Surfactant Lipidomics of Alveolar Lavage Fluid in Mice Based on Ultra-High-Performance Liquid Chromatography Coupled to Hybrid Quadrupole-Exactive Orbitrap Mass Spectrometry
Abstract
1. Introduction
2. Results
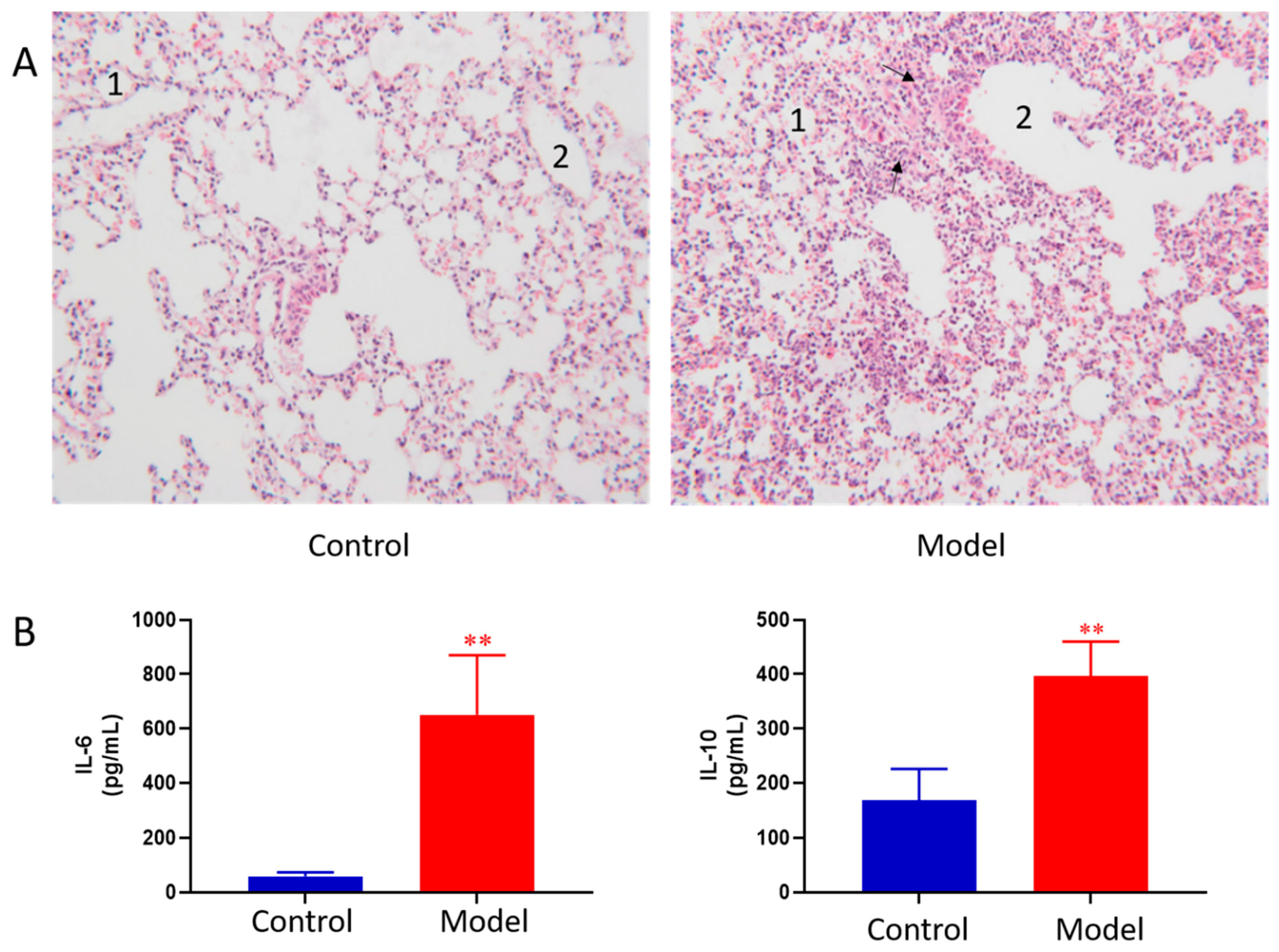
2.1. Evaluation of Mouse Model of Acute Lung Injury
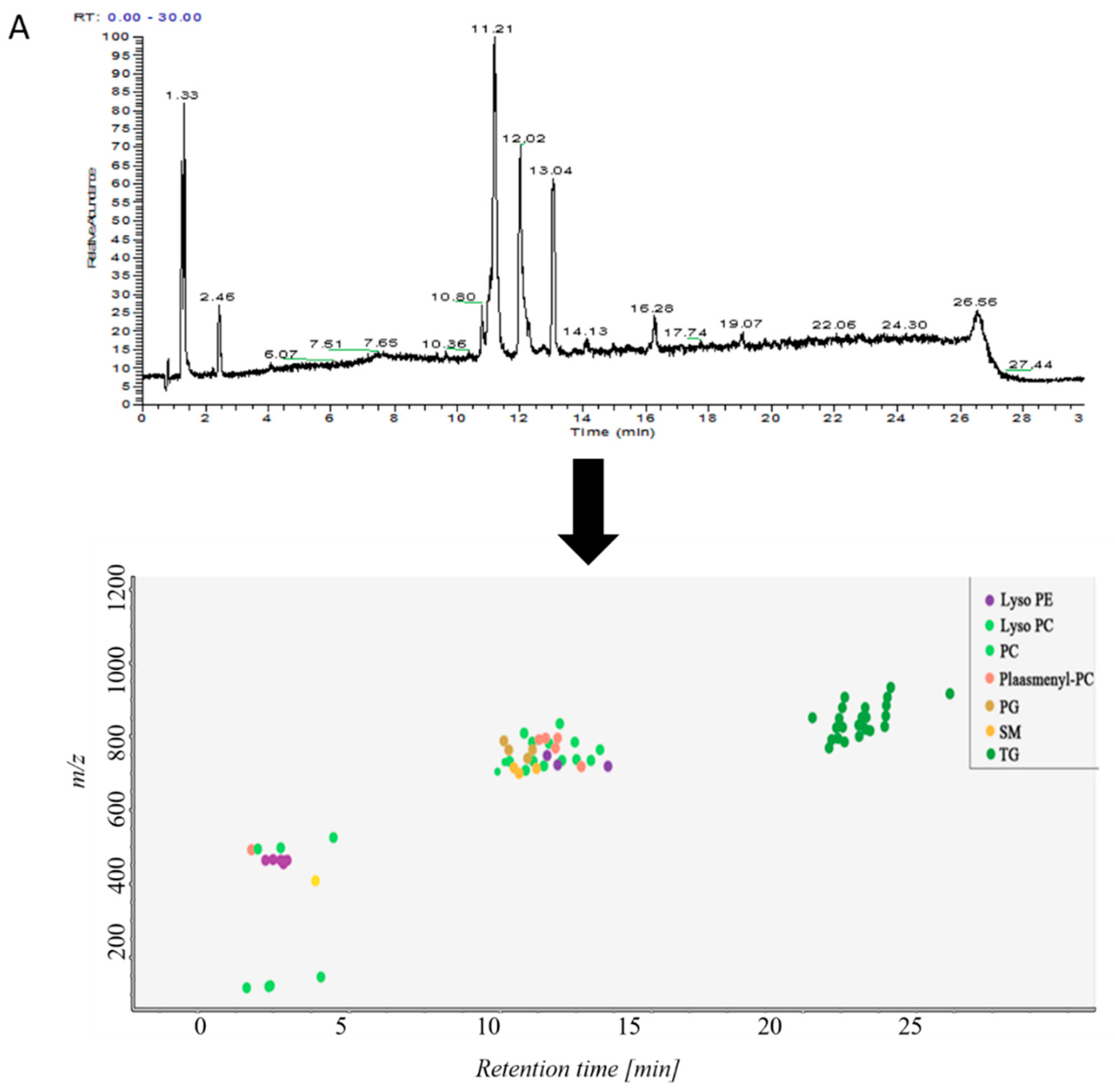
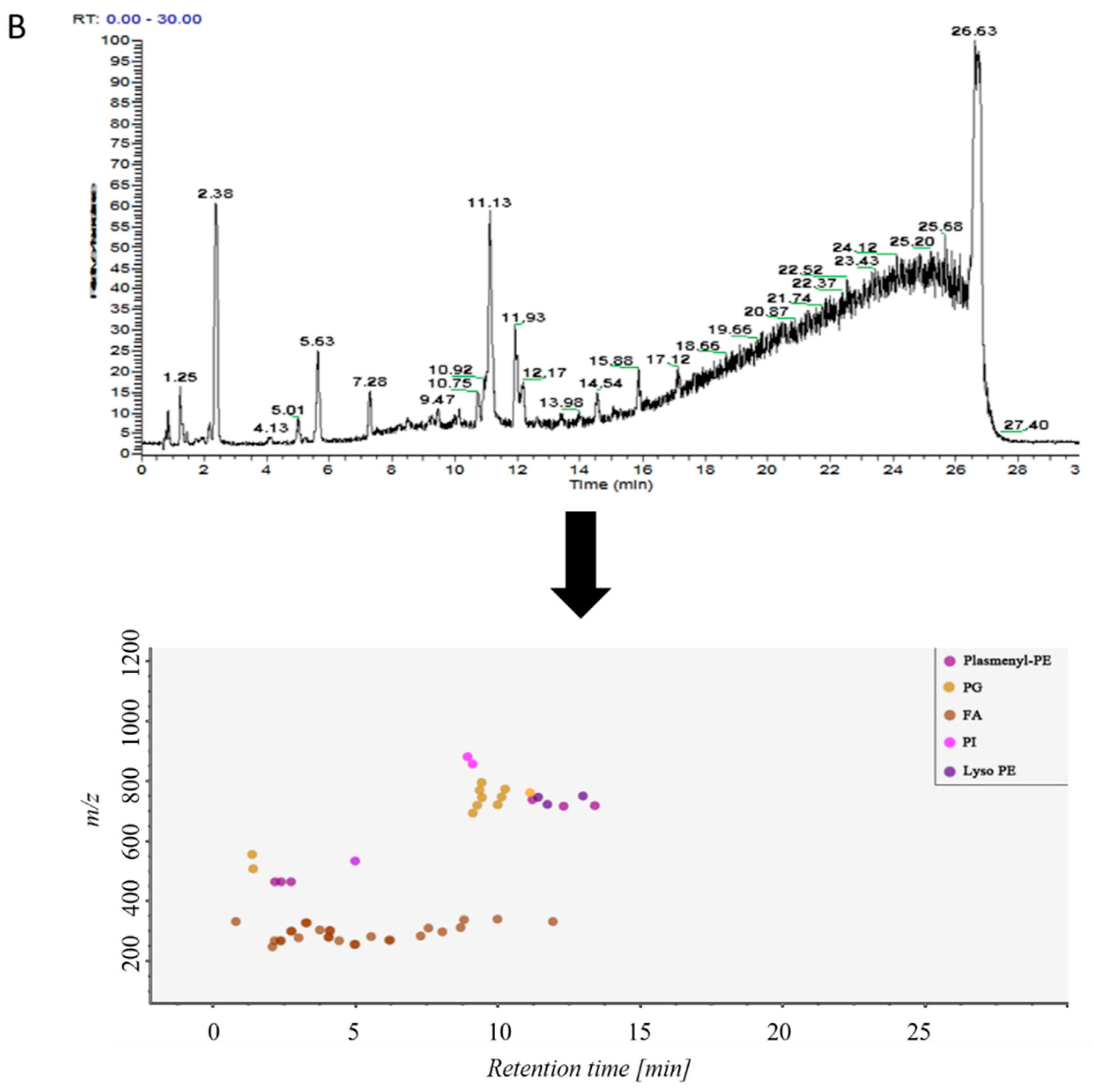
2.2. Analysis of Lipid Components in Alveolar Lavage Fluid
2.3. Liposomics of Alveolar Lavage Fluid in Mice with Acute Lung Injury
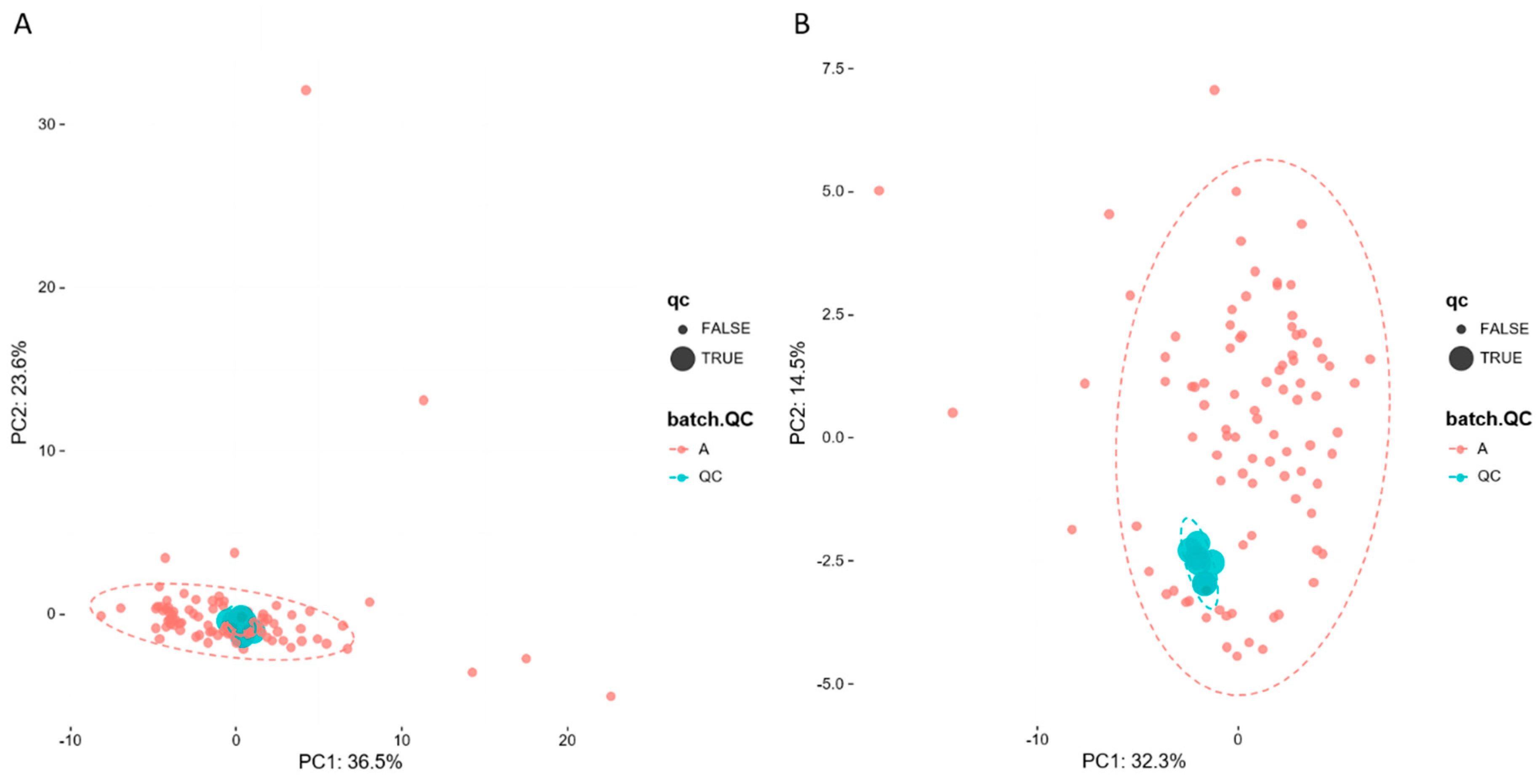
2.3.1. UHPLC-Q-Exactive Orbitrap MS Method Validation
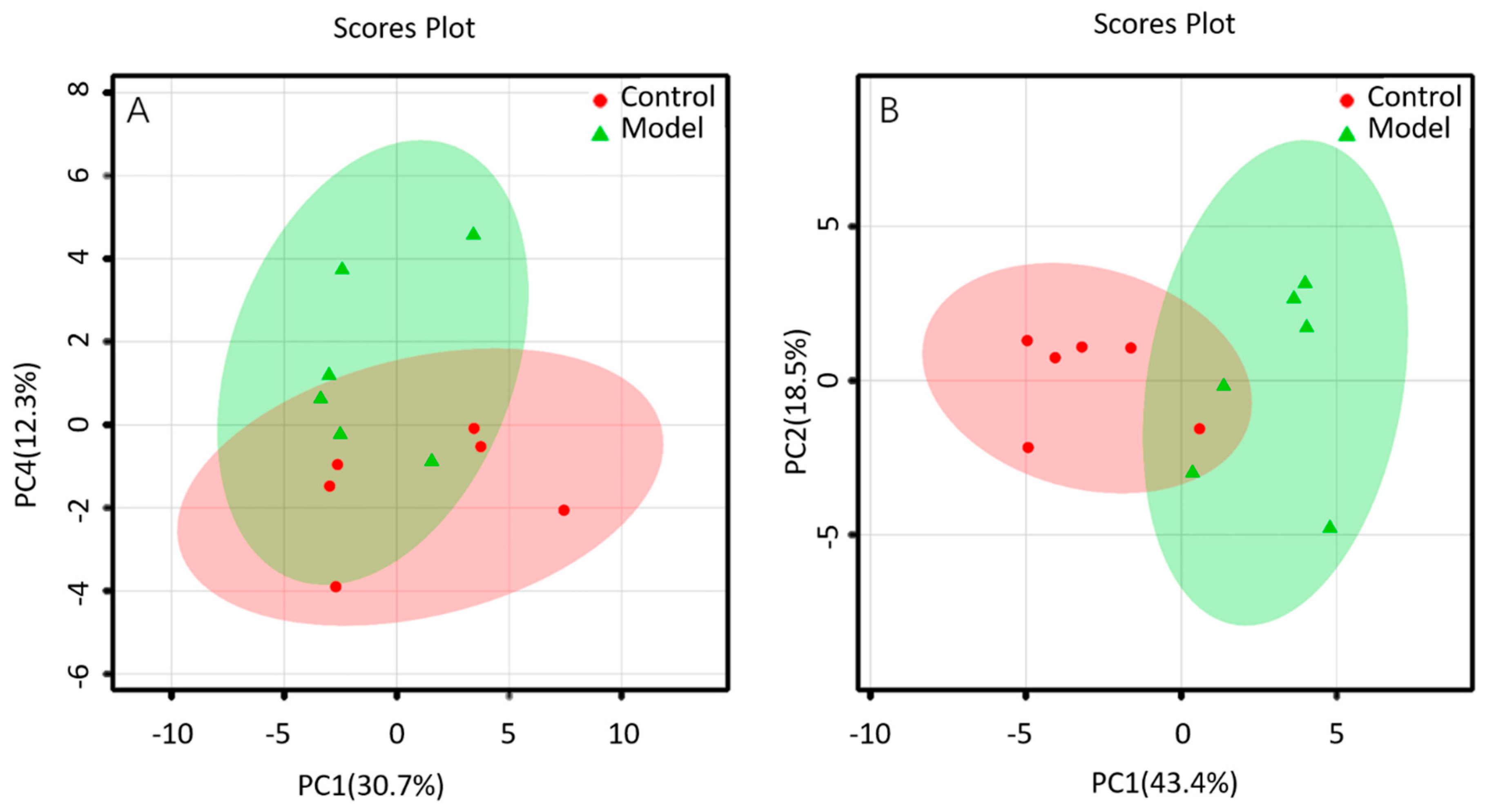
2.3.2. Non-Targeted Lipidomics Metabolic Analysis
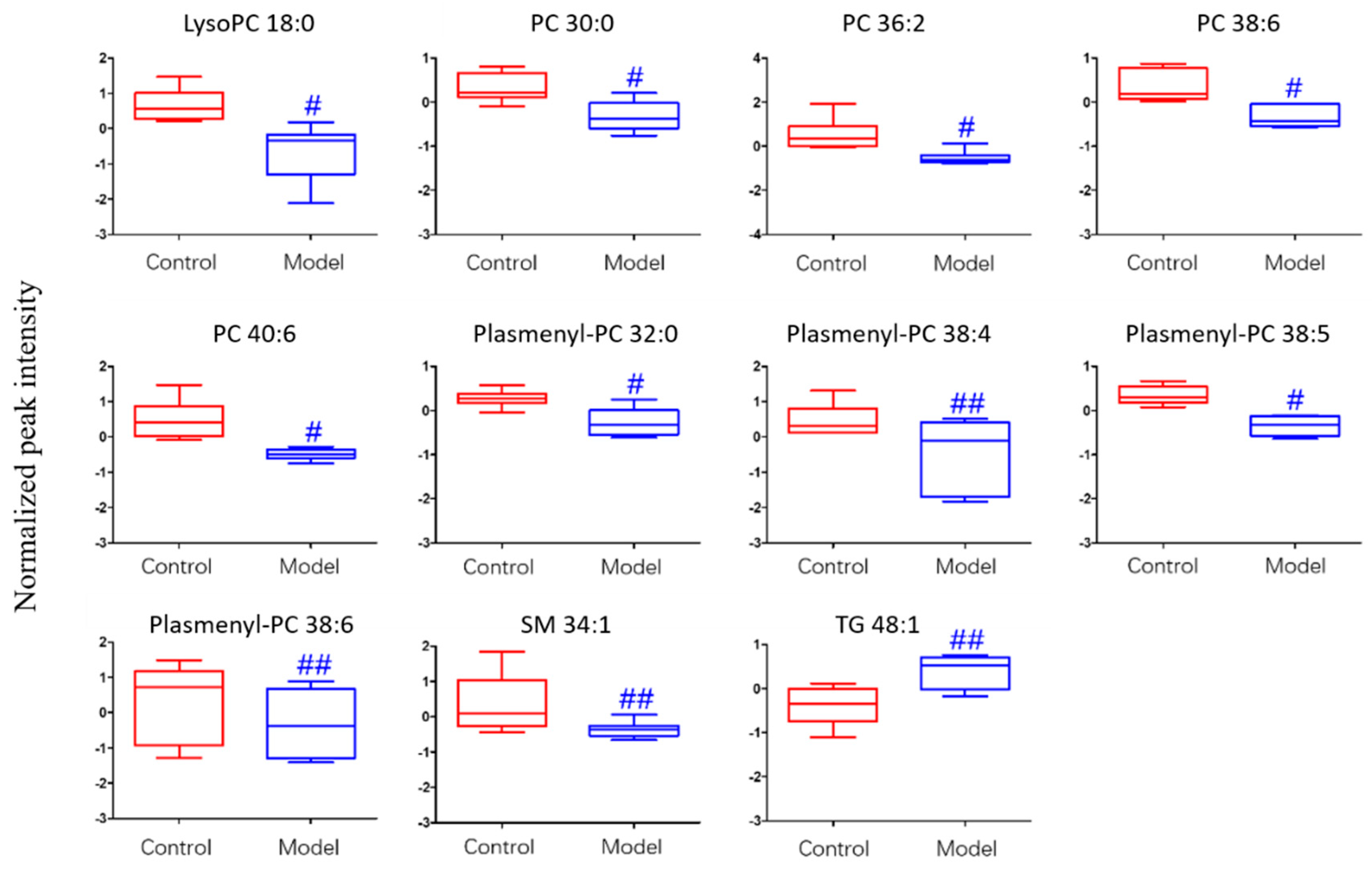
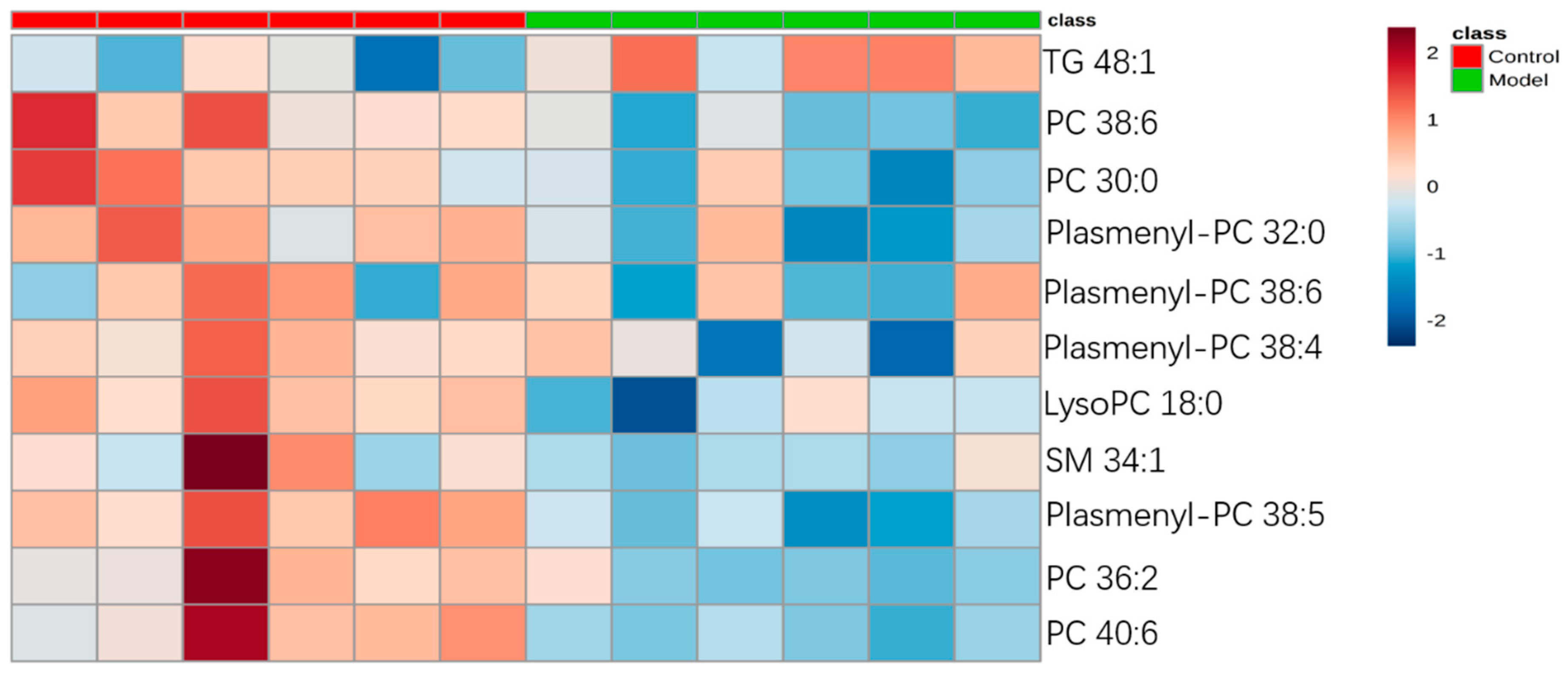
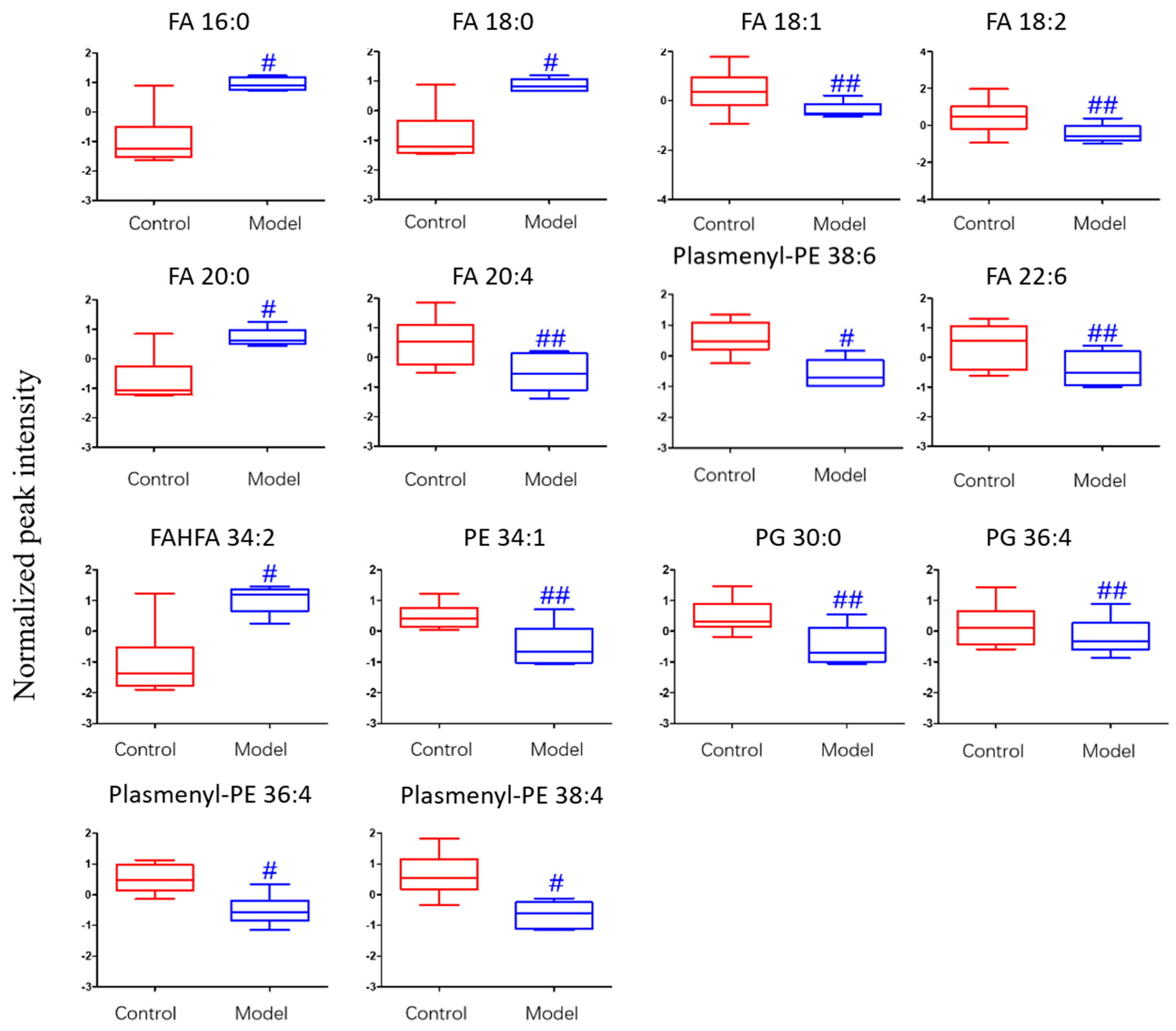
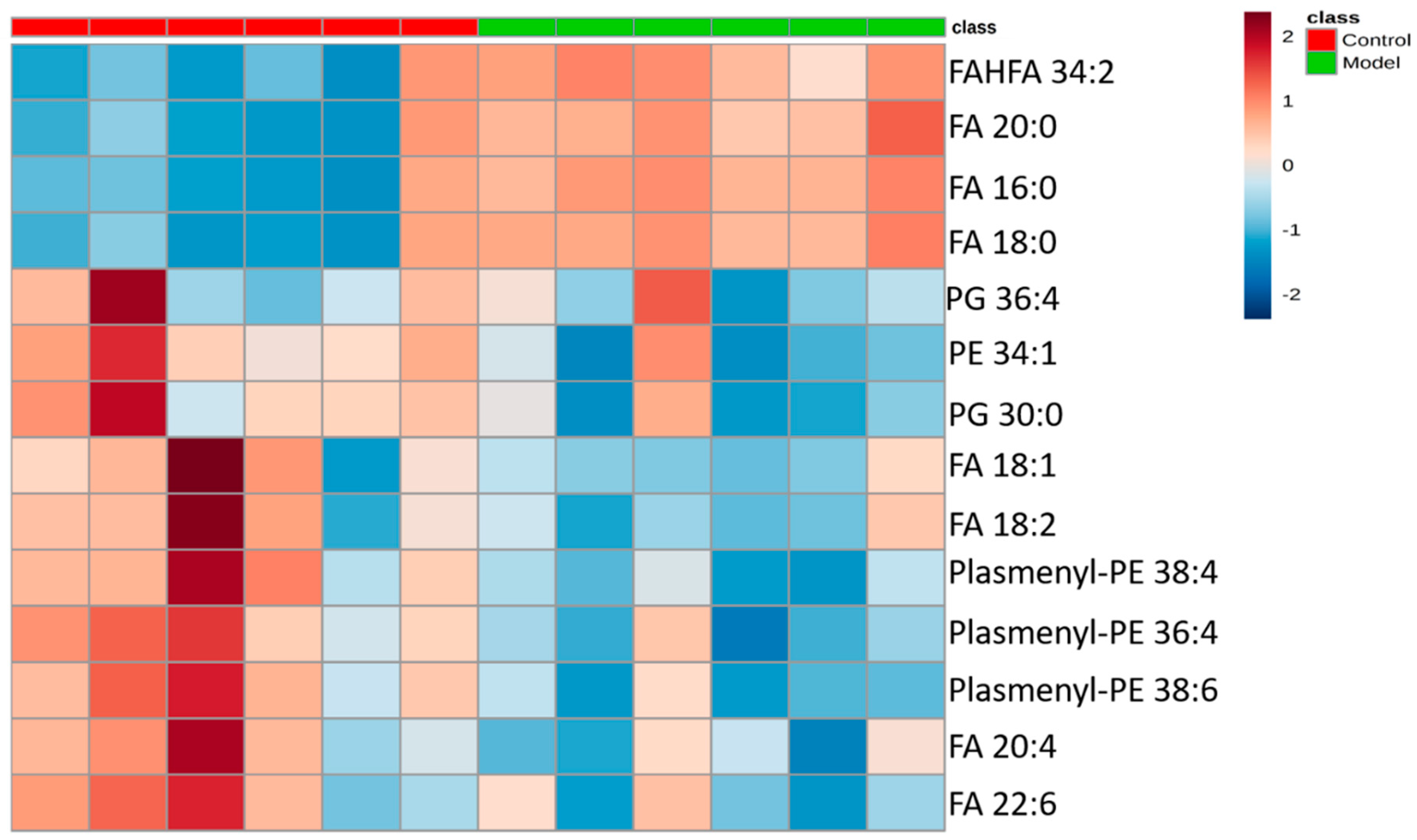
2.3.3. Analysis of Differential Lipid Metabolites
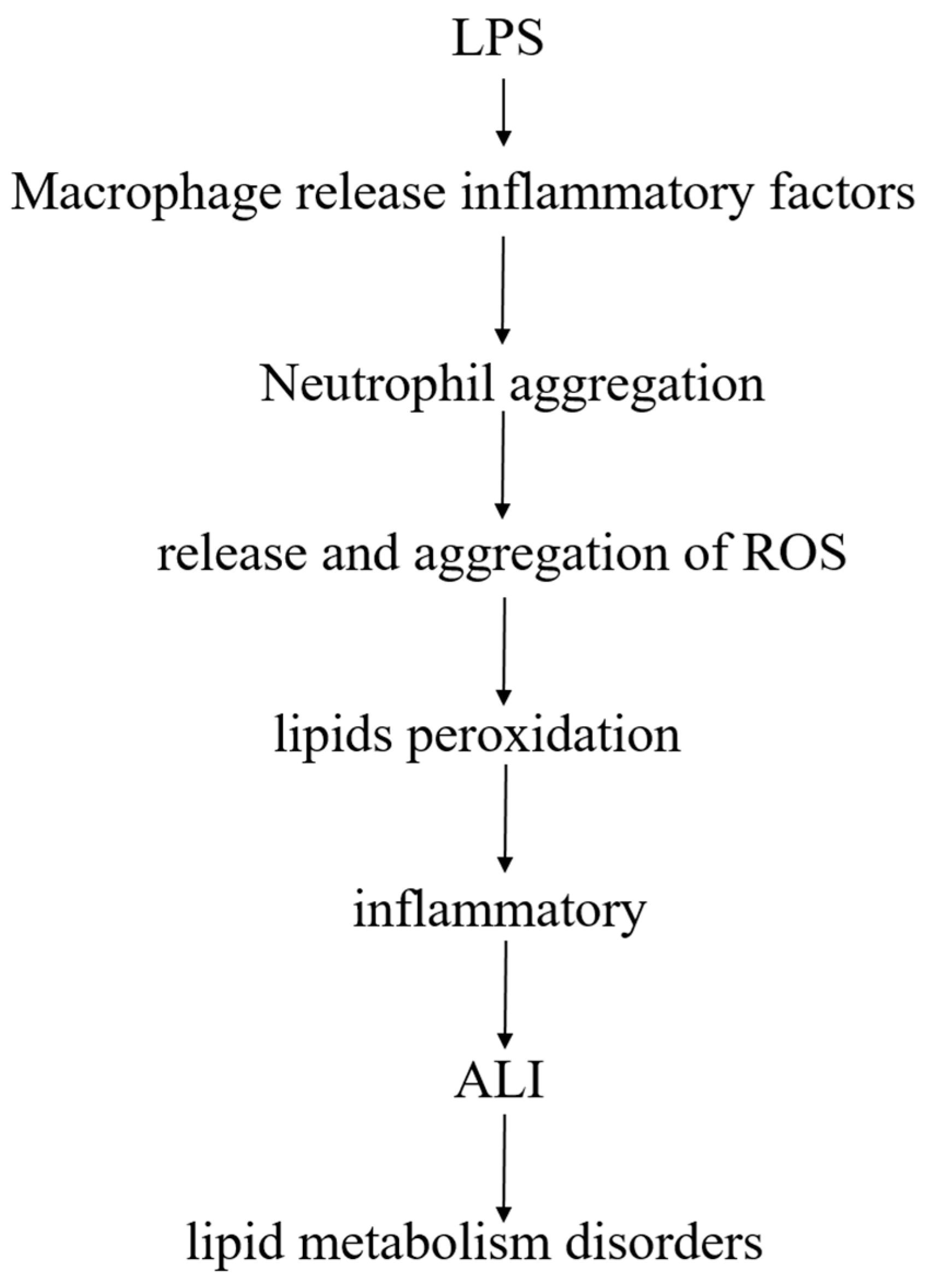
3. Discussion
4. Materials and Methods
4.1. Instruments and Chemicals
4.2. Animals
4.3. Sample Collection
4.4. Histopathology
4.5. Measurements of Inflammatory Mediators
4.6. Sample Preparation
4.7. Untargeted Lipidomic Analysis
4.8. Data Processing
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Zhao, Y.Y.; Cheng, X.L.; Lin, R.C. Chapter One–Lipidomics Applications for Discovering Biomarkers of Diseases in Clinical Chemistry. Int. Rev. Cell Mol. Biol. 2014, 313, 1–26. [Google Scholar] [CrossRef]
- Han, X. Lipidomics for studying metabolism. Nat. Rev. Endocrinol. 2016, 12, 668–679. [Google Scholar] [CrossRef]
- Jurowski, K.; Kochan, K.; Walczak, J.; Baranska, M.; Piekoszewski, W.; Buszewski, B. Analytical Techniques in Lipidomics: State of the Art. Crit. Rev. Anal. Chem. 2017, 47, 418–437. [Google Scholar] [CrossRef]
- Buas, M.F.; Gu, H.; Djukovic, D.; Zhu, J.; Drescher, C.W.; Urban, N.; Raftery, D.; Li, C.I. Identification of novel candidate plasma metabolite biomarkers for distinguishing serous ovarian carcinoma and benign serous ovarian tumors. Gynecol. Oncol. 2016, 140, 138–144. [Google Scholar] [CrossRef]
- Wang, W.; Xin, J.; Yang, X.; Lam, S.M.; Shui, G.; Wang, Y.; Huang, X. Lipid-gene regulatory network reveals coregulations of triacylglycerol with phosphatidylinositol/lysophosphatidylinositol and with hexosyl-ceramide. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2019, 1864, 168–180. [Google Scholar] [CrossRef]
- Ren, F.P.; Jiang, Y.W. Bronchoalveolar lavage fluid application value of tumor markers detection in lung cancer. Chin. J. Diagn. Pathol. 2017, 24, 774–777. [Google Scholar] [CrossRef]
- Voelker, D.R.; Numata, M. Phospholipid regulation of innate immunity and respiratory viral infection. J. Biol. Chem. 2019, 294, 4282–4289. [Google Scholar] [CrossRef] [PubMed]
- Echaide, M.; Autilio, C.; Arroyo, R.; Perez-Gil, J. Restoring pulmonary surfactant membranes and films at the respiratory surface. Biochim. Biophys. Acta Biomembr. 2017, 1859, 1725–1739. [Google Scholar] [CrossRef]
- Fessler, M.B.; Summer, R.S. Surfactant Lipids at the Host-Environment Interface. Metabolic Sensors, Suppressors, and Effectors of Inflammatory Lung Disease. Am. J. Respir. Cell Mol. Biol. 2016, 54, 624–635. [Google Scholar] [CrossRef] [PubMed]
- Walmsley, S.; Cruickshank-Quinn, C.; Quinn, K.; Zhang, X.; Petrache, I.; Bowler, R.P.; Reisdorph, R.; Reisdorph, N. A prototypic small molecule database for bronchoalveolar lavage-based metabolomics. Sci. Data 2018, 5, 180060. [Google Scholar] [CrossRef]
- Xie, L.; Yin, X.J. New progress in research on neonatal pulmonary surfactant. Chin. J. Child Health Care 2012, 20, 235–237. [Google Scholar]
- Matute-Bello, G.; Downey, G.; Moore, B.B.; Groshong, S.D.; Matthay, M.A.; Slutsky, A.S.; Kuebler, W.M.; Acute Lung Injury in Animals Study Group. An official American Thoracic Society workshop report: Features and measurements of experimental acute lung injury in animals. Am. J. Respir. Cell Mol. Biol. 2011, 44, 725–738. [Google Scholar] [CrossRef]
- Zimmermann, K.K.; Spassov, S.G.; Strosing, K.M.; Ihle, P.M.; Engelstaedter, H.; Hoetzel, A.; Faller, S. Hydrogen Sulfide Exerts Anti-oxidative and Anti-inflammatory Effects in Acute Lung Injury. Inflammation 2018, 41, 249–259. [Google Scholar] [CrossRef]
- Griese, M.; Kirmeier, H.G.; Liebisch, G.; Rauch, D.; Stuckler, F.; Schmitz, G.; Zarbock, R.; Kids-Lung-Register, I.-B.w.g.o.t. Surfactant lipidomics in healthy children and childhood interstitial lung disease. PLoS ONE 2015, 10, e0117985. [Google Scholar] [CrossRef]
- Chen, Y.H.; Lv, J.W. Building the acute lung injury mouse model by LPS direct and indirect challenge. Chin. J. Clin. Anatomy 2015, 33, 439–443. [Google Scholar]
- Shan, J.; Qian, W.; Kang, A.; Peng, L.; Xie, T.; Lin, L.; Di, L.; Xiao, P.; Zhou, W. Lipid profile perturbations in the plasma and lungs of mice with LPS-induced acute lung injury revealed by UHPLC-ESI-Q Exactive HF MS analysis. J. Pharm. Biomed. Anal. 2019, 162, 242–248. [Google Scholar] [CrossRef]
- Jiao, G.Y.; Nie, Z.W. A Rat Model of Acute Lung Injury Induced by Lipopolysaccharide. Acta Laboratorium. Animalis Scientia Sinica 2007, 15, 292–295. [Google Scholar]
- Miyoshi, M.; Iizuka, N.; Sakai, S.; Fujiwara, M.; Aoyama-Ishikawa, M.; Maeshige, N.; Hamada, Y.; Takahashi, M.; Usami, M. Oral tributyrin prevents endotoxin-induced lipid metabolism disorder. Clin. Nutr. ESPEN 2015, 10, e83–e88. [Google Scholar] [CrossRef]
- Vaz, F.M.; Pras-Raves, M.; Bootsma, A.H.; van Kampen, A.H. Principles and practice of lipidomics. J. Inherit. Metab. Dis. 2015, 38, 41–52. [Google Scholar] [CrossRef]
- Zhao, Y.Y.; Miao, H.; Cheng, X.L.; Wei, F. Lipidomics: Novel insight into the biochemical mechanism of lipid metabolism and dysregulation-associated disease. Chem. Biol. Interact. 2015, 240, 220–238. [Google Scholar] [CrossRef]
- Tsugawa, H.; Cajka, T.; Kind, T.; Ma, Y.; Higgins, B.; Ikeda, K.; Kanazawa, M.; VanderGheynst, J.; Fiehn, O.; Arita, M. MS-DIAL: Data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 2015, 12, 523–526. [Google Scholar] [CrossRef]
- Han, S.; Mallampalli, R.K. The Role of Surfactant in Lung Disease and Host Defense against Pulmonary Infections. Ann. Am. Thorac. Soc. 2015, 12, 765–774. [Google Scholar] [CrossRef]
- Wang, M.; Wang, C.; Han, R.H.; Han, X. Novel advances in shotgun lipidomics for biology and medicine. Prog. Lipid Res. 2016, 61, 83–108. [Google Scholar] [CrossRef]
- Shan, J.J.; Qian, W.J. High-resolution lipidomics reveals dysregulation of lipid metabolism in respiratory syncytial virus pneumonia mice. RSC Adv. 2018, 8, 29368–29377. [Google Scholar] [CrossRef]
- Furse, S.; de Kroon, A.I. Phosphatidylcholine’s functions beyond that of a membrane brick. Mol. Membr. Biol. 2015, 32, 117–119. [Google Scholar] [CrossRef]
- Agassandian, M.; Mallampalli, R.K. Surfactant phospholipid metabolism. Biochim. Biophys. Acta 2013, 1831, 612–625. [Google Scholar] [CrossRef]
- Xie, T.; Du, L.N. Metabonomics Study on the Effect of Polygonum Cuspidatumon Anti-respiratory Syncytial Virus (RSV) Basing on Lipid Metabolism. World Chin. Med. 2016, 11, 1670–1673. [Google Scholar] [CrossRef]
- Du, L.N.; Shan, J.J. Lipidomics study on antiviral effects of baical skullcap root extraction based on liquid chromatography-mass spectrography. Chin. J. Tradit. Chin. Med. Pharm. 2015, 1728–1733. [Google Scholar]
- Wang, Y.; Roller, J.; Slotta, J.E.; Zhang, S.; Luo, L.; Rahman, M.; Syk, I.; Menger, M.D.; Thorlacius, H. Distinct patterns of leukocyte recruitment in the pulmonary microvasculature in response to local and systemic inflammation. Am. J. Physiol. Lung Cell Mol. Physiol. 2013, 304, L298–L305. [Google Scholar] [CrossRef]
- Kapoor, K.; Singla, E.; Sahu, B.; Naura, A.S. PARP inhibitor, olaparib ameliorates acute lung and kidney injury upon intratracheal administration of LPS in mice. Mol. Cell Biochem. 2015, 400, 153–162. [Google Scholar] [CrossRef]
- Cajka, T.; Fiehn, O. Comprehensive analysis of lipids in biological systems by liquid chromatography-mass spectrometry. Trends Analyt. Chem. 2014, 61, 192–206. [Google Scholar] [CrossRef]
- Chen, S.; Hoene, M.; Li, J.; Li, Y.; Zhao, X.; Häring, H.U.; Schleicher, E.D.; Weigert, C.; Xu, G.; Lehmann, R. Simultaneous extraction of metabolome and lipidome with methyl tert-butyl ether from a single small tissue sample for ultra-high performance liquid chromatography/mass spectrometry. J. Chromatogr. A 2013, 1298, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.Y.; Kuo, C.H.; Tseng, Y.J. Batch Normalizer: A fast total abundance regression calibration method to simultaneously adjust batch and injection order effects in liquid chromatography/time-of-flight mass spectrometry-based metabolomics data and comparison with current calibration methods. Anal. Chem. 2013, 85, 1037–1046. [Google Scholar] [CrossRef]
- Fan, S.; Kind, T.; Cajka, T.; Hazen, S.L.; Tang, W.H.W.; Kaddurah-Daouk, R.; Irvin, M.R.; Arnett, D.K.; Barupal, D.K.; Fiehn, O. Systematic Error Removal Using Random Forest for Normalizing Large-Scale Untargeted Lipidomics Data. Anal. Chem. 2019, 91, 3590–3596. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, R.; Zhang, Y.; Qian, W.; Peng, L.; Lin, L.; Xu, J.; Xie, T.; Ji, J.; Zhan, X.; Shan, J. Surfactant Lipidomics of Alveolar Lavage Fluid in Mice Based on Ultra-High-Performance Liquid Chromatography Coupled to Hybrid Quadrupole-Exactive Orbitrap Mass Spectrometry. Metabolites 2019, 9, 80. https://doi.org/10.3390/metabo9040080
Yang R, Zhang Y, Qian W, Peng L, Lin L, Xu J, Xie T, Ji J, Zhan X, Shan J. Surfactant Lipidomics of Alveolar Lavage Fluid in Mice Based on Ultra-High-Performance Liquid Chromatography Coupled to Hybrid Quadrupole-Exactive Orbitrap Mass Spectrometry. Metabolites. 2019; 9(4):80. https://doi.org/10.3390/metabo9040080
Chicago/Turabian StyleYang, Rui, Ying Zhang, Wenjuan Qian, Linxiu Peng, Lili Lin, Jia Xu, Tong Xie, Jianjian Ji, Xiuqin Zhan, and Jinjun Shan. 2019. "Surfactant Lipidomics of Alveolar Lavage Fluid in Mice Based on Ultra-High-Performance Liquid Chromatography Coupled to Hybrid Quadrupole-Exactive Orbitrap Mass Spectrometry" Metabolites 9, no. 4: 80. https://doi.org/10.3390/metabo9040080
APA StyleYang, R., Zhang, Y., Qian, W., Peng, L., Lin, L., Xu, J., Xie, T., Ji, J., Zhan, X., & Shan, J. (2019). Surfactant Lipidomics of Alveolar Lavage Fluid in Mice Based on Ultra-High-Performance Liquid Chromatography Coupled to Hybrid Quadrupole-Exactive Orbitrap Mass Spectrometry. Metabolites, 9(4), 80. https://doi.org/10.3390/metabo9040080





