Abstract
Hsp90 is a ubiquitous molecular chaperone involved in many cell signaling pathways, and its interactions with specific chaperones and cochaperones determines which client proteins to fold. Hsp90 has been shown to be involved in the promotion and maintenance of proper protein complex assembly either alone or in association with other chaperones such as the R2TP chaperone complex. Hsp90-R2TP acts through several mechanisms, such as by controlling the transcription of protein complex subunits, stabilizing protein subcomplexes before their incorporation into the entire complex, and by recruiting adaptors that facilitate complex assembly. Despite its many roles in protein complex assembly, detailed mechanisms of how Hsp90-R2TP assembles protein complexes have yet to be determined, with most findings restricted to proteomic analyses and in vitro interactions. This review will discuss our current understanding of the function of Hsp90-R2TP in the assembly, stabilization, and activity of the following seven classes of protein complexes: L7Ae snoRNPs, spliceosome snRNPs, RNA polymerases, PIKKs, MRN, TSC, and axonemal dynein arms.
Keywords:
molecular chaperones; Hsp90; R2TP; PAQosome; TTT; snoRNP; snRNP; RNA polymerase; PIKK; TSC; dynein arm 1. Overview of Hsp90 Structure and Its Function with R2TP
The Hsp90 molecular chaperone is a central regulator of protein homeostasis in eukaryotes under normal and stressed conditions. Hsp90 is involved in the final stages of client protein folding and maturation. In mammals, there are two cytoplasmic Hsp90 isoforms, Hsp90α and Hsp90β, while in yeast, Hsp82 and Hsc82 are the inducible and constitutively expressed Hsp90 isoforms, respectively [1]. Hsp90 isoforms (referred to here as Hsp90) exist as dynamic homodimers, with each protomer comprised of three domains: an N-terminal domain, the site of ATP binding and hydrolysis [2]; a middle domain, which interacts with Hsp90 substrates; and a C-terminal domain, which forms the Hsp90 dimerization interface (Figure 1A) [3]. The C-terminal domain also contains a MEEVD motif, which is important for interactions with Hsp90 cochaperones that contain TPR domains (see Table 1 for nomenclature). Hsp90 substrates are called clients, and the current set of Hsp90 clients includes steroid hormone receptors, kinases, transcription factors, E3 ubiquitin ligases, and many others that share no common features in terms of sequence, structure, or function [4]. Hsp90-mediated client folding and stabilization is a regulated process that requires the association and release of chaperones and cochaperones. Hsp90 client loading is largely dependent on Hsp70, which binds to nascent or partially folded polypeptides with exposed hydrophobic residues [5,6], and Hop, which functions as an adaptor between Hsp70 and Hsp90 [7].
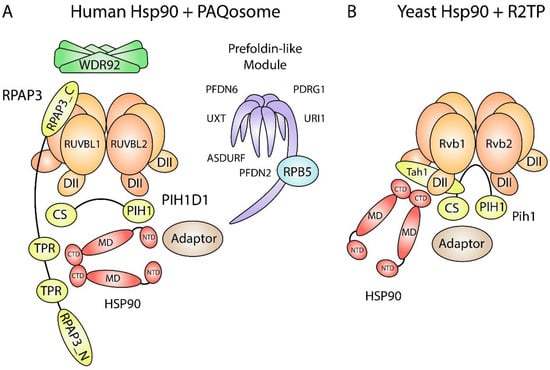
Figure 1.
Schematic of Hsp90 and PAQosome subunits. (A) Human Hsp90 interacts with R2TP through the TPR domains on RPAP3. The RPAP3 C-terminal domain binds to the ATPase side of RUVBL2 and tethers Hsp90 and PIH1D1 to the rest of the R2TP complex. Some Hsp90 and RUVBL1/2 clients are recruited through adaptors. The PIH1 domain in PIH1D1 binds to proteins that contain a DpSDD/E motif. WDR92 and the prefoldin-like module (UPC) may also act as Hsp90 adaptors since they associate with human R2TP. CS, CHORD domain-containing protein Sgt1 domain; CTD, C-terminal domain; DII, Domain II; MD, middle domain; NTD, N-terminal domain; PIH1, Pih1 homology domain; RPAP3_C, RPAP3 C-terminal domain; RPAP3_N, RPAP3 N-terminal domain; TPR, tetratricopeptide domain. (B) Yeast Hsp90 interacts with R2TP through the RPAP3 yeast orthologue Tah1. Tah1 is much smaller than RPAP3, which gives yeast R2TP an open basket structure for client binding. An orthologous prefoldin-like module (UPC) and an orthologue for WDR92 are absent in yeast.

Table 1.
Nomenclature.
In addition to stabilizing tertiary structure, Hsp90 and its cochaperones stabilize the quaternary structure of various macromolecular complexes. In 2005, our group identified Tah1 and Pih1 as Hsp90 interactors in yeast [8]. Tah1 and Pih1 form a heterodimer and interact with AAA+ proteins Rvb1 and Rvb2 to form the R2TP chaperone complex that is conserved in higher eukaryotes including humans. Most notably, the R2TP complex is involved in the assembly of L7Ae ribonucleoproteins [9,10,11], RNA polymerases [12], and PIKK complexes [13]. In humans, R2TP associates with RNA polymerase subunit RPB5, WD40 repeat protein WDR92, and the Unconventional Prefoldin Complex (UPC), comprising of URI1, UXT, PDRG1, PFDN2, PFDN6, and ASDURF [14,15,16]. Altogether, these 12 proteins constitute the PAQosome, Particle for Arrangement of Quaternary Structure (Figure 1A) [17]. The PAQosome is the largest and most intricate chaperone interacting with Hsp90. The R2TP complex is involved in all PAQosome-mediated pathways as the catalytic component, whereas the function of the other subunits is mostly unknown. WDR92 has a specialized role in dynein arm assembly [18], RPB5 likely bridges the interactions between the PAQosome and RNA polymerases, and the UPC may regulate R2TP in response to cell growth and proliferation [19]. Moreover, URI1 mediates nuclear and cytoplasmic shuttling of RNAP subunits, and it has been suggested to do so as part of the PAQosome [20,21]. Thus, PAQosome assembly may occur in the cytoplasm with URI1 facilitating its transport into the nucleus and vice-versa (Figure 2).
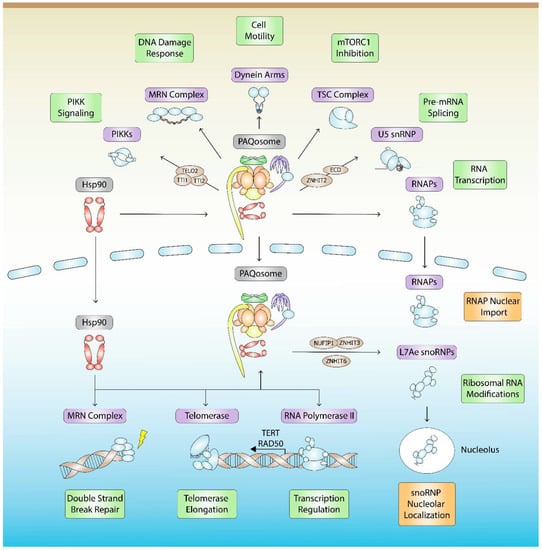
Figure 2.
Hsp90- and PAQosome-mediated quaternary assembly and stabilization pathways. Hsp90 together with the PAQosome are involved in the assembly, stabilization, function (green), or localization (orange) of at least seven classes of protein complexes (purple), which include L7Ae snoRNPs, spliceosome snRNPs, RNA Polymerases, PIKKs, MRN, TSC, and dynein arms. RNAs within each RNP complex that are mentioned in the text are listed. R2TP/PAQosome assembly factors (brown) are shown.
Within the PAQosome, RPAP3 and PIH1D1 are proposed to function as scaffolds for Hsp90 and its diverse client proteins. RPAP3 contains an RPAP3_N domain that mediates interactions with substrates enriched with helical-type domains [22]; two TPR domains, whereby TPR2 has high affinity for Hsp90 [23]; an intrinsically disordered region that makes contacts with RUVBL1 [22]; and an RPAP3_C domain that binds to the ATPase side of RUVBL2 [24]. PIH1D1 contains an N-terminal PIH1 domain that binds DpSDD/E motifs on clients [25,26] and a C-terminal CHORD and Sgt1 (CS) domain that binds RPAP3 [24,27]. Although it has been proposed that PIH1D1 binds to and regulates RUVBL2 ATPase activity as a nucleotide exchange factor, our group has shown that, within the R2TP complex, PIH1D1 binds exclusively to RPAP3 and that PIH1D1 has little effect on RUVBL1/2 ATPase activity and nucleotide binding affinity [22]. Interestingly, although our model suggests that PIH1D1 only interacts with RPAP3 within the R2TP complex, we have identified R2T and R2P complexes in vitro and in cellulo [22]. The significance of these findings in regard to Hsp90 function has yet to be determined.
In yeast, Tah1 is much smaller than RPAP3 and contains two TPR repeats followed by a C-helix and an unstructured region [28,29]. The TPR domain binds the Hsp90 C-terminal MEEVD motif, while the unstructured region binds Pih1. Yeast Pih1 is slightly larger than PIH1D1 and contains an N-terminal PIH1 domain, which also recruits clients with DpSDD/E motifs, and a C-terminal CS domain that binds Tah1 [26,28,30]. The Tah1-Pih1 dimer binds to the Rvb1/2 hexamer DII domains to form the R2TP complex [31,32]. Yeast R2TP forms an open basket that accommodates client proteins and Hsp90 (Figure 1B).
Although it has been established that human Hsp90 interacts with R2TP through RPAP3 [23,24], the details of Hsp90-mediated protein complex assembly are limited, with most of our knowledge restricted to proteomics and in vitro interaction analyses. This review will discuss our current understanding of Hsp90-R2TP in higher metazoans and its roles in protein complex assembly, stabilization, function, or localization for seven classes of protein complexes: L7Ae snoRNPs, spliceosome snRNPs, RNA polymerases, PIKKs, MRN, TSC, and axonemal dynein arms (Figure 2). Of note, human Hsp90 in these studies may refer to either isoform, Hsp90α or Hsp90β, since they have nearly identical structural and functional similarities that cannot be easily distinguished from one another.
2. snoRNP Biogenesis
Eukaryotic ribosomal RNA (rRNA) processing occurs in the nucleolus, which contains numerous small nucleolar RNAs (snoRNA). Most snoRNAs function as sequence-specific guides during rRNA modification [33], while others are involved in folding and cleavage events [34,35]. There are two major families of snoRNAs: box C/D and box H/ACA. Box C/D snoRNAs direct ribose 2′-O-methylation within rRNA and certain spliceosome small nuclear RNA (snRNA) [36,37], and box H/ACA snoRNAs direct the isomerization of uridine to pseudouridine [38]. They are classified based on conserved sequence motifs and their association with common core proteins. Mature snoRNP complexes are comprised of snoRNA and four common core proteins, namely, fibrillarin, NOP56, NOP58, and 15.5K for box C/D snoRNPs, and NHP2, NOP10, GAR1, and NAP57 for box H/ACA snoRNPs (Figure 3). During snoRNP biogenesis, Hsp90 stabilizes NOP58, 15.5K, and NHP2 [9].
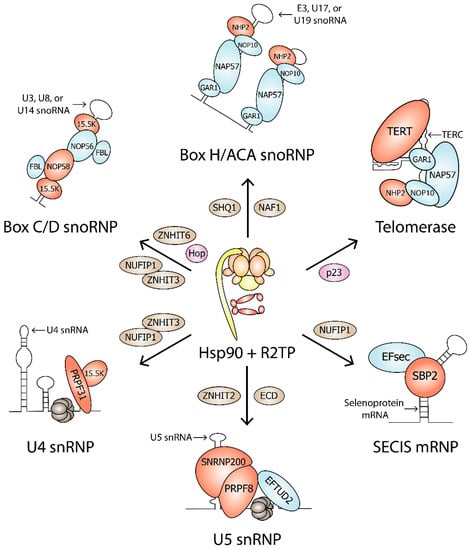
Figure 3.
Hsp90 clients in RNP complexes. Hsp90, Hsp90 cochaperones (pink), R2TP, and assembly factors (brown) are involved in the biogenesis of Box C/D snoRNP, Box H/ACA snoRNP, Telomerase, U4 snRNP, U5 snRNP, and SECIS mRNP. Hsp90 clients are shown in red. Protein complex components that are not Hsp90 clients are shown in blue.
2.1. Box C/D snoRNP Assembly
2.1.1. Role of Hsp90 in Box C/D snoRNP Assembly
Regardless of their snoRNA component, box C/D snoRNPs have a highly conserved asymmetric arrangement of core proteins [39,40,41,42]. The 15.5K protein is part of the L7Ae family of ribosomal proteins and was first identified as a component of the U4/U6.U5 tri-snRNP that binds directly to the 5′ stem loop of U4 snRNA [43], which has a similar primary and secondary structure to box C/D and C′/D′ motifs [44]. 15.5K binding to the box C/D motif is essential for the recruitment of assembly factors RUVBL1 and RUVBL2 and other core proteins including NOP56, NOP58, and fibrillarin [45]. NOP56 and NOP58 are two paralogous proteins that contain NOP and coiled-coil (CC) domains [46]. The NOP domain exhibits RNA and protein binding, which allows NOP56 and NOP58 binding to 15.5K-box C/D snoRNA complexes [47], while the CC domain enables NOP56-NOP58 heterodimerization across the C/D and C′/D′ motifs [39,42]. The NOP56 and NOP58 N-terminal domains together recruit one copy of fibrillarin to the snoRNP complex [39].
Box C/D snoRNP formation requires Hsp90. In HeLa cell extracts, Hsp90, RPAP3, and PIH1D1 coprecipitated with precursor and mature forms of ectopically expressed rat U3 snoRNA [9]. HEK293 cells expressing rat U3 snoRNA and treated with geldanamycin, an Hsp90 inhibitor that blocks the ATP binding site [2,48], had less U3 snoRNA accumulation [9]. These findings were the first to indicate a role for Hsp90 in box C/D snoRNP biogenesis.
Subsequent experiments have suggested that the role of Hsp90 in box C/D snoRNP biogenesis is to stabilize core protein NOP58. When HEK293 cell lines expressing GFP-tagged proteins were treated with geldanamycin, NOP58 and 15.5K failed to accumulate, while a mild effect was seen for NOP56 [9]. NOP58 mutants, NOP58-K310A-A313R, and NOP58-A283P, which cannot assemble into mature snoRNPs, showed stronger interactions with Hsp90 than NOP58-WT [49]. NOP58-A283P also associated with the Hsp70-Hsp90 adaptor Hop [49]. Therefore, NOP58 is likely stabilized through the Hsp70-Hop-Hsp90 pathway during its maturation and assembly into snoRNPs. Interestingly, Hsp90 also stabilized the L7Ae protein SBP2, suggesting that Hsp90 is also involved in SECIS mRNP biogenesis (Figure 3) [9]. SECIS mRNPs associate with selenoprotein mRNAs for translational recoding of a UGA codon that enables the insertion of selenocysteine [50].
2.1.2. Role of R2TP in Box C/D snoRNP Assembly
NOP58 can be stabilized by other chaperones, namely the RUVBL1/2 complex and NOPCHAP1 [49]. RUVBL1 and RUVBL2 were identified as box C/D snoRNP biogenesis factors from an early study identifying mouse U14 snoRNA interactors [51]. Subsequent studies have shown that RUVBL1/2 interact with precursor and mature forms of rat U3 and human U8 snoRNA [9,52,53]. NOPCHAP1 was identified as a snoRNP assembly factor through Stable Isotope Labeling with Amino Acids in Cell Culture (SILAC) experiments, which showed that NOPCHAP1 and RUVBL1/2 associated with nascent NOP58 [54]. NOPCHAP1 binds to NOP58 through the CC-NOP fragment, while it binds to RUVBL1 through the DII domain [49]. The interaction between NOP58 and RUVBL1 is weak, but in the presence of NOPCHAP1, it is enhanced 20-fold [49]. RUVBL1 binds to NOPCHAP1 in the absence of ATP since ATPγS, a non-hydrolyzable ATP analogue, abolished NOPCHAP1 binding [49]. The interaction between NOPCHAP1 and RUVBL1/2 is likely transient and may serve only to direct NOP58 to RUVBL1/2. Interestingly, WT HEK293T cells treated with geldanamycin and NOPCHAP1 KO cells displayed similar levels of reduced NOP58, indicating that Hsp90 and NOPCHAP1 may act on the same pathway [49]. A caveat to consider is that geldanamycin may have additional binding targets that affect the viability of Hsp90 clients.
RUVBL1/2 may also act as a NOP58 chaperone as part of the R2TP complex. HeLa cell extracts separated on linear glycerol gradients showed the assembly factor NUFIP1 and core proteins NOP58 and fibrillarin to be in the same fractions as RUVBL1, RUVBL2, and RPAP3 [9]. Also, pulldown assays in rabbit reticulocyte lysates showed that PIH1D1 directly interacts with NOP56 and NOP58 [9], and that RUVBL1 and RUVBL2 interact with all four box C/D snoRNP proteins [55]. Moreover, the R2TP complex is involved in Cajal body and nucleolar localization of pre-snoRNPs and mature snoRNPs, respectively. In HeLa cells transfected with siRNA, depletion of RUVBL1 and RUVBL2 caused reductions of Cajal body and nucleolar U3 snoRNA [55].
Hsp90 inhibition in HEK293 cells resulted in the disappearance of both NOP58 and 15.5K [9], but the link between Hsp90 ATPase activity and 15.5K stabilization remains unclear. Hsp90 may indirectly stabilize 15.5K by stabilizing NOP58 first. Rather than interacting with Hsp90, 15.5K can bind RUVBL1, RUVBL2, and RUVBL1/2 in the presence of ATP [55,56]. Additionally, RUVBL1/2 was shown to bridge the interaction between 15.5K and core proteins NOP56 and NOP58 [55], which may be important for 15.5K stability. Taken together, Hsp70, Hop, Hsp90, R2TP, and NOPCHAP1 stabilize NOP58, and RUVBL1/2 subsequently recruits 15.5K to NOP56-NOP58, thereby stabilizing 15.5K.
2.1.3. R2TP-Associated Box C/D snoRNP Assembly Factors
The R2TP complex is a highly interactive chaperone complex that works together with other box C/D snoRNP assembly factors, namely, NUFIP1, ZNHIT3, and ZNHIT6 [54]. NUFIP1 acts mainly as a tethering protein, joining 15.5K with NOP56, NOP58, and fibrillarin [9,56]. However, NUFIP1 may also regulate R2TP function during snoRNP assembly since it interacts directly with RUVBL1, RUVBL2, and PIH1D1 [9,56]. For example, the coprecipitation of 15.5K with either NOP56 or NOP58 was enhanced with RUVBL1/2, but was repressed with both RUVBL1/2 and NUFIP1 [55].
NUFIP1 forms a heterodimer with ZNHIT3, an assembly factor belonging to the zf-HIT family, which are often observed in complexes containing RUVBL1 and RUVBL2 [54,57,58,59,60]. ZNHIT3 is required for NUFIP1 stability since siRNA-mediated depletion of ZNHIT3 in HeLa cells resulted in similar decreases in NUFIP1 levels [60]. ZNHIT3 was unable to bind precursor or mature forms of rat U3 snoRNA, but it was able to bind U3 snoRNA mutants that had decreased affinity for NOP56, NOP58, and fibrillarin [54].
Finally, in the presence of ATP, ZNHIT6 interacts with the RUVBL1/2 complex, but not with individual RUVBL1 and RUVBL2 proteins [55]. In addition, ZNHIT6 binds 15.5K, but not NOP56, NOP58, or fibrillarin [56].
2.2. Box H/ACA snoRNP Assembly
The assembly of box H/ACA snoRNPs has been well-established. The pseudouridine synthase NAP57 (also named dyskerin) and core proteins NOP10 and NHP2 form a trimer that binds directly to H/ACA RNA in the absence of GAR1 [61]. Early yeast genetic depletion studies have demonstrated that the core trimer is required for H/ACA RNA stability and that all four core proteins are essential for cell viability [62,63,64,65,66]. The assembly of mammalian H/ACA snoRNPs requires two assembly factors, NAF1 and SHQ1, which are needed for H/ACA RNA accumulation without being part of the mature particles [67,68]. NAF1 is structurally similar to GAR1 [69]. During snoRNP biogenesis, NAF1 and the core trimer associate with H/ACA RNA at the site of transcription [70]. Upon snoRNP maturation, snoRNP particles localize to Cajal bodies or nucleoli where GAR1 replaces NAF1 [68,70]. SHQ1 functions as a NAP57 chaperone by acting as an RNA placeholder, thereby protecting NAP57 from nonspecific RNA binding before its association with H/ACA RNA and other core RNP proteins [71].
2.2.1. Role of Hsp90 in Box H/ACA snoRNP Assembly
Hsp90 is involved in H/ACA snoRNP biogenesis since Hsp90 inhibition led to defects in H/ACA RNA production and core protein stability [9]. HEK293 cells treated with geldanamycin showed decreased levels of telomerase H/ACA RNA [9], which is consistent with TERT, the reverse transcriptase in the telomerase complex, being an Hsp90 client [72]. In addition, geldanamycin-treated cells showed a complete loss of core protein NHP2, indicating NHP2 as a potential Hsp90 client [9]. NAP57 levels were unaffected by geldanamycin [9], presumably because it was stabilized by SHQ1 [73].
Telomerase is a box H/ACA snoRNP complex that synthesizes the G-rich DNA at the 3´-ends of linear chromosomes [74]. In addition to the four box H/ACA core proteins, human telomerase contains reverse transcriptase TERT and telomerase RNA component TERC (Figure 3). Hsp90 and its cochaperone p23 bind TERT, and blocking this interaction inhibits the proper assembly of active telomerase in vitro [72]. TERT and TERC could bind to each other without Hsp90-p23, although this complex was inactive [75]. In addition, Hsp90 inhibitors geldanamycin and novobiocin inhibited telomerase even after telomerase was assembled [75]. Unlike most of their clients, Hsp90 and p23 remain associated with active telomerase [76]. In mammalian cells, Hsp90 regulates TERT expression. In SCC4 cells, a telomerase-positive oral cancer cell line, coprecipitation experiments showed an in vivo interaction between Hsp90 and the TERT promoter. Geldanamycin exposure decreased telomerase activity, TERT promoter activity, and TERT mRNA expression [77]. Additionally, in cerebral endothelial cells, siRNA-mediated Hsp90 depletion inhibited telomerase activity and decreased telomerase protein expression [78].
2.2.2. Role of R2TP in Box H/ACA snoRNP Assembly
Hsp90 chaperone function on telomerase may depend on R2TP, since RUVBL1 and RUVBL2 were reported to interact with TERT and NAP57 [79]. During telomerase assembly, RUVBL1 and RUVBL2 may stabilize NAP57 since depletion of RUVBL1 and RUVBL2 led to a significant reduction of NAP57 steady-state levels [79]. Moreover, RUVBL1 and RUVBL2 associated with TERC in HeLa cell extracts, and RUVBL1 ATPase activity was essential for TERC maintenance [79]. In addition, RUVBL1 and RUVBL2 are involved in the production of other H/ACA RNAs. RUVBL1- or RUVBL2-siRNA knockdown in HeLa cells caused a reduction in the H/ACA RNAs E3 and U17/E1 [10].
Regarding H/ACA snoRNP complex assembly, knockdown of RUVBL1 and RUVBL2 in HeLa cells resulted in a loss of NHP2 and NAP57 [10]. NHP2 makes a direct interaction with NUFIP1 [9], suggesting that NUFIP1 could bridge the interaction between NHP2 and RUVBL1/2 to mediate NHP2 assembly or stability. Similar to its role in box C/D snoRNPs, NUFIP1 may also be involved in bridging interactions between H/ACA snoRNA and core proteins. NUFIP1 coprecipitated with U19 H/ACA snoRNA, and its depletion reduced the levels of U19 and telomerase RNA [9].
During snoRNP assembly, NAP57 is stabilized by the RUVBL1/2 complex and SHQ1 [73,79], and NAP57 requires the R2TP complex to dissociate from SHQ1 [10]. SHQ1 exerts a clamp-like grip on NAP57 through binding to NAP57 in trans: the n-terminal CS domain of SHQ1 binds to the surface that is opposite from the RNA binding surface where the c-terminal SHQ1-specific domain binds [10]. NAP57 recruits the R2TP complex through its unstructured C-terminus [10]. RUVBL1, RUVBL2, and PIH1D1 bind to the same domain on NAP57 as SHQ1. RUVBL1 and RUVBL2 also bind to the CS domain of SHQ1 and remove it from NAP57 through an ATP-independent mechanism [10]. ATP binding and hydrolysis may only be required for the release of RUVBL1/2 from NAP57 after SHQ1 has been removed.
3. Spliceosome snRNP Assembly
The spliceosome is a molecular machine that catalyzes splicing, an essential post-transcriptional modification that removes introns from pre-mRNA. Spliceosomes are comprised of the Prp19 complex, U1 snRNP, U2 snRNP, and the U4/U6.U5 tri-RNP, with each snRNP having their own snRNA component and associated proteins. The spliceosome associates with more than 300 different proteins [80]. Proteomic analyses of purified spliceosomes have shown that the complex is highly conserved, with more than 85% of yeast proteins having a direct human orthologue [81]. Proteomic analyses in yeast and human cells have revealed a role for Hsp90 and R2TP in U4 and U5 snRNP assembly [54,58,59,82].
3.1. U4 snRNP Assembly
3.1.1. Similarities between U4 snRNP and Box C/D snoRNP Complexes
The U4 snRNP is comprised of U4 snRNA, L7Ae protein 15.5K, and splicing component PRPF31 (Figure 3). In addition, 15.5K binds to the 5′ stem-loop of U4 snRNA in a manner similar to the box C/D motif [43,45], enabling PRPF31 recruitment [83]. Without 15.5K, PRPF31 weakly associates with U4 RNA [82]. The association between PRPF31 and 15.5K is essential because an A216P mutation in PRPF31, which abolished the PRPF31-15.5K interaction, resulted in PRPF31 cytoplasmic accumulation, indicating the prevention of PRPF31 incorporation into mature spliceosomes within the nucleus [82].
U4 snRNP shares a few similarities with box C/D snoRNP. In addition to both containing the RNA binding component 15.5K, the U4 snRNP splicing component PRPF31 is homologous to box C/D core proteins NOP56 and NOP58. These three proteins each have a NOP domain, which binds to preformed 15.5K-RNA complexes, and a CC domain, which mediate protein–protein interactions within RNP complexes [47]. Furthermore, U4 snRNA and box C/D snoRNA both associate with the assembly factor NUFIP1, but unlike box C/D snoRNA, U4 snRNA is not dependent on NUFIP1 for its assembly and maturation [9].
3.1.2. Role of Hsp90 and R2TP in U4 snRNP Assembly
Similar to box C/D snoRNP assembly, U4 snRNP assembly and stabilization is mediated by chaperones R2TP, Hsp90, and Hsp70. Co-IP experiments using antibodies against Hsp90, RUVBL1, RUVBL2, RPAP3, and PIH1D1 showed that each associated with U4 snRNA [9]. HEK293 cells treated with geldanamycin showed an almost complete loss of U4 snRNA, a moderate decrease of 15.5K, and a mild effect on PRPF31 [9]. In PRPF31, an A216P mutation prevents its nuclear localization [84], and a K243A/A246R double mutation prevents its interaction with 15.5K [82]. SILAC-IP experiments using HeLa cells expressing PRPF31-A216P or PRPF31-K243A/A246R showed that both PRPF31 mutants were enriched with Hop and Hsp70 [49]. Hsp90 was also present at low levels [49]. These findings show that PRPF31 binds to Hsp70 in the cytoplasm and suggest that the Hsp70-Hop-Hsp90 pathway mediates the PRPF31-15.5K interaction.
The assembly factors NUFIP1 and ZNHIT3 together with the R2TP complex are also involved in U4 snRNP biogenesis. In mammalian cells, NUFIP1 can mediate the interaction between 15.5K and PRPF31, and it may do so with help from the R2TP complex since NUFIP1 also binds to RUVBL1, RUVBL2, and PIH1D1 [9,56]. Moreover, coprecipitation experiments in HEK293T cells showed that NUFIP1, ZNHIT3, and RUVBL1 each associate with U4 snRNA and PRPF31, suggesting that they can mediate the interaction between U4 snRNA and PRPF31 [82]. Indeed, NUFIP1 knockout cells had a two-fold reduction in binding between PRPF31 and U4 snRNA [82].
3.2. U5 snRNP Assembly
3.2.1. Role of Hsp90 and R2TP in U5 snRNP Assembly
U5 snRNP is recruited to the spliceosome as part of the U4/U6.U5 tri-snRNP and is comprised of U5 snRNA, GTPase EFTUD2, helicase SNRNP200, and mRNA processing factor PRPF8 (Figure 3). The Hsp90/R2TP complex is mostly involved in the stabilization and assembly of PRPF8 into mature U5 snRNP particles. SILAC experiments showed that Hsp90 associates with PRPF8 and EFTUD2, as well as with assembly factors AAR2 and ECD [59]. In HeLa cells, Hsp90 ATPase activity was shown to stabilize PRPF8 and SNRNP200, but not EFTUD2 [59]. Hsp90 was also shown to mediate the interaction between PRPF8 and cytoplasmic RPAP3 [59].
PRPF8 is stabilized by the R2TP complex and U5 snRNP assembly factors. In vitro pulldown experiments have shown FLAG-tagged PRPF8 to simultaneously co-elute with purified RUVBL1-RUVBL2, RPAP3-PIH1D1, AAR2, ECD, and ZNHIT2 [85]. PRPF8 can make direct interactions with RUVBL1-RUVBL2 and RPAP3-PIH1D1. The PRPF8-RUVBL1/2 interaction is stronger than the PRPF8-RPAP3-PIH1D1 interaction [85]. PRPF8 mutants that cannot be integrated into mature U5 snRNPs associate more strongly with R2TP than WT PRPF8 [59]. PRPF8 binding to R2TP and AAR2 chaperones was increased in the absence of PIH1D1, suggesting that the formation of the R2TP complex through PIH1D1 binding is important for the release of PRPF8 from R2TP and AAR2 [59].
EFTUD2 may also be stabilized by the R2TP complex and AAR2. EFTUD2 has a DSDED motif, suggesting that it binds to PIH1D1; however, the PIH1D1 N-terminal domain was not sufficient to bind EFTUD2 [59]. Although, mutations in the EFTUD2 DSDED motif did affect EFTUD2 binding with AAR2, RUVBL1/2, and ZNHIT2 [59]. In addition, when PIH1D1, RUVBL2, and ZNHIT2 were depleted, there was less EFTUD2 [59]. EFTUD2 may be recruited to the R2TP complex through RPAP3 since EFTUD2 was shown to interact with ectopically expressed FLAG-RPAP3 in HEK293 lysates [58].
3.2.2. R2TP-Associated U5 snRNP Assembly Factors
ZNHIT2 and the R2TP complex mediate U5 snRNP subunit interactions during U5 snRNP assembly. SILAC experiments showed that ZNHIT2 interacts with R2TP, EFTUD2, PRPF8, SNRNP200, and yeast two-hybrid experiments confirmed direct interactions between ZNHIT2-EFTUD2 and ZNHIT2-RUVBL1 [59]. When ZNHIT2 is knocked out, the interactions between RPAP3-EFTUD2 and RPAP3-PRPF8 were absent [58]. ZNHIT2 is also needed to bridge the binding of RUVBL1/2 with EFTUD2 and PRPF8 [58]. The RUVBL1/2-ZNHIT2 cryo-EM structure shows that the RUVBL1/2 DII domains interact with the ZNHIT2 C-terminal end [85], which is in contrast to another study that showed that the ZNHIT2 HIT domain was essential for binding [58]. Rather than mediating the RUVBL1/2-ZNHIT2 interaction, the HIT domain may regulate the conformation and nucleotide state of RUVBL1/2 [85]. Through binding to the DII domains, ZNHIT2 disrupts the RUVBL1/2 dodecamer [85]. When ZNHIT2 was bound to the hexamer, RUVBL1 still had ADP bound while RUVBL2 was in the apo state [85]. Interestingly, the intrinsically low ATPase activity of RUVBL1/2 hexamers with one Walker B mutant, in either RUVBL1 or RUVBL2, was significantly increased with ZNHIT2 present, suggesting that ZNHIT2 affects the activity of both RUVBL1 and RUVBL2 subunits [85].
ECD is another adaptor protein involved in U5 snRNP biogenesis. In vitro pulldowns showed that ECD co-eluted with GST-ZNHIT2 and RUVBL1/2, and its association with this complex was enhanced when RPAP3 and PIH1D1 were added [85]. ECD can bind RUVBL1 and PIH1D1, either through its DpSDD motif or another uncharacterized binding site [25,86].
4. Hsp90- and R2TP-Mediated RNA Polymerase Assembly and Localization
The eukaryotic RNA polymerases, RNAP I, RNAP II, and RNAP III, are multiprotein complexes that synthesize ribosomal, messenger, and transfer RNA, respectively. The three RNA polymerases are structurally related. Within each complex, the two largest subunits form the catalytic core, while the smaller subunits are located on the periphery. They are also related through having five common subunits: RPB5, RPB6, RPB8, RPB10, and RPB12. Large-scale proteomic screens identified Hsp90, R2TP, and prefoldins as RNAP II interactors (Figure 1) [14,16,87,88]. RNAP II is assembled in the cytoplasm by Hsp90 and R2TP and then imported into the nucleus through URI1 [12,20]. To further analyze the interactions of RNAP II subunits during assembly, Boulon and colleagues performed triple-SILAC purifications on U2OS cells treated with α-amanitin, a small molecule that binds RPB1 and induces its degradation [12,89]. Their findings revealed the presence of two subcomplexes: RPB1-RPB8 and RPB2-RPB3-RPB10-RPB11-RPB12. In addition, each subcomplex associated with a specific set of assembly factors, such as RPAP2, GPN2, GPN3, and GrinL1A. RPB1-RPB8 also associated with R2TP/Prefoldin components RPAP3, PFDN2, and UXT [12].
RPB1 is the largest subunit in RNAP II and interacts with many RNAP II subunits and assembly factors. Coprecipitation and yeast two-hybrid experiments showed that Hsp90 interacts with RPB1 and with the TPR2 domain on RPAP3 [12]. RPB1 interacts with RPAP3 outside of the TPR2 domain [12], implying that RPAP3 stabilizes RPB1 by tethering the interaction between Hsp90 and RPB1. Indeed, long-term RPAP3 depletion in U2OS cells resulted in RPB1 loss [12]. Also, RPAP3 depletion resulted in RPB1 cytoplasmic accumulation in mouse intestinal epithelium cells and crypt base columnar stem cells [90]. Another study showed that RNAP II assembly in melanoma cells was dependent on RPB1 interacting with URI1 [91], but the role of the prefoldin-like module during RNAP II assembly is uncharacterized.
The depletion of RNAP subunits leads to the accumulation of unstable cytoplasmic RPB1. Boulon and colleagues showed that siRNA-mediated depletion of any RNAP II subunit in U2OS cells resulted in RPB1 cytoplasmic accumulation [12]. When cells were treated with geldanamycin, there was a significant decrease of RPB1 in RPB2-, RPB3-, RPB5-, RPB8-, RPB10-, RPB11-, and RPB12-depleted cells, but no significant changes of RPB1 in RPB4-, RPB6-, RPB7-, and RPB9-depleted cells [12]. Hsp90 is essential for stabilizing RPB1, however, Hsp90 binding to RPB1 occurred independent of its ATPase activity [12]. To stabilize RPB1, Hsp90 ATPase activity may mediate interactions between RPB1 and RNAP II subunits RPB5, RPB8, and the subcomplex RPB2-RPB3-RPB10-RPB11-RPB12. The remaining RNAP II subunits, RPB4, RPB6, RPB7, and RPB9, are likely nonessential for RPB1 stability and may be integrated at a later stage. Taken together, these findings show that Hsp90 and R2TP stabilize RPB1 by mediating its interactions with other RNAP II subunits.
R2TP may also be involved in RNAP I and RNAP III assembly since RPAP3-based purifications showed interactions with RPA1 and RPC1, the two largest subunits of RNAP I and RNAP III, respectively [12,16]. Depletion of RPA135, the second largest subunit in RNAP I, increased the interaction between RPA1 and RPAP3, demonstrating that RPAP3 preferentially binds to RPA1 when it is unassembled [12].
5. PIKK Complex Assembly and Stabilization
Phosphatidylinositol 3-kinase-related kinases (PIKKs) belong to the Ser/Thr kinase family and are required for cell proliferation, metabolism, and differentiation. The PIKK family is comprised of ATM, ATR, and DNA-PKcs, which are involved in DNA damage sensing, signaling, and repair (Figure 4); mTOR, a central regulator of cell metabolism, growth, and survival (Figure 4); SMG1, involved in nonsense-mediated mRNA decay; and TRRAP, a pseudokinase that lacks catalytic activity but functions as a large protein interaction hub. Although they have diverse functions, PIKKs share a common domain architecture where their N-termini carry long arrays of HEAT repeats [92], and their C-termini phosphorylate target proteins using a region related to the domain of PI3 kinase [93]. The PIKKs oligomerize with other proteins to form complexes, yet none of their binding partners or target proteins are common to all family members. During PIKK complex assembly and function, they each depend on Hsp90, the TTT complex, and the R2TP complex [13,94,95].
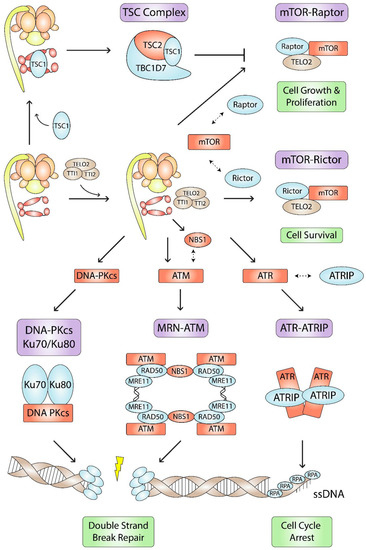
Figure 4.
Hsp90- and R2TP-mediated PIKK, MRN, and TSC complex assembly pathways. Hsp90, R2TP, and the TTT (brown) are involved in the assembly, stabilization, or function (green) of several complexes (purple) involved in cell metabolism and DNA damage responses. Hsp90-R2TP stabilizes its clients (red) and mediates interactions (dashed double-sided arrows) between its clients and other complex subunits (blue).
5.1. Role of Hsp90-TTT in PIKK Complex Assembly
The TTT complex, comprising of TELO2, TTI1, and TTI2, was discovered through a large-scale proteomic analysis identifying Tel2 interactors in fission yeast [96,97]. Each component of the TTT complex is mutually dependent on each other for their stability [98]. TTI1 provides a platform for TELO2 and TTI2 to bind to its central and C-terminal regions, respectively [99]. Functional studies in yeast, C. elegans, and mammalian cells show that components of the TTT complex are essential for proper PIKK signaling pathways, namely, the DNA damage response [94,98,100,101,102,103,104,105,106], metabolic stress [107,108,109,110,111,112,113], nonsense-mediated mRNA decay [114,115], and transcriptional regulation [116,117]. The TTT complex recognizes and stabilizes PIKKs cotranslationally before mediating their assembly into larger complexes [95,111,118].
The TTT complex functions as an Hsp90 cochaperone. Co-IP experiments coupled with MALDI-TOF using HeLa S3 cell extracts revealed that FLAG-tagged TTT subunits associate with PIKKs, R2TP subunits, and Hsps [118]. TTI2 was shown to associate with Hsp90 [118]. Co-IPs in HEK293 cells showed that TELO2 and TTI1 could also associate with Hsp90, and Hsp90 ATPase activity was needed to stabilize TELO2 and TTI1 [94]. Thus, Hsp90 ATPase activity is essential for PIKK stabilization and proper PIKK signaling [94,119]. Hsp90 inhibition in HeLa cells interfered with TELO2-ATR and TELO2-mTOR interactions, which decreased the association of ATR with ATRIP, and mTOR with Raptor and Rictor [118]. These findings have important implications on overall cell metabolism. ATR-ATRIP interact with RPA on ssDNA to initiate cell cycle arrest [120]. mTOR and Raptor are part of the mTORC1 complex, which is involved in cell growth and proliferation, and mTOR and Rictor are part of the mTORC2 complex, which is involved in cell survival (Figure 4) [121]. Hsp90 inhibition coupled with glutaminase inhibition has been shown to be an effective therapeutic strategy against mTORC1-driven tumors [122]. Furthermore, Hsp90 inhibition also interfered with TELO2-ATM and TELO2-DNA-PKcs interactions [118]. Similar to ATR and mTOR, ATM and DNA-PKcs interactions with TELO2 may mediate ATM-MRN and DNA-PKcs-Ku70/Ku80 interactions. The ATM-MRN and DNA-PKcs-Ku70/Ku80 complexes are both involved in DNA double-strand break repair (Figure 4) [123,124,125,126].
5.2. Role of R2TP-TTT in PIKK Complex Assembly
In addition to Hsp90, the RUVBL1/2 complex also functions as a TTT cochaperone. The mechanism of RUVBL1/2-mediated mTORC1 complex assembly and activation has recently been elucidated. The cryo-EM structure of the human R2TP-TTT complex shows that TTT binds simultaneously with DII domains from consecutive RUVBL1 and RUVBL2 subunits of the RUVBL1/2 hexameric ring [127]. RUVBL1/2 and TTT cooperate to recruit mTOR to the complex. In vitro binding experiments showed that the human TTT complex coprecipitates with a mTOR C-terminal fragment, and the RUVBL1/2 complex coprecipitates with an mTOR N-terminal fragment [127]. Although TTT complex binding to RUVBL1/2 inhibited RUVBL1/2 ATPase activity in vitro [127], RUVBL1 ATPase activity is essential for TTT complex formation and mTOR complex activation in cellulo [112]. When endogenous RUVBL1 was knocked out in TSC2−/− MEFs, which have high mTORC1-driven translation, and rescued with a RUVBL1 ATPase-activity deficient mutant, mTORC1 complex dimerization was inhibited [112]. RUVBL1 ATPase inhibition prevented mTOR-TELO2, TTI1-TELO2, and RUVBL1-TELO2 interactions, indicating disassembly of the RUVBL1/2-mTOR-TTT complex [112]. Interestingly, piperlongumine, a cancer therapeutic for mTORC1-addicted cells, targets RUVBL1/2 to prevent the formation of the RUVBL1/2-TTT complex [128].
Several scaffold and adaptor proteins mediate RUVBL1/2-TTT-PIKK complex assembly. WAC was identified as an adaptor between RUVBL1/2 and TTT during energy-dependent mTORC1 dimerization [108]. Co-IPs in HeLa cell lysates showed that URI1 associates with TELO2, TTI1, TTI2, Hsp90, RUVBL1, RUVBL2, RPAP3, and PIH1D1 [94], suggesting that it may bridge interactions between TTT, Hsp90, and R2TP. Moreover, PIH1D1 within the R2TP complex acts as an adaptor between RUVBL1/2 and TTT. PIH1D1 binds to two constitutively phosphorylated serine residues (S487 and S491) on TELO2 [13]. Co-IPs using TELO2 knockout MEFs rescued with TELO2 S487A and S491A mutants showed compromised association between TELO2 with PIH1D1 and RUVBL1 [13]. In addition, compared to MEF knockout cells rescued with WT TELO2, cells rescued with the TELO2 substitution mutants had a complete reduction of SMG1, a significant reduction of mTOR, and a minor reduction of ATM, ATR, and DNA-PKcs [13]. Inhibition of CK2, the kinase that phosphorylates TELO2 [13], also reduced levels of endogenous SMG1 [114]. Further analysis of the PIH1D1 binding motif, DpSDD, on TELO2 showed that it was highly conserved from yeast to humans, suggesting a mechanism by which PIH1D1 recognizes its substrates [25]. As mentioned above, the DpSDD motif is present in other proteins involved in PAQosome-mediated assembly pathways [25], including the E3 ligase UBR5, which interacts with the H/ACA ribonucleoprotein complex and regulates ribosomal RNA biogenesis [129]; RNAP II subunit RPB1, the largest subunit in the RNAP II complex; EFTUD2, the splicing component of U5 snRNP; and ECD, an adaptor protein for U5 snRNP complex assembly [130].
6. Hsp90- and R2TP-Mediated MRN Complex Stabilization
The MRN complex is involved in sensing, processing, and repairing DNA strand breaks (DSBs). The complex is comprised of the nuclease MRE11, ATPase RAD50, and PIKK scaffold NBS1 (Figure 4). During the DNA damage response, the MRN complex binds to DSBs, recruits and activates the PIKKs ATM and ATR, and facilitates DNA repair by homologous recombination and non-homologous end-joining [131,132,133,134,135]. Hypomorphic mutations in MRE11, NBS1, and RAD50 cause ataxia-telangiectasia-like disease [136], Nijmegen breakage syndrome [137], and Nijmegen breakage syndrome-like disorder [138], respectively. Both ataxia-telangiectasia-like disease and Nijmegen breakage syndrome are characterized by genomic instability, hypersensitivity to radiation, and increased susceptibility to cancer.
MRE11 is a conserved 70–90 kDa dimeric protein that has endo- and exonuclease activity against single- and double-stranded DNA [139,140,141]. MRE11 stability was shown to be dependent on its interaction with PIH1D1 [142]. PIH1D1 interacts with MRE11 at S558/S561 or S688/S689 when both serines of each site are phosphorylated, with the latter being the major binding site [142]. Cells expressing MRE11 mutated at S688/S689 had reduced levels of stable MRE11 compared to WT cells [142]. In addition, RPE1, U2OS, and HCT116 cells treated with siRNA against PIH1D1 had reduced levels of MRE11 and slightly reduced levels of RAD50 and NBS1 [142].
RAD50 is a 150 kDa protein that contains an ABC-type ATPase domain that binds and unwinds dsDNA termini [143,144]. Hsp90 ATPase activity is essential for RAD50 expression [145]. HO-8910 ovarian cancer cells treated with 17-AAG had significantly reduced levels of RAD50 [145]. Hsp90 is also important for RAD50-mediated BRCA1 recruitment to DSBs. BRCA1, a tumor suppressor protein linked to breast and ovarian cancer, interacts with RAD50 in vitro and in vivo and co-localizes with RAD50, MRE11, and NBS1 in irradiation-induced foci [146]. In MCF7 breast cancer cells, 17-AAG decreased BRCA1 protein levels in a dose- and time-dependent manner and impaired irradiation-induced homologous recombination and non-homologous end joining [147].
NBS1 is an 85 kDa protein containing two BRCT domains that bind pSDpTD motifs on interacting proteins, including repair and checkpoint proteins at DSBs [148,149]. Hsp90α stabilizes NBS1 and ATM, but not MRE11 and RAD50 [150]. Hsp90 also stabilizes the interaction between NBS1 and ATM and is needed for MRN translocation to nuclear foci after irradiation [151]. Upon irradiation-induced ATM activation, ATM phosphorylates both NBS1 and Hsp90α, and pNBS1 dissociates from pHsp90 and translocates to DSBs [150,152]. When PIKKs phosphorylate Hsp90 at Thr 7, Hsp90α also translocates to DSBs [153]. By contrast, another study showed that when ATM phosphorylates Hsp90 at Thr 5 and Thr 7, Hsp90α is not significantly recruited to DSBs [150]. In addition to regulating Hsp90 localization, Hsp90 phosphorylation is essential for MRN stabilization since Cdc7-Dbf4-mediated phosphorylation of S164 on Hsp90 was required for stabilizing the Hsp90-TELO2-MRN complex and resulted in enhanced ATM/ATR signaling [154].
7. Hsp90- and R2TP-Mediated TSC Complex Stabilization
The TSC complex, comprising of tumor-suppressor proteins TSC1, TSC2, and TBC1D7, inhibits the mTORC1 complex, which controls cell growth and proliferation (Figure 4) [155]. Loss-of-function mutations in TSC1 or TSC2 have been linked to tuberous sclerosis, a rare genetic disorder that causes tumor growth in multiple organs and neurological symptoms [156]. Within the TSC complex, TSC1 binds and stabilizes both TSC2 and TBC1D7 [157,158,159]. To inactivate mTORC1, TSC2, which contains a GAP domain, catalyzes the conversion of Rheb-GTP to Rheb-GDP [160,161].
TSC1 was reported to be an Hsp90 cochaperone that inhibits Hsp90 ATPase activity (Figure 4) [162]. TSC1 enables TSC2 binding to Hsp90, which prevents TSC2 ubiquitin-mediated proteasomal degradation [162]. TSC1 binding to Hsp90 was also important for stability and activity of kinase client proteins such as c-Src, CDK4, and Ulk1, as well as non-kinase client proteins such as glucocorticoid receptor and folliculin [162]. In bladder cancer cells, TSC1 facilitated Hsp90 acetylation at K407/K419, which increased its binding affinity for Hsp90 inhibitor ganetespib [163]. In contrast to bladder cancer cells, however, CAL-72 and PEER cells, which have a complete loss of TSC1 and reduced TSC2 expression, were also sensitized to ganetespib with IC50 values of 22 and 3 nM, respectively [164]. In addition, hepatocellular cancer cell lines SNU-398, SNU-878, and SNU-886, which have a complete loss of TSC2 and normal TSC1 expression, had IC50 values of 9, 14, and 35 nM, respectively [164]. Nevertheless, these findings show that TSC1 and TSC2 influence Hsp90 activity.
The TSC complex may be stabilized through the PAQosome. Co-IP experiments in HeLa cells showed that FLAG-tagged URI1 and RPAP3 interacted with endogenous TSC1 and TSC2 [58]. TAP-MS of each TSC complex subunit demonstrated high confidence interactions with RUVBL1, RUVBL2, RPAP3, PIH1D1, WDR92, and URI1 [58]. A SILAC proteomic analysis using the N-terminal domain of PIH1D1 showed that it associated with all three subunits of the TSC complex [59]. The significance of these interactions is unknown. The PAQosome may act as a loading dock that stabilizes each TSC subunit before combining them into a single complex. Moreover, the PAQosome may scaffold TSC1, to regulate Hsp90 ATPase activity, or it may scaffold TSC2, to facilitate loading onto Hsp90 [162].
8. Axonemal Dynein Arm Assembly
Motile cilia are small microtubule-based organelles required for fluid transport and cell motility in many organisms. In humans, motile cilia are essential for the generation of left-right asymmetry during embryonic development, sperm motility, and the movement of fluid in the respiratory tract, brain ventricular system, and oviducts [165]. Motile cilia contain a 9 + 2 axoneme comprised of nine outer doublet microtubules and a pair of central microtubules. Between each outer doublet, there are several multiprotein complexes, which include the inner dynein arms (IDA) and outer dynein arms (ODA). Dynein is a AAA+ ATPase that mediates microtubule sliding and subsequent ciliary movement [166]. Before being incorporated into the axoneme, dynein arms are preassembled in the cytoplasm [167,168].
8.1. DNAAFs Form Complexes with Hsp90
DNAAFs were discovered through genetic analyses of families with primary ciliary dyskinesia and mutation studies in animals [168,169,170,171,172,173,174,175,176,177,178,179]. Although most of their functions are still being investigated, it is clear that DNAAFs work together with Hsp90 to mediate IDA and ODA assembly [180]. DNAAF2, DNAAF4, DNAAF6, and DNAAF11 have domains that associate with Hsp90, including PIH1, CS, and TPR domains, while DNAAF1, DNAAF3, DNAAF5, and DNAAF7 lack Hsp90 association domains.
In vertebrates, the PIH1 domain is present in at least four proteins: DNAAF2, DNAAF6, PIH1D1, and PIH1D2 [181,182,183]. Each protein has been shown to be involved in ciliary dynein arm assembly [182]. DNAAF2 and DNAAF6 each contain an N-terminal PIH1 domain followed by a CS domain. In mouse testis extracts, DNAAF2 coprecipitated with Hsp70 but not Hsp90 [175], whereas DNAAF6 coprecipitated with both Hsp70 and Hsp90 [181]. In addition, a yeast two-hybrid analysis showed that DNAAF6 interacts with Hsp90, DNAAF2, and DNAAF4 [176].
DNAAF4 contains a C-terminal TPR domain, and a yeast two-hybrid screen showed that it interacts with Hsp70 and Hsp90 C-termini via the EEVD motif that binds TPR domains [184]. These interactions were confirmed through coprecipitation experiments in mouse trachea tissues [177]. In addition, DNAAF4 coprecipitated with DNAAF2 in HEK293 cells [177]. Based on their domains, DNAAF2 and DNAAF4 may form R2TP-like complexes that mediate Hsp90 involvement in dynein arm assembly (Figure 5) [26]. Moreover, TTC12 has recently emerged as another dynein arm assembly factor, and it contains a stretch of three TPR domains [185], suggesting that it may also be involved in forming R2TP-like complexes.
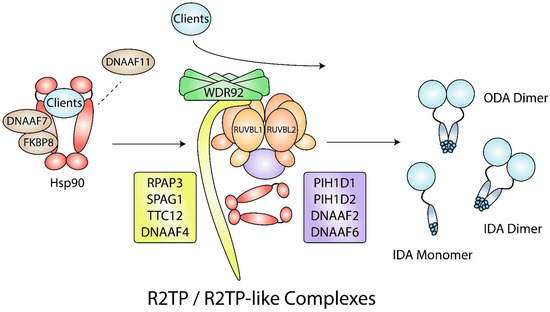
Figure 5.
Hsp90- and R2TP/R2TP-like complex-mediated dynein arm assembly. During dynein arm assembly, DNAAF7 and FKBP8 act as Hsp90 cochaperones that are required for the folding of dynein arm clients. DNAAF11 may be needed for client release from Hsp90 to WDR92, R2TP, and R2TP-like complexes. Other clients may not require Hsp90 for folding and may interact with WDR92, R2TP, and R2TP-like complexes directly. R2TP-like complexes contain the RUVBL1/2 hexamer and may have a combination of RPAP3-like (yellow) and PIH1D1-like (purple) proteins. IDA, inner dynein arm; ODA; outer dynein arm.
DNAAF1 and DNAAF7, which lack Hsp90 binding domains, have been linked to Hsp90. Streptavidin-II/FLAG tandem affinity purification coupled with mass spectrometry (SF-TAP/MS) experiments using HEK293 lysates showed that DNAAF1 associates with several Hsps, including Hsp70 and Hsp90 [186]. Although DNAAF7 lacks an Hsp90 binding domain, endogenous DNAAF7 coprecipitations from P30 mouse testes, P7 mouse oviducts, and primary ciliated HEK293 cells revealed the presence of Hsp90 [187]. Hsp90 may have an indirect interaction with DNAAF7 through FKBP8, an immunophilin belonging to the FK506-binding protein family, thereby forming a DNAAF7-FKBP8-Hsp90 complex. FKBP8 contains a TPR domain that interacts with Hsp90 [188], and it was present in endogenous DNAAF7 coprecipitations from P30 mouse testes and differentiating human tracheal epithelial cultures [187]. There have been no reports linking DNAAF3 and DNAAF5 to Hsp90. Aside from it being essential for dynein arm assembly, little is known about DNAAF3 function, but it may have a role similar to DNAAF1 and DNAAF2 [168]. Coprecipitation experiments using human bronchial epithelial tissues showed that DNAAF5 does not interact with Hsp70 or Hsp90 [169].
DNAAF11 (formerly named LRRC6) is another essential protein for dynein arm assembly [189,190]. HEK293T cells co-expressing DNAAF7 and DNAAF11 and treated with protein synthesis inhibitor cycloheximide for 48 h had 44.4% of its DNAAF11 remaining, while DNAAF11 expressed alone had 7.8% remaining [191], indicating that DNAAF7 is needed to stabilize DNAAF11. DNAAF11 may interact with Hsp90 directly through its CS domain, or indirectly through its interactors DNAAF7 and RUVBL2 [173,178,187,192]. To release client proteins from Hsp90, p23 binding and ATP hydrolysis is required [193]. Thus, DNAAF11 binding to Hsp90 may promote the release of dynein arms from DNAAF7-FKBP8-Hsp90 to other chaperone complexes, including R2TP and R2TP-like complexes (Figure 5) [187].
8.2. R2TP and R2TP-like Complexes Are Dynein Arm Assembly Factors
The R2TP complex may be involved in late-stage dynein arm assembly. Similar to DNAAFs, the catalytic components of R2TP, RUVBL1 and RUVBL2, were demonstrated to be involved in dynein arm assembly through mutational analyses in animal models. Inducible deletion of RUVBL1 in mouse oviducts resulted in the absence of outer dynein arms and the appearance of undefined protein clusters [194]. Streptavidin-II/FLAG tandem affinity purification (SF-TAP) using HEK293 cell lysates showed that DNAAF1 interacts with RUVBL1 and RUVBL2, and that the RUVBL1 interaction was reduced with mutant DNAAF1 [186]. RUVBL1 knockdown in hTERT-RPE1 cells showed increased co-localization between intraflagellar transport protein IFT1 and DNAAF1, suggesting that RUVBL1 mediates DNAAF1 transport or localization [186]. In zebrafish, RUVBL1 and RUVBL2 are enriched in cytoplasmic puncta in zebrafish ciliated tissues, and cilia motility is lost in zebrafish with either RUVBL1 or RUVBL2 mutants [192,195]. RUVBL2 interacts with DNAAF11, which has a similar domain composition to DNAAF1 [192]. The RUVBL2-DNAAF11 complex was essential for dynein arm assembly in zebrafish [192]. Altogether, these findings suggest the presence of cytoplasmic R2TP-like complexes that mediate dynein arm assembly.
In a conditional mouse model, loss of RUVBL1 resulted in immotile spermatozoa due to reduced ODA components, DNAI1 and DNAI2 [195]. In mouse testes, RUVBL2 interacted with Hsp90, suggesting that RUVBL2 scaffolds DNAI1 and DNAI2 to Hsp90 [195]. RUVBL1 may also scaffold IDA and ODA components to Hsp90 through the R2TP-like complex R2SP, comprising of RUVBL1, RUVBL2, SPAG1, and PIH1D2 [196]. Both SPAG1 and RPAP3 contain RPAP3_C and TPR domains, while PIH1D1 and PIH1D2 both contain N-terminal PIH1 and C-terminal CS domains. In zebrafish, SPAG1 null mutations resulted in dorsal body curvature and hydrocephalus, indications of primary ciliary dyskinesia [197], while double-null mutations in PIH1D2 and DNAAF2 resulted in abnormal sperm motility [182]. RUVBL2, SPAG1, and PIH1D2 were found to be ubiquitously expressed in all human tissues and had moderate to high enrichment in the testes [196]. The R2SP complex was shown to facilitate the formation of liprin-α2 complexes [196], which are involved in synaptic vesicle release [198]. Interestingly, PIH1 domain-containing proteins DNAAF2 and DNAAF6 were also enriched in the testes, suggesting the presence of multiple R2TP-like complexes [196].
In addition to R2TP and R2TP-like complexes, proper dynein arm assembly requires WDR92 (recently renamed DNAAF10), which is also highly expressed in human testes [199]. In Chlamydomonas, experiments using insertion and truncation mutants showed that WDR92 is needed to stabilize ODA and IDA heavy chains during preassembly [200,201]. Co-IPs using HEK293 cells and in vitro pulldowns showed that WDR92 interacts directly with RPAP3 [200,202], suggesting that a WDR92-R2TP complex is needed for proper dynein arm assembly (Figure 5). In addition, Drosophila WDR92 was shown to interact with CG18472, the closest Drosophila orthologue of human SPAG1 [197,203]. A proteomic analysis also supports a possible interaction between human WDR92 and SPAG1 [58]. These findings suggest the possibility of a WDR92-R2SP complex.
RUVBL1 and RUVBL2 were recently demonstrated to be involved in the synthesis of cytoplasmic cilia, in which the axoneme is exposed to the cytoplasm [204,205]. Cytoplasmic cilia are found in male gametes, including human and Drosophila sperm. While investigating Drosophila spermiogenesis, Fingerhut and colleagues identified a novel RNP granule located at the axoneme distal end, the site of ciliogenesis [205]. The RNP granule contained RUVBL1 and RUVBL2, as well as mRNA that encodes axonemal dynein arms. By localizing translation, dynein arms can be integrated into the axoneme directly from the cytoplasm. RUVBL1 and RUVBL2 were essential for dynein arm integration and subsequent spermatozoa motility. Similar to their involvement with other RNPs, RUVBL1 and RUVBL2 were also essential for RNP granule formation [205].
9. Concluding Remarks
These studies demonstrate that Hsp90 and R2TP have many diverse and essential roles in macromolecular complex assembly that often complement each other. Before complex assembly, Hsp90 initiates subunit expression (e.g., RAD50, TERT transcription) and regulates RNA levels (e.g., U3 snoRNA, U4 snRNA). During complex assembly, Hsp90 stabilizes clients that compose multisubunit complexes (e.g., L7Ae proteins 15.5K, NHP2, SBP2; RNAP subunits RPA1, RPB1, RPC1; PIKK proteins ATM, ATR, DNA-PKcs, mTOR, SMG-1, TRRAP), while R2TP mediates important interactions between Hsp90 and clients (e.g., Hsp90-RPB1), adaptors and clients (e.g., TELO2-mTOR, TELO2-ATR), and complex subunits (e.g., 15.5K-NOP56, 15.5K-NOP58). After complex assembly, Hsp90 is critical for the function of some complexes (e.g., telomerase elongation, MRN-mediated recruitment of BRCA1 to DSBs), and R2TP together with its associated prefoldin-like module are involved in the localization of assembled protein complexes (i.e., R2TP-mediated Cajal body and nucleolar localization of snoRNPs, URI1-mediated nuclear localization of RNAP II). Thus, in addition to the canonical role of Hsp90 in client stabilization, these findings highlight the additional roles of Hsp90, together with R2TP, in quaternary complex assembly. Determining how Hsp90 integrates its clients into multiprotein complexes may facilitate the discovery of novel therapeutic drug targets. For example, inhibiting Hsp90-mediated TELO2-mTOR interactions may be an effective adjuvant against mTORC1-driven tumors. Thus, the role of Hsp90 during complex assembly and how it functions with its chaperones and cochaperones, especially TTT and R2TP, should be further investigated.
Author Contributions
Conceptualization, J.L. and W.A.H.; writing—original draft preparation, J.L.; writing—review and editing, W.A.H.; visualization, J.L.; supervision, W.A.H.; funding acquisition, W.A.H. All authors have read and agreed to the published version of the manuscript.
Funding
Research in the PI’s lab on this project is funded by the Canadian Institutes of Health Research project grant (PJT-173491). The APC was funded by Canadian Institutes of Health Research project grant (PJT-173491).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
J.L. held an Ontario Graduate Scholarship (2019–2020) and currently holds a fellowship from the Centre for Pharmaceutical Oncology at the University of Toronto (2021–2022). The work in WAH’s group on this subject is supported by the Canadian Institutes of Health Research project grant (PJT-173491).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Gupta, R.S. Phylogenetic analysis of the 90 kD heat shock family of protein sequences and an examination of the relationship among animals, plants, and fungi species. Mol. Biol. Evol. 1995, 12, 1063–1073. [Google Scholar] [CrossRef]
- Prodromou, C.; Roe, S.M.; O’Brien, R.; Ladbury, J.E.; Piper, P.W.; Pearl, L.H. Identification and structural characterization of the ATP/ADP-binding site in the Hsp90 molecular chaperone. Cell 1997, 90, 65–75. [Google Scholar] [CrossRef]
- Harris, S.F.; Shiau, A.K.; Agard, D.A. The crystal structure of the carboxy-terminal dimerization domain of htpG, the Escherichia coli Hsp90, reveals a potential substrate binding site. Structure 2004, 12, 1087–1097. [Google Scholar] [CrossRef] [PubMed]
- Schopf, F.H.; Biebl, M.M.; Buchner, J. The HSP90 chaperone machinery. Nat. Rev. Mol. Cell Biol. 2017, 18, 345–360. [Google Scholar] [CrossRef] [PubMed]
- Gragerov, A.; Zeng, L.; Zhao, X.; Burkholder, W.; Gottesman, M.E. Specificity of DnaK-peptide binding. J. Mol. Biol. 1994, 235, 848–854. [Google Scholar] [CrossRef]
- Rudiger, S.; Germeroth, L.; Schneider-Mergener, J.; Bukau, B. Substrate specificity of the DnaK chaperone determined by screening cellulose-bound peptide libraries. EMBO J. 1997, 16, 1501–1507. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Smith, D.F. Hop as an adaptor in the heat shock protein 70 (Hsp70) and hsp90 chaperone machinery. J. Biol. Chem. 1998, 273, 35194–35200. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Davey, M.; Hsu, Y.C.; Kaplanek, P.; Tong, A.; Parsons, A.B.; Krogan, N.; Cagney, G.; Mai, D.; Greenblatt, J.; et al. Navigating the chaperone network: An integrative map of physical and genetic interactions mediated by the hsp90 chaperone. Cell 2005, 120, 715–727. [Google Scholar] [CrossRef] [PubMed]
- Boulon, S.; Marmier-Gourrier, N.; Pradet-Balade, B.; Wurth, L.; Verheggen, C.; Jady, B.E.; Rothe, B.; Pescia, C.; Robert, M.C.; Kiss, T.; et al. The Hsp90 chaperone controls the biogenesis of L7Ae RNPs through conserved machinery. J. Cell Biol. 2008, 180, 579–595. [Google Scholar] [CrossRef] [PubMed]
- Machado-Pinilla, R.; Liger, D.; Leulliot, N.; Meier, U.T. Mechanism of the AAA+ ATPases pontin and reptin in the biogenesis of H/ACA RNPs. RNA 2012, 18, 1833–1845. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Kakihara, Y.; Gribun, A.; Huen, J.; Yang, G.; Khanna, M.; Costanzo, M.; Brost, R.L.; Boone, C.; Hughes, T.R.; et al. Molecular chaperone Hsp90 stabilizes Pih1/Nop17 to maintain R2TP complex activity that regulates snoRNA accumulation. J. Cell Biol. 2008, 180, 563–578. [Google Scholar] [CrossRef] [PubMed]
- Boulon, S.; Pradet-Balade, B.; Verheggen, C.; Molle, D.; Boireau, S.; Georgieva, M.; Azzag, K.; Robert, M.C.; Ahmad, Y.; Neel, H.; et al. HSP90 and its R2TP/Prefoldin-like cochaperone are involved in the cytoplasmic assembly of RNA polymerase II. Mol. Cell 2010, 39, 912–924. [Google Scholar] [CrossRef] [PubMed]
- Horejsi, Z.; Takai, H.; Adelman, C.A.; Collis, S.J.; Flynn, H.; Maslen, S.; Skehel, J.M.; de Lange, T.; Boulton, S.J. CK2 phospho-dependent binding of R2TP complex to TEL2 is essential for mTOR and SMG1 stability. Mol. Cell 2010, 39, 839–850. [Google Scholar] [CrossRef]
- Cloutier, P.; Al-Khoury, R.; Lavallee-Adam, M.; Faubert, D.; Jiang, H.; Poitras, C.; Bouchard, A.; Forget, D.; Blanchette, M.; Coulombe, B. High-resolution mapping of the protein interaction network for the human transcription machinery and affinity purification of RNA polymerase II-associated complexes. Methods 2009, 48, 381–386. [Google Scholar] [CrossRef] [PubMed]
- Cloutier, P.; Poitras, C.; Faubert, D.; Bouchard, A.; Blanchette, M.; Gauthier, M.S.; Coulombe, B. Upstream ORF-Encoded ASDURF Is a Novel Prefoldin-like Subunit of the PAQosome. J. Proteome Res. 2020, 19, 18–27. [Google Scholar] [CrossRef] [PubMed]
- Jeronimo, C.; Forget, D.; Bouchard, A.; Li, Q.; Chua, G.; Poitras, C.; Therien, C.; Bergeron, D.; Bourassa, S.; Greenblatt, J.; et al. Systematic analysis of the protein interaction network for the human transcription machinery reveals the identity of the 7SK capping enzyme. Mol. Cell 2007, 27, 262–274. [Google Scholar] [CrossRef]
- Houry, W.A.; Bertrand, E.; Coulombe, B. The PAQosome, an R2TP-Based Chaperone for Quaternary Structure Formation. Trends Biochem. Sci. 2018, 43, 4–9. [Google Scholar] [CrossRef] [PubMed]
- Patel-King, R.S.; King, S.M. A prefoldin-associated WD-repeat protein (WDR92) is required for the correct architectural assembly of motile cilia. Mol. Biol. Cell 2016, 27, 1204–1209. [Google Scholar] [CrossRef] [PubMed]
- Lynham, J.; Houry, W.A. The Multiple Functions of the PAQosome: An R2TP- and URI1 Prefoldin-Based Chaperone Complex. Adv. Exp. Med. Biol. 2018, 1106, 37–72. [Google Scholar] [CrossRef] [PubMed]
- Miron-Garcia, M.C.; Garrido-Godino, A.I.; Garcia-Molinero, V.; Hernandez-Torres, F.; Rodriguez-Navarro, S.; Navarro, F. The prefoldin bud27 mediates the assembly of the eukaryotic RNA polymerases in an rpb5-dependent manner. PLoS Genet. 2013, 9, e1003297. [Google Scholar] [CrossRef] [PubMed]
- Mita, P.; Savas, J.N.; Ha, S.; Djouder, N.; Yates, J.R., 3rd; Logan, S.K. Analysis of URI nuclear interaction with RPB5 and components of the R2TP/prefoldin-like complex. PLoS ONE 2013, 8, e63879. [Google Scholar] [CrossRef] [PubMed]
- Seraphim, T.V.; Nano, N.; Cheung, Y.W.S.; Aluksanasuwan, S.; Colleti, C.; Mao, Y.Q.; Bhandari, V.; Young, G.; Holl, L.; Phanse, S.; et al. Assembly principles of the human R2TP chaperone complex reveal the presence of R2T and R2P complexes. Structure 2022, 30, 156–171.e112. [Google Scholar] [CrossRef] [PubMed]
- Henri, J.; Chagot, M.E.; Bourguet, M.; Abel, Y.; Terral, G.; Maurizy, C.; Aigueperse, C.; Georgescauld, F.; Vandermoere, F.; Saint-Fort, R.; et al. Deep Structural Analysis of RPAP3 and PIH1D1, Two Components of the HSP90 Co-chaperone R2TP Complex. Structure 2018, 26, 1196–1209.e1198. [Google Scholar] [CrossRef] [PubMed]
- Martino, F.; Pal, M.; Munoz-Hernandez, H.; Rodriguez, C.F.; Nunez-Ramirez, R.; Gil-Carton, D.; Degliesposti, G.; Skehel, J.M.; Roe, S.M.; Prodromou, C.; et al. RPAP3 provides a flexible scaffold for coupling HSP90 to the human R2TP co-chaperone complex. Nat. Commun. 2018, 9, 1501. [Google Scholar] [CrossRef]
- Horejsi, Z.; Stach, L.; Flower, T.G.; Joshi, D.; Flynn, H.; Skehel, J.M.; O’Reilly, N.J.; Ogrodowicz, R.W.; Smerdon, S.J.; Boulton, S.J. Phosphorylation-dependent PIH1D1 interactions define substrate specificity of the R2TP cochaperone complex. Cell Rep. 2014, 7, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Pal, M.; Morgan, M.; Phelps, S.E.; Roe, S.M.; Parry-Morris, S.; Downs, J.A.; Polier, S.; Pearl, L.H.; Prodromou, C. Structural basis for phosphorylation-dependent recruitment of Tel2 to Hsp90 by Pih1. Structure 2014, 22, 805–818. [Google Scholar] [CrossRef]
- Munoz-Hernandez, H.; Pal, M.; Rodriguez, C.F.; Fernandez-Leiro, R.; Prodromou, C.; Pearl, L.H.; Llorca, O. Structural mechanism for regulation of the AAA-ATPases RUVBL1-RUVBL2 in the R2TP co-chaperone revealed by cryo-EM. Sci. Adv. 2019, 5, eaaw1616. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, B.; Ugwu, F.; Zhao, R.; Orti, L.; Makhnevych, T.; Pineda-Lucena, A.; Houry, W.A. Structure of minimal tetratricopeptide repeat domain protein Tah1 reveals mechanism of its interaction with Pih1 and Hsp90. J. Biol. Chem. 2012, 287, 5698–5709. [Google Scholar] [CrossRef]
- Millson, S.H.; Vaughan, C.K.; Zhai, C.; Ali, M.M.; Panaretou, B.; Piper, P.W.; Pearl, L.H.; Prodromou, C. Chaperone ligand-discrimination by the TPR-domain protein Tah1. Biochem. J. 2008, 413, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Back, R.; Dominguez, C.; Rothe, B.; Bobo, C.; Beaufils, C.; Morera, S.; Meyer, P.; Charpentier, B.; Branlant, C.; Allain, F.H.; et al. High-resolution structural analysis shows how Tah1 tethers Hsp90 to the R2TP complex. Structure 2013, 21, 1834–1847. [Google Scholar] [CrossRef]
- Rivera-Calzada, A.; Pal, M.; Munoz-Hernandez, H.; Luque-Ortega, J.R.; Gil-Carton, D.; Degliesposti, G.; Skehel, J.M.; Prodromou, C.; Pearl, L.H.; Llorca, O. The Structure of the R2TP Complex Defines a Platform for Recruiting Diverse Client Proteins to the HSP90 Molecular Chaperone System. Structure 2017, 25, 1145–1152.e1144. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Yu, G.; He, H.; Zhao, Y.; Liu, P.; Marshall, A.G.; Demeler, B.; Stagg, S.M.; Li, H. Pih1p-Tah1p Puts a Lid on Hexameric AAA+ ATPases Rvb1/2p. Structure 2017, 25, 1519–1529.e1514. [Google Scholar] [CrossRef]
- Filipowicz, W.; Pelczar, P.; Pogacic, V.; Dragon, F. Structure and biogenesis of small nucleolar RNAs acting as guides for ribosomal RNA modification. Acta Biochim. Pol. 1999, 46, 377–389. [Google Scholar] [CrossRef]
- Borovjagin, A.V.; Gerbi, S.A. U3 small nucleolar RNA is essential for cleavage at sites 1, 2 and 3 in pre-rRNA and determines which rRNA processing pathway is taken in Xenopus oocytes. J. Mol. Biol. 1999, 286, 1347–1363. [Google Scholar] [CrossRef] [PubMed]
- Dutca, L.M.; Gallagher, J.E.; Baserga, S.J. The initial U3 snoRNA: Pre-rRNA base pairing interaction required for pre-18S rRNA folding revealed by in vivo chemical probing. Nucleic Acids Res. 2011, 39, 5164–5180. [Google Scholar] [CrossRef]
- Jady, B.E.; Kiss, T. A small nucleolar guide RNA functions both in 2’-O-ribose methylation and pseudouridylation of the U5 spliceosomal RNA. EMBO J. 2001, 20, 541–551. [Google Scholar] [CrossRef] [PubMed]
- Tycowski, K.T.; You, Z.H.; Graham, P.J.; Steitz, J.A. Modification of U6 spliceosomal RNA is guided by other small RNAs. Mol. Cell 1998, 2, 629–638. [Google Scholar] [CrossRef]
- Ganot, P.; Bortolin, M.-L.; Kiss, T. Site-Specific Pseudouridine Formation in Preribosomal RNA Is Guided by Small Nucleolar RNAs. Cell 1997, 89, 799–809. [Google Scholar] [CrossRef]
- Aittaleb, M.; Rashid, R.; Chen, Q.; Palmer, J.R.; Daniels, C.J.; Li, H. Structure and function of archaeal box C/D sRNP core proteins. Nat. Struct. Biol. 2003, 10, 256–263. [Google Scholar] [CrossRef] [PubMed]
- Cahill, N.M.; Friend, K.; Speckmann, W.; Li, Z.H.; Terns, R.M.; Terns, M.P.; Steitz, J.A. Site-specific cross-linking analyses reveal an asymmetric protein distribution for a box C/D snoRNP. EMBO J. 2002, 21, 3816–3828. [Google Scholar] [CrossRef]
- Lin, J.; Lai, S.; Jia, R.; Xu, A.; Zhang, L.; Lu, J.; Ye, K. Structural basis for site-specific ribose methylation by box C/D RNA protein complexes. Nature 2011, 469, 559–563. [Google Scholar] [CrossRef]
- Qu, G.; van Nues, R.W.; Watkins, N.J.; Maxwell, E.S. The spatial-functional coupling of box C/D and C’/D’ RNPs is an evolutionarily conserved feature of the eukaryotic box C/D snoRNP nucleotide modification complex. Mol. Cell Biol. 2011, 31, 365–374. [Google Scholar] [CrossRef]
- Nottrott, S.; Hartmuth, K.; Fabrizio, P.; Urlaub, H.; Vidovic, I.; Ficner, R.; Luhrmann, R. Functional interaction of a novel 15.5kD [U4/U6.U5] tri-snRNP protein with the 5’ stem-loop of U4 snRNA. EMBO J. 1999, 18, 6119–6133. [Google Scholar] [CrossRef]
- Watkins, N.J.; Segault, V.; Charpentier, B.; Nottrott, S.; Fabrizio, P.; Bachi, A.; Wilm, M.; Rosbash, M.; Branlant, C.; Luhrmann, R. A common core RNP structure shared between the small nucleoar box C/D RNPs and the spliceosomal U4 snRNP. Cell 2000, 103, 457–466. [Google Scholar] [CrossRef]
- Watkins, N.J.; Dickmanns, A.; Luhrmann, R. Conserved stem II of the box C/D motif is essential for nucleolar localization and is required, along with the 15.5K protein, for the hierarchical assembly of the box C/D snoRNP. Mol. Cell Biol. 2002, 22, 8342–8352. [Google Scholar] [CrossRef]
- Gautier, T.; Berges, T.; Tollervey, D.; Hurt, E. Nucleolar KKE/D repeat proteins Nop56p and Nop58p interact with Nop1p and are required for ribosome biogenesis. Mol. Cell Biol. 1997, 17, 7088–7098. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Li, P.; Dybkov, O.; Nottrott, S.; Hartmuth, K.; Luhrmann, R.; Carlomagno, T.; Wahl, M.C. Binding of the human Prp31 Nop domain to a composite RNA-protein platform in U4 snRNP. Science 2007, 316, 115–120. [Google Scholar] [CrossRef]
- Stebbins, C.E.; Russo, A.A.; Schneider, C.; Rosen, N.; Hartl, F.U.; Pavletich, N.P. Crystal structure of an Hsp90-geldanamycin complex: Targeting of a protein chaperone by an antitumor agent. Cell 1997, 89, 239–250. [Google Scholar] [CrossRef]
- Abel, Y.; Paiva, A.C.F.; Bizarro, J.; Chagot, M.E.; Santo, P.E.; Robert, M.C.; Quinternet, M.; Vandermoere, F.; Sousa, P.M.F.; Fort, P.; et al. NOPCHAP1 is a PAQosome cofactor that helps loading NOP58 on RUVBL1/2 during box C/D snoRNP biogenesis. Nucleic Acids Res. 2021, 49, 1094–1113. [Google Scholar] [CrossRef]
- Berry, M.J.; Banu, L.; Chen, Y.Y.; Mandel, S.J.; Kieffer, J.D.; Harney, J.W.; Larsen, P.R. Recognition of UGA as a selenocysteine codon in type I deiodinase requires sequences in the 3′ untranslated region. Nature 1991, 353, 273–276. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.R.; Kuhn, J.F.; Shanab, G.M.; Maxwell, E.S. Box C/D snoRNA-associated proteins: Two pairs of evolutionarily ancient proteins and possible links to replication and transcription. RNA 2000, 6, 861–879. [Google Scholar] [CrossRef]
- Watkins, N.J.; Lemm, I.; Ingelfinger, D.; Schneider, C.; Hossbach, M.; Urlaub, H.; Luhrmann, R. Assembly and maturation of the U3 snoRNP in the nucleoplasm in a large dynamic multiprotein complex. Mol. Cell 2004, 16, 789–798. [Google Scholar] [CrossRef] [PubMed]
- Watkins, N.J.; Lemm, I.; Luhrmann, R. Involvement of nuclear import and export factors in U8 box C/D snoRNP biogenesis. Mol. Cell Biol. 2007, 27, 7018–7027. [Google Scholar] [CrossRef][Green Version]
- Bizarro, J.; Charron, C.; Boulon, S.; Westman, B.; Pradet-Balade, B.; Vandermoere, F.; Chagot, M.E.; Hallais, M.; Ahmad, Y.; Leonhardt, H.; et al. Proteomic and 3D structure analyses highlight the C/D box snoRNP assembly mechanism and its control. J. Cell Biol. 2014, 207, 463–480. [Google Scholar] [CrossRef] [PubMed]
- McKeegan, K.S.; Debieux, C.M.; Watkins, N.J. Evidence that the AAA+ proteins TIP48 and TIP49 bridge interactions between 15.5K and the related NOP56 and NOP58 proteins during box C/D snoRNP biogenesis. Mol. Cell Biol. 2009, 29, 4971–4981. [Google Scholar] [CrossRef]
- McKeegan, K.S.; Debieux, C.M.; Boulon, S.; Bertrand, E.; Watkins, N.J. A dynamic scaffold of pre-snoRNP factors facilitates human box C/D snoRNP assembly. Mol. Cell Biol. 2007, 27, 6782–6793. [Google Scholar] [CrossRef]
- Sardiu, M.E.; Cai, Y.; Jin, J.; Swanson, S.K.; Conaway, R.C.; Conaway, J.W.; Florens, L.; Washburn, M.P. Probabilistic assembly of human protein interaction networks from label-free quantitative proteomics. Proc. Natl. Acad. Sci. USA 2008, 105, 1454–1459. [Google Scholar] [CrossRef] [PubMed]
- Cloutier, P.; Poitras, C.; Durand, M.; Hekmat, O.; Fiola-Masson, E.; Bouchard, A.; Faubert, D.; Chabot, B.; Coulombe, B. R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein. Nat. Commun. 2017, 8, 15615. [Google Scholar] [CrossRef] [PubMed]
- Malinová, A.; Cvačková, Z.; Matějů, D.; Hořejší, Z.; Abéza, C.; Vandermoere, F.; Bertrand, E.; Staněk, D.; Verheggen, C. Assembly of the U5 snRNP component PRPF8 is controlled by the HSP90/R2TP chaperones. J. Cell Biol. 2017, 216, 1579–1596. [Google Scholar] [CrossRef] [PubMed]
- Rothe, B.; Saliou, J.M.; Quinternet, M.; Back, R.; Tiotiu, D.; Jacquemin, C.; Loegler, C.; Schlotter, F.; Pena, V.; Eckert, K.; et al. Protein Hit1, a novel box C/D snoRNP assembly factor, controls cellular concentration of the scaffolding protein Rsa1 by direct interaction. Nucleic Acids Res. 2014, 42, 10731–10747. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Meier, U.T. Architecture and assembly of mammalian H/ACA small nucleolar and telomerase ribonucleoproteins. EMBO J. 2004, 23, 1857–1867. [Google Scholar] [CrossRef]
- Dez, C.; Henras, A.; Faucon, B.; Lafontaine, D.; Caizergues-Ferrer, M.; Henry, Y. Stable expression in yeast of the mature form of human telomerase RNA depends on its association with the box H/ACA small nucleolar RNP proteins Cbf5p, Nhp2p and Nop10p. Nucleic Acids Res. 2001, 29, 598–603. [Google Scholar] [CrossRef]
- Girard, J.P.; Lehtonen, H.; Caizergues-Ferrer, M.; Amalric, F.; Tollervey, D.; Lapeyre, B. GAR1 is an essential small nucleolar RNP protein required for pre-rRNA processing in yeast. EMBO J. 1992, 11, 673–682. [Google Scholar] [CrossRef] [PubMed]
- Henras, A.; Henry, Y.; Bousquet-Antonelli, C.; Noaillac-Depeyre, J.; Gelugne, J.P.; Caizergues-Ferrer, M. Nhp2p and Nop10p are essential for the function of H/ACA snoRNPs. EMBO J. 1998, 17, 7078–7090. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Middleton, K.; Yoon, H.J.; Fouquet, C.; Carbon, J. An essential yeast protein, CBF5p, binds in vitro to centromeres and microtubules. Mol. Cell Biol. 1993, 13, 4884–4893. [Google Scholar] [CrossRef] [PubMed]
- Kolodrubetz, D.; Burgum, A. Sequence and genetic analysis of NHP2: A moderately abundant high mobility group-like nuclear protein with an essential function in Saccharomyces cerevisiae. Yeast 1991, 7, 79–90. [Google Scholar] [CrossRef]
- Grozdanov, P.N.; Roy, S.; Kittur, N.; Meier, U.T. SHQ1 is required prior to NAF1 for assembly of H/ACA small nucleolar and telomerase RNPs. RNA 2009, 15, 1188–1197. [Google Scholar] [CrossRef] [PubMed]
- Hoareau-Aveilla, C.; Bonoli, M.; Caizergues-Ferrer, M.; Henry, Y. hNaf1 is required for accumulation of human box H/ACA snoRNPs, scaRNPs, and telomerase. RNA 2006, 12, 832–840. [Google Scholar] [CrossRef] [PubMed]
- Leulliot, N.; Godin, K.S.; Hoareau-Aveilla, C.; Quevillon-Cheruel, S.; Varani, G.; Henry, Y.; Van Tilbeurgh, H. The box H/ACA RNP assembly factor Naf1p contains a domain homologous to Gar1p mediating its interaction with Cbf5p. J. Mol. Biol. 2007, 371, 1338–1353. [Google Scholar] [CrossRef]
- Darzacq, X.; Kittur, N.; Roy, S.; Shav-Tal, Y.; Singer, R.H.; Meier, U.T. Stepwise RNP assembly at the site of H/ACA RNA transcription in human cells. J. Cell Biol. 2006, 173, 207–218. [Google Scholar] [CrossRef]
- Walbott, H.; Machado-Pinilla, R.; Liger, D.; Blaud, M.; Rety, S.; Grozdanov, P.N.; Godin, K.; van Tilbeurgh, H.; Varani, G.; Meier, U.T.; et al. The H/ACA RNP assembly factor SHQ1 functions as an RNA mimic. Genes Dev. 2011, 25, 2398–2408. [Google Scholar] [CrossRef] [PubMed]
- Holt, S.E.; Aisner, D.L.; Baur, J.; Tesmer, V.M.; Dy, M.; Ouellette, M.; Trager, J.B.; Morin, G.B.; Toft, D.O.; Shay, J.W.; et al. Functional requirement of p23 and Hsp90 in telomerase complexes. Genes Dev. 1999, 13, 817–826. [Google Scholar] [CrossRef] [PubMed]
- Godin, K.S.; Walbott, H.; Leulliot, N.; van Tilbeurgh, H.; Varani, G. The box H/ACA snoRNP assembly factor Shq1p is a chaperone protein homologous to Hsp90 cochaperones that binds to the Cbf5p enzyme. J. Mol. Biol. 2009, 390, 231–244. [Google Scholar] [CrossRef] [PubMed]
- Morin, G.B. The human telomere terminal transferase enzyme is a ribonucleoprotein that synthesizes TTAGGG repeats. Cell 1989, 59, 521–529. [Google Scholar] [CrossRef]
- Keppler, B.R.; Grady, A.T.; Jarstfer, M.B. The biochemical role of the heat shock protein 90 chaperone complex in establishing human telomerase activity. J. Biol. Chem. 2006, 281, 19840–19848. [Google Scholar] [CrossRef] [PubMed]
- Forsythe, H.L.; Jarvis, J.L.; Turner, J.W.; Elmore, L.W.; Holt, S.E. Stable association of hsp90 and p23, but Not hsp70, with active human telomerase. J. Biol. Chem. 2001, 276, 15571–15574. [Google Scholar] [CrossRef] [PubMed]
- Kim, R.H.; Kim, R.; Chen, W.; Hu, S.; Shin, K.H.; Park, N.H.; Kang, M.K. Association of hsp90 to the hTERT promoter is necessary for hTERT expression in human oral cancer cells. Carcinogenesis 2008, 29, 2425–2431. [Google Scholar] [CrossRef] [PubMed]
- Chiu, W.T.; Shen, S.C.; Yang, L.Y.; Chow, J.M.; Wu, C.Y.; Chen, Y.C. Inhibition of HSP90-dependent telomerase activity in amyloid beta-induced apoptosis of cerebral endothelial cells. J. Cell Physiol. 2011, 226, 2041–2051. [Google Scholar] [CrossRef] [PubMed]
- Venteicher, A.S.; Meng, Z.; Mason, P.J.; Veenstra, T.D.; Artandi, S.E. Identification of ATPases pontin and reptin as telomerase components essential for holoenzyme assembly. Cell 2008, 132, 945–957. [Google Scholar] [CrossRef] [PubMed]
- Rappsilber, J.; Ryder, U.; Lamond, A.I.; Mann, M. Large-scale proteomic analysis of the human spliceosome. Genome Res. 2002, 12, 1231–1245. [Google Scholar] [CrossRef] [PubMed]
- Fabrizio, P.; Dannenberg, J.; Dube, P.; Kastner, B.; Stark, H.; Urlaub, H.; Luhrmann, R. The evolutionarily conserved core design of the catalytic activation step of the yeast spliceosome. Mol. Cell 2009, 36, 593–608. [Google Scholar] [CrossRef] [PubMed]
- Bizarro, J.; Dodre, M.; Huttin, A.; Charpentier, B.; Schlotter, F.; Branlant, C.; Verheggen, C.; Massenet, S.; Bertrand, E. NUFIP and the HSP90/R2TP chaperone bind the SMN complex and facilitate assembly of U4-specific proteins. Nucleic Acids Res. 2015, 43, 8973–8989. [Google Scholar] [CrossRef]
- Nottrott, S.; Urlaub, H.; Luhrmann, R. Hierarchical, clustered protein interactions with U4/U6 snRNA: A biochemical role for U4/U6 proteins. EMBO J. 2002, 21, 5527–5538. [Google Scholar] [CrossRef] [PubMed]
- Deery, E.C.; Vithana, E.N.; Newbold, R.J.; Gallon, V.A.; Bhattacharya, S.S.; Warren, M.J.; Hunt, D.M.; Wilkie, S.E. Disease mechanism for retinitis pigmentosa (RP11) caused by mutations in the splicing factor gene PRPF31. Hum. Mol. Genet. 2002, 11, 3209–3219. [Google Scholar] [CrossRef] [PubMed]
- Serna, M.; Gonzalez-Corpas, A.; Cabezudo, S.; Lopez-Perrote, A.; Degliesposti, G.; Zarzuela, E.; Skehel, J.M.; Munoz, J.; Llorca, O. CryoEM of RUVBL1-RUVBL2-ZNHIT2, a complex that interacts with pre-mRNA-processing-splicing factor 8. Nucleic Acids Res. 2022, 50, 1128–1146. [Google Scholar] [CrossRef] [PubMed]
- Mir, R.A.; Lovelace, J.; Schafer, N.P.; Simone, P.D.; Kellezi, A.; Kolar, C.; Spagnol, G.; Sorgen, P.L.; Band, H.; Band, V.; et al. Biophysical characterization and modeling of human Ecdysoneless (ECD) protein supports a scaffolding function. AIMS Biophys 2016, 3, 195–208. [Google Scholar] [CrossRef] [PubMed]
- Cloutier, P.; Coulombe, B. New insights into the biogenesis of nuclear RNA polymerases? Biochem. Cell Biol. 2010, 88, 211–221. [Google Scholar] [CrossRef]
- Forget, D.; Lacombe, A.A.; Cloutier, P.; Al-Khoury, R.; Bouchard, A.; Lavallee-Adam, M.; Faubert, D.; Jeronimo, C.; Blanchette, M.; Coulombe, B. The protein interaction network of the human transcription machinery reveals a role for the conserved GTPase RPAP4/GPN1 and microtubule assembly in nuclear import and biogenesis of RNA polymerase II. Mol. Cell Proteom. 2010, 9, 2827–2839. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, V.T.; Giannoni, F.; Dubois, M.F.; Seo, S.J.; Vigneron, M.; Kedinger, C.; Bensaude, O. In vivo degradation of RNA polymerase II largest subunit triggered by alpha-amanitin. Nucleic Acids Res. 1996, 24, 2924–2929. [Google Scholar] [CrossRef] [PubMed]
- Maurizy, C.; Abeza, C.; Lemmers, B.; Gabola, M.; Longobardi, C.; Pinet, V.; Ferrand, M.; Paul, C.; Bremond, J.; Langa, F.; et al. The HSP90/R2TP assembly chaperone promotes cell proliferation in the intestinal epithelium. Nat. Commun. 2021, 12, 4810. [Google Scholar] [CrossRef]
- Frischknecht, L.; Britschgi, C.; Galliker, P.; Christinat, Y.; Vichalkovski, A.; Gstaiger, M.; Kovacs, W.J.; Krek, W. BRAF inhibition sensitizes melanoma cells to alpha-amanitin via decreased RNA polymerase II assembly. Sci. Rep. 2019, 9, 7779. [Google Scholar] [CrossRef]
- Perry, J.; Kleckner, N. The ATRs, ATMs, and TORs are giant HEAT repeat proteins. Cell 2003, 112, 151–155. [Google Scholar] [CrossRef]
- Lavin, M.F.; Khanna, K.K.; Beamish, H.; Spring, K.; Watters, D.; Shiloh, Y. Relationship of the ataxia-telangiectasia protein ATM to phosphoinositide 3-kinase. Trends Biochem. Sci. 1995, 20, 382–383. [Google Scholar] [CrossRef]
- Izumi, N.; Yamashita, A.; Hirano, H.; Ohno, S. Heat shock protein 90 regulates phosphatidylinositol 3-kinase-related protein kinase family proteins together with the RUVBL1/2 and Tel2-containing co-factor complex. Cancer Sci. 2012, 103, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Toullec, D.; Elias-Villalobos, A.; Faux, C.; Noly, A.; Lledo, G.; Seveno, M.; Helmlinger, D. The Hsp90 cochaperone TTT promotes cotranslational maturation of PIKKs prior to complex assembly. Cell Rep. 2021, 37, 109867. [Google Scholar] [CrossRef]
- Hayashi, T.; Hatanaka, M.; Nagao, K.; Nakaseko, Y.; Kanoh, J.; Kokubu, A.; Ebe, M.; Yanagida, M. Rapamycin sensitivity of the Schizosaccharomyces pombe tor2 mutant and organization of two highly phosphorylated TOR complexes by specific and common subunits. Genes Cells 2007, 12, 1357–1370. [Google Scholar] [CrossRef] [PubMed]
- Shevchenko, A.; Roguev, A.; Schaft, D.; Buchanan, L.; Habermann, B.; Sakalar, C.; Thomas, H.; Krogan, N.J.; Shevchenko, A.; Stewart, A.F. Chromatin Central: Towards the comparative proteome by accurate mapping of the yeast proteomic environment. Genome Biol. 2008, 9, R167. [Google Scholar] [CrossRef] [PubMed]
- Hurov, K.E.; Cotta-Ramusino, C.; Elledge, S.J. A genetic screen identifies the Triple T complex required for DNA damage signaling and ATM and ATR stability. Genes Dev. 2010, 24, 1939–1950. [Google Scholar] [CrossRef]
- Kim, Y.; Park, J.; Joo, S.Y.; Kim, B.G.; Jo, A.; Lee, H.; Cho, Y. Structure of the Human TELO2-TTI1-TTI2 Complex. J. Mol. Biol. 2022, 434, 167370. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.; Alpi, A.; Hengartner, M.O.; Gartner, A.C. Elegans RAD-5/CLK-2 defines a new DNA damage checkpoint protein. Curr. Biol. 2001, 11, 1934–1944. [Google Scholar] [CrossRef]
- Anderson, C.M.; Korkin, D.; Smith, D.L.; Makovets, S.; Seidel, J.J.; Sali, A.; Blackburn, E.H. Tel2 mediates activation and localization of ATM/Tel1 kinase to a double-strand break. Genes Dev. 2008, 22, 854–859. [Google Scholar] [CrossRef]
- Goto, G.H.; Ogi, H.; Biswas, H.; Ghosh, A.; Tanaka, S.; Sugimoto, K. Two separate pathways regulate protein stability of ATM/ATR-related protein kinases Mec1 and Tel1 in budding yeast. PLoS Genet. 2017, 13, e1006873. [Google Scholar] [CrossRef] [PubMed]
- Rao, F.; Cha, J.; Xu, J.; Xu, R.; Vandiver, M.S.; Tyagi, R.; Tokhunts, R.; Koldobskiy, M.A.; Fu, C.; Barrow, R.; et al. Inositol pyrophosphates mediate the DNA-PK/ATM-p53 cell death pathway by regulating CK2 phosphorylation of Tti1/Tel2. Mol. Cell 2014, 54, 119–132. [Google Scholar] [CrossRef] [PubMed]
- Shikata, M.; Ishikawa, F.; Kanoh, J. Tel2 is required for activation of the Mrc1-mediated replication checkpoint. J. Biol. Chem. 2007, 282, 5346–5355. [Google Scholar] [CrossRef] [PubMed]
- Takai, H.; Wang, R.C.; Takai, K.K.; Yang, H.; de Lange, T. Tel2 regulates the stability of PI3K-related protein kinases. Cell 2007, 131, 1248–1259. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.J.; Khan, S.; Didier, A.C.; Wozniak, M.; Liu, Y.; Singh, A.; Nakamura, T.M. A tel2 Mutation That Destabilizes the Tel2-Tti1-Tti2 Complex Eliminates Rad3(ATR) Kinase Signaling in the DNA Replication Checkpoint and Leads to Telomere Shortening in Fission Yeast. Mol. Cell Biol. 2019, 39, e00175-19. [Google Scholar] [CrossRef] [PubMed]
- Brown, M.C.; Gromeier, M. MNK controls mTORC1: Substrate association through regulation of TELO2 binding with mTORC1. Cell Rep. 2017, 18, 1444–1457. [Google Scholar] [CrossRef] [PubMed]
- David-Morrison, G.; Xu, Z.; Rui, Y.N.; Charng, W.L.; Jaiswal, M.; Yamamoto, S.; Xiong, B.; Zhang, K.; Sandoval, H.; Duraine, L.; et al. WAC Regulates mTOR Activity by Acting as an Adaptor for the TTT and Pontin/Reptin Complexes. Dev. Cell 2016, 36, 139–151. [Google Scholar] [CrossRef]
- Fernández-Sáiz, V.; Targosz, B.-S.; Lemeer, S.; Eichner, R.; Langer, C.; Bullinger, L.; Reiter, C.; Slotta-Huspenina, J.; Schroeder, S.; Knorn, A.-M.; et al. SCFFbxo9 and CK2 direct the cellular response to growth factor withdrawal via Tel2/Tti1 degradation and promote survival in multiple myeloma. Nat. Cell Biol. 2012, 15, 72. [Google Scholar] [CrossRef]
- Hoffman, K.S.; Duennwald, M.L.; Karagiannis, J.; Genereaux, J.; McCarton, A.S.; Brandl, C.J. Saccharomyces cerevisiae Tti2 Regulates PIKK Proteins and Stress Response. G3 2016, 6, 1649–1659. [Google Scholar] [CrossRef]
- Kaizuka, T.; Hara, T.; Oshiro, N.; Kikkawa, U.; Yonezawa, K.; Takehana, K.; Iemura, S.; Natsume, T.; Mizushima, N. Tti1 and Tel2 are critical factors in mammalian target of rapamycin complex assembly. J. Biol. Chem. 2010, 285, 20109–20116. [Google Scholar] [CrossRef]
- Kim, S.G.; Hoffman, G.R.; Poulogiannis, G.; Buel, G.R.; Jang, Y.J.; Lee, K.W.; Kim, B.Y.; Erikson, R.L.; Cantley, L.C.; Choo, A.Y.; et al. Metabolic stress controls mTORC1 lysosomal localization and dimerization by regulating the TTT-RUVBL1/2 complex. Mol. Cell 2013, 49, 172–185. [Google Scholar] [CrossRef]
- Rozario, D.; Siede, W. Saccharomyces cerevisiae Tel2 plays roles in TORC signaling and telomere maintenance that can be mutationally separated. Biochem. Biophys. Res. Commun. 2012, 417, 1182–1187. [Google Scholar] [CrossRef]
- Ahn, S.; Kim, J.; Hwang, J. CK2-mediated TEL2 phosphorylation augments nonsense-mediated mRNA decay (NMD) by increase of SMG1 stability. Biochim. Biophys. Acta 2013, 1829, 1047–1055. [Google Scholar] [CrossRef]
- Guo, Y.; Tocchini, C.; Ciosk, R. CLK-2/TEL2 is a conserved component of the nonsense-mediated mRNA decay pathway. PLoS ONE 2021, 16, e0244505. [Google Scholar] [CrossRef] [PubMed]
- Detilleux, D.; Raynaud, P.; Pradet-Balade, B.; Helmlinger, D. The TRRAP transcription cofactor represses interferon-stimulated genes in colorectal cancer cells. Elife 2022, 11, e69705. [Google Scholar] [CrossRef] [PubMed]
- Elias-Villalobos, A.; Toullec, D.; Faux, C.; Seveno, M.; Helmlinger, D. Chaperone-mediated ordered assembly of the SAGA and NuA4 transcription co-activator complexes in yeast. Nat. Commun. 2019, 10, 5237. [Google Scholar] [CrossRef]
- Takai, H.; Xie, Y.; de Lange, T.; Pavletich, N.P. Tel2 structure and function in the Hsp90-dependent maturation of mTOR and ATR complexes. Genes Dev. 2010, 24, 2019–2030. [Google Scholar] [CrossRef] [PubMed]
- Izumi, N.; Yamashita, A.; Iwamatsu, A.; Kurata, R.; Nakamura, H.; Saari, B.; Hirano, H.; Anderson, P.; Ohno, S. AAA+ proteins RUVBL1 and RUVBL2 coordinate PIKK activity and function in nonsense-mediated mRNA decay. Sci. Signal. 2010, 3, ra27. [Google Scholar] [CrossRef] [PubMed]
- Zou, L.; Elledge, S.J. Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science 2003, 300, 1542–1548. [Google Scholar] [CrossRef]
- Foster, K.G.; Fingar, D.C. Mammalian target of rapamycin (mTOR): Conducting the cellular signaling symphony. J. Biol. Chem. 2010, 285, 14071–14077. [Google Scholar] [CrossRef]
- Li, J.; Csibi, A.; Yang, S.; Hoffman, G.R.; Li, C.; Zhang, E.; Yu, J.J.; Blenis, J. Synthetic lethality of combined glutaminase and Hsp90 inhibition in mTORC1-driven tumor cells. Proc. Natl. Acad. Sci. USA 2015, 112, E21–E29. [Google Scholar] [CrossRef] [PubMed]
- Uziel, T.; Lerenthal, Y.; Moyal, L.; Andegeko, Y.; Mittelman, L.; Shiloh, Y. Requirement of the MRN complex for ATM activation by DNA damage. EMBO J. 2003, 22, 5612–5621. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Paull, T.T. ATM activation by DNA double-strand breaks through the Mre11-Rad50-Nbs1 complex. Science 2005, 308, 551–554. [Google Scholar] [CrossRef] [PubMed]
- Singleton, B.K.; Torres-Arzayus, M.I.; Rottinghaus, S.T.; Taccioli, G.E.; Jeggo, P.A. The C terminus of Ku80 activates the DNA-dependent protein kinase catalytic subunit. Mol. Cell Biol. 1999, 19, 3267–3277. [Google Scholar] [CrossRef]
- Sibanda, B.L.; Chirgadze, D.Y.; Ascher, D.B.; Blundell, T.L. DNA-PKcs structure suggests an allosteric mechanism modulating DNA double-strand break repair. Science 2017, 355, 520–524. [Google Scholar] [CrossRef]
- Pal, M.; Munoz-Hernandez, H.; Bjorklund, D.; Zhou, L.; Degliesposti, G.; Skehel, J.M.; Hesketh, E.L.; Thompson, R.F.; Pearl, L.H.; Llorca, O.; et al. Structure of the TELO2-TTI1-TTI2 complex and its function in TOR recruitment to the R2TP chaperone. Cell Rep. 2021, 36, 109317. [Google Scholar] [CrossRef]
- Shin, S.H.; Lee, J.S.; Zhang, J.M.; Choi, S.; Boskovic, Z.V.; Zhao, R.; Song, M.; Wang, R.; Tian, J.; Lee, M.H.; et al. Synthetic lethality by targeting the RUVBL1/2-TTT complex in mTORC1-hyperactive cancer cells. Sci. Adv. 2020, 6, eaay9131. [Google Scholar] [CrossRef]
- Saez, I.; Gerbracht, J.V.; Koyuncu, S.; Lee, H.J.; Horn, M.; Kroef, V.; Denzel, M.S.; Dieterich, C.; Gehring, N.H.; Vilchez, D. The E3 ubiquitin ligase UBR5 interacts with the H/ACA ribonucleoprotein complex and regulates ribosomal RNA biogenesis in embryonic stem cells. FEBS Lett. 2020, 594, 175–188. [Google Scholar] [CrossRef]
- Erkelenz, S.; Stankovic, D.; Mundorf, J.; Bresser, T.; Claudius, A.K.; Boehm, V.; Gehring, N.H.; Uhlirova, M. Ecd promotes U5 snRNP maturation and Prp8 stability. Nucleic Acids Res. 2021, 49, 1688–1707. [Google Scholar] [CrossRef]
- Buis, J.; Wu, Y.; Deng, Y.; Leddon, J.; Westfield, G.; Eckersdorff, M.; Sekiguchi, J.M.; Chang, S.; Ferguson, D.O. Mre11 nuclease activity has essential roles in DNA repair and genomic stability distinct from ATM activation. Cell 2008, 135, 85–96. [Google Scholar] [CrossRef]
- Milman, N.; Higuchi, E.; Smith, G.R. Meiotic DNA double-strand break repair requires two nucleases, MRN and Ctp1, to produce a single size class of Rec12 (Spo11)-oligonucleotide complexes. Mol. Cell Biol. 2009, 29, 5998–6005. [Google Scholar] [CrossRef]
- You, Z.; Chahwan, C.; Bailis, J.; Hunter, T.; Russell, P. ATM activation and its recruitment to damaged DNA require binding to the C terminus of Nbs1. Mol. Cell Biol. 2005, 25, 5363–5379. [Google Scholar] [CrossRef] [PubMed]
- Zhong, H.; Bryson, A.; Eckersdorff, M.; Ferguson, D.O. Rad50 depletion impacts upon ATR-dependent DNA damage responses. Hum. Mol. Genet. 2005, 14, 2685–2693. [Google Scholar] [CrossRef][Green Version]
- Zhuang, J.; Jiang, G.; Willers, H.; Xia, F. Exonuclease function of human Mre11 promotes deletional nonhomologous end joining. J. Biol. Chem. 2009, 284, 30565–30573. [Google Scholar] [CrossRef] [PubMed]
- Stewart, G.S.; Maser, R.S.; Stankovic, T.; Bressan, D.A.; Kaplan, M.I.; Jaspers, N.G.; Raams, A.; Byrd, P.J.; Petrini, J.H.; Taylor, A.M. The DNA double-strand break repair gene hMRE11 is mutated in individuals with an ataxia-telangiectasia-like disorder. Cell 1999, 99, 577–587. [Google Scholar] [CrossRef]
- Varon, R.; Vissinga, C.; Platzer, M.; Cerosaletti, K.M.; Chrzanowska, K.H.; Saar, K.; Beckmann, G.; Seemanova, E.; Cooper, P.R.; Nowak, N.J.; et al. Nibrin, a novel DNA double-strand break repair protein, is mutated in Nijmegen breakage syndrome. Cell 1998, 93, 467–476. [Google Scholar] [CrossRef]
- Waltes, R.; Kalb, R.; Gatei, M.; Kijas, A.W.; Stumm, M.; Sobeck, A.; Wieland, B.; Varon, R.; Lerenthal, Y.; Lavin, M.F.; et al. Human RAD50 deficiency in a Nijmegen breakage syndrome-like disorder. Am. J. Hum. Genet. 2009, 84, 605–616. [Google Scholar] [CrossRef]
- Paull, T.T.; Gellert, M. The 3’ to 5’ exonuclease activity of Mre 11 facilitates repair of DNA double-strand breaks. Mol. Cell 1998, 1, 969–979. [Google Scholar] [CrossRef]
- Paull, T.T.; Gellert, M. Nbs1 potentiates ATP-driven DNA unwinding and endonuclease cleavage by the Mre11/Rad50 complex. Genes Dev. 1999, 13, 1276–1288. [Google Scholar] [CrossRef]
- Williams, R.S.; Moncalian, G.; Williams, J.S.; Yamada, Y.; Limbo, O.; Shin, D.S.; Groocock, L.M.; Cahill, D.; Hitomi, C.; Guenther, G.; et al. Mre11 dimers coordinate DNA end bridging and nuclease processing in double-strand-break repair. Cell 2008, 135, 97–109. [Google Scholar] [CrossRef] [PubMed]
- von Morgen, P.; Burdova, K.; Flower, T.G.; O’Reilly, N.J.; Boulton, S.J.; Smerdon, S.J.; Macurek, L.; Horejsi, Z. MRE11 stability is regulated by CK2-dependent interaction with R2TP complex. Oncogene 2017, 36, 4943–4950. [Google Scholar] [CrossRef] [PubMed]
- de Jager, M.; van Noort, J.; van Gent, D.C.; Dekker, C.; Kanaar, R.; Wyman, C. Human Rad50/Mre11 is a flexible complex that can tether DNA ends. Mol. Cell 2001, 8, 1129–1135. [Google Scholar] [CrossRef]
- Hopfner, K.P.; Karcher, A.; Shin, D.S.; Craig, L.; Arthur, L.M.; Carney, J.P.; Tainer, J.A. Structural biology of Rad50 ATPase: ATP-driven conformational control in DNA double-strand break repair and the ABC-ATPase superfamily. Cell 2000, 101, 789–800. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, Q.; Wu, D.; Chen, Q.; Gong, G.; He, L.; Wu, X. Lamin-A interacting protein Hsp90 is required for DNA damage repair and chemoresistance of ovarian cancer cells. Cell Death Dis. 2021, 12, 786. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Q.; Chen, C.F.; Li, S.; Chen, Y.; Wang, C.C.; Xiao, J.; Chen, P.L.; Sharp, Z.D.; Lee, W.H. Association of BRCA1 with the hRad50-hMre11-p95 complex and the DNA damage response. Science 1999, 285, 747–750. [Google Scholar] [CrossRef]
- Stecklein, S.R.; Kumaraswamy, E.; Behbod, F.; Wang, W.; Chaguturu, V.; Harlan-Williams, L.M.; Jensen, R.A. BRCA1 and HSP90 cooperate in homologous and non-homologous DNA double-strand-break repair and G2/M checkpoint activation. Proc. Natl. Acad. Sci. USA 2012, 109, 13650–13655. [Google Scholar] [CrossRef]
- Lloyd, J.; Chapman, J.R.; Clapperton, J.A.; Haire, L.F.; Hartsuiker, E.; Li, J.; Carr, A.M.; Jackson, S.P.; Smerdon, S.J. A supramodular FHA/BRCT-repeat architecture mediates Nbs1 adaptor function in response to DNA damage. Cell 2009, 139, 100–111. [Google Scholar] [CrossRef] [PubMed]
- Williams, R.S.; Dodson, G.E.; Limbo, O.; Yamada, Y.; Williams, J.S.; Guenther, G.; Classen, S.; Glover, J.N.; Iwasaki, H.; Russell, P.; et al. Nbs1 flexibly tethers Ctp1 and Mre11-Rad50 to coordinate DNA double-strand break processing and repair. Cell 2009, 139, 87–99. [Google Scholar] [CrossRef]
- Pennisi, R.; Antoccia, A.; Leone, S.; Ascenzi, P.; di Masi, A. Hsp90alpha regulates ATM and NBN functions in sensing and repair of DNA double-strand breaks. FEBS J. 2017, 284, 2378–2395. [Google Scholar] [CrossRef] [PubMed]
- Dote, H.; Burgan, W.E.; Camphausen, K.; Tofilon, P.J. Inhibition of hsp90 compromises the DNA damage response to radiation. Cancer Res. 2006, 66, 9211–9220. [Google Scholar] [CrossRef]
- Elaimy, A.L.; Ahsan, A.; Marsh, K.; Pratt, W.B.; Ray, D.; Lawrence, T.S.; Nyati, M.K. ATM is the primary kinase responsible for phosphorylation of Hsp90alpha after ionizing radiation. Oncotarget 2016, 7, 82450–82457. [Google Scholar] [CrossRef]
- Quanz, M.; Herbette, A.; Sayarath, M.; de Koning, L.; Dubois, T.; Sun, J.S.; Dutreix, M. Heat shock protein 90alpha (Hsp90alpha) is phosphorylated in response to DNA damage and accumulates in repair foci. J. Biol. Chem. 2012, 287, 8803–8815. [Google Scholar] [CrossRef]
- Cheng, A.N.; Fan, C.C.; Lo, Y.K.; Kuo, C.L.; Wang, H.C.; Lien, I.H.; Lin, S.Y.; Chen, C.H.; Jiang, S.S.; Chang, I.S.; et al. Cdc7-Dbf4-mediated phosphorylation of HSP90-S164 stabilizes HSP90-HCLK2-MRN complex to enhance ATR/ATM signaling that overcomes replication stress in cancer. Sci. Rep. 2017, 7, 17024. [Google Scholar] [CrossRef]
- Huang, J.; Manning, B.D. The TSC1-TSC2 complex: A molecular switchboard controlling cell growth. Biochem. J. 2008, 412, 179–190. [Google Scholar] [CrossRef]
- Henske, E.P.; Jozwiak, S.; Kingswood, J.C.; Sampson, J.R.; Thiele, E.A. Tuberous sclerosis complex. Nat. Rev. Dis. Primers 2016, 2, 16035. [Google Scholar] [CrossRef]
- Benvenuto, G.; Li, S.; Brown, S.J.; Braverman, R.; Vass, W.C.; Cheadle, J.P.; Halley, D.J.; Sampson, J.R.; Wienecke, R.; DeClue, J.E. The tuberous sclerosis-1 (TSC1) gene product hamartin suppresses cell growth and augments the expression of the TSC2 product tuberin by inhibiting its ubiquitination. Oncogene 2000, 19, 6306–6316. [Google Scholar] [CrossRef]
- Nakashima, A.; Yoshino, K.; Miyamoto, T.; Eguchi, S.; Oshiro, N.; Kikkawa, U.; Yonezawa, K. Identification of TBC7 having TBC domain as a novel binding protein to TSC1-TSC2 complex. Biochem. Biophys. Res. Commun. 2007, 361, 218–223. [Google Scholar] [CrossRef]
- Sato, N.; Koinuma, J.; Ito, T.; Tsuchiya, E.; Kondo, S.; Nakamura, Y.; Daigo, Y. Activation of an oncogenic TBC1D7 (TBC1 domain family, member 7) protein in pulmonary carcinogenesis. Genes Chromosomes Cancer 2010, 49, 353–367. [Google Scholar] [CrossRef]
- Inoki, K.; Li, Y.; Xu, T.; Guan, K.L. Rheb GTPase is a direct target of TSC2 GAP activity and regulates mTOR signaling. Genes Dev. 2003, 17, 1829–1834. [Google Scholar] [CrossRef]
- Zhang, Y.; Gao, X.; Saucedo, L.J.; Ru, B.; Edgar, B.A.; Pan, D. Rheb is a direct target of the tuberous sclerosis tumour suppressor proteins. Nat. Cell Biol. 2003, 5, 578–581. [Google Scholar] [CrossRef]
- Woodford, M.R.; Sager, R.A.; Marris, E.; Dunn, D.M.; Blanden, A.R.; Murphy, R.L.; Rensing, N.; Shapiro, O.; Panaretou, B.; Prodromou, C.; et al. Tumor suppressor Tsc1 is a new Hsp90 co-chaperone that facilitates folding of kinase and non-kinase clients. EMBO J. 2017, 36, 3650–3665. [Google Scholar] [CrossRef]
- Woodford, M.R.; Hughes, M.; Sager, R.A.; Backe, S.J.; Baker-Williams, A.J.; Bratslavsky, M.S.; Jacob, J.M.; Shapiro, O.; Wong, M.; Bratslavsky, G.; et al. Mutation of the co-chaperone Tsc1 in bladder cancer diminishes Hsp90 acetylation and reduces drug sensitivity and selectivity. Oncotarget 2019, 10, 5824–5834. [Google Scholar] [CrossRef] [PubMed]
- Mrozek, E.M.; Bajaj, V.; Guo, Y.; Malinowska, I.A.; Zhang, J.; Kwiatkowski, D.J. Evaluation of Hsp90 and mTOR inhibitors as potential drugs for the treatment of TSC1/TSC2 deficient cancer. PLoS ONE 2021, 16, e0248380. [Google Scholar] [CrossRef] [PubMed]
- Fliegauf, M.; Benzing, T.; Omran, H. When cilia go bad: Cilia defects and ciliopathies. Nat. Rev. Mol. Cell Biol. 2007, 8, 880–893. [Google Scholar] [CrossRef]
- Gibbons, I.R.; Rowe, A.J. Dynein: A Protein with Adenosine Triphosphatase Activity from Cilia. Science 1965, 149, 424–426. [Google Scholar] [CrossRef]
- Fok, A.K.; Wang, H.; Katayama, A.; Aihara, M.S.; Allen, R.D. 22S axonemal dynein is preassembled and functional prior to being transported to and attached on the axonemes. Cell Motil. Cytoskelet. 1994, 29, 215–224. [Google Scholar] [CrossRef] [PubMed]
- Mitchison, H.M.; Schmidts, M.; Loges, N.T.; Freshour, J.; Dritsoula, A.; Hirst, R.A.; O’Callaghan, C.; Blau, H.; Al Dabbagh, M.; Olbrich, H.; et al. Mutations in axonemal dynein assembly factor DNAAF3 cause primary ciliary dyskinesia. Nat. Genet. 2012, 44, 381–389. [Google Scholar] [CrossRef]
- Diggle, C.P.; Moore, D.J.; Mali, G.; zur Lage, P.; Ait-Lounis, A.; Schmidts, M.; Shoemark, A.; Garcia Munoz, A.; Halachev, M.R.; Gautier, P.; et al. HEATR2 plays a conserved role in assembly of the ciliary motile apparatus. PLoS Genet. 2014, 10, e1004577. [Google Scholar] [CrossRef] [PubMed]
- Duquesnoy, P.; Escudier, E.; Vincensini, L.; Freshour, J.; Bridoux, A.M.; Coste, A.; Deschildre, A.; de Blic, J.; Legendre, M.; Montantin, G.; et al. Loss-of-function mutations in the human ortholog of Chlamydomonas reinhardtii ODA7 disrupt dynein arm assembly and cause primary ciliary dyskinesia. Am. J. Hum. Genet. 2009, 85, 890–896. [Google Scholar] [CrossRef]
- Guo, Z.; Chen, W.; Huang, J.; Wang, L.; Qian, L. Clinical and genetic analysis of patients with primary ciliary dyskinesia caused by novel DNAAF3 mutations. J. Hum. Genet. 2019, 64, 711–719. [Google Scholar] [CrossRef] [PubMed]
- Horani, A.; Druley, T.E.; Zariwala, M.A.; Patel, A.C.; Levinson, B.T.; Van Arendonk, L.G.; Thornton, K.C.; Giacalone, J.C.; Albee, A.J.; Wilson, K.S.; et al. Whole-exome capture and sequencing identifies HEATR2 mutation as a cause of primary ciliary dyskinesia. Am. J. Hum. Genet. 2012, 91, 685–693. [Google Scholar] [CrossRef]
- Moore, D.J.; Onoufriadis, A.; Shoemark, A.; Simpson, M.A.; zur Lage, P.I.; de Castro, S.C.; Bartoloni, L.; Gallone, G.; Petridi, S.; Woollard, W.J.; et al. Mutations in ZMYND10, a gene essential for proper axonemal assembly of inner and outer dynein arms in humans and flies, cause primary ciliary dyskinesia. Am. J. Hum. Genet. 2013, 93, 346–356. [Google Scholar] [CrossRef]
- Olcese, C.; Patel, M.P.; Shoemark, A.; Kiviluoto, S.; Legendre, M.; Williams, H.J.; Vaughan, C.K.; Hayward, J.; Goldenberg, A.; Emes, R.D.; et al. X-linked primary ciliary dyskinesia due to mutations in the cytoplasmic axonemal dynein assembly factor PIH1D3. Nat. Commun. 2017, 8, 14279. [Google Scholar] [CrossRef] [PubMed]
- Omran, H.; Kobayashi, D.; Olbrich, H.; Tsukahara, T.; Loges, N.T.; Hagiwara, H.; Zhang, Q.; Leblond, G.; O’Toole, E.; Hara, C.; et al. Ktu/PF13 is required for cytoplasmic pre-assembly of axonemal dyneins. Nature 2008, 456, 611. [Google Scholar] [CrossRef] [PubMed]
- Paff, T.; Loges, N.T.; Aprea, I.; Wu, K.; Bakey, Z.; Haarman, E.G.; Daniels, J.M.A.; Sistermans, E.A.; Bogunovic, N.; Dougherty, G.W.; et al. Mutations in PIH1D3 Cause X-Linked Primary Ciliary Dyskinesia with Outer and Inner Dynein Arm Defects. Am. J. Hum. Genet. 2017, 100, 160–168. [Google Scholar] [CrossRef] [PubMed]
- Tarkar, A.; Loges, N.T.; Slagle, C.E.; Francis, R.; Dougherty, G.W.; Tamayo, J.V.; Shook, B.; Cantino, M.; Schwartz, D.; Jahnke, C.; et al. DYX1C1 is required for axonemal dynein assembly and ciliary motility. Nat. Genet. 2013, 45, 995–1003. [Google Scholar] [CrossRef]
- Zariwala, M.A.; Gee, H.Y.; Kurkowiak, M.; Al-Mutairi, D.A.; Leigh, M.W.; Hurd, T.W.; Hjeij, R.; Dell, S.D.; Chaki, M.; Dougherty, G.W.; et al. ZMYND10 is mutated in primary ciliary dyskinesia and interacts with LRRC6. Am. J. Hum. Genet. 2013, 93, 336–345. [Google Scholar] [CrossRef] [PubMed]
- Zur Lage, P.; Xi, Z.; Lennon, J.; Hunter, I.; Chan, W.K.; Bolado Carrancio, A.; von Kriegsheim, A.; Jarman, A.P. The Drosophila orthologue of the primary ciliary dyskinesia-associated gene, DNAAF3, is required for axonemal dynein assembly. Biol. Open 2021, 10, bio058812. [Google Scholar] [CrossRef]
- Fabczak, H.; Osinka, A. Role of the Novel Hsp90 Co-Chaperones in Dynein Arms’ Preassembly. Int. J. Mol. Sci. 2019, 20, 6174. [Google Scholar] [CrossRef]
- Dong, F.; Shinohara, K.; Botilde, Y.; Nabeshima, R.; Asai, Y.; Fukumoto, A.; Hasegawa, T.; Matsuo, M.; Takeda, H.; Shiratori, H.; et al. Pih1d3 is required for cytoplasmic preassembly of axonemal dynein in mouse sperm. J. Cell Biol. 2014, 204, 203–213. [Google Scholar] [CrossRef]
- Yamaguchi, H.; Oda, T.; Kikkawa, M.; Takeda, H. Systematic studies of all PIH proteins in zebrafish reveal their distinct roles in axonemal dynein assembly. Elife 2018, 7, e36979. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, R.; Hirono, M.; Kamiya, R. Discrete PIH proteins function in the cytoplasmic preassembly of different subsets of axonemal dyneins. J. Cell Biol. 2010, 190, 65–71. [Google Scholar] [CrossRef]
- Chen, Y.; Zhao, M.; Wang, S.; Chen, J.; Wang, Y.; Cao, Q.; Zhou, W.; Liu, J.; Xu, Z.; Tong, G.; et al. A novel role for DYX1C1, a chaperone protein for both Hsp70 and Hsp90, in breast cancer. J. Cancer Res. Clin. Oncol. 2009, 135, 1265–1276. [Google Scholar] [CrossRef] [PubMed]
- Thomas, L.; Bouhouche, K.; Whitfield, M.; Thouvenin, G.; Coste, A.; Louis, B.; Szymanski, C.; Bequignon, E.; Papon, J.F.; Castelli, M.; et al. TTC12 Loss-of-Function Mutations Cause Primary Ciliary Dyskinesia and Unveil Distinct Dynein Assembly Mechanisms in Motile Cilia Versus Flagella. Am. J. Hum. Genet. 2020, 106, 153–169. [Google Scholar] [CrossRef]
- Hartill, V.L.; van de Hoek, G.; Patel, M.P.; Little, R.; Watson, C.M.; Berry, I.R.; Shoemark, A.; Abdelmottaleb, D.; Parkes, E.; Bacchelli, C.; et al. DNAAF1 links heart laterality with the AAA+ ATPase RUVBL1 and ciliary intraflagellar transport. Hum. Mol. Genet. 2018, 27, 529–545. [Google Scholar] [CrossRef]
- Mali, G.R.; Yeyati, P.L.; Mizuno, S.; Dodd, D.O.; Tennant, P.A.; Keighren, M.A.; Zur Lage, P.; Shoemark, A.; Garcia-Munoz, A.; Shimada, A.; et al. ZMYND10 functions in a chaperone relay during axonemal dynein assembly. Elife 2018, 7, e34389. [Google Scholar] [CrossRef]
- Okamoto, T.; Nishimura, Y.; Ichimura, T.; Suzuki, K.; Miyamura, T.; Suzuki, T.; Moriishi, K.; Matsuura, Y. Hepatitis C virus RNA replication is regulated by FKBP8 and Hsp90. EMBO J. 2006, 25, 5015–5025. [Google Scholar] [CrossRef] [PubMed]
- Horani, A.; Ferkol, T.W.; Shoseyov, D.; Wasserman, M.G.; Oren, Y.S.; Kerem, B.; Amirav, I.; Cohen-Cymberknoh, M.; Dutcher, S.K.; Brody, S.L.; et al. LRRC6 mutation causes primary ciliary dyskinesia with dynein arm defects. PLoS ONE 2013, 8, e59436. [Google Scholar] [CrossRef] [PubMed]
- Kott, E.; Duquesnoy, P.; Copin, B.; Legendre, M.; Dastot-Le Moal, F.; Montantin, G.; Jeanson, L.; Tamalet, A.; Papon, J.F.; Siffroi, J.P.; et al. Loss-of-function mutations in LRRC6, a gene essential for proper axonemal assembly of inner and outer dynein arms, cause primary ciliary dyskinesia. Am. J. Hum. Genet. 2012, 91, 958–964. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.J.; Noh, S.H.; Han, S.M.; Choi, W.I.; Kim, H.Y.; Yu, S.; Lee, J.S.; Rim, J.H.; Lee, M.G.; Hildebrandt, F.; et al. ZMYND10 stabilizes intermediate chain proteins in the cytoplasmic pre-assembly of dynein arms. PLoS Genet. 2018, 14, e1007316. [Google Scholar] [CrossRef]
- Zhao, L.; Yuan, S.; Cao, Y.; Kallakuri, S.; Li, Y.; Kishimoto, N.; DiBella, L.; Sun, Z. Reptin/Ruvbl2 is a Lrrc6/Seahorse interactor essential for cilia motility. Proc. Natl. Acad. Sci. USA 2013, 110, 12697–12702. [Google Scholar] [CrossRef]
- Li, J.; Richter, K.; Buchner, J. Mixed Hsp90-cochaperone complexes are important for the progression of the reaction cycle. Nat. Struct. Mol. Biol. 2011, 18, 61–66. [Google Scholar] [CrossRef] [PubMed]
- Dafinger, C.; Rinschen, M.M.; Borgal, L.; Ehrenberg, C.; Basten, S.G.; Franke, M.; Hohne, M.; Rauh, M.; Gobel, H.; Bloch, W.; et al. Targeted deletion of the AAA-ATPase Ruvbl1 in mice disrupts ciliary integrity and causes renal disease and hydrocephalus. Exp. Mol. Med. 2018, 50, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhao, L.; Yuan, S.; Zhang, J.; Sun, Z. Axonemal dynein assembly requires the R2TP complex component Pontin. Dev. 2017, 144, 4684–4693. [Google Scholar] [CrossRef]
- Maurizy, C.; Quinternet, M.; Abel, Y.; Verheggen, C.; Santo, P.E.; Bourguet, M.; Paiva, A.C.F.; Bragantini, B.; Chagot, M.E.; Robert, M.C.; et al. The RPAP3-Cterminal domain identifies R2TP-like quaternary chaperones. Nat. Commun. 2018, 9, 2093. [Google Scholar] [CrossRef] [PubMed]
- Knowles, M.R.; Ostrowski, L.E.; Loges, N.T.; Hurd, T.; Leigh, M.W.; Huang, L.; Wolf, W.E.; Carson, J.L.; Hazucha, M.J.; Yin, W.; et al. Mutations in SPAG1 cause primary ciliary dyskinesia associated with defective outer and inner dynein arms. Am. J. Hum. Genet. 2013, 93, 711–720. [Google Scholar] [CrossRef] [PubMed]
- Spangler, S.A.; Schmitz, S.K.; Kevenaar, J.T.; de Graaff, E.; de Wit, H.; Demmers, J.; Toonen, R.F.; Hoogenraad, C.C. Liprin-alpha2 promotes the presynaptic recruitment and turnover of RIM1/CASK to facilitate synaptic transmission. J. Cell Biol. 2013, 201, 915–928. [Google Scholar] [CrossRef] [PubMed]
- Saeki, M.; Irie, Y.; Ni, L.; Yoshida, M.; Itsuki, Y.; Kamisaki, Y. Monad, a WD40 repeat protein, promotes apoptosis induced by TNF-alpha. Biochem. Biophys. Res. Commun. 2006, 342, 568–572. [Google Scholar] [CrossRef]
- Liu, G.; Wang, L.; Pan, J. Chlamydomonas WDR92 in association with R2TP-like complex and multiple DNAAFs to regulate ciliary dynein preassembly. J. Mol. Cell Biol. 2019, 11, 770–780. [Google Scholar] [CrossRef] [PubMed]
- Patel-King, R.S.; Sakato-Antoku, M.; Yankova, M.; King, S.M. WDR92 is required for axonemal dynein heavy chain stability in cytoplasm. Mol. Biol. Cell 2019, 30, 1834–1845. [Google Scholar] [CrossRef]
- Itsuki, Y.; Saeki, M.; Nakahara, H.; Egusa, H.; Irie, Y.; Terao, Y.; Kawabata, S.; Yatani, H.; Kamisaki, Y. Molecular cloning of novel Monad binding protein containing tetratricopeptide repeat domains. FEBS Lett. 2008, 582, 2365–2370. [Google Scholar] [CrossRef]
- zur Lage, P.; Stefanopoulou, P.; Styczynska-Soczka, K.; Quinn, N.; Mali, G.; von Kriegsheim, A.; Mill, P.; Jarman, A.P. Ciliary dynein motor preassembly is regulated by Wdr92 in association with HSP90 co-chaperone, R2TP. J. Cell Biol. 2018, 217, 2583–2598. [Google Scholar] [CrossRef] [PubMed]
- Avidor-Reiss, T.; Ha, A.; Basiri, M.L. Transition Zone Migration: A Mechanism for Cytoplasmic Ciliogenesis and Postaxonemal Centriole Elongation. Cold Spring Harb. Perspect. Biol. 2017, 9, a028142. [Google Scholar] [CrossRef] [PubMed]
- Fingerhut, J.M.; Yamashita, Y.M. mRNA localization mediates maturation of cytoplasmic cilia in Drosophila spermatogenesis. J. Cell Biol. 2020, 219, e202003084. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).