A Ribosome Interaction Surface Sensitive to mRNA GCN Periodicity
Abstract
1. Introduction
2. Materials and Methods
2.1. System for Study
2.2. Specifications of MD Simulations
2.3. Subsystem Data Collection
3. Results
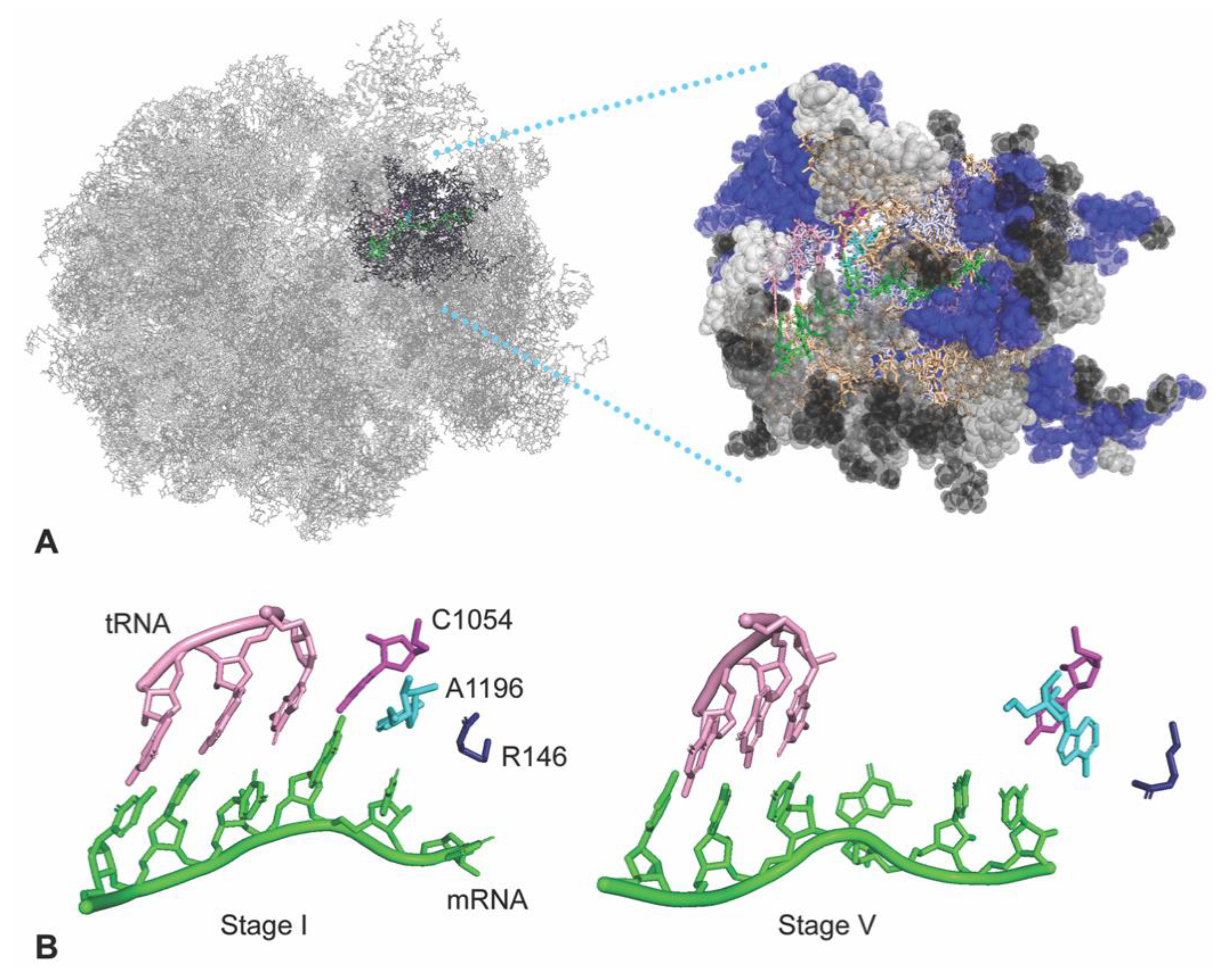
3.1. A Subsystem of the Ribosome
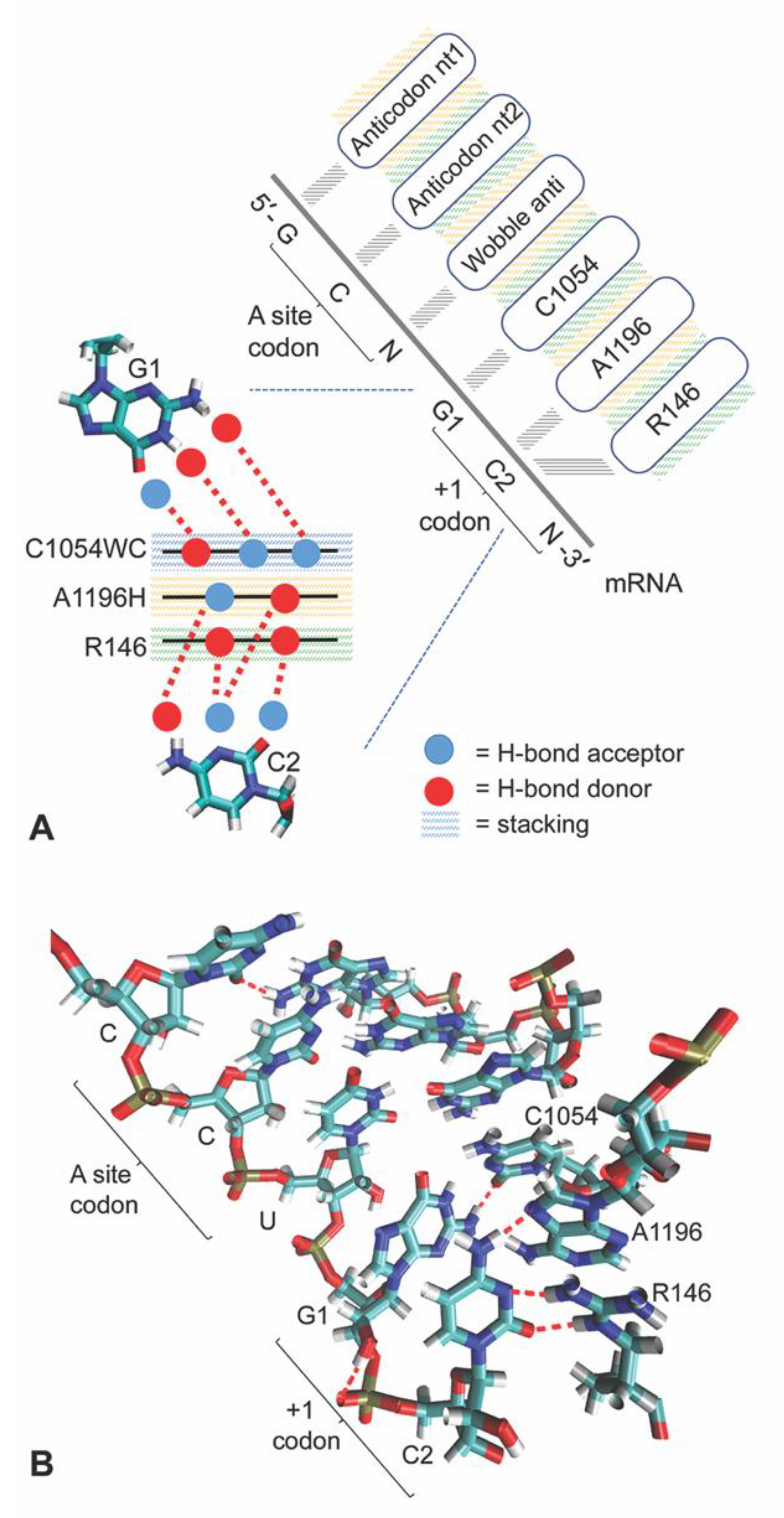
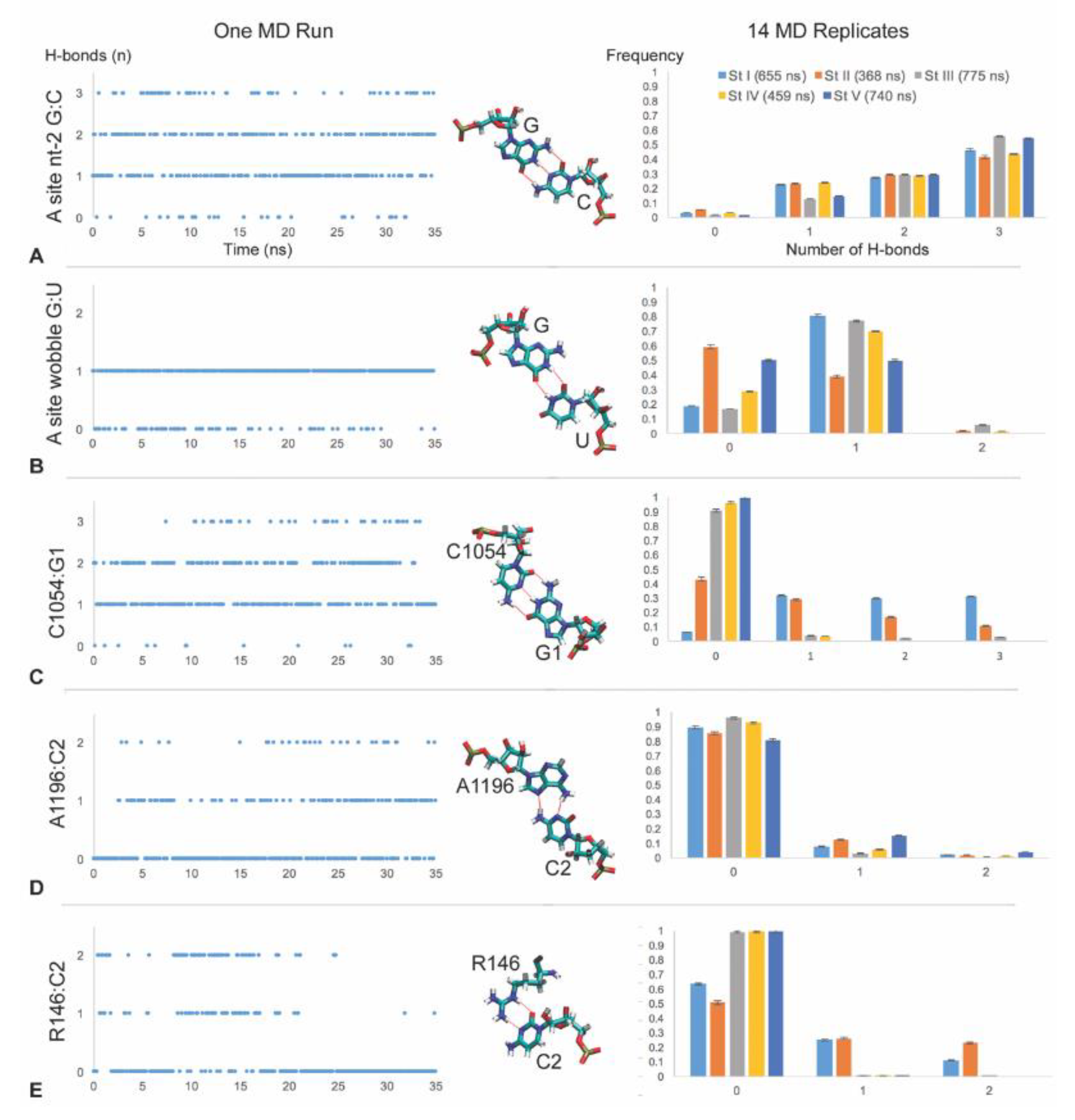
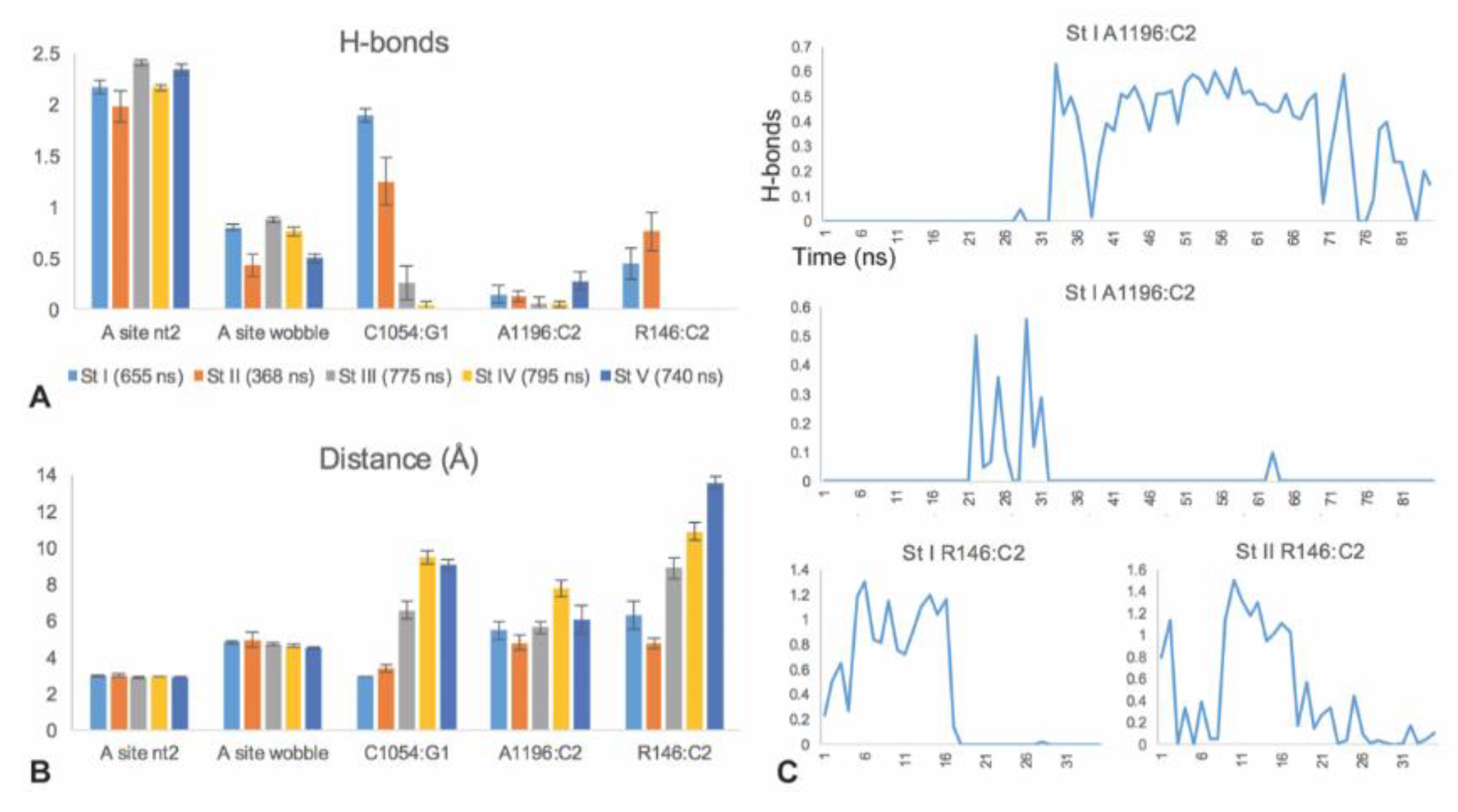
3.2. C1054 Base Pairs to the mRNA
3.3. A1196 and Rps3 R146 Hydrogen Bond to the C2 mRNA Nucleotide
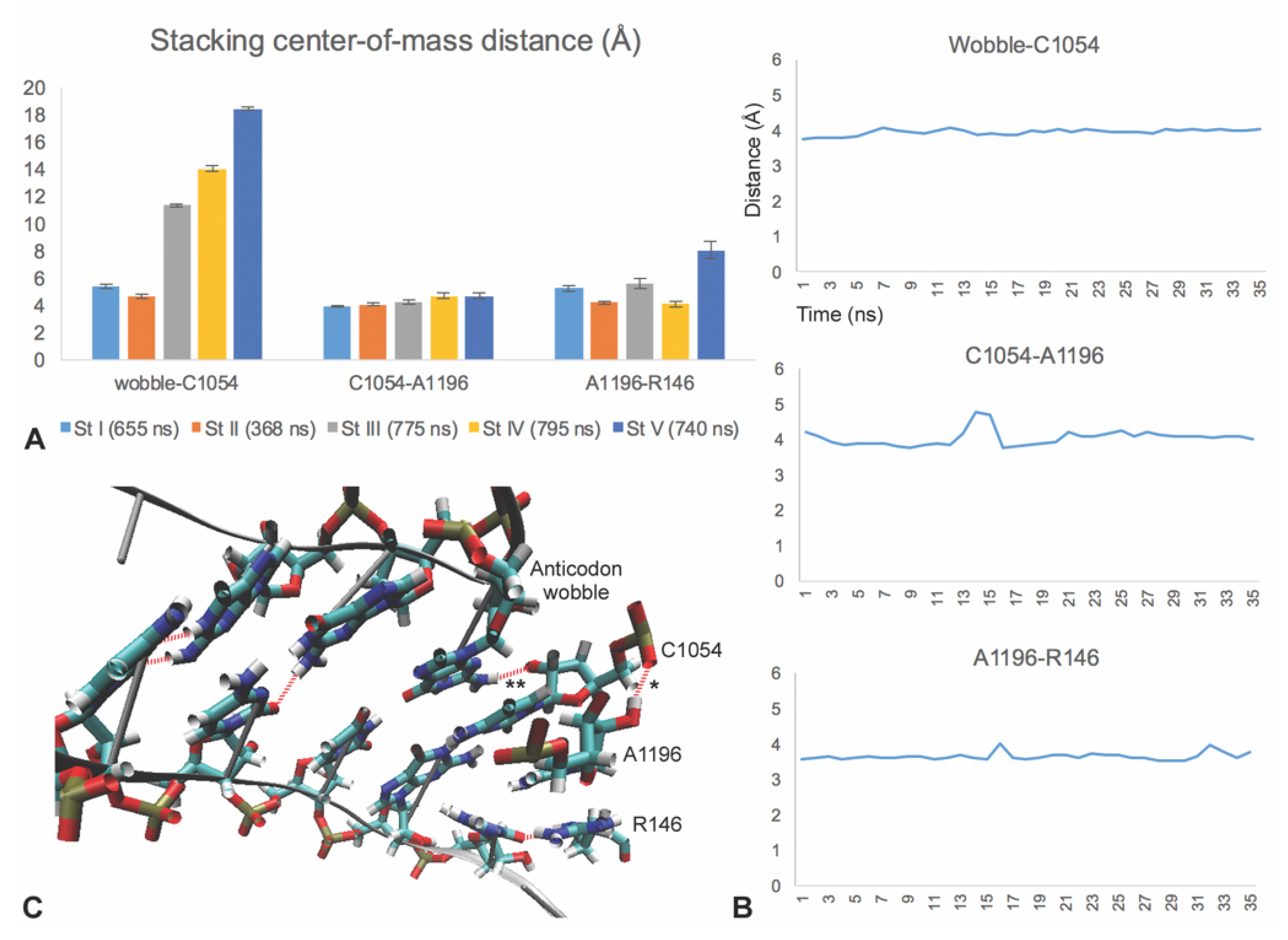
3.4. C1054-A1196-R146 Stacking Defines an mRNA Interaction Surface
3.5. The A-Site tRNA Anchors the C1054-A1196-R146 Interaction Surface
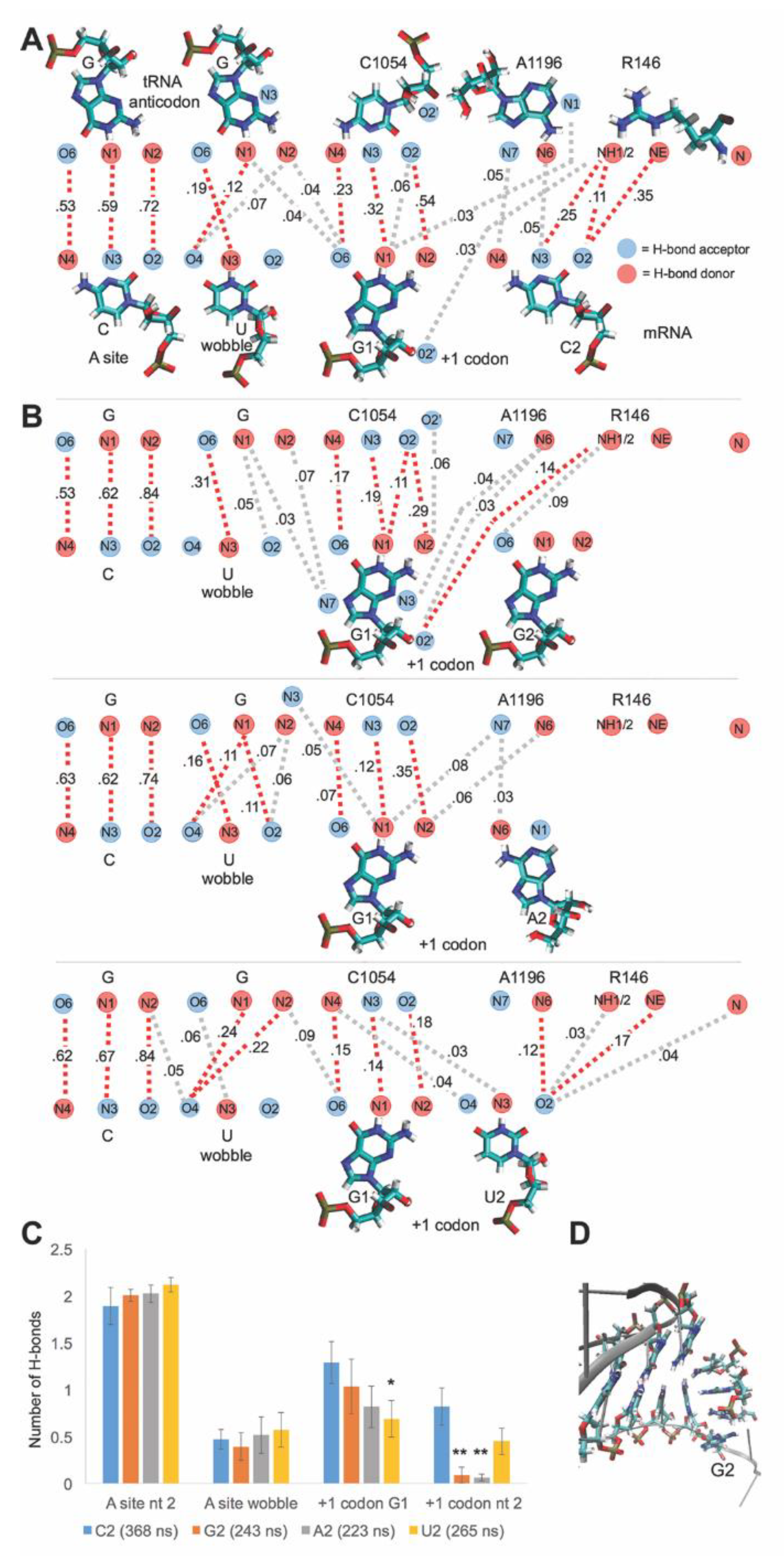
3.6. Sequence Sensitivity of the C1054-A1196-R146 mRNA Interaction
4. Discussion
4.1. The C1054-A1196-R146 (CAR) Interaction Surface
4.2. Wobble Anticodon Nucleotide Anchors the C1054-A1196-R146 Interaction Surface
4.3. Regulation of the mRNA–CAR Interaction
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ratje, A.H.; Loerke, J.; Mikolajka, A.; Brunner, M.; Hildebrand, P.W.; Starosta, A.L.; Donhofer, A.; Connell, S.R.; Fucini, P.; Mielke, T.; et al. Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites. Nature 2010, 468, 713–716. [Google Scholar] [CrossRef] [PubMed]
- Spahn, C.M.; Gomez-Lorenzo, M.G.; Grassucci, R.A.; Jorgensen, R.; Andersen, G.R.; Beckmann, R.; Penczek, P.A.; Ballesta, J.P.; Frank, J. Domain movements of elongation factor eEF2 and the eukaryotic 80S ribosome facilitate tRNA translocation. EMBO J. 2004, 23, 1008–1019. [Google Scholar] [CrossRef] [PubMed]
- Dever, T.E.; Dinman, J.D.; Green, R. Translation elongation and recoding in eukaryotes. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Shoji, S.; Walker, S.E.; Fredrick, K. Ribosomal translocation: One step closer to the molecular mechanism. ACS Chem. Biol. 2009, 4, 93–107. [Google Scholar] [CrossRef] [PubMed]
- Abeyrathne, P.D.; Koh, C.S.; Grant, T.; Grigorieff, N.; Korostelev, A.A. Ensemble cryo-EM uncovers inchworm-like translocation of a viral IRES through the ribosome. Elife 2016, 5. [Google Scholar] [CrossRef]
- Tuller, T.; Carmi, A.; Vestsigian, K.; Navon, S.; Dorfan, Y.; Zaborske, J.; Pan, T.; Dahan, O.; Furman, I.; Pilpel, Y. An evolutionarily conserved mechanism for controlling the efficiency of protein translation. Cell 2010, 141, 344–354. [Google Scholar] [CrossRef]
- Verma, M.; Choi, J.; Cottrell, K.A.; Lavagnino, Z.; Thomas, E.N.; Pavlovic-Djuranovic, S.; Szczesny, P.; Piston, D.W.; Zaher, H.S.; Puglisi, J.D.; et al. A short translational ramp determines the efficiency of protein synthesis. Nat. Commun. 2019, 10, 5774. [Google Scholar] [CrossRef]
- Barr, W.A.; Sheth, R.B.; Kwon, J.; Cho, J.; Glickman, J.W.; Hart, F.; Chatterji, O.K.; Voelkel-Meiman, K.; Krizanc, D.; Thayer, K.M.; et al. GCN sensitive protein translation in yeast. bioRxiv 2020. [Google Scholar] [CrossRef]
- Lagunez-Otero, J.; Trifonov, E.N. mRNA periodical infrastructure complementary to the proof-reading site in the ribosome. J. Biomol. Struct. Dyn. 1992, 10, 455–464. [Google Scholar] [CrossRef]
- Mendoza, L.; Mondragon, M.; Lagunez-Otero, J. Interaction of the 530 ribosomal site with regions of mRNA. Biosystems 1998, 46, 293–298. [Google Scholar] [CrossRef]
- DeLano, W.L. The PyMOL Molecular Graphics System, Version 2.3; Schrödinger, LLC: New York, NY, USA; Available online: https://pymol.org/2/support.html (accessed on 3 May 2020).
- Hsin, J.; Arkhipov, A.; Yin, Y.; Stone, J.E.; Schulten, K. Using VMD: An introductory tutorial. Curr. Protoc. Bioinform. 2008, 14. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visuual molecular dynamics. J. Molec. Graph. 1986, 14, 33–38. [Google Scholar] [CrossRef]
- Leontis, N.B.; Westhof, E. Geometric nomenclature and classification of RNA base pairs. RNA 2001, 7, 499–512. [Google Scholar] [CrossRef] [PubMed]
- Case, D.A.; Babin, V.; Berryman, J.T.; Betz, R.M.; Cai, Q.; Cerutti, D.S.; Cheatham, T.E., III; Darden, T.A.; Duke, R.E.; Gohlke, H.; et al. Amber 14; University of California: San Francisco, CA, USA, 2014. [Google Scholar]
- Case, D.A.; Ben-Shalom, I.Y.; Brozell, S.R.; Cerutti, D.S.; Cheatham, T.E., III; Cruzeiro, V.W.D.; Darden, T.A.; Duke, R.E.; Ghoreishi, D.; Gilson, M.K.; et al. Amber 18; University of California: San Francisco, CA, USA, 2018. [Google Scholar]
- Salomon-Ferrer, R.; Gotz, A.W.; Poole, D.; Le Grand, S.; Walker, R.C. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. explicit solvent particle mesh ewald. J. Chem. Theory. Comput. 2013, 9, 3878–3888. [Google Scholar] [CrossRef] [PubMed]
- Kollman, P.A.; Massova, I.; Reyes, C.; Kuhn, B.; Huo, S.; Chong, L.; Lee, M.; Lee, T.; Duan, Y.; Wang, W.; et al. Calculating structures and free energies of complex molecules: Combining molecular mechanics and continuum models. Acc. Chem. Res. 2000, 33, 889–897. [Google Scholar] [CrossRef]
- Wang, J.; Cieplak, P.; Kollman, P.A. How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J. Comput. Chem. 2000, 21, 1049–1074. [Google Scholar] [CrossRef]
- Salomon-Ferrer, R.; Case, D.A.; Walker, R.C. An overview of the Amber biomolecular simulation package. Interdiscip. Rev. Comput. Mol. Sci. 2013, 3, 198–210. [Google Scholar] [CrossRef]
- Zgarbova, M.; Otyepka, M.; Sponer, J.; Mladek, A.; Banas, P.; Cheatham, T.E., III; Jurecka, P. Refinement of the Cornell et al. Nucleic acids force field based on reference quantum chemical calculations of glycosidic torsion profiles. J. Chem. Theory. Comput. 2011, 7, 2886–2902. [Google Scholar] [CrossRef]
- Goetz, A.W.; Williamson, M.J.; Xu, D.; Poole, D.; Le Grand, S.; Walker, R.C. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. generalized born. J. Chem. Theory. Comput. 2012, 8, 1542–1555. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Jenson, C. Temperature dependence of TIP3P, SPC, and TIP4P water from NPT Monte Carlo simulations: Seeking temperatures of maximum density. J. Comput. Chem. 1998, 19, 1179–1186. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089. [Google Scholar] [CrossRef]
- Berendsen, H.J.C.; Postma, J.P.M.; van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984, 81, 3684–3690. [Google Scholar] [CrossRef]
- Roe, D.R.; Cheatham, T.E., III. PTRAJ and CPPTRAJ: Software for processing and analysis of molecular synamics trajectory data. J. Chem. Theory. Comput. 2013, 9, 3084–3095. [Google Scholar] [CrossRef] [PubMed]
- Sergiev, P.V.; Lavrik, I.N.; Wlasoff, V.A.; Dokudovskaya, S.S.; Dontsova, O.A.; Bogdanov, A.A.; Brimacombe, R. The path of mRNA through the bacterial ribosome: A site-directed crosslinking study using new photoreactive derivatives of guanosine and uridine. RNA 1997, 3, 464–475. [Google Scholar] [PubMed]
- Bhangu, R.; Juzumiene, D.; Wollenzien, P. Arrangement of messenger RNA on Escherichia coli ribosomes with respect to 10 16S rRNA cross-linking sites. Biochemistry 1994, 33, 3063–3070. [Google Scholar] [CrossRef] [PubMed]
- Juzumiene, D.I.; Shapkina, T.G.; Wollenzien, P. Distribution of cross-links between mRNA analogues and 16 S rRNA in Escherichia coli 70 S ribosomes made under equilibrium conditions and their response to tRNA binding. J. Biol. Chem. 1995, 270, 12794–12800. [Google Scholar] [CrossRef]
- Matassova, A.B.; Rodnina, M.V.; Wintermeyer, W. Elongation factor G-induced structural change in helix 34 of 16S rRNA related to translocation on the ribosome. RNA 2001, 7, 1879–1885. [Google Scholar]
- McClory, S.P.; Devaraj, A.; Fredrick, K. Distinct functional classes of ram mutations in 16S rRNA. RNA 2014, 20, 496–504. [Google Scholar] [CrossRef]
- Konstantinidis, T.C.; Patsoukis, N.; Georgiou, C.D.; Synetos, D. Translational fidelity mutations in 18S rRNA affect the catalytic activity of ribosomes and the oxidative balance of yeast cells. Biochemistry 2006, 45, 3525–3533. [Google Scholar] [CrossRef]
- Dong, J.; Aitken, C.E.; Thakur, A.; Shin, B.S.; Lorsch, J.R.; Hinnebusch, A.G. Rps3/uS3 promotes mRNA binding at the 40S ribosome entry channel and stabilizes preinitiation complexes at start codons. Proc. Natl. Acad. Sci. USA 2017, 114, E2126–E2135. [Google Scholar] [CrossRef]
- Llacer, J.L.; Hussain, T.; Marler, L.; Aitken, C.E.; Thakur, A.; Lorsch, J.R.; Hinnebusch, A.G.; Ramakrishnan, V. Conformational differences between open and closed states of the eukaryotic translation initiation complex. Mol. Cell. 2015, 59, 399–412. [Google Scholar] [CrossRef]
- Young, B.D.; Weiss, D.I.; Zurita-Lopez, C.I.; Webb, K.J.; Clarke, S.G.; McBride, A.E. Identification of methylated proteins in the yeast small ribosomal subunit: A role for SPOUT methyltransferases in protein arginine methylation. Biochemistry 2012, 51, 5091–5104. [Google Scholar] [CrossRef]
- Lv, F.; Zhang, T.; Zhou, Z.; Gao, S.; Wong, C.C.; Zhou, J.Q.; Ding, J. Structural basis for Sfm1 functioning as a protein arginine methyltransferase. Cell Discov. 2015, 1, 15037. [Google Scholar] [CrossRef]
- Armstrong, C.T.; Mason, P.E.; Anderson, J.L.; Dempsey, C.E. Arginine side chain interactions and the role of arginine as a gating charge carrier in voltage sensitive ion channels. Sci. Rep. 2016, 6, 21759. [Google Scholar] [CrossRef]
- Vernon, R.M.; Chong, P.A.; Tsang, B.; Kim, T.H.; Bah, A.; Farber, P.; Lin, H.; Forman-Kay, J.D. Pi-Pi contacts are an overlooked protein feature relevant to phase separation. Elife 2018, 7. [Google Scholar] [CrossRef]
- Kumar, K.; Woo, S.M.; Siu, T.; Cortopassi, W.A.; Duarte, F.; Paton, R.S. Cation-pi interactions in protein-ligand binding: Theory and data-mining reveal different roles for lysine and arginine. Chem. Sci. 2018, 9, 2655–2665. [Google Scholar] [CrossRef]
- Gamble, C.E.; Brule, C.E.; Dean, K.M.; Fields, S.; Grayhack, E.J. Adjacent codons act in concert to modulate translation efficiency in yeast. Cell 2016, 166, 679–690. [Google Scholar] [CrossRef]
- Letzring, D.P.; Dean, K.M.; Grayhack, E.J. Control of translation efficiency in yeast by codon-anticodon interactions. Rna 2010, 16, 2516–2528. [Google Scholar] [CrossRef]
- Lin, Y.; May, G.E.; Kready, H.; Nazzaro, L.; Mao, M.; Spealman, P.; Creeger, Y.; McManus, C.J. Impacts of uORF codon identity and position on translation regulation. Nucleic Acids Res. 2019, 47, 9358–9367. [Google Scholar] [CrossRef]
- Ran, W.; Higgs, P.G. The influence of anticodon-codon interactions and modified bases on codon usage bias in bacteria. Mol. Biol. Evol. 2010, 27, 2129–2140. [Google Scholar] [CrossRef]
- Stadler, M.; Fire, A. Wobble base-pairing slows in vivo translation elongation in metazoans. RNA 2011, 17, 2063–2073. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.T.; Dyavaiah, M.; DeMott, M.S.; Taghizadeh, K.; Dedon, P.C.; Begley, T.J. A quantitative systems approach reveals dynamic control of tRNA modifications during cellular stress. PLoS Genet. 2010, 6, e1001247. [Google Scholar] [CrossRef] [PubMed]
- Gu, C.; Begley, T.J.; Dedon, P.C. tRNA modifications regulate translation during cellular stress. FEBS Lett. 2014, 588, 4287–4296. [Google Scholar] [CrossRef]
- Hou, Y.M.; Gamper, H.; Yang, W. Post-transcriptional modifications to tRNA--a response to the genetic code degeneracy. RNA 2015, 21, 642–644. [Google Scholar] [CrossRef]
- Fournier, C.T.; Cherny, J.J.; Truncali, K.; Robbins-Pianka, A.; Lin, M.S.; Krizanc, D.; Weir, M.P. Amino termini of many yeast proteins map to downstream start codons. J. Proteome Res. 2012, 11, 5712–5719. [Google Scholar] [CrossRef]
- Lycette, B.E.; Glickman, J.; Roth, S.J.; Cram, A.E.; Kim, T.H.; Krizanc, D.; Weir, M.P. N-terminal peptide detection with optimized peptide-spectrum matching and streamlined sequence libraries. J. Proteome Res. 2016, 15, 2891–2899. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Scopino, K.; Williams, E.; Elsayed, A.; Barr, W.A.; Krizanc, D.; Thayer, K.M.; Weir, M.P. A Ribosome Interaction Surface Sensitive to mRNA GCN Periodicity. Biomolecules 2020, 10, 849. https://doi.org/10.3390/biom10060849
Scopino K, Williams E, Elsayed A, Barr WA, Krizanc D, Thayer KM, Weir MP. A Ribosome Interaction Surface Sensitive to mRNA GCN Periodicity. Biomolecules. 2020; 10(6):849. https://doi.org/10.3390/biom10060849
Chicago/Turabian StyleScopino, Kristen, Elliot Williams, Abdelrahman Elsayed, William A. Barr, Daniel Krizanc, Kelly M. Thayer, and Michael P. Weir. 2020. "A Ribosome Interaction Surface Sensitive to mRNA GCN Periodicity" Biomolecules 10, no. 6: 849. https://doi.org/10.3390/biom10060849
APA StyleScopino, K., Williams, E., Elsayed, A., Barr, W. A., Krizanc, D., Thayer, K. M., & Weir, M. P. (2020). A Ribosome Interaction Surface Sensitive to mRNA GCN Periodicity. Biomolecules, 10(6), 849. https://doi.org/10.3390/biom10060849





