New Therapeutic Strategies for Osteoarthritis by Targeting Sialic Acid Receptors
Abstract
1. Introduction
2. Materials and Methods
2.1. Cartilage Processing and Primary Culture
2.2. Tissue Culture
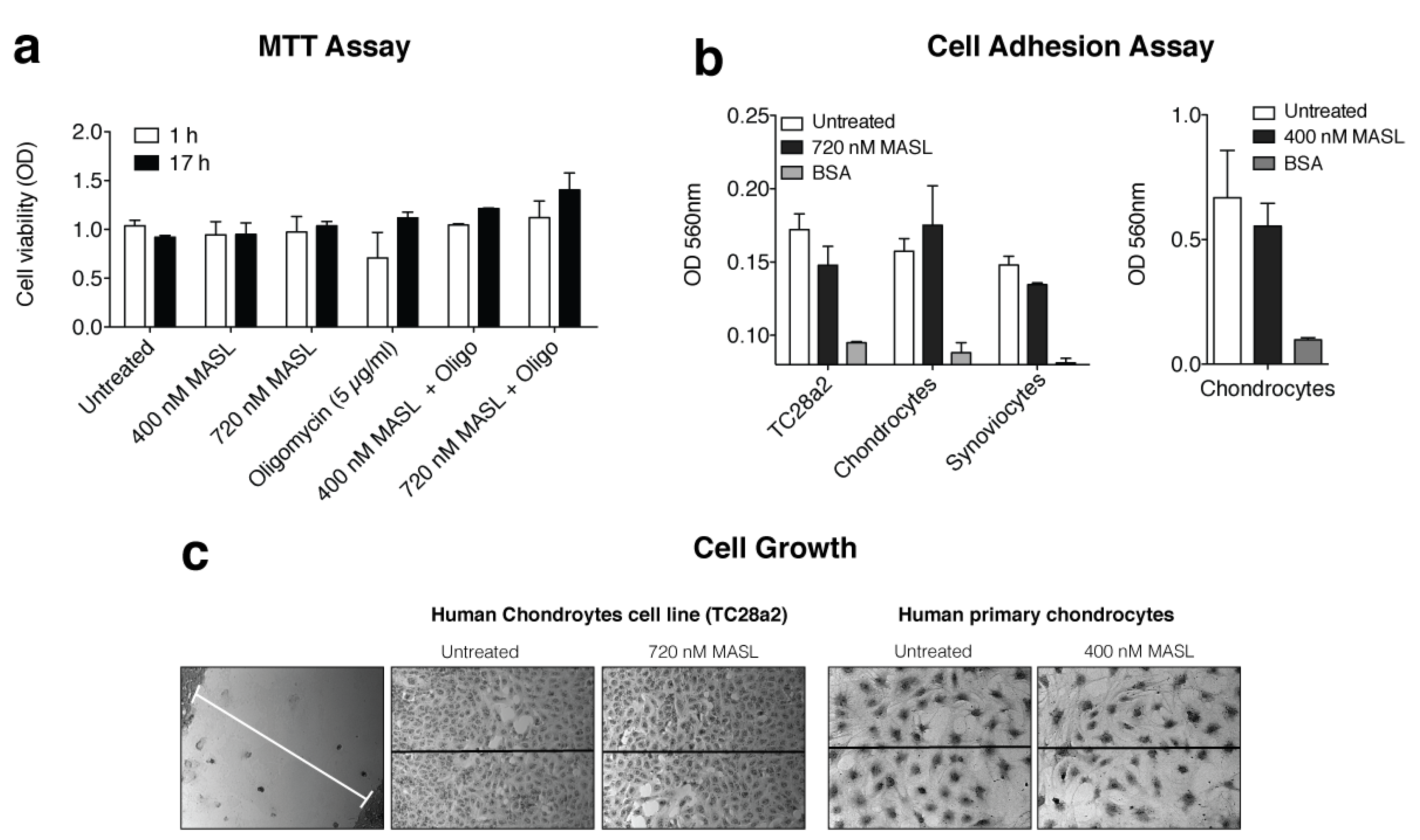
2.3. Cell Viability Assay
2.4. Adhesion Assay
2.5. Cell Growth and Migration
2.6. Animals, Treatments, and Histological Analysis
2.7. Immunohistochemistry and Immunofluorescence
2.8. Western Blot
2.9. Lectin-Binding Analysis
2.10. Quantitative RT-PCR
2.11. Computational Modelling of the MASL Proteins (MAL and MAH)
2.12. Molecular Dynamics (MD) Simulations
2.13. Computational Modelling of the Ligand Neu5Acα(2–3)Galβ-(1–3)[Neu5Acα(2–6)]GalNAc
2.14. Docking Calculations
2.15. Statistical Analysis
3. Results
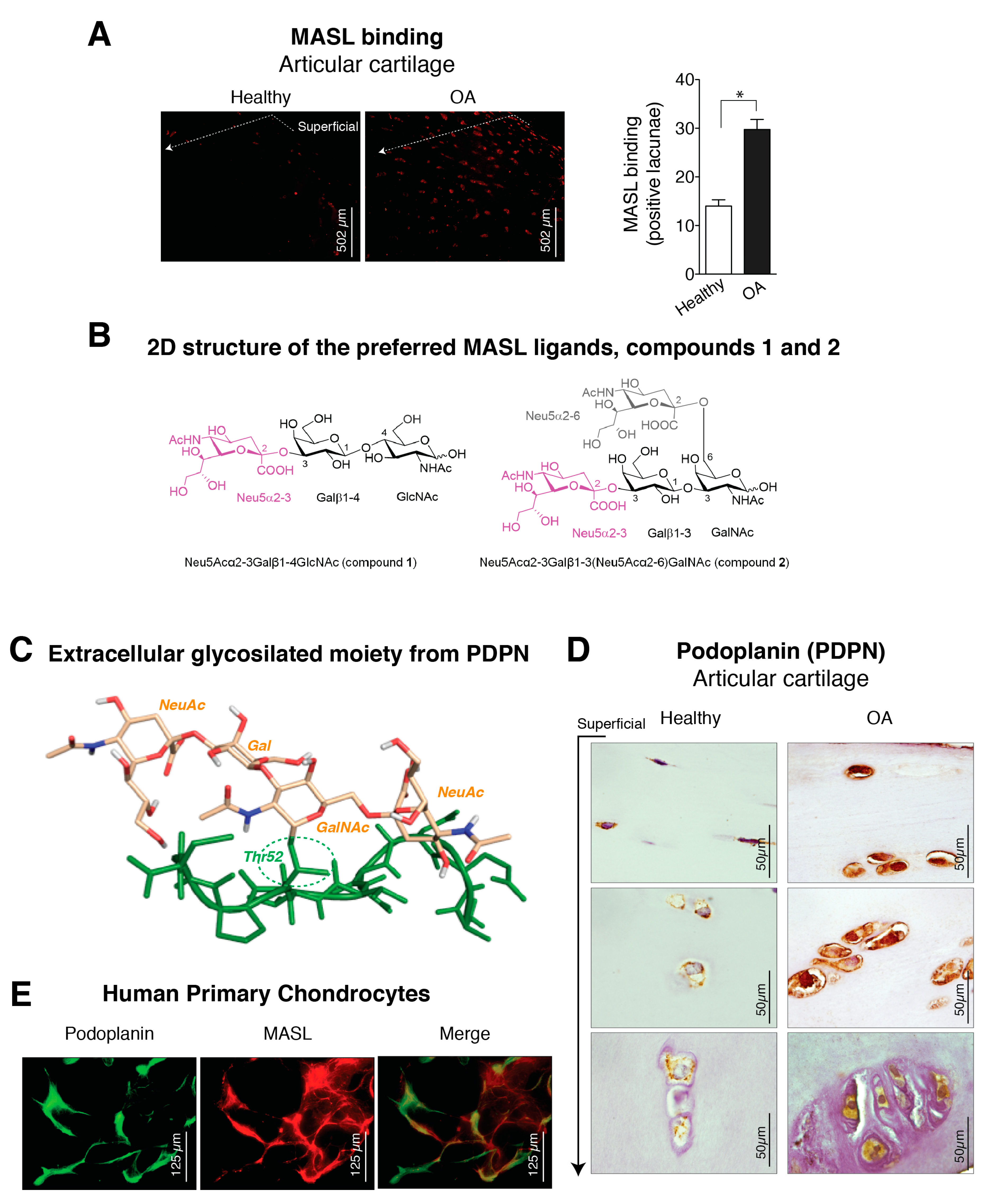
3.1. α-2,3-Sialylated Glycoproteins are Induced in Arthritic Chondrocytes
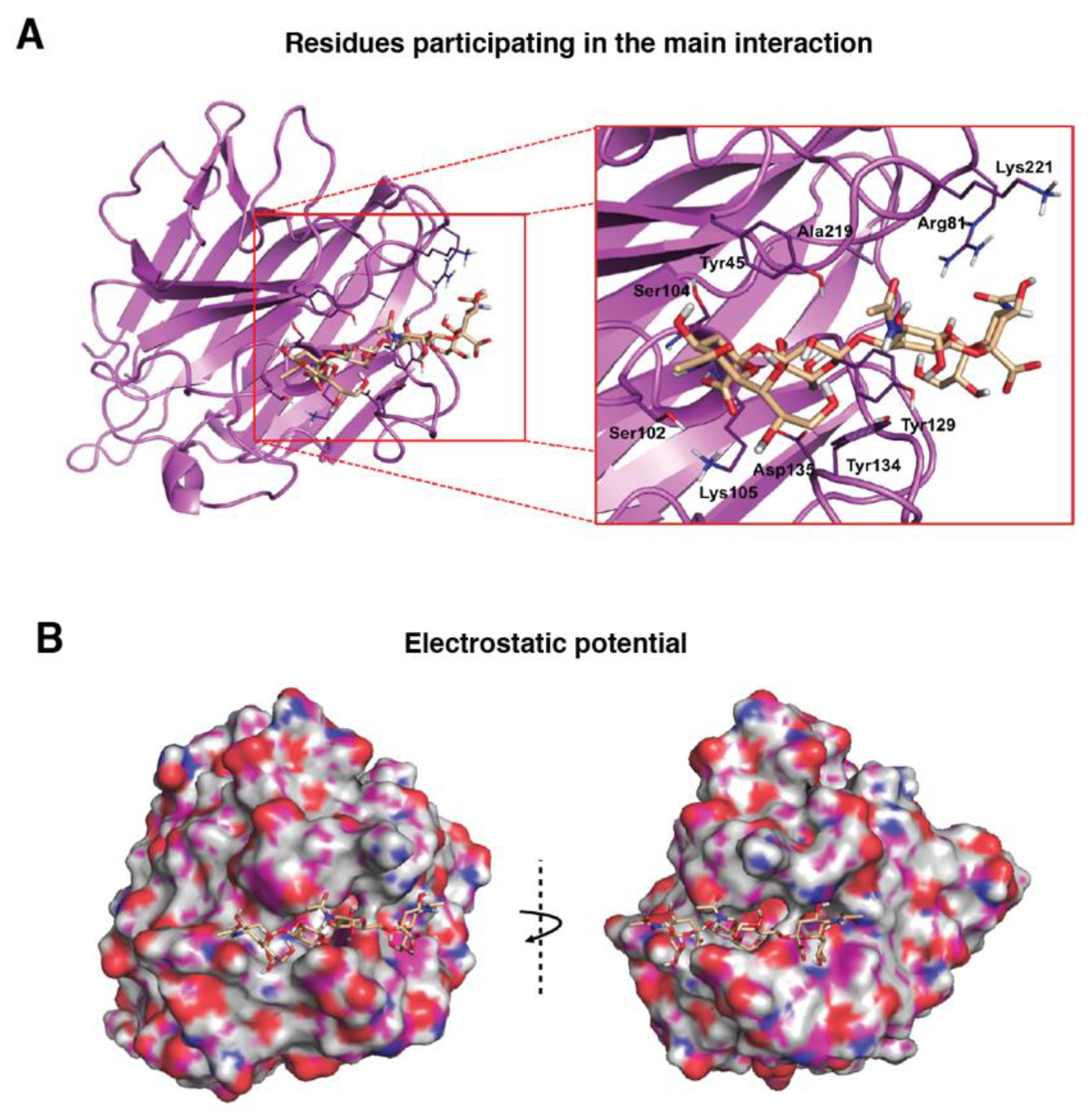
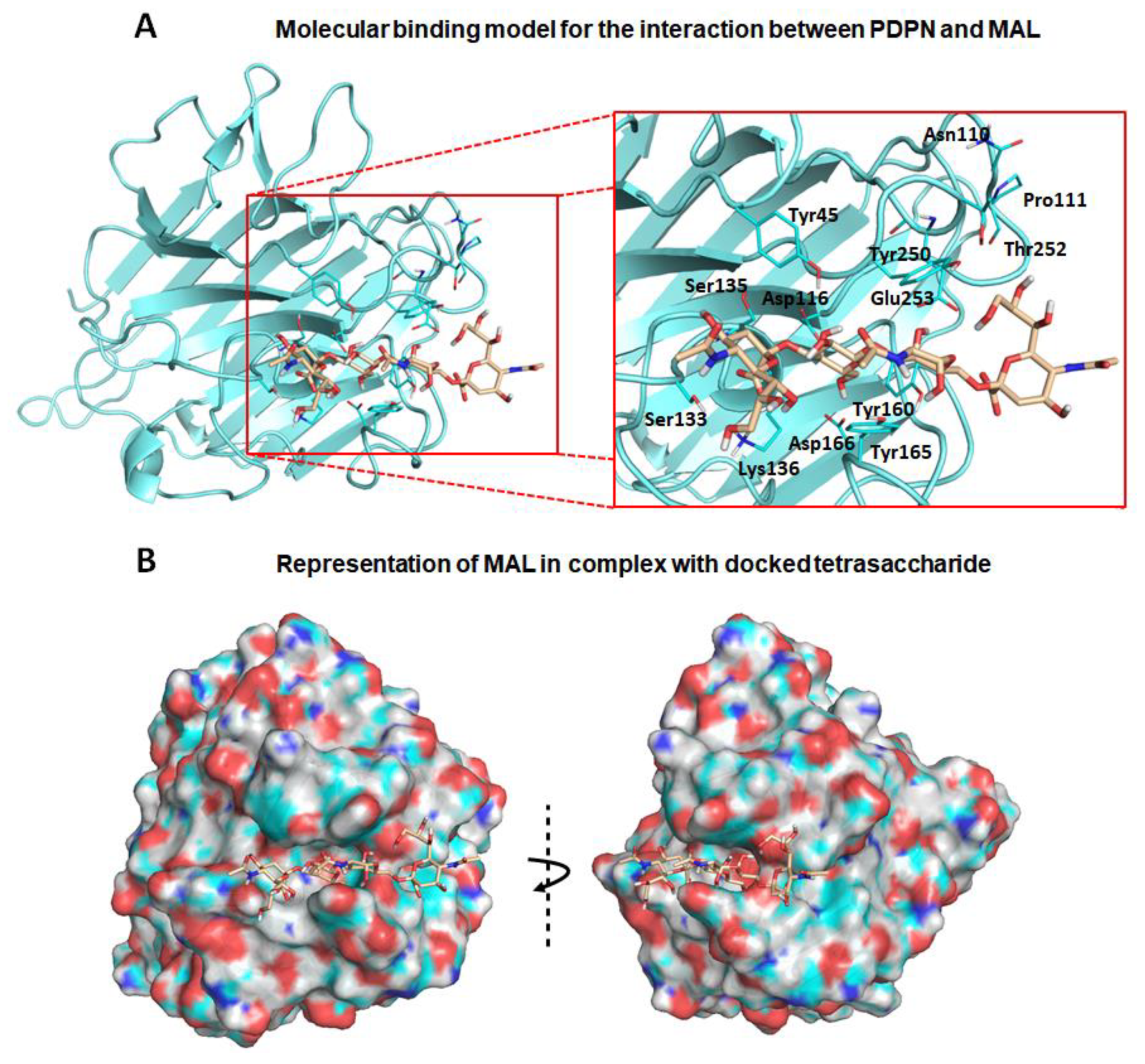
3.2. Molecular Binding Characteristics of the Interaction of PDPN with MASL
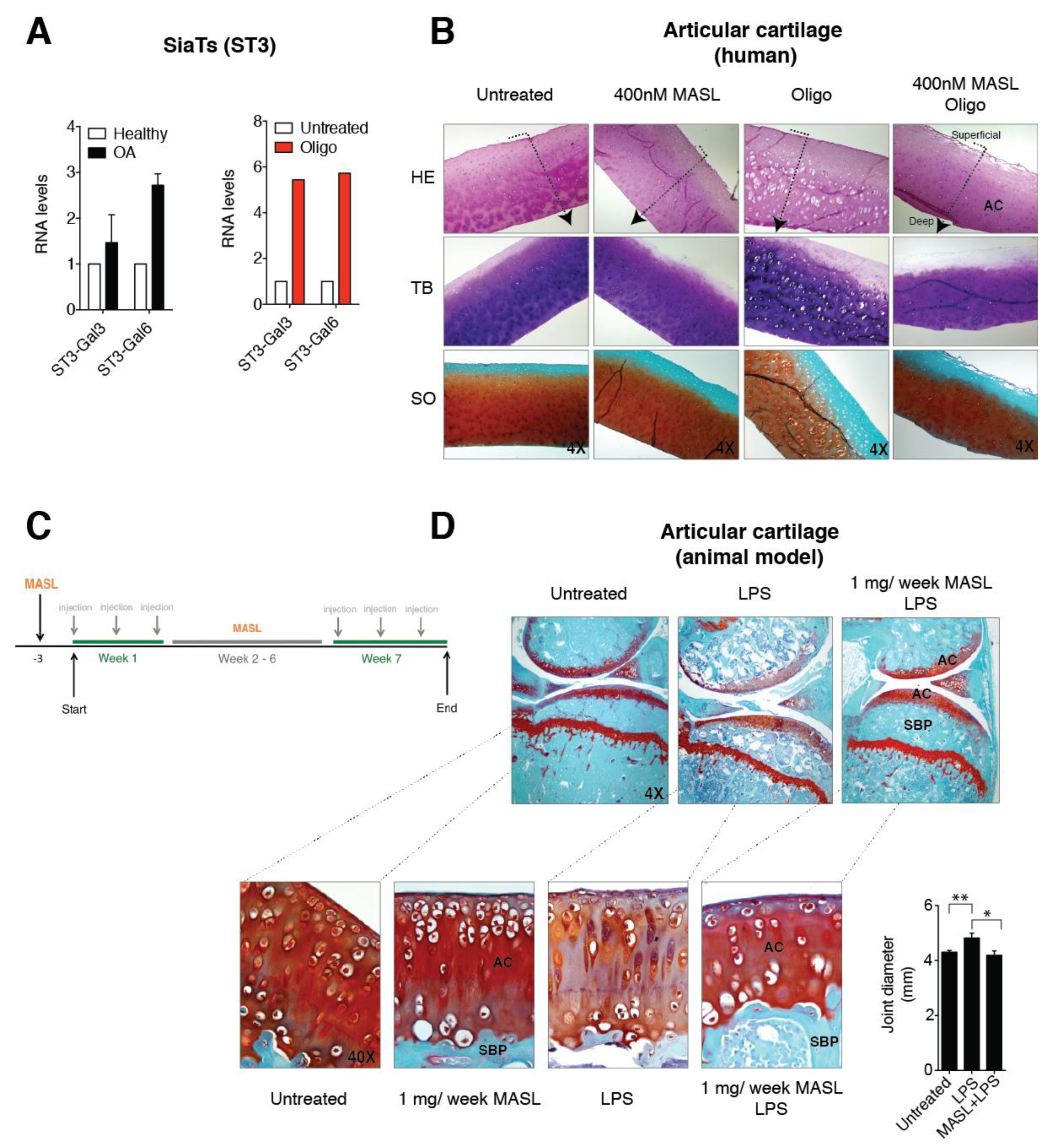
3.3. α-2,3-SiaT Isoforms are Overexpressed in Osteoarthritic Chondrocytes
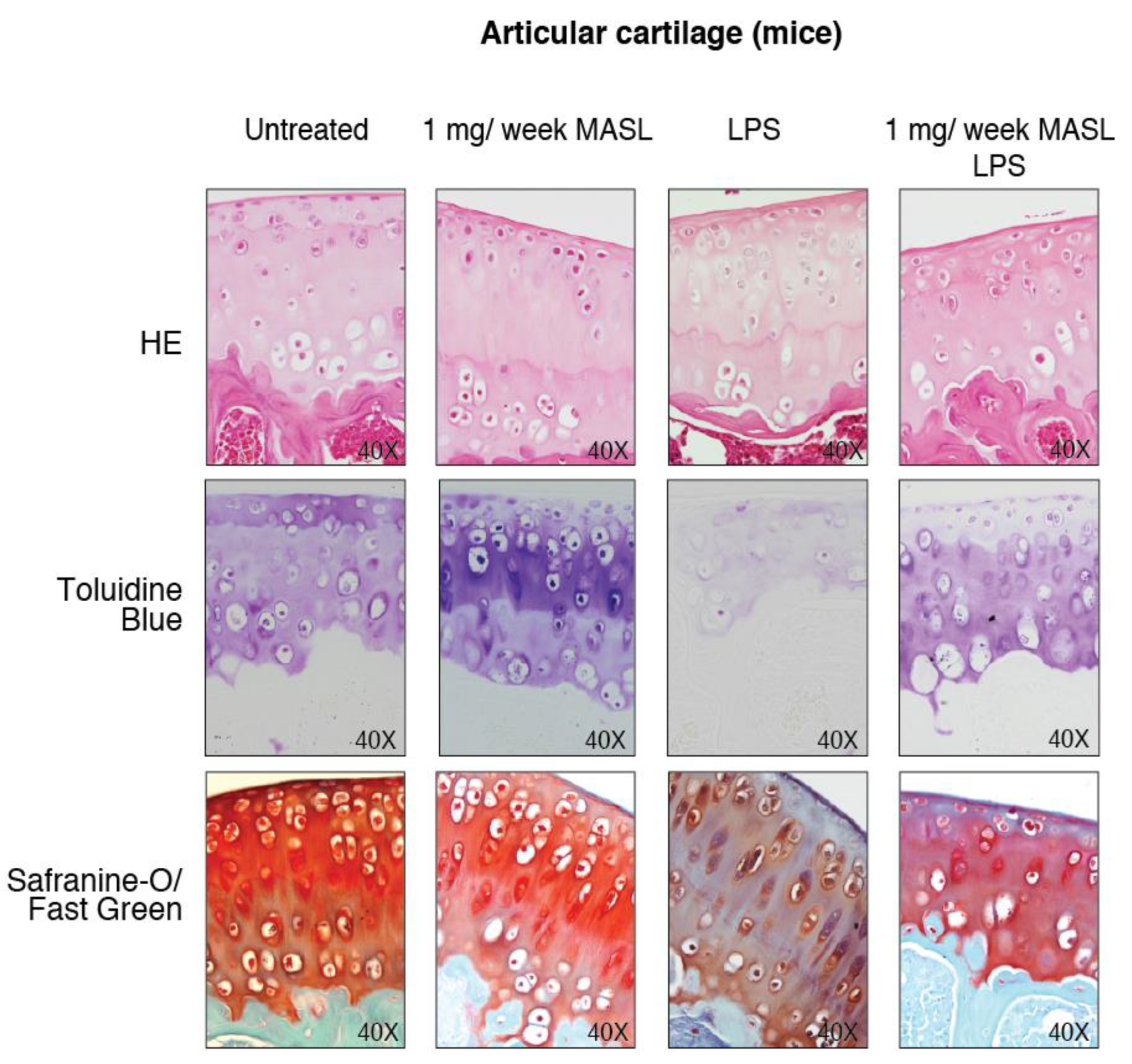
3.4. Effect of MASL in an Ex Vivo and In Vivo Model of OA
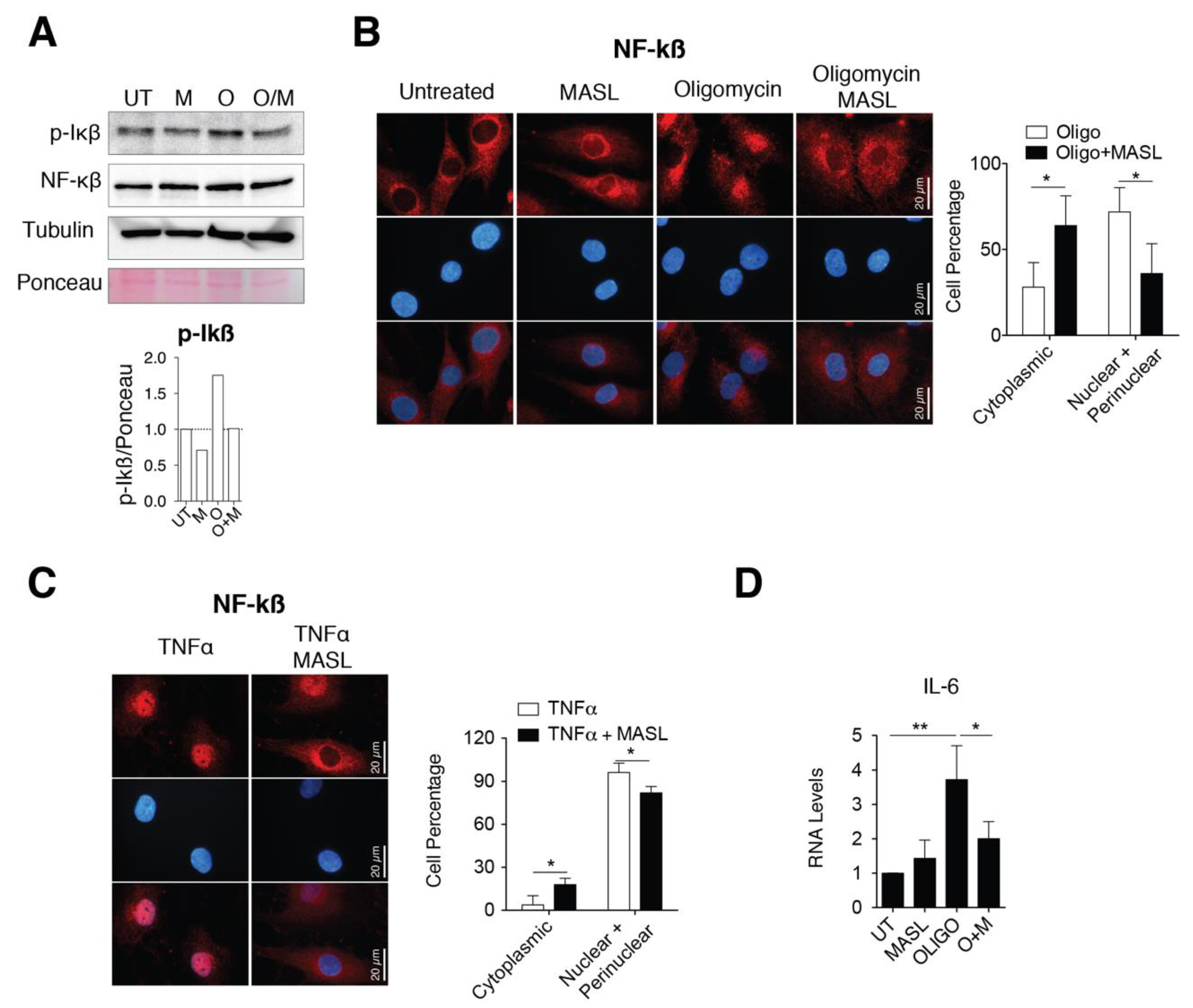
3.5. MASL Prevents Activation of NF-kB Pathway
4. Discussion
5. Conclusions
6. Patents
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A

References
- Loeser, R.F.; Goldring, S.R.; Scanzello, C.R.; Goldring, M.B. Osteoarthritis: A disease of the joint as an organ. Arthritis Rheum. 2012, 64, 1697–1707. [Google Scholar] [CrossRef] [PubMed]
- Moremen, K.W.; Tiemeyer, M.; Nairn, A.V. Vertebrate protein glycosylation: Diversity, synthesis and function. Nat. Rev. Mol. Cell Biol. 2012, 13, 448–462. [Google Scholar] [CrossRef] [PubMed]
- Turner, G.A. N-glycosylation of serum proteins in disease and its investigation using lectins. Clin. Chim. Acta 1992, 208, 149–171. [Google Scholar] [CrossRef]
- Toegel, S.; Pabst, M.; Wu, S.Q.; Grass, J.; Goldring, M.B.; Chiari, C.; Kolb, A.; Altmann, F.; Viernstein, H.; Unger, F.M. Phenotype-related differential alpha-2,6- or alpha-2,3-sialylation of glycoprotein N-glycans in human chondrocytes. Osteoarthr. Cartil. 2010, 18, 240–248. [Google Scholar] [CrossRef]
- Schultz, M.J.; Swindall, A.F.; Bellis, S.L. Regulation of the metastatic cell phenotype by sialylated glycans. Cancer Metastasis Rev. 2012, 31, 501–518. [Google Scholar] [CrossRef]
- Dube, D.H.; Bertozzi, C.R. Glycans in cancer and inflammation--potential for therapeutics and diagnostics. Nat. Rev. Drug Discov. 2005, 4, 477–488. [Google Scholar] [CrossRef]
- Schauer, R. Chemistry, metabolism, and biological functions of sialic acids. Adv. Carbohydr. Chem. Biochem. 1982, 40, 131–234. [Google Scholar]
- Rao, F.V.; Rich, J.R.; Rakic, B.; Buddai, S.; Schwartz, M.F.; Johnson, K.; Bowe, C.; Wakarchuk, W.W.; Defrees, S.; Withers, S.G.; et al. Structural insight into mammalian sialyltransferases. Nat. Struct. Mol. Biol. 2009, 16, 1186–1188. [Google Scholar]
- Harduin-Lepers, A.; Vallejo-Ruiz, V.; Krzewinski-Recchi, M.A.; Samyn-Petit, B.; Julien, S.; Delannoy, P. The human sialyltransferase family. Biochimie 2001, 83, 727–737. [Google Scholar] [CrossRef]
- Toegel, S.; Bieder, D.; Andre, S.; Altmann, F.; Walzer, S.M.; Kaltner, H.; Hofstaetter, J.G.; Windhager, R.; Gabius, H.J. Glycophenotyping of osteoarthritic cartilage and chondrocytes by RT-qPCR, mass spectrometry, histochemistry with plant/human lectins and lectin localization with a glycoprotein. Arthritis Res. Ther. 2013, 15, R147. [Google Scholar] [CrossRef]
- Toegel, S.; Bieder, D.; Andre, S.; Kayser, K.; Walzer, S.M.; Hobusch, G.; Windhager, R.; Gabius, H.J. Human osteoarthritic knee cartilage: Fingerprinting of adhesion/growth-regulatory galectins in vitro and in situ indicates differential upregulation in severe degeneration. Histochem. Cell Biol. 2014, 142, 373–388. [Google Scholar] [CrossRef] [PubMed]
- Varki, A.; Etzler, M.E.; Cummings, R.D.; Esko, J.D. Discovery and Classification of Glycan-Binding Proteins. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring: Harbor, NY, USA, 2009. [Google Scholar]
- Solis, D.; Bovin, N.V.; Davis, A.P.; Jimenez-Barbero, J.; Romero, A.; Roy, R.; Smetana, K., Jr.; Gabius, H.J. A guide into glycosciences: How chemistry, biochemistry and biology cooperate to crack the sugar code. Biochim. Biophys. Acta 2015, 1850, 186–235. [Google Scholar] [CrossRef] [PubMed]
- Gabius, H.J. The magic of the sugar code. Trends Biochem. Sci. 2015, 40, 341. [Google Scholar] [CrossRef] [PubMed]
- Reesink, H.L.; Bonnevie, E.D.; Liu, S.; Shurer, C.R.; Hollander, M.J.; Bonassar, L.J.; Nixon, A.J. Galectin-3 Binds to Lubricin and Reinforces the Lubricating Boundary Layer of Articular Cartilage. Sci. Rep. 2016, 6, 25463. [Google Scholar] [CrossRef] [PubMed]
- Rabinovich, G.A.; Toscano, M.A. Turning ‘sweet’ on immunity: Galectin-glycan interactions in immune tolerance and inflammation. Nat. Rev. Immunol. 2009, 9, 338–352. [Google Scholar] [CrossRef]
- RodrIguez, E.; Schetters, S.T.T.; Van Kooyk, Y. The tumour glyco-code as a novel immune checkpoint for immunotherapy. Nat. Rev. Immunol. 2018, 18, 204–211. [Google Scholar] [CrossRef]
- Zhou, J.Y.; Oswald, D.M.; Oliva, K.D.; Kreisman, L.S.C.; Cobb, B.A. The Glycoscience of Immunity. Trends Immunol. 2018, 39, 523–535. [Google Scholar] [CrossRef]
- Cueni, L.N.; Detmar, M. Galectin-8 interacts with podoplanin and modulates lymphatic endothelial cell functions. Exp. Cell Res. 2009, 315, 1715–1723. [Google Scholar] [CrossRef]
- Chen, W.S.; Cao, Z.; Sugaya, S.; Lopez, M.J.; Sendra, V.G.; Laver, N.; Leffler, H.; Nilsson, U.J.; Fu, J.; Song, J.; et al. Pathological lymphangiogenesis is modulated by galectin-8-dependent crosstalk between podoplanin and integrin-associated VEGFR-3. Nat. Commun. 2016, 7, 11302. [Google Scholar] [CrossRef] [PubMed]
- Hasan, S.S.; Ashraf, G.M.; Banu, N. Galectins—Potential targets for cancer therapy. Cancer Lett. 2007, 253, 25–33. [Google Scholar] [CrossRef]
- Pusztai, A.; Bardocz, S.; Ewen, S.W. Uses of plant lectins in bioscience and biomedicine. Front. Biosci. 2008, 13, 1130–1140. [Google Scholar] [CrossRef] [PubMed]
- Coulibaly, F.Y. Bi-Botti, Current status of lectin-based cancer diagnosis and therapy. AIMS Mol. Sci. 2017, 4, 1–27. [Google Scholar]
- Geisler, C.; Jarvis, D.L. Effective glycoanalysis with Maackia amurensis lectins requires a clear understanding of their binding specificities. Glycobiology 2011, 21, 988–993. [Google Scholar] [CrossRef] [PubMed]
- Xiong, L.; Jianfang, L.; Dong, W.; Weining, W.; Zheng, C. Chromone and flavonoids from Maackia amurensis. Asian J. Tradit. Med. 2009, 4, 98–103. [Google Scholar]
- Wang, W.C.; Cummings, R.D. The immobilized leukoagglutinin from the seeds of Maackia amurensis binds with high affinity to complex-type Asn-linked oligosaccharides containing terminal sialic acid-linked alpha-2,3 to penultimate galactose residues. J. Biol. Chem. 1988, 263, 4576–4585. [Google Scholar]
- Imberty, A.; Gautier, C.; Lescar, J.; Perez, S.; Wyns, L.; Loris, R. An unusual carbohydrate binding site revealed by the structures of two Maackia amurensis lectins complexed with sialic acid-containing oligosaccharides. J. Biol. Chem. 2000, 275, 17541–17548. [Google Scholar] [CrossRef]
- Lehmann, K.; Janda, E.; Pierreux, C.E.; Ryt√∂maa, M.; Schulze, A.; McMahon, M.; Hill, C.S.; Beug, H.; Downward, J. Raf induces TGFbeta production while blocking its apoptotic but not invasive responses: A mechanism leading to increased malignancy in epithelial cells. Genes Dev. 2000, 14, 2610–2622. [Google Scholar] [CrossRef]
- Kawaguchi, T.; Matsumoto, I.; Osawa, T. Studies on hemagglutinins from Maackia amurensis seeds. J. Biol. Chem. 1974, 249, 2786–2792. [Google Scholar]
- Ochoa-Alvarez, J.A.; Krishnan, H.; Shen, Y.; Acharya, N.K.; Han, M.; McNulty, D.E.; Hasegawa, H.; Hyodo, T.; Senga, T.; Geng, J.G.; et al. Plant lectin can target receptors containing sialic acid, exemplified by podoplanin, to inhibit transformed cell growth and migration. PLoS One 2012, 7, e41845. [Google Scholar] [CrossRef]
- Del Rey, M.J.; Izquierdo, E.; Faré, R.; Miranda, V.; Criado, G.; Cañete, J.D.; Pablos, J.L. Podoplanin-Mediated Interaction Of Rheumatoid Arthritis Synovial Fibroblasts With Platelets Modulates IL-8 Expression. Arthritis Rheum. 2012, 64 (Suppl. 10), 1183. [Google Scholar]
- Ekwall, A.K.; Eisler, T.; Anderberg, C.; Jin, C.; Karlsson, N.; Brisslert, M.; Bokarewa, M.I. The tumour-associated glycoprotein podoplanin is expressed in fibroblast-like synoviocytes of the hyperplastic synovial lining layer in rheumatoid arthritis. Arthritis Res. Ther. 2011, 13, R40. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, Y.; Uga, H.; Tanaka, S.; Kadowaki, M.; Ikeda, M.; Saegusa, J.; Morinobu, A.; Kumagai, S.; Kurata, H. Podoplanin is an inflammatory protein upregulated in Th17 cells in SKG arthritic joints. Mol. Immunol. 2013, 54, 199–207. [Google Scholar] [CrossRef] [PubMed]
- Faré, R.; Izquierdo, E.; Del Rey, M.J.; Usategui, A.; Celis, R.; Cañete, J.D.; Pablos, J.L. Podoplanin Expression In Rheumatoid Stroma Correlates With Lymphoid Neogenesis And Is Downregulated By Anti-TNF- Therapy. Arthritis Rheum. 2012, 64 (Suppl. 10), 1193. [Google Scholar]
- Wicki, A.; Christofori, G. The potential role of podoplanin in tumour invasion. Br. J. Cancer 2007, 96, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Astarita, J.L.; Acton, S.E.; Turley, S.J. Podoplanin: Emerging functions in development, the immune system, and cancer. Front. Immunol. 2012, 3, 283. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, H.; Ochoa-Alvarez, J.A.; Shen, Y.; Nevel, E.; Lakshminarayanan, M.; Williams, M.C.; Ramirez, M.I.; Miller, W.T.; Goldberg, G.S. Serines in the intracellular tail of podoplanin (PDPN) regulate cell motility. J. Biol. Chem. 2013, 288, 12215–12221. [Google Scholar] [CrossRef]
- Bertozzi, C.C.; Schmaier, A.A.; Mericko, P.; Hess, P.R.; Zou, Z.; Chen, M.; Chen, C.Y.; Xu, B.; Lu, M.M.; Zhou, D.; et al. Platelets regulate lymphatic vascular development through CLEC-2-SLP-76 signaling. Blood 2010, 116, 661–670. [Google Scholar] [CrossRef]
- Watson, S.P.; Herbert, J.M.; Pollitt, A.Y. GPVI and CLEC-2 in hemostasis and vascular integrity. J. Thromb. Haemost. 2010, 8, 1456–1467. [Google Scholar] [CrossRef]
- Mourao-Sa, D.; Robinson, M.J.; Zelenay, S.; Sancho, D.; Chakravarty, P.; Larsen, R.; Plantinga, M.; Van Rooijen, N.; Soares, M.P.; Lambrecht, B.; et al. CLEC-2 signaling via Syk in myeloid cells can regulate inflammatory responses. Eur. J. Immunol. 2011, 41, 3040–3053. [Google Scholar] [CrossRef]
- Suzuki-Inoue, K.; Inoue, O.; Ozaki, Y. Novel platelet activation receptor CLEC-2: From discovery to prospects. J. Thromb. Haemost. 2011, 9 (Suppl. 1), 44–55. [Google Scholar] [CrossRef]
- Case, D.A.; Darden, T.A.; Cheatham, T.E., III; Simmerling, C.L.; Wang, J.; Duke, R.E.; Luo, R.; Walker, R.C.; Zhang, W.; Merz, K.M.; et al. AMBER 12; University of California: San Francisco, CA, USA, 2012. [Google Scholar]
- Mayan, M.D.; Blanco, F.J.; Carpintero-Fernandez, P.; Goldberg, G.S. Compositions and Methods to Treat Inflammatory Joint Diseases. U.S. Patent PCT/US2014/045229; (Publication number WO 2015/003049), 2 July 2014. [Google Scholar]
- Mayan, M.D.; Gago-Fuentes, R.; Carpintero-Fernandez, P.; Fernandez-Puente, P.; Filgueira-Fernandez, P.; Goyanes, N.; Valiunas, V.; Brink, P.R.; Goldberg, G.S.; Blanco, F.J. Articular chondrocyte network mediated by gap junctions: Role in metabolic cartilage homeostasis. Ann. Rheum. Dis. 2015, 74, 275–284. [Google Scholar] [CrossRef] [PubMed]
- Mayan, M.D.; Carpintero-Fernandez, P.; Gago-Fuentes, R.; Martinez-de-Ilarduya, O.; Wang, H.Z.; Valiunas, V.; Brink, P.; Blanco, F.J. Human articular chondrocytes express multiple gap junction proteins: Differential expression of connexins in normal and osteoarthritic cartilage. Am. J. Pathol. 2013, 182, 1337–1346. [Google Scholar] [CrossRef] [PubMed]
- Levkowitz, G.; Waterman, H.; Ettenberg, S.A.; Katz, M.; Tsygankov, A.Y.; Alroy, I.; Lavi, S.; Iwai, K.; Reiss, Y.; Ciechanover, A.; et al. Ubiquitin ligase activity and tyrosine phosphorylation underlie suppression of growth factor signaling by c-Cbl/Sli-1. Mol. Cell 1999, 4, 1029–1040. [Google Scholar] [CrossRef]
- Case, D.B.V.; Berryman, J.; Betz, M.R.; Cai, Q.; Cerutti, S.D.; Cheatham, T.; Darden, T.; Duke, R.; Gohlke, H.; Götz, A.; et al. AMBER 2014; University of California: San Francisco, CA, USA, 2014. [Google Scholar]
- Nagae, M.; Morita-Matsumoto, K.; Kato, M.; Kaneko, M.K.; Kato, Y.; Yamaguchi, Y. A platform of C-type lectin-like receptor CLEC-2 for binding O-glycosylated podoplanin and nonglycosylated rhodocytin. Structure 2014, 22, 1711–1721. [Google Scholar] [CrossRef] [PubMed]
- Van Damme, E.J.; Van Leuven, F.; Peumans, W.J. Isolation, characterization and molecular cloning of the bark lectins from Maackia amurensis. Glycoconj. J. 1997, 14, 449–456. [Google Scholar] [CrossRef]
- Yamamoto, K.; Konami, Y.; Irimura, T. Sialic acid-binding motif of Maackia amurensis lectins. J. Biochem 1997, 121, 756–761. [Google Scholar] [CrossRef]
- Yamamoto, K.; Ishida, C.; Saito, M.; Konami, Y.; Osawa, T.; Irimura, T. Cloning and sequence analysis of the Maackia amurensis haemagglutinin cDNA. Glycoconj. J. 1994, 11, 572–575. [Google Scholar] [CrossRef]
- Konami, Y.; Yamamoto, K.; Osawa, T.; Irimura, T. Strong affinity of Maackia amurensis hemagglutinin (MAH) for sialic acid-containing Ser/Thr-linked carbohydrate chains of N-terminal octapeptides from human glycophorin A. FEBS Lett. 1994, 342, 334–338. [Google Scholar] [CrossRef]
- Knibbs, R.N.; Goldstein, I.J.; Ratcliffe, R.M.; Shibuya, N. Characterization of the carbohydrate binding specificity of the leukoagglutinating lectin from Maackia amurensis. Comparison with other sialic acid-specific lectins. J. Biol. Chem. 1991, 266, 83–88. [Google Scholar]
- M’kacher, R.; El Maalouf, E.; Terzoudi, G.; Ricoul, M.; Heidingsfelder, L.; Karachristou, I.; Laplagne, E.; Hempel, W.M.; Colicchio, B.; Dieterlen, A.; et al. Detection and Automated Scoring of Dicentric Chromosomes in Nonstimulated Lymphocyte Prematurely Condensed Chromosomes After Telomere and Centromere Staining. Int. J. Radiat. Oncol. 2015, 91, 640–649. [Google Scholar] [CrossRef]
- Kaneko, M.K.; Kato, Y.; Kameyama, A.; Ito, H.; Kuno, A.; Hirabayashi, J.; Kubota, T.; Amano, K.; Chiba, Y.; Hasegawa, Y.; et al. Functional glycosylation of human podoplanin: Glycan structure of platelet aggregation-inducing factor. FEBS Lett. 2007, 581, 331–336. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Xie, G.; Wang, W. Reactive oxygen species: The 2-edged sword of osteoarthritis. Am. J. Med. Sci. 2012, 344, 486–490. [Google Scholar] [CrossRef] [PubMed]
- Kundu, S.; Ghosh, P.; Datta, S.; Ghosh, A.; Chattopadhyay, S.; Chatterjee, M. Oxidative stress as a potential biomarker for determining disease activity in patients with rheumatoid arthritis. Free Radic. Res. 2012, 46, 1482–1489. [Google Scholar] [CrossRef] [PubMed]
- Rosas, L.E.; Elgamal, O.A.; Mo, X.; Phelps, M.A.; Schmittgen, T.D.; Papenfuss, T.L. In vitro immunotoxicity assessment of culture-derived extracellular vesicles in human monocytes. J. Immunotoxicol. 2016, 13, 652–665. [Google Scholar] [CrossRef] [PubMed]
- Rigoglou, S.; Papavassiliou, A.G. The NF-kappaB signalling pathway in osteoarthritis. Int. J. Biochem. Cell Biol. 2013, 45, 2580–2584. [Google Scholar] [CrossRef] [PubMed]
- Olivotto, E.; Otero, M.; Marcu, K.B.; Goldring, M.B. Pathophysiology of osteoarthritis: Canonical NF-kappaB/IKKbeta-dependent and kinase-independent effects of IKKalpha in cartilage degradation and chondrocyte differentiation. RMD Open 2015, 1 (Suppl. 1), e000061. [Google Scholar] [CrossRef]
- Onder, L.; Nindl, V.; Scandella, E.; Chai, Q.; Cheng, H.W.; Caviezel-Firner, S.; Novkovic, M.; Bomze, D.; Maier, R.; Mair, F.; et al. Alternative NF-kappaB signaling regulates mTEC differentiation from podoplanin-expressing precursors in the cortico-medullary junction. Eur. J. Immunol. 2015, 45, 2218–2231. [Google Scholar] [CrossRef]
- Lawrence, T. The nuclear factor NF-kappaB pathway in inflammation. Cold Spring Harb. Perspect. Biol. 2009, 1, a001651. [Google Scholar] [CrossRef]
- Glasson, S.S.; Chambers, M.G.; Van Den Berg, W.B.; Little, C.B. The OARSI histopathology initiative—Recommendations for histological assessments of osteoarthritis in the mouse. Osteoarthr. Cartil. 2010, 18 (Suppl. 3), S17–S23. [Google Scholar] [CrossRef]
- Chambers, M.G.; Bayliss, M.T.; Mason, R.M. Chondrocyte cytokine and growth factor expression in murine osteoarthritis. Osteoarthr. Cartil. 1997, 5, 301–308. [Google Scholar] [CrossRef]
- Toegel, S.; Weinmann, D.; Andre, S.; Walzer, S.M.; Bilban, M.; Schmidt, S.; Chiari, C.; Windhager, R.; Krall, C.; Bennani-Baiti, I.M.; et al. Galectin-1 Couples Glycobiology to Inflammation in Osteoarthritis through the Activation of an NF-kappaB-Regulated Gene Network. J Immunol 2016, 196, 1910–1921. [Google Scholar] [CrossRef] [PubMed]
- Osorio, J. Osteoarthritis: Galectin-1 damages cartilage via inflammation. Nat. Rev. Rheumatol. 2016, 12, 132. [Google Scholar] [CrossRef] [PubMed]
- Pabst, M.; Wu, S.Q.; Grass, J.; Kolb, A.; Chiari, C.; Viernstein, H.; Unger, F.M.; Altmann, F.; Toegel, S. IL-1beta and TNF-alpha alter the glycophenotype of primary human chondrocytes in vitro. Carbohydr. Res. 2010, 345, 1389–1393. [Google Scholar] [CrossRef] [PubMed]
- Honma, M.; Minami-Hori, M.; Takahashi, H.; Iizuka, H. Podoplanin expression in wound and hyperproliferative psoriatic epidermis: Regulation by TGF-beta and STAT-3 activating cytokines, IFN-gamma, IL-6, and IL-22. J. Dermatol. Sci. 2012, 65, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Renart, J.; Carrasco-Ramirez, P.; Fernandez-Munoz, B.; Martin-Villar, E.; Montero, L.; Yurrita, M.M.; Quintanilla, M. New insights into the role of podoplanin in epithelial-mesenchymal transition. Int. Rev. Cell Mol. Biol. 2015, 317, 185–239. [Google Scholar] [PubMed]
- Martin-Villar, E.; Borda-d’Agua, B.; Carrasco-Ramirez, P.; Renart, J.; Parsons, M.; Quintanilla, M.; Jones, G.E. Podoplanin mediates ECM degradation by squamous carcinoma cells through control of invadopodia stability. Oncogene 2015, 34, 4531–4544. [Google Scholar] [CrossRef]
- Krishnan, H.; Retzbach, E.P.; Ramirez, M.I.; Liu, T.; Li, H.; Miller, W.T.; Goldberg, G.S. PKA and CDK5 can phosphorylate specific serines on the intracellular domain of podoplanin (PDPN) to inhibit cell motility. Exp. Cell Res. 2015, 335, 115–122. [Google Scholar] [CrossRef]
- Ochoa-Alvarez, J.A.; Krishnan, H.; Pastorino, J.G.; Nevel, E.; Kephart, D.; Lee, J.J.; Retzbach, E.P.; Shen, Y.; Fatahzadeh, M.; Baredes, S.; et al. Antibody and lectin target podoplanin to inhibit oral squamous carcinoma cell migration and viability by distinct mechanisms. Oncotarget 2015, 6, 9045–9060. [Google Scholar] [CrossRef]
- Kato, Y.; Kunita, A.; Abe, S.; Ogasawara, S.; Fujii, Y.; Oki, H.; Fukayama, M.; Nishioka, Y.; Kaneko, M.K. The chimeric antibody chLpMab-7 targeting human podoplanin suppresses pulmonary metastasis via ADCC and CDC rather than via its neutralizing activity. Oncotarget 2015, 6, 36003. [Google Scholar] [CrossRef]
- Tsuneki, M.; Maruyama, S.; Yamazaki, M.; Cheng, J.; Saku, T. Podoplanin expression profiles characteristic of odontogenic tumor-specific tissue architectures. Pathol. Res. Prac. 2012, 208, 140–146. [Google Scholar] [CrossRef]
- Shindo, K.; Aishima, S.; Ohuchida, K.; Fujiwara, K.; Fujino, M.; Mizuuchi, Y.; Hattori, M.; Mizumoto, K.; Tanaka, M.; Oda, Y. Podoplanin expression in cancer-associated fibroblasts enhances tumor progression of invasive ductal carcinoma of the pancreas. Mol. Cancer 2013, 12, 168. [Google Scholar] [CrossRef] [PubMed]
- Inoue, H.; Miyazaki, Y.; Kikuchi, K.; Yoshida, N.; Ide, F.; Ohmori, Y.; Tomomura, A.; Sakashita, H.; Kusama, K. Podoplanin promotes cell migration via the EGF-Src-Cas pathway in oral squamous cell carcinoma cell lines. J. Oral Sci. 2012, 54, 241–250. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Del Rey, M.J.; Fare, R.; Izquierdo, E.; Usategui, A.; Rodriguez-Fernandez, J.L.; Suarez-Fueyo, A.; Canete, J.D.; Pablos, J.L. Clinicopathological correlations of podoplanin (gp38) expression in rheumatoid synovium and its potential contribution to fibroblast platelet crosstalk. PLoS ONE 2014, 9, e99607. [Google Scholar] [CrossRef] [PubMed]
- Talmon, G.; Wake, L.; Muirhead, D. Podoplanin and clusterin are reliable markers of nonneoplastic synovium at various sites. Int. J. Surg. Pathol. 2013, 21, 587–590. [Google Scholar] [CrossRef]
- Varki, A.; Freeze, H.H. Glycans in Acquired Human Diseases. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring: Harbor, NY, USA, 2009. [Google Scholar]
- Freeze, H.H. Genetic Disorders of Glycan Degradation. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring: Harbor, NY, USA, 2009. [Google Scholar]
- Notredame, C.; Higgins, D.G.; Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 2000, 302, 205–217. [Google Scholar] [CrossRef]
- Kaku, H.; Mori, Y.; Goldstein, I.J.; Shibuya, N. Monomeric, monovalent derivative of Maackia amurensis leukoagglutinin. Preparation and application to the study of cell surface glycoconjugates by flow cytometry. J. Biol. Chem. 1993, 268, 13237–13241. [Google Scholar]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 2015, 10, 845–858. [Google Scholar] [CrossRef]
| Treatment | Grade |
|---|---|
| Control | 0.5 |
| 1 mg MASL | 0.5 |
| LPS | 2 |
| 1 mg MASL + LPS | 0.5 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carpintero-Fernandez, P.; Varela-Eirin, M.; Lacetera, A.; Gago-Fuentes, R.; Fonseca, E.; Martin-Santamaria, S.; Mayan, M.D. New Therapeutic Strategies for Osteoarthritis by Targeting Sialic Acid Receptors. Biomolecules 2020, 10, 637. https://doi.org/10.3390/biom10040637
Carpintero-Fernandez P, Varela-Eirin M, Lacetera A, Gago-Fuentes R, Fonseca E, Martin-Santamaria S, Mayan MD. New Therapeutic Strategies for Osteoarthritis by Targeting Sialic Acid Receptors. Biomolecules. 2020; 10(4):637. https://doi.org/10.3390/biom10040637
Chicago/Turabian StyleCarpintero-Fernandez, Paula, Marta Varela-Eirin, Alessandra Lacetera, Raquel Gago-Fuentes, Eduardo Fonseca, Sonsoles Martin-Santamaria, and Maria D. Mayan. 2020. "New Therapeutic Strategies for Osteoarthritis by Targeting Sialic Acid Receptors" Biomolecules 10, no. 4: 637. https://doi.org/10.3390/biom10040637
APA StyleCarpintero-Fernandez, P., Varela-Eirin, M., Lacetera, A., Gago-Fuentes, R., Fonseca, E., Martin-Santamaria, S., & Mayan, M. D. (2020). New Therapeutic Strategies for Osteoarthritis by Targeting Sialic Acid Receptors. Biomolecules, 10(4), 637. https://doi.org/10.3390/biom10040637





