Circulating microRNA as a Biomarker for Coronary Artery Disease
Abstract
:1. Introduction
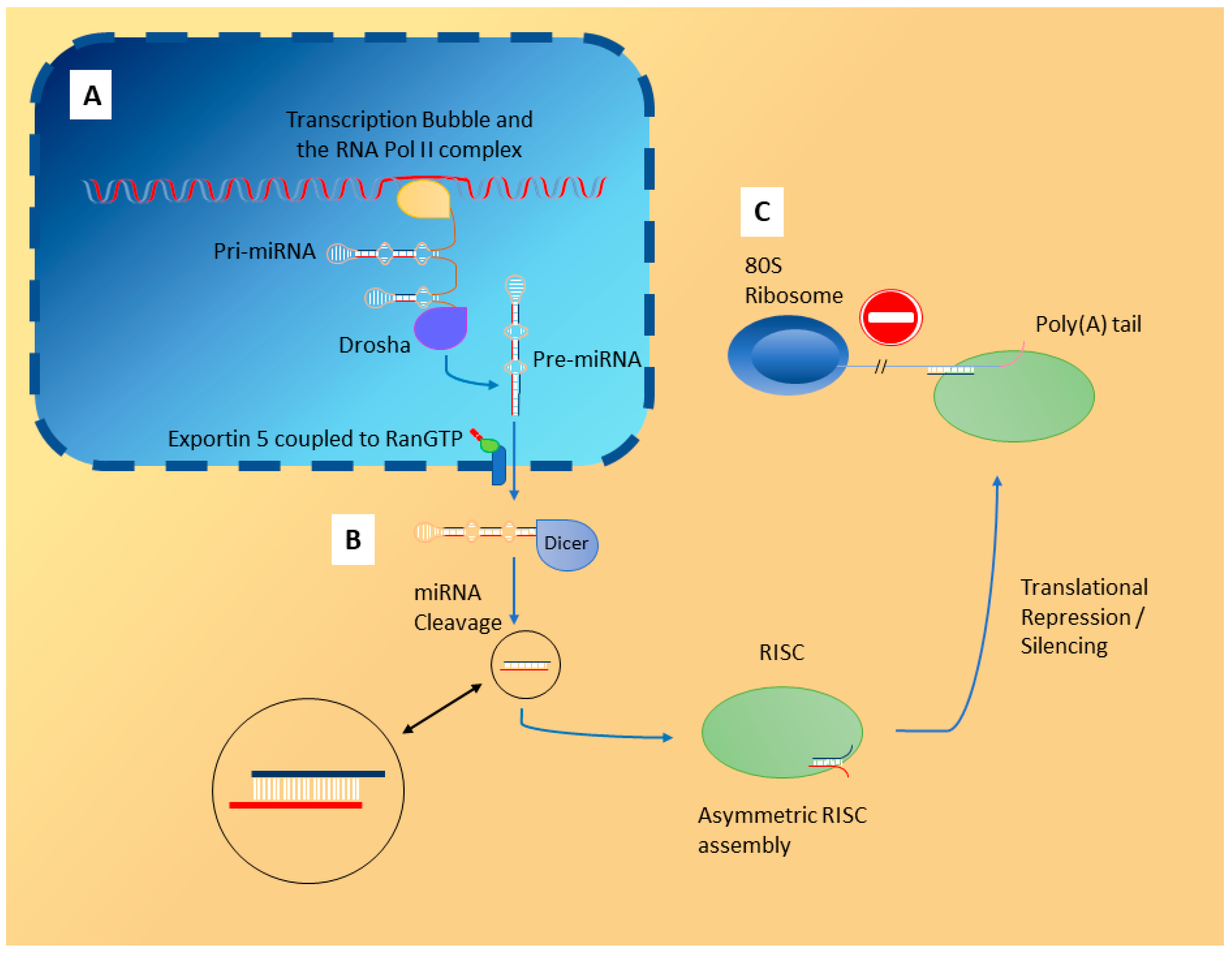
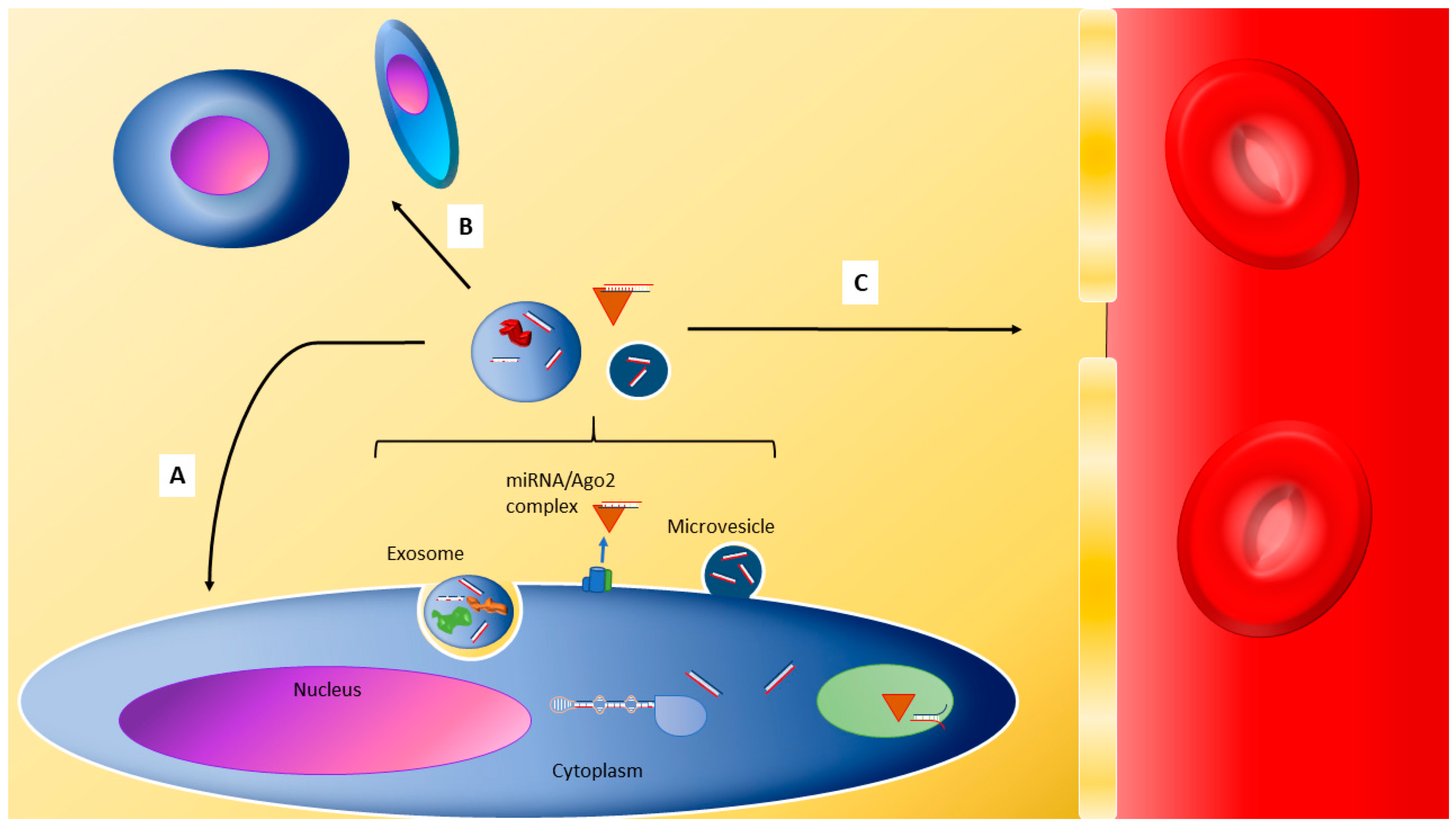
2. Localisation of miRNA
3. Physiological Roles of miRNA and Their Clinical Relevance
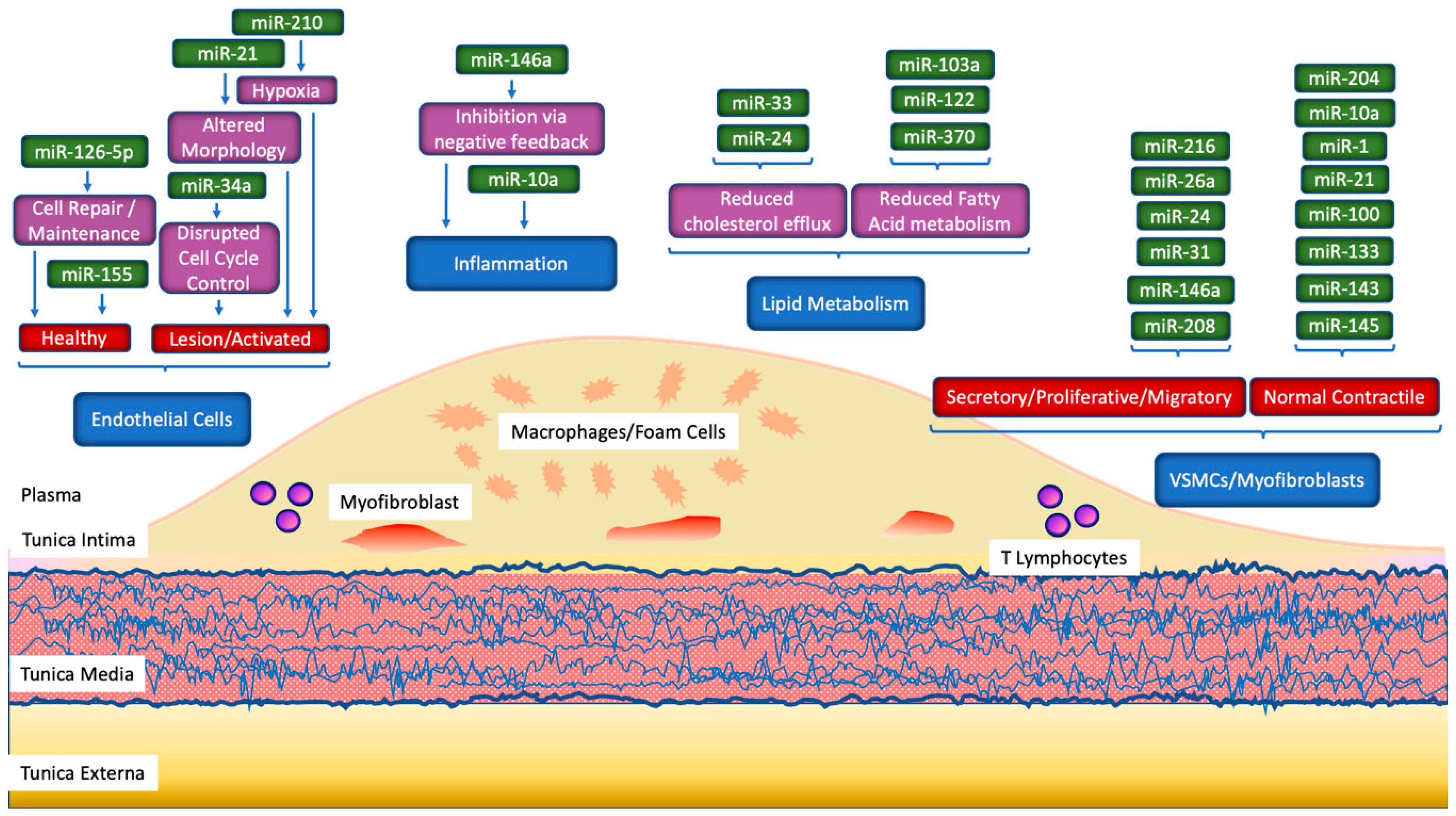
4. Coronary Artery Disease (CAD) Pathophysiology
5. CAD Biomarkers and miRNA
5.1. Localised Changes in miRNA Profiles
5.2. Changes in Circulating miRNA
6. miRNA in CAD Pathophysiology
6.1. Lipid Metabolism
6.2. Inflammation
7. Pitfalls in Assessing miRNA as Biomarker Targets
7.1. Confounding Factors
7.2. Measuring Serum and Plasma miRNA
8. Validity of miRNA as Biomarkers
9. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- 2016 Diagnostic Biomarkers. In BEST (Biomarkers, EndpointS, and other Tools) Resource; Food and Drug Administration (US): Silver Spring, MD, USA; National Institutes of Health (US): Bethesda, MD, USA, 2018; pp. 3–5.
- Salzano, A.; Marra, A.M.; D’Assante, R.; Arcopinto, M.; Bossone, E.; Suzuki, T.; Cittadini, A. Biomarkers and Imaging: Complementary or Subtractive? Heart Fail. Clin. 2019, 15, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Mordi, I.R.; Badar, A.A.; John Irving, R.; Weir-McCall, J.R.; Houston, J.G.; Lang, C.C. Efficacy of noninvasive cardiac imaging tests in diagnosis and management of stable coronary artery disease. Vasc. Health Risk Manag. 2017, 13, 427–437. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vogel, R.A. Biomarkers of High-Grade Coronary Stenosis: Searching for Seventies. J. Am. Coll. Cardiol. 2017, 69, 1157–1159. [Google Scholar] [CrossRef] [PubMed]
- Carthew, R.W.; Sontheimer, E.J. Origins and Mechanisms of miRNAs and siRNAs. Cell 2009, 136, 642–655. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodriguez, A.; Griffiths-Jones, S.; Ashurst, J.L.; Bradley, A. Identification of mammalian microRNA host genes and transcription units. Genome Res. 2004, 14, 1902–1910. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartel, D.P. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef] [Green Version]
- Cortez, M.A.; Bueso-Ramos, C.; Ferdin, J.; Lopez-Berestein, G.; Sood, A.K.; Calin, G.A. MicroRNAs in body fluids--the mix of hormones and biomarkers. Nat. Rev. Clin. Oncol. 2011, 8, 467–477. [Google Scholar] [CrossRef] [Green Version]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.H.; Lee, M.J.; Galas, D.J.; Wang, K. The microRNA spectrum in 12 body fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 2, 1000. [Google Scholar] [CrossRef] [Green Version]
- Sohel, M.H. Extracellular/Circulating MicroRNAs: Release Mechanisms, Functions and Challenges. ALS 2016, 10, 175–186. [Google Scholar] [CrossRef] [Green Version]
- Yuan, A.; Farber, E.L.; Rapoport, A.L.; Tejada, D.; Deniskin, R.; Akhmedov, N.B.; Farber, D.B. Transfer of microRNAs by embryonic stem cell microvesicles. PLoS ONE 2009, 4, e4722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zernecke, A.; Bidzhekov, K.; Noels, H.; Shagdarsuren, E.; Gan, L.; Denecke, B.; Hristov, M.; Köppel, T.; Jahantigh, M.N.; Lutgens, E.; et al. Delivery of MicroRNA-126 by Apoptotic Bodies Induces CXCL12-Dependent Vascular Protection. Sci. Signal 2009, 2, 81. [Google Scholar] [CrossRef] [PubMed]
- Turchinovich, A.; Weiz, L.; Langheinz, A.; Burwinkel, B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 2011, 39, 7223–7233. [Google Scholar] [CrossRef] [PubMed]
- Hessvik, N.P.; Llorente, A. Current knowledge on exosome biogenesis and release. Cell. Mol. Life Sci. 2018, 75, 193–208. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ståhl, A.L.; Johansson, K.; Mossberg, M.; Kahn, R.; Karpman, D. Exosomes and microvesicles in normal physiology, pathophysiology, and renal diseases. Pediatr. Nephrol. 2019, 34, 11–30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tricarico, C.; Clancy, J.; D’Souza-Schorey, C. Biology and biogenesis of shed microvesicles. Small GTPases 2017, 8, 220–232. [Google Scholar] [CrossRef] [Green Version]
- Pegtel, D.M.; Cosmopoulos, K.; Thorley-Lawson, D.A.; Van Eijndhoven, M.A.J.; Hopmans, E.S.; Lindenberg, J.L.; De Gruijl, T.D.; Würdinger, T.; Middeldorp, J.M. Functional delivery of viral miRNAs via exosomes. Proc. Natl. Acad. Sci. USA 2010, 107, 6328–6333. [Google Scholar] [CrossRef] [Green Version]
- Stoorvogel, W. Functional transfer of microRNA by exosomes. Blood 2012, 119, 646–648. [Google Scholar] [CrossRef]
- Pfeffer, S.; Zavolan, M.; Grässer, F.A.; Chien, H.; Russo, J.J.; Ju, J.; John, B.; Enright, A.J.; Marks, D.; Sander, C.; et al. Identification of Virus-Encoded MicroRNAs. Science (80-) 2004, 304, 734–736. [Google Scholar] [CrossRef]
- Xia, T.; O’Hara, A.; Araujo, I.; Barreto, J.; Carvalho, E.; Sapucaia, J.B.; Ramos, J.C.; Luz, E.; Pedroso, C.; Manrique, M.; et al. EBV microRNAs in primary lymphomas and targeting of CXCL-11 by ebv-mir-BHRF1-3. Cancer Res. 2008, 68, 1436–1442. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buck, A.H.; Coakley, G.; Simbari, F.; Mcsorley, H.J.; Quintana, J.F.; Le, T.; Kumar, S.; Abreu-goodger, C.; Lear, M.; Harcus, Y.; et al. Exosomes secreted by nematode parasites transfer small RNAs to mammalian cells and modulate innate immunity. Nat. Commun. 2014, 5, 5488. [Google Scholar] [CrossRef] [PubMed]
- Mause, S.F.; Weber, C. Microparticles: Protagonists of a novel communication network for intercellular information exchange. Circ. Res. 2010, 107, 1047–1057. [Google Scholar] [CrossRef]
- Vickers, K.C.; Palmisano, B.T.; Shoucri, B.M.; Shamburek, R.D.; Remaley, A.T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 2011, 13, 423–435. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Niculescu, L.S.; Simionescu, N.; Sanda, G.M.; Carnuta, M.G.; Stancu, C.S.; Popescu, A.C.; Popescu, M.R.; Vlad, A.; Dimulescu, D.R.; Simionescu, M.; et al. MiR-486 and miR-92a Identified in Circulating HDL Discriminate between Stable and Vulnerable Coronary Artery Disease Patients. PLoS ONE 2015, 10, e0140958. [Google Scholar] [CrossRef] [PubMed]
- Auber, M.; Fröhlich, D.; Drechsel, O.; Karaulanov, E.; Krämer-Albers, E.M. Serum-free media supplements carry miRNAs that co-purify with extracellular vesicles. J. Extracell. Vesicles 2019, 8, 1656042. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jeppesen, D.K.; Fenix, A.M.; Franklin, J.L.; Higginbotham, J.N.; Zhang, Q.; Zimmerman, L.J.; Liebler, D.C.; Ping, J.; Liu, Q.; Evans, R.; et al. Reassessment of Exosome Composition. Cell 2019, 177, 428–445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Albanese, M.; Chen, Y.-F.A.; Hüls, C.; Gärtner, K.; Tagawa, T.; Keppler, O.T.; Göbel, C.; Zeidler, R.; Hammerschmidt, W. Micro RNAs Are Minor Constituents of Extracellular Vesicles and Are Hardly Delivered to Target Cells. Available online: https://www.biorxiv.org/content/10.1101/2020.05.20.106393v1.abstract (accessed on 26 July 2020).
- O’Brien, K.; Breyne, K.; Ughetto, S.; Laurent, L.C.; Breakefield, X.O. RNA delivery by extracellular vesicles in mammalian cells and its applications. Nat. Rev. Mol. Cell Biol. 2020, 21, 585–606. [Google Scholar] [CrossRef]
- Park, C.Y.; Choi, Y.S.; McManus, M.T. Analysis of microRNA knockouts in mice. Hum. Mol. Genet. 2010, 19, 169–175. [Google Scholar] [CrossRef] [Green Version]
- Bernstein, E.; Kim, S.Y.; Carmell, M.A.; Murchison, E.P.; Alcorn, H.; Li, M.Z.; Mills, A.A.; Elledge, S.J.; Anderson, K.V.; Hannon, G.J. Dicer is essential for mouse development. Nat. Genet. 2003, 35, 215–217. [Google Scholar] [CrossRef]
- London, L. Motivator and Barriers to Latina’s Participation in Clinical Trials. Contemp. C 2015, 40, 3–14. [Google Scholar] [CrossRef] [Green Version]
- Van Rooij, E.; Quiat, D.; Johnson, B.A.; Sutherland, L.B.; Qi, X.; Richardson, J.A.; Kelm, R.J.; Olson, E.N. Expression and Muscle Performance. Dev. Cell 2009, 17, 662–673. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Rooij, E.; Sutherland, L.B.; Qi, X.; Richardson, J.A.; Hill, J.; Olson, E.N. Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 2007, 316, 575–579. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xin, M.; Small, E.M.; Sutherland, L.B.; Qi, X.; McAnally, J.; Plato, C.F.; Richardson, J.A.; Bassel-Duby, R.; Olson, E.N. MicroRNAs miR-143 and miR-145 modulate cytoskeletal dynamics and responsiveness of smooth muscle cells to injury. Genes Dev. 2009, 23, 2166–2178. [Google Scholar] [CrossRef] [Green Version]
- Zhao, W.; Zhao, S.-P.; Zhao, Y.-H. MicroRNA-143/-145 in Cardiovascular Diseases. Biomed. Res. Int. 2015, 2015, 531740. [Google Scholar] [CrossRef]
- Bang, C.; Batkai, S.; Dangwal, S.; Gupta, S.K.; Foinquinos, A.; Holzmann, A.; Just, A.; Remke, J.; Zimmer, K.; Zeug, A.; et al. Cardiac fibroblast-derived microRNA passenger strand-enriched exosomes mediate cardiomyocyte hypertrophy. J. Clin. Invest. 2014, 124, 2136–2146. [Google Scholar] [CrossRef]
- Fu, F.; Jiang, W.; Zhou, L. Circulating Exosomal miR-17-5p and miR-92a-3p Predict Pathologic Stage and Grade of Colorectal Cancer. Transl. Oncol. 2018, 11, 221–232. [Google Scholar] [CrossRef]
- Liu, T.; Zhang, Q.; Zhang, J.; Li, C.; Miao, Y.R.; Lei, Q.; Li, Q.; Guo, A.Y. EVmiRNA: A database of miRNA profiling in extracellular vesicles. Nucleic Acids Res. 2019, 47, 89–93. [Google Scholar] [CrossRef] [Green Version]
- Yang, S.Y.; Wang, Y.Q.; Gao, H.M.; Wang, B.; He, Q. The clinical value of circulating MIR-99a in plasma of patients with acute myocardial infarction. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 5193–5197. [Google Scholar]
- Schulte, C.; Barwari, T.; Joshi, A.; Theofilatos, K.; Zampetaki, A.; Barallobre-Barreiro, J.; Singh, B.; Sörensen, N.A.; Neumann, J.T.; Zeller, T.; et al. Comparative analysis of circulating noncoding rnas versus protein biomarkers in the detection of myocardial injury. Circ. Res. 2019, 125, 328–340. [Google Scholar] [CrossRef]
- Mendell, J.T.; Olson, E.N. MicroRNAs in stress signaling and human disease. Cell 2012, 148, 1172–1187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bonneau, E.; Neveu, B.; Kostantin, E.; Tsongalis, G.J.; De Guire, V. How close are miRNAs from clinical practice? A perspective on the diagnostic and therapeutic market. Electron. J. Int. Fed. Clin. Chem. Lab. Med. 2019, 30, 114–127. [Google Scholar]
- Bajan, S.; Hutvagner, G. RNA-Based Therapeutics: From Antisense Oligonucleotides to miRNAs. Cells 2020, 9, 137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mitra, S.; Deshmukh, A.; Sachdeva, R.; Lu, J.; Mehta, J.L. Oxidized Low-Density Lipoprotein and Atherosclerosis Implications in Antioxidant Therapy. Am. J. Med. Sci. 2011, 342, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Linton, M.F.; Yancey, P.G.; Davies, S.S.; Jerome, W.G.; Linton, E.F.; Vickers, K.C. The Role of Lipids and Lipoproteins in Atherosclerosis; MDText: South Dartmouth, MA, USA, 2000. [Google Scholar]
- Bairey Merz, C.N.; Pepine, C.J.; Walsh, M.N.; Fleg, J.L.; Camici, P.G.; Chilian, W.M.; Clayton, J.A.; Cooper, L.S.; Crea, F.; Di Carli, M.; et al. Ischemia and No Obstructive Coronary Artery Disease (INOCA). Circulation 2017, 135, 1075–1092. [Google Scholar] [CrossRef] [PubMed]
- Tse, K.; Tse, H.; Sidney, J.; Sette, A.; Ley, K. T cells in atherosclerosis. Int. Immunol. 2013, 25, 615–622. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Golledge, J.; Norman, P.E. Atherosclerosis and abdominal aortic aneurysm: Cause, response, or common risk factors? Arterioscler. Thromb. Vasc. Biol. 2010, 30, 1075–1077. [Google Scholar] [CrossRef] [Green Version]
- Koroleva, I.A.; Nazarenko, M.S.; Kucher, A.N. Role of microRNA in development of instability of atherosclerotic plaques. Biochemistry 2017, 82, 1380–1390. [Google Scholar] [CrossRef]
- Sharma, S.; Jackson, P.G.; Makan, J. Cardiac troponins. J. Clin. Pathol. 2004, 57, 1025–1026. [Google Scholar] [CrossRef] [Green Version]
- Omland, T.; de Lemos, J.A.; Sabatine, M.S.; Christophi, C.A.; Rice, M.M.; Jablonski, K.A.; Tjora, S.; Domanski, M.J.; Gersh, B.J.; Rouleau, J.L.; et al. A sensitive cardiac troponin T assay in stable coronary artery disease. N. Engl. J. Med. 2009, 361, 2538–2547. [Google Scholar] [CrossRef] [Green Version]
- Speidl, W.S.; Graf, S.; Hornykewycz, S.; Nikfardjam, M.; Niessner, A.; Zorn, G.; Wojta, J.; Huber, K. High-sensitivity C-reactive protein in the prediction of coronary events in patients with premature coronary artery disease. Am. Heart J. 2002, 144, 449–455. [Google Scholar] [CrossRef]
- Sara, J.D.S.; Prasad, M.; Zhang, M.; Lennon, R.J.; Herrmann, J.; Lerman, L.O.; Lerman, A. High-sensitivity C-reactive protein is an independent marker of abnormal coronary vasoreactivity in patients with non-obstructive coronary artery disease. Am. Heart J. 2017, 190, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Tahhan, A.S.; Sandesara, P.; Hayek, S.S.; Hammadah, M.; Alkhoder, A.; Kelli, H.M.; Topel, M.; O’Neal, W.T.; Ghasemzadeh, N.; Ko, Y.A.; et al. High-sensitivity troponin I levels and coronary artery disease severity, progression, and long-term outcomes. J. Am. Heart Assoc. 2018, 7, e007914. [Google Scholar] [CrossRef] [Green Version]
- Zimmerli, L.U.; Schiffer, E.; Zürbig, P.; Good, D.M.; Kellmann, M.; Mouls, L.; Pitt, A.R.; Coon, J.J.; Schmieder, R.E.; Peter, K.H.; et al. Urinary proteomic biomarkers in coronary artery disease. Mol. Cell. Proteomics 2008, 7, 290–298. [Google Scholar] [CrossRef] [Green Version]
- Ibrahim, N.E.; Januzzi, J.L.; Magaret, C.A.; Gaggin, H.K.; Rhyne, R.F.; Gandhi, P.U.; Kelly, N.; Simon, M.L.; Motiwala, S.R.; Belcher, A.M.; et al. A Clinical and Biomarker Scoring System to Predict the Presence of Obstructive Coronary Artery Disease. J. Am. Coll. Cardiol. 2017, 69, 1147–1156. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandra, Y.; Devanna, P.; Limana, F.; Straino, S.; Di Carlo, A.; Brambilla, P.G.; Rubino, M.; Carena, M.C.; Spazzafumo, L.; De Simone, M.; et al. Circulating microRNAs are new and sensitive biomarkers of myocardial infarction. Eur. Heart J. 2010, 31, 2765–2773. [Google Scholar] [CrossRef] [Green Version]
- Condorelli, G.; Latronico, M.V.G.; Dorn, G.W. MicroRNAs in heart disease: Putative novel therapeutic targets? Eur. Heart J. 2010, 31, 649–658. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fichtlscherer, S.; De Rosa, S.; Fox, H.; Schwietz, T.; Fischer, A.; Liebetrau, C.; Weber, M.; Hamm, C.W.; Röxe, T.; Müller-Ardogan, M.; et al. Circulating microRNAs in patients with coronary artery disease. Circ. Res. 2010, 107, 677–684. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raitoharju, E.; Lyytikäinen, L.-P.; Levula, M.; Oksala, N.; Mennander, A.; Tarkka, M.; Klopp, N.; Illig, T.; Kähönen, M.; Karhunen, P.J.; et al. miR-21, miR-210, miR-34a, and miR-146a/b are up-regulated in human atherosclerotic plaques in the Tampere Vascular Study. Atherosclerosis 2011, 219, 211–217. [Google Scholar] [CrossRef]
- Raitoharju, E.; Oksala, N.; Lehtimäki, T. MicroRNAs in the atherosclerotic plaque. Clin. Chem. 2013, 59, 1708–1721. [Google Scholar] [CrossRef] [Green Version]
- Chan, M.C.; Hilyard, A.C.; Wu, C.; Davis, B.N.; Hill, N.S.; Lal, A.; Lieberman, J.; Lagna, G.; Hata, A. Molecular basis for antagonism between PDGF and the TGF b family of signalling pathways by control of miR-24 expression. EMBO J. 2009, 29, 559–573. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davis, B.N.; Hilyard, A.C.; Nguyen, P.H.; Lagna, G.; Hata, A. Induction of MicroRNA-221 by platelet-derived growth factor signaling is critical for modulation of vascular smooth muscle phenotype. J. Biol. Chem. 2009, 284, 3728–3738. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Cheng, Y.; Chen, X.; Yang, J.; Xu, L.; Zhang, C. MicroRNA-31 regulated by the extracellular regulated kinase is involved in vascular smooth muscle cell growth via large tumor suppressor homolog 2. J. Biol. Chem. 2011, 286, 42371–42380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Wang, Y.; Wang, X.; Zhang, Y.; Eisner, G.M.; Asico, L.D.; Jose, P.A.; Zeng, C. Insulin promotes vascular smooth muscle cell proliferation via microRNA-208-mediated downregulation of p21. J. Hypertens. 2011, 152, 66. [Google Scholar] [CrossRef]
- Leeper, N.J.; Raiesdana, A.; Kojima, Y.; Chun, H.J.; Azuma, J.; Maegdefessel, L.; Kundu, R.K.; Quertermous, T.; Tsao, P.S.; Spin, J.M. MicroRNA-26a is a novel regulator of vascular smooth muscle cell function. J. Cell. Physiol. 2011, 226, 1035–1043. [Google Scholar] [CrossRef] [Green Version]
- Hutcheson, R.; Chaplin, J.; Hutcheson, B.; Borthwick, F.; Proctor, S.; Gebb, S.; Jadhav, R.; Smith, E.; Russell, J.C.; Rocic, P. miR-21 normalizes vascular smooth muscle proliferation and improves coronary collateral growth in metabolic syndrome. FASEB J. 2014, 28, 4088–4099. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, J.; Shao, G.; Chen, X.; Yang, X.; Huang, X.; Peng, P.; Ba, Y. miRNA 206 and miRNA 574-5p are highly expression in coronary artery disease. Biosci. Rep. 2016, 36, e00295. [Google Scholar] [CrossRef] [PubMed]
- Li, H.Y.; Zhao, X.; Liu, Y.Z.; Meng, Z.; Wang, D.; Yang, F.; Shi, Q.W. Plasma MicroRNA-126-5p is Associated with the Complexity and Severity of Coronary Artery Disease in Patients with Stable Angina Pectoris. Cell. Physiol. Biochem. 2016, 39, 837–846. [Google Scholar] [CrossRef]
- Wang, H.W.; Lo, H.H.; Chiu, Y.L.; Chang, S.J.; Huang, P.H.; Liao, K.H.; Tasi, C.F.; Wu, C.H.; Tsai, T.N.; Cheng, C.C.; et al. Dysregulated miR-361-5p/VEGF axis in the plasma and endothelial progenitor cells of patients with coronary artery disease. PLoS ONE 2014, 9, e98070. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Ji, Y.; Cai, S.; Ding, W. MiR-206 suppresses the progression of coronary artery disease by modulating vascular endothelial growth factor (VEGF) expression. Med. Sci. Monit. 2016, 22, 5011–5020. [Google Scholar] [CrossRef] [Green Version]
- Adachi, T.; Nakanishi, M.; Otsuka, Y.; Nishimura, K.; Hirokawa, G.; Goto, Y.; Nonogi, H.; Iwai, N. Plasma microRNA 499 as a biomarker of acute myocardial infarction. Clin. Chem. 2010, 56, 1183–1185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Corsten, M.F.; Dennert, R.; Jochems, S.; Kuznetsova, T.; Devaux, Y.; Hofstra, L.; Wagner, D.R.; Staessen, J.A.; Heymans, S.; Schroen, B. Circulating MicroRNA-208b and MicroRNA-499 reflect myocardial damage in cardiovascular disease. Circ. Cardiovasc. Genet. 2010, 3, 499–506. [Google Scholar] [CrossRef] [PubMed]
- Ji, X.; Takahashi, R.; Hiura, Y.; Hirokawa, G.; Fukushima, Y.; Iwai, N. Plasma miR-208 as a biomarker of myocardial injury. Clin. Chem. 2009, 55, 1944–1949. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ai, J.; Zhang, R.; Li, Y.; Pu, J.; Lu, Y.; Jiao, J.; Li, K.; Yu, B.; Li, Z.; Wang, R.; et al. Circulating microRNA-1 as a potential novel biomarker for acute myocardial infarction. Biochem. Biophys. Res. Commun. 2010, 391, 73–77. [Google Scholar] [CrossRef] [PubMed]
- Hoekstra, M.; van der Lans, C.A.C.; Halvorsen, B.; Gullestad, L.; Kuiper, J.; Aukrust, P.; van Berkel, T.J.C.; Biessen, E.A.L. The peripheral blood mononuclear cell microRNA signature of coronary artery disease. Biochem. Biophys. Res. Commun. 2010, 394, 792–797. [Google Scholar] [CrossRef]
- Takahashi, Y.; Satoh, M.; Minami, Y.; Tabuchi, T.; Itoh, T.; Nakamura, M. Expression of miR-146a/b is associated with the Toll-like receptor 4 signal in coronary artery disease: Effect of renin-angiotensin system blockade and statins on miRNA-146a/b and Toll-like receptor 4 levels. Clin. Sci. 2010, 119, 395–405. [Google Scholar] [CrossRef] [Green Version]
- Satoh, M.; Tabuchi, T.; Minami, Y.; Takahashi, Y.; Itoh, T.; Nakamura, M. Expression of let-7i is associated with Toll-like receptor 4 signal in coronary artery disease: Effect of statins on let-7i and Toll-like receptor 4 signal. Immunobiology 2012, 217, 533–539. [Google Scholar] [CrossRef]
- Willeit, P.; Zampetaki, A.; Dudek, K.; Kaudewitz, D.; King, A.; Kirkby, N.S.; Crosby-Nwaobi, R.; Prokopi, M.; Drozdov, I.; Langley, S.R.; et al. Circulating MicroRNAs as novel biomarkers for platelet activation. Circ. Res. 2013, 112, 595–600. [Google Scholar] [CrossRef] [Green Version]
- Das, A.; Samidurai, A.; Salloum, F.N. Deciphering Non-coding RNAs in Cardiovascular Health and Disease. Front. Cardiovasc. Med. 2018, 5, 73. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Zhang, L.; Wang, Y.; Ding, H.; Xue, S.; Qi, H.; Li, P. MicroRNAs or long noncoding RNAs in diagnosis and prognosis of coronary artery disease. Aging Dis. 2019, 10, 353–366. [Google Scholar] [CrossRef] [Green Version]
- Elmén, J.; Lindow, M.; Silahtaroglu, A.; Bak, M.; Christensen, M.; Lind-Thomsen, A.; Hedtjärn, M.; Hansen, J.B.; Hansen, H.F.; Straarup, E.M.; et al. Antagonism of microRNA-122 in mice by systemically administered LNA-antimiR leads to up-regulation of a large set of predicted target mRNAs in the liver. Nucleic Acids Res. 2008, 36, 1153–1162. [Google Scholar] [CrossRef] [Green Version]
- Dong, J.; Liang, Y.Z.; Zhang, J.; Wu, L.J.; Wang, S.; Hua, Q.; Yan, Y.X. Potential role of lipometabolism-related microRNAs in peripheral blood mononuclear cells as biomarkers for coronary artery disease. J. Atheroscler. Thromb. 2017, 24, 430–441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moore, K.J.; Rayner, K.J.; Suárez, Y.; Fernández-Hernando, C. The role of microRNAs in cholesterol efflux and hepatic lipid metabolism. Annu. Rev. Nutr. 2011, 31, 49–63. [Google Scholar] [CrossRef] [Green Version]
- Marquart, T.J.; Allen, R.M.; Ory, D.S.; Baldán, Á. miR-33 links SREBP-2 induction to repression of sterol transporters. Proc. Natl. Acad. Sci. USA 2010, 107, 12228–12232. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iliopoulos, D.; Drosatos, K.; Hiyama, Y.; Goldberg, I.J.; Zannis, V.I. MicroRNA-370 controls the expression of MicroRNA-122 and Cpt1α and affects lipid metabolism. J. Lipid Res. 2010, 51, 1513–1523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Faccini, J.; Ruidavets, J.B.; Cordelier, P.; Martins, F.; Maoret, J.J.; Bongard, V.; Ferrières, J.; Roncalli, J.; Elbaz, M.; Vindis, C. Circulating MIR-155, MIR-145 and let-7c as diagnostic biomarkers of the coronary artery disease. Sci. Rep. 2017, 7, 42916. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.; Kong, D.; Chen, H.; Liu, S.; Hu, H.; Wu, T.; Wang, J.; Chen, W.; Ning, Y.; Li, Y.; et al. MiR-155 acts as an anti-inflammatory factor in atherosclerosis-Associated foam cell formation by repressing calcium-regulated heat stable protein 1. Sci. Rep. 2016, 6, 21789. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, H.S.; Sivachandran, N.; Lau, A.; Boudreau, E.; Zhao, J.L.; Baltimore, D.; Delgado-Olguin, P.; Cybulsky, M.I.; Fish, J.E. MicroRNA-146 represses endothelial activation by inhibiting pro-inflammatory pathways. EMBO Mol. Med. 2013, 5, 949–966. [Google Scholar] [CrossRef]
- Huang, R.S.; Gamazon, E.R.; Ziliak, D.; Wen, Y.; Im, H.K.; Zhang, W.; Wing, C.; Duan, S.; Bleibel, W.K.; Cox, N.J.; et al. Population differences in microRNA expression and biological implications. RNA Biol. 2011, 8, 692–701. [Google Scholar] [CrossRef] [Green Version]
- Rawlings-Goss, R.A.; Campbell, M.C.; Tishkoff, S.A. Global population-specific variation in miRNA associated with cancer risk and clinical biomarkers. BMC Med. Genomics 2014, 7, 53. [Google Scholar] [CrossRef] [Green Version]
- Meder, B.; Backes, C.; Haas, J.; Leidinger, P.; Stähler, C.; Großmann, T.; Vogel, B.; Frese, K.; Giannitsis, E.; Katus, H.A.; et al. Influence of the confounding factors age and sex on microRNA profiles from peripheral blood. Clin. Chem. 2014, 60, 1200–1208. [Google Scholar] [CrossRef] [PubMed]
- Simon, L.M.; Edelstein, L.C.; Nagalla, S.; Woodley, A.B.; Chen, E.S.; Kong, X.; Ma, L.; Fortina, P.; Kunapuli, S.; Holinstat, M.; et al. Human platelet microRNA-mRNA networks associated with age and gender revealed by integrated plateletomics. Blood 2014, 123, 37–45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sayed, A.S.M.; Xia, K.; Li, F.; Deng, X.; Salma, U.; Li, T.; Deng, H.; Yang, D.; Haoyang, Z.; Yang, T.L.; et al. The diagnostic value of circulating microRNAs for middle-aged (40–60-year-old) coronary artery disease patients. Clinics 2015, 70, 257–263. [Google Scholar] [CrossRef]
- Olivieri, F.; Bonafè, M.; Spazzafumo, L.; Gobbi, M.; Prattichizzo, F.; Recchioni, R.; Marcheselli, F.; La Sala, L.; Galeazzi, R.; Rippo, M.R.; et al. Age- and glycemia-related miR-126-3p levels in plasma and endothelial cells. Aging 2014, 6, 771–786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ultimo, S.; Zauli, G.; Martelli, A.M.; Vitale, M.; McCubrey, J.A.; Capitani, S.; Neri, L.M. Cardiovascular disease-related miRNAs expression: Potential role as biomarkers and effects of training exercise. Oncotarget 2018, 9, 17238–17254. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gupta, S.K.; Foinquinos, A.; Thum, S.; Remke, J.; Zimmer, K.; Bauters, C.; de Groote, P.; Boon, R.A.; de Windt, L.J.; Preissl, S.; et al. Preclinical Development of a MicroRNA-Based Therapy for Elderly Patients With Myocardial Infarction. J. Am. Coll. Cardiol. 2016, 68, 1557–1571. [Google Scholar] [CrossRef] [Green Version]
- De Lucia, C.; Komici, K.; Borghetti, G.; Femminella, G.D.; Bencivenga, L.; Cannavo, A.; Corbi, G.; Ferrara, N.; Houser, S.R.; Koch, W.J.; et al. MicroRNA in cardiovascular aging and age-related cardiovascular diseases. Front. Med. Lausanne 2017, 4, 74. [Google Scholar] [CrossRef] [Green Version]
- Van Almen, G.C.; Verhesen, W.; van Leeuwen, R.E.W.; van de Vrie, M.; Eurlings, C.; Schellings, M.W.M.; Swinnen, M.; Cleutjens, J.P.M.; van Zandvoort, M.A.M.J.; Heymans, S.; et al. MicroRNA-18 and microRNA-19 regulate CTGF and TSP-1 expression in age-related heart failure. Aging Cell 2011, 10, 769–779. [Google Scholar] [CrossRef] [Green Version]
- Zhou, M.; Cai, J.; Tang, Y.; Zhao, Q. MiR-17-92 cluster is a novel regulatory gene of cardiac ischemic/reperfusion injury. Med. Hypotheses 2013, 81, 108–110. [Google Scholar] [CrossRef]
- Zhong, Z.; Hou, J.; Zhang, Q.; Zhong, W.; Li, B.; Li, C.; Liu, Z.; Yang, M.; Zhao, P. Circulating microRNA expression profiling and bioinformatics analysis of dysregulated microRNAs of patients with coronary artery disease. Medicine 2018, 97, e11428. [Google Scholar] [CrossRef]
- Wang, K.; Yuan, Y.; Cho, J.H.; McClarty, S.; Baxter, D.; Galas, D.J. Comparing the MicroRNA spectrum between serum and plasma. PLoS ONE 2012, 7, e41561. [Google Scholar] [CrossRef] [PubMed]
- Blondal, T.; Jensby Nielsen, S.; Baker, A.; Andreasen, D.; Mouritzen, P.; Wrang Teilum, M.; Dahlsveen, I.K. Assessing sample and miRNA profile quality in serum and plasma or other biofluids. Methods 2013, 59, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Kroh, E.M.; Parkin, R.K.; Mitchell, P.S.; Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 2010, 50, 298–301. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sunderland, N.; Skroblin, P.; Barwari, T.; Huntley, R.P.; Lu, R.; Joshi, A.; Lovering, R.C.; Mayr, M. MicroRNA Biomarkers and Platelet Reactivity: The Clot Thickens. Circ. Res. 2017, 120, 418–435. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peirson, S.N.; Butler, J.N. RNA Extraction From Mammalian Tissues. In Methods in Molecular Biology; Rosato, E., Ed.; Humana Press: Totowa, NJ, USA, 2007; pp. 315–327. ISBN 978-1-59745-257-1. [Google Scholar]
- Glinge, C.; Clauss, S.; Boddum, K.; Jabbari, R.; Jabbari, J.; Risgaard, B.; Tomsits, P.; Hildebrand, B.; Kääb, S.; Wakili, R.; et al. Stability of Circulating Blood-Based MicroRNAs—Pre-Analytic Methodological Considerations. PLoS ONE 2017, 12, e0167969. [Google Scholar] [CrossRef]
- Mayeux, R. Biomarkers: Potential Uses and Limitations. NeuroRx 2004, 1, 182–188. [Google Scholar] [CrossRef]
- Jansen, F.; Yang, X.; Proebsting, S.; Hoelscher, M.; Przybilla, D.; Baumann, K.; Schmitz, T.; Dolf, A.; Endl, E.; Franklin, B.S.; et al. MicroRNA expression in circulating microvesicles predicts cardiovascular events in patients with coronary artery disease. J. Am. Heart Assoc. 2014, 3, e001249. [Google Scholar] [CrossRef] [Green Version]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef] [Green Version]
- Hayes, J.; Peruzzi, P.P.; Lawler, S. MicroRNAs in cancer: Biomarkers, functions and therapy. Trends Mol. Med. 2014, 20, 460–469. [Google Scholar] [CrossRef]
- Hongyan, Z.; Guo-chang, F. Extracellular/circulating microRNAs and their potential role in cardiovascular disease. Am. J. Cardiovasc. Dis. 2011, 1, 138–149. [Google Scholar]
- Trzybulska, D.; Vergadi, E.; Tsatsanis, C. MiRNA and other non-coding RNAs as promising diagnostic markers. Electron. J. Int. Fed. Clin. Chem. Lab. Med. 2018, 29, 221–226. [Google Scholar]
- Pogribny, I.P. MicroRNAs as biomarkers for clinical studies. Exp. Biol. Med. 2018, 243, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Kreth, S.; Hübner, M.; Hinske, L.C. MicroRNAs as clinical biomarkers and therapeutic tools in perioperative medicine. Anesth. Analg. 2018, 126, 670–681. [Google Scholar] [CrossRef]
- Wang, J.; Yan, Y.; Song, D.; Liu, B. Reduced Plasma miR-146a is a Predictor of Poor Coronary Collateral Circulation in Patients with Coronary Artery Disease. Biomed Res. Int. 2016, 2016, 4285942. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Lee, C.; Song, J.; Lu, C.; Liu, J.; Cui, Y.; Liang, H.; Cao, C.; Zhang, F.; Chen, H. Circulating microRNAs as potential biomarkers for coronary plaque rupture. Oncotarget 2017, 8, 48145–48156. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.D.; Yang, Y.J.; Wang, L.Y.; Qiao, S.B.; Lu, X.F.; Wu, Y.J.; Xu, B.; Li, H.F.; Gu, D.F. Elevated plasma miRNA-122, -140-3p, -720, -2861, and -3149 during early period of acute coronary syndrome are derived from peripheral blood mononuclear cells. PLoS ONE 2017, 12, e0184256. [Google Scholar] [CrossRef] [Green Version]
- Jansen, F.; Schäfer, L.; Wang, H.; Schmitz, T.; Flender, A.; Schueler, R.; Hammerstingl, C.; Nickenig, G.; Sinning, J.; Werner, N. Kinetics of Circulating MicroRNAs in Response to Cardiac Stress in Patients With Coronary Artery Disease. J. Am. Heart Assoc. 2017, 6, e005270. [Google Scholar] [CrossRef]
- Soeki, T.; Yamaguchi, K.; Niki, T.; Uematsu, E.; Bando, S.; Matsuura, T.; Ise, T.; Kusunose, K.; Hotchi, J.; Tobiume, T.; et al. Plasma microRNA-100 is associated with coronary plaque vulnerability. Circ. J. 2015, 79, 413–418. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.Z.; Zhong, Q.; Huang, Y.Q. Elevated plasma miR-29a levels are associated with increased carotid intima-media thickness in atherosclerosis patients. Tohoku J. Exp. Med. 2017, 241, 183–188. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Lian, Y.; Wen, X.; Guo, J.; Wang, Z.; Jiang, S.; Hu, Y. Expression of miR-126 and its potential function in coronary artery disease. Afr. Health Sci. 2017, 17, 474–480. [Google Scholar] [CrossRef] [Green Version]
- Al-Kafaji, G.; Al-Mahroos, G.; Abdulla Al-Muhtaresh, H.; Sabry, M.A.; Abdul Razzak, R.; Salem, A.H. Circulating endothelium-enriched microRNA-126 as a potential biomarker for coronary artery disease in type 2 diabetes mellitus patients. Biomarkers 2017, 22, 268–278. [Google Scholar] [CrossRef] [PubMed]
- Al-Muhtaresh, H.A.; Salem, A.H.; Al-Kafaji, G. Upregulation of Circulating Cardiomyocyte-Enriched miR-1 and miR-133 Associate with the Risk of Coronary Artery Disease in Type 2 Diabetes Patients and Serve as Potential Biomarkers. J. Cardiovasc. Transl. Res. 2019, 12, 347–357. [Google Scholar] [CrossRef]
- Boon, R.A. Non-coding RNAs in cardiovascular health and disease. Non Coding RNA Res. 2018, 3, 99. [Google Scholar] [CrossRef]
- Wang, H.W.; Huang, T.S.; Lo, H.H.; Huang, P.H.; Lin, C.C.; Chang, S.J.; Liao, K.H.; Tsai, C.H.; Chan, C.H.; Tsai, C.F.; et al. Deficiency of the MicroRNA-31-MicroRNA-720 pathway in the plasma and endothelial progenitor cells from patients with coronary artery disease. Arterioscler. Thromb. Vasc. Biol. 2014, 34, 857–869. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Li, H.-H.; Yang, R.; Yang, B.-J.; Gao, Z.-Y. Association between circulating microRNA-208a and severity of coronary heart disease. Scand. J. Clin. Lab. Investig. 2017, 77, 379–384. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Qu, G.; Han, C.; Wang, Y.; Sun, T.; Li, F.; Wang, J.; Luo, S. MiR-34a, miR-21 and miR-23a as potential biomarkers for coronary artery disease: A pilot microarray study and confirmation in a 32 patient cohort. Exp. Mol. Med. 2015, 47, 138. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, H.; Guddeti, R.R.; Matsuzawa, Y.; Liu, L.P.; Su, L.X.; Guo, D.; Nie, S.P.; Du, J.; Zhang, M. Plasma levels of microRNA-145 are associated with severity of coronary artery disease. PLoS ONE 2015, 10, e0123477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ren, J.; Zhang, J.; Xu, N.; Han, G.; Geng, Q.; Song, J.; Li, S.; Zhao, J.; Chen, H. Signature of circulating MicroRNAs As potential biomarkers in vulnerable coronary artery disease. PLoS ONE 2013, 8, e80738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.; Xu, L.; Hu, Q.; Yang, S.; Zhang, B.; Jiang, H. MiR-17-5p as circulating biomarkers for the severity of coronary atherosclerosis in coronary artery disease. Int. J. Cardiol. 2015, 197, 123–124. [Google Scholar] [CrossRef] [PubMed]
- Lin, B.; Feng, D.-G.; Wang, F.; Wang, J.-X.; Xu, C.-G.; Zhao, H.; Cheng, Z.-Y. MiR-365 participates in coronary atherosclerosis through regulating IL-6. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 5186–5192. [Google Scholar]
- Cipollone, F.; Felicioni, L.; Sarzani, R.; Ucchino, S.; Spigonardo, F.; Mandolini, C.; Malatesta, S.; Bucci, M.; Mammarella, C.; Santovito, D.; et al. A unique MicroRNA signature associated with plaque instability in humans. Stroke 2011, 42, 2556–2563. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Kim, C.W.; Simmons, R.D.; Jo, H. Role of flow-sensitive microRNAs in endothelial dysfunction and atherosclerosis—“Mechanosensitive Athero-miRs”. Arter. Thromb Vasc Biol. 2014, 22, 313–333. [Google Scholar] [CrossRef] [Green Version]
- Tian, F.J.; An, L.N.; Wang, G.K.; Zhu, J.Q.; Li, Q.; Zhang, Y.Y.; Zeng, A.; Zou, J.; Zhu, R.F.; Han, X.S.; et al. Elevated microRNA-155 promotes foam cell formation by targeting HBP1 in atherogenesis. Cardiovasc. Res. 2014, 103, 100–110. [Google Scholar] [CrossRef]
- Horie, T.; Baba, O.; Kuwabara, Y.; Chujo, Y.; Watanabe, S.; Kinoshita, M.; Horiguchi, M.; Nakamura, T.; Chonabayashi, K.; Hishizawa, M.; et al. MicroRNA-33 deficiency reduces the progression of atherosclerotic plaque in ApoE−/− mice. J. Am. Heart Assoc. 2012, 1, e003376. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, Y.; Shi, C.; Manduchi, E.; Civelek, M.; Davies, P.F. MicroRNA-10a regulation of proinflammatory phenotype in athero-susceptible endothelium in vivo and in vitro. Proc. Natl. Acad. Sci. USA 2010, 107, 13450–13455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shan, Z.; Qin, S.; Li, W.; Wu, W.; Yang, J.; Chu, M.; Li, X.; Huo, Y.; Schaer, G.L.; Wang, S.; et al. An Endocrine Genetic Signal Between Blood Cells and Vascular Smooth Muscle Cells: Role of MicroRNA-223 in Smooth Muscle Function and Atherogenesis. J. Am. Coll. Cardiol. 2015, 65, 2526–2537. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bidzhekov, K.; Gan, L.; Denecke, B.; Rostalsky, A.; Hristov, M.; Koeppel, T.A.; Zernecke, A.; Weber, C. microRNA expression signatures and parallels between monocyte subsets and atherosclerotic plaque in humans. Thromb. Haemost. 2012, 107, 619–625. [Google Scholar] [CrossRef] [PubMed]
- Jansen, F.; Yang, X.; Hoelscher, M.; Cattelan, A.; Schmitz, T.; Proebsting, S.; Wenzel, D.; Vosen, S.; Franklin, B.S.; Fleischmann, B.K.; et al. Endothelial microparticle-mediated transfer of microRNA-126 promotes vascular endothelial cell repair via spred1 and is abrogated in glucose-damaged endothelial microparticles. Circulation 2013, 128, 2026–2038. [Google Scholar] [CrossRef] [Green Version]
- Schulte, C.; Molz, S.; Appelbaum, S.; Karakas, M.; Ojeda, F.; Lau, D.M.; Hartmann, T.; Lackner, K.J.; Westermann, D.; Schnabel, R.B.; et al. MiRNA-197 and miRNA-223 predict cardiovascular death in a cohort of patients with symptomatic coronary artery disease. PLoS ONE 2015, 10, e0145930. [Google Scholar] [CrossRef] [Green Version]
- Hulsmans, M.; Sinnaeve, P.; Van Der Schueren, B.; Mathieu, C.; Janssens, S.; Holvoet, P. Decreased miR-181a expression in monocytes of obese patients is associated with the occurrence of metabolic syndrome and coronary artery disease. J. Clin. Endocrinol. Metab. 2012, 97, 1213–1218. [Google Scholar] [CrossRef] [Green Version]
| References | miRNA | Quantitative Effect | Outcome | Sample Type | miRNA Identification/Quantification Method | Cell Lines/Study Population | Acute/Chronic Disease Status |
|---|---|---|---|---|---|---|---|
| Wang et al., 2016 [118] | miRNA-146a | Upregulated | This miRNA may be a potential biomarker for poor coronary collateral circulation in CAD patients. | Plasma | qRT-PCR | Human patients | Chronic (1-month cut-off) |
| Li et al., 2017 [119] | miRNA-155-5p miRNA-483-5p miRNA-451a | MiRNA-155-5p and miRNA-483-5p are upregulated; miRNA-451a is down-regulated | Potential biomarkers for the early detection of atherosclerotic plaque rupture. | Plasma | qRT-PCR | Human patients | Stable CAD |
| Zhao et al., 2015 [37] | miRNA-143 miRNA-145 | Contested | Altered in CAD. Potentially released from vascular walls. | Plasma | (Review article) | (Review article) | (Review article) |
| Li et al., 2017 [120] | miRNA-122 miRNA-140-3p miRNA-720 miRNA-2861 miRNA-3149 | Upregulated | Elevated during the early stages of ACS. | Plasma | qRT-PCR | Bama male minipigs and human patients | Minipigs: normal and acute MI. Human patients: Stable angina, unstable angina and acute MI. |
| Jansen et al., 2017 [121] | miRNA-21 miRNA-126-3p miRNA-222 | Upregulated | These miRNAs increased in concentration following periods of cardiac stress in patients with stenosed coronary arteries. | Plasma | qRT-PCR | Human patients | Stable CAD |
| Soeki et al., 2015 [122] | miRNA-100 | - | Associated with coronary plaque instability. Potentially released from plaques. | Plasma | qRT-PCR | Human patients | Unknown |
| Liu et al., 2017 [123] | miRNA-29a | Upregulated | Moderates expression of mRNAs of extracellular matrix proteins. Associated with atherosclerosis and intima-media thickness of carotid arteries. | Plasma | qRT-PCR | Human patients | Unknown |
| Wang et al., 2017 [124] | miRNA-126 | Downregulated | A potential biomarker for CAD. Inversely correlated to placenta growth factor. | Plasma | qRT-PCR | Human patients | CAD for 15–24 months |
| Al-Kafaji et al., 2017 [125] | miRNA-126 | Downregulated | A potential biomarker for CAD. Inversely correlated with LDL concentration. | Plasma | qRT-PCR | Human patients | Type 2 diabetics, some with CAD diagnoses |
| Al-Muhtaresh et al., 2019 [126] | miRNA-1 miRNA-133 | Upregulated | Potential biomarkers. Both correlate with LDL-C levels; miR-1 is known to negatively regulate Bcl2 [127]. | Plasma | qRT-PCR | Human patients | Type 2 diabetics, some with CAD diagnoses |
| Zernecke et al., 2009 [14] | miRNA-126 | - | Released from apoptotic bodies derived from endothelial cells from atherosclerotic plaques. Reduces inflammatory activity/plaque development. | Plasma/Plaque | qRT-PCR | Human aortic smooth muscle cell culture. Human atherosclerotic plaques. ApoE−/− murine endothelial cell cultures. HUVEC cell line | Unknown |
| Wang et al., 2014 [128] | miRNA-31 miRNA-720 | Downregulated | Potential biomarkers for early CAD. | Plasma/endothelial progenitor cells | qRT-PCR | Human patients | Unknown CAD |
| Zhang et al., 2017 [129] | miRNA-208a | - | Significant association with Gensini score, and by extension the severity of atherosclerosis. Potential biomarker for CAD severity. | Plasma | qRT-PCR | Human patients | Unknown CAD |
| Jansen et al., 2014 [111] | miRNA-126 miRNA-199a | - | The levels of these miRNA, which occur in circulating microvesicles, are potentially prognostic for major adverse cardiovascular events in patients with stable CAD. | Plasma | qRT-PCR | Human patients | Stable CAD |
| Han et al., 2015 [130] | miRNA-21 miRNA-23a miRNA-30a miRNA-34a miRNA-106b | Upregulated | These miRNAs occur at higher levels in ApoE−/− mice, which models hypercholesterolaemia. MiRNA-21, -23a, and -34a are potential biomarkers for CAD. MiRNA-21 has been linked to CAD-derived ACS. | Plasma | qRT-PCR and miRNA microarrays | ApoE−/− mice and human CAD patients | Unknown |
| Zhou et al., 2016 [70] | miRNA-206 miRNA-564-5p | Upregulated | Potential biomarkers for CAD | Plasma | qRT-PCR and miRNA microarrays | Human patients | Unknown |
| Sayed et al., 2015 [96] | miRNA-149 miRNA-424 miRNA-765 | MiRNA-149 and miRNA-424 were upregulated, miRNA-765 was downregulated | Potential biomarkers for CAD in middle-aged patients | Plasma | qRT-PCR | Human patients | Stable and unstable CAD |
| Gao et al., 2015 [131] | miRNA-145 | Downregulated | This miRNA regulates VSMC fate, inhibiting proliferation. It is the modal miRNA in healthy vessel walls, though in atherosclerotic plaques it may not even be detected. Plasma concentration levels are significantly reduced in CAD patients, and those with three-vessel disease have a significantly lower quantity as well. Potential biomarker for CAD. | Plasma/plaque | qRT-PCR | Human patients | Unknown (patients diagnosed with CAD for more than a year) |
| Ren et al., 2013 [132] | miRNA-106b/25 cluster miRNA-17/92a cluster miRNA-21/590-5p cluster miRNA-126 miRNA-451 | Upregulated in patients with unstable angina, though there is evidence that miRNA-17/92a was actually downregulated in CAD patients [83] | These miRNAs are elevated in CAD patients relative to those with stable AP. MiRNA-17/92a is involved in angiogenesis, which further complicates plaques. Increased miRNA-21 can yield increased MMP activity, which can hinder plaque progression. Potential biomarkers for CAD. | Plasma | qRT-PCR | Human patients | CAD and unstable angina |
| Chen et al., 2015 [133] | miRNA-17-5p | Upregulated | Potential biomarker for early CAD. | Plasma | qRT-PCR | Human patients | Unknown |
| Faccini et al., 2017 [89] | miRNA-155 miRNA-145 let-7c | Downregulated | Potential biomarkers for CAD | Plasma | qRT-PCR and miRNA microarrays | Human patients | Unknown |
| Koroleva et al., 2017 [51] | miRNA-21 miRNA-100 miRNA-127 miRNA-133 miRNA-143/145 miRNA-221/222 miRNA-494 | All upregulated apart from miRNA-221/222, which was downregulated | The expression of these miRNA may influence plaque stability: miRNA-21, -143, and -221 are pro-stability; miRNA-100, -127, -133, and -494 are pro-instability. | Plaque | (Review article) | (Review article) | (Review article) |
| Lin et al., 2016 [134] | miRNA-365 | Downregulated | Regulation of the inflammatory response, specifically IL-6 activity, such that IL-6 expression increases as miRNA-365 expression decreases. | Plaque, serum, and circulating monocytes | qRT-PCR | Human patients | Unknown (patients with atherosclerosis) |
| Cipollone et al., 2011 [135] | miRNA-100 miRNA-127 miRNA-145 miRNA-133a/b | Upregulated | The expression of these miRNA varies with plaque stability. MiRNA-133 is relevant to stroke-related proteins and is thought to be vascular smooth muscle-specific. | Plaque | qRT-PCR | Human patients | Unknown |
| Kumar et al., 2014 [136] | miRNA-712 miRNA-205 | Upregulated in atherosclerosis | These miRNA target and reduce expression of metalloproteinase inhibitor 3 (TIMP3), increasing the activity of matrix metalloproteinases (MMPs), which affects inflammatory processes and VSMC/leukocyte migration in atherosclerosis. | Endothelial cells (Plaque) | Review (qRT-PCR, microarrays, and fluorescent in situ hybridisation) | Review (mice (C57BL/6 and ApoE−/−)) | Review (unknown) |
| Tian et al., 2014 [137] | miRNA-155 | Upregulated | Raised inflammatory response and foam cell differentiation. | Monocytes (plaque) | qRT-PCR | ApoE−/− mice | Unknown |
| Horie et al., 2012 [138] | miRNA-33 | - | Deficiency in ApoE knockout mice suppressed atherogenesis/plaque progression. | Monocytes/macrophages (plaque) | qRT-PCR | ApoE−/− mice | Unknown |
| Fang et al., 2010 [139] | miRNA-10a | Downregulated | Expression levels were reduced in endothelial cells that are thought to be pre-atherosclerotic, affecting inflammation signalling. | Endothelial cells (plaque) | qRT-PCR, miRNA microarrays, and fluorescent in situ hybridisation | Adult pigs | Unknown |
| Zernecke et al., 2009 [14] | miRNA-126 | - | Released from apoptotic bodies derived from endothelial cells from atherosclerotic plaques. MiRNAs reduce inflammatory activity/plaque development. | Plasma/plaque | qRT-PCR | Human aortic smooth muscle cell culture. Human atherosclerotic plaques. ApoE−/− murine endothelial cell cultures. HUVEC cell line. | Unknown |
| Raitoharju et al., 2011 [62] | miRNA-21 miRNA-34a miRNA-146a miRNA-146b-5p miRNA-210 | Upregulated | These miRNAs were upregulated in plaques compared to left internal thoracic arteries that were not atherosclerotic. This has been linked to VSMC changes seen in atherogenesis. | Plaque | miRNA microarrays and qRT-PCR | Human patients | Unknown |
| Shan et al., 2015 [140] | miRNA-223 | Upregulated | This miRNAs seems to be secreted from cells in the circulation. Their levels are elevated in the serum and atherosclerotic lesions in apolipoprotein-E knockout mice. | Plaque serum/blood cells | qRT-PCR | Sprague–Dawley rat VSMC cultures and C67BL/6 murine platelets | Unknown |
| Bidzhekov et al., 2012 [141] | miRNA-26b miRNA30e-5p miRNA-105 miRNA125a-5p miRNA-520b | MiRNA-26b, -30e-5p, and -125a-5p were upregulated. MiRNA-105 and miRNA-520b were downregulated. | These miRNAs had altered expression in CAD patients relative to healthy controls. | Plaque, monocytes | qRT-PCR and miRNA microarrays | Human patients | Unknown |
| Jansen et al., 2013 [142] | miRNA-126 | Downregulated | Circulating levels of miRNA-126 decreased in CAD patients. | Circulating microparticles | qRT-PCR | Mice and human patients | Stable CAD since 2003 |
| Schulte et al., 2015 [143] | miRNA-197 miRNA-223 | - | Strong prognostic value in CAD patients for cardiac death. | Serum | qRT-PCR | Human patients | Unknown CAD |
| Hulsmans et al., 2012 [144] | miRNA-181a | Downregulated | Potential biomarker for CAD, as well as metabolic syndrome | Monocytes | qRT-PCR and miRNA microarrays | Human patients | Unknown |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fazmin, I.T.; Achercouk, Z.; Edling, C.E.; Said, A.; Jeevaratnam, K. Circulating microRNA as a Biomarker for Coronary Artery Disease. Biomolecules 2020, 10, 1354. https://doi.org/10.3390/biom10101354
Fazmin IT, Achercouk Z, Edling CE, Said A, Jeevaratnam K. Circulating microRNA as a Biomarker for Coronary Artery Disease. Biomolecules. 2020; 10(10):1354. https://doi.org/10.3390/biom10101354
Chicago/Turabian StyleFazmin, Ibrahim T., Zakaria Achercouk, Charlotte E. Edling, Asri Said, and Kamalan Jeevaratnam. 2020. "Circulating microRNA as a Biomarker for Coronary Artery Disease" Biomolecules 10, no. 10: 1354. https://doi.org/10.3390/biom10101354