The Aldehyde Dehydrogenase ALDH2*2 Allele, Associated with Alcohol Drinking Behavior, Dates Back to Prehistoric Times
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Subjects
2.2. Genome-Wide Genotype Calling and Assessment of Associations
2.3. Obtaining Genotypes from a Public-Domain Database to Augment This Study
2.4. Obtaining Allele-Carrying Haplotypes from Densely Genotyped Persons
2.5. Mutation Age Estimation Using Haplotype Length Distributions
3. Results
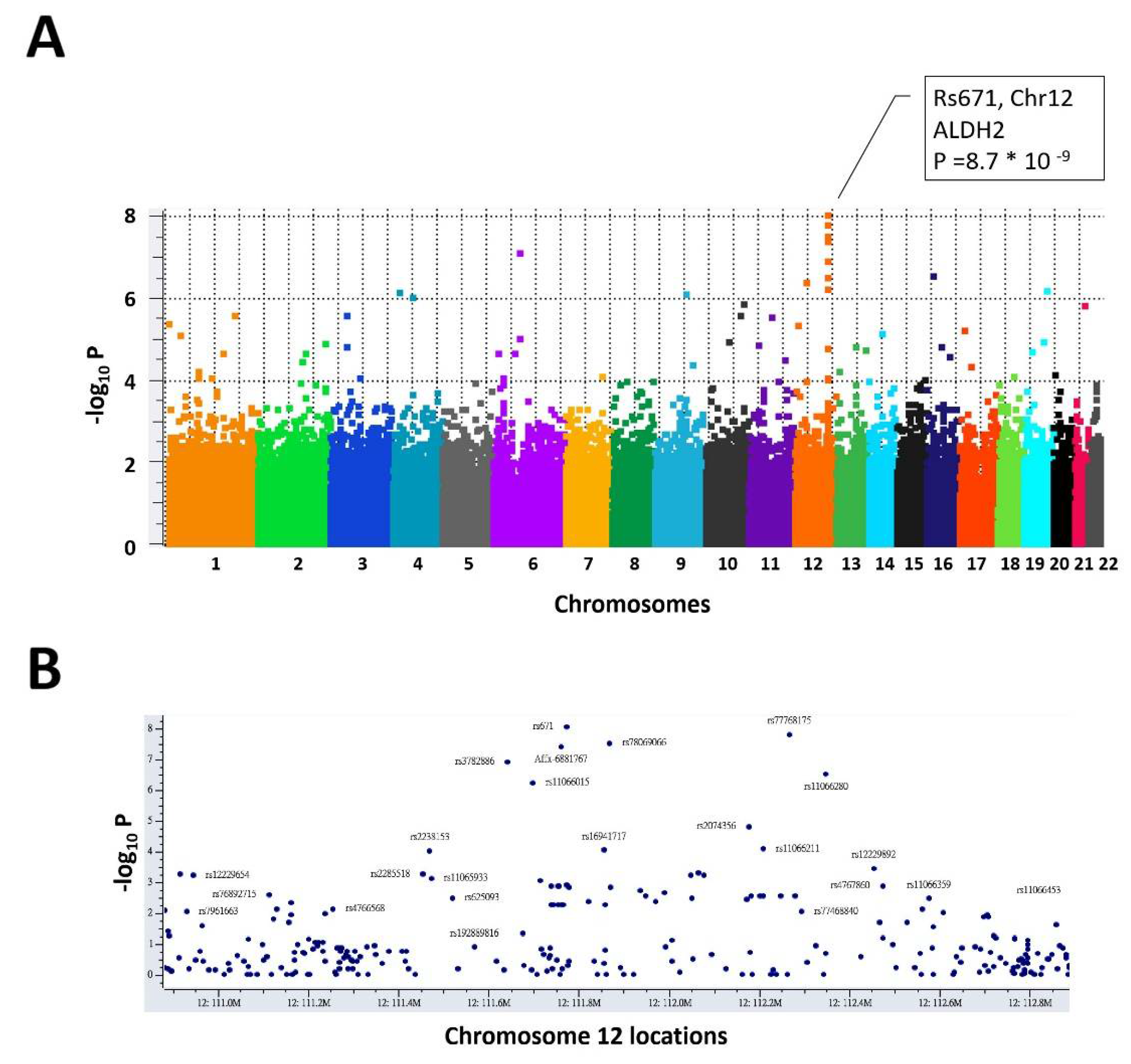
3.1. The Genomic Variant rs671 G > A Is the Leading Variant Associated with Alcohol Consumption Behavior
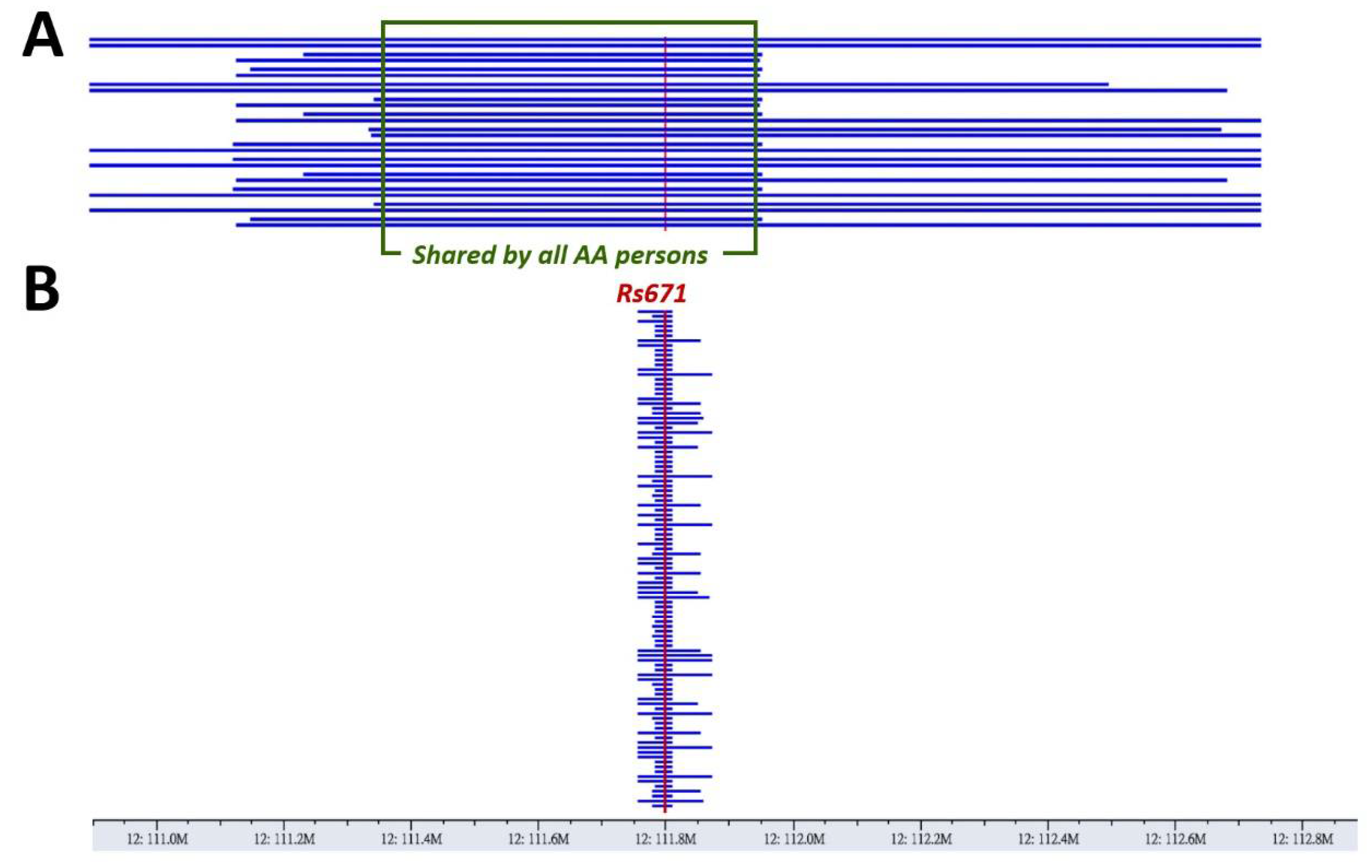
3.2. A Haplotype Shared by All ALDH2*2-Carrying Individuals Indicates Its Identity by Descent
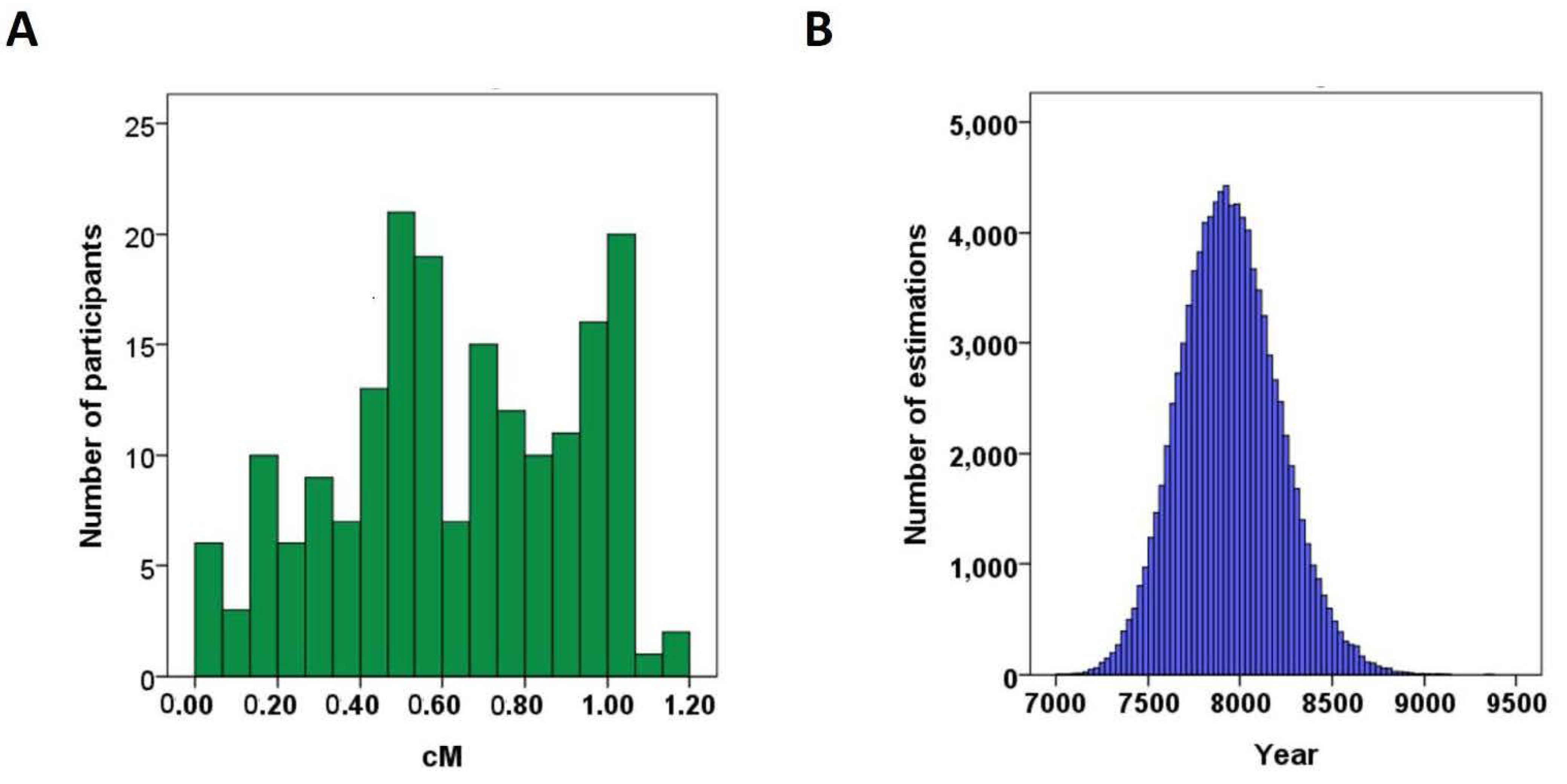
3.3. Age of the ALDH2*2 Allele Was Estimated to Be 7935 Years Ago
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- McGovern, P.E.; Zhang, J.; Tang, J.; Zhang, Z.; Hall, G.R.; Moreau, R.A.; Nunez, A.; Butrym, E.D.; Richards, M.P.; Wang, C.-S.; et al. Fermented beverages of pre- and proto-historic China. Proc. Natl. Acad. Sci. USA 2004, 101, 17593–17598. [Google Scholar] [CrossRef]
- Hines, L.M.; Rimm, E.B. Moderate alcohol consumption and coronary heart disease: A review. Postgrad. Med. J. 2001, 77, 747–752. [Google Scholar] [CrossRef]
- Goldwater, D.; Karlamangla, A.; Merkin, S.S.; Seeman, T. Compared to non-drinkers, individuals who drink alcohol have a more favorable multisystem physiologic risk score as measured by allostatic load. PLoS ONE 2019, 14, e0223168. [Google Scholar] [CrossRef]
- Koppes, L.L.; Dekker, J.M.; Hendriks, H.F.; Bouter, L.; Heine, R.J. Moderate Alcohol Consumption Lowers the Risk of Type 2 Diabetes: A meta-analysis of prospective observational studies. Diabetes Care 2005, 28, 719–725. [Google Scholar] [CrossRef]
- Mukamal, K.J.; Maclure, M.; Muller, J.E.; Sherwood, J.B.; Mittleman, M. Prior Alcohol Consumption and Mortality Following Acute Myocardial Infarction. JAMA 2001, 285, 1965–1970. [Google Scholar] [CrossRef]
- Mukamal, K.J.; Chung, H.; Jenny, N.S.; Kuller, L.H.; Longstreth, W.T.; Mittleman, M.; Burke, G.L.; Cushman, M.; Psaty, B.M.; Siscovick, D.S. Alcohol Consumption and Risk of Coronary Heart Disease in Older Adults: The Cardiovascular Health Study. J. Am. Geriatr. Soc. 2005, 54, 30–37. [Google Scholar] [CrossRef]
- Muntwyler, J.; Hennekens, C.H.; Buring, J.E.; Gaziano, J.M. Mortality and light to moderate alcohol consumption after myocardial infarction. Lancet 1998, 352, 1882–1885. [Google Scholar] [CrossRef]
- Solomon, C.G.; Hu, F.B.; Stampfer, M.J.; Colditz, G.A.; Speizer, F.E.; Rimm, E.B.; Willett, W.C.; Manson, J.E. Moderate alcohol consumption and risk of coronary heart disease among women with type 2 diabetes mellitus. Circulation 2000, 102, 494–499. [Google Scholar] [CrossRef]
- Esch, T.; Stefano, G.B. The neurobiology of pleasure, reward processes, addiction and their health implications. Neuro Endocrinol. Lett. 2004, 25, 235–251. [Google Scholar]
- Mbarek, H.; Milaneschi, Y.; Fedko, I.O.; Hottenga, J.-J.; de Moor, M.; Jansen, R.; Gelernter, J.; Sherva, R.; Willemsen, G.; Boomsma, D.I.; et al. The genetics of alcohol dependence: Twin and SNP-based heritability, and genome-wide association study based on AUDIT scores. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2015, 168, 739–748. [Google Scholar] [CrossRef]
- Jorgenson, E.; Thai, K.K.; Hoffmann, T.J.; Sakoda, L.C.; Kvale, M.N.; Banda, Y.; Schaefer, C.; Risch, N.; Mertens, J.; Weisner, C.; et al. Genetic contributors to variation in alcohol consumption vary by race/ethnicity in a large multi-ethnic genome-wide association study. Mol. Psychiatry 2017, 22, 1359–1367. [Google Scholar] [CrossRef]
- Sun, Y.; Chang, S.; Wang, F.; Sun, H.; Ni, Z.; Yue, W.; Zhou, H.; Gelernter, J.; Malison, R.T.; Kalayasiri, R.; et al. Genome-wide association study of alcohol dependence in male Han Chinese and cross-ethnic polygenic risk score comparison. Transl. Psychiatry 2019, 9, 249. [Google Scholar] [CrossRef]
- Clarke, T.-K.; Adams, M.J.; Davies, G.; Howard, D.; Hall, L.S.; Padmanabhan, S.; Murray, A.; Smith, B.; Campbell, A.; Hayward, C.; et al. Genome-wide association study of alcohol consumption and genetic overlap with other health-related traits in UK Biobank (N = 112,117). Mol. Psychiatry 2017, 22, 1376–1384. [Google Scholar] [CrossRef]
- Macgregor, S.; Lind, P.A.; Bucholz, K.K.; Hansell, N.K.; Madden, P.A.; Richter, M.M.; Montgomery, G.W.; Martin, N.G.; Heath, A.C.; Whitfield, J.B. Associations of ADH and ALDH2 gene variation with self report alcohol reactions, consumption and dependence: An integrated analysis. Hum. Mol. Genet. 2009, 18, 580–593. [Google Scholar] [CrossRef] [PubMed]
- Quillen, E.; Chen, X.-D.; Almasy, L.; Yang, F.; He, H.; Li, X.; Wang, X.-Y.; Liu, T.-Q.; Hao, W.; Deng, H.-W.; et al. ALDH2is associated to alcohol dependence and is the major genetic determinant of “daily maximum drinks” in a GWAS study of an isolated rural chinese sample. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2014, 165, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Roige, S.; Palmer, A.A.; Fontanillas, P.; Elson, S.L.; Adams, M.J.; Howard, D.; Edenberg, H.J.; Davies, G.; Crist, R.C.; Deary, I.J.; et al. Genome-Wide Association Study Meta-Analysis of the Alcohol Use Disorders Identification Test (AUDIT) in Two Population-Based Cohorts. Am. J. Psychiatry 2019, 176, 107–118. [Google Scholar] [CrossRef] [PubMed]
- Kranzler, H.R.; Zhou, H.; Kember, R.; Smith, R.V.; Justice, A.C.; Damrauer, S.; Tsao, P.S.; Klarin, D.; Baras, A.; Reid, J.; et al. Author Correction: Genome-wide association study of alcohol consumption and use disorder in 274,424 individuals from multiple populations. Nat. Commun. 2019, 10, 4050. [Google Scholar] [CrossRef]
- Budas, G.R.; Disatnik, M.-H.; Mochly-Rosen, D. Aldehyde Dehydrogenase 2 in Cardiac Protection: A New Therapeutic Target? Trends Cardiovasc. Med. 2009, 19, 158–164. [Google Scholar] [CrossRef]
- Yang, H.; Song, Z.; Yang, G.-P.; Zhang, B.-K.; Chen, M.; Wu, T.; Guo, R. The ALDH2 rs671 Polymorphism Affects Post-Stroke Epilepsy Susceptibility and Plasma 4-HNE Levels. PLoS ONE 2014, 9, e109634. [Google Scholar] [CrossRef] [PubMed]
- Hemani, G.; Bowden, J.; Smith, G.D. Evaluating the potential role of pleiotropy in Mendelian randomization studies. Hum. Mol. Genet. 2018, 27, R195–R208. [Google Scholar] [CrossRef]
- Chen, C.-H.; Ferreira, J.C.B.; Gross, E.R.; Mochly-Rosen, D. Targeting Aldehyde Dehydrogenase 2: New Therapeutic Opportunities. Physiol. Rev. 2014, 94, 1–34. [Google Scholar] [CrossRef] [PubMed]
- Peng, G.-S.; Chen, Y.-C.; Wang, M.-F.; Lai, C.-L.; Yin, S.-J. ALDH2*2 but not ADH1B*2 is a causative variant gene allele for Asian alcohol flushing after a low-dose challenge. Pharm. Genom. 2014, 24, 607–617. [Google Scholar] [CrossRef]
- Kato, N.; Takeuchi, F.; Tabara, Y.; Kelly, T.N.; Go, M.J.; Sim, X.; Tay, W.T.; Chen, C.-H.; Zhang, Y.; Yamamoto, K.; et al. Meta-analysis of genome-wide association studies identifies common variants associated with blood pressure variation in east Asians. Nat. Genet. 2011, 43, 531–538. [Google Scholar] [CrossRef]
- Mizuno, Y.; Hokimoto, S.; Harada, E.; Kinoshita, K.; Nakagawa, K.; Yoshimura, M.; Ogawa, H.; Yasue, H. Variant Aldehyde Dehydrogenase 2 (ALDH2*2) Is a Risk Factor for Coronary Spasm and ST-Segment Elevation Myocardial Infarction. J. Am. Heart Assoc. 2016, 5. [Google Scholar] [CrossRef]
- Xia, C.-L.; Chu, P.; Liu, Y.-X.; Qu, X.-L.; Gao, X.-F.; Wang, Z.-M.; Dong, J.; Chen, S.-L.; Zhang, J.-X. ALDH2 rs671 polymorphism and the risk of heart failure with preserved ejection fraction (HFpEF) in patients with cardiovascular diseases. J. Hum. Hypertens. 2019, 34, 16–23. [Google Scholar] [CrossRef]
- Chen, J.; Huang, W.; Cheng, C.-H.; Zhou, L.; Jiang, G.-B.; Hu, Y.-Y. Association Between Aldehyde dehydrogenase-2 Polymorphisms and Risk of Alzheimer’s Disease and Parkinson’s Disease: A Meta-Analysis Based on 5315 Individuals. Front. Neurol. 2019, 10, 290. [Google Scholar] [CrossRef]
- Dakeishi, M.; Murata, K.; Sasaki, M.; Tamura, A.; Iwata, T. Association of alcohol dehydrogenase 2 and aldehyde dehydrogenase 2 genotypes with fasting plasma glucose levels in Japanese male and female workers. Alcohol Alcohol. 2008, 43, 143–147. [Google Scholar] [CrossRef] [PubMed]
- Sakiyama, M.; Matsuo, H.; Nakaoka, H.; Yamamoto, K.; Nakayama, A.; Nakamura, T.; Kawai, S.; Okada, R.; Ooyama, H.; Shimizu, T.; et al. Identification of rs671, a common variant of ALDH2, as a gout susceptibility locus. Sci. Rep. 2016, 6, 25360. [Google Scholar] [CrossRef] [PubMed]
- Takeshima, K.; Nishiwaki, Y.; Suda, Y.; Niki, Y.; Sato, Y.; Kobayashi, T.; Miyamoto, K.; Uchida, H.; Inokuchi, W.; Tsuji, T.; et al. A missense single nucleotide polymorphism in the ALDH2 gene, rs671, is associated with hip fracture. Sci. Rep. 2017, 7, 428. [Google Scholar] [CrossRef] [PubMed]
- Dong, Q.; Ren, G.; Zhang, K.; Liu, D.; Dou, Q.; Hao, D. Genetic polymorphisms of ALDH2 are associated with lumbar disc herniation in a Chinese Han population. Sci. Rep. 2018, 8, 10379. [Google Scholar] [CrossRef]
- Park, B.; Kim, J.-H.; Lee, E.-G.; Jung, S.-Y.; Lee, S.Y.; Kang, H.-S.; Han, J.H. Role of aldehyde dehydrogenases, alcohol dehydrogenase 1B genotype, alcohol consumption, and their combination in breast cancer in East-Asian women. Sci. Rep. 2020, 10, 6564. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Wang, C.; Shen, L.; Gu, D.; Xu, Z.; Zhang, X.; Xu, Y.; Chen, J. Clinical significance of ALDH2 rs671 polymorphism in esophageal cancer: Evidence from 31 case-control studies. OncoTargets Ther. 2015, 8, 649–659. [Google Scholar] [CrossRef]
- Chien, J.; Liu, J.; Lee, M.; Jen, C.; Batrla-Utermann, R.; Lu, S.; Wang, L.; You, S.; Yang, H.; Chen, C. Risk and Predictors of Hepatocellular Carcinoma for Chronic Hepatitis b Patients with Newly Developed Cirrhosis. J. Gastroenterol. Hepatol. 2016, 31, 1971–1977. [Google Scholar] [CrossRef]
- Luo, H.-R.; Wu, G.-S.; Pakstis, A.J.; Tong, L.; Oota, H.; Kidd, K.K.; Zhang, Y.-P. Origin and dispersal of atypical aldehyde dehydrogenase ALDH2⁎487Lys. Gene 2009, 435, 96–103. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Wu, G. Could ALDH2*2 be the reason for low incidence and mortality of ovarian cancer for East Asia women? Oncotarget 2017, 9, 12503–12512. [Google Scholar] [CrossRef]
- Chang, J.S.; Hsiao, J.-R.; Chen, C.-H. ALDH2 polymorphism and alcohol-related cancers in Asians: A public health perspective. J. Biomed. Sci. 2017, 24, 19. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-H.; Ferreira, J.C.B.; Joshi, A.U.; Stevens, M.C.; Li, S.-J.; Hsu, J.H.-M.; Maclean, R.; Ferreira, N.D.; Cervantes, P.R.; Martinez, D.D.; et al. Novel and prevalent non-East Asian ALDH2 variants; Implications for global susceptibility to aldehydes’ toxicity. EBioMedicine 2020, 55, 102753. [Google Scholar] [CrossRef]
- Saunders, J.B.; Aasland, O.G.; Babor, T.F.; De La Fuente, J.R.; Grant, M. Development of the Alcohol Use Disorders Identification Test (AUDIT): WHO Collaborative Project on Early Detection of Persons with Harmful Alcohol Consumption-II. Addiction 1993, 88, 791–804. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.-C.; Tsai, Y.-F.; Chen, C.-Y.; Liu, C.-Y. Alcohol Use Disorders Identification Test (AUDIT): Establishment of Cut-off Scores in a Hospitalized Chinese Population. Alcohol. Clin. Exp. Res. 2005, 29, 53–57. [Google Scholar] [CrossRef] [PubMed]
- Higgins-Biddle, J.C.; Babor, T.F. A review of the Alcohol Use Disorders Identification Test (AUDIT), AUDIT-C, and USAUDIT for screening in the United States: Past issues and future directions. Am. J. Drug Alcohol. Abus. 2018, 44, 578–586. [Google Scholar] [CrossRef]
- Slatkin, M.; Rannala, B. Estimating Allele Age. Ann. Rev. Genom. Human Genet. 2020, 1, 225–249. [Google Scholar] [CrossRef] [PubMed]
- Gandolfo, L.C.; Bahlo, M.; Speed, T.P. Dating Rare Mutations from Small Samples with Dense Marker Data. Genetics 2014, 197, 1315–1327. [Google Scholar] [CrossRef]
- Liang, K.-H.; Lu, Y.-H.; Niu, C.-W.; Chang, S.-K.; Chen, Y.-R.; Cheng, C.-Y.; Hsu, T.; Yang, C.-F.; Nakamura, K.; Niu, D.-M. The Fabry disease-causing mutation, GLA IVS4+919G>A, originated in Mainland China more than 800 years ago. J. Hum. Genet. 2020, 65, 619–625. [Google Scholar] [CrossRef]
- Efron, B. Bootstrap Methods: Another Look at the Jackknife; Springer Series in Statistics; Springer: Berlin/Heidelberg, Germany, 1992; pp. 569–593. [Google Scholar]
- Hong, E.P.; Park, J.W. Sample Size and Statistical Power Calculation in Genetic Association Studies. Genom. Inf. 2012, 10, 117–122. [Google Scholar] [CrossRef]
- Li, H.; Borinskaya, S.; Yoshimura, K.; Kal’Ina, N.; Marusin, A.; Stepanov, V.; Qin, Z.; Khaliq, S.; Lee, M.-Y.; Yang, Y.; et al. Refined Geographic Distribution of the OrientalALDH2*504Lys(nee487Lys) Variant. Ann. Hum. Genet. 2009, 73, 335–345. [Google Scholar] [CrossRef] [PubMed]
- Goebel, T.; Waters, M.R.; O’Rourke, D.H. The Late Pleistocene Dispersal of Modern Humans in the Americas. Science 2008, 319, 1497–1502. [Google Scholar] [CrossRef]
- Gruhn, R. Evidence grows that peopling of the Americas began more than 20,000 years ago. Nat. Cell Biol. 2020, 584, 47–48. [Google Scholar] [CrossRef]
| AUDIT Score ≥ 8 | AUDIT Score = 0 | P | |
|---|---|---|---|
| Patient number | 71 | 126 | |
| Gender | |||
| Male | 65 (91.55%) | 114 (90.48%) | 0.802 * |
| Female | 6 (8.45%) | 12 (9.52%) | |
| Age, years, Mean ± SD | 53.72 ± 9.93 | 56.97 ± 12.53 | 0.062 ‡ |
| BMI, kg/m2, Mean ± SD | 25.6 ± 4.0 | 25.8 ± 3.7 | 0.643 ‡ |
| <24, | 25 (35.2%) | 41 (32.5%) | 0.597 * |
| 24–27 | 24 (33.8%) | 37 (29.4%) | |
| >27 | 22 (31.0%) | 48 (38.1%) | |
| Coffee | |||
| never | 24 (33.8%) | 36 (28.6%) | 0.483 * |
| social | 19 (26.8%) | 44 (34.9%) | |
| more than sometimes | 28 (39.4%) | 46 (36.5%) | |
| Tea | |||
| never | 15 (21.1%) | 36 (28.6%) | 0.401 * |
| social | 24 (33.8%) | 44 (34.9%) | |
| more than sometimes | 32 (45.1%) | 46 (36.5%) | |
| Education level | |||
| No education | 1 (1.4%) | 4 (3.2%) | 0.609 ‡ |
| Elementary school | 8 (11.3%) | 23 (18.3%) | |
| Junior high school | 15 (21.1%) | 19 (15.1%) | |
| Senior high school | 34 (47.9%) | 34 (27.0%) | |
| College | 11 (15.5%) | 38 (30.2%) | |
| Higher education | 2 (2.8%) | 8 (6.3%) | |
| Exercise | |||
| little | 23 (33.8%) | 41 (31.9%) | 0.699 * |
| Sometimes (≤2 days per week) | 17 (25.0%) | 26 (23.7%) | |
| Frequent (≥3 days per week) | 28 (41.2%) | 59 (44.4%) |
| SNP | Chr | Gene | Allele | Case | Control | MAF | Fisher’s Exact P | Hardy–Weinberg Equilibrium P | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Homozygous Major | Heterozygous | Homozygous Minor | Homozygous Major | Heterozygous | Homozygous Minor | Case | Control | ||||||
| rs116369005 | 6 | Intergenic | G/C | 38 | 26 | 0 | 113 | 7 | 0 | 0.090 | 7.54 × 10−8 | 0.125 | 0.947 |
| rs671 | 12 | ALDH2 | G/A | 56 | 15 | 0 | 47 | 66 | 13 | 0.272 | 8.75 × 10−9 | 0.609 | 0.345 |
| rs77768175 | 12 | HECTD4 | A/G | 56 | 15 | 0 | 48 | 65 | 13 | 0.269 | 1.56 × 10−8 | 0.609 | 0.416 |
| rs78069066 | 12 | MAPKAPK5/TMEM116 | G/A | 54 | 17 | 0 | 45 | 68 | 13 | 0.282 | 2.96 × 10−8 | 0.519 | 0.225 |
| rs3782886 | 12 | BRAP | T/C | 53 | 18 | 0 | 46 | 67 | 13 | 0.282 | 1.20 × 10−7 | 0.473 | 0.282 |
| rs11066280 | 12 | HECTD4 | T/A | 52 | 18 | 1 | 43 | 70 | 13 | 0.294 | 3.05 × 10−7 | 0.923 | 0.136 |
| rs541300736 | 16 | TEKT5 | G/A | 36 | 25 | 0 | 112 | 8 | 0 | 0.091 | 2.62 × 10−7 | 0.132 | 0.931 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, C.-L.; Chien, R.-N.; Chen, L.-W.; Huang, T.-S.; Shyu, Y.-C.; Yeh, C.-T.; Liang, K.-H. The Aldehyde Dehydrogenase ALDH2*2 Allele, Associated with Alcohol Drinking Behavior, Dates Back to Prehistoric Times. Biomolecules 2021, 11, 1376. https://doi.org/10.3390/biom11091376
Lin C-L, Chien R-N, Chen L-W, Huang T-S, Shyu Y-C, Yeh C-T, Liang K-H. The Aldehyde Dehydrogenase ALDH2*2 Allele, Associated with Alcohol Drinking Behavior, Dates Back to Prehistoric Times. Biomolecules. 2021; 11(9):1376. https://doi.org/10.3390/biom11091376
Chicago/Turabian StyleLin, Chih-Lang, Rong-Nan Chien, Li-Wei Chen, Ting-Shuo Huang, Yu-Chiau Shyu, Chau-Ting Yeh, and Kung-Hao Liang. 2021. "The Aldehyde Dehydrogenase ALDH2*2 Allele, Associated with Alcohol Drinking Behavior, Dates Back to Prehistoric Times" Biomolecules 11, no. 9: 1376. https://doi.org/10.3390/biom11091376
APA StyleLin, C.-L., Chien, R.-N., Chen, L.-W., Huang, T.-S., Shyu, Y.-C., Yeh, C.-T., & Liang, K.-H. (2021). The Aldehyde Dehydrogenase ALDH2*2 Allele, Associated with Alcohol Drinking Behavior, Dates Back to Prehistoric Times. Biomolecules, 11(9), 1376. https://doi.org/10.3390/biom11091376






