Consistency of the Tools That Predict the Impact of Single Nucleotide Variants (SNVs) on Gene Functionality: The BRCA1 Gene
Abstract
1. Introduction
2. Materials and Methods
2.1. P-Tools
2.2. Datasets
2.3. Inner Consistency Analysis
2.4. Outer Consistency Analysis
3. Results and Discussion
3.1. Inner Consistency Results
3.2. Outer Consistency
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A. P-Tools Main Features
Appendix B. Outer Consistency
| Tools | Acc. | Prec. | Recall | MCC | F1-Score |
|---|---|---|---|---|---|
| Pph2 | 0.75 | 0.38 | 0.44 | 0.25 | 0.41 |
| Prov | 0.79 | 0.75 | 0.01 | 0.06 | 0.02 |
| Gvgd | 0.36 | 0.27 | 0.96 | 0.16 | 0.42 |
| Cupd | 0.67 | 0.28 | 0.29 | 0.07 | 0.28 |
| Cupt | 0.41 | 0.25 | 0.29 | 0.08 | 0.38 |
| Pthr | 0.63 | 0.35 | 0.82 | 0.33 | 0.49 |
| Strm | 0.40 | 0.26 | 0.93 | 0.19 | 0.41 |
| Tools | Acc. | Prec. | Recall | MCC | F1-Score |
|---|---|---|---|---|---|
| Pph2 | 0.86 | 0.14 | 0.39 | 0.17 | 0.20 |
| Prov | 0.94 | 0.00 | 0.00 | 0.00 | 0.00 |
| Gvgd | 0.08 | 0.06 | 0.98 | 0.01 | 0.11 |
| Cupd | 0.72 | 0.32 | 0.37 | 0.19 | 0.37 |
| Cupt | 0.40 | 0.22 | 0.85 | 0.12 | 0.35 |
| Pthr | 0.75 | 0.17 | 0.85 | 0.31 | 0.28 |
| Strm | 0.07 | 0.07 | 0.95 | 0.11 | 0.13 |
References
- Tsui, L.; Dorfman, R. The Cystic Fibrosis Gene: A Molecular Genetic Perspective. Cold Spring Harbor Perspect. Med. 2013, 3, a009472. [Google Scholar] [CrossRef] [PubMed]
- Sharma, N.; Cutting, G. The genetics and genomics of cystic fibrosis. J. Cyst. Fibrosis 2019, 19 (Suppl. 1), S5–S9. [Google Scholar] [CrossRef] [PubMed]
- Gregersen, N. Protein misfolding disorders: Pathogenesis and intervention. J. Inherit. Metab. Dis. 2006, 29, 456–470. [Google Scholar] [CrossRef] [PubMed]
- Menzies, F.M.; Moreau, K.; Rubinsztein, D.C. Protein misfolding disorders and macroautophagy. Curr. Opin. Cell Biol. 2011, 23, 190–197. [Google Scholar] [CrossRef] [PubMed]
- Abecasis, G.R.; Auton, A.; Brooks, L.D.; DePristo, M.A.; Durbin, R.M.; Handsaker, R.E.; Kang, H.M.; Marth, G.T.; McVean, G.A. An integrated map of genetic variation from 1,092 human genomes. Nature 2012, 492, 56–65. [Google Scholar]
- Ng, P.C.; Henikoff, S. Predicting the Effects of Amino Acid Substitutions on Protein Function. Ann. Rev. Genom. Hum. Genet. 2006, 7, 61–80. [Google Scholar] [CrossRef]
- Li, M.M.; Datto, M.; Duncavage, E.J.; Kulkarni, S.; Lindeman, N.I.; Roy, S.; Tsimberidou, A.M.; Vnencak-Jones, C.L.; Wolff, D.J.; Younes, A.; et al. Standards and Guidelines for the Interpretation and Reporting of Sequence Variants in Cancer: A Joint Consensus Recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J. Mol. Diagn. 2017, 19, 4–23. [Google Scholar] [CrossRef]
- Mi, H.; Muruganujan, A.; Thomas, P.D. PANTHER in 2013: Modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 2013, 41, D377–D386. [Google Scholar] [CrossRef]
- Quan, L.; Lv, Q.; Zhang, Y. STRUM: structure-based prediction of protein stability changes upon single-point mutation. Bioinformatics 2016, 32, 2936–2946. [Google Scholar] [CrossRef]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A method and server for predicting damaging missense mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef]
- Castellana, S.; Mazza, T. Congruency in the prediction of pathogenic missense mutations: state-of-the-art web-based tools. Brief. Bioinform. 2013, 14, 448–459. [Google Scholar] [CrossRef] [PubMed]
- Thusberg, J.; Olatubosun, A.; Vihinen, M. Performance of mutation pathogenicity prediction methods on missense variants. Hum. Mutat. 2011, 32, 358–368. [Google Scholar] [CrossRef]
- Hicks, S.; Wheeler, D.A.; Plon, S.E.; Kimmel, M. Prediction of missense mutation functionality depends on both the algorithm and sequence alignment employed. Hum. Mutat. 2011, 32, 661–668. [Google Scholar] [CrossRef] [PubMed]
- Urnov, F.D. Biological techniques: Edit the genome to understand it. Nature 2014, 513, 40–41. [Google Scholar] [CrossRef] [PubMed]
- Findlay, G.M.; Daza, R.M.; Martin, B.; Zhang, M.D.; Leith, A.P.; Gasperini, M.; Janizek, J.D.; Huang, X.; Starita, L.M.; Shendure, J. Accurate classification of BRCA1 variants with saturation genome editing. Nature 2018, 562, 217–222. [Google Scholar] [CrossRef] [PubMed]
- Starita, L.M.; Islam, M.M.; Banerjee, T.; Adamovich, A.I.; Gullingsrud, J.; Fields, S.; Shendure, J.; Parvin, J.D. A Multiplex Homology-Directed DNA Repair Assay Reveals the Impact of More Than 1000 BRCA1 Missense Substitution Variants on Protein Function. Am. J. Hum. Genet. 2018, 103, 498–508. [Google Scholar] [CrossRef]
- Ransburgh, D.J.; Chiba, N.; Ishioka, C.; Toland, A.E.; Parvin, J.D. Identification of breast tumor mutations in BRCA1 that abolish its function in homologous DNA recombination. Cancer Res. 2010, 70, 988–995. [Google Scholar] [CrossRef]
- Turnbull, C.; Sud, A.; Houlston, R.S. Cancer genetics, precision prevention and a call to action. Nat. Genet. 2018, 50, 1212–1218. [Google Scholar] [CrossRef]
- Choi, Y.; Sims, G.E.; Murphy, S.; Miller, J.R.; Chan, A.P. Predicting the Functional Effect of Amino Acid Substitutions and Indels. PLoS ONE 2012, 7, e46688. [Google Scholar] [CrossRef]
- Tavtigian, S.; Deffenbaugh, A.; Yin, L.; Judkins, T.; Scholl, T.; Samollow, P.; Silva, D.; Zharkikh, A.; Thomas, A. Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J. Med. Genet. 2006, 43, 295–305. [Google Scholar] [CrossRef]
- Parthiban, V.; Gromiha, M.M.; Schomburg, D. CUPSAT: Prediction of protein stability upon point mutations. Nucleic Acids Res. 2006, 34, 239–242. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017. [Google Scholar]
- Puka, L.; Kendall’s, T. International Encyclopedia of Statistical Science; Springer: Berlin/Heidelberg, Germany, 2011; pp. 713–715. [Google Scholar]
- Dodge, Y. Spearman Rank Correlation Coefficient. In The Concise Encyclopedia of Statistics; Springer: New York, NY, USA, 2008; pp. 502–505. [Google Scholar]
- Punta, M.; Coggill, P.C.; Eberhardt, R.Y.; Mistry, J.; Tate, J.; Boursnell, C.; Pang, N.; Forslund, K.; Ceric, G.; Clements, J.; et al. The Pfam protein families database. Nucleic Acids Res. 2012, 40, 290–301. [Google Scholar] [CrossRef] [PubMed]
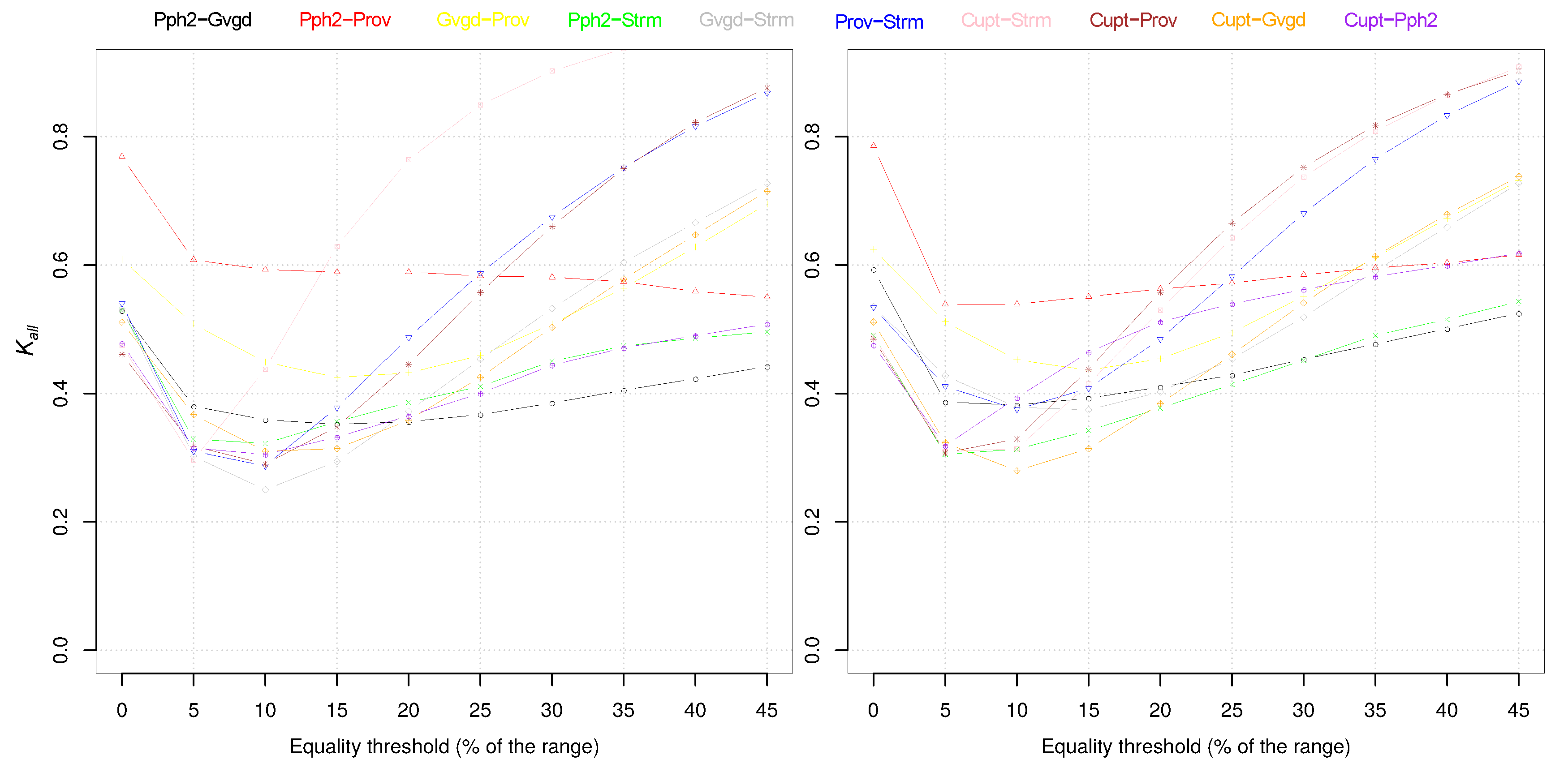
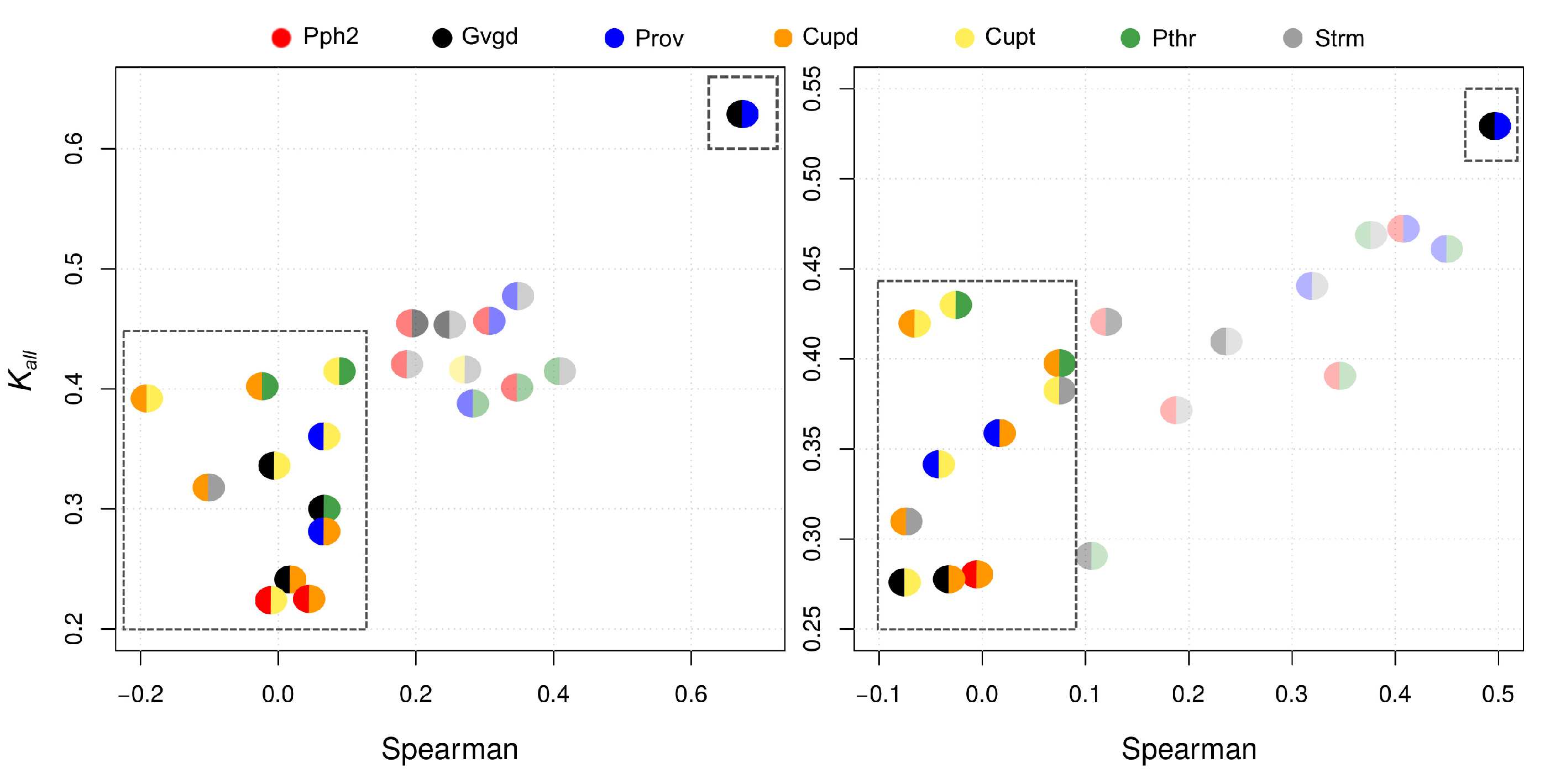
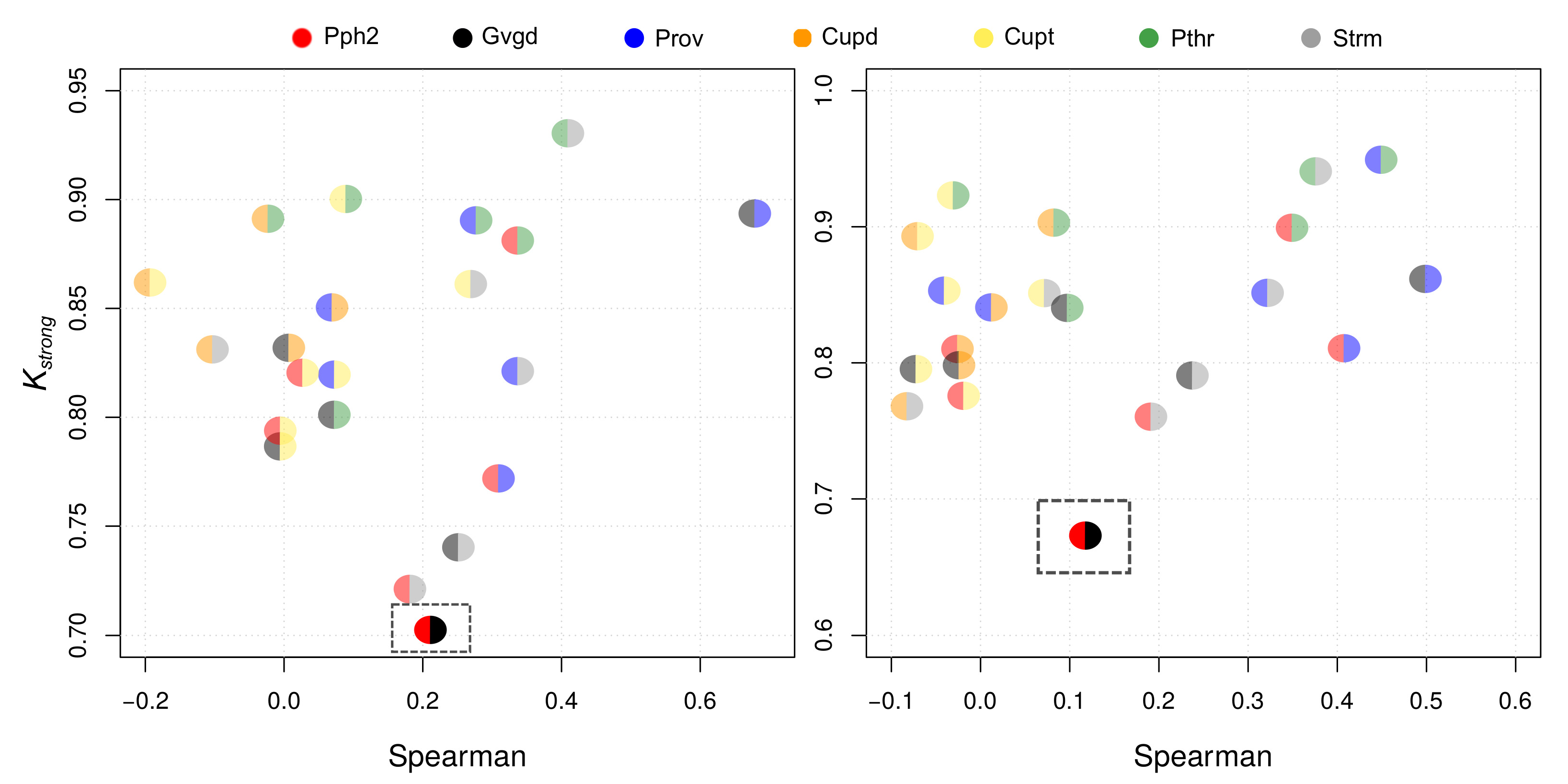
| Pph2 | Prov | Gvgd | Cupd | Cupt | Pthr | Strm | |
|---|---|---|---|---|---|---|---|
| Pph2 | 0.46 | 0.45 | 0.23 | 0.33 | 0.40 | 0.42 | |
| Prov | 0.47 | 0.62 | 0.29 | 0.36 | 0.39 | 0.47 | |
| Gvgd | 0.42 | 0.52 | 0.24 | 0.33 | 0.30 | 0.45 | |
| Cupd | 0.28 | 0.36 | 0.28 | 0.39 | 0.40 | 0.32 | |
| Cupt | 0.28 | 0.34 | 0.28 | 0.42 | 0.41 | 0.41 | |
| Pthr | 0.37 | 0.46 | 0.29 | 0.39 | 0.43 | 0.41 | |
| Strm | 0.39 | 0.44 | 0.41 | 0.31 | 0.38 | 0.47 |
| Pph2 | Prov | Gvgd | Cupd | Cupt | Pthr | Strm | |
|---|---|---|---|---|---|---|---|
| Pph2 | 0.77 | 0.70 | 0.82 | 0.79 | 0.88 | 0.72 | |
| Prov | 0.81 | 0.89 | 0.85 | 0.82 | 0.89 | 0.82 | |
| Gvgd | 0.67 | 0.85 | 0.83 | 0.79 | 0.80 | 0.74 | |
| Cupd | 0.81 | 0.84 | 0.80 | 0.86 | 0.89 | 0.83 | |
| Ccupt | 0.78 | 0.85 | 0.79 | 0.89 | 0.90 | 0.86 | |
| Pthr | 0.90 | 0.95 | 0.84 | 0.90 | 0.92 | 0.93 | |
| Strm | 0.76 | 0.85 | 0.79 | 0.77 | 0.85 | 0.94 |
| Pph2 | Prov | Gvgd | Cupd | Cupt | Pthr | Strm | |
|---|---|---|---|---|---|---|---|
| Pph2 | 0.31 | 0.20 | 0.03 | 0.00 | 0.34 | 0.18 | |
| Prov | 0.41 | 0.68 | 0.07 | 0.07 | 0.28 | 0.34 | |
| Gvgd | 0.12 | 0.50 | 0.01 | 0.00 | 0.07 | 0.25 | |
| Cupd | −0.03 | 0.01 | −0.03 | −0.19 | −0.02 | −0.10 | |
| Cupt | −0.02 | −0.04 | −0.07 | −0.07 | 0.09 | 0.27 | |
| Pthr | 0.35 | 0.45 | 0.09 | 0.08 | −0.03 | 0.41 | |
| Strm | 0.19 | 0.32 | 0.24 | −0.08 | 0.07 | 0.38 |
| BRCA1-SGE | BRCA1-HDR | |||||||
|---|---|---|---|---|---|---|---|---|
| Tools | TP (#387) | TN (#1405) | MCC | F1-Score | TP (#59) | TN (#977) | MCC | F1-Score |
| Pph2 | 115 | 871 | 0.25 | 0.41 | 12 | 555 | 0.17 | 0.20 |
| Prov | 3 | 1404 | 0.06 | 0.02 | 0 | 977 | 0.00 | 0.00 |
| Gvgd | 343 | 198 | 0.16 | 0.42 | 55 | 21 | 0.01 | 0.11 |
| Cupd | 111 | 1031 | 0.07 | 0.28 | 25 | 195 | 0.19 | 0.37 |
| Cupt | 305 | 393 | 0.08 | 0.38 | 50 | 71 | 0.12 | 0.35 |
| Pthr | 319 | 803 | 0.33 | 0.49 | 51 | 723 | 0.31 | 0.28 |
| Strm | 114 | 111 | 0.19 | 0.41 | 56 | 244 | 0.11 | 0.13 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Murillo, J.; Spetale, F.; Guillaume, S.; Bulacio, P.; Garcia Labari, I.; Cailloux, O.; Destercke, S.; Tapia, E. Consistency of the Tools That Predict the Impact of Single Nucleotide Variants (SNVs) on Gene Functionality: The BRCA1 Gene. Biomolecules 2020, 10, 475. https://doi.org/10.3390/biom10030475
Murillo J, Spetale F, Guillaume S, Bulacio P, Garcia Labari I, Cailloux O, Destercke S, Tapia E. Consistency of the Tools That Predict the Impact of Single Nucleotide Variants (SNVs) on Gene Functionality: The BRCA1 Gene. Biomolecules. 2020; 10(3):475. https://doi.org/10.3390/biom10030475
Chicago/Turabian StyleMurillo, Javier, Flavio Spetale, Serge Guillaume, Pilar Bulacio, Ignacio Garcia Labari, Olivier Cailloux, Sebastien Destercke, and Elizabeth Tapia. 2020. "Consistency of the Tools That Predict the Impact of Single Nucleotide Variants (SNVs) on Gene Functionality: The BRCA1 Gene" Biomolecules 10, no. 3: 475. https://doi.org/10.3390/biom10030475
APA StyleMurillo, J., Spetale, F., Guillaume, S., Bulacio, P., Garcia Labari, I., Cailloux, O., Destercke, S., & Tapia, E. (2020). Consistency of the Tools That Predict the Impact of Single Nucleotide Variants (SNVs) on Gene Functionality: The BRCA1 Gene. Biomolecules, 10(3), 475. https://doi.org/10.3390/biom10030475







