Metabolomics of Small Intestine Neuroendocrine Tumors and Related Hepatic Metastases
Abstract
1. Introduction
2. Results
2.1. Patient Population and Tumor Characteristics
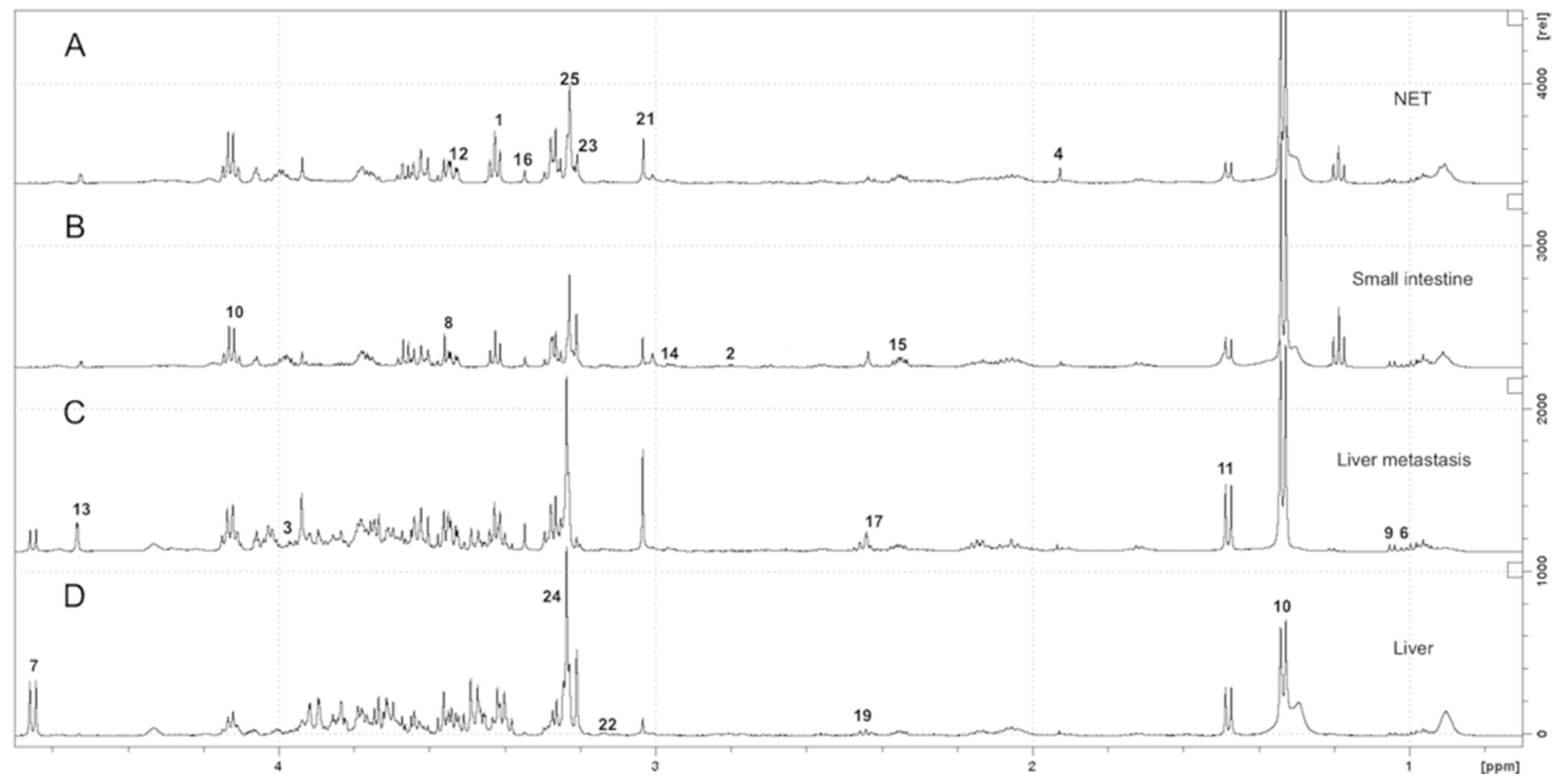
2.2. HRMAS NMR Spectra
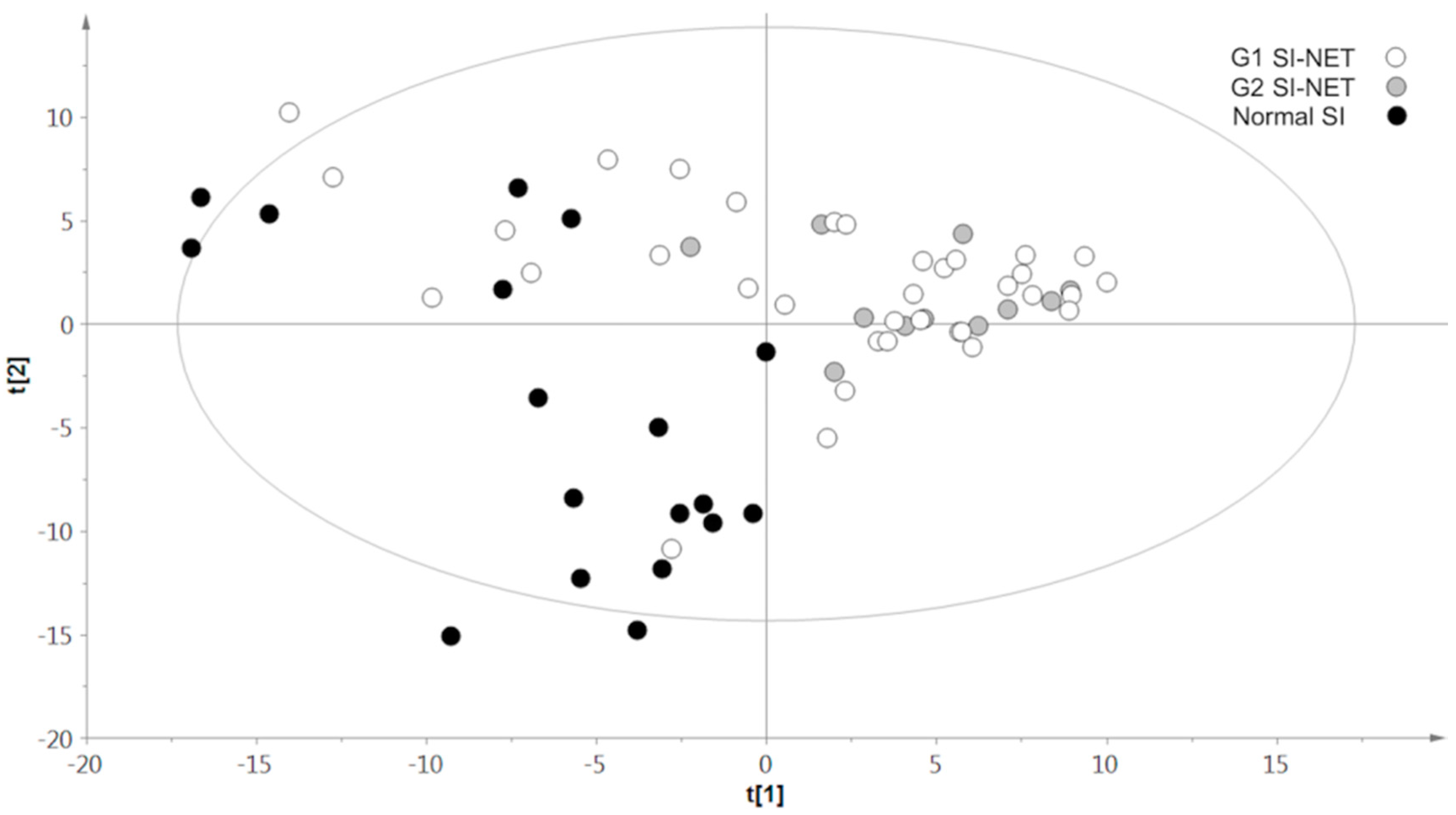
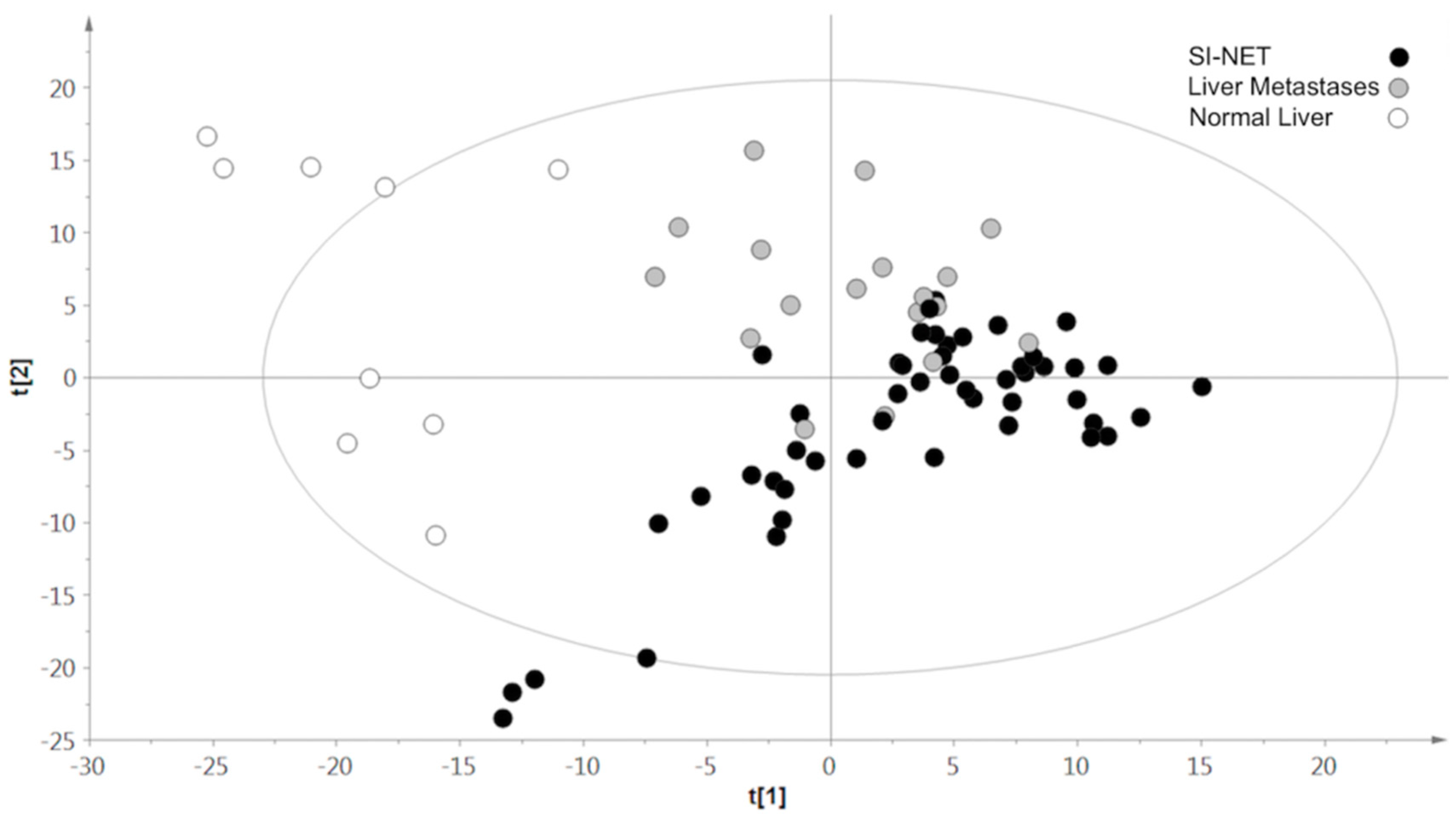
2.3. PLS-DA Results
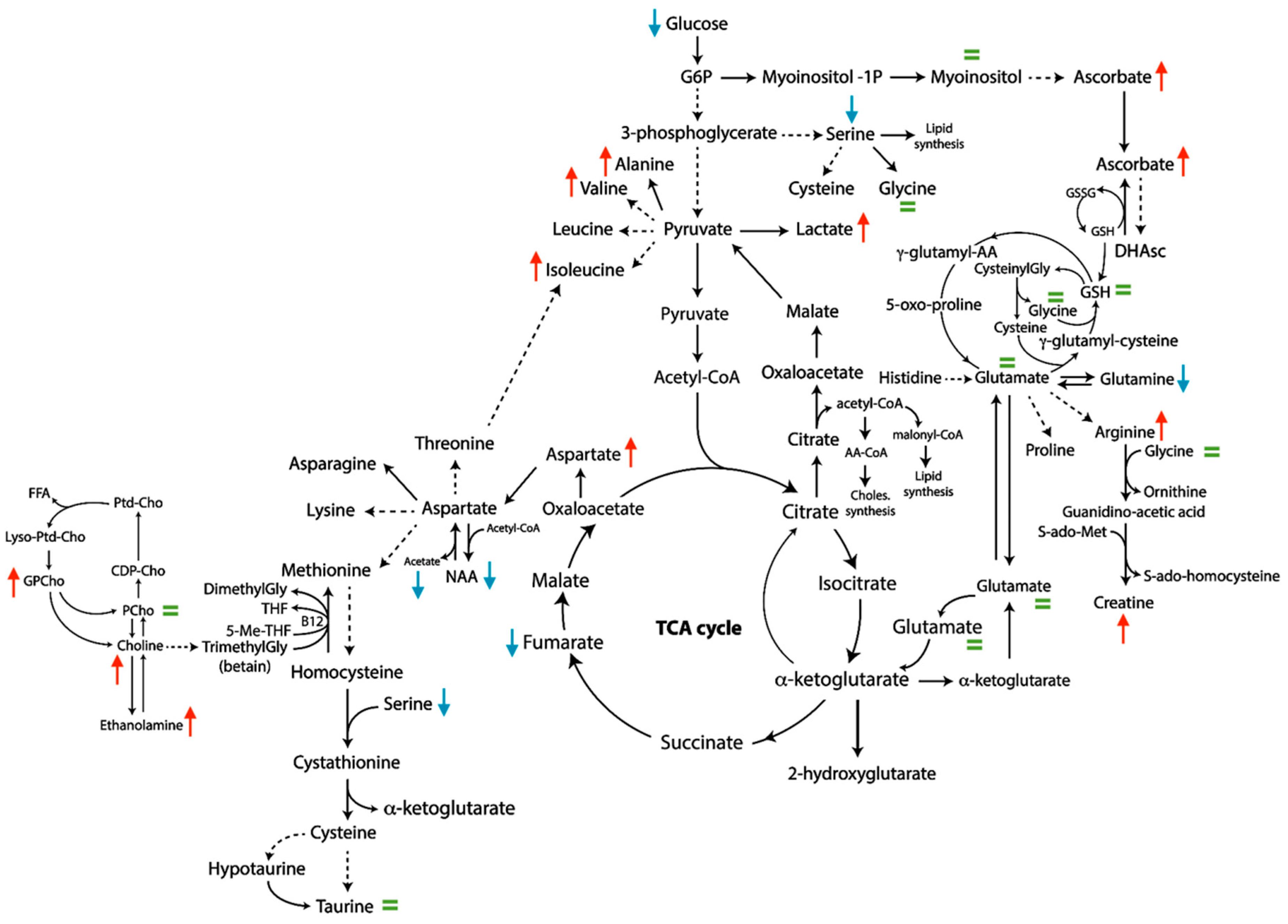
2.4. Univariate and ADEMA Network Analysis
3. Discussion
4. Materials and Methods
4.1. Patient Population
4.2. Tissue Sample Preparation
4.3. HRMAS NMR Technical Features
4.4. Metabolite Quantification Procedure
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Dasari, A.; Shen, C.; Halperin, D.; Zhao, B.; Zhou, S.; Xu, Y.; Shih, T.; Yao, J.C. Trends in the Incidence, Prevalence, and Survival Outcomes in Patients With Neuroendocrine Tumors in the United States. JAMA Oncol. 2017, 3, 1335–1342. [Google Scholar] [CrossRef] [PubMed]
- Keck, K.J.; Maxwell, J.E.; Utria, A.F.; Bellizzi, A.M.; Dillon, J.S.; O’Dorisio, T.M.; Howe, J.R. The Distal Predilection of Small Bowel Neuroendocrine Tumors. Ann. Surg. Oncol. 2018, 25, 3207–3213. [Google Scholar] [CrossRef] [PubMed]
- Halperin, D.M.; Shen, C.; Dasari, A.; Xu, Y.; Chu, Y.; Zhou, S.; Shih, Y.C.; Yao, J.C. Frequency of carcinoid syndrome at neuroendocrine tumour diagnosis: A population-based study. Lancet Oncol. 2017, 18, 525–534. [Google Scholar] [CrossRef]
- Klimstra, D.S.; Modlin, I.R.; Coppola, D.; Lloyd, R.V.; Suster, S. The pathologic classification of neuroendocrine tumors: A review of nomenclature, grading, and staging systems. Pancreas 2010, 39, 707–712. [Google Scholar] [CrossRef] [PubMed]
- Warburg, O. On the origin of cancer cells. Science 1956, 123, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Lindsey KBoroughs, L.K.; DeBerardinis, R.J. Metabolic pathways promoting cancer cell survival and growth. Nat. Cell Biol. 2015, 17, 351–359. [Google Scholar]
- Luengo, A.; Gui, D.Y.; Vander Heiden, M.J. Targeting Metabolism for Cancer Therapy. Cell Chem. Biol. 2017, 24, 1161–1180. [Google Scholar] [CrossRef]
- Muir, A.; Danai, L.V.; Vander Heiden, M.J. Microenvironmental regulation of cancer cell metabolism: Implications for experimental design and translational studies. Dis. Models Mech. 2018, 11, dmm035758. [Google Scholar] [CrossRef]
- Wishart, D.S. Exploring the Human Metabolome by Nuclear Magnetic Resonance Spectroscopy and Mass Spectroscopy; Lutz, N.W., Ed.; Cambridge University Press: Cambridge, UK, 2013. [Google Scholar]
- Beger, R.D.; Dunn, W.; Schmidt, M.A.; Gross, S.S.; Kirwan, J.A.; Cascante, M.; Brennan, L.; Wishart, D.S.; Oresic, M.; Hankemeier, T.; et al. Metabolomics enables precision medicine: “A White Paper, Community Perspective”. Metabolomics 2016, 12, 149. [Google Scholar] [CrossRef]
- Beckonert, O.; Coen, M.; Keun, H.C.; Wang, Y.; Ebbels, T.M.; Holmes, E.; Lindon, J.C.; Nicholson, J.K. High-resolution magic-angle-spinning NMR spectroscopy for metabolic profiling of intact tissues. Nat. Protoc. 2010, 5, 1019–1032. [Google Scholar] [CrossRef]
- Piotto, M.; Moussallieh, F.M.; Imperiale, A.; Benahmed, M.A.; Detour, J.; Bellocq, J.P.; Namer, J.I.; Elbayed, K. Reproducible sample preparation and spectrum acquisition techniques for metabolic profiling of human tissues by proton high-resolution magic angle spinning nuclear magnetic resonance. In Metabolomic Nuclear Magnetic Resonance Spectroscopy Techniques for Body Tissue Analysis; Lutz, N.W., Ed.; Cambridge University Press: Cambridge, UK, 2013; pp. 496–524. [Google Scholar]
- Wishart, D.S.; Jewison, T.; Guo, A.C.; Wilson, M.; Knox, C.; Liu, Y.; Djoumbou, Y.; Mandal, R.; Aziat, F.; Dong, E.; et al. HMDB 3.0—The Human Metabolome Database in 2013. Nucleic Acids Res. 2013, 41, D801–D807. [Google Scholar] [CrossRef] [PubMed]
- Ruhland, E.; Bund, C.; Outilaft, H.; Piotto, M.; Namer, I.J. A metabolic database for biomedical studies of biopsy specimens by high-resolution magic angle spinning nuclear MR: A qualitative and quantitative tool. Magn. Reson. Med. 2019, 82, 62–83. [Google Scholar] [CrossRef] [PubMed]
- Imperiale, A.; Moussallieh, F.M.; Roche, P.; Battini, S.; Cicek, A.E.; Sebag, F.; Brunaud, L.; Barlier, A.; Elbayed, K.; Loundou, A.; et al. Metabolome profiling by HRMAS NMR spectroscopy of pheochromocytomas and paragangliomas detects SDH deficiency: Clinical and pathophysiological implications. Neoplasia 2015, 17, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Battini, S.; Faitot, F.; Imperiale, A.; Cicek, A.E.; Heimburger, C.; Averous, G.; Bachellier, P.; Namer, I.J. Metabolomics approaches in pancreatic adenocarcinoma: Tumor metabolism profiling predicts clinical outcome of patients. BMC Med. 2017, 15, 56. [Google Scholar] [CrossRef] [PubMed]
- Worley, B.; Powers, R. Multivariate Analysis in Metabolomics. Curr. Metab. 2013, 1, 92–107. [Google Scholar]
- Cicek, A.E.; Bederman, I.; Henderson, L.; Drumm, M.L.; Ozsoyoglu, G. ADEMA: An algorithm to determine expected metabolite level alterations using mutual information. PLoS Comput. Biol. 2013, 9, e1002859. [Google Scholar] [CrossRef] [PubMed]
- Kinross, J.M.; Drymousis, P.; Jiménez, B.; Frilling, A. Metabonomic profiling: A novel approach in neuroendocrine neoplasias. Surgery 2013, 154, 1185–1193. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Letouzé, E.; Martinelli, C.; Loriot, C.; Burnichon, N.; Abermil, N.; Ottolenghi, C.; Janin, M.; Menara, M.; Nguyen, A.T.; Benit, P.; et al. SDH mutations establish a hypermethylator phenotype in paraganglioma. Cancer Cell 2013, 23, 739–752. [Google Scholar] [CrossRef]
- Garrigue, P.; Bodin-Hullin, A.; Balasse, L.; Fernandez, S.; Essamet, W.; Dignat-George, F.; Pacak, K.; Guillet, B.; Taieb, D. The Evolving Role of Succinate in Tumor Metabolism: An 18 F-FDG–Based Study. J. Nucl. Med. 2017, 58, 1749–1755. [Google Scholar] [CrossRef]
- Trettera, L.; Patocsb, A.; Chinopoulos, C. Succinate, an intermediate in metabolism, signal transduction, ROS, hypoxia, and tumorigenesis. Biochim. Biophys. Acta. 2016, 1857, 1086–1101. [Google Scholar] [CrossRef] [PubMed]
- Rosado, J.O.; Salvador, M.; Bonatto, D. Importance of the trans-sulfuration pathway in cancer prevention and promotion. Mol. Cell. Biochem. 2007, 301, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Redmond, H.P.; Stapleton, P.P.; Neary, P.; Bouchier-Hayes, D. Immunonutrition: The role of taurine. Nutrition 1998, 14, 599–604. [Google Scholar] [CrossRef]
- Eagle, H.; Oyama, V.I.; Levy, M.; Freeman, A.E. Myo-inositol as an essential growth factor for normal and malignant human cells in tissue culture. J. Biol. Chem. 1957, 226, 191–205. [Google Scholar] [CrossRef] [PubMed]
- Croze, M.L.; Soulage, C.O. Potential role and therapeutic interests of myo-inositol in metabolic diseases. Biochimie 2013, 95, 1811–1827. [Google Scholar] [CrossRef] [PubMed]
- Sei, Y.; Zhao, X.; Forbes, J.; Szymczak, S.; Li, Q.; Trivedi, A.; Voellinger, M.; Joy, G.; Feng, J.; Whatley, M.; et al. A Hereditary Form of Small Intestinal Carcinoid Associated With a Germline Mutation in Inositol Polyphosphate Multikinase. Gastroenterology 2015, 149, 67–78. [Google Scholar] [CrossRef]
- Deroose, C.M.; Hindié, E.; Kebebew, E.; Goichot, B.; Pacak, K.; Taïeb, D.; Imperiale, A. Molecular imaging of gastroenteropancreatic neuroendocrine tumors: Current status and future directions. J. Nucl. Med. 2016, 57, 1949–1956. [Google Scholar] [CrossRef]
- Mattaini, K.R.; Sullivan, M.R.; Vander Heiden, M.G. The importance of serine metabolism in cancer. J. Cell Biol. 2016, 214, 249–257. [Google Scholar] [CrossRef]
- Jain, M.; Nilsson, R.; Sharma, S.; Madhusudhan, N.; Kitami, T.; Souza, A.L.; Kafri, R.; Kirschner, M.W.; Clish, C.B.; Mootha, V.K. Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation. Science 2012, 336, 1040–1044. [Google Scholar] [CrossRef]
- Hosios, A.M.; Hecht, V.C.; Danai, L.V.; Johnson, M.O.; Rathmell, J.C.; Steinhauser, M.L.; Manalis, S.R.; Vander Heiden, M.G. Amino acids rather than glucose account for the majority of cell mass in proliferating mammalian cells. Dev. Cell 2016, 36, 540–549. [Google Scholar] [CrossRef]
- Dupuy, F.; Tabariès, S.; Andrzejewski, S.; Dong, Z.; Blagih, J.; Annis, M.G.; Omeroglu, A.; Gao, D.; Leung, S.; Amir, E.; et al. PDK1-Dependent Metabolic Reprogramming Dictates Metastatic Potential in Breast Cancer. Cell Metab. 2015, 22, 577–589. [Google Scholar] [CrossRef] [PubMed]
- Brand, A.; Singer, K.; Koehl, G.E.; Kolitzus, M.; Schoenhammer, G.; Thiel, A.; Matos, C.; Bruss, C.; Klobuch, S.; Peter, K.; et al. Lactic Acid Production Blunts Tumor Immunosurveillance by T and NK Cells. Cell Metab. 2016, 24, 657–671. [Google Scholar] [CrossRef] [PubMed]
- Colegio, O.R.; Chu, N.Q.; Szabo, A.L.; Chu, T.; Rhebergen, A.M.; Jairam, V.; Cyrus, N.; Brokowski, C.E.; Eisenbarth, S.C.; Phillips, G.M.; et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 2014, 513, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, L.B.; Luengo, A.; Danai, L.V.; Bush, L.N.; Diehl, F.F.; Hosios, A.M.; Lau, A.N.; Elmiligy, S.; Malstrom, S.; Lewis, C.A.; et al. Aspartate is an endogenous metabolic limitation for tumour growth. Nat. Cell Biol. 2018, 20, 782–788. [Google Scholar] [CrossRef]
- Jordan, K.W.; He, W.; Halpern, E.F.; Wu, C.L.; Cheng, L.L. Evaluation of Tissue Metabolites with High Resolution Magic Angle Spinning MR Spectroscopy Human Prostate Samples After Three-Year Storage at -80 degrees C. Biomark. Insights 2007, 2, 147–154. [Google Scholar] [CrossRef]
- Wider, G.; Dreier, L. Measuring protein concentrations by NMR spectroscopy. J. Am. Chem. Soc. 2006, 128, 2571–2576. [Google Scholar] [CrossRef]
- Wold, S.; Ruhe, A.; Wold, H.; Dunn, W.J.J. The collinearity problem in linear regression. The partial least squares (PLS) approach to generalized inverses. SIAM J. Sci. Stat. Comput. 1984, 5, 735–743. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. Data, information, knowledge and principle: Back to metabolism in KEGG. Nucleic Acids Res. 2014, 42, D199–D205. [Google Scholar] [CrossRef]
- Selway, Z.Z. Metabolism at a Glance, 3rd ed.; Blackwell Publishing: Malden, MI, USA, 2014. [Google Scholar]
| Metabolite | Tumor Grade | Perineural-Invasion | Angio-Invasion | Parietal Infiltration | Tumor Secretion | Metastases at Diagnosis | Primary T Multifocality |
|---|---|---|---|---|---|---|---|
| G1 (35) vs. G2 (11) | Yes (13) vs. No (8) | Yes (16) vs. No (8) | Deep (38) vs. Superficial (5) | Yes (13) vs. No (33) | Yes (31) vs. No (15) | Yes (16) vs. No (30) | |
| Taurine | = | up | up | up | = | = | up |
| Aspartate | up | up | = | down | down | up | up |
| Serine | up | down | down | down | down | down | down |
| Acetate | up | up | down | down | down | down | down |
| NAA | up | down | down | up | up | down | down |
| Isoleucine | up | down | down | up | down | up | down |
| Glucose | up | down | = | down | down | down | = |
| Glycine | down | = | down | down | = | = | = |
| Valine | up | down | down | = | down | up | down |
| Lactate | = | up | up | up | up | up | up |
| Alanine | = | = | down | up | down | up | down |
| Myoinositol | = | = | = | up | = | = | up |
| Ascorbate | up | = | = | down | up | up | = |
| GSH | = | = | = | = | = | = | = |
| Glutamate | = | = | down | up | = | = | = |
| Scylloinositol | = | down | = | up | down | = | up |
| Succinate | up | down | = | = | = | = | up |
| Fumarate | = | up | down | down | up | down | down |
| Glutamine | up | = | down | up | up | down | = |
| Arginine | up | = | down | up | up | up | up |
| Creatine | up | = | down | down | down | up | up |
| Ethalonamine | up | down | down | = | = | up | = |
| Choline | = | up | up | = | up | up | up |
| GPC | = | up | up | = | = | up | up |
| PC | = | = | = | down | up | = | up |
| Tyrosine | up | down | down | up | up | down | down |
| Tryptophane | up | up | up | up | = | up | = |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Imperiale, A.; Poncet, G.; Addeo, P.; Ruhland, E.; Roche, C.; Battini, S.; Cicek, A.E.; Chenard, M.P.; Hervieu, V.; Goichot, B.; et al. Metabolomics of Small Intestine Neuroendocrine Tumors and Related Hepatic Metastases. Metabolites 2019, 9, 300. https://doi.org/10.3390/metabo9120300
Imperiale A, Poncet G, Addeo P, Ruhland E, Roche C, Battini S, Cicek AE, Chenard MP, Hervieu V, Goichot B, et al. Metabolomics of Small Intestine Neuroendocrine Tumors and Related Hepatic Metastases. Metabolites. 2019; 9(12):300. https://doi.org/10.3390/metabo9120300
Chicago/Turabian StyleImperiale, Alessio, Gilles Poncet, Pietro Addeo, Elisa Ruhland, Colette Roche, Stephanie Battini, A. Ercument Cicek, Marie Pierrette Chenard, Valérie Hervieu, Bernard Goichot, and et al. 2019. "Metabolomics of Small Intestine Neuroendocrine Tumors and Related Hepatic Metastases" Metabolites 9, no. 12: 300. https://doi.org/10.3390/metabo9120300
APA StyleImperiale, A., Poncet, G., Addeo, P., Ruhland, E., Roche, C., Battini, S., Cicek, A. E., Chenard, M. P., Hervieu, V., Goichot, B., Bachellier, P., Walter, T., & Namer, I. J. (2019). Metabolomics of Small Intestine Neuroendocrine Tumors and Related Hepatic Metastases. Metabolites, 9(12), 300. https://doi.org/10.3390/metabo9120300






