Proteomic and Bioinformatic Profiling of Transporters in Higher Plant Mitochondria
Abstract
1. Introduction
2. The Experimental Proteome and Transportome in Plant Mitochondria
2.1. The Experimental Mitochondrial Proteome
2.2. The Experimental Mitochondrial Transportome
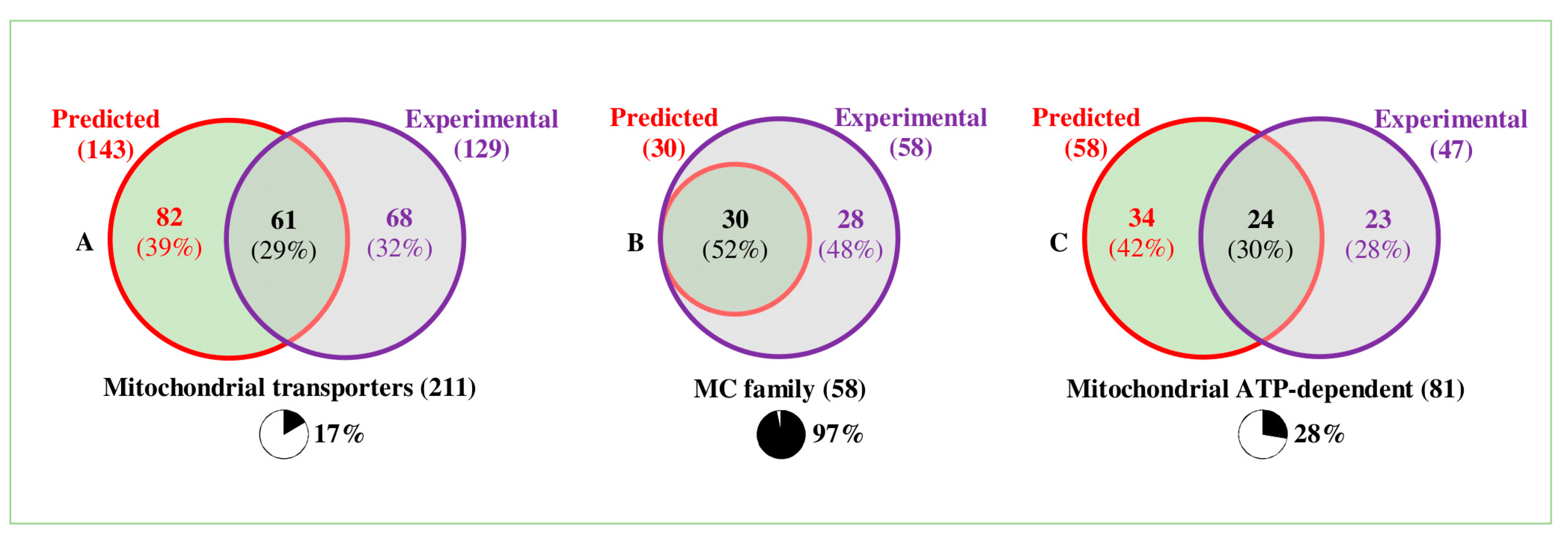
3. Mitochondrial Transporters—Predictions
4. The Different Transporter Classes and Families
4.1. ATP-Dependent Transporters
4.1.1. ABC Transporters
4.1.2. The Mitochondrial Protein Translocator (MPT) Family
4.1.3. F-ATPases
4.1.4. P-ATPases
4.2. Secondary Transporters
4.2.1. The Mitochondrial Carrier (MC) Family
4.2.2. Transport of Inorganic Ions
4.3. (Ion) Channels
4.3.1. Porin (VDAC or VIC)
4.3.2. Aquaporin
4.3.3. Ion Channels
5. Posttranslational Modifications of Transporters
6. Physiological Changes in the Mitochondrial Transporters
6.1. Changes in Rice during Development
6.2. Changes in Maize during Development
7. Conclusions
Supplementary Materials
Funding
Conflicts of Interest
Abbreviations
| ABC | ATP-binding cassette |
| DAP | days after pollination |
| DTC | dicarboxylate/tricarboxylate carrier |
| GO | gene ontology |
| IMM | inner mitochondrial membrane |
| MC | mitochondrial carrier |
| MCF | MC family |
| MPT | mitochondrial protein translocase |
| OMM | outer mitochondrial membrane |
| PTM | posttranslational modification |
| TCA | tricarboxylic acid |
| TIM | translocase inner membrane |
| TOM | translocase outer membrane |
| UCP | uncoupling protein |
| VDAC | voltage-dependent anion channel |
| VIC | voltage-gated ion channel |
References
- Stein, W.D.; Litman, T. Channels, Carriers, and Pumps—An Introduction to Membrane Transport, 2nd ed.; Academic Press: San. Diego, CA, USA, 2015. [Google Scholar]
- Møller, I.M.; Rasmusson, A.G.; Browse, J.J. Chapter 12 Plant Respiration and Lipid Metabolism. In Plant Physiology, 6th Ed.; Taiz, L., Zeiger, E., Murphy, A., Møller, I.M., Eds.; Sinauer Associates: Sunderland, MA, USA, 2014; pp. 317–352. [Google Scholar]
- Li-Beisson, Y.; Thelen, J.J.; Fedosejevs, E.; Harwood, J.L. The lipid biochemistry of eukaryotic algae. Progr. Lipid Res. 2019, 74, 31–68. [Google Scholar] [CrossRef] [PubMed]
- Igamberdiev, A.U. Citrate valve integrates mitochondria into photosynthetic metabolism. Mitochondrion. 2020, 52, 218–230. [Google Scholar] [CrossRef] [PubMed]
- Vanlerberghe, G.C.; Dahal, K.; Alber, N.A.; Chadee, A. Photosynthesis, respiration and growth: A carbon and energy balancing act for alternative oxidase. Mitochondrion 2020, 52, 197–211. [Google Scholar] [CrossRef] [PubMed]
- Møller, I.M.; Sweetlove, L.J. ROS signaling—Specificity is required. Trends Plant Sci. 2010, 15, 370–374. [Google Scholar] [CrossRef] [PubMed]
- Tran, H.C.; Van Aken, O. Mitochondrial unfolded protein-related responses across kingdoms: Similar problems, different regulators. Mitochondrion 2020, 53, 166–177. [Google Scholar] [CrossRef] [PubMed]
- Rao, R.S.P.; Salvato, F.; Thal, B.; Eubel, H.; Thelen, J.J.; Møller, I.M. The proteome of higher plant mitochondria. Mitochondrion. 2017, 33, 22–37. [Google Scholar] [CrossRef] [PubMed]
- Jeandard, D.; Smirnova, A.; Tarassov, I.; Barrey, E.; Smirnov, A.; Entelis, N. Import of non-coding RNAs into human mitochondria: A critical review andemerging approaches. Cells 2019, 8, 286. [Google Scholar] [CrossRef] [PubMed]
- Warren, J.M.; Sloan, D.B. Interchangeable parts: The evolutionarily dynamic tRNA population in plant mitochondria. Mitochondrion 2020, 52, 144–156. [Google Scholar] [CrossRef]
- Homblé, F.; Krammer, E.-M.; Prévost, M. Plant VDAC: Facts and speculations. Biochim. Biophys. Acta (BBA)—Biomembr. 2012, 1818, 1486–1501. [Google Scholar] [CrossRef]
- Hemono, M.; Ubrig, E.; Azeredo, K.; Salinas-Giegé, T.; Drouard, L.; Duchêne, A.-M. Arabidopsis voltage-dependent anion channels (VDACs): Overlapping and specific functions in mitochondria. Cells 2020, 9, 1023. [Google Scholar] [CrossRef]
- Lee, C.P.; Millar, A.H. The plant mitochondrial transportome: Balancing metabolic demands with energetic constraints. Trends Plant Sci. 2016, 21, 662–676. [Google Scholar] [CrossRef]
- Zhang, N.; Rao, R.S.P.; Salvato, F.; Havelund, J.F.; Møller, I.M.; Thelen, J.J.; Xu, D. MU-LOC: A machine-learning method for predicting mitochondrially localized proteins in plants. Front. Plant Sci. 2018, 9, 634. [Google Scholar] [CrossRef] [PubMed]
- Senkler, J.; Senkler, M.; Eubel, H.; Hildebrandt, T.; Lengwenus, C.; Schertl, P.; Schwarzländer, M.; Wagner, S.; Wittig, I.; Braun, H.P. The mitochondrial complexome of Arabidopsis thaliana. Plant J. 2017, 89, 1079–1092. [Google Scholar] [CrossRef] [PubMed]
- Mower, J.P. Variation in protein gene and intron content among land plant mitogenomes. Mitochondrion 2020, 52, 203–213. [Google Scholar] [CrossRef] [PubMed]
- George, R.; Walsh, P.; Beddoe, T.; Lithgow, T. The nascent polypeptide-associated complex (NAC) promotes interaction of ribosomes with the mitochondrial surface in vivo. FEBS Lett. 2002, 516, 213–216. [Google Scholar] [CrossRef]
- Salvato, F.; Havelund, J.F.; Chen, M.; Rao, R.S.P.; Rogowska-Wrzesinska, A.; Jensen, O.N.; Gang, D.R.; Thelen, J.J.; Møller, I.M. The potato tuber mitochondrial proteome. Plant Physiol. 2014, 164, 637–653. [Google Scholar] [CrossRef]
- Fuchs, P.; Rugen, N.; Carrie, C.; Elsässer, M.; Finkemeier, I.; Giese, J.; Hildebrandt, T.M.; Kühn, K.; Maurino, V.G.; Ruberti, C.; et al. Single organelle function and organization as estimated from Arabidopsis mitochondrial proteomics. Plant J. 2020, 101, 420–441. [Google Scholar] [CrossRef]
- Millar, A.H.; Heazlewood, J.L. Genomic and proteomic analysis of mitochondrial carrier proteins in Arabidopsis. Plant Physiol. 2003, 131, 443–453. [Google Scholar] [CrossRef]
- Barrera, N.P.; Robinson, C.V. Advances in the mass spectrometry of membrane proteins: From individual proteins to intact complexes. Annu. Rev. Biochem. 2011, 80, 247–271. [Google Scholar] [CrossRef]
- Calabrese, A.N.; Radford, S.E. Mass spectrometry-enabled structural biology of membrane proteins. Methods 2018, 147, 187–205. [Google Scholar] [CrossRef]
- Schey, K.L.; Grey, A.C.; Nicklay, J.J. Mass spectrometry of membrane proteins: A focus on aquaporins. Biochemistry 2013, 52, 3807–3817. [Google Scholar] [CrossRef] [PubMed]
- Taylor, N.L.; Howell, K.A.; Heazlewood, J.L.; Tan, T.Y.; Narsai, R.; Huang, S.; Whelan, J.; Millar, A.H. Analysis of the rice mitochondrial carrier family reveals anaerobic accumulation of a basic amino acid carrier involved in arginine metabolism during seed germination. Plant Physiol. 2010, 154, 691–704. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.Q.; Wang, Y.; Zhang, Q.; Møller, I.M.; Song, S.Q. Changes in the maize embryo mitochondrial proteome during seed development. Physiol. Plant. 2018, 163, 552–572. [Google Scholar] [CrossRef] [PubMed]
- Elbourne, L.D.H.; Tetu, S.G.; Hassan, K.A.; Paulsen, I.T. TransportDB 2.0: A database for exploring membrane transporters in sequenced genomes from all domains of life. Nucleic Acids Res. 2016, 45, D320–D324. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.U.; Song, W.Y.; Hong, D.; Ko, D.; Yamaoka, Y.; Jang, S.; Yim, S.; Lee, E.; Khare, D.; Kim, K.; et al. Plant ABC transporters enable many unique aspects of a terrestrial plant’s lifestyle. Mol. Plant. 2016, 9, 338–355. [Google Scholar] [CrossRef]
- Palmieri, F.; Pierri, C.L.; De Grassi, A.; Nunes-Nesi, A.; Fernie, A.R. Evolution, structure and function of mitochondrial carriers: A review with new insights. Plant J. 2011, 66, 161–181. [Google Scholar] [CrossRef]
- Haferkamp, I.; Schmitz-Esser, S. The plant mitochondrial carrier family: Functional and evolutionary aspects. Front. Plant Sci. 2012, 3, 2. [Google Scholar] [CrossRef]
- Carrie, C.; Small, I. A reevaluation of dual-targeting of proteins to mitochondria and chloroplasts. Biochim. Biophys. Acta. 2013, 1833, 253–259. [Google Scholar] [CrossRef]
- Langner, U.; Baudisch, U.; Klösgen, R.B. Organelle import of proteins with dual targeting properties into mitochondria and chloroplasts takes place by the general import pathways. Plant Signal. Behav. 2014, 9, e29301. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Sanchez-Fernandez, R.; Lyver, E.R.; Dancis, A.; Rea, P.A. Functional characterization of AtATM1, AtATM2, and AtATM3, a subfamily of Arabidopsis half-molecule ATP-binding cassette implicated in iron homeostasis. J. Biol. Chem. 2007, 282, 21561–21571. [Google Scholar] [CrossRef]
- Balk, J.; Schaedler, T.A. Iron cofactor assembly in plants. Annu. Rev. Plant Biol. 2014, 65, 125–153. [Google Scholar] [CrossRef] [PubMed]
- Ye, H.; Rouault, T.A. Human iron-sulfur cluster assembly, cellular iron homeostasis, and disease. Biochemistry 2010, 49, 4945–4956. [Google Scholar] [CrossRef] [PubMed]
- Ghifari, A.S.; Gill-Hille, M.; Murcha, M.W. Plant mitochondrial protein import: The ins and outs. Biochemical J. 2018, 475, 2191–2208. [Google Scholar] [CrossRef]
- Toleco, M.R.; Naake, T.; Zhang, Y.; Heazlewood, J.L.; Fernie, A.R. Plant mitochondrial carriers: Molecular gatekeepers that help to regulate plant central carbon metabolism. Plants. 2020, 9, 117. [Google Scholar] [CrossRef] [PubMed]
- Møller, I.M.; Igamberdiev, A.U.; Bykova, N.V.; Finkemeier, I.; Rasmusson, A.G.; Schwarzländer, M. Matrix redox physiology governs the regulation of plant mitochondrial metabolism through post-translational protein modifications. Plant Cell. 2020, 32, 573–594. [Google Scholar] [CrossRef]
- Bartoli, C.G.; Pastori, G.M.; Foyer, C.H. Ascorbate biosynthesis in mitochondria is linked to the electron transport chain between complexes III and IV. Plant Physiol. 2000, 123, 335–344. [Google Scholar] [CrossRef]
- Scalera, V.; Giangregorio, N.; De Leonardis, S.; Console, L.; Carulli, E.S.; Tonazzi, A. Characterization of a novel mitochondrial ascorbate transporter from rat liver and potato mitochondria. Front. Mol. Biosci. 2018, 5, 58. [Google Scholar] [CrossRef]
- Miyaji, T.; Kuromori, T.; Takeuchi, Y.; Yamaji, N.; Yokosho, K.; Shimazawa, A.; Sugimoto, E.; Omote, H.; Ma, J.F.; Shinozaki, K.; et al. MAtPHT4;4 is a chloroplast-localized ascorbate transporter in Arabidopsis. Nat. Commun. 2015, 6, 5928. [Google Scholar] [CrossRef]
- Hebelstrup, K.H.; Møller, I.M. Mitochondrial signaling in plants under hypoxia: Use of Reactive Oxygen Species (ROS) and Reactive Nitrogen Species (RNS). In Reactive Oxygen and Nitrogen Species Signaling and Communication in Plants; Taiz, L., Zeiger, E., Murphy, A., Møller, I.M., Eds.; Springer: Berlin, Germany, 2015; pp. 63–77. [Google Scholar]
- Sugiura, M.; Georgescu, M.N.; Takahashi, M. A nitrite transporter associated with nitrite uptake by higher plant chloroplasts. Plant Cell Physiol. 2007, 48, 1022–1035. [Google Scholar] [CrossRef]
- Bienert, G.P.; Møller, A.L.B.; Kristiansen, K.A.; Schulz, A.; Møller, I.M.; Schjoerring, J.K.; Jahn, J.P. Specific aquaporins facilitate the diffusion of hydrogen peroxide across membranes. J. Biol. Chem. 2007, 282, 1183–1192. [Google Scholar] [CrossRef]
- Maurel, C.; Boursiac, Y.; Luu, D.T.; Santoni, V.; Shahzad, Z.; Verdoucq, L. Aquaporins in plants. Physiol Rev. 2015, 95, 1321–1358. [Google Scholar] [CrossRef] [PubMed]
- Day, D.A.; Hanson, J.B. Pyruvate and malate transport and oxidation in corn mitochondrial. Plant Physiol. 1977, 59, 630–635. [Google Scholar] [CrossRef] [PubMed]
- Vothknecht, U.; Szabò, I. Channels and transporters for inorganic ions in plant mitochondria: Prediction and facts. Mitochondrion 2020, 153, 224–233. [Google Scholar] [CrossRef] [PubMed]
- Carraretto, L.; Teardo, E.; Checchetto, V.; Finazzi, G.; Uozumi, N.; Szabo, I. Ion channels in plant bioenergetic organelles, chloroplasts and mitochondria: From molecular identification to function. Mol. Plant 2016, 9, 371–395. [Google Scholar] [CrossRef]
- Teardo, E.; Carraretto, L.; Wagner, S.; Formentin, E.; Behera, S.; De Bortoli, S.; Larosa, V.; Fuchs, P.; Schiavo, F.L.; Raffaello, A.; et al. Physiological characterization of a plant mitochondrialcalcium uniporter in vitro and in vivo. Plant Physiol. 2017, 173, 1355–1370. [Google Scholar] [CrossRef]
- Elter, A.; Hartel, A.; Sieben, C.; Hertel, B.; Fischer-Schliebs, E.; Lüttge, U.; Moroni, A.; Thiel, G. A plant homolog of animal chloride intracellular channels (CLICs) generates an ion conductance inheterologous systems. J. Biol. Chem. 2007, 282, 8786–8792. [Google Scholar] [CrossRef]
- Nakagami, H.; Sugiyama, N.; Mochida, K.; Daudi, A.; Yoshida, Y.; Toyoda, T.; Tomita, M.; Ishihama, Y.; Shirasu, K. Large-scale comparative phosphoproteomics identifiesconserved phosphorylationsitesinplants. Plant Physiol. 2010, 153, 1161–1174. [Google Scholar] [CrossRef]
- Meyer, L.J.; Gao, J.; Xu, D.; Thelen, J.J. Phosphoproteomic analysis of seed maturation in Arabidopsis, rapeseed, and soybean. Plant Physiol. 2012, 159, 517–528. [Google Scholar] [CrossRef]
- Havelund, J.F.; Thelen, J.J.; Møller, I.M. Biochemistry, proteomics, and phosphoproteomics of plant mitochondria from non-photosynthetic cells. Front. Plant Sci. 2013, 4, 51. [Google Scholar] [CrossRef]
- König, A.C.; Hartl, M.; Boersema, P.J.; Mann, M.; Finkemeier, I. The mitochondrial lysine acetylome of Arabidopsis. Mitochondrion. 2014, 19, 252–260. [Google Scholar] [CrossRef]
- Winger, A.M.; Taylor, N.L.; Heazlewood, J.L.; Day, D.A.; Millar, A.H. The cytotoxic lipid peroxidation product 4-hydroxy-2-nonenal covalently modifies a selective range of proteins linked to respiratory function in plant mitochondria. J. Biol. Chem. 2007, 282, 37436. [Google Scholar] [CrossRef]
- Møller, I.M.; Rogowska-Wrzesinska, A.; Rao, R.S.P. Protein carbonylation and metal-catalyzed protein oxidation in a cellular perspective. J. Proteomics. 2011, 74, 2228–2242. [Google Scholar] [CrossRef] [PubMed]
- Møller, I.M.; Jensen, P.E.; Hansson, A. Oxidative modifications to cellular components in plants. Annu. Rev. Plant Biol. 2007, 58, 459–481. [Google Scholar] [CrossRef]
- Smakowska, E.; Skibior-Blaszczyk, R.; Czarna, M.; Kolodziejczak, M.; Kwasniak-Owczarek, M.; Parys, K.; Funk, C.; Janska, H. Lack of FTSH4 protease affects protein carbonylation, mitochondrial morphology, and phospholipid content in mitochondria of Arabidopsis: New insights into a complex interplay. Plant Physiol. 2016, 171, 2516–2535. [Google Scholar] [CrossRef] [PubMed]
| A. thaliana | O. sativa | H. sapiens | M. musculus | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TransportDB # | Experimental * | % | Predicted # | % | TransportDB # | Experimental * | % | Predicted # | % | TransportDB # | Predicted # | % | TransportDB # | Predicted # | % | |
| Whole proteome | 26,091 | - | - | - | - | 55,890 | - | - | - | - | 37,742 | - | - | 34,966 | - | - |
| ATP-dependent | ||||||||||||||||
| ABC transporters | 124 | 6 | 5 | 13 | 11 | 129 | 0 | 0 | 11 | 9 | 77 | 10 | 13 | 65 | 5 | 8 |
| F-ATPase | 44 | 16 | 36 | 17 | 39 | 50 | 7 | 14 | 13 | 26 | 60 | 25 | 42 | 58 | 19 | 33 |
| MPT family | 25 | 19 | 76 | 10 | 40 | 30 | 2 | 7 | 10 | 33 | 29 | 6 | 21 | 40 | 6 | 15 |
| P-type ATPases | 50 | 2 | 4 | 4 | 8 | 45 | 0 | 0 | 3 | 7 | 68 | 6 | 9 | 50 | 0 | 0 |
| Other ATP-dependent | 50 | 4 | 8 | 14 | 28 | 30 | 0 | 0 | 10 | 33 | 19 | 4 | 21 | 34 | 4 | 12 |
| Secondary transporters | ||||||||||||||||
| MC family | 60 | 58 | 97 | 30 | 50 | 61 | 50 | 82 | 33 | 54 | 63 | 25 | 40 | 64 | 21 | 33 |
| Other families | 722 | 13 | 2 | 34 | 5 | 799 | 0 | 0 | 50 | 6 | 511 | 39 | 8 | 473 | 28 | 6 |
| Ion channels | 151 | 11 | 7 | 15 | 10 | 137 | 0 | 0 | 10 | 7 | 588 | 9 | 2 | 470 | 8 | 2 |
| Unclassified | 53 | 0 | 0 | 6 | 11 | 4 | 0 | 0 | 0 | 0 | 52 | 5 | 10 | 36 | 2 | 6 |
| Total (Transporters) | 1279 | 129 | 10 | 143 | 11 | 1285 | 59 | 5 | 140 | 11 | 1467 | 129 | 9 | 1290 | 93 | 7 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Møller, I.M.; Rao, R.S.P.; Jiang, Y.; Thelen, J.J.; Xu, D. Proteomic and Bioinformatic Profiling of Transporters in Higher Plant Mitochondria. Biomolecules 2020, 10, 1190. https://doi.org/10.3390/biom10081190
Møller IM, Rao RSP, Jiang Y, Thelen JJ, Xu D. Proteomic and Bioinformatic Profiling of Transporters in Higher Plant Mitochondria. Biomolecules. 2020; 10(8):1190. https://doi.org/10.3390/biom10081190
Chicago/Turabian StyleMøller, Ian Max, R. Shyama Prasad Rao, Yuexu Jiang, Jay J. Thelen, and Dong Xu. 2020. "Proteomic and Bioinformatic Profiling of Transporters in Higher Plant Mitochondria" Biomolecules 10, no. 8: 1190. https://doi.org/10.3390/biom10081190
APA StyleMøller, I. M., Rao, R. S. P., Jiang, Y., Thelen, J. J., & Xu, D. (2020). Proteomic and Bioinformatic Profiling of Transporters in Higher Plant Mitochondria. Biomolecules, 10(8), 1190. https://doi.org/10.3390/biom10081190






