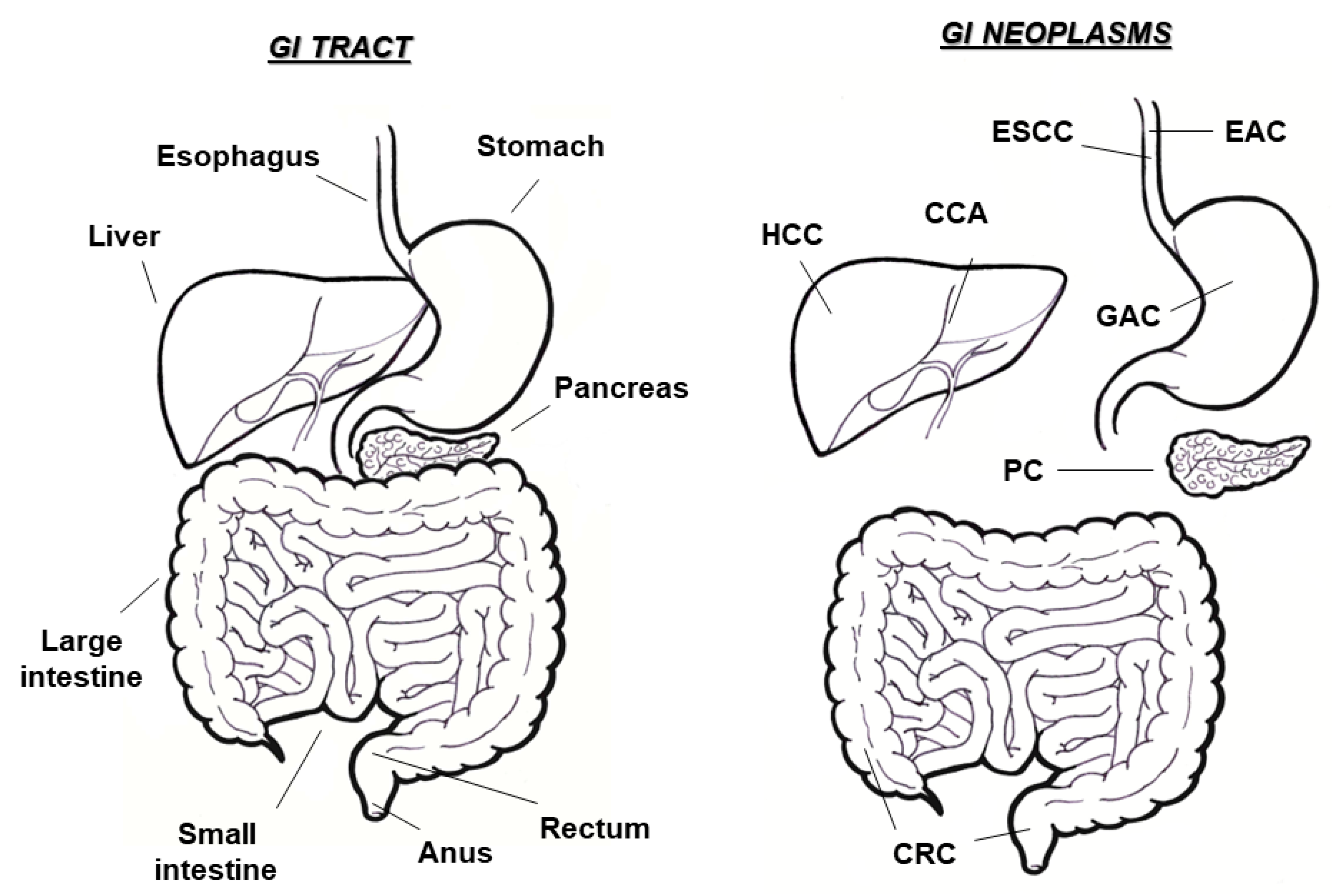
1. Gastrointestinal Cancers
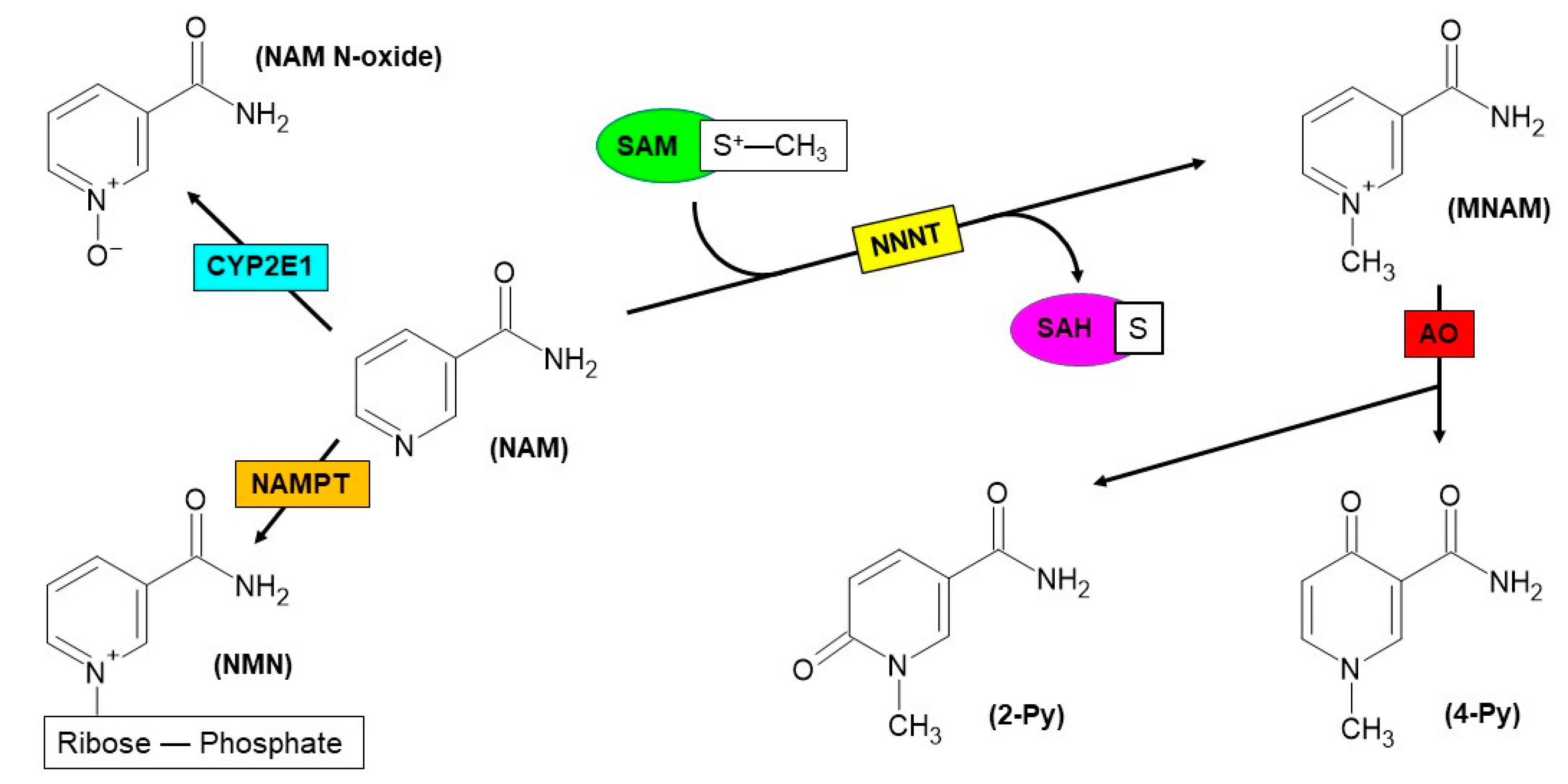
2. Nicotinamide N-Methyltransferase
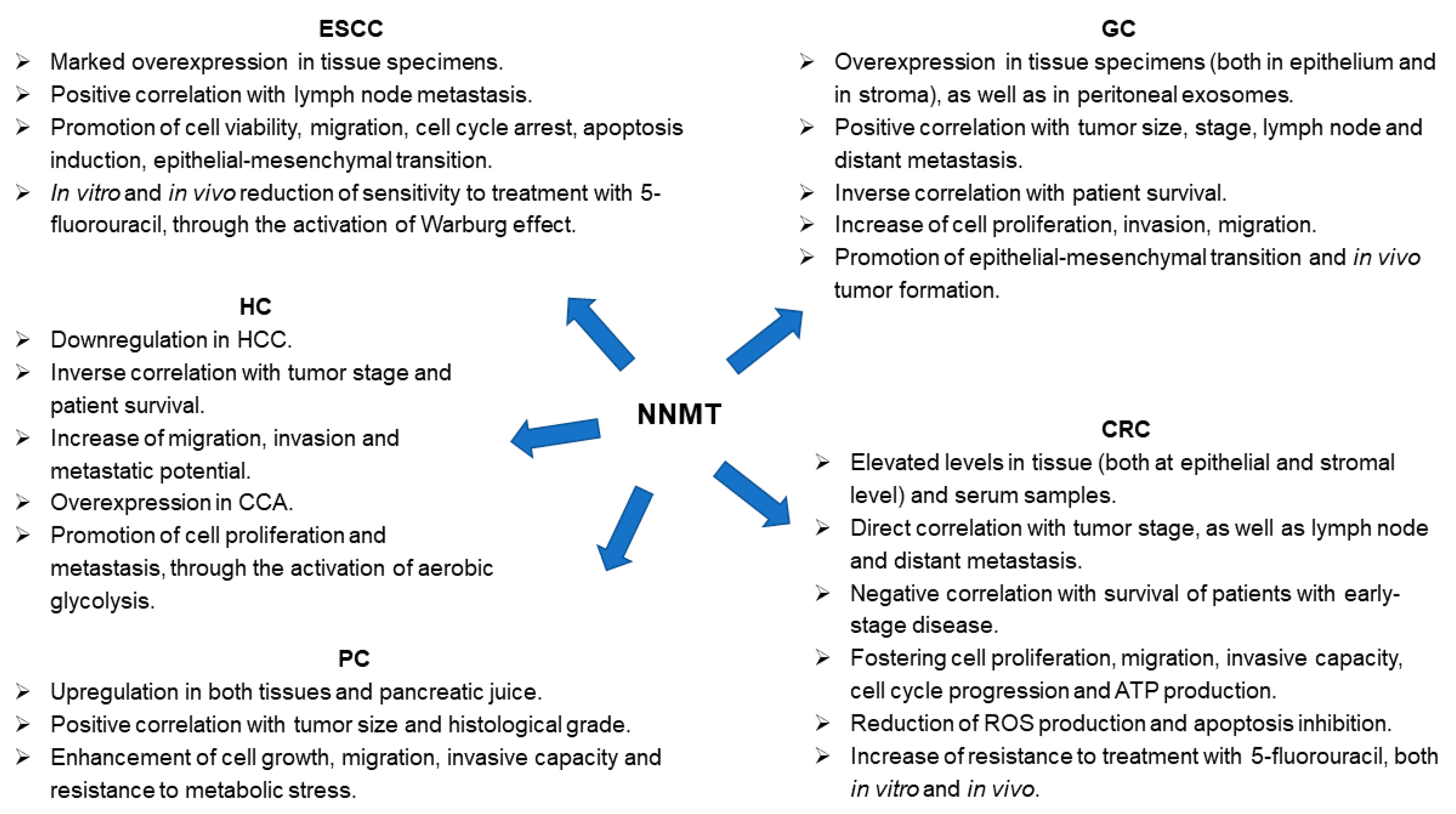
3. NNMT and ESCC
4. NNMT and GC
5. NNMT and CRC
6. NNMT and HC
7. NNMT and PC
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Ferlay, J.; Soerjomataram, I.; Dikshit, R.; Eser, S.; Mathers, C.; Rebelo, M.; Parkin, D.M.; Forman, D.; Bray, F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 2015, 136, E359–E386. [Google Scholar] [CrossRef] [PubMed]
- Rahib, L.; Smith, B.D.; Aizenberg, R.; Rosenzweig, A.B.; Fleshman, J.M.; Matrisian, L.M. Projecting cancer incidence and deaths to 2030: The unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014, 74, 2913–2921. [Google Scholar] [CrossRef] [PubMed]
- Torre, L.A.; Bray, F.; Siegel, R.L.; Ferlay, J.; Lortet-Tieulent, J.; Jemal, A. Global cancer statistics, 2012. CA Cancer J. Clin. 2015, 65, 87–108. [Google Scholar] [CrossRef]
- Coleman, H.G.; Xie, S.H.; Lagergren, J. The Epidemiology of Esophageal Adenocarcinoma. Gastroenterology 2018, 154, 390–405. [Google Scholar] [CrossRef]
- Arnold, M.; Soerjomataram, I.; Ferlay, J.; Forman, D. Global incidence of oesophageal cancer by histological subtype in 2012. Gut 2015, 64, 381–387. [Google Scholar] [CrossRef]
- Malhotra, G.K.; Yanala, U.; Ravipati, A.; Follet, M.; Vijayakumar, M.; Are, C. Global trends in esophageal cancer. J. Surg. Oncol. 2017, 115, 564–579. [Google Scholar] [CrossRef]
- Thrumurthy, S.G.; Chaudry, M.A.; Hochhauser, D.; Mughal, M. The diagnosis and management of gastric cancer. BMJ 2013, 347, f6367. [Google Scholar] [CrossRef]
- Van Cutsem, E.; Sagaert, X.; Topal, B.; Haustermans, K.; Prenen, H. Gastric cancer. Lancet 2016, 388, 2654–2664. [Google Scholar] [CrossRef]
- Ajani, J.A.; Lee, J.; Sano, T.; Janjigian, Y.Y.; Fan, D.; Song, S. Gastric adenocarcinoma. Nat. Rev. Dis. Primers 2017, 3, 17036. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Garborg, K.; Holme, Ø.; Løberg, M.; Kalager, M.; Adami, H.O.; Bretthauer, M. Current status of screening for colorectal cancer. Ann. Oncol. 2013, 24, 1963–1972. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Fedewa, S.A.; Ahnen, D.J.; Meester, R.G.S.; Barzi, A.; Jemal, A. Colorectal cancer statistics, 2017. CA Cancer J. Clin. 2017, 67, 177–193. [Google Scholar] [CrossRef] [PubMed]
- Misiakos, E.P.; Karidis, N.P.; Kouraklis, G. Current treatment for colorectal liver metastases. World J. Gastroenterol. 2011, 17, 4067–4075. [Google Scholar] [CrossRef]
- Giovannucci, E. Modifiable risk factors for colon cancer. Gastroenterol. Clin. N. Am. 2002, 31, 925–943. [Google Scholar] [CrossRef]
- Abraham, C.; Medzhitov, R. Interactions between the host innate immune system and microbes in inflammatory bowel disease. Gastroenterology 2011, 140, 1729–1737. [Google Scholar] [CrossRef]
- M’Koma, A.E. Inflammatory bowel disease: An expanding global health problem. Clin. Med. Insights Gastroenterol. 2013, 6, 33–47. [Google Scholar] [CrossRef]
- Mogoantă, S.S.; Vasile, I.; Totolici, B.; Neamţu, C.; Streba, L.; Busuioc, C.J.; Mateescu, G.O. Colorectal cancer—Clinical and morphological aspects. Rom. J. Morphol. Embryol. 2014, 55, 103–110. [Google Scholar]
- Mancebo, A.; Varela, M.; González-Diéguez, M.L.; Navascués, C.A.; Cadahía, V.; Mesa-Álvarez, A.; Rodrigo, L.; Rodríguez, M. Incidence and risk factors associated with hepatocellular carcinoma surveillance failure. J. Gastroenterol. Hepatol. 2018, 33, 1524–1529. [Google Scholar] [CrossRef]
- El-Serag, H.B.; Rudolph, K.L. Hepatocellular carcinoma: Epidemiology and molecular carcinogenesis. Gastroenterology 2007, 132, 2557–2576. [Google Scholar] [CrossRef]
- Razumilava, N.; Gores, G.J. Cholangiocarcinoma. Lancet 2014, 383, 2168–2179. [Google Scholar] [CrossRef]
- Jiang, W.; Zeng, Z.C.; Tang, Z.Y.; Fan, J.; Sun, H.C.; Zhou, J.; Zeng, M.S.; Zhang, B.H.; Ji, Y.; Chen, Y.X. A prognostic scoring system based on clinical features of intrahepatic cholangiocarcinoma: The Fudan score. Ann. Oncol. 2011, 22, 1644–1652. [Google Scholar] [CrossRef] [PubMed]
- Ribero, D.; Pinna, A.D.; Guglielmi, A.; Ponti, A.; Nuzzo, G.; Giulini, S.M.; Aldrighetti, L.; Calise, F.; Gerunda, G.E.; Tomatis, M.; et al. Italian Intrahepatic Cholangiocarcinoma Study Group. Surgical Approach for Long-term Survival of Patients with Intrahepatic Cholangiocarcinoma: A Multi-institutional Analysis of 434 Patients. Arch. Surg. 2012, 147, 1107–1113. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, J.; Xia, Y.; Gong, R.; Wang, K.; Yan, Z.; Wan, X.; Liu, G.; Wu, D.; Shi, L.; et al. Prognostic nomogram for intrahepatic cholangiocarcinoma after partial hepatectomy. J. Clin. Oncol. 2013, 31, 1188–1195. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef]
- Kamisawa, T.; Wood, L.D.; Itoi, T.; Takaori, K. Pancreatic cancer. Lancet 2016, 388, 73–85. [Google Scholar] [CrossRef]
- Soreide, K. Sweet Predictions Speak Volumes for Early Detection of Pancreatic Cancer. Gastroenterology 2018, 155, 265–268. [Google Scholar] [CrossRef]
- Vennin, C.; Murphy, K.J.; Morton, J.P.; Cox, T.R.; Pajic, M.; Timpson, P. Reshaping the Tumor Stroma for Treatment of Pancreatic Cancer. Gastroenterology 2018, 154, 820–838. [Google Scholar] [CrossRef]
- Pantel, K.; Brakenhoff, R.H. Dissecting the metastatic cascade. Nat. Rev. Cancer 2004, 4, 448–456. [Google Scholar] [CrossRef]
- Rini, J.; Szumlanski, C.; Guerciolini, R.; Weinshilboum, R.M. Human liver nicotinamide N-methyltransferase: Ion-pairing radiochemical assay, biochemical properties and individual variation. Clin. Chim. Acta 1990, 186, 359–374. [Google Scholar] [CrossRef]
- Aksoy, S.; Szumlanski, C.L.; Weinshilboum, R.M. Human liver nicotinamide N-methyltransferase. cDNA cloning, expression, and biochemical characterization. J. Biol. Chem. 1994, 269, 14835–14840. [Google Scholar] [CrossRef]
- Aksoy, S.; Brandriff, B.F.; Ward, A.; Little, P.F.; Weinshilboum, R.M. Human nicotinamide N-methyltransferase gene: Molecular cloning, structural characterization and chromosomal localization. Genomics 1995, 29, 555–561. [Google Scholar] [CrossRef] [PubMed]
- Felsted, R.L.; Chaykin, S. N1-methylnicotinamide oxidation in a number of mammals. J. Biol. Chem. 1967, 242, 1274–1279. [Google Scholar] [CrossRef]
- Okamoto, H.; Ishikawa, A.; Yoshitake, Y.; Kodama, N.; Nishimuta, M.; Fukuwatari, T.; Shibata, K. Diurnal variations in human urinary excretion of nicotinamide catabolites: Effects of stress on the metabolism of nicotinamide. Am. J. Clin. Nutr 2003, 77, 406–410. [Google Scholar] [CrossRef]
- Wang, W.; Yang, C.; Wang, T.; Deng, H. Complex roles of nicotinamide N-methyltransferase in cancer progression. Cell Death Dis. 2022, 13, 267. [Google Scholar] [CrossRef]
- Makarov, M.V.; Trammell, S.A.J.; Migaud, M.E. The chemistry of the vitamin B3 metabolome. Biochem. Soc. Trans. 2019, 47, 131–147. [Google Scholar] [CrossRef]
- Zhang, J. Are poly(ADP-ribosyl)ation by PARP-1 and deacetylation by Sir2 linked? Bioessays 2003, 25, 808–814. [Google Scholar] [CrossRef]
- Riederer, M.; Erwa, W.; Zimmermann, R.; Frank, S.; Zechner, R. Adipose tissue as a source of nicotinamide N-methyltransferase and homocysteine. Atherosclerosis 2009, 204, 412–417. [Google Scholar] [CrossRef]
- Real, A.M.; Hong, S.; Pissios, P. Nicotinamide N-oxidation by CYP2E1 in human liver microsomes. Drug Metab. Dispos. 2013, 41, 550–553. [Google Scholar] [CrossRef]
- Peng, Y.; Sartini, D.; Pozzi, V.; Wilk, D.; Emanuelli, M.; Yee, V.C. Structural basis of substrate recognition in human nicotinamide N-methyltransferase. Biochemistry 2011, 50, 7800–7808. [Google Scholar] [CrossRef]
- Loring, H.S.; Thompson, P.R. Kinetic Mechanism of Nicotinamide N-Methyltransferase. Biochemistry 2018, 57, 5524–5532. [Google Scholar] [CrossRef] [PubMed]
- Neelakantan, H.; Wang, H.Y.; Vance, V.; Hommel, J.D.; McHardy, S.F.; Watowich, S.J. Structure-Activity Relationship for Small Molecule Inhibitors of Nicotinamide N-Methyltransferase. J. Med. Chem. 2017, 60, 5015–5028. [Google Scholar] [CrossRef] [PubMed]
- van Haren, M.J.; Taig, R.; Kuppens, J.; Sastre Toraño, J.; Moret, E.E.; Parsons, R.B.; Sartini, D.; Emanuelli, M.; Martin, N.I. Inhibitors of nicotinamide N-methyltransferase designed to mimic the methylation reaction transition state. Org. Biomol. Chem. 2017, 15, 6656–6667. [Google Scholar] [CrossRef] [PubMed]
- Neelakantan, H.; Vance, V.; Wetzel, M.D.; Wang, H.L.; McHardy, S.F.; Finnerty, C.C.; Hommel, J.D.; Watowich, S.J. Selective and membrane-permeable small molecule inhibitors of nicotinamide N-methyltransferase reverse high fat diet-induced obesity in mice. Biochem. Pharmacol. 2018, 147, 141–152. [Google Scholar] [CrossRef]
- Kannt, A.; Rajagopal, S.; Kadnur, S.V.; Suresh, J.; Bhamidipati, R.K.; Swaminathan, S.; Hallur, M.S.; Kristam, R.; Elvert, R.; Czech, J.; et al. A small molecule inhibitor of Nicotinamide N-methyltransferase for the treatment of metabolic disorders. Sci. Rep. 2018, 8, 3660. [Google Scholar] [CrossRef]
- Babault, N.; Allali-Hassani, A.; Li, F.; Fan, J.; Yue, A.; Ju, K.; Liu, F.; Vedadi, M.; Liu, J.; Jin, J. Discovery of Bisubstrate Inhibitors of Nicotinamide N-Methyltransferase (NNMT). J. Med. Chem. 2018, 61, 1541–1551. [Google Scholar] [CrossRef]
- Ruf, S.; Hallur, M.S.; Anchan, N.K.; Swamy, I.N.; Murugesan, K.R.; Sarkar, S.; Narasimhulu, L.K.; Putta, V.P.R.K.; Shaik, S.; Chandrasekar, D.V.; et al. Novel nicotinamide analog as inhibitor of nicotinamide N-methyltransferase. Bioorg. Med. Chem. Lett. 2018, 28, 922–925. [Google Scholar] [CrossRef]
- Lee, H.Y.; Suciu, R.M.; Horning, B.D.; Vinogradova, E.V.; Ulanovskaya, O.A.; Cravatt, B.F. Covalent inhibitors of nicotinamide N-methyltransferase (NNMT) provide evidence for target engagement challenges in situ. Bioorg. Med. Chem. Lett. 2018, 28, 2682–2687. [Google Scholar] [CrossRef]
- Neelakantan, H.; Brightwell, C.R.; Graber, T.G.; Maroto, R.; Wang, H.L.; McHardy, S.F.; Papaconstantinou, J.; Fry, C.S.; Watowich, S.J. Small molecule nicotinamide N-methyltransferase inhibitor activates senescent muscle stem cells and improves regenerative capacity of aged skeletal muscle. Biochem. Pharmacol. 2019, 163, 481–492. [Google Scholar] [CrossRef]
- Gao, Y.; van Haren, M.J.; Moret, E.E.; Rood, J.J.M.; Sartini, D.; Salvucci, A.; Emanuelli, M.; Craveur, P.; Babault, N.; Jin, J.; et al. Bisubstrate Inhibitors of Nicotinamide N-Methyltransferase (NNMT) with Enhanced Activity. J. Med. Chem. 2019, 62, 6597–6614. [Google Scholar] [CrossRef]
- Policarpo, R.L.; Decultot, L.; May, E.; Kuzmič, P.; Carlson, S.; Huang, D.; Chu, V.; Wright, B.A.; Dhakshinamoorthy, S.; Kannt, A.; et al. High-Affinity Alkynyl Bisubstrate Inhibitors of Nicotinamide N-Methyltransferase (NNMT). J. Med. Chem. 2019, 62, 9837–9873. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Li, L.; Diaz, K.; Iyamu, I.D.; Yadav, R.; Noinaj, N.; Huang, R. Novel Propargyl-Linked Bisubstrate Analogues as Tight-Binding Inhibitors for Nicotinamide N-Methyltransferase. J. Med. Chem. 2019, 62, 10783–10797. [Google Scholar] [CrossRef] [PubMed]
- Kannt, A.; Rajagopal, S.; Hallur, M.S.; Swamy, I.; Kristam, R.; Dhakshinamoorthy, S.; Czech, J.; Zech, G.; Schreuder, H.; Ruf, S. Novel Inhibitors of Nicotinamide-N-Methyltransferase for the Treatment of Metabolic Disorders. Molecules 2021, 26, 991. [Google Scholar] [CrossRef] [PubMed]
- Akar, S.; Duran, T.; Azzawri, A.A.; Koçak, N.; Çelik, Ç.; Yıldırım, H.İ. Small molecule inhibitor of nicotinamide N-methyltransferase shows anti-proliferative activity in HeLa cells. J. Obstet. Gynaecol. 2021, 41, 1240–1245. [Google Scholar] [CrossRef]
- Sabnis, R.W. Novel Pyrimidine-5-carboxamide Compounds as NNMT Inhibitors for Treating Diabetes. ACS Med. Chem. Lett. 2021, 12, 538–539. [Google Scholar] [CrossRef]
- Hayashi, K.; Uehara, S.; Yamamoto, S.; Cary, D.R.; Nishikawa, J.; Ueda, T.; Ozasa, H.; Mihara, K.; Yoshimura, N.; Kawai, T.; et al. Macrocyclic Peptides as a Novel Class of NNMT Inhibitors: A SAR Study Aimed at Inhibitory Activity in the Cell. ACS Med. Chem. Lett. 2021, 12, 1093–1101. [Google Scholar] [CrossRef]
- Iyamu, I.D.; Huang, R. Mechanisms and inhibitors of nicotinamide N-methyltransferase. RSC Med. Chem. 2021, 12, 1254–1261. [Google Scholar] [CrossRef]
- van Haren, M.J.; Zhang, Y.; Thijssen, V.; Buijs, N.; Gao, Y.; Mateuszuk, L.; Fedak, F.A.; Kij, A.; Campagna, R.; Sartini, D.; et al. Macrocyclic peptides as allosteric inhibitors of nicotinamide N-methyltransferase (NNMT). RSC Chem. Biol. 2021, 2, 1546–1555. [Google Scholar] [CrossRef]
- van Haren, M.J.; Gao, Y.; Buijs, N.; Campagna, R.; Sartini, D.; Emanuelli, M.; Mateuszuk, L.; Kij, A.; Chlopicki, S.; Escudé Martinez de Castilla, P.; et al. Esterase-Sensitive Prodrugs of a Potent Bisubstrate Inhibitor of Nicotinamide N-Methyltransferase (NNMT) Display Cellular Activity. Biomolecules 2021, 11, 1357. [Google Scholar] [CrossRef]
- Gao, Y.; van Haren, M.J.; Buijs, N.; Innocenti, P.; Zhang, Y.; Sartini, D.; Campagna, R.; Emanuelli, M.; Parsons, R.B.; Jespers, W.; et al. Potent Inhibition of Nicotinamide N-Methyltransferase by Alkene-Linked Bisubstrate Mimics Bearing Electron Deficient Aromatics. J. Med. Chem. 2021, 64, 12938–12963. [Google Scholar] [CrossRef]
- Gao, Y.; Martin, N.I.; van Haren, M.J. Nicotinamide N-methyl transferase (NNMT): An emerging therapeutic target. Drug Discov. Today 2021, 26, 2699–2706. [Google Scholar] [CrossRef] [PubMed]
- Iyamu, I.D.; Vilseck, J.Z.; Yadav, R.; Noinaj, N.; Huang, R. Exploring Unconventional SAM Analogues To Build Cell-Potent Bisubstrate Inhibitors for Nicotinamide N-Methyltransferase. Angew. Chem. Int. Ed. Engl. 2022, 61, e202114813. [Google Scholar] [CrossRef]
- Kushavah, U.; Panigrahi, L.; Ahmed, S.; Siddiqi, M.I. Ligand-based in silico identification and biological evaluation of potential inhibitors of nicotinamide N-methyltransferase. Mol. Divers. 2022, in press. [Google Scholar] [CrossRef] [PubMed]
- Sartini, D.; Muzzonigro, G.; Milanese, G.; Pierella, F.; Rossi, V.; Emanuelli, M. Identification of nicotinamide N-methyltransferase as a novel tumor marker for renal clear cell carcinoma. J. Urol. 2006, 176, 2248–2254. [Google Scholar] [CrossRef] [PubMed]
- Sartini, D.; Muzzonigro, G.; Milanese, G.; Pozzi, V.; Vici, A.; Morganti, S.; Rossi, V.; Mazzucchelli, R.; Montironi, R.; Emanuelli, M. Upregulation of tissue and urinary nicotinamide N-methyltransferase in bladder cancer: Potential for the development of a urine-based diagnostic test. Cell Biochem. Biophys. 2013, 65, 473–483. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Gui, M.; Zhu, M.; Long, Z.; Huang, L.; Zhou, J.; He, L.; Zhong, K. Nicotinamide N-methyltransferase is overexpressed in prostate cancer and correlates with prolonged progression-free and overall survival times. Oncol. Lett. 2014, 8, 1175–1180. [Google Scholar] [CrossRef]
- Ganzetti, G.; Sartini, D.; Campanati, A.; Rubini, C.; Molinelli, E.; Brisigotti, V.; Cecati, M.; Pozzi, V.; Campagna, R.; Offidani, A.; et al. Nicotinamide N-methyltransferase: Potential involvement in cutaneous malignant melanoma. Melanoma Res. 2018, 28, 82–88. [Google Scholar] [CrossRef]
- Pompei, V.; Salvolini, E.; Rubini, C.; Lucarini, G.; Molinelli, E.; Brisigotti, V.; Pozzi, V.; Sartini, D.; Campanati, A.; Offidani, A.; et al. Nicotinamide N-methyltransferase in nonmelanoma skin cancers. Eur. J. Clin. Investig. 2019, 49, e13175. [Google Scholar] [CrossRef]
- Xu, J.; Moatamed, F.; Caldwell, J.S.; Walker, J.R.; Kraiem, Z.; Taki, K.; Brent, G.A.; Hershman, J.M. Enhanced expression of nicotinamide N-methyltransferase in human papillary thyroid carcinoma cells. J. Clin. Endocrinol. Metab. 2003, 88, 4990–4996. [Google Scholar] [CrossRef][Green Version]
- Akar, S.; Harmankaya, İ.; Uğraş, S.; Çelik, Ç. Nicotinamide N-methyltransferase expression and its association with phospho-Akt, p53 expression, and survival in high-grade endometrial cancer. Turk. J. Med. Sci. 2019, 49, 1547–1554. [Google Scholar] [CrossRef]
- Akar, S.; Harmankaya, İ.; Uğraş, S.; Çelik, Ç. Expression and Clinical Significance of Nicotinamide N-Methyltransferase in Cervical Squamous Cell Carcinoma. Int. J. Gynecol. Pathol. 2020, 39, 289–295. [Google Scholar] [CrossRef] [PubMed]
- Harmankaya, İ.; Akar, S.; Uğraş, S.; Güler, A.H.; Ezveci, H.; Aydoğdu, M.; Çelik, Ç. Nicotinamide N-methyltransferase overexpression may be associated with poor prognosis in ovarian cancer. J. Obstet. Gynaecol. 2020, 41, 248–253. [Google Scholar] [CrossRef] [PubMed]
- Sartini, D.; Santarelli, A.; Rossi, V.; Goteri, G.; Rubini, C.; Ciavarella, D.; Lo Muzio, L.; Emanuelli, M. Nicotinamide N-methyltransferase upregulation inversely correlates with lymph node metastasis in oral squamous cell carcinoma. Mol. Med. 2007, 13, 415–421. [Google Scholar] [CrossRef] [PubMed]
- Emanuelli, M.; Santarelli, A.; Sartini, D.; Ciavarella, D.; Rossi, V.; Pozzi, V.; Rubini, C.; Lo Muzio, L. Nicotinamide N-methyltransferase upregulation correlates with tumour differentiation in oral squamous cell carcinoma. Histol. Histopathol. 2010, 25, 15–20. [Google Scholar] [PubMed]
- Sartini, D.; Pozzi, V.; Renzi, E.; Morganti, S.; Rocchetti, R.; Rubini, C.; Santarelli, A.; Lo Muzio, L.; Emanuelli, M. Analysis of tissue and salivary nicotinamide N-methyltransferase in oral squamous cell carcinoma: Basis for the development of a noninvasive diagnostic test for early-stage disease. Biol. Chem. 2012, 393, 505–511. [Google Scholar] [CrossRef]
- Win, K.T.; Lee, S.W.; Huang, H.Y.; Lin, L.C.; Lin, C.Y.; Hsing, C.H.; Chen, L.T.; Li, C.F. Nicotinamide N-methyltransferase overexpression is associated with Akt phosphorylation and indicates worse prognosis in patients with nasopharyngeal carcinoma. Tumour Biol. 2013, 34, 3923–3931. [Google Scholar] [CrossRef]
- Tomida, M.; Mikami, I.; Takeuchi, S.; Nishimura, H.; Akiyama, H. Serum levels of nicotinamide N-methyltransferase in patients with lung cancer. J. Cancer Res. Clin. Oncol. 2009, 135, 1223–1229. [Google Scholar] [CrossRef]
- Sartini, D.; Morganti, S.; Guidi, E.; Rubini, C.; Zizzi, A.; Giuliante, R.; Pozzi, V.; Emanuelli, M. Nicotinamide N-methyltransferase in non-small cell lung cancer: Promising results for targeted anti-cancer therapy. Cell Biochem. Biophys. 2013, 67, 865–873. [Google Scholar] [CrossRef]
- Markert, J.M.; Fuller, C.M.; Gillespie, G.Y.; Bubien, J.K.; McLean, L.A.; Hong, R.L.; Lee, K.; Gullans, S.R.; Mapstone, T.B.; Benos, D.J. Differential gene expression profiling in human brain tumors. Physiol. Genomics 2001, 5, 21–33. [Google Scholar] [CrossRef]
- D’Andrea, F.P.; Safwat, A.; Kassem, M.; Gautier, L.; Overgaard, J.; Horsman, M.R. Cancer stem cell overexpression of nicotinamide N-methyltransferase enhances cellular radiation resistance. Radiother. Oncol. 2011, 99, 373–378. [Google Scholar] [CrossRef]
- Pozzi, V.; Sartini, D.; Rocchetti, R.; Santarelli, A.; Rubini, C.; Morganti, S.; Giuliante, R.; Calabrese, S.; Di Ruscio, G.; Orlando, F.; et al. Identification and Characterization of Cancer Stem Cells from Head and Neck Squamous Cell Carcinoma Cell Lines. Cell Physiol. Biochem. 2015, 36, 784–798. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.; Kim, L.J.; Wang, X.; Wu, Q.; Sanvoranart, T.; Hubert, C.G.; Prager, B.C.; Wallace, L.C.; Jin, X.; Mack, S.C.; et al. Nicotinamide metabolism regulates glioblastoma stem cell maintenance. JCI Insight 2017, 2, e90019. [Google Scholar] [CrossRef] [PubMed]
- Pozzi, V.; Salvolini, E.; Lucarini, G.; Salvucci, A.; Campagna, R.; Rubini, C.; Sartini, D.; Emanuelli, M. Cancer stem cell enrichment is associated with enhancement of nicotinamide N-methyltransferase expression. IUBMB Life 2020, 72, 1415–1425. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Zhang, L.; Wang, W.; Ma, S.; Liu, H.; Zang, X.; Zhang, Y.; Guan, F. Downregulation of nicotinamide N-methyltransferase inhibits migration and epithelial-mesenchymal transition of esophageal squamous cell carcinoma via Wnt/beta-catenin pathway. Mol. Cell. Biochem. 2019, 460, 93–103. [Google Scholar] [CrossRef]
- Cui, Y.; Yang, D.; Wang, W.; Zhang, L.; Liu, H.; Ma, S.; Guo, W.; Yao, M.; Zhang, K.; Li, W.; et al. Nicotinamide N-methyltransferase decreases 5-fluorouracil sensitivity in human esophageal squamous cell carcinoma through metabolic reprogramming and promoting the Warburg effect. Mol. Carcinog. 2020, 59, 940–954. [Google Scholar] [CrossRef]
- Jang, J.S.; Cho, H.Y.; Lee, Y.J.; Ha, W.S.; Kim, H.W. The differential proteome profile of stomach cancer: Identification of the biomarker candidates. Oncol. Res. 2004, 14, 491–499. [Google Scholar] [CrossRef]
- Wang, X.; Zhi, Q.; Liu, S.; Xue, S.L.; Shen, C.; Li, Y.; Wu, C.; Tang, Z.; Chen, W.; Song, J.L.; et al. Identification of specific biomarkers for gastric adenocarcinoma by ITRAQ proteomic approach. Sci. Rep. 2016, 6, 38871. [Google Scholar] [CrossRef]
- Chen, C.; Wang, X.; Huang, X.; Yong, H.; Shen, J.; Tang, Q.; Zhu, J.; Ni, J.; Feng, Z. Nicotinamide N-methyltransferase: A potential biomarker for worse prognosis in gastric carcinoma. Am. J. Cancer Res. 2016, 6, 649–663. [Google Scholar]
- Wen, F.; Huang, J.; Lu, X.; Huang, W.; Wang, Y.; Bai, Y.; Ruan, S.; Gu, S.; Chen, X.; Shu, P. Identification and prognostic value of metabolism-related genes in gastric cancer. Aging 2020, 12, 17647–17661. [Google Scholar] [CrossRef]
- Wu, M.; Hu, W.; Wang, G.; Yao, Y.; Yu, X.F. Nicotinamide N-Methyltransferase Is a Prognostic Biomarker and Correlated with Immune Infiltrates in Gastric Cancer. Front. Genet. 2020, 11, 580299. [Google Scholar] [CrossRef]
- Zhang, L.; Song, M.; Zhang, F.; Yuan, H.; Chang, W.; Yu, G.; Niu, Y. Accumulation of Nicotinamide N-Methyltransferase (NNMT) in Cancer-associated Fibroblasts: A Potential Prognostic and Predictive Biomarker for Gastric Carcinoma. J. Histochem. Cytochem. 2021, 69, 165–176. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.; Zeng, M.; Pan, H.; Liu, H.; He, Y. Nicotinamide N-methyltransferase promotes epithelial-mesenchymal transition in gastric cancer cells by activating transforming growth factor-β1 expression. Oncol. Lett. 2018, 15, 4592–4598. [Google Scholar] [CrossRef] [PubMed]
- Zhu, A.K.; Shan, Y.Q.; Zhang, J.; Liu, X.C.; Ying, R.C.; Kong, W.C. Exosomal NNMT from peritoneum lavage fluid promotes peritoneal metastasis in gastric cancer. Kaohsiung J. Med. Sci. 2021, 37, 305–313. [Google Scholar] [CrossRef]
- Lim, B.H.; Cho, B.I.; Kim, Y.N.; Kim, J.W.; Park, S.T.; Lee, C.W. Overexpression of nicotinamide N-methyltransferase in gastric cancer tissues and its potential post-translational modification. Exp. Mol. Med. 2006, 38, 455–465. [Google Scholar] [CrossRef][Green Version]
- Roessler, M.; Rollinger, W.; Palme, S.; Hagmann, M.L.; Berndt, P.; Engel, A.M.; Schneidinger, B.; Pfeffer, M.; Andres, H.; Karl, J.; et al. Identification of nicotinamide N-methyltransferase as a novel serum tumor marker for colorectal cancer. Clin. Cancer Res. 2005, 11, 6550–6557. [Google Scholar] [CrossRef]
- Tomida, M.; Ohtake, H.; Yokota, T.; Kobayashi, Y.; Kurosumi, M. Stat3 up-regulates expression of nicotinamide N-methyltransferase in human cancer cells. J. Cancer Res. Clin. Oncol. 2008, 134, 551–559. [Google Scholar] [CrossRef]
- Song, M.; Li, Y.; Miao, M.; Zhang, F.; Yuan, H.; Cao, F.; Chang, W.; Shi, H.; Song, C. High stromal nicotinamide N-methyltransferase (NNMT) indicates poor prognosis in colorectal cancer. Cancer Med. 2020, 9, 2030–2038. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Tong, Q.; Zhang, Y.; Yuan, S.; Gao, Y.; Deng, K.; Wang, Y.; Lu, J.; Xie, X.; Zhang, Z.; et al. Overexpression of Nicotinamide N-methyltransferase mainly covers stroma of colorectal cancer and correlates with unfavorable survival by its product 1-MNA. J. Cancer 2021, 12, 6170–6181. [Google Scholar] [CrossRef]
- Ogawa, M.; Tanaka, A.; Namba, K.; Shia, J.; Wang, J.Y.; Roehrl, M.H.A. Tumor stromal nicotinamide N-methyltransferase overexpression as a prognostic biomarker for poor clinical outcome in early-stage colorectal cancer. Sci. Rep. 2022, 12, 2767. [Google Scholar] [CrossRef]
- Xie, X.; Yu, H.; Wang, Y.; Zhou, Y.; Li, G.; Ruan, Z.; Li, F.; Wang, X.; Liu, H.; Zhang, J. Nicotinamide N-methyltransferase enhances the capacity of tumorigenesis associated with the promotion of cell cycle progression in human colorectal cancer cells. Arch. Biochem. Biophys. 2014, 564, 52–66. [Google Scholar] [CrossRef]
- Xie, X.; Liu, H.; Wang, Y.; Zhou, Y.; Yu, H.; Li, G.; Ruan, Z.; Li, F.; Wang, X.; Zhang, J. Nicotinamide N-methyltransferase enhances resistance to 5-fluorouracil in colorectal cancer cells through inhibition of the ASK1-p38 MAPK pathway. Oncotarget 2016, 7, 45837–45848. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Kong, B.; Tong, Q.; Li, Y.; Chen, L.; Zeng, J.; Yu, H.; Xie, X.; Zhang, J. Vanillin downregulates NNMT and attenuates NNMT-related resistance to 5-fluorouracil via ROS-induced cell apoptosis in colorectal cancer cells. Oncol. Rep. 2021, 45, 110. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Fang, S.; Shao, X.; Li, Y.; Tong, Q.; Kong, B.; Chen, L.; Wang, Y.; Yang, J.; Yu, H.; et al. Curcumin Reverses NNMT-Induced 5-Fluorouracil Resistance via Increasing ROS and Cell Cycle Arrest in Colorectal Cancer Cells. Biomolecules 2021, 11, 1295. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Hong, S.J.; Lim, E.K.; Yu, Y.S.; Kim, S.W.; Roh, J.H.; Do, I.G.; Joh, J.W.; Kim, D.S. Expression of nicotinamide N-methyltransferase in hepatocellular carcinoma is associated with poor prognosis. J. Exp. Clin. Cancer Res. 2009, 28, 20. [Google Scholar] [CrossRef]
- Li, J.; You, S.; Zhang, S.; Hu, Q.; Wang, F.; Chi, X.; Zhao, W.; Xie, C.; Zhang, C.; Yu, Y.; et al. Elevated N-methyltransferase expression induced by hepatic stellate cells contributes to the metastasis of hepatocellular carcinoma via regulation of the CD44v3 isoform. Mol. Oncol. 2019, 13, 1993–2009. [Google Scholar] [CrossRef]
- Lu, S.; Ke, S.; Wang, C.; Xu, Y.; Li, Z.; Song, K.; Bai, M.; Zhou, M.; Yu, H.; Yin, B.; et al. NNMT promotes the progression of intrahepatic cholangiocarcinoma by regulating aerobic glycolysis via the EGFR-STAT3 axis. Oncogenesis 2022, 11, 39. [Google Scholar] [CrossRef]
- Rogers, C.D.; Fukushima, N.; Sato, N.; Shi, C.; Prasad, N.; Hustinx, S.R.; Matsubayashi, H.; Canto, M.; Eshleman, J.R.; Hruban, R.H.; et al. Differentiating pancreatic lesions by microarray and QPCR analysis of pancreatic juice RNAs. Cancer Biol. Ther. 2006, 5, 1383–1389. [Google Scholar] [CrossRef]
- Xu, Y.; Liu, P.; Zheng, D.H.; Wu, N.; Zhu, L.; Xing, C.; Zhu, J. Expression profile and prognostic value of NNMT in patients with pancreatic cancer. Oncotarget 2016, 7, 19975–19981. [Google Scholar] [CrossRef]
- Yu, T.; Wang, Y.T.; Chen, P.; Li, Y.H.; Chen, Y.X.; Zeng, H.; Yu, A.M.; Huang, M.; Bi, H.C. Effects of nicotinamide N-methyltransferase on PANC-1 cells proliferation, metastatic potential and survival under metabolic stress. Cell Physiol. Biochem. 2015, 35, 710–721. [Google Scholar] [CrossRef]
- Parsons, R.B.; Facey, P.D. Nicotinamide N-Methyltransferase: An Emerging Protagonist in Cancer Macro(r)evolution. Biomolecules 2021, 11, 1418. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pozzi, V.; Campagna, R.; Sartini, D.; Emanuelli, M. Nicotinamide N-Methyltransferase as Promising Tool for Management of Gastrointestinal Neoplasms. Biomolecules 2022, 12, 1173. https://doi.org/10.3390/biom12091173
Pozzi V, Campagna R, Sartini D, Emanuelli M. Nicotinamide N-Methyltransferase as Promising Tool for Management of Gastrointestinal Neoplasms. Biomolecules. 2022; 12(9):1173. https://doi.org/10.3390/biom12091173
Chicago/Turabian StylePozzi, Valentina, Roberto Campagna, Davide Sartini, and Monica Emanuelli. 2022. "Nicotinamide N-Methyltransferase as Promising Tool for Management of Gastrointestinal Neoplasms" Biomolecules 12, no. 9: 1173. https://doi.org/10.3390/biom12091173
APA StylePozzi, V., Campagna, R., Sartini, D., & Emanuelli, M. (2022). Nicotinamide N-Methyltransferase as Promising Tool for Management of Gastrointestinal Neoplasms. Biomolecules, 12(9), 1173. https://doi.org/10.3390/biom12091173








