Targeted Clinical Metabolite Profiling Platform for the Stratification of Diabetic Patients
Abstract
:1. Introduction
2. Results
2.1. Sample Preparation
2.2. LC-MS
2.3. Method Validation
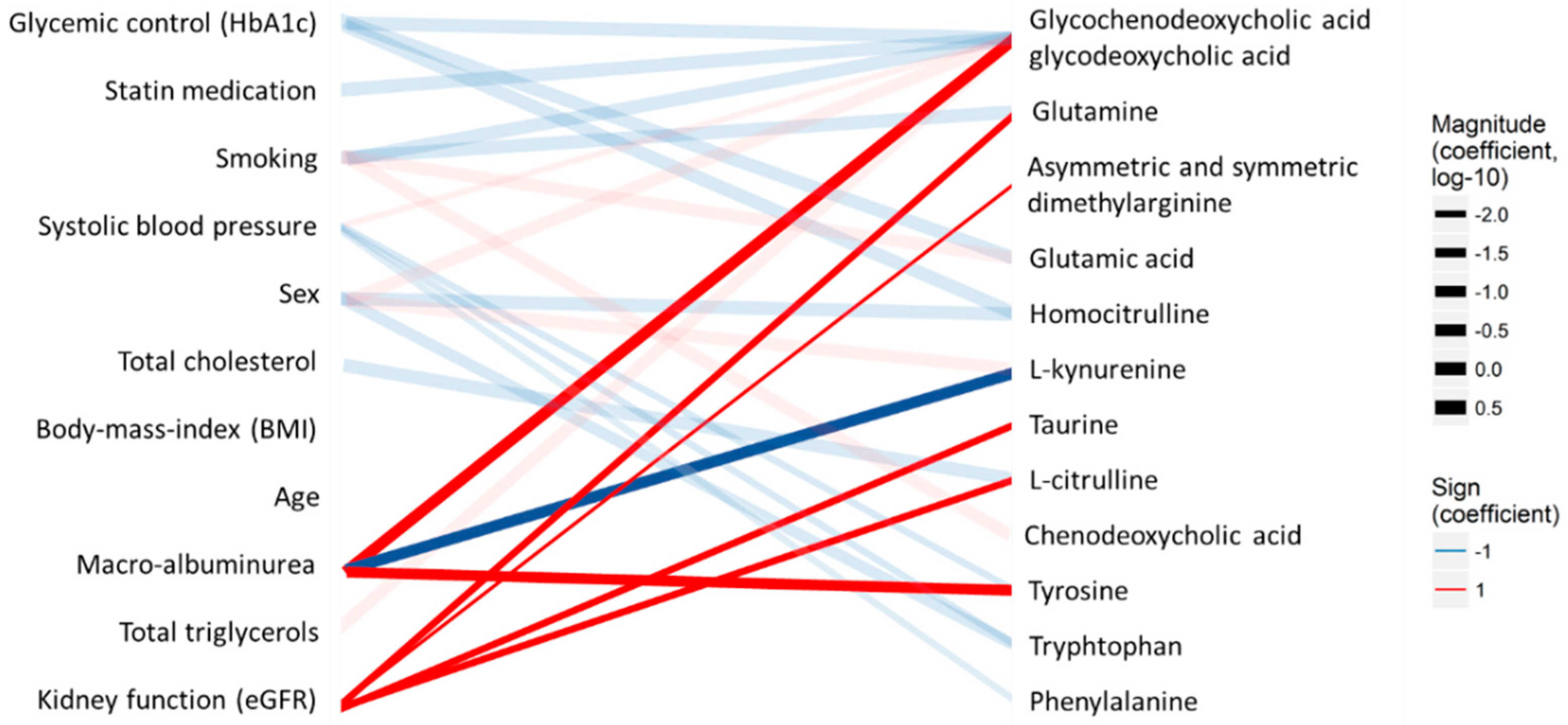
2.4. Feasibility of the Method for the Analysis of Samples from a Diabetes Cohort
3. Discussion
4. Materials and Methods
4.1. Chemicals and Standard Solutions
4.2. Samples
4.3. Sample Preparation
4.4. Ultra High-Performance Liquid Chromatography (UHPLC)-Mass Spectrometry
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tuso, P. Prediabetes and Lifestyle Modification: Time to Prevent a Preventable Disease. Perm. J. 2014, 18, 88–93. [Google Scholar] [CrossRef] [PubMed]
- Bansal, N. Prediabetes diagnosis and treatment: A review. World J. Diabetes 2015, 6, 296–303. [Google Scholar] [CrossRef] [PubMed]
- Kristine, F.; Adam, H.; Thomas, P.J.S. Heterogeneity of Pre-diabetes and Type 2 Diabetes: Implications for Prediction, Prevention and Treatment Responsiveness. Curr. Diabetes Rev. 2016, 12, 30–41. [Google Scholar]
- Chatterjee, S.; Khunti, K.; Davies, M.J. Type 2 diabetes. Lancet 2017, 389, 2239–2251. [Google Scholar] [CrossRef]
- Tillin, T.; Hughes, A.D.; Wang, Q.; Würtz, P.; Ala-Korpela, M.; Sattar, N.; Forouhi, N.G.; Godsland, I.F.; Eastwood, S.V.; McKeigue, P.M.; et al. Diabetes risk and amino acid profiles: Cross-sectional and prospective analyses of ethnicity, amino acids and diabetes in a South Asian and European cohort from the SABRE (Southall And Brent REvisited) Study. Diabetologia 2015, 58, 968–979. [Google Scholar] [CrossRef]
- Würtz, P.; Soininen, P.; Kangas, A.J.; Rönnemaa, T.; Lehtimäki, T.; Kähönen, M.; Viikari, J.S.; Raitakari, O.T.; Ala-Korpela, M. Branched-Chain and Aromatic Amino Acids Are Predictors of Insulin Resistance in Young Adults. Diabetes Care 2013, 36, 648–655. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, T.J.; Larson, M.G.; Vasan, R.S.; Cheng, S.; Rhee, E.P.; McCabe, E.; Lewis, G.D.; Fox, C.S.; Jacques, P.F.; Fernandez, C. Metabolite profiles and the risk of developing diabetes. Nat. Med. 2011, 17, 448. [Google Scholar] [CrossRef]
- Stančáková, A.; Civelek, M.; Saleem, N.K.; Soininen, P.; Kangas, A.J.; Cederberg, H.; Paananen, J.; Pihlajamäki, J.; Bonnycastle, L.L.; Morken, M.A.; et al. Hyperglycemia and a Common Variant of GCKR Are Associated With the Levels of Eight Amino Acids in 9369 Finnish Men. Diabetes 2012, 61, 1895–1902. [Google Scholar] [CrossRef]
- Suvitaival, T.; Mantere, O.; Kieseppä, T.; Mattila, I.; Pöhö, P.; Hyötyläinen, T.; Suvisaari, J.; Orešič, M. Serum metabolite profile associates with the development of metabolic co-morbidities in first-episode psychosis. Transl. Psychiatry 2016, 6, e951. [Google Scholar] [CrossRef]
- Floegel, A.; Stefan, N.; Yu, Z.; Mühlenbruch, K.; Drogan, D.; Joost, H.G.; Fritsche, A.; Häring, H.U; de Hrabě Angelis, M.; Peters, A. Identification of Serum Metabolites Associated With Risk of Type 2 Diabetes Using a Targeted Metabolomic Approach. Diabetes 2013, 62, 639–648. [Google Scholar] [CrossRef] [Green Version]
- Welsh, P.; Rankin, N.; Li, Q.; Mark, P.B.; Würtz, P.; Ala-Korpela, M.; Marre, M.; Poulter, N.; Hamet, P.; Chalmers, J. Circulating amino acids and the risk of macrovascular, microvascular and mortality outcomes in individuals with type 2 diabetes: Results from the ADVANCE trial. Diabetologia 2018, 61, 1581–1591. [Google Scholar] [CrossRef] [PubMed]
- Tavares, G.; Venturini, G.; Padilha, K.; Zatz, R.; Pereira, A.C.; Thadhani, R.; Rhee, E.P.; Titan, S.M.O. 1,5-Anhydroglucitol predicts CKD progression in macroalbuminuric diabetic kidney disease: Results from non-targeted metabolomics. Metabolomics 2018, 14, 39. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Zheng, Y.; Nettleton, J.A.; Alexander, D.; Coresh, J.; Boerwinkle, E. Serum metabolomic profiling and incident CKD among African Americans. Clin. J. Am. Soc. Nephrol. 2014, 9, 1410–1417. [Google Scholar] [CrossRef] [PubMed]
- Rhee, S.Y.; Jung, E.S.; Park, H.M.; Jeong, S.J.; Kim, K.; Chon, S.; Yu, S.Y.; Woo, J.T.; Lee, C.H. Plasma glutamine and glutamic acid are potential biomarkers for predicting diabetic retinopathy. Metabolomics 2018, 14, 89. [Google Scholar] [CrossRef] [PubMed]
- Lajer, M.; Tarnow, L.; Jorsal, A.; Teerlink, T.; Parving, H.H.; Rossing, P. Plasma Concentration of Asymmetric Dimethylarginine (ADMA) Predicts Cardiovascular Morbidity and Mortality in Type 1 Diabetic Patients With Diabetic Nephropathy. Diabetes Care 2008, 31, 747–752. [Google Scholar] [CrossRef] [Green Version]
- Zobel, E.H.; von Scholten, B.J.; Reinhard, H.; Persson, F.; Teerlink, T.; Hansen, T.W.; Parving, H.H.; Jacobsen, P.K.; Rossing, P. Symmetric and asymmetric dimethylarginine as risk markers of cardiovascular disease, all-cause mortality and deterioration in kidney function in persons with type 2 diabetes and microalbuminuria. Cardiovasc. Diabetol. 2017, 16, 88. [Google Scholar] [CrossRef]
- Yousri, N.A.; Mook-Kanamori, D.O.; Selim, M.M.; Takiddin, A.H.; Al-Homsi, H.; Al-Mahmoud, K.A.; Karoly, E.D.; Krumsiek, J.; Do, K.T.; Neumaier, U. A systems view of type 2 diabetes-associated metabolic perturbations in saliva, blood and urine at different timescales of glycaemic control. Diabetologia 2015, 58, 1855–1867. [Google Scholar] [CrossRef] [Green Version]
- Gall, W.E.; Beebe, K.; Lawton, K.A.; Adam, K.P.; Mitchell, M.W.; Nakhle, P.J.; Ryals, J.A.; Milburn, M.V.; Nannipieri, M.; Camastra, S. α-Hydroxybutyrate Is an Early Biomarker of Insulin Resistance and Glucose Intolerance in a Nondiabetic Population. PLoS ONE 2010, 5, e10883. [Google Scholar] [CrossRef]
- Suvitaival, T.; Bondia-Pons, I.; Yetukuri, L.; Pöhö, P.; Nolan, J.J.; Hyötyläinen, T.; Kuusisto, J.; Orešič, M. Lipidome as a predictive tool in progression to type 2 diabetes in Finnish men. Metabolism 2018, 78, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Wang-Sattler, R.; Yu, Z.; Herder, C.; Messias, A.C.; Floegel, A.; He, Y.; Heim, K.; Campillos, M.; Holzapfel, C.; Thorand, B. Novel biomarkers for pre-diabetes identified by metabolomics. Mol. Syst. Biol. 2012, 8, 615. [Google Scholar] [CrossRef]
- Carter, T.C.; Rein, D.; Padberg, I.; Peter, E.; Rennefahrt, U.; David, D.E.; McManus, V.; Stefanski, E.; Martin, S.; Schatz, P. Validation of a metabolite panel for early diagnosis of type 2 diabetes. Metabolism 2016, 65, 1399–1408. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haeusler, R.A.; Astiarraga, B.; Camastra, S.; Accili, D.; Ferrannini, E. Human Insulin Resistance Is Associated With Increased Plasma Levels of 12α-Hydroxylated Bile Acids. Diabetes 2013, 62, 4184–4191. [Google Scholar] [CrossRef] [PubMed]
- Sone, H.; Shimano, H.; Ebinuma, H.; Takahashi, A.; Yano, Y.; Iida, K.T.; Suzuki, H.; Toyoshima, H.; Kawakami, Y.; Okuda, Y.; et al. Physiological changes in circulating mannose levels in normal, glucose-intolerant, and diabetic subjects. Metabolism 2003, 52, 1019–1027. [Google Scholar] [CrossRef]
- Wang, T.J.; Ngo, D.; Psychogios, N.; Dejam, A.; Larson, M.G.; Vasan, R.S.; Ghorbani, A.; O’Sullivan, J.; Cheng, S.; Rhee, E.P.; et al. 2-Aminoadipic acid is a biomarker for diabetes risk. J. Clin. Investig. 2013, 123, 4309–4317. [Google Scholar] [CrossRef] [PubMed]
- Klein, M.S.; Shearer, J. Metabolomics and Type 2 Diabetes: Translating Basic Research into Clinical Application. J. Diabetes Res. 2016, 2016, 10. [Google Scholar] [CrossRef] [PubMed]
- Barrios, C.; Beaumont, M.; Pallister, T.; Villar, J.; Goodrich, J.K.; Clark, A.; Pascual, J.; Ley, R.E.; Spector, T.D.; Bell, J.T.; et al. Gut-Microbiota-Metabolite Axis in Early Renal Function Decline. PLoS ONE 2015, 10, e0134311. [Google Scholar] [CrossRef]
- Kim, S.H.; Lee, K.A.; Jin, H.Y.; Baek, H.S.; Park, T.S. The Relationship between Anemia and the Initiation of Dialysis in Patients with Type 2 Diabetic Nephropathy. Diabetes Metab. J. 2015, 39, 240–246. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Zhang, S.; Wang, G. Metabolomic biomarkers in diabetic kidney diseases—A systematic review. J. Diabetes Complicat. 2015, 29, 1345–1351. [Google Scholar] [CrossRef]
- Percival, B.C.; Grootveld, M.; Gibson, M.; Osman, Y.; Molinari, M.; Jafari, F.; Sahota, T.; Martin, M.; Casanova, F.; Mather, M.L.; et al. Low-Field, Benchtop NMR Spectroscopy as a Potential Tool for Point-of-Care Diagnostics of Metabolic Conditions: Validation, Protocols and Computational Models. High Throughput. 2018, 8, 2. [Google Scholar] [CrossRef]
- Theilade, S.; Lajer, M.; Persson, F.; Joergensen, C.; Rossing, P. Arterial Stiffness Is Associated With Cardiovascular, Renal, Retinal, and Autonomic Disease in Type 1 Diabetes. Diabetes Care 2013, 36, 715–721. [Google Scholar] [CrossRef] [Green Version]
- Du, M.-R.; Yan, L.; Li, N.S.; Wang, Y.J.; Zhou, T.; Jiang, J.L. Asymmetric dimethylarginine contributes to retinal neovascularization of diabetic retinopathy through EphrinB2 pathway. Vasc. Pharmacol. 2018, 108, 46–56. [Google Scholar] [CrossRef] [PubMed]
- Larsen, M.B.; Henriksen, J.E.; Grauslund, J.; Peto, T. Prevalence and risk factors for diabetic retinopathy in 17 152 patients from the island of Funen, Denmark. Acta Ophthalmol. 2017, 95, 778–786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arneth, B.; Arneth, R.; Shams, M. Metabolomics of Type 1 and Type 2 Diabetes. Int. J. Mol. Sci. 2019, 20, 2467. [Google Scholar] [CrossRef] [PubMed]
- Campion, C.G.; Sanchez-Ferras, O.; Batchu, S.N. Potential Role of Serum and Urinary Biomarkers in Diagnosis and Prognosis of Diabetic Nephropathy. Can. J. Kidney Health Dis. 2017, 4. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Feunang, Y.D.; Marcu, A.; Guo, A.C.; Liang, K.; Vázquez-Fresno, R.; Sajed, T.; Johnson, D.; Li, C.; Karu, N.; et al. HMDB 4.0: The human metabolome database for 2018. Nucleic Acids Res. 2018, 46, D608–D617. [Google Scholar] [CrossRef]
- Theilade, S.; Hansen, T.W.; Goetze, J.P.; Rossing, P. Increased Plasma Concentrations of Midregional Proatrial Natriuretic Peptide Is Associated With Risk of Cardiorenal Dysfunction in Type 1 Diabetes. Am. J. Hypertens. 2015, 28, 772–779. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to MultipleTesting. J. R. Stat. Soc. Ser. B 1995, 57. [Google Scholar]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer-Verlag: Berlin/Heidelberg, Germany, 2016. [Google Scholar]
- Leung, K.S.-Y.; Fong, B.M.-W.J.A.; Chemistry, B. LC–MS/MS in the routine clinical laboratory: Has its time come? Anal. Bioanal. Chem. 2014, 406, 2289–2301. [Google Scholar] [CrossRef]
- Hashemipour, S.; Charkhchian, M.; Javadi, A.; Afaghi, A.; Hajiaghamohamadi, A.A.; Bastani, A.; Hajmanoochehri, F.; Ziaee, A. Urinary total protein as the predictor of albuminuria in diabetic patients. Int. J. Endocrinol. Metab. 2012, 10, 523–526. [Google Scholar] [CrossRef]
- Kar, D.; Gillies, C.; Nath, M.; Khunti, K.; Davies, M.J.; Seidu, S. Association of smoking and cardiometabolic parameters with albuminuria in people with type 2 diabetes mellitus: A systematic review and meta-analysis. Acta Diabetol. 2019, 56, 839–850. [Google Scholar] [CrossRef] [PubMed]
- Song, K.-H.; Jeong, J.S.; Kim, M.K.; Kwon, H.S.; Baek, K.H.; Ko, S.H.; Ahn, Y.B. Discordance in risk factors for the progression of diabetic retinopathy and diabetic nephropathy in patients with type 2 diabetes mellitus. J. Diabetes Investig. 2019, 10, 745–752. [Google Scholar] [CrossRef] [PubMed]
- O’Seaghdha, C.M.; Hwang, S.J.; Upadhyay, A.; Meigs, J.B.; Fox, C.S. Predictors of Incident Albuminuria in the Framingham Offspring Cohort. Am. J. Kidney Dis. 2010, 56, 852–860. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chuang, W.-H.; Arundhathi, A.; Lu, C.; Chen, Ch.; Wu, Wa.; Susanto, H.; Purnomo, J.D.T.; Wang, C. Altered plasma acylcarnitine and amino acid profiles in type 2 diabetic kidney disease. Metabolomics 2016, 12, 108. [Google Scholar] [CrossRef]
- Guasch-Ferre, M.; Hruby, A.; Toledo, E.; Clish, C.B.; Martínez-González, M.A.; Salas-Salvadó, J.; Hu, F.B. Metabolomics in Prediabetes and Diabetes: A Systematic Review and Meta-analysis. Diabetes Care 2016, 39, 14. [Google Scholar] [CrossRef] [PubMed]
- Abbiss, H.; Maker, G.L.; Trengove, R.D. Metabolomics Approaches for the Diagnosis and Understanding of Kidney Diseases. Metabolites 2019, 9, 34. [Google Scholar] [CrossRef]
- Fliser, D.; Kronenberg, F.; Kielstein, J.T.; Morath, C.; Bode-Böger, S.M.; Haller, H.; Ritz, E. Asymmetric Dimethylarginine and Progression of Chronic Kidney Disease: The Mild to Moderate Kidney Disease Study. J. Am. Soc. Nephrol. 2005, 16, 2456–2461. [Google Scholar] [CrossRef] [Green Version]
- Prawitt, J.; Caron, S.; Staels, B.J.C.D.R. Bile Acid Metabolism and the Pathogenesis of Type 2 Diabetes. Curr. Diab. Rep. 2011, 11, 160. [Google Scholar] [CrossRef]
- MacLean, B.; Tomazela, D.M.; Shulman, N.; Chambers, M.; Finney, G.L.; Frewen, B.; Kern, R.; Tabb, D.L.; Liebler, D.C.; MacCoss, M.J. Skyline: An open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 2010, 26, 966–968. [Google Scholar] [CrossRef]
- RC Team. R: A Language and Environment for Statistical Computing, R Foundation for Statistical Computing. 2017. Available online: https://www.R-project.org/ (accessed on 13 September 2019).

| Compound | Abbreviation | Group | Vendor | Solvent, Stock Solution |
|---|---|---|---|---|
| L-Glutamine | Gln | Amino acids + related metabolites | Sigma-Aldrich | H2O |
| Glycine | Gly | 0.1 M HCl | ||
| L-Alanine | Ala | |||
| L-Leucine | Leu | |||
| L-Isoleucine | Ile | |||
| L-Phenylalanine | Phe | |||
| L-Tryptophan | Trp | |||
| L-Tyrosine | Tyr | |||
| L-Glutamic Acid | Glu | |||
| L-Citrulline | Cit | |||
| L-Homocitrulline | HCit | SCB | ||
| Asymmetric dimethylarginine | ADMA | |||
| Symmetric dimethylarginine | SDMA | |||
| DL-2-Aminoadipic Acid | AADA | Sigma-Aldrich | ||
| L-Kynurenine | Kynu | |||
| Taurine | Taurine | |||
| Deoxycholic Acid | DCA | Bile acids | Sigma-Aldrich | MeOH |
| Glycochenodeoxycholic Acid | GCDCA | |||
| Glycodeoxycholic Acid | GDCA | |||
| Glycocholic Acid | GCA | |||
| Taurodeoxycholic Acid | TDCA | |||
| Taurochenodeoxycholic Acid | TCDCA | |||
| Deoxychenocholic Acid | CDCA | |||
| Cholic Acid | CA | |||
| Taurocholic Acid | TCA | |||
| Glycoursodeoxycholic Acid | GUDCA | CIL | ||
| Ursodeoxycholic Acid | UDCA | |||
| Tauroursodeoxycholic Acid | TUDCA | |||
| Creatinine | Crea | Other metabolites | Sigma-Aldrich | 10% MeOH |
| Indoxyl Sulfate | IndS | |||
| N-methyl-nicotinamide | N-MNA | SCB | ||
| Gamma-butyrobetaine | GBB | |||
| Azelaic Acid | AzelA | Small organic acids | Sigma-Aldrich | MeOH |
| L-3-hydroxybutyric Acid | β-OHB | 10% MeOH | ||
| R-2-hydroxybutyric Acid | α(R)-OHB | |||
| S-2-hydroxybutyric Acid | α(S)-OHB |
| Compound | Molecular Weight (MW) | Ion Transition | Polarity | Fragmentor Voltage (V) | Collision Energy (V) | Cell Accelerator Voltage (V) |
|---|---|---|---|---|---|---|
| AADA | 161.2 | 330.2–160.1 | Negative | 150 | 10 | 1 |
| ADMA and SDMA | 202.3 | 371.2–201.2 * | Negative | 150 | 5 | 5 |
| 371.2–156.1 | Negative | 150 | 20 | 1 | ||
| Ala | 89.1 | 258.1–88.1 | Negative | 100 | 15 | 3 |
| AzelA | 188.2 | 187.2–169 | Negative | 150 | 10 | 1 |
| 187.2–125.2 * | Negative | 150 | 15 | 1 | ||
| β-OHB | 104.1 | 103.2–59.2 | Negative | 100 | 5 | 1 |
| CA | 408.6 | 407.3–407.3 * | Negative | 250 | 0 | 1 |
| 407.3–343.3 | Negative | 250 | 35 | 3 | ||
| CDCA | 392.6 | 391.3–391.3 | Negative | 250 | 0 | 3 |
| Cit | 175.2 | 344.4–174.2 | Negative | 150 | 4 | 7 |
| Crea | 113.1 | 114.1–86.2 | Positive | 150 | 11 | 4 |
| 114.1–44.1 * | Positive | 150 | 15 | 4 | ||
| DCA | 392.6 | 391.2–345.3 * | Negative | 200 | 35 | 4 |
| 391.2–327.2 | Negative | 200 | 40 | 4 | ||
| GBB | 146.2 | 147.2–88.1 * | Positive | 100 | 16 | 1 |
| 147.2–60.2 | Positive | 100 | 13 | 1 | ||
| GCA | 465.6 | 464.3–402.1 | Negative | 250 | 40 | 4 |
| 464.3–74.1 * | Negative | 250 | 45 | 7 | ||
| GCDCA | 449.6 | 448.3–386.3 | Negative | 150 | 40 | 2 |
| GDCA | 449.6 | 448.3–402.1 | Negative | 250 | 40 | 2 |
| GCDCA and GDCA | 449.6 | 448.3–74.2 | Negative | 200 | 55 | 2 |
| Gln | 146.1 | 315.3–145.1 | Negative | 100 | 9 | 6 |
| Glu | 147.1 | 316.1–146.1 | Negative | 100 | 6 | 6 |
| Gly | 75.1 | 244.1–74.1 | Negative | 200 | 7 | 4 |
| GUDCA | 449.6 | 448.3–386 | Negative | 250 | 40 | 2 |
| 448.3–74.1 * | Negative | 250 | 45 | 2 | ||
| HCit | 189.2 | 358.3–188.1 | Negative | 200 | 10 | 1 |
| 358.3–145 * | Negative | 150 | 25 | 2 | ||
| IndS | 213.2 | 212–132 * | Negative | 100 | 15 | 2 |
| 212–80 | Negative | 100 | 20 | 2 | ||
| Kynu | 208.2 | 377–316.1 | Negative | 150 | 5 | 2 |
| 377–207 * | Negative | 150 | 5 | 5 | ||
| Leu and Ile | 131.2 | 300.2–130.2 | Negative | 100 | 10 | 1 |
| N-MNA | 136.2 | 137.1–108.1 | Positive | 100 | 15 | 2 |
| 137.1–80.2 * | Positive | 100 | 26 | 2 | ||
| Phe | 165.2 | 334.2–164 | Negative | 100 | 10 | 1 |
| Taurine | 125.2 | 294.1–124.1 * | Negative | 100 | 10 | 2 |
| 294.1–80.1 | Negative | 100 | 55 | 2 | ||
| TCA | 515.7 | 514.3–123.8 | Negative | 300 | 65 | 5 |
| 514.3–80.2 * | Negative | 300 | 95 | 1 | ||
| TDCA and TCDCA | 499.3 | 498.3–107.1 | Negative | 250 | 80 | 1 |
| 498.3–80.1 * | Negative | 300 | 90 | 1 | ||
| Trp | 204.2 | 373.2–203.1 | Negative | 150 | 7 | 2 |
| TUDCA | 499.7 | 498.3–107.1 | Negative | 300 | 65 | 5 |
| 498.3–80.1 * | Negative | 300 | 85 | 1 | ||
| Tyr | 181.2 | 350.2–180.1 | Negative | 100 | 7 | 5 |
| AADA-d3 | 164.2 | 333.2–145.2 | Negative | 100 | 20 | 2 |
| ADMA-d7 | 209.8 | 378–208.3 | Negative | 100 | 10 | 5 |
| Ala-d4 | 93.1 | 262.1–92.1 | Negative | 100 | 5 | 6 |
| α-OHB-d3 | 107.1 | 106.1–59.1 | Negative | 100 | 10 | 1 |
| AzelA-d14 | 202.3 | 201.2–137.2 | Negative | 150 | 10 | 2 |
| β-OHB-d4 | 108.1 | 107.1–59.1 | Negative | 100 | 5 | 1 |
| CA-d4 | 412.3 | 411.3–411.3 | Negative | 250 | 0 | 3 |
| CDCA-d4 and DCA-d4 | 396.6 | 395.2–395.2 | Negative | 300 | 0 | 4 |
| Cit-d4 | 179.2 | 348.1–135.1 | Negative | 100 | 25 | 2 |
| Crea-d5 | 118.2 | 119.2–49.3 | Positive | 100 | 20 | 1 |
| GBB-d9 | 154.7 | 155.2–87.3 | Positive | 100 | 15 | 6 |
| GCA-d4 | 469.6 | 468.3–74.1 | Negative | 250 | 45 | 1 |
| GCDCA-d4 and GUDCA-d4 | 453.6 | 452.3–74.1 | Negative | 250 | 40 | 1 |
| GDCA-d6 | 455.7 | 454.3–408.2 | Negative | 250 | 55 | 4 |
| Gln-d5 | 151.2 | 320.1–150.1 | Negative | 100 | 5 | 1 |
| Glu-d5 | 152.1 | 321.1–151.1 | Negative | 100 | 5 | 1 |
| Gly-13C,d2 | 78.1 | 247–77.1 | Negative | 100 | 5 | 7 |
| HCit-2H4 | 193.2 | 362.2–192.2 | Negative | 100 | 5 | 6 |
| IndS-d4 | 217.3 | 216–136.1 | Negative | 100 | 15 | 2 |
| Kynu-13C6 | 214.2 | 383.1–195.8 | Negative | 100 | 10 | 6 |
| Leu-d10 and Ile-d10 | 141.2 | 310.1–140 | Negative | 125 | 10 | 2 |
| N-MNA-d4 | 140.2 | 141.2–84.2 | Positive | 100 | 20 | 7 |
| Phe-d5 | 170.2 | 339.1–169.1 | Negative | 150 | 5 | 1 |
| Taurine-d4 | 129.2 | 298.3–128.2 | Negative | 100 | 10 | 3 |
| TCA-d4 | 519.7 | 518.3–80 | Negative | 340 | 100 | 7 |
| TCDCA-d9 | 508.3 | 507.4–80.1 | Negative | 300 | 95 | 1 |
| Trp-d8 | 212.3 | 381.2–211.2 | Negative | 100 | 10 | 5 |
| TUDCA-d4 | 503.7 | 502.3–80.1 | Negative | 300 | 100 | 1 |
| Tyr-d7 | 188.2 | 357.1–187.2 | Negative | 100 | 10 | 1 |
| UDCA-d4 | 396.6 | 395.3–395.3 | Negative | 250 | 0 | 4 |
| Compound | Linearity (R2) Range (LLOQ-ULOQ) (ng mL−1) | LOD (ng/mL−1) | %RSD_Rt, Intra-Day | %RSD_Area, Intra-Day (N = 4) | %RSD_Rt, Inter-Day | %RSD_Area, Inter-Day (N = 15) | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| 100 ng mL−1 | 1000 ng mL−1 | 10,000 ng mL−1 | 100 ng mL−1 | 1000 ng mL−1 | 10,000 ng mL−1 | |||||
| AADA | 0.984 5000–75,000 | 500 | 0.2 (N = 4) | - | - | 9.1 | 0.1 (N = 15) | - | - | 8.7 |
| ADMA and SDMA | 0.992 2500–50,000 | 500 | 0.2 (N = 8) | - | 5.6 | 0.8 | 0.2 (N = 30) | - | 8.5 | 4.2 |
| Ala | 0.996 500–50,000 | <2.5 | 0.2 (N = 8) | - | 4.5 | 3.0 | 0.1 (N = 30) | - | 9.8 | 13.6 |
| AzelA | 0.995 500–10,000 | <2.5 | 0.5 (N = 8) | - | 11.4 | 3.9 | - | - | 15.8 | 8.4 |
| β-OHB | 0.970 2500–75,000 | 75 | 0.6 (N = 4) | - | - | 20.9 | 1.2 (N = 15) | - | - | 24.5 |
| CA | 0.996 10–10,000 | 7.5 | 0.2 (N = 12) | 2.4 | 3.1 | 5.2 | 0.7 (N = 45) | 20.8 | 18.1 | 20.2 |
| CDCA | 0.999 25–2500 | 7.5 | 0.2 (N = 8) | 4.0 | 4.9 | - | 0.2 (N = 45) | 4.3 | 5.1 | 14.2 |
| Cit | 0.984 500–10,000 | 250 | 0.2 (N = 8) | - | 7.7 | 6.7 | 0.2 (N = 30) | - | 9.1 | 8.3 |
| Crea | 0.973 250–7500 | 25 | 0.8 (N = 4) | - | 17.8 | - | 0.0 (N = 15) | - | 3.5 | - |
| DCA | 0.996 5–2500 | 2.5 | 0.2 (N = 8) | 5.8 | 6.1 | - | 0.3 (N = 30) | 4.3 | 8.3 | - |
| GBB | 0.974 250–10,000 | 50 | 0.5 (N = 8) | - | 18.7 | 15.9 | 1.5 (N = 30) | - | 27.3 | 28.5 |
| GCA | 0.997 50–25,000 | 25 | 0.1 (N = 12) | 4.6 | 4.2 | 4.2 | 0.4 (N = 45) | 6.8 | 5.1 | 7.3 |
| GCDCA and GDCA | 0.997 25–2500 | <2.5 | 0.2 (N = 8) | 1.9 | 4.2 | - | 0.5 (N = 30) | 16.4 | 16.1 | - |
| Gln | 0.987 750–50,000 | 5 | 0.1 (N = 8) | - | 5.0 | 7.7 | 0.5 (N = 30) | - | 10.5 | 11.5 |
| Glu | 0.990 750–75,000 | 500 | 0.2 (N = 8) | - | 13.9 | 10.7 | 0.3 (N = 30) | - | 10.9 | 5.2 |
| Gly | 0.993 7500–75,000 | 1000 | 0.03 (N = 4) | - | - | 16.2 | 0.6 (N = 15) | - | - | 19.6 |
| GUDCA | 0.994 75–10,000 | 25 | 0.1 (N = 12) | 5.0 | 9.0 | 10.6 | 0.3 (N = 45) | 13.1 | 10.9 | 6.2 |
| HCit | 0.995 500–25,000 | 250 | 0.2 (N = 8) | - | 8.3 | 2.6 | 0.5 (N = 30) | - | 11.1 | 16.4 |
| IndS | 0.986 5000–75,000 | 750 | 0.3 (N = 4) | - | - | 11.3 | 0.3 (N = 15) | - | - | 15.4 |
| Kynu | 0.993 500–75,000 | 250 | 0.2 (N = 8) | - | 11.2 | 7.4 | 0.4 (N = 30) | - | 7.7 | 4.4 |
| Leu and Ile | 0.997 25–75,000 | <2.5 | 0.4 (N = 12) | 4.6 | 4.3 | 1.5 | 0.5 (N = 45) | 13.0 | 14.0 | 5.7 |
| N-MNA | 0.998 25–10,000 | <2.5 | 0.5 (N = 12) | 1.6 | 6.4 | 3.7 | 1.0 (N = 45) | 20.1 | 18.5 | 6.5 |
| Phe | 0.995 250–25,000 | <2.5 | 0.4 (N = 0.4) | - | 5.9 | 6.6 | 0.4 (N = 30) | - | 10.1 | 4.6 |
| Taurine | 0.994 250–25,000 | 10 | 0.2 (N = 8) | - | 8.3 | 5.7 | 0.5 (N = 30) | - | 8.4 | 8.7 |
| TCA | 0.983 2500–25,000 | <2.5 | 0.1 (N = 4) | - | - | 4.5 | 0.3 (N = 15) | - | - | 15.5 |
| TDCA and TCDCA | 0.984 1000–25,000 | 10 | 0.7 (N = 8) | - | 0.4 | 5.7 | 0.7 (N = 30) | - | 2.6 | 4.2 |
| Trp | 0.996 25–25,000 | 25 | 0.4 (N = 12) | 9.0 | 2.9 | 4.7 | 0.5 (N = 45) | 18.8 | 5.4 | 5.3 |
| TUDCA | 0.990 250–10,000 | 10 | 0.1 (N = 8) | - | 5.5 | 4.5 | 0.7 (N = 30) | - | 1.8 | 3.0 |
| Tyr | 0.992 50–75,000 | 25 | 0.2 (N = 12) | 10.3 | 9.1 | 4.4 | 0.3 (N = 45) | 5.7 | 8.4 | 3.4 |
| UDCA | 0.991 50–50,000 | 25 | 0.2 (N = 12) | 1.5 | 3.5 | 3.3 | 0.2 (N = 45) | 3.5 | 10.3 | 5.6 |
| Metabolite Name | Normo-Albuminuria, Mean c (Standard Deviation) | Macro-Albuminuria, Mean c (Standard Deviation) | p Value | adj. p Value |
|---|---|---|---|---|
| Glycochenodeoxycholic Acid and Glycodeoxycholic Acid | 4.33 (11.74) | 2.10 (6.58) | 0.00012 | 0.0021 |
| L-Kynurenine | 383.23 (249.28) | 309.03 (86.53) | 0.00043 | 0.0034 |
| Tyrosine | 6185.75 (1865.87) | 7012.51 (2076.69) | 0.00057 | 0.0034 |
| Tryptophan | 5913.04 (1705.38) | 6388.34 (1346.28) | 0.031 | 0.14 |
| Asymmetric dimethylarginine and Symmetric Dimethylarginine | 165.73 (51.07) | 153.35 (18.60) | 0.26 | 0.57 |
| Leucine and Isoleucine | 6393.48 (3159.65) | 7303.02 (3656.17) | 0.28 | 0.57 |
| Chenodeoxycholic Acid | 1101.07 (7.10) | 1099.58 (6.38) | 0.29 | 0.57 |
| Glycine | 9696.30 (5174.24) | 10,313.80 (3604.96) | 0.32 | 0.58 |
| Glutamine | 31,651.43 (8920.90) | 29,020.85 (6798.27) | 0.4 | 0.63 |
| L-Citrulline | 2235.88 (1160.64) | 2253.08 (852.27) | 0.42 | 0.63 |
| Alanine | 16,925.72 (4875.55) | 16,087.19 (3345.81) | 0.58 | 0.75 |
| Indoxyl Sulfate | 907.87 (493.53) | 920.80 (561.30) | 0.6 | 0.75 |
| Homocitrulline | 11.36 (25.76) | 10.21 (20.90) | 0.62 | 0.75 |
| Taurine | 4741.00 (2046.23) | 4128.35 (1424.84) | 0.77 | 0.86 |
| Phenylalanine | 9337.50 (2600.13) | 8949.64 (2062.99) | 0.86 | 0.91 |
| Glutamic Acid | 8164.60 (3588.71) | 9304.01 (7562.67) | 0.93 | 0.93 |
| Internal Standard | Abbreviation | Group | Vendor | Solvent, Stock Solution | Concentration in ISTD MIX (ng mL−1) |
|---|---|---|---|---|---|
| d5-Glutamine | d5-Gln | Amino acids + related metabolites | CIL | H2O | 30,000 |
| d10-L-Leucine | d10-Leu | CDN | 0.1 M HCl | 5000 | |
| 2H4-L-Homocitrulline | 2H4-HCit | Alsachim | |||
| Glycine-1-13C,2,2-d2 | 13C, d2-Gly | Sigma-Aldrich | |||
| d4-DL-Alanine | d4-Ala | ||||
| d5-L-Glutamic Acid | d5-Glu | ||||
| d10-Isoleucine | d10-Ile | CIL | |||
| d5-L-Phenylalanine | d5-Phe | 500 | |||
| d8-Tryptophan | d8-Trp | 5000 | |||
| d7-Tyrosine | d7-Tyr | ||||
| d4-Citrulline | d4-Cit | 500 | |||
| d3-L-2-Aminoadipic Acid | d3-AADA | 10,000 | |||
| d7-Asymmetric dimethylarginine | d7-ADMA | 5000 | |||
| 13C6-Kynurenine | 13C6-Kynu | Alsachim | 30,000 | ||
| d4-Taurine | d4-Taurine | 500 | |||
| d4-Deoxycholic Acid | d4-DCA | Bile acids | CDN | MeOH | 500 |
| d4-Glycocholic Acid | d4-GCA | 250 | |||
| d4-Deoxychenocholic Acid | d4-CDCA | 500 | |||
| d4-Glycoursodeoxycholic Acid | d4-GUDCA | 5000 | |||
| d4-Cholic Acid | d4-CA | 500 | |||
| d4-Ursodeoxycholic Acid | d4-UDCA | 250 | |||
| d4-Glychochenodeoxycholic Acid | d4-GCDCA | CIL | 5000 | ||
| d6-Glycodeoxycholic Acid | d6-GDCA | 30,000 | |||
| d9-Taurochenodeoxycholic Acid | d9-TCDCA | 500 | |||
| d4-Taurocholic Acid | d4-TCA | ||||
| d4-Tauroursodeoxycholic Acid | d4-TUDCA | 250 | |||
| d5-Creatinine | d5-Crea | Polar metabolites | CDN | 10% MeOH | 10,000 |
| d4-N-methyl-nicotinamide | d4-N-MNA | 250 | |||
| d9-Gamma-butyrobetaine | d9-GBB | 500 | |||
| d4-Indoxyl Sulfate | d4-IndS | Sigma-Aldrich | 5000 | ||
| d14-Azelaic Acid | d14-AzelA | Small organic acids | CDN | MeOH | 5000 |
| d4-3-Hydroxybutyric Acid | d4-β-OHB | 10% MeOH | 100,000 | ||
| d3-2-Hydroxybutyric Acid | d3-α-OHB |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ahonen, L.; Jäntti, S.; Suvitaival, T.; Theilade, S.; Risz, C.; Kostiainen, R.; Rossing, P.; Orešič, M.; Hyötyläinen, T. Targeted Clinical Metabolite Profiling Platform for the Stratification of Diabetic Patients. Metabolites 2019, 9, 184. https://doi.org/10.3390/metabo9090184
Ahonen L, Jäntti S, Suvitaival T, Theilade S, Risz C, Kostiainen R, Rossing P, Orešič M, Hyötyläinen T. Targeted Clinical Metabolite Profiling Platform for the Stratification of Diabetic Patients. Metabolites. 2019; 9(9):184. https://doi.org/10.3390/metabo9090184
Chicago/Turabian StyleAhonen, Linda, Sirkku Jäntti, Tommi Suvitaival, Simone Theilade, Claudia Risz, Risto Kostiainen, Peter Rossing, Matej Orešič, and Tuulia Hyötyläinen. 2019. "Targeted Clinical Metabolite Profiling Platform for the Stratification of Diabetic Patients" Metabolites 9, no. 9: 184. https://doi.org/10.3390/metabo9090184
APA StyleAhonen, L., Jäntti, S., Suvitaival, T., Theilade, S., Risz, C., Kostiainen, R., Rossing, P., Orešič, M., & Hyötyläinen, T. (2019). Targeted Clinical Metabolite Profiling Platform for the Stratification of Diabetic Patients. Metabolites, 9(9), 184. https://doi.org/10.3390/metabo9090184