Identification of Polyunsaturated Fatty Acids Synthesis Pathways in the Toxic Dinophyte Alexandrium minutum Using 13C-Labelling
Abstract
1. Introduction
2. Materials and Methods
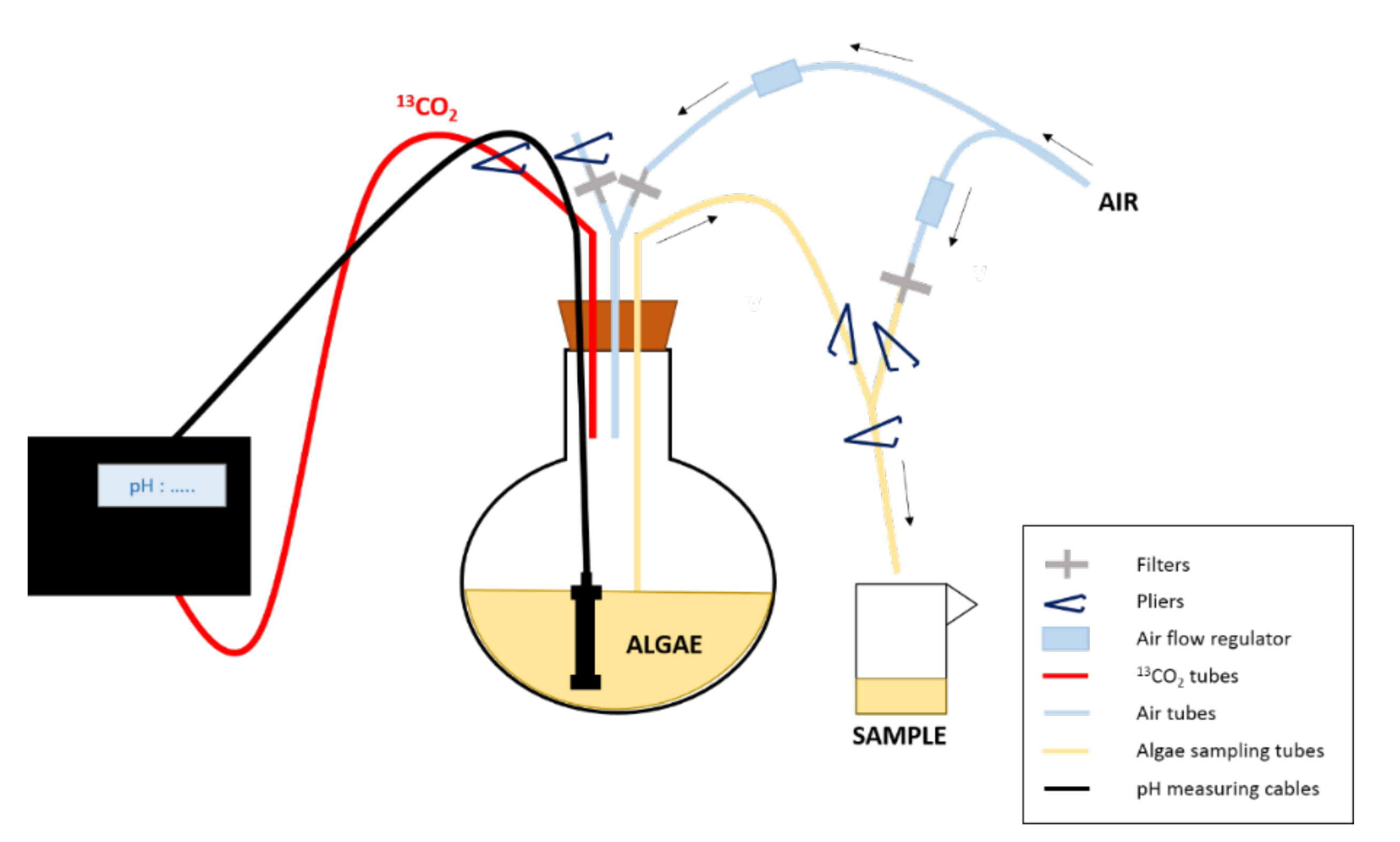
2.1. Algal Culture and Isotopic Labelling
2.2. Sample Collection
2.3. Flow Cytometry Analysis
2.4. POC Concentration and Bulk Carbon Isotopic Composition
2.5. DIC Concentration and Bulk Carbon Isotopic Composition
2.6. Isotopic Data Processing
2.7. Fatty Acid Analysis
2.7.1. Lipid Extraction
2.7.2. Separation of Neutral and Polar Lipids
2.7.3. Fatty Acid Analysis by Gas Chromatography (GC-FID and GC-c-IRMS)
2.7.4. Compound Specific Isotope Data Processing
2.8. Statistical Analysis
3. Results
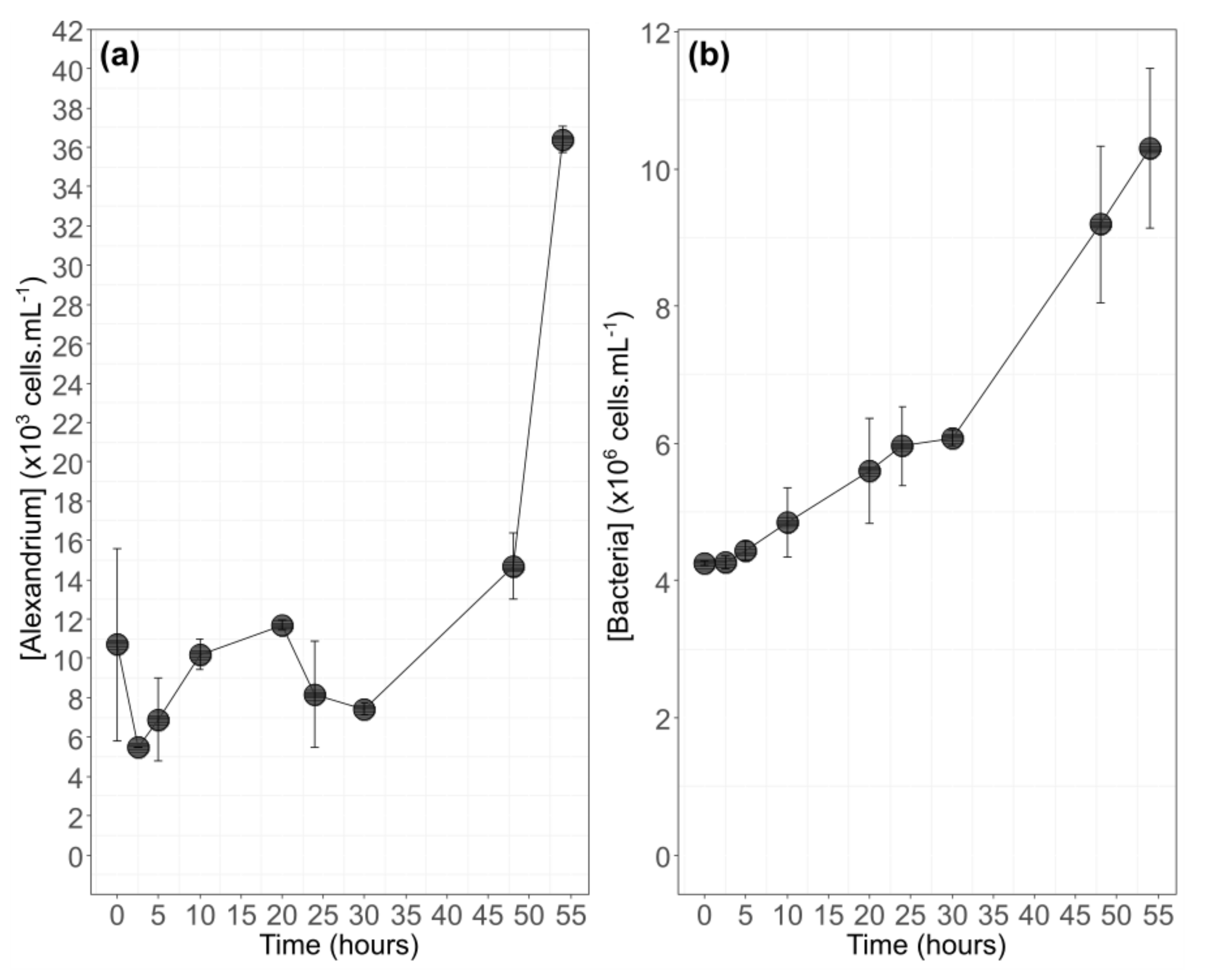
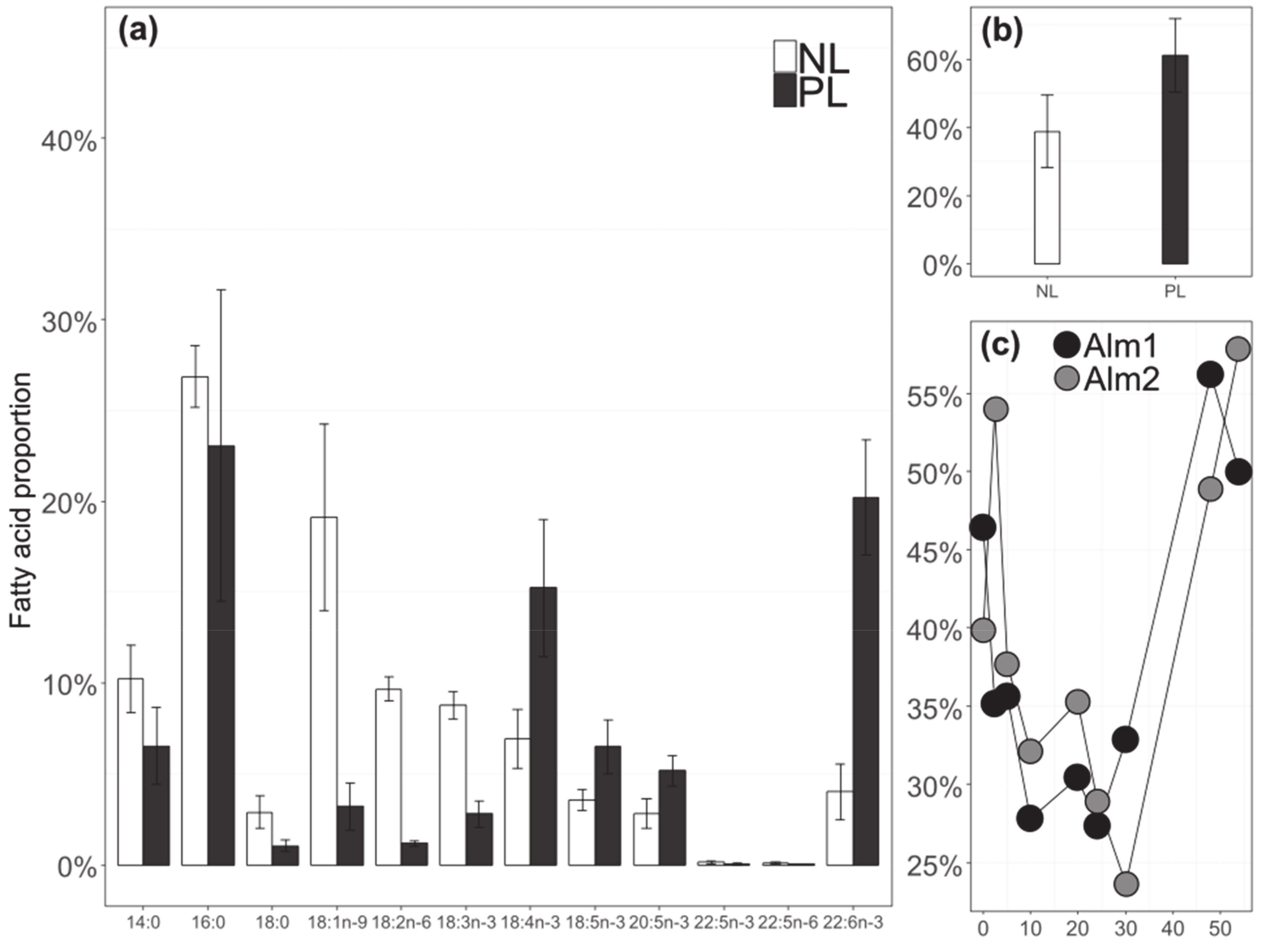
3.1. Algal Physiology during Growth
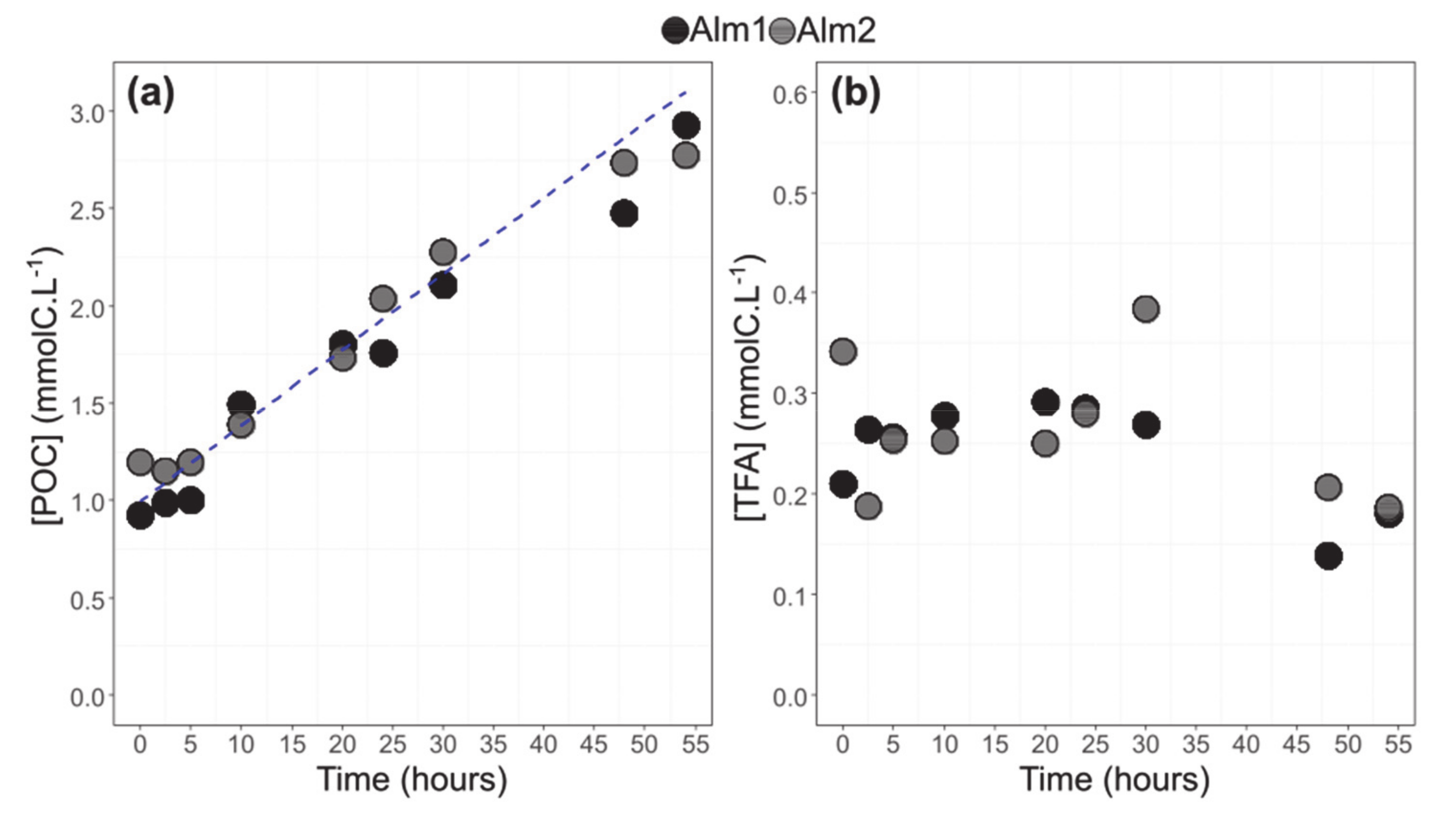
3.2. POC and TFA Concentrations
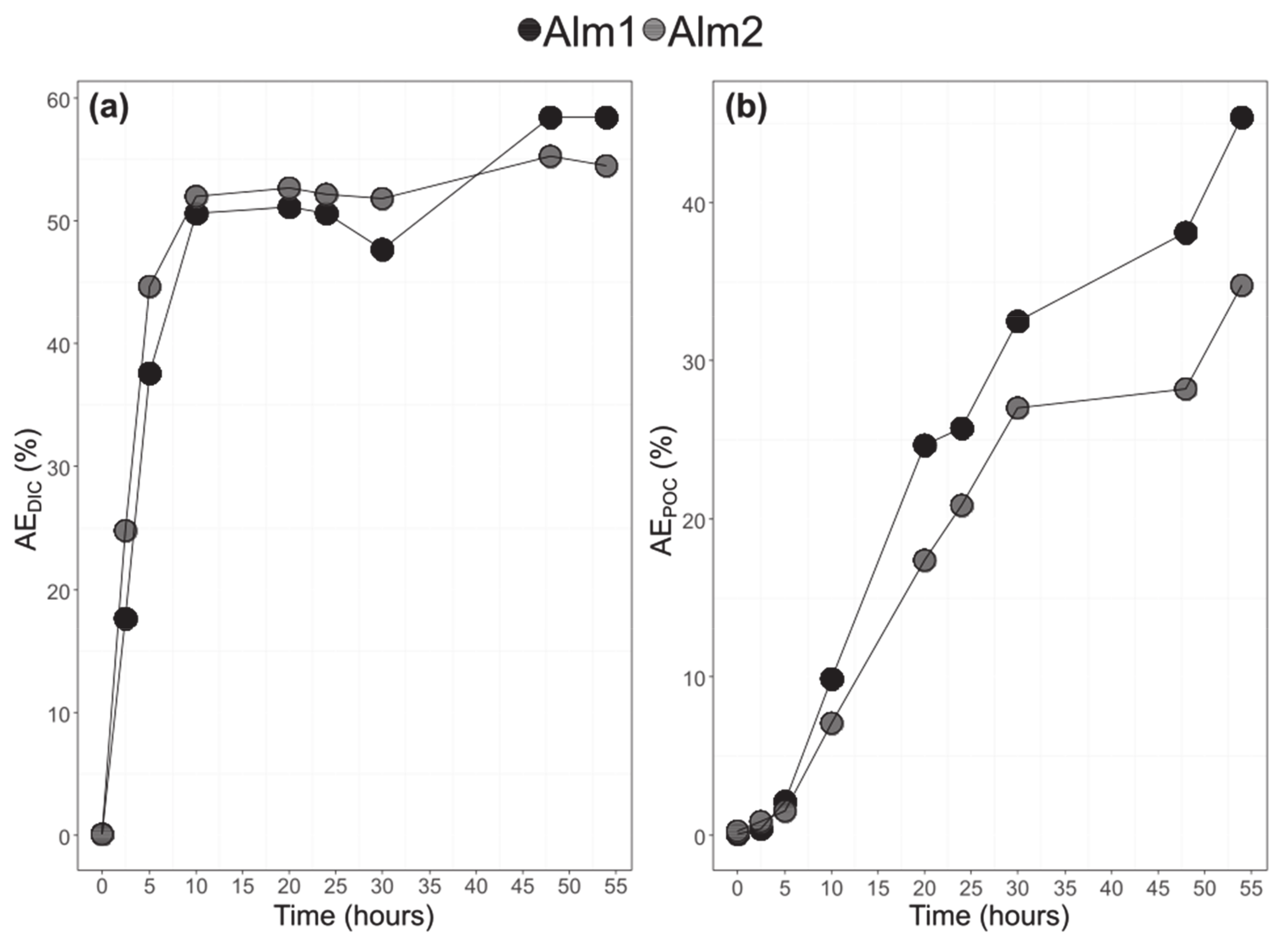
3.3. Bulk POC and DIC and Their 13C-Labelling
3.4. Bulk POC and DIC and Their 13C-Labelling
3.5. Relationship between FA
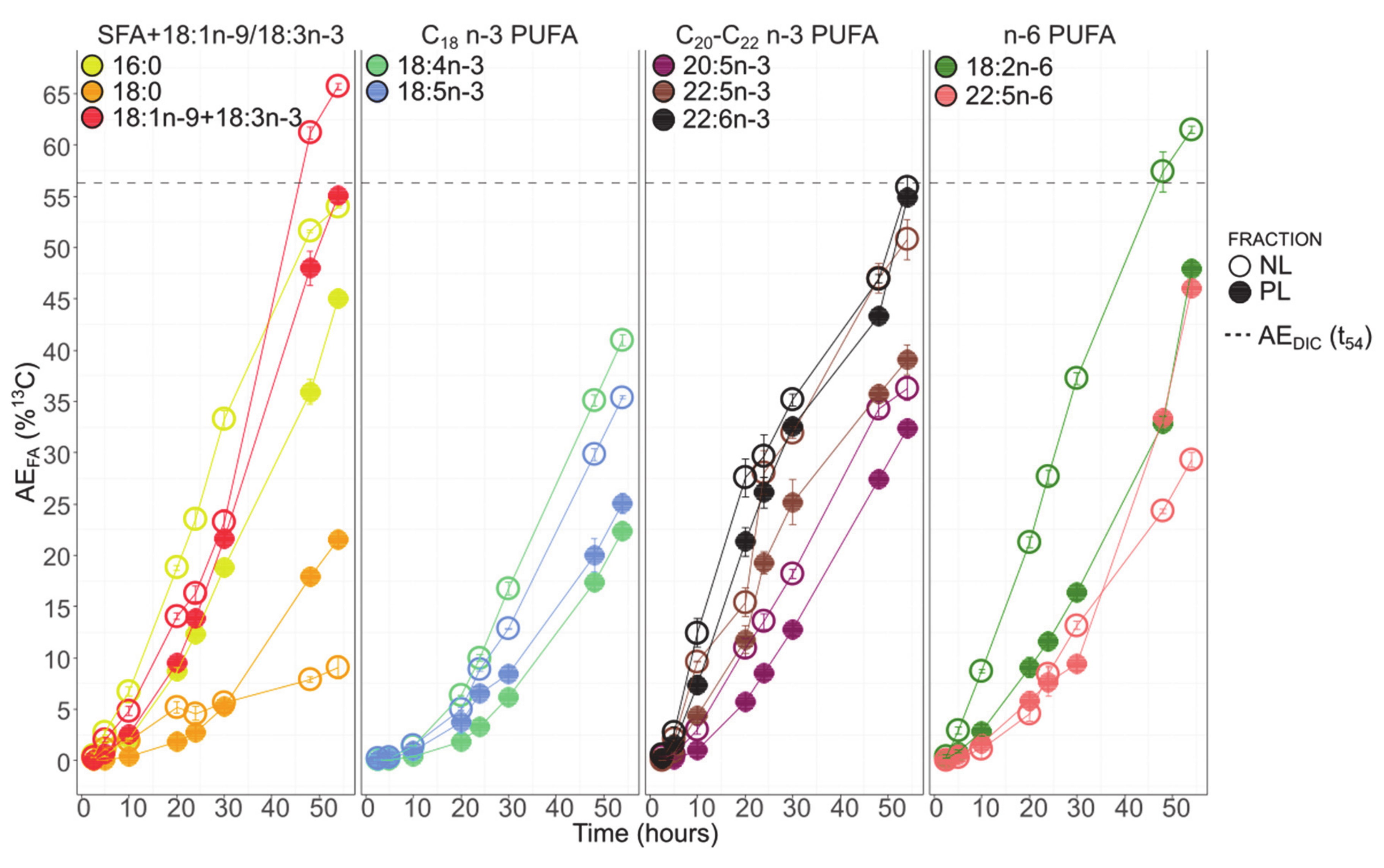
3.6. Fatty Acids 13C-Enrichment
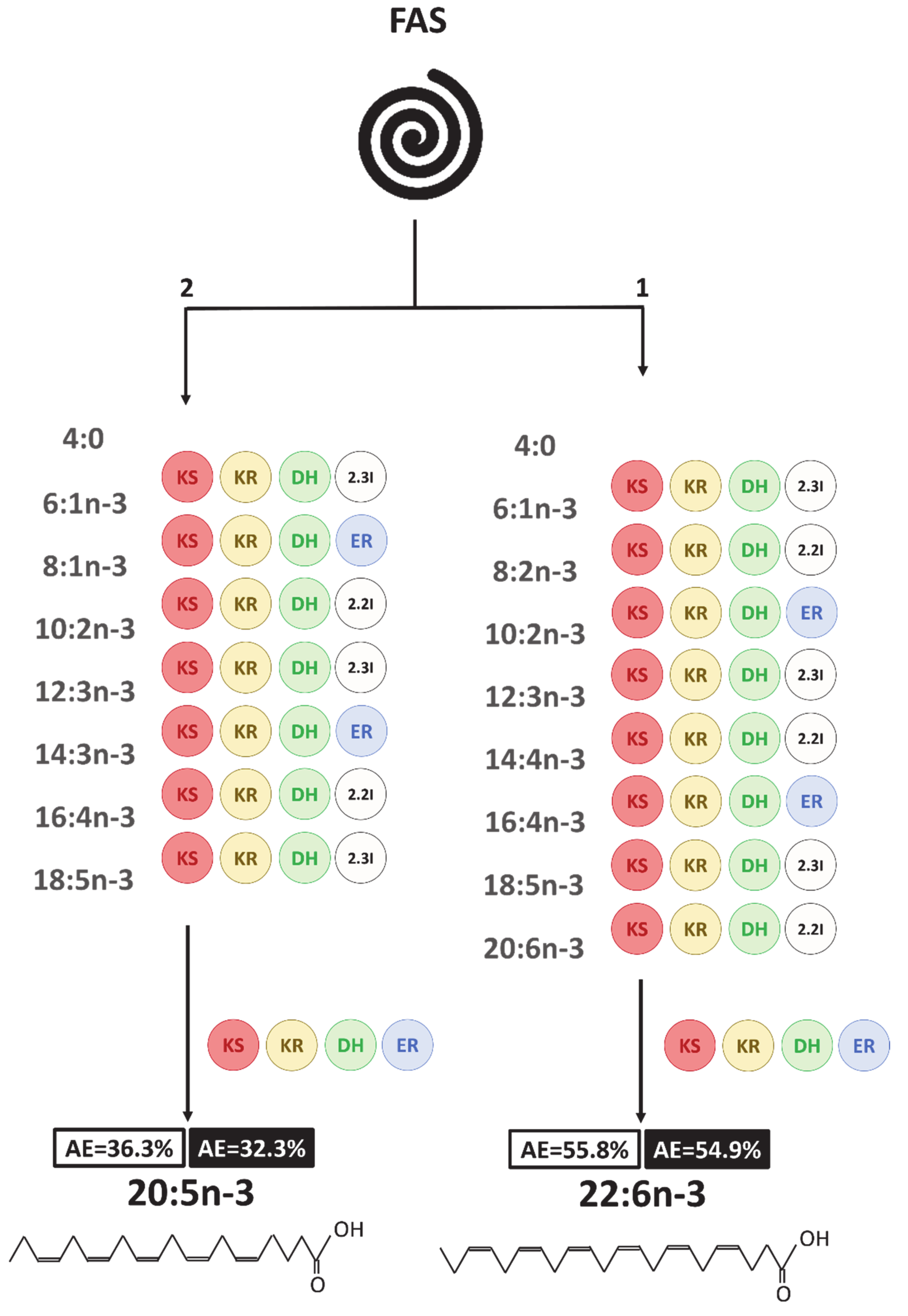
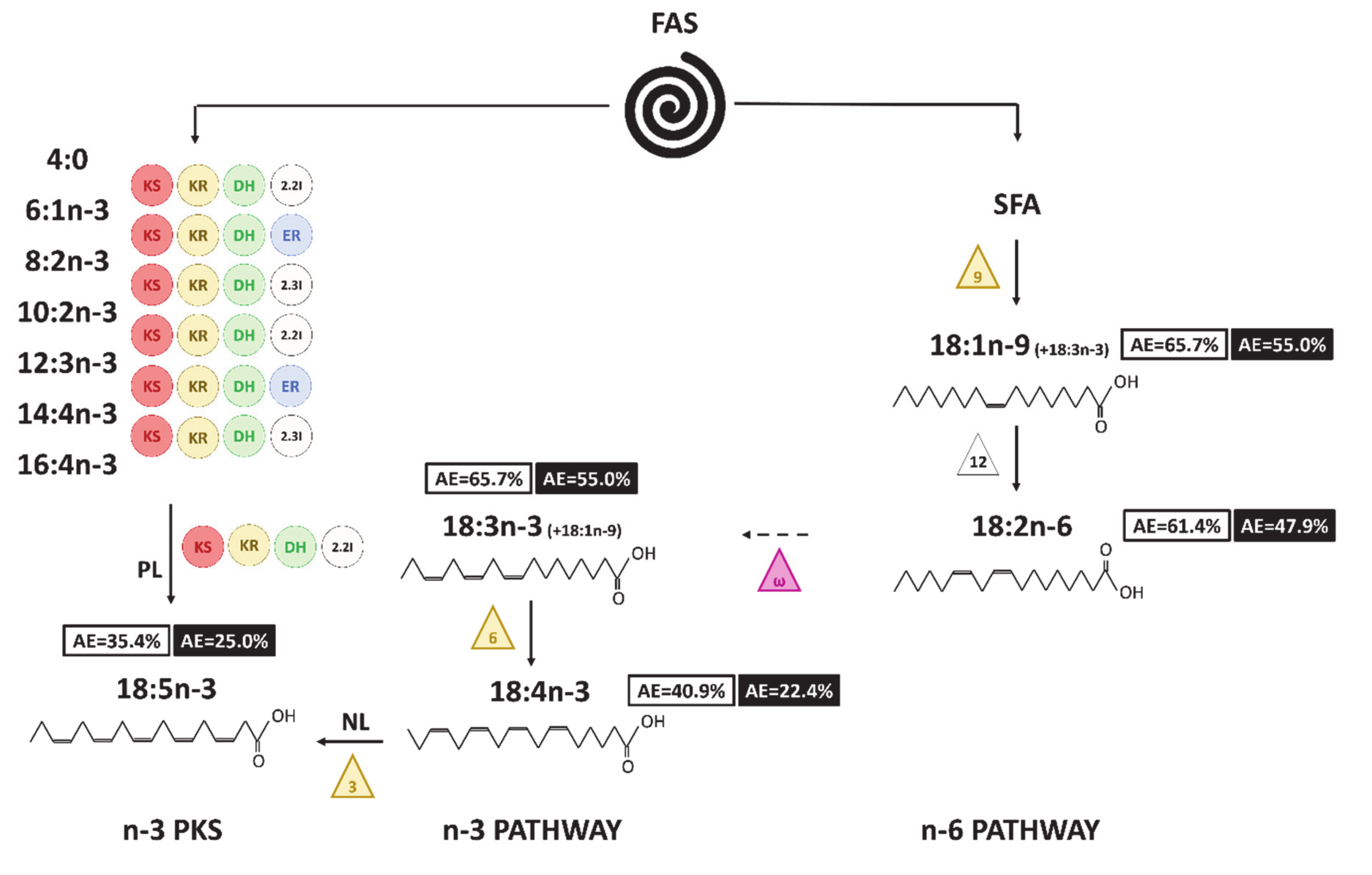
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gladyshev, M.I.; Sushchik, N.N.; Makhutova, O.N. Production of EPA and DHA in aquatic ecosystems and their transfer to the land. Prostaglandins Other Lipid Mediat. 2013, 107, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Hixson, S.M.; Arts, M.T. Climate warming is predicted to reduce omega-3, long-chain, polyunsaturated fatty acid production in phytoplankton. Glob. Chang. Biol. 2016, 22, 2744–2755. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Bunn, S.E.; Brett, M.T.; Kainz, M. Polyunsaturated fatty acids in stream food webs—high dissimilarity among producers and consumers. Freshw. Biol. 2017, 62, 1325–1334. [Google Scholar] [CrossRef]
- Cook, H.W.; McMaster, C.R. Fatty acid desaturation and chain elongation in eukaryotes. In Biochemistry of Lipids, Lipoproteins and Membranes; Vance, D.E., Vance, J.E., Eds.; Elsevier: Halifax, NS, Canada, 2002; Volume 4, p. 24. [Google Scholar]
- Tocher, D.R. Omega-3 long-chain polyunsaturated fatty acids and aquaculture in perspective. Aquaculture 2015, 449, 94–107. [Google Scholar] [CrossRef]
- Arts, M.T.; Ackman, R.G.; Holub, B.J. “Essential fatty acids” in aquatic ecosystems: A crucial link between diet and human health and evolution. Can. J. Fish. Aquat. Sci. 2001, 58, 122–137. [Google Scholar] [CrossRef]
- Mozzaffarian, D.; Wu, J.H.Y. (n-3) Fatty Acids and Cardiovascular Health: Are Effects of EPA and DHA Shared or Complementary? J. Nutr. 2012, 142, 614S–625S. [Google Scholar] [CrossRef]
- Dyall, S.C. Long-chain omega-3 fatty acids and the brain: A review of the independent and shared effects of EPA, DPA and DHA. Front. Aging Neurosci. 2015, 7, 1–16. [Google Scholar] [CrossRef]
- Hamed, I.; Özogul, F.; Özogul, Y.; Regenstein, J.M. Marine Bioactive Compounds and Their Health Benefits: A Review. Compr. Rev. Food Sci. Food Saf. 2015, 14, 446–465. [Google Scholar] [CrossRef]
- Gil, A.; Ramirez, M.; Gil, M. Role of long-chain polyunsaturated fatty acids in infant nutrition. Eur. J. Clin. Nutr. 2003, 57, S31–S34. [Google Scholar] [CrossRef]
- Li, J.J.; Huang, C.J.; Xie, D. Anti-obesity effects of conjugated linoleic acid, docosahexaenoic acid, and eicosapentaenoic acid. Mol. Nutr. Food Res. 2008, 52, 631–645. [Google Scholar] [CrossRef]
- Shidfar, F.; Keshavarz, A.; Hosseyni, S.; Ameri, A.; Yarahmadi, S. Effects of omega-3 fatty acid supplements on serum lipids, apolipoproteins and malondialdehyde in type 2 diabetes patients. East. Mediterr. Health J. 2008, 14, 9. [Google Scholar]
- Friedberg, C.E.; Janssen, M.J.F.M.; Heine, R.J.; Grobbee, D.E. Fish Oil and Glycemic Control in Diabetes: A meta-analysis. Diabetes Care 1998, 21, 494–500. [Google Scholar] [CrossRef] [PubMed]
- Tocher, D.; Betancor, M.B.; Sprague, M.; Olsen, R.E.; Napier, J.A. Omega-3 Long-Chain Polyunsaturated Fatty Acids, EPA and DHA: Bridging the Gap between Supply and Demand. Nutrients 2019, 11, 89. [Google Scholar] [CrossRef] [PubMed]
- Tocher, D. Issues surrounding fish as a source of omega-3 long-chain polyunsaturated fatty acids. Lipid Technol. 2009, 21, 13–16. [Google Scholar] [CrossRef]
- Kang, J.X. Omega-3: A link between global climate change and human health. Biotechnol. Adv. 2011, 29, 388–390. [Google Scholar] [CrossRef]
- Arao, T.; Yamada, M. Biosynthesis of polyunsaturated fatty acids in the marine diatom, Phaeodactylum tricornutum. Phytochemistry 1994, 35, 1177–1181. [Google Scholar] [CrossRef]
- Metz, J.G.; Roessler, P.; Facciotti, D.; Levering, C.; Dittrich, F.; Lassner, M.; Valentine, R.; Lardizabal, K.; Domergue, F.; Yamada, A.; et al. Production of polyunsaturated fatty acids by polyketide synthases in both prokaryotes and eukaryotes. Science 2001, 293, 290–293. [Google Scholar] [CrossRef]
- Bell, M.V.; Tocher, D. Biosynthesis of polyunsaturated fatty acids in aquatic ecosystems: General pathways and new directions. In Lipids in Aquatic Ecosystems; Arts, M.T., Brett, M.T., Kainz, M., Eds.; Springer: New York, NY, USA, 2009; pp. 211–239. [Google Scholar]
- Domergue, F.; Lerchl, J.; Zähringer, U.; Heinz, E. Cloning and functional characterization of Phaeodactylum tricornutum front-end desaturases involved in eicosapentaenoic acid biosynthesis. Eur. J. Biochem. 2002, 269, 4105–4113. [Google Scholar] [CrossRef]
- Ratledge, C. Fatty acid biosynthesis in microorganisms being used for Single Cell Oil production. Biochimie 2004, 86, 807–815. [Google Scholar] [CrossRef]
- Armenta, R.E.; Valentine, M.C. Single-Cell Oils as a Source of Omega-3 Fatty Acids: An Overview of Recent Advances. J. Am. Oil Chem. Soc. 2013, 90, 167–182. [Google Scholar] [CrossRef]
- Eichholz, K.; Beszteri, B.; John, U. Putative monofunctional type I polyketide synthase units: A dinoflagellate-specific feature? PLoS ONE 2012, 7, e48624. [Google Scholar] [CrossRef] [PubMed]
- Kimura, K.; Okuda, S.; Nakayama, K.; Shikata, T.; Takahashi, F.; Yamaguchi, H.; Skamoto, S.; Yamaguchi, M.; Tomaru, Y. RNA sequencing revealed numerous polyketide synthase genes in the harmful dinoflagellate Karenia mikimotoi. PLoS ONE 2015, 10, 16. [Google Scholar] [CrossRef] [PubMed]
- Monroe, E.A.; van Dolah, F.M. The Toxic Dinoflagellate Karenia brevis Encodes Novel Type I-like Polyketide Synthases Containing Discrete Catalytic Domains. Protist 2008, 159, 471–482. [Google Scholar] [CrossRef]
- Rein, K.S.; Gibbs, P.D.L.; Palacios, A.; Abiy, L.; Dickey, R.; Snyder, R.V.; Lopez, J.V. Polyketide Synthase Genes from Marine Dinoflagellates. Mar. Biotechnol. 2003, 5, 1–12. [Google Scholar] [CrossRef]
- Freitag, M.; Beszteri, S.; Vogel, H.; John, U. Effects of physiological shock treatments on toxicity and polyketide synthase gene expression in Prymnesium parvum (Prymnesiophyceae). Eur. J. Phycol. 2011, 46, 193–201. [Google Scholar] [CrossRef]
- John, U.; Beszteri, S.; Glöckner, G.; Singh, R.; Medlin, L.; Cembella, A.D. Genomic characterisation of the ichthyotoxic prymnesiophyte Chrysochromulina polylepis, and the expression of polyketide synthase genes in synchronized cultures. Eur. J. Phycol. 2010, 45, 215–229. [Google Scholar] [CrossRef]
- Murray, S.A.; Garby, T.; Hoppenrath, M.; Neilan, B.A.; Salamin, N. Genetic Diversity, Morphological Uniformity and Polyketide Production in Dinoflagellates (Amphidinium, Dinoflagellata). PLoS ONE 2012, 7, e38253. [Google Scholar] [CrossRef]
- Kohli, G.S.; John, U.; van Dolah, F.M.; Murray, S.A. Evolutionary distinctiveness of fatty acid and polyketide synthesis in eukaryotes. ISME J. 2016, 10, 1877–1890. [Google Scholar] [CrossRef]
- Cock, J.M.; Sterck, L.; Rouzé, P.; Scornet, D.; Allen, A.E.; Amoutziaz, G.; Anthouard, V.; Artiguenave, F.; Aury, J.M.; Badger, J.H.; et al. The Ectocarpus genome and the independent evolution of multicellularity in brown algae. Nature 2010, 465, 617–621. [Google Scholar] [CrossRef]
- Jenke-Kodama, H.; Sandmann, A. Evolutionary Implications of Bacterial Polyketide Synthases. Mol. Biol. Evol. 2005, 22, 2027–2039. [Google Scholar] [CrossRef]
- Remize, M.; Planchon, F.; Loh, A.N.; Le Grand, F.; Bideau, A.; Le Goic, N.; Fleury, E.; Miner, P.; Corvaisier, R.; Volety, A.; et al. Study of Synthesis Pathways of the Essential Polyunsaturated Fatty Acid 20:5n-3 in the Diatom Chaetoceros muelleri Using 13C-Isotope Labeling. Biomolecules 2020, 10, 797. [Google Scholar] [CrossRef] [PubMed]
- Galloway, A.W.E.; Winder, M. Partitioning the Relative Importance Phylogeny and Environmental Conditions on Phytoplankton Fatty Acids. PLoS ONE 2015, 10, e0130053. [Google Scholar] [CrossRef]
- Taylor, F.J.R.; Hoppenrath, M.; Saldarriaga, J.F. Dinoflagellate diversity and distribution. Biodivers. Conserv. 2008, 17, 407–418. [Google Scholar] [CrossRef]
- Landsberg, J.H. The Effects of Harmful Algal Blooms on Aquatic Organisms. Rev. Fish. Sci. 2002, 10, 113–390. [Google Scholar] [CrossRef]
- Jang, S.H.; Jeong, H.J.; Chon, J.K. De novo transcriptome of the newly described phototrophic dinoflagellate Yihiella yeosuensis: Comparison between vegetative cells and cysts. Mar. Biol. 2019, 166. [Google Scholar] [CrossRef]
- Kim, Y.O.; Park, M.H.; Han, M.S. Role of cyst germination in the bloom initiation of Alexandrium tamarense (Dinophyceae) in Masan Bay, Korea. Aquat. Microb. Ecol. 2002, 29, 279–286. [Google Scholar] [CrossRef]
- Saito, K.; Drgon, T.; Krupatkina, D.N.; Drgonova, J.; Terlizzi, D.E.; Mercer, N.; Vasta, G.R. Effect of Biotic and Abiotic Factors on In Vitro Proliferation, Encystment, and Excystment of Pfiesteria piscicida. Appl. Environ. Microbiol. 2007, 73, 6410–6420. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Deng, Y.; Hu, Z.; Shang, L.; Peng, Q.; Tang, Y.Z. Transcriptomic Analyses of Scrippsiella trochoidea Reveals Processes Regulating Encystment and Dormancy in the Life Cycle of a Dinoflagellate, with a Particular Attention to the Role of Abscisic Acid. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef]
- Garcés, E.; Bravo, I.; Vila, M.; Figueroa, R.I.; Maso, M.; Sampedro, N. Relationship between vegetative cells and cyst production during Alexandrium minutum bloom in Arenys de Mar harbour (NW Mediterranean). J. Plankton Res. 2004, 26, 637–645. [Google Scholar] [CrossRef]
- Matantseva, O. Cellular mechanisms of dinoflagellate cyst development and ecdysis—many questions to answer. Protistology 2019, 13. [Google Scholar] [CrossRef]
- Olli, K. Temporary cyst formation of Heterocapsa triquetra (Dinophyceae) in natural populations. Mar. Biol. 2004, 145. [Google Scholar] [CrossRef]
- Binder, B.J.; Anderson, D.M. Biochemical composition and metabolic activity of Scrippsiella trochoidea (Dinophycea) resting cysts. J. Phycol. 1990, 26, 289–298. [Google Scholar] [CrossRef]
- Kellmann, R.; Stüken, A.; Orr, R.J.S.; Svendsen, H.M.; Jakobsen, K.S. Biosynthesis and Molecular Genetics of Polyketides in Marine Dinoflagellates. Mar. Drugs 2010, 8, 1011–1048. [Google Scholar] [CrossRef] [PubMed]
- John, U.; Beszteri, B.; Derelle, E.; van de Peer, Y.; Read, B.; Moreau, H.; Cembella, A. Novel insights into evolution of protistan polyketide synthases through phylogenomic analysis. Protist 2008, 159, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Sellem, F.; Pesando, D.; Bodennec, G.; El Abed, A.; Girard, J.P. Toxic effects of Gymnodinium cf. mikimotoi unsaturated fatty acids to gametes and embryos of the sea urchin Paracentrotus lividus. Water Res. 2000, 34, 550–556. [Google Scholar] [CrossRef]
- Zhao, Y.; Tang, X.; Zhao, X.; Wang, Y.; Lin, S. Effect of various nitrogen conditions on population growth, temporary cysts and cellular biochemical compositions of Karenia mikimotoi. PLoS ONE 2017, 12, e0171996. [Google Scholar] [CrossRef] [PubMed]
- De Swaaf, M.E.; de Rijk, T.C.; van der Meer, P.; Eggink, G.; Sijtsma, L. Analysis of docosahexaenoic acid biosynthesis in Crypthecodinium cohnii by 13C labelling and desaturase inhibitor experiments. J. Biotechnol. 2003, 103, 21–29. [Google Scholar] [CrossRef]
- Ecker, J.; Scherer, M.; Schimtz, G.; Liebisch, G. A rapid GC–MS method for quantification of positional and geometric isomers of fatty acid methyl esters. J. Chromatogr. B 2012, 897, 98–104. [Google Scholar] [CrossRef]
- Van den Meersche, K.; Middelburg, J.J.; Soetaert, K.; van Rijswijk, P.; Boschker, H.T.S.; Heip, C.H.R. Carbon-nitrogen coupling and algal-bacterial interaction during an experimental bloom: Modeling a 13C tracer experiment. Limnol. Oceanogr. 2004, 49, 862–878. [Google Scholar] [CrossRef]
- Zhang, H.; Wu, C.; Wu, Q.; Dai, J.; Song, Y.; Chen, W.N. Metabolic Flux Analysis of Lipid Biosynthesis in the Yeast Yarrowia lipolytica Using 13C-Labled Glucose and Gas Chromatography-Mass Spectrometry. PLoS ONE 2016, 11, 14. [Google Scholar] [CrossRef]
- Zhao, J.; Shimizu, K. Metabolic flux analysis of Escherichia coli K12 grown on 13C-labeled acetate and glucose using GC-MS and powerful flux calculation method. J. Biotechnol. 2003, 101, 101–117. [Google Scholar] [CrossRef]
- Xiong, W.; Liu, L.; Wu, C.; Yang, C.; Wu, Q. 13C-Tracer and Gas Chromatography-Mass Spectrometry Analyses Reveal Metabolic Flux Distribution in the Oleaginous Microalga Chlorella protothecoides. Plant Physiol. 2010, 154, 1001–1011. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Diao, J.; Sun, T.; Shi, M.; Liu, L.; Wang, F.; Chen, L.; Zhang, W. 13C Metabolic Flux Analysis of Enhanced Lipid Accumulation Modulated by Ethanolamine in Crypthecodinium cohnii. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Martzolff, A.; Cahoreau, E.; Cogne, G.; Peyriga, L.; Portais, J.C.; Dechandol, E.; Le Grand, F.; Massou, S.; Gonçalves, O.; Pruvost, J.; et al. Photobioreactor design for isotopic non-stationnary 13C-metabolic flux analysis (INST 13C-MFA) under photoautotrophic conditions. Biotechnol. Bioeng. 2012, 109, 3030–3040. [Google Scholar] [CrossRef] [PubMed]
- Grosse, J.; van Breugel, P.; Boschker, H.T.S. Tracing carbon fixation in phytoplankton-compound specific and total 13C incorporation rates: 13C uptake into macromolecules. Limnol. Oceanogr. Methods 2015, 13, 288–302. [Google Scholar] [CrossRef]
- Menzel, R.; Ngosong, C.; Ruess, L. Isotopologue profiling enables insights into dietary routing and metabolism of trophic biomarker fatty acids. Chemoecology 2017, 27, 101–114. [Google Scholar] [CrossRef]
- Wei, X.; Shi, B.; Koo, I.; Yin, X.; Lorkiewicz, P.; Suhail, H.; Rattan, R.; Giri, S.; McClain, C.J.; Zhang, X. Analysis of stable isotope assisted metabolomics data acquired by GC-MS. Anal. Chim. Acta 2017, 980, 25–32. [Google Scholar] [CrossRef]
- Lima, V.F.; Perez de Souza, L.; Williams, T.C.R.; Fernie, A.R.; Daloso, D.M. Gas chromatography-mass spectrometry based 13C labeling studies in plant metabolomics. In Plant Metabolomics; Antonio, C., Ed.; Springer: New York, NY, USA, 2018; Volume 1778, pp. 47–58. [Google Scholar]
- Andersen, R.A. Algal Culturing Techniques; Elsevier: Amsterdam, The Netherlands, 2005; p. 578. [Google Scholar]
- González-Fernández, C.; Toullec, J.; Lambert, C.; Le Goïc, N.; Seoane, M.; Moriceau, B.; Huvet, A.; Berchel, M.; Vincent, D.; Courcot, L.; et al. Do transparent exopolymeric particles (TEP) affect the toxicity of nanoplastics on Chaetoceros neogracile? Environ. Pollut. 2019, 250, 873–882. [Google Scholar] [CrossRef]
- Tracy, B.P.; Gaida, S.M.; Papoutsakis, E.T. Flow cytometry for bacteria: Enabling metabolic engineering, synthetic biology and the elucidation of complex phenotypes. Curr. Opin. Biotechnol. 2010, 21, 85–99. [Google Scholar] [CrossRef]
- Le Goïc, N.; Hegaret, H.; Boulais, M.; Béguel, J.P.; Lambert, C.; Fabioux, C.; Soudant, P. Flow cytometric assessment of morphology, viability, and production of Reactive Oxygen Species of Crassostrea gigas oocytes. Application to toxic Dinoflagellate (Alexandrium minutum) exposure. Cytom. Part A 2014, 85A, 1049–1056. [Google Scholar] [CrossRef]
- Seoane, M.; González-Fernández, C.; Soudant, P.; Huvet, A.; Esperanza, M.; Cid, A.; Paul-Pont, I. Polystyrene microbeads modulate the energy metabolism of the marine diatom Chaetoceros neogracile. Environ. Pollut. 2019, 251, 362–371. [Google Scholar] [CrossRef]
- Lelong, A.; Haberkorn, H.; Le Goïc, N.; Hégaret, H.; Soudant, P. A New Insight into Allelopathic Effects of Alexandrium minutum on Photosynthesis and Respiration of the Diatom Chaetoceros neogracile Revealed by Photosynthetic-performance Analysis and Flow Cytometry. Microb. Ecol. 2011, 62, 919–930. [Google Scholar] [CrossRef] [PubMed]
- United States Environmental Protection Agency; U.S.E. Methods for the Determination of Chemical Substances in Marine and Estuarine Environmental Matrices; U.S Environmental Protection Agency: Washington, DC, USA, 1997; p. 199. [Google Scholar]
- Assayag, N.; Rivé, K.; Ader, M.; Jézéquel, D.; Agrinier, P. Improved method for isotopic and quantitative analysis of dissolved inorganic carbon in natural water samples. Rapid Commun. Mass Spectrom. 2006, 20, 2243–2251. [Google Scholar] [CrossRef]
- Larsson, C.; Ahlstrøm, Ø.; Junghans, P.; Jensen, R.B.; Blache, D.; Tauson, A.-H. The oral [13C]bicarbonate technique for measurement of short-term energy expenditure of sled dogs and their physiological response to diets with different fat:carbohydrate ratios. J. Nutr. Sci. 2015, 4, 10. [Google Scholar] [CrossRef] [PubMed]
- Le Grand, F.; Soudant, P.; Siah, A.; Tremblay, P.; Marty, Y.; Kraffe, E. Disseminated Neoplasia in the Soft-Shell Clam Mya arenaria: Membrane Lipid Composition and Functional Parameters of Circulating Cells. Lipids 2014, 49, 807–818. [Google Scholar] [CrossRef] [PubMed]
- Mathieu-Resuge, M.; Schaal, G.; Kraffe, E.; Corvaisier, R.; Lebeau, O.; Lluch-Costa, S.E.; Salgado García, R.L.; Kainz, M.J.; Le Grand, F. Different particle sources in a bivalve species of a coastal lagoon: Evidence from stable isotopes, fatty acids, and compound-specific stable isotopes. Mar. Biol. 2019, 166, 89–101. [Google Scholar] [CrossRef]
- Mansour, M.P.; Volkman, J.K.; Blackburn, S.I. The effect of growth phase on the lipid class, fatty acid and sterol composition in the marine dinoflagellate, Gymnodinium sp. in batch culture. Phytochemistry 2003, 63, 145–153. [Google Scholar] [CrossRef]
- Mooney, B.D.; Nichols, P.D.; de Salas, M.F.; Hallegraeff, G.M. Lipid, fatty acid, and sterol composition of eight species of Kareniaceae (Dinophyta): Chemotaxonomy and putative lipid phycotoxins. J. Phycol. 2007, 43, 101–111. [Google Scholar] [CrossRef]
- Leblond, J.D.; Chapman, P.J. Lipid class distribution of highly unsaturated long chain fatty acids in marine dinoflagellates. J. Phycol. 2000, 36, 1103–1108. [Google Scholar] [CrossRef]
- Khozin-Goldberg, I. Lipid metabolism in microalgae. In The Physiology of Microalgae; Borowitska, M.A., Beardall, J., Raven, J.A., Eds.; Springer: Cham, Switzerland, 2016; pp. 413–484. [Google Scholar]
- Li, Y.; Horsman, M.; Wang, B.; Wu, N.; Lan, C.Q. Effects of nitrogen sources on cell growth and lipid accumulation of green alga Neochloris oleoabundans. Appl. Microbiol. Biotechnol. 2008, 81, 629–636. [Google Scholar] [CrossRef]
- Liang, Y.; Beardall, J.; Heraud, P. Effects of nitrogen source and UV radiation on the growth, chlorophyll fluorescence and fatty acid composition of Phaeodactylum tricornutum and Chaetoceros muelleri (Bacillariophyceae). J. Photochem. Photobiol. B 2006, 82, 161–172. [Google Scholar] [CrossRef]
- Leblond, J.D.; Lasiter, A.D. Mono- and digalactosyldiacylglycerol composition of dinoflagellates. II. Lepidodinium chlorophorum, Karenia brevis, and Kryptoperidinium foliaceum, three dinoflagellates with aberrant plastids. Eur. J. Phycol. 2009, 44, 199–205. [Google Scholar] [CrossRef][Green Version]
- Henderson, J.; Leftley, J.W.; Sargent, J.R. Lipid composition and biosynthesis in the marine dinoflagellate Crypthecodinium cohnii. Phytochemistry 1988, 27, 1679–1683. [Google Scholar] [CrossRef]
- Pei, G.; Li, X.; Liu, J.; Wang, F.; Chen, L.; Zhang, W. De novo transcriptomic and metabolomic analysis of docosahexaenoic acid (DHA)-producing Crypthecodinium cohnii during fed-batch fermentation. Algal Res. 2017, 26, 380–391. [Google Scholar] [CrossRef]
- Kong, F.; Romero, I.T.; Warakanont, J.; Li-Beisson, Y. Lipid catabolism in microalgae. New Phytol. 2018, 218, 1340–1348. [Google Scholar] [CrossRef] [PubMed]
- Řezanka, T.; Lukavský, J.; Nedbalová, L.; Sigler, K. Lipidomic profile in three species of dinoflagellates (Amphidinium carterae, Cystodinium sp. and Peridinium aciculiferum) containing very long chain polyunsaturated fatty acids. Phytochemistry 2017, 139, 88–97. [Google Scholar] [CrossRef]
- Lichtlé, C.; Dubacq, J.P. Lipid modification related to encystment and excystment of Cryptomonas rufescens skuja (Cryptophyceae). J. Phycol. 1984, 20, 8–12. [Google Scholar] [CrossRef]
- Xu, J.; Kazachkov, M.; Jia, Y.; Zheng, Z.; Zou, J. Expression of a type 2 diacylglycerol acyltransferase from Thalassiosira pseudonana in yeast leads to incorporation of docosahexaenoic acid β-oxidation intermediates into triacylglycerol. FEBS J. 2013, 280, 6162–6172. [Google Scholar] [CrossRef]
- Becker, K.W.; Collins, J.R.; Durham, B.P.; Groussman, R.D.; White, A.E.; Fredricks, H.F.; Ossolinski, J.E.; Repeta, D.J.; Carini, P.; Armbrust, E.V.; et al. Daily changes in phytoplankton lipidomes reveal mechanisms of energy storage in the open ocean. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef]
- Dahmen, J.L.; Khadka, M.; Dodson, V.J.; Leblond, J.D. Mono- and digalactosyldiacylglycerol composition of dinoflagellates. VI. Biochemical and genomic comparison of galactolipid biosynthesis between Chromera velia (Chromerida), a photosynthetic alveolate with red algal plastid ancestry, and the dinoflagellate, Lingulodinium polyedrum. Eur. J. Biochem. 2013, 48, 268–277. [Google Scholar] [CrossRef]
- Joseph, J.D. Identification of 3, 6, 9, 12, 15-octadecapentaenoic acid in laboratory-cultured photosynthetic dinoflagellates. Lipids 1975, 10, 395–403. [Google Scholar] [CrossRef]
- Adolf, J.E.; Place, A.R.; Stoecker, D.K.; Harding, L.W. Modulation of polyunsaturated fatty acids in mixotrophic Karlodinium veneficum (Dinophyceae) and its prey, Storeatula major (Cryptophyceae). J. Phycol. 2007, 43, 1259–1270. [Google Scholar] [CrossRef]
- Tsai, C.H.; Warakanont, J.; Takeuchi, T.; Sears, B.B.; Moellering, E.R.; Benning, C. The protein Compromised Hydrolysis of Triacylglycerols 7 (CHT7) acts as a repressor of cellular quiescence in Chlamydomonas. Proc. Natl. Acad. Sci. USA 2014, 111, 15833–15838. [Google Scholar] [CrossRef] [PubMed]
- Da Costa, E.; Silva, J.; Mendonça, S.; Abreu, M.; Domingues, M. Lipidomic Approaches towards Deciphering Glycolipids from Microalgae as a Reservoir of Bioactive Lipids. Mar. Drugs 2016, 14, 101–128. [Google Scholar] [CrossRef] [PubMed]
- Boudière, L.; Michaud, M.; Petroutsos, D.; Rébeillé, F.; Falconet, D.; Bastien, O.; Roy, S.; Finazzi, G.; Rolland, N.; Jouhet, J.; et al. Glycerolipids in photosynthesis: Composition, synthesis and trafficking. Biochim. Biophys. Acta Bioenerg. 2014, 1837, 470–480. [Google Scholar] [CrossRef] [PubMed]
| Description | Nature | Analysis | δ13C (‰) | SD |
|---|---|---|---|---|
| IAEA-CH6 | Sucrose (C12H22O11) | 13C-POC | −10.45 | 0.03 |
| IAEA-600 | Caffeine (C8H10N4O2) | 13C-POC | −27.77 | 0.04 |
| Acetanilide | Acetanilide (C8H9NO) | 13C-POC | +29.53 | 0.01 |
| CA21 (in-house std) | Calcium carbonate (CaCO3) | 13C-DIC | +1476 | |
| Na2CO3 (in-house std) | Sodium carbonate | 13C-DIC | −6.8805 | |
| NaHCO3 (in-house std) | Sodium bicarbonate | 13C-DIC | −5.9325 |
| (h) | FL1-SYTOX ALIVE | FL1-BODIPY | FL3 | SSC | FSC | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 100 | ± | 0 | 107 | ± | 18 | 547 | ± | 5 | 627 | ± | 13 | 164 | ± | 5 |
| 2.5 | 100 | ± | 0 | 115 | ± | 11 | 549 | ± | 17 | 639 | ± | 8 | 161 | ± | 5 |
| 5 | 99 | ± | 1 | 118 | ± | 3 | 574 | ± | 25 | 639 | ± | 7 | 162 | ± | 0 |
| 10 | 100 | ± | 0 | 158 | ± | 25 | 599 | ± | 9 | 676 | ± | 0 | 164 | ± | 1 |
| 20 | 100 | ± | 0 | 130 | ± | 22 | 601 | ± | 15 | 653 | ± | 15 | 141 | ± | 11 |
| 24 | 100 | ± | 0 | 90 | ± | 7 | 606 | ± | 12 | 730 | ± | 22 | 159 | ± | 7 |
| 30 | 100 | ± | 1 | 96 | ± | 10 | 534 | ± | 3 | 707 | ± | 26 | 167 | ± | 5 |
| 48 | 100 | ± | 0 | 58 | ± | 9 | 554 | ± | 14 | 697 | ± | 15 | 162 | ± | 1 |
| 54 | 100 | ± | 0 | 81 | ± | 3 | 522 | ± | 2 | 602 | ± | 6 | 181 | ± | 5 |
| Fatty acid B/Fatty acid A | Neutral Lipids | Polar Lipids | ||||||
|---|---|---|---|---|---|---|---|---|
| Alm1 | Alm2 | Alm1 | Alm2 | |||||
| Mean | SD | Mean | SD | Mean | SD | Mean | SD | |
| 18:5n-3/18:4n-3 | 0.56 | 0.16 | 0.53 | 0.15 | 1.67 | 0.64 | 1.70 | 0.55 |
| 20:5n-3/18:5n-3 | 1.28 | 0.38 | 1.48 | 0.61 | 2.04 | 0.56 | 2.20 | 0.59 |
| 22:5n-3/20:5n-3 | 1.94 | 0.85 | 1.81 | 0.62 | 2.24 | 1.32 | 2.15 | 1.00 |
| 22:6n-3/20:5n-3 | 2.32 | 1.04 | 2.25 | 0.99 | 3.33 | 2.15 | 3.32 | 2.21 |
| 22:6n-3/22:5n-3 | 1.21 | 0.30 | 1.23 | 0.29 | 1.45 | 0.23 | 1.47 | 0.29 |
| 22:6n-3/22:5n-6 * | 3.22 | 1.87 | 3.89 | 1.61 | 2.74 | 1.27 | 3.15 | 1.51 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Remize, M.; Planchon, F.; Loh, A.N.; Le Grand, F.; Lambert, C.; Bideau, A.; Bidault, A.; Corvaisier, R.; Volety, A.; Soudant, P. Identification of Polyunsaturated Fatty Acids Synthesis Pathways in the Toxic Dinophyte Alexandrium minutum Using 13C-Labelling. Biomolecules 2020, 10, 1428. https://doi.org/10.3390/biom10101428
Remize M, Planchon F, Loh AN, Le Grand F, Lambert C, Bideau A, Bidault A, Corvaisier R, Volety A, Soudant P. Identification of Polyunsaturated Fatty Acids Synthesis Pathways in the Toxic Dinophyte Alexandrium minutum Using 13C-Labelling. Biomolecules. 2020; 10(10):1428. https://doi.org/10.3390/biom10101428
Chicago/Turabian StyleRemize, Marine, Frédéric Planchon, Ai Ning Loh, Fabienne Le Grand, Christophe Lambert, Antoine Bideau, Adeline Bidault, Rudolph Corvaisier, Aswani Volety, and Philippe Soudant. 2020. "Identification of Polyunsaturated Fatty Acids Synthesis Pathways in the Toxic Dinophyte Alexandrium minutum Using 13C-Labelling" Biomolecules 10, no. 10: 1428. https://doi.org/10.3390/biom10101428
APA StyleRemize, M., Planchon, F., Loh, A. N., Le Grand, F., Lambert, C., Bideau, A., Bidault, A., Corvaisier, R., Volety, A., & Soudant, P. (2020). Identification of Polyunsaturated Fatty Acids Synthesis Pathways in the Toxic Dinophyte Alexandrium minutum Using 13C-Labelling. Biomolecules, 10(10), 1428. https://doi.org/10.3390/biom10101428





