JMJ Histone Demethylases Balance H3K27me3 and H3K4me3 Levels at the HSP21 Locus during Heat Acclimation in Arabidopsis
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Normal Growth and Heat Stress Conditions
2.3. Estradiol Treatment
2.4. Chromatin Immunoprecipitation (ChIP)
3. Results
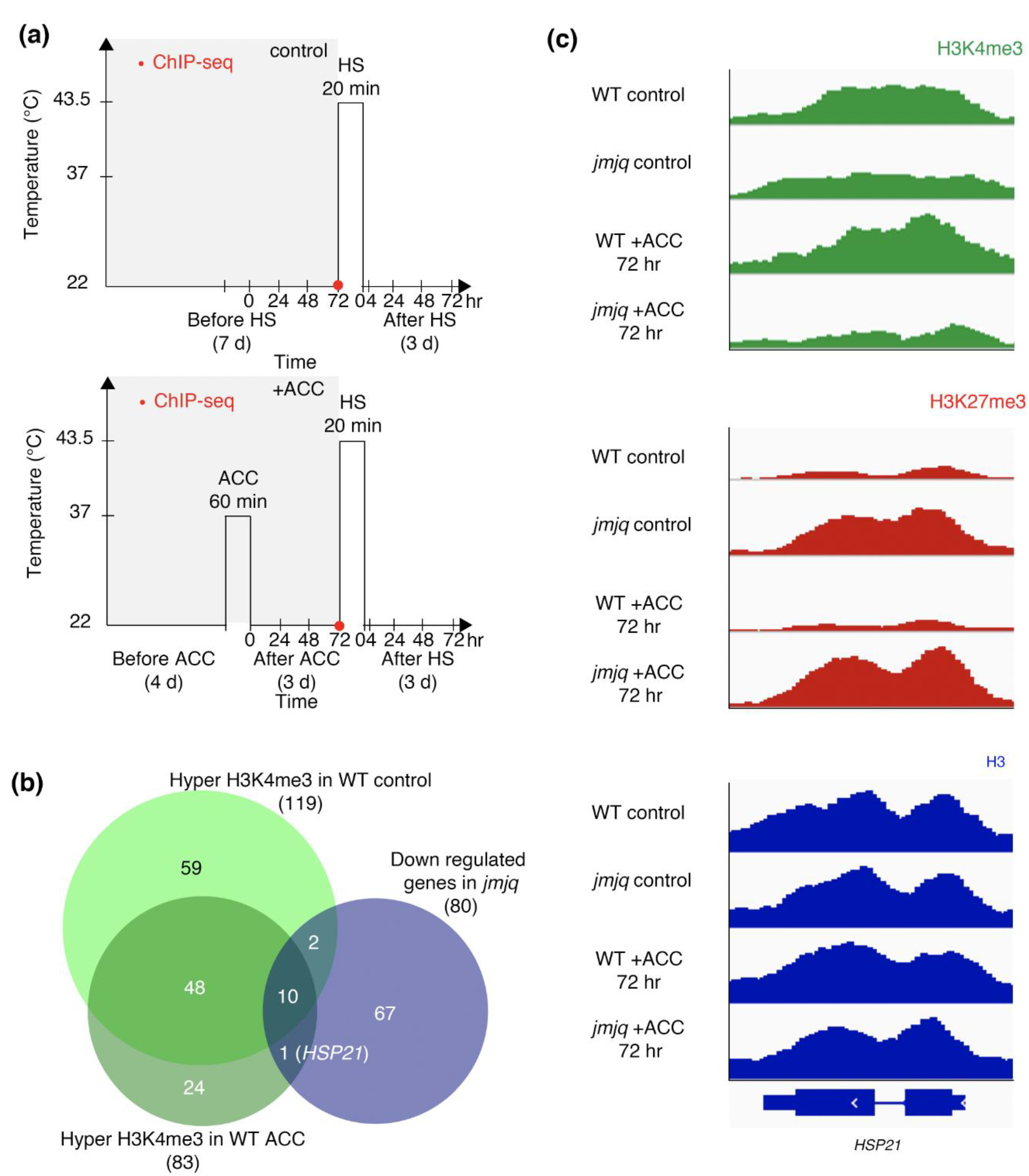
3.1. Acclimation-Dependent Sustained H3K4me3 Methylation at the HSP21 Locus
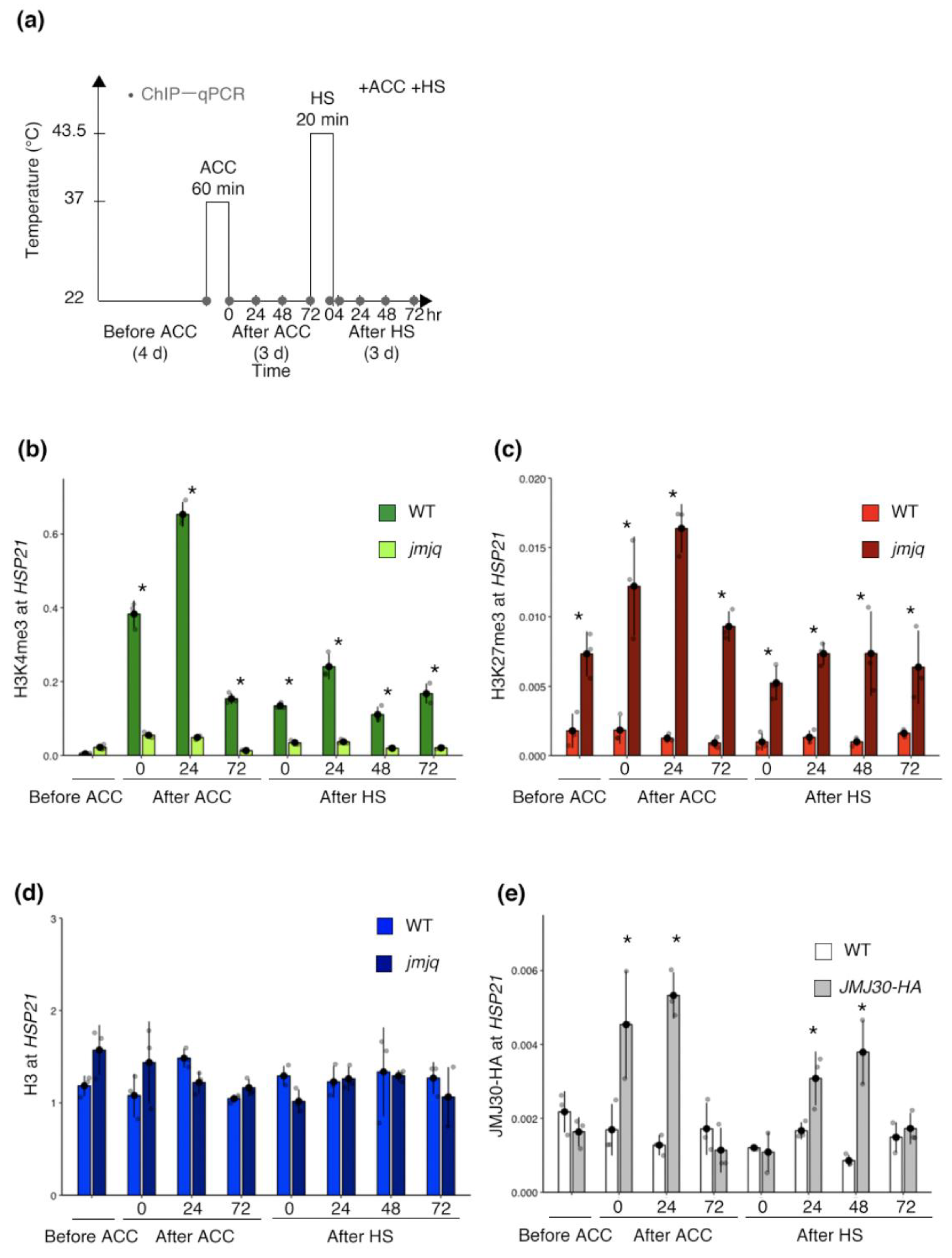
3.2. Proper Maintenance of the Balance between H3K27me3 and H3K4me3 by JMJs
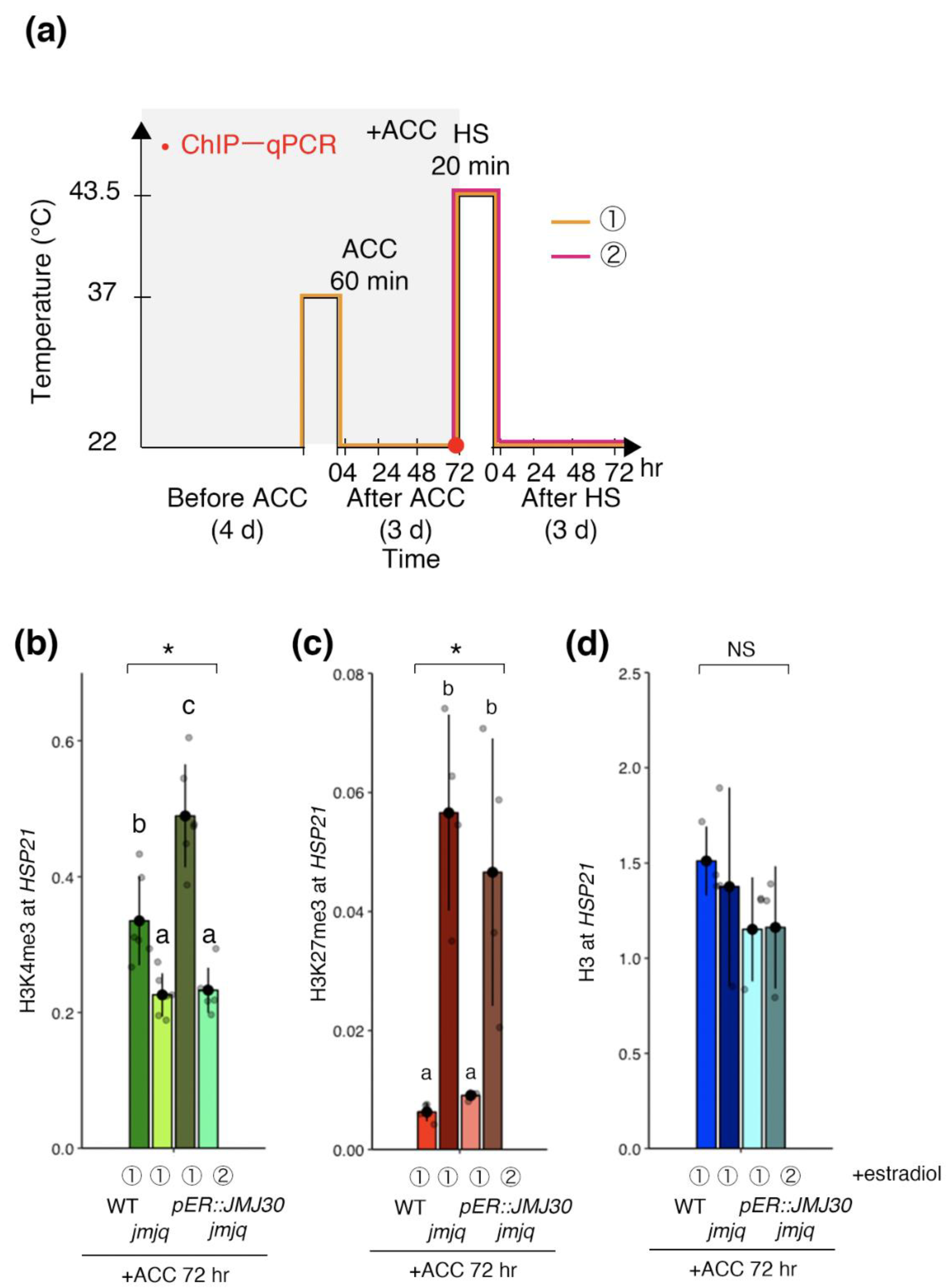
3.3. Manipulation of Balance between H3K27me3 and H3K4me3 by JMJ30 Induction
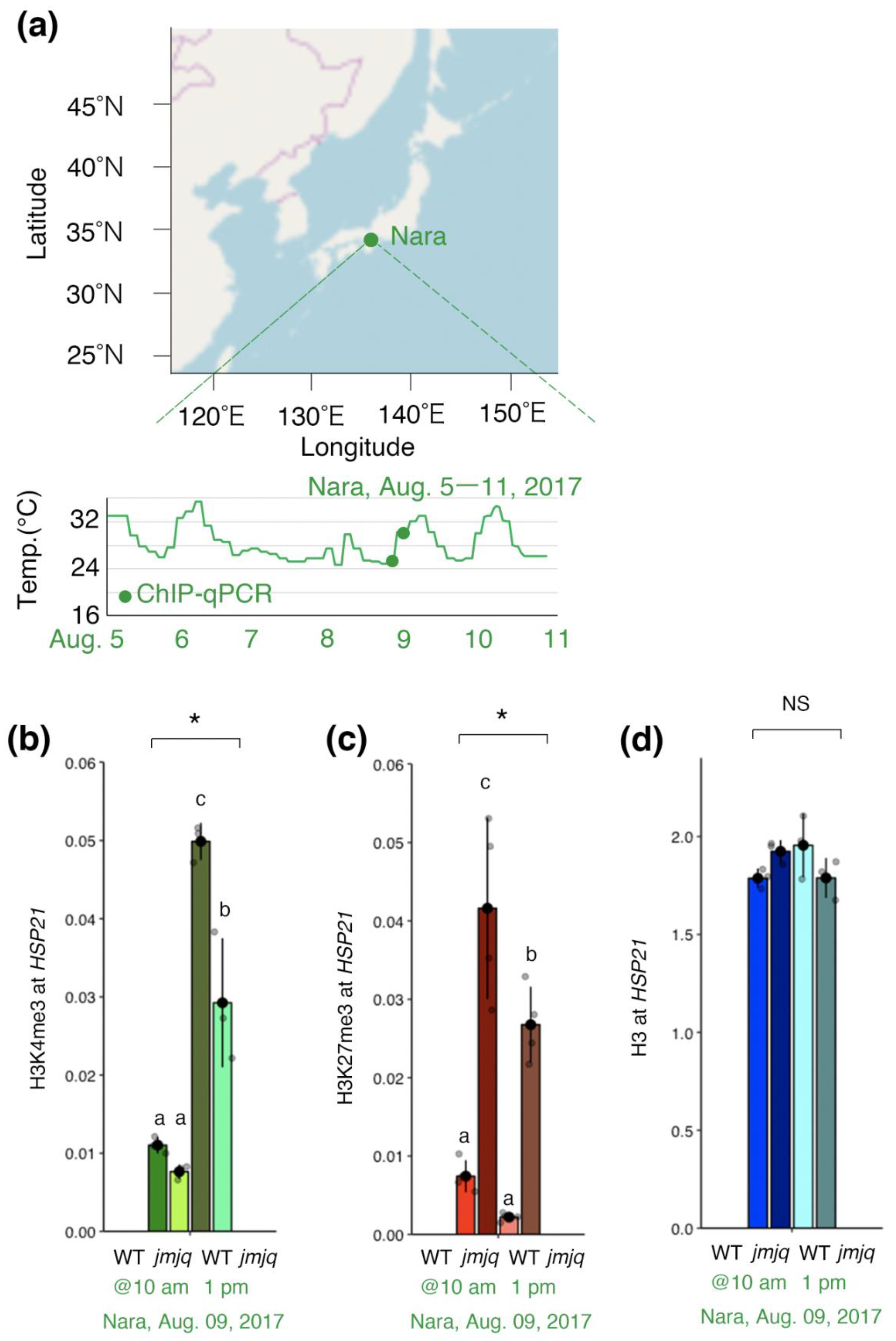
3.4. Maintenance of Balance between H3K27me3 and H3K4me3 by JMJs under Field Conditions
4. Discussion
4.1. Transcriptional and Epigenetic Regulation of the HSP21 Gene by H3K27me3 Demethylases during Heat Acclimation
4.2. Transcriptional and Epigenetic Regulation of the HSP21 Gene under Natural Conditions
4.3. The Redundant JMJ Network Regulating HSP Genes
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pandey, P.; Irulappan, V.; Bagavathiannan, M.V.; Senthil-Kumar, M. Impact of combined abiotic and biotic stresses on plant growth and avenues for crop improvement by exploiting physio-morphological traits. Front. Plant Sci. 2017, 8, 537. [Google Scholar] [CrossRef]
- Lämke, J.; Bäurle, I. Epigenetic and chromatin-based mechanisms in environmental stress adaptation and stress memory in plants. Genome Biol. 2017, 18, 124. [Google Scholar] [CrossRef]
- Ohama, N.; Sato, H.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Transcriptional regulatory network of plant heat stress response. Trends Plant Sci. 2017, 22, 53–65. [Google Scholar] [CrossRef]
- AI-Whaibi, M.H. Plant heat-shock proteins: A mini review. J. King Saud Univ. Sci. 2011, 2, 139–150. [Google Scholar] [CrossRef]
- Xiao, J.; Lee, U.S.; Wagner, D. Tug of war: Adding and removing histone lysine methylation in Arabidopsis. Curr. Opin. Plant Biol. 2016, 34, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Law, J.A.; Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 2005, 6, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Charng, Y.Y.; Liu, H.C.; Liu, N.Y.; Chi, W.T.; Wang, C.N.; Chang, S.H.; Wang, T.T. A heat-inducible transcription factor, HsfA2, is required for extension of acquired thermotolerance in Arabidopsis. Plant Physiol. 2006, 143, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Nishizawa, A.; Yabuta, Y.; Yoshida, E.; Maruta, T.; Yoshimura, K.; Shigeoka, S. Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J. 2006, 48, 535–547. [Google Scholar] [CrossRef]
- Lämke, J.; Brzezinka, K.; Altmann, S.; Bäurle, I. A hit-and-run heat shock factor governs sustained histone methylation and transcriptional stress memory. EMBO J. 2016, 35, 162–175. [Google Scholar] [CrossRef]
- Guenther, M.G.; Levine, S.S.; Boyer, L.A.; Jaenisch, R.; Young, R.A. A chromatin landmark and transcription initiation at most promoters in human cells. Cell 2006, 130, 77–88. [Google Scholar] [CrossRef]
- Zhang, X.; Bernatavichute, Y.V.; Cokus, S.; Pellegrini, M.; Jacobsen, S.E. Genome-wide analysis of mono-, di- and trimethylation of histone H3 lysine 4 in Arabidopsis thaliana. Genome Biol. 2009, 10, R62. [Google Scholar] [CrossRef]
- Crevillén, P.; Yang, H.; Cui, X.; Greeff, C.; Trick, M.; Qiu, Q.; Cao, X.; Dean, C. Epigenetic reprogramming that prevents transgenerational inheritance of the vernalized state. Nature 2014, 515, 587–590. [Google Scholar] [CrossRef]
- Lu, F.; Cui, X.; Zhang, S.; Jenuwein, T.; Cao, X. Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat. Genet. 2011, 43, 715–719. [Google Scholar] [CrossRef]
- Cui, X.; Lu, F.; Qiu, Q.; Zhou, B.; Gu, L.; Zhang, S.; Kang, Y.; Cui, X.; Ma, X.; Yao, Q.; et al. REF6 recognizes a specific DNA sequence to demethylate H3K27me3 and regulate organ boundary formation in Arabidopsis. Nat. Genet. 2016, 48, 694–699. [Google Scholar] [CrossRef] [PubMed]
- Yan, W.; Chen, D.; Smaczniak, C.; Engelhorn, J.; Liu, H.; Yang, W.; Graf, A.; Carles, C.C.; Zhou, D.X.; Kaufmann, K. Dynamic and spatial restriction of Polycomb activity by plant histone demethylases. Nat. Plants 2018, 4, 681–689. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Ichihashi, Y.; Suzuki, T.; Shibata, A.; Shirasu, K.; Yamaguchi, N.; Ito, T. Abscisic acid-dependent histone demethylation during post-germination growth arrest in Arabidopsis. Plant Cell Environ. 2019, 42, 2198–2214. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Yamaguchi, N.; Ito, T. Histone demethylases control root elongation in response to stress-signaling hormone abcisic acid. Plant Signal. Behav. 2019, 14, 1604019. [Google Scholar] [CrossRef] [PubMed]
- Gan, E.-S.; Xu, Y.; Wong, J.-Y.; Goh, J.G.; Sun, B.; Wee, W.-Y.; Huang, J.; Ito, T. Jumonji demethylases moderate precocious flowering at elevated temperature via regulation of FLC in Arabidopsis. Nat. Commun. 2014, 5, 5098. [Google Scholar] [CrossRef]
- Yamaguchi, N.; Matsubara, S.; Yoshimizu, K.; Seki, M.; Hamada, K.; Kamitani, M.; Kurita, Y.; Nomura, Y.; Nagashima, K.; Inagaki, S.; et al. Removal of repressive histone marks creates epigenetic memory of recurring heat in Arabidopsis. bioRxiv 2020. [Google Scholar] [CrossRef]
- Sedaghatmehr, M.; Mueller-Roeber, B.; Balazadeh, S. The plastid metalloprotease FtsH6 and small heat shock protein HSP21 jointly regulate thermomemory in Arabidopsis. Nat. Commun. 2016, 7, 12439. [Google Scholar] [CrossRef]
- Li, Y.; Li, Y.; Liu, Y.; Wu, Y.; Xie, Q. The sHSP22 heat shock protein requires the ABI1 protein phosphatase to modulate polar auxin transport and downstream responses. Plant Physiol. 2018, 176, 2406–2425. [Google Scholar] [CrossRef] [PubMed]
- Rutsdottir, G.; Härmark, J.; Weide, Y.; Hebert, H.; Rasmussen, M.I.; Wernersson, S.; Respondek, M.; Akke, M.; Højrup, P.; Koeck, P.J.B.; et al. Structural model of dodecameric heat-shock protein Hsp21: Flexible N-terminal arms interact with client proteins while C-terminal tails maintain the dodecamer and chaperone activity. J. Biol. Chem. 2017, 292, 8103–8121. [Google Scholar] [CrossRef]
- Zhong, L.; Zhou, W.; Wang, H.; Ding, S.; Lu, Q.; Wen, X.; Peng, L.; Zhang, L.; Lu, C. Chloroplast small heat shock protein HSP21 interacts with plastid nucleoid protein pTAC5 and is essential for chloroplast development in Arabidopsis under heat stress. Plant Cell 2013, 25, 2925–2943. [Google Scholar] [CrossRef] [PubMed]
- Pfalz, J.; Liere, K.; Kandlbinder, A.; Dietz, K.J.; Oelmüller, R. pTAC2, -6, and -12 are components of the transcriptionally active plastid chromosome that are required for plastid gene expression. Plant Cell 2006, 18, 176–197. [Google Scholar] [CrossRef] [PubMed]
- Pfalz, J.; Pfannschmidt, T. Essential nucleoid proteins in early chloroplast development. Trends Plant Sci. 2013, 18, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Yu, E.; Fan, C.; Yang, Q.; Li, X.; Wan, B.; Dong, Y.; Wang, X.; Zhou, Y. Identification of heat responsive genes in Brassica napus siliques at the seed-filling stage through transcriptional profiling. PLoS ONE 2014, 9, e101914. [Google Scholar] [CrossRef]
- Sun, A.-Q.; Yi, S.-Y.; Yang, J.-Y.; Zhao, C.-M.; Liu, J. Identification and characterization of a heat-inducible ftsH gene from tomato (Lycopersicon esculentum Mill.). Plant Sci. 2006, 170, 551–562. [Google Scholar] [CrossRef]
- Xue, G.-P.; Drenth, J.; McIntyre, C.L. TaHsfA6f is a transcriptional activator that regulates a suite of heat stress protection genes in wheat (Triticum aestivum L.) including previously unknown Hsf targets. J. Exp. Bot. 2015, 66, 1025–1039. [Google Scholar] [CrossRef]
- Johnson, S.M.; Lim, F.L.; Finkler, A.; Fromm, H.; Slabas, A.R.; Knight, M.R. Transcriptomic analysis of Sorghum bicolor responding to combined heat and drought stress. BMC Genom. 2014, 15, 456. [Google Scholar] [CrossRef]
- Busch, W.; Wunderlich, M.; Schöffl, F. Identification of novel heat shock factor-dependent genes and biochemical pathways in Arabidopsis thaliana. Plant J. 2005, 41, 1–14. [Google Scholar] [CrossRef]
- Brzezinka, K.; Altmann, S.; Czesnick, H.; Nicolas, P.; Gorka, M.; Benke, E.; Kabelitz, T.; Jähne, F.; Graf, A.; Kappel, C.; et al. Arabidopsis FORGETTER1 mediates stress-induced chromatin memory through nucleosome remodeling. eLife 2016, 5, e17061. [Google Scholar] [CrossRef]
- Baile, F.; Merini, W.; Hidalgo, I.; Calonje, M. Dissection of PRC1 and PRC2 recruitment in Arabidopsis connects EAR repressome to PRC2 anchoring. bioRxiv 2020. [Google Scholar] [CrossRef]
- Pelayo, M.A.; Yamaguchi, N.; Ito, T. One factor, many systems: The floral homeotic protein AGAMOUS and its epigenetic regulatory mechanisms. Curr. Opin. Plant Biol. 2021, 61, 102009. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Hu, H.; Ren, H.; Yang, Z.; Qiu, Q.; Qi, W.; Liu, X.; Chen, X.; Cui, X.; Li, S.; et al. The Arabidopsis H3K27me3 demethylase JUMONJI 13 is a temperature and photoperiod dependent flowering repressor. Nat. Commun. 2019, 10, 1303. [Google Scholar] [CrossRef] [PubMed]
- Keyzor, C.; Mermaz, B.; Trigazis, E.; Jo, S.; Song, J. Histone demethylases ELF6 and JMJ13 antagonistically regulate self-fertility in arabidopsis. Front. Plant Sci. 2021, 12, 640135. [Google Scholar] [CrossRef]
- Nagano, A.J.; Sato, Y.; Mihara, M.; Antonio, B.A.; Motoyama, R.; Itoh, H.; Nagamura, Y.; Izawa, T. Deciphering and prediction of transcriptome dynamics under fluctuating field conditions. Cell 2012, 151, 1358–1369. [Google Scholar] [CrossRef] [PubMed]
- Nagano, A.J.; Kawagoe, T.; Sugisaka, J.; Honjo, M.N.; Iwayama, K.; Kudoh, H. Annual transcriptome dynamics in natural environments reveals plant seasonal adaptation. Nat. Plants 2019, 5, 74–83. [Google Scholar] [CrossRef]
- Nishio, H.; Nagano, A.J.; Ito, T.; Suzuki, Y.; Kudoh, H. Seasonal plasticity and diel stability of H3K27me3 in natural fluctuating environments. Nat. Plants 2020, 6, 1091–1097. [Google Scholar] [CrossRef]
- Nishio, H.; Buzas, D.M.; Nagano, A.J.; Iwayama, K.; Ushio, M.; Kudoh, H. Repressive chromatin modification underpins the long-term expression trend of a perennial flowering gene in nature. Nat. Commun. 2020, 11, 2065. [Google Scholar] [CrossRef]
- Hepworth, J.; Dean, C. Flowering locus C’s lessons: Conserved chromatin switches underpinning developmental timing and adaptation. Plant Physiol. 2015, 168, 1237–1245. [Google Scholar] [CrossRef]
- Tamada, Y.; Yun, J.Y.; Woo, S.C.; Amasino, R.M. ARABIDOPSIS TRITHORAX-RELATED7 is required for methylation of lysine 4 of histone H3 and for transcriptional activation of FLOWERING LOCUS C. Plant Cell 2009, 21, 3257–3269. [Google Scholar] [CrossRef] [PubMed]
- Yun, J.Y.; Tamada, Y.; Kang, Y.E.; Amasino, R.M. Arabidopsis trithorax-related3/SET domain GROUP2 is required for the winter-annual habit of Arabidopsis thaliana. Plant Cell Physiol. 2012, 53, 834–846. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.Q.; Luo, J.H.; Cui, Z.H.; Xue, M.; Wang, L.; Zhang, X.Y.; Pawlowski, W.P.; He, Y. ATX3, ATX4, and ATX5 encode putative H3K4 methyltransferases and are critical for plant development. Plant Physiol. 2017, 174, 1795–1806. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamaguchi, N.; Ito, T. JMJ Histone Demethylases Balance H3K27me3 and H3K4me3 Levels at the HSP21 Locus during Heat Acclimation in Arabidopsis. Biomolecules 2021, 11, 852. https://doi.org/10.3390/biom11060852
Yamaguchi N, Ito T. JMJ Histone Demethylases Balance H3K27me3 and H3K4me3 Levels at the HSP21 Locus during Heat Acclimation in Arabidopsis. Biomolecules. 2021; 11(6):852. https://doi.org/10.3390/biom11060852
Chicago/Turabian StyleYamaguchi, Nobutoshi, and Toshiro Ito. 2021. "JMJ Histone Demethylases Balance H3K27me3 and H3K4me3 Levels at the HSP21 Locus during Heat Acclimation in Arabidopsis" Biomolecules 11, no. 6: 852. https://doi.org/10.3390/biom11060852
APA StyleYamaguchi, N., & Ito, T. (2021). JMJ Histone Demethylases Balance H3K27me3 and H3K4me3 Levels at the HSP21 Locus during Heat Acclimation in Arabidopsis. Biomolecules, 11(6), 852. https://doi.org/10.3390/biom11060852






