Characterization of a Virally Encoded Flavodoxin That Can Drive Bacterial Cytochrome P450 Monooxygenase Activity
Abstract
1. Importance
2. Introduction
3. Materials and Methods
3.1. Bioinformatic and Phylogenetic Analysis
3.2. Cloning, Heterologous Expression and Purification of 0305φ8-36 Flavodoxin
3.3. Reconstitution of Streptomyces coelicolor CYP170A1 Activity with 0305φ8-36 Flavodoxin
3.4. General Methods
4. Results
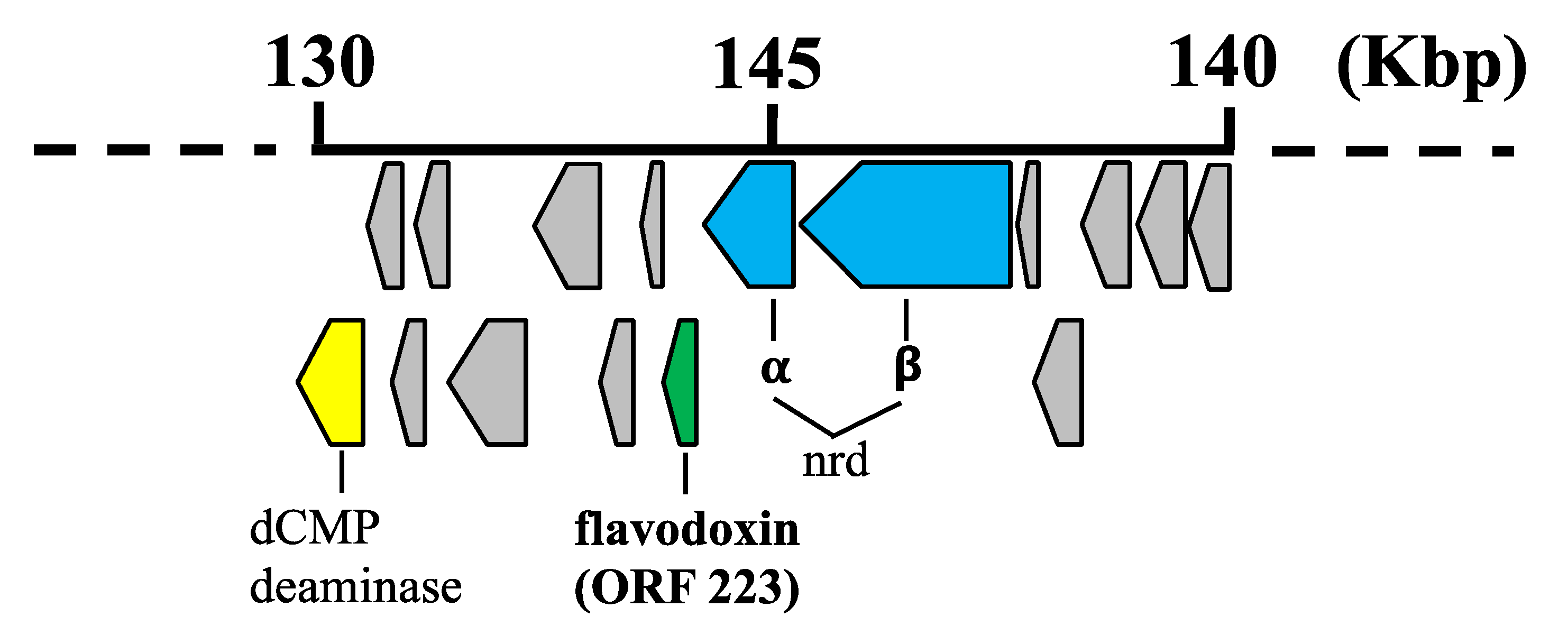
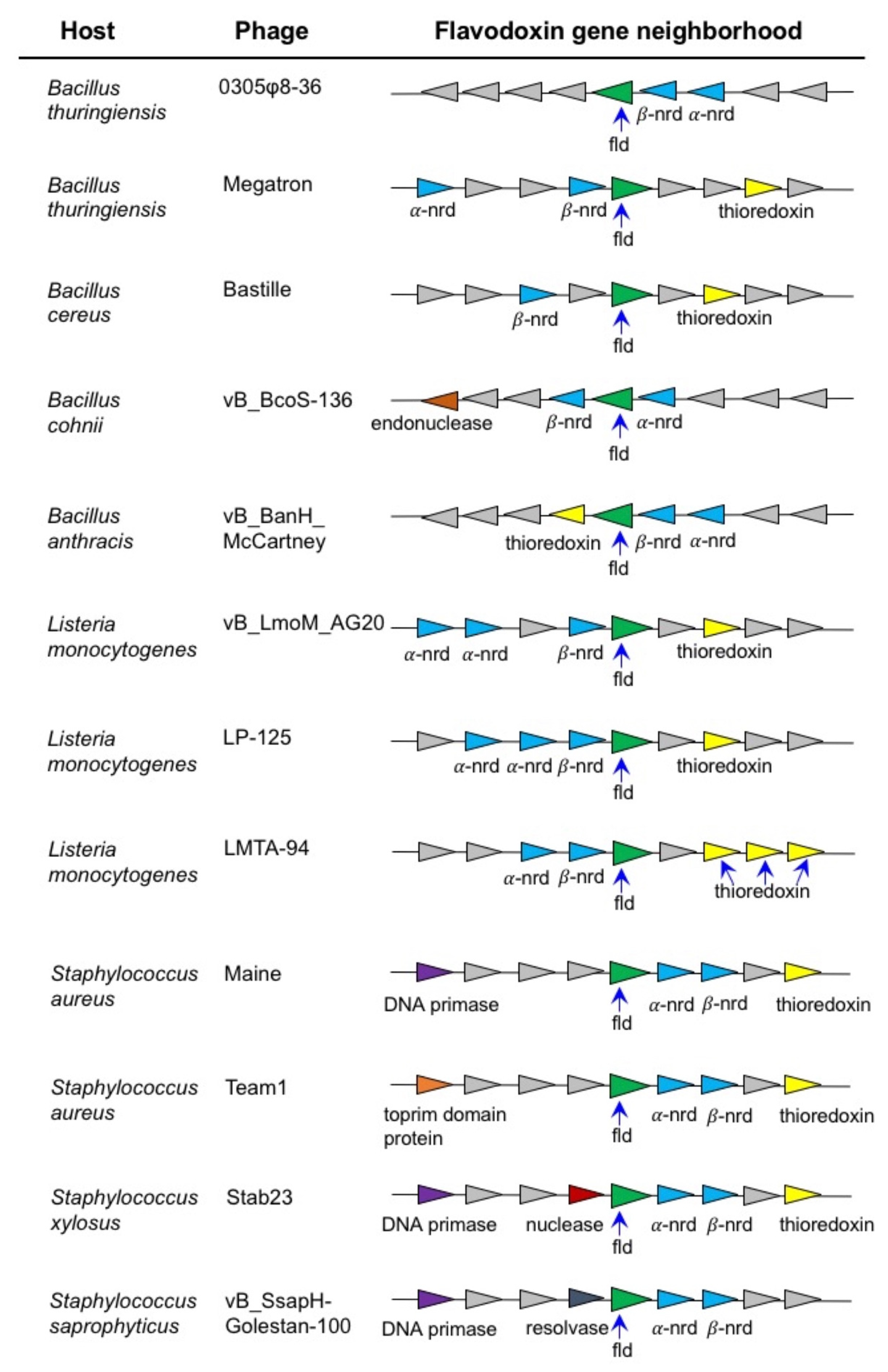
4.1. Bioinformatic and Phylogenetic Analysis of 0305φ8-36 ORF223
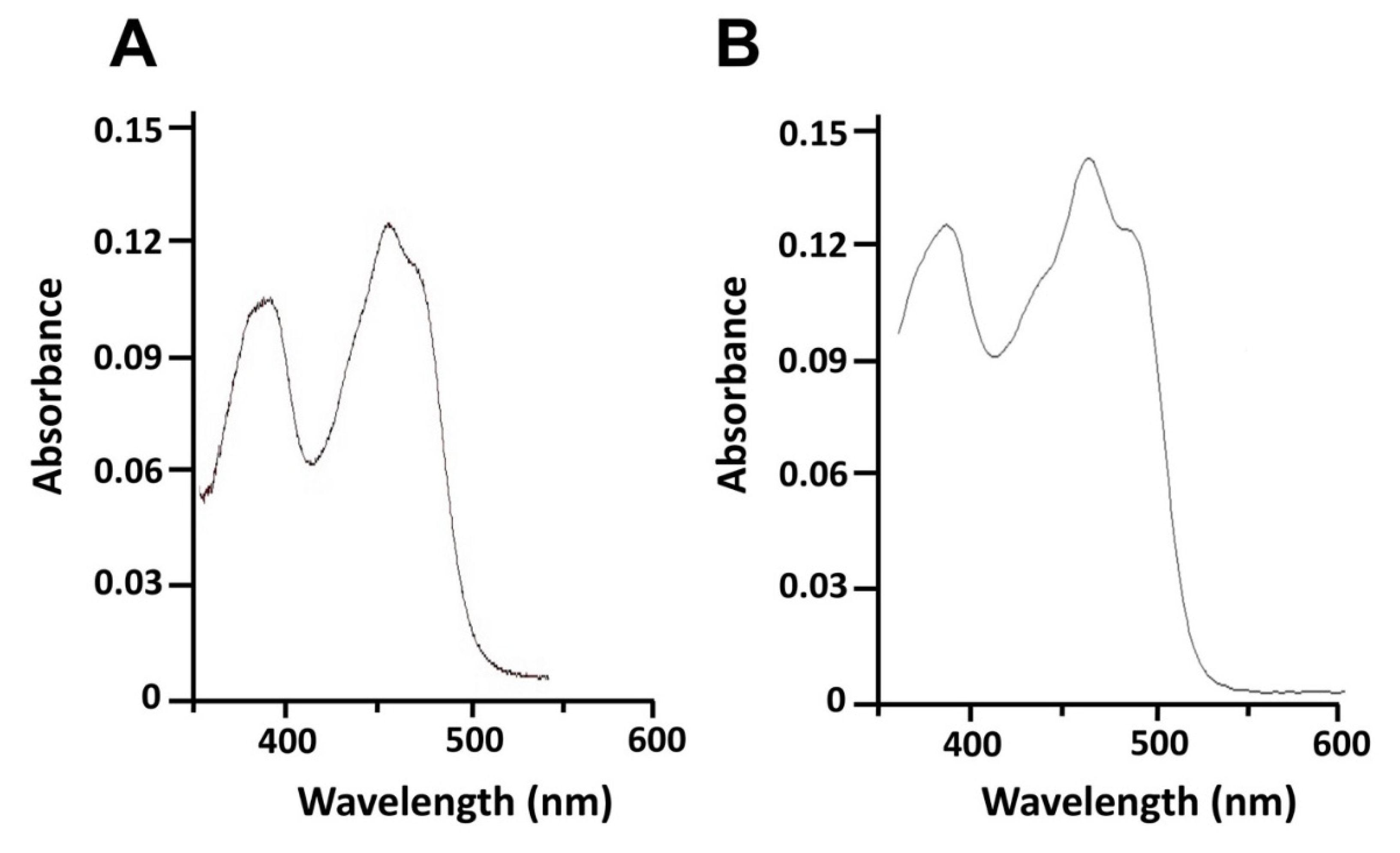
4.2. Expression, Purification and Spectral Analysis of Recombinant 0305φ8-36 ORF223
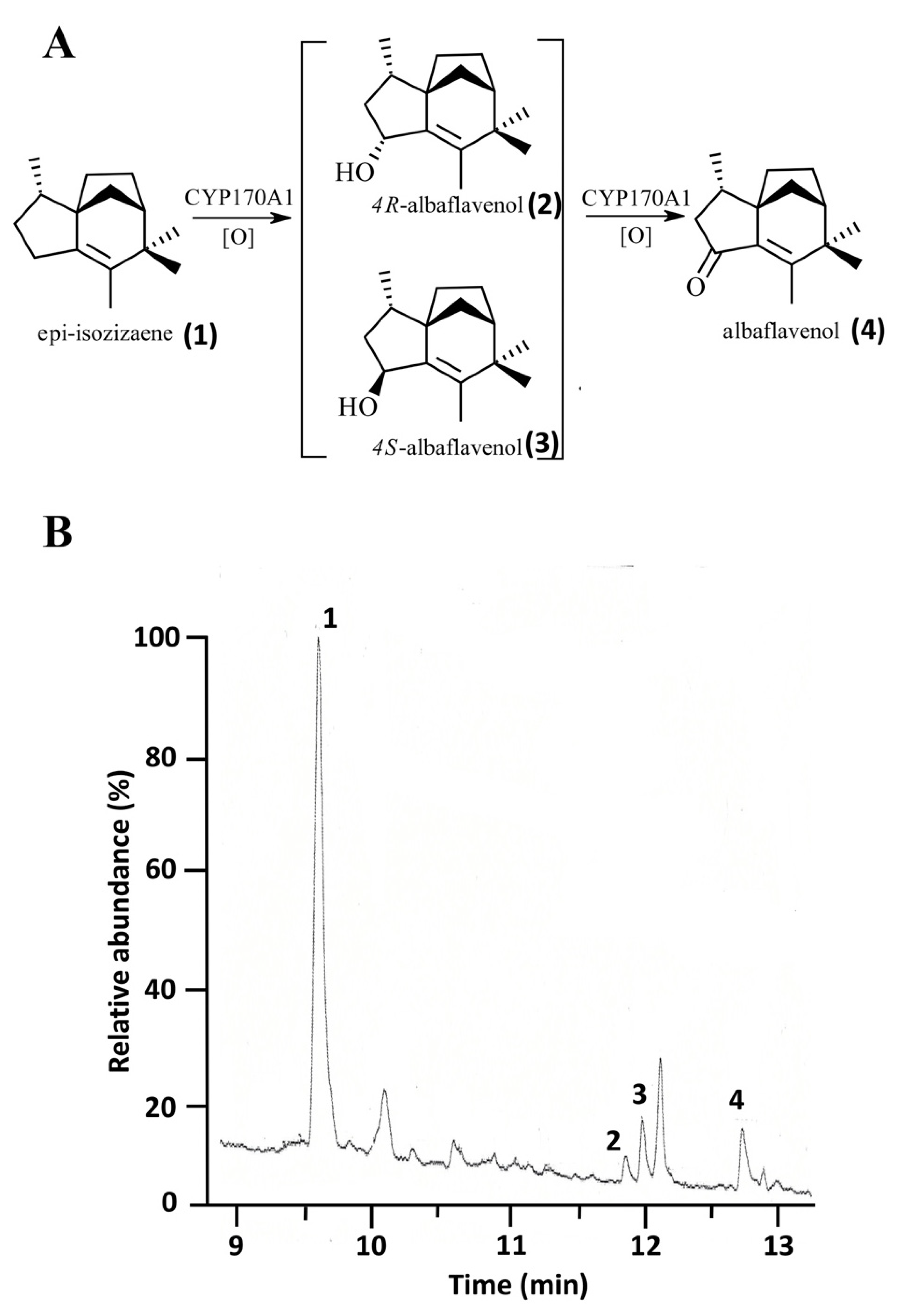
4.3. Characterization of the Catalytic Activity of 0305φ8-36 Flavodoxin in P450-Mediated Monooxygenation
5. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sancho, J. Flavodoxins: Sequence, folding, binding, function and beyond. Cell Mol. Life Sci. 2006, 63, 855–864. [Google Scholar] [CrossRef] [PubMed]
- Smillie, R.M. Isolation of two proteins with chloroplast ferredoxin activity from a blue-green alga. Biochem. Biophys. Res. Commun. 1965, 20, 621–629. [Google Scholar] [CrossRef]
- Knight, E.; D’Eustachio, A.J.; Hardy, R.W. Flavodoxin: A flavoprotein with ferredoxin activity from Clostrium pasteurianum. Biochim. Biophys. Acta 1966, 113, 626–628. [Google Scholar] [CrossRef]
- Anderson, R.F. Energetics of the one-electron reduction steps of riboflavin, FMN and FAD to their fully reduced forms. Biochim. Biophys. Acta 1983, 722, 158–162. [Google Scholar] [CrossRef]
- Mayhew, S.G. The effects of pH and semiquinone formation on the oxidation-reduction potentials of flavin mononucleotide. A reappraisal. Eur. J. Biochem. 1999, 265, 698–702. [Google Scholar] [CrossRef] [PubMed]
- Gaudu, P.; Weiss, B. Flavodoxin mutants of Escherichia coli K-12. J. Bacteriol. 2000, 182, 1788–1793. [Google Scholar] [CrossRef] [PubMed]
- Freigang, J.; Diederichs, K.; Schafer, K.P.; Welte, W.; Paul, R. Crystal structure of oxidized flavodoxin, an essential protein in Helicobacter pylori. Protein Sci. 2002, 11, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Modi, S.; Smith, G.; Paine, M.; McDonagh, P.D.; Wolf, C.R.; Tew, D.; Lian, L.Y.; Roberts, G.C.; Driessen, H.P. Crystal structure of the FMN-binding domain of human cytochrome P450 reductase at 1.93 A resolution. Protein Sci. 1999, 8, 298–306. [Google Scholar] [CrossRef]
- Lopez-Llano, J.; Maldonado, S.; Jain, S.; Lostao, A.; Godoy-Ruiz, R.; Sanchez-Ruiz, J.M.; Cortijo, M.; Fernandez-Recio, J.; Sancho, J. The long and short flavodoxins: II. The role of the differentiating loop in apoflavodoxin stability and folding mechanism. J. Biol. Chem. 2004, 279, 47184–47191. [Google Scholar] [CrossRef] [PubMed]
- Fukuyama, K.; Matsubara, H.; Rogers, L.J. Crystal structure of oxidized flavodoxin from a red alga Chondrus crispus refined at 1.8 A resolution. Description of the flavin mononucleotide binding site. J. Mol. Biol. 1992, 225, 775–789. [Google Scholar] [CrossRef]
- Peleato, M.L.; Ayora, S.; Inda, L.A.; Gomez-Moreno, C. Isolation and characterization of two different flavodoxins from the eukaryote Chlorella fusca. Biochem. J. 1994, 302, 807–811. [Google Scholar] [CrossRef] [PubMed]
- Dwivedi, B.; Xue, B.; Lundin, D.; Edwards, R.A.; Breitbart, M. A bioinformatic analysis of ribonucleotide reductase genes in phage genomes and metagenomes. BMC Evol. Biol. 2013, 13, 33. [Google Scholar] [CrossRef] [PubMed]
- Harrison, A.O.; Moore, R.M.; Polson, S.W.; Wommack, K.E. Reannotation of the Ribonucleotide Reductase in a Cyanophage Reveals Life History Strategies Within the Virioplankton. Front. Microbiol. 2019, 10, 134. [Google Scholar] [CrossRef]
- Lamb, D.C.; Follmer, A.H.; Goldstone, J.V.; Nelson, D.R.; Warrilow, A.G.; Price, C.L.; True, M.Y.; Kelly, S.L.; Poulos, T.L.; Stegeman, J.J. On the occurrence of cytochrome P450 in viruses. Proc. Natl. Acad. Sci. USA 2019, 116, 12343–12352. [Google Scholar] [CrossRef] [PubMed]
- Hardies, S.C.; Thomas, J.A.; Serwer, P. Comparative genomics of Bacillus thuringiensis phage 0305phi8-36: Defining patterns of descent in a novel ancient phage lineage. Virol. J. 2007, 4, 97. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.A.; Hardies, S.C.; Rolando, M.; Hayes, S.J.; Lieman, K.; Carroll, C.A.; Weintraub, S.T.; Serwer, P. Complete genomic sequence and mass spectrometric analysis of highly diverse, atypical Bacillus thuringiensis phage 0305phi8-36. Virology 2007, 368, 405–421. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Soding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef]
- Chang, J.M.; Di Tommaso, P.; Notredame, C. TCS: A new multiple sequence alignment reliability measure to estimate alignment accuracy and improve phylogenetic tree reconstruction. Mol. Biol. Evol. 2014, 31, 1625–1637. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl. Acids Symp. Series 1999, 41, 95–98. [Google Scholar]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, 320–324. [Google Scholar] [CrossRef]
- Kozlov, A.; Darriba, D.; Flouri, T.; Morel, B.; Stamatakis, A. RAxML-NG: A fast, scalable, and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 2019, 35, 4453–4455. [Google Scholar] [CrossRef] [PubMed]
- Lemoine, F.; Domelevo Entfellner, J.B.; Wilkinson, E.; Correia, D.; Felipe, M.D.; De Oliveira, T.; Gascuel, O. Renewing Felsenstein’s phylogenetic bootstrap in the era of big data. Nature 2018, 556, 452–456. [Google Scholar] [CrossRef] [PubMed]
- Lamb, D.C.; Lei, L.; Warrilow, A.G.; Lepesheva, G.I.; Mullins, J.G.; Waterman, M.R.; Kelly, S.L. The first virally encoded cytochrome P450. J. Virol. 2009, 83, 8266–8269. [Google Scholar] [CrossRef] [PubMed]
- Lamb, D.C.; Lei, L.; Zhao, B.; Yuan, H.; Jackson, C.J.; Warrilow, A.G.; Skaug, T.; Dyson, P.J.; Dawson, E.S.; Kelly, S.L.; et al. Streptomyces coelicolor A3(2) CYP102 protein, a novel fatty acid hydroxylase encoded as a heme domain without an N-terminal redox partner. Appl. Environ. Microbiol. 2010, 76, 1975–1980. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jenkins, C.M.; Waterman, M.R. Flavodoxin and NADPH-flavodoxin reductase from Escherichia coli support bovine cytochrome P450c17 hydroxylase activities. J. Biol. Chem. 1994, 269, 27401–27408. [Google Scholar] [CrossRef]
- Zhao, B.; Lin, X.; Lei, L.; Lamb, D.C.; Kelly, S.L.; Waterman, M.R.; Cane, D.E. Biosynthesis of the sesquiterpene antibiotic albaflavenone in Streptomyces coelicolor A3(2). J. Biol. Chem. 2008, 283, 8183–8189. [Google Scholar] [CrossRef]
- Omura, T.; Sato, R. The Carbon Monoxide-Binding Pigment of Liver Microsomes. I. Evidence for Its Hemoprotein Nature. J. Biol. Chem. 1964, 239, 2370–2378. [Google Scholar] [CrossRef]
- Jenkins, C.M.; Waterman, M.R. NADPH-flavodoxin reductase and flavodoxin from Escherichia coli: Characteristics as a soluble microsomal P450 reductase. Biochemistry 1998, 37, 6106–6113. [Google Scholar] [CrossRef]
- Zhao, B.; Guengerich, F.P.; Bellamine, A.; Lamb, D.C.; Izumikawa, M.; Lei, L.; Podust, L.M.; Sundaramoorthy, M.; Kalaitzis, J.A.; Reddy, L.M.; et al. Binding of two flaviolin substrate molecules, oxidative coupling, and crystal structure of Streptomyces coelicolor A3(2) cytochrome P450 158A2. J. Biol. Chem. 2005, 280, 11599–11607. [Google Scholar] [CrossRef]
- Bravo, A.; Likitvivatanavong, S.; Gill, S.S.; Soberon, M. Bacillus thuringiensis: A story of a successful bioinsecticide. Insect Biochem. Mol. 2011, 41, 423–431. [Google Scholar] [CrossRef]
- Sauder, A.B.; Quinn, M.R.; Brouillette, A.; Caruso, S.; Cresawn, S.; Erill, I.; Lewis, L.; Loesser-Casey, K.; Pate, M.; Scott, C.; et al. Genomic characterization and comparison of seven Myoviridae bacteriophage infecting Bacillus thuringiensis. Virology 2016, 489, 243–251. [Google Scholar] [CrossRef] [PubMed]
- Erill, I.; Caruso, S.M.; 2015 UMBC Phage Hunters. Genome sequence of Bacillus cereus group phage SalinJah. Genome Announc. 2016, 4, e00953-16. [Google Scholar] [CrossRef] [PubMed]
- Erill, I.; Caruso, S.M. Genome sequences of two Bacillus cereus group Bacteriophages, Eyuki and AvesoBmore. Genome Announc. 2015, 3, e01199-15. [Google Scholar] [CrossRef] [PubMed]
- Ting, J.H.; Smyth, T.B.; Chamakura, K.R.; Kuty Everett, G.F. Complete genome of Bacillus thuringiensis myophage BigBertha. Genome Announc. 2013, 1, e00853-13. [Google Scholar] [CrossRef]
- Maroun, J.W.; Whitcher, K.J.; Chamakura, K.R.; Kuty Everett, G.F. Complete genome of Bacillus thuringiensis myophage Spock. Genome Announc. 2013, 1, e00863-13. [Google Scholar] [CrossRef]
- Breslin, E.F.; Cornell, J.; Schuhmacher, Z.; Himelright, M.; Andos, A.; Childs, A.; Clem, A.; Gerber, M.; Gordillo, A.; Harb, L.; et al. Complete genome sequence of Bacillus phage Belinda from Grand Cayman Island. Genome Announc. 2016, 4, e00571-16. [Google Scholar] [CrossRef]
- Denes, T.; Vongkamjan, K.; Ackermann, H.W.; Moreno Switt, A.I.; Wiedmann, M.; den Bakker, H.C. Comparative genomic and morphological analyses of Listeria phages isolated from farm environments. Appl. Environ. Microbiol. 2014, 80, 4616–4625. [Google Scholar] [CrossRef]
- Peters, T.L.; Song, Y.; Bryan, D.W.; Hudson, L.K.; Denes, T.G. Mutant and recombinant phages selected from in vitro coevolution conditions overcome phage-resistant Listeria monocytogenes. Appl. Environ. Microbiol. 2020, 86, e02138-20. [Google Scholar] [CrossRef]
- Duperier, J.; Bulpitt, M.; Bispo, F.; Greguske, E. Genome annotations of two Bacillus phages, Tomato and Baseball Field. Microbiol. Resour. Announc. 2021, 10, e01196-20. [Google Scholar] [CrossRef]
- Peters, T.L.; Hudson, L.K.; Song, Y.; Denes, T.G. Complete genome sequences of two Listeria phages of the genus Pecentumvirus. Microbiol. Resour. Announc. 2019, 8, e01229-19. [Google Scholar] [CrossRef]
- Kostyk, N.; Chigbu, O.; Cochran, E.; Davis, J.; Essig, J.; Do, L.; Farooque, N.; Gbadamosi, Z.; Gnanodayan, A.; Hale, A.; et al. Complete genome sequences of Bacillus cereus group phages AaronPhadgers, ALPS, Beyonphe, Bubs, KamFam, OmnioDeoPrimus, Phireball, PPIsBest, YungSlug, and Zainny. Microbiol. Resour. Announc. 2021, 10, e0030021. [Google Scholar] [CrossRef] [PubMed]
- Nordmann, B.; Schilling, T.; Hoppert, M.; Hertel, R. Complete genome sequence of the virus isolate vB_BthM-Goe5 infecting Bacillus thuringiensis. Arch Virol. 2019, 164, 1485–1488. [Google Scholar] [CrossRef] [PubMed]
- Foltz, S.; Johnson, A.A. Complete genome sequences of nine Bacillus cereus group phages. Genome Announc. 2016, 4, e00473-16. [Google Scholar] [CrossRef] [PubMed]
- Flounlacker, K.; Miller, R.; Marquez, D.; Johnson, A.; the 2015–2016 VCU Phage Hunters. complete genome sequences of Bacillus phages DirtyBetty and Kida. Genome Announc. 2017, 5, e01385-16. [Google Scholar] [CrossRef]
- Erill, I.; Caruso, S.M. Complete genome sequence of Bacillus cereus group phage TsarBomba. Genome Announc. 2015, 3, e01178-15. [Google Scholar] [CrossRef]
- Lee, M.; Puglisi, K.M.; UMBC STEM-BUILD Cohort 1; Erill, I.; Caruso, S.M. Complete genome sequences of HonestAbe, Anthony, and Taffo16, three cluster C Bacillus cereus group bacteriophages. Genome Announc. 2018, 6, e00493-18. [Google Scholar] [CrossRef]
- Greguske, E.; Nadeau, A.; Fitzmeyer, E.; Fucikova, K. Genome sequence of Bacillus phage Saddex. Microbiol. Resour. Announc. 2018, 7, e01044-18. [Google Scholar] [CrossRef]
- Kent, B.; Raymond, T.; Mosier, P.D.; Johnson, A.A.; the 2016–2017 VCU Phage Hunters. Complete genome sequences of Bacillus phages Janet and OTooleKemple52. Genome Announc. 2018, 6, e00083-18. [Google Scholar] [CrossRef]
- Moreland, R.; Korn, A.; Newkirk, H.; Liu, M.; Gill, J.J.; Cahill, J.; Ramsey, J. Complete genome sequence of Staphylococcus aureus myophage Maine. Microbiol. Resour. Announc. 2019, 8, e01050-19. [Google Scholar] [CrossRef]
- Korn, A.M.; Hillhouse, A.E.; Sun, L.; Gill, J.J. Comparative genomics of three novel jumbo bacteriophages infecting Staphylococcus aureus. J. Virol. 2021, 95, e0239120. [Google Scholar] [CrossRef]
- Gutiérrez, D.; Vandenheuvel, D.; Martínez, B.; Rodríguez, A.; Lavigne, R.; García, P. Two phages, phiIPLA-RODI and phiIPLA-C1C, lyse mono- and dual-species Staphylococcal biofilms. Appl. Environ. Microbiol. 2015, 81, 3336–3348. [Google Scholar] [CrossRef] [PubMed]
- El Haddad, L.; Ben Abdallah, N.; Plante, P.L.; Dumaresq, J.; Katsarava, R.; Labrie, S.; Corbeil, J.; St-Gelais, D.; Moineau, S. Improving the safety of Staphylococcus aureus polyvalent phages by their production on a Staphylococcus xylosus strain. PLoS ONE 2014, 9, e102600. [Google Scholar]
- Alves, D.R.; Gaudion, A.; Bean, J.E.; Perez Esteban, P.; Arnot, T.C.; Harper, D.R.; Kot, W.; Hansen, L.H.; Enright, M.C.; Jenkins, A.T. Combined use of bacteriophage K and a novel bacteriophage to reduce Staphylococcus aureus biofilm formation. Appl. Environ. Microbiol. 2014, 80, 6694–6703. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Liu, X.; Lu, R.; Li, Y.; Song, J.; Lei, L.; Sun, C.; Feng, X.; Du, C.; Yu, H.; et al. Complete genome sequence of Staphylococcus aureus bacteriophage GH15. J. Virol. 2012, 86, 8914–8915. [Google Scholar] [CrossRef]
- Crane, A.; Abaidoo, J.; Beltran, G.; Fry, D.; Furey, C.; Green, N.; Johal, R.; La Rosa, B.; Jimenez, C.L.; Luong, L.; et al. The complete genome sequence of the Staphylococcus bacteriophage Metroid. G3 2020, 10, 2975–2979. [Google Scholar] [CrossRef]
- Jun, J.W.; Giri, S.S.; Kim, H.J.; Chi, C.; Yun, S.; Kim, S.G.; Kim, S.W.; Kang, J.W.; Park, S.C. Complete genome sequence of the novel bacteriophage pSco-10 Infecting Staphylococcus cohnii. Genome Announc. 2017, 5, e01032-17. [Google Scholar] [CrossRef]
- Sáez Moreno, D.; Visram, Z.; Mutti, M.; Restrepo-Córdoba, M.; Hartmann, S.; Kremers, A.I.; Tišáková, L.; Schertler, S.; Wittmann, J.; Kalali, B.; et al. ε2-phages are naturally bred and have a vastly improved host range in Staphylococcus aureus over wild type phages. Pharmaceuticals 2021, 14, 325. [Google Scholar] [CrossRef]
- Oduor, J.M.O.; Kadija, E.; Nyachieo, A.; Mureithi, M.W.; Skurnik, M. Bioprospecting Staphylococcus phages with therapeutic and bio-control potential. Viruses 2020, 12, 133. [Google Scholar] [CrossRef]
- Abatángelo, V.; Peressutti Bacci, N.; Boncompain, C.A.; Amadio, A.F.; Carrasco, S.; Suárez, C.A.; Morbidoni, H.R. Broad-range lytic bacteriophages that kill Staphylococcus aureus local field strains. PLoS ONE 2017, 12, e0181671. [Google Scholar]
- Hoover, D.M.; Jarrett, J.T.; Sands, R.H.; Dunham, W.R.; Ludwig, M.L.; Matthews, R.G. Interaction of Escherichia coli cobalamin-dependent methionine synthase and its physiological partner flavodoxin: Binding of flavodoxin leads to axial ligand dissociation from the cobalamin cofactor. Biochemistry 1997, 36, 127–138. [Google Scholar] [CrossRef]
- Green, A.J.; Rivers, S.L.; Cheeseman, M.; Reid, G.A.; Quaroni, L.G.; Macdonald, I.D.; Chapman, S.K.; Munro, A.W. Expression, purification and characterization of cytochrome P450 Biol: A novel P450 involved in biotin synthesis in Bacillus subtilis. J. Biol. Inorg. Chem. 2001, 6, 523–533. [Google Scholar] [CrossRef]
- Gangeswaran, R.; Eady, R.R. Flavodoxin 1 of Azotobacter vinelandii: Characterization and role in electron donation to purified assimilatory nitrate reductase. Biochem. J. 1996, 317, 103–108. [Google Scholar] [CrossRef]
- Prakash, D.; Iyer, P.R.; Suharti, S.; Walters, K.A.; Santiago-Martinez, M.G.; Golbeck, J.H.; Murakami, K.S.; Ferry, J.G. Structure and function of an unusual flavodoxin from the domain Archaea. Proc. Natl. Acad. Sci. USA 2019, 116, 25917–25922. [Google Scholar] [CrossRef]
- Jackson, R.G.; Rylott, E.L.; Fournier, D.; Hawari, J.; Bruce, N.C. Exploring the biochemical properties and remediation applications of the unusual explosive-degrading P450 system XplA/B. Proc. Natl. Acad. Sci. USA 2007, 104, 16822–16827. [Google Scholar] [CrossRef]
- Liu, L.; Schmid, R.D.; Urlacher, V.B. Cloning, expression, and characterization of a self-sufficient cytochrome P450 monooxygenase from Rhodococcus ruber DSM 44319. Appl. Microbiol. Biotechnol. 2006, 72, 876–882. [Google Scholar] [CrossRef]
- Hawkes, D.B.; Adams, G.W.; Burlingame, A.L.; Ortiz de Montellano, P.R.; De Voss, J.J. Cytochrome P450(cin) (CYP176A), isolation, expression, and characterization. J. Biol. Chem. 2002, 277, 27725–27732. [Google Scholar] [CrossRef]
- Jordan, A.; Reichard, P. Ribonucleotide reductases. Annu. Rev. Biochem. 1998, 67, 71–98. [Google Scholar] [CrossRef]
- Sheppard, A.E.; Poehlein, A.; Rosenstiel, P.; Liesegang, H.; Schulenburg, H. Complete Genome Sequence of Bacillus thuringiensis Strain 407 Cry. Genome Announc. 2013, 1, e00158-12. [Google Scholar] [CrossRef]
- Mthethwa, B.C.; Chen, W.; Ngwenya, M.L.; Kappo, A.P.; Syed, P.R.; Karpoormath, R.; Yu, J.H.; Nelson, D.R.; Syed, K. Comparative Analyses of Cytochrome P450s and Those Associated with Secondary Metabolism in Bacillus Species. Int. J. Mol. Sci. 2018, 19, 3623. [Google Scholar] [CrossRef]
- Szaleniec, M.; Wojtkiewicz, A.M.; Bernhardt, R.; Borowski, T.; Donova, M. Bacterial steroid hydroxylases: Enzyme classes, their functions and comparison of their catalytic mechanisms. Appl. Microbiol. Biotechnol. 2018, 102, 8153–8171. [Google Scholar] [CrossRef]
- Grose, J.H.; Jensen, G.L.; Burnett, S.H.; Breakwell, D.P. Genomic comparison of 93 Bacillus phages reveals 12 clusters, 14 singletons and remarkable diversity. BMC Genom. 2014, 15, 855. [Google Scholar]
- Furuya, T.; Shibata, D.; Kino, K. Phylogenetic analysis of Bacillus P450 monooxygenases and evaluation of their activity towards steroids. Steroids 2009, 74, 906–912. [Google Scholar] [CrossRef]
- Padayachee, T.; Nzuza, N.; Chen, W.; Nelson, D.R.; Syed, K. Impact of lifestyle on cytochrome P450 monooxygenase repertoire is clearly evident in the bacterial phylum Firmicutes. Sci. Rep. 2020, 10, 13982. [Google Scholar] [CrossRef]
- Jiang, Y.; Li, Z.; Wang, C.; Zhou, Y.J.; Xu, H.; Li, S. Biochemical characterization of three new α-olefin-producing P450 fatty acid decarboxylases with a halophilic property. Biotechno. Biofuels 2019, 12, 79. [Google Scholar] [CrossRef]
- Vardi, A.; Haramaty, L.; Van Mooy, B.A.; Fredricks, H.F.; Kimmance, S.A.; Larsen ABidle, K.D. Host-virus dynamics and subcellular controls of cell fate in a natural coccolithophore population. Proc. Natl. Acad. Sci. USA 2012, 109, 19327–19332. [Google Scholar] [CrossRef]
- Monier, A.; Chambouvet, A.; Milner, D.S.; Attah, V.; Terrado, R.; Lovejoy, C.; Moreau, H.; Santoro, A.E.; Derelle, E.; Richards, T.A. Host-derived viral transporter protein for nitrogen uptake in infected marine phytoplankton. Proc. Natl. Acad. Sci. USA 2017, 114, 489–498. [Google Scholar] [CrossRef]
- Moniruzzaman, M.; Martinez-Gutierrez, C.A.; Weinheimer, A.R.; Aylward, F.O. Dynamic genome evolution and complex virocell metabolism of globally-distributed giant viruses. Nat. Commun. 2020, 11, 1710. [Google Scholar] [CrossRef]
- Blanc-Mathieu, R.; Dahle, H.; Hofgaard, A.; Brandt, D.; Ban, H.; Kalinowski, J.; Ogata, H.; Sandaa, R.A. A persistent giant algal virus, with a unique morphology, encodes an unprecedented number of genes involved in energy metabolism. J. Virol. 2021, 95, e02446-20. [Google Scholar] [CrossRef]
- Brahim Belhaouari, D.; Pires De Souza, G.A.; Lamb, D.C.; Kelly, S.L.; Goldstone, J.V.; Stegeman, J.J.; Colson, P.; La Scola, B.; Aherfi, S. Metabolic arsenal of giant viruses: Host hijack or self-use? eLife 2022, 11, e78674. [Google Scholar] [CrossRef]


| Class | Phage | Phage Host | Sample Location | Predicted Phage ORFs | Flavodoxin aa | Genbank Identifier | Ref. |
|---|---|---|---|---|---|---|---|
| Bacillus | 0305φ8-36 | Bacillus thuringiensis | soil | 246 | 148 | ABS83781.1 | [16] |
| Hakuna | Bacillus thuringiensis | soil | 294 | 159 | YP_009036585.1 | [31] | |
| Megatron | Bacillus thuringiensis | soil | 290 | 159 | YP_009036208.1 | [31] | |
| Riley | Bacillus thuringiensis | soil | 290 | 149 | YP_009055891.1 | [31] | |
| CAM003 | Bacillus thuringiensis | soil | 287 | 150 | YP_009037037.1 | [31] | |
| Evoli | Bacillus thuringiensis | soil | 294 | 150 | YP_009035659.1 | [31] | |
| Hoody T | Bacillus thuringiensis | soil | 270 | 153 | YP_009035330.1 | [31] | |
| Troll | Bacillus thuringiensis | soil | 289 | 149 | YP_008430917.1 | [31] | |
| SalinJah | Bacillus cereus | soil | 292 | 149 | ANH50605.1 | [32] | |
| Eyuki | Bacillus cereus | soil | 300 | 159 | YP_009212082.1 | [33] | |
| AvesoBmore | Bacillus thuringiensis | soil | 301 | 149 | YP_009206486.1 | [33] | |
| BigBertha | Bacillus thuringiensis | soil | 291 | 153 | YP_008771155.1 | [34] | |
| Spock | Bacillus thuringiensis | soil | 283 | 149 | YP_008770352.1 | [35] | |
| Belinda | Bacillus thuringiensis | soil | 295 | 159 | ANM46069.1 | [36] | |
| Tomato | Bacillus thuringiensis | soil | 200 | 159 | QLF85935.1 | [37] | |
| AaronPhadgers | Bacillus thuringiensis | soil | 301 | 159 | ASR78796.1 | [38] | |
| Beyonphe | Bacillus cereus | water | 300 | 149 | QDH49824.1 | [38] | |
| Bubs | Bacillus thuringiensis | soil | 302 | 159 | ASR78601.1 | [38] | |
| Kamfam | Bacillus thuringiensis | soil | 293 | 150 | AXQ67322.1 | [38] | |
| PIPsBest | Bacillus thuringiensis | soil | 301 | 159 | ASR78379.1 | [38] | |
| ALPS | Bacillus thuringiensis | soil | 295 | 159 | QDH50122.1 | [38] | |
| Phireball | Bacillus thuringiensis | soil | 299 | 159 | QDH49415.1 | [38] | |
| OmnioDeoPrimus | Bacillus thuringiensis | soil | 299 | 159 | AXQ67477.1 | [38] | |
| Zainny | Bacillus thuringiensis | soil | 303 | 159 | ASR79375.1 | [38] | |
| vB_BthM-Goe5 | Bacillus thuringiensis | soil | 272 | 150 | AZF89241.1 | [39] | |
| SageFayge | Bacillus thuringiensis | soil | 300 | 159 | YP_009280944.1 | [40] | |
| Nemo | Bacillus thuringiensis | soil | 301 | 159 | YP_009287019.1 | [40] | |
| Nigalana | Bacillus thuringiensis | soil | 302 | 159 | YP_009282535.1 | [40] | |
| DIGNKC | Bacillus thuringiensis | soil | 291 | 159 | AMW62869.1 | [40] | |
| Zuko | Bacillus thuringiensis | soil | 294 | 159 | AMW62552.1 | [40] | |
| Phrodo | Bacillus thuringiensis | soil | 288 | 149 | YP_009290006.1 | [40] | |
| NotTheCreek | Bacillus thuringiensis | soil | 296 | 159 | YP_009284467.1 | [40] | |
| Juglone | Bacillus thuringiensis | soil | 293 | 149 | AMW61744.1 | [40] | |
| Vinny | Bacillus thuringiensis | soil | 297 | 150 | AMW61891.1 | [40] | |
| DirtyBetty | Bacillus thuringiensis | soil | 302 | 159 | YP_009285086.1 | [41] | |
| Kida | Bacillus thuringiensis | soil | 304 | 159 | YP_009279309.1 | [41] | |
| TsarBomba | Bacillus thuringiensis | soil | 247 | 151 | YP_009206942.1 | [42] | |
| HonestAbe | Bacillus thuringiensis | soil | 286 | 159 | AUV57777.1 | [43] | |
| Taffo16 | Bacillus thuringiensis | soil | 284 | 149 | ASZ75860.1 | [43] | |
| Anthony | Bacillus thuringiensis | soil | 279 | 150 | ASU00988.1 | [43] | |
| Saddex | Bacillus thuringiensis | soil | 208 | 159 | AXF41931.1 | [44] | |
| Janet | Bacillus thuringiensis | soil | 285 | 150 | ASR79938.1 | [45] | |
| OTooleKemple52 | Bacillus thuringiensis | soil | 291 | 153 | ASR79614.1 | [45] | |
| SBP8a | Bacillus cereus | soil | 298 | 146 | AOZ62389.1 | DS * | |
| QCM8 | Bacillus cereus | soil | 288 | 151 | AOZ62044.1 | DS * | |
| BJ4 | Bacillus cereus | soil | 298 | 146 | AOZ61763.1 | DS * | |
| Bastille | Bacillus cereus | soil | 273 | 150 | YP_006907364.1 | DS * | |
| Chotacabras | Bacillus thuringiensis | soil | 285 | 149 | QEM43180.1 | DS * | |
| vB_BcoS-136 | Bacillus cohnii | lake sediment | 238 | 146 | AYP68319.1 | DS * | |
| Flapjack | Bacillus thuringiensis | soil | 288 | 149 | ARQ95040.1 | DS * | |
| vB_BanH_ RonSwanson | Bacillus anthracis | soil | - | 202 | UGO50429.1 | DS * | |
| vB_BanH_ Emiliahah | Bacillus anthracis | soil | - | 174 | UGO49205.1 | DS * | |
| vB_BanH_JarJar | Bacillus anthracis | soil | - | 202 | UGO48939.1 | DS * | |
| vB_BanH_ McCartney | Bacillus anthracis | soil | - | 174 | UGO47691.1 | DS * | |
| vB_BanH_ Abinadi | Bacillus anthracis | soil | - | 159 | UGO46389.1 | DS * | |
| Anthos | Bacillus thuringiensis | soil | 290 | 149 | QPY77365.1 | DS * | |
| Smudge | Bacillus thuringiensis | soil | 292 | 159 | ANI24755.1 | DS * | |
| BC-T25 | Bacillus sp. | soil | 236 | 151 | QEG04194.1 | DS * | |
| Listeria | vB_LmoM_AG20 | Listeria monocytogenes | abbatoir | 178 | 141 | YP_007676786.1 | DS * |
| LP-124 | Listeria monocytogenes | silage | 188 | 150 | YP_009784573.1 | [46] | |
| LP-064 | Listeria monocytogenes | silage | 188 | 150 | YP_009592673.1 | [46] | |
| LP-083-2 | Listeria monocytogenes | silage | 189 | 150 | YP_009044596.1 | [46] | |
| LP-048 | Listeria monocytogenes | silage | 177 | 150 | YP_009042933.1 | [46] | |
| LP-125 | Listeria monocytogenes | silage | 189 | 150 | YP_008240106.1 | [46] | |
| LP-Mix_6.2 | Listeria monocytogenes | laboratory | 197 | 150 | QNL32088.1 | [47] | |
| LP-Mix_6.1 | Listeria monocytogenes | laboratory | 195 | 150 | QNL31890.1 | [47] | |
| LP-039 | Listeria monocytogenes | silage | 201 | 150 | QEP53123.2 | [48] | |
| LP-066 | Listeria monocytogenes | silage | 189 | 150 | QDK04972.2 | [48] | |
| LMTA-94 | Listeria monocytogenes | laboratory | 189 | 150 | AID17150.1 | DS | |
| Staphylococcus | Maine | Staphylococcus aureus | pig barn swab | 219 | 130 | QEM41386.1 | [49] |
| MarsHill | Staphylococcus aureus | pig barn swab | 262 | 134 | QQM14610.1 | [50] | |
| Madawaska | Staphylococcus aureus | pig barn swab | 264 | 134 | QQO92731.1 | [50] | |
| Machias | Staphylococcus aureus | pig barn swab | 263 | 140 | QQO92468.1 | [50] | |
| phiIPLA-RODI | Staphylococcus aureus | sewage | 213 | 139 | AJA42083.1 | [51] | |
| phiIPLA-C1C | Staphylococcus aureus | sewage | 203 | 132 | YP_009214514.1 | [51] | |
| Team1 | Staphylococcus aureus | hospital | 217 | 143 | YP_009098273.1 | [52] | |
| MCE-2014 | Staphylococcus aureus | sewage | 204 | 130 | P_009098058.1 | [53] | |
| GH15 | Staphylococcus aureus | sewage | 214 | 130 | YP_007002257.1 | [54] | |
| Metroid | Staphylococcus sp. | soil | 254 | 143 | KE56191.1 | [55] | |
| pSco-10 | Staphylococcus cohnii | duck feces | 131 | 130 | ANH50479.1 | [56] | |
| BT3 | Staphylococcus aureus | animal | 232 | 139 | QVD58098.1 | [57] | |
| Stab23 | Staphylococcus xylosus | sewage | 247 | 130 | VEV88569.1 | [58] | |
| vB_Sau_S24 | Staphylococcus aureus | soil | 209 | 130 | ARM69410.1 | [59] | |
| vB_Sau_Clo6 | Staphylococcus aureus | sewage | 213 | 130 | ARM69197.1 | [59] | |
| vB_SsapH-Golestan-100 | Staphylococcus saprophyticus | - | 192 | 133 | BDA81541.1 | DS * | |
| phiSA_BS1 | Staphylococcus sp. | dairy farm | 200 | 129 | YP_009799552.1 | DS * | |
| vB_SsapH-Golestan-105-M | Staphylococcus saprophyticus | - | 203 | 149 | BDA82285.1 | DS * | |
| SA3 | Staphylococcus aureus | sewage | 223 | 130 | ASZ78055.1 | DS * | |
| PALS_1 | Staphylococcus aureus | animal | 191 | 130 | QDJ97591.1 | DS * |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lamb, D.C.; Goldstone, J.V.; Zhao, B.; Lei, L.; Mullins, J.G.L.; Allen, M.J.; Kelly, S.L.; Stegeman, J.J. Characterization of a Virally Encoded Flavodoxin That Can Drive Bacterial Cytochrome P450 Monooxygenase Activity. Biomolecules 2022, 12, 1107. https://doi.org/10.3390/biom12081107
Lamb DC, Goldstone JV, Zhao B, Lei L, Mullins JGL, Allen MJ, Kelly SL, Stegeman JJ. Characterization of a Virally Encoded Flavodoxin That Can Drive Bacterial Cytochrome P450 Monooxygenase Activity. Biomolecules. 2022; 12(8):1107. https://doi.org/10.3390/biom12081107
Chicago/Turabian StyleLamb, David C., Jared V. Goldstone, Bin Zhao, Li Lei, Jonathan G. L. Mullins, Michael J. Allen, Steven L. Kelly, and John J. Stegeman. 2022. "Characterization of a Virally Encoded Flavodoxin That Can Drive Bacterial Cytochrome P450 Monooxygenase Activity" Biomolecules 12, no. 8: 1107. https://doi.org/10.3390/biom12081107
APA StyleLamb, D. C., Goldstone, J. V., Zhao, B., Lei, L., Mullins, J. G. L., Allen, M. J., Kelly, S. L., & Stegeman, J. J. (2022). Characterization of a Virally Encoded Flavodoxin That Can Drive Bacterial Cytochrome P450 Monooxygenase Activity. Biomolecules, 12(8), 1107. https://doi.org/10.3390/biom12081107







