FoodPro: A Web-Based Tool for Evaluating Covariance and Correlation NMR Spectra Associated with Food Processes
Abstract
:1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. Accumulation of Experimental Data
3.2. Database Design
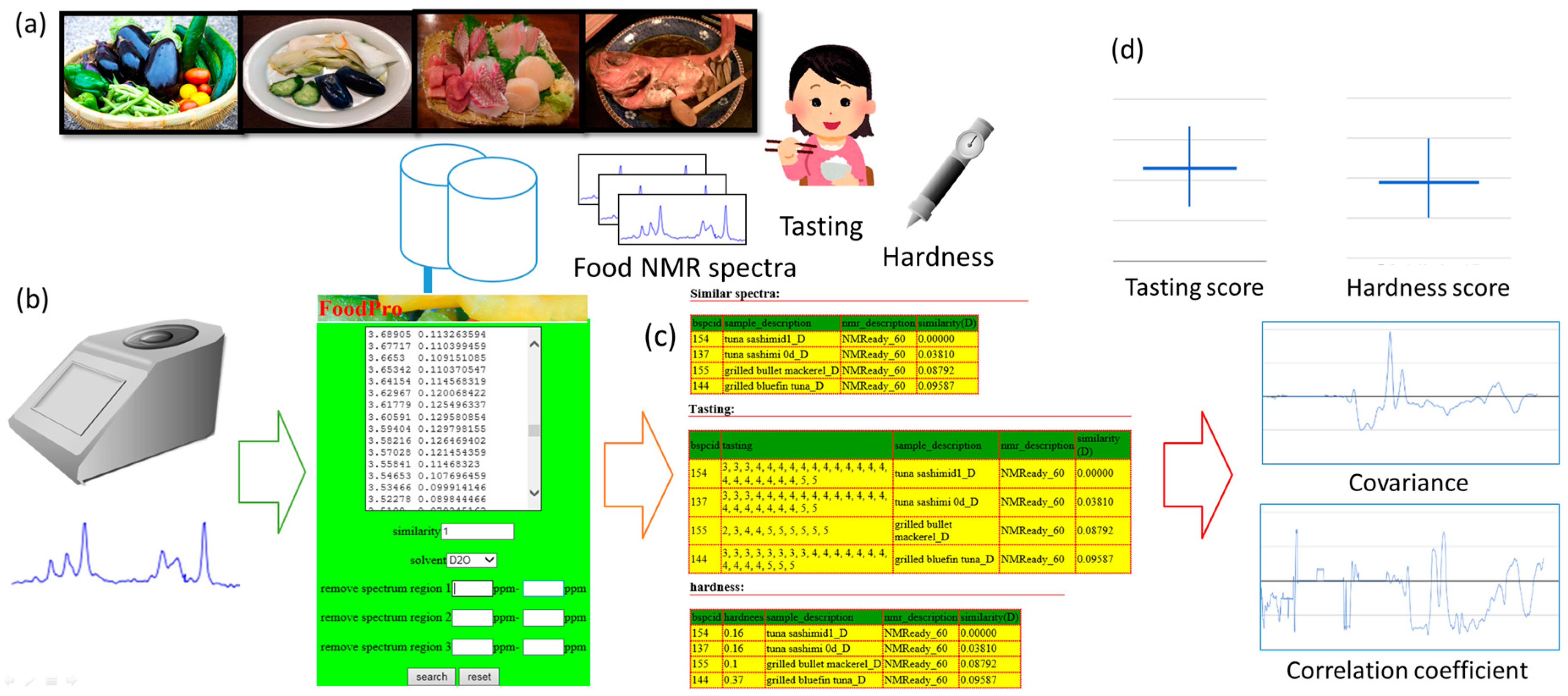
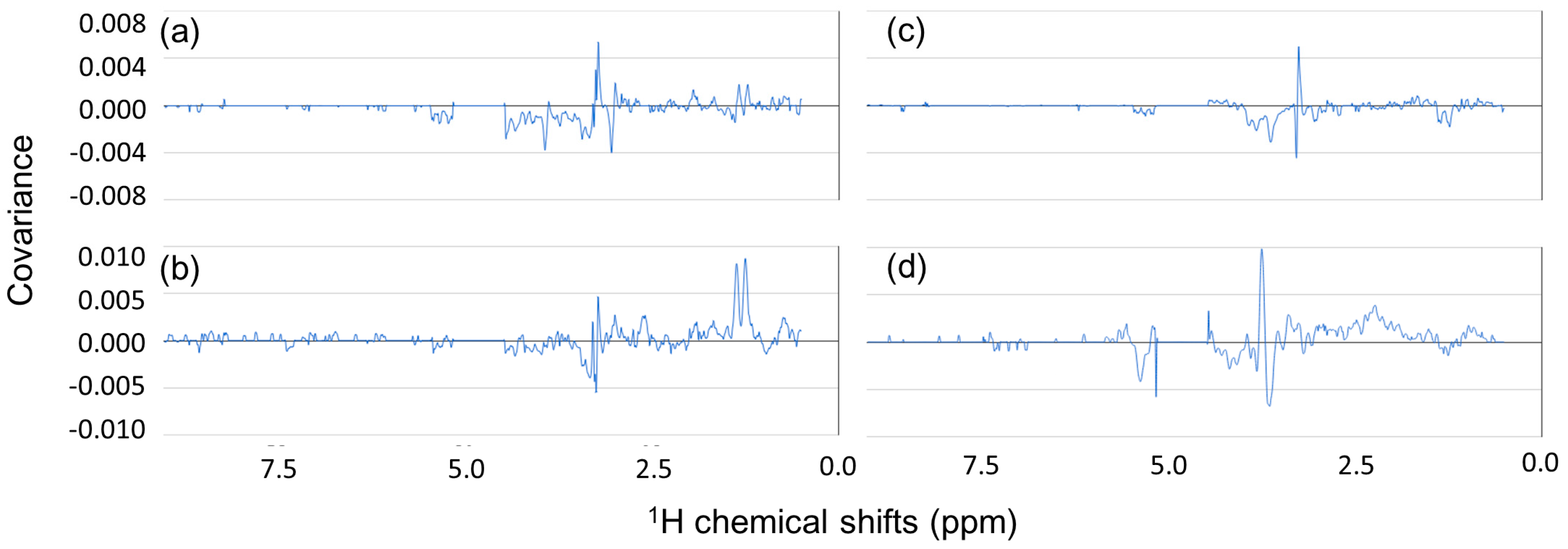
3.3. Computation of Covariance and Correlation Spectra for Tasting and Hardness
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Erikson, U.; Standal, I.B.; Aursand, I.G.; Veliyulin, E.; Aursand, M. Use of NMR in fish processing optimization: A review of recent progress. Magn. Reson. Chem. 2012, 50, 471–480. [Google Scholar] [CrossRef] [PubMed]
- Melito, H.S.; Daubert, C.R. Rheological innovations for characterizing food material properties. Annu. Rev. Food Sci. Technol. 2011, 2, 153–179. [Google Scholar] [CrossRef] [PubMed]
- Colnago, L.A.; Azeredo, R.B.; Marchi Netto, A.; Andrade, F.D.; Venancio, T. Rapid analyses of oil and fat content in agri-food products using continuous wave free precession time domain NMR. Magn. Reson. Chem. 2011, 49 (Suppl. S1), S113–S120. [Google Scholar] [CrossRef] [PubMed]
- Creusot, N.; Gruppen, H. Enzyme-induced aggregation and gelation of proteins. Biotechnol. Adv. 2007, 25, 597–601. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.J.; Gruppen, H.; Maes, D.; Boots, J.W.; Wierenga, P.A. Factors causing compositional changes in soy protein hydrolysates and effects on cell culture functionality. J. Agric. Food Chem. 2013, 61, 10613–10625. [Google Scholar] [CrossRef] [PubMed]
- Larsen, F.H.; Engelsen, S.B. Insight into the Functionality of Microbial Exopolysaccharides by NMR Spectroscopy and Molecular Modeling. Front. Microbiol. 2015, 6, 1374. [Google Scholar] [CrossRef] [PubMed]
- Larsen, F.H.; Byg, I.; Damager, I.; Diaz, J.; Engelsen, S.B.; Ulvskov, P. Residue specific hydration of primary cell wall potato pectin identified by solid-state 13C single-pulse MAS and CP/MAS NMR spectroscopy. Biomacromolecules 2011, 12, 1844–1850. [Google Scholar] [CrossRef] [PubMed]
- Larsen, F.H.; Schobitz, M.; Schaller, J. Hydration properties of regioselectively etherified celluloses monitored by 2H and 13C solid-state MAS NMR spectroscopy. Carbohydr. Polym. 2012, 89, 640–647. [Google Scholar] [CrossRef] [PubMed]
- Mochida, K.; Furuta, T.; Ebana, K.; Shinozaki, K.; Kikuchi, J. Correlation exploration of metabolic and genomic diversity in rice. BMC Genom. 2009, 10, 568. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, S.; Toh, H.; Hase, K.; Oshima, K.; Nakanishi, Y.; Yoshimura, K.; Tobe, T.; Clarke, J.M.; Topping, D.L.; Suzuki, T.; et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 2011, 469, 543–547. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, Y.; Fukuda, S.; Chikayama, E.; Kimura, Y.; Ohno, H.; Kikuchi, J. Dynamic omics approach identifies nutrition-mediated microbial interactions. J. Proteome Res. 2011, 10, 824–836. [Google Scholar] [CrossRef] [PubMed]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Dumas, M.E.; Maibaum, E.C.; Teague, C.; Ueshima, H.; Zhou, B.; Lindon, J.C.; Nicholson, J.K.; Stamler, J.; Elliott, P.; Chan, Q.; et al. Assessment of analytical reproducibility of 1H NMR spectroscopy based metabonomics for large-scale epidemiological research: The INTERMAP Study. Anal. Chem. 2006, 78, 2199–2208. [Google Scholar] [CrossRef] [PubMed]
- Viant, M.R.; Bearden, D.W.; Bundy, J.G.; Burton, I.W.; Collette, T.W.; Ekman, D.R.; Ezernieks, V.; Karakach, T.K.; Lin, C.Y.; Rochfort, S.; et al. International NMR-based environmental metabolomics intercomparison exercise. Environ. Sci. Technol. 2009, 43, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Ward, J.L.; Baker, J.M.; Miller, S.J.; Deborde, C.; Maucourt, M.; Biais, B.; Rolin, D.; Moing, A.; Moco, S.; Vervoort, J.; et al. An inter-laboratory comparison demonstrates that [H]-NMR metabolite fingerprinting is a robust technique for collaborative plant metabolomic data collection. Metabolomics 2010, 6, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, M.; Ohta, Y.; Licang, S.; Motoyama, N.; Kikuchi, J. Profiling contents of water-soluble metabolites and mineral nutrients to evaluate the effects of pesticides and organic and chemical fertilizers on tomato fruit quality. Food Chem. 2015, 169, 387–395. [Google Scholar] [CrossRef] [PubMed]
- Tomita, S.; Nemoto, T.; Matsuo, Y.; Shoji, T.; Tanaka, F.; Nakagawa, H.; Ono, H.; Kikuchi, J.; Ohnishi-Kameyama, M.; Sekiyama, Y. A NMR-based, non-targeted multistep metabolic profiling revealed L-rhamnitol as a metabolite that characterised apples from different geographic origins. Food Chem. 2015, 174, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, S.; Date, Y.; Akama, M.; Kikuchi, J. Comparative metabolomic and ionomic approach for abundant fishes in estuarine environments of Japan. Sci. Rep. 2014, 4, 7005. [Google Scholar] [CrossRef] [PubMed]
- Misawa, T.; Wei, F.; Kikuchi, J. Application of Two-Dimensional Nuclear Magnetic Resonance for Signal Enhancement by Spectral Integration Using a Large Data Set of Metabolic Mixtures. Anal. Chem. 2016, 88, 6130–6134. [Google Scholar] [CrossRef] [PubMed]
- Ito, K.; Sakata, K.; Date, Y.; Kikuchi, J. Integrated Analysis of Seaweed Components during Seasonal Fluctuation by Data Mining Across Heterogeneous Chemical Measurements with Network Visualization. Anal. Chem. 2014, 86, 1098–1105. [Google Scholar] [CrossRef] [PubMed]
- Brennan, L. NMR-based metabolomics: From sample preparation to applications in nutrition research. Prog. Nucl. Magn. Reson. Spectrosc. 2014, 83, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Ellis, D.I.; Brewster, V.L.; Dunn, W.B.; Allwood, J.W.; Golovanov, A.P.; Goodacre, R. Fingerprinting food: Current technologies for the detection of food adulteration and contamination. Chem. Soc. Rev. 2012, 41, 5706–5727. [Google Scholar] [CrossRef] [PubMed]
- Valentini, M.; Ritota, M.; Cafiero, C.; Cozzolino, S.; Leita, L.; Sequi, P. The HRMAS-NMR tool in foodstuff characterisation. Magn. Reson. Chem. 2011, 49 (Suppl. S1), S121–S125. [Google Scholar] [CrossRef] [PubMed]
- Gudjonsdottir, M.; Jonsson, A.; Bergsson, A.B.; Arason, S.; Rustad, T. Shrimp processing assessed by low field nuclear magnetic resonance, near infrared spectroscopy, and physicochemical measurements—The effect of polyphosphate content and length of prebrining on shrimp muscle. J. Food Sci. 2011, 76, E357–E367. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Cai, S.; Zhang, Z.; Chen, Z. High-resolution two-dimensional J-resolved NMR spectroscopy for biological systems. Biophys. J. 2014, 106, 2061–2070. [Google Scholar] [CrossRef] [PubMed]
- Pages, G.; Gerdova, A.; Williamson, D.; Gilard, V.; Martino, R.; Malet-Martino, M. Evaluation of a benchtop cryogen-free low-field 1H NMR spectrometer for the analysis of sexual enhancement and weight loss dietary supplements adulterated with pharmaceutical substances. Anal. Chem. 2014, 86, 11897–11904. [Google Scholar] [CrossRef] [PubMed]
- Parker, T.; Limer, E.; Watson, A.D.; Defernez, M.; Williamson, D.; Kemsley, E.K. 60 MHz H NMR spectroscopy for the analysis of edible oils. Trends Anal. Chem. 2014, 57, 147–158. [Google Scholar] [CrossRef] [PubMed]
- Rolletschek, H.; Fuchs, J.; Friedel, S.; Borner, A.; Todt, H.; Jakob, P.M.; Borisjuk, L. A novel noninvasive procedure for high-throughput screening of major seed traits. Plant Biotechnol. J. 2015, 13, 188–199. [Google Scholar] [CrossRef] [PubMed]
- Silva Elipe, M.V.; Milburn, R.R. Monitoring chemical reactions by low-field benchtop NMR at 45 MHz: Pros and cons. Magn. Reson. Chem. 2016, 54, 437–443. [Google Scholar] [CrossRef] [PubMed]
- FoodPro. Available online: http://emar.riken.jp/FoodPro/ (accessed on 18 October 2016).
- Shiokawa, Y.; Misawa, T.; Date, Y.; Kikuchi, J. Application of Market Basket Analysis for the Visualization of Transaction Data Based on Human Lifestyle and Spectroscopic Measurements. Anal. Chem. 2016, 88, 2714–2719. [Google Scholar] [CrossRef] [PubMed]
- Mannina, L.; Sobolev, A.P.; Capitani, D. Applications of NMR metabolomics to the study of foodstuffs: Truffle, kiwifruit, lettuce, and sea bass. Electrophoresis 2012, 33, 2290–2313. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.S. NMR-based metabolomics in wine science. Magn. Reson. Chem. 2011, 49 (Suppl. S1), S13–S21. [Google Scholar] [CrossRef] [PubMed]
- Wei, F.; Ito, K.; Sakata, K.; Date, Y.; Kikuchi, J. Pretreatment and integrated analysis of spectral data reveal seaweed similarities based on chemical diversity. Anal. Chem. 2015, 87, 2819–2826. [Google Scholar] [CrossRef] [PubMed]
- Spraul, M.; Schutz, B.; Humpfer, E.; Mortter, M.; Schafer, H.; Koswig, S.; Rinke, P. Mixture analysis by NMR as applied to fruit juice quality control. Magn. Reson. Chem. 2009, 47 (Suppl. S1), S130–S137. [Google Scholar] [CrossRef] [PubMed]
- Shumilina, E.; Ciampa, A.; Capozzi, F.; Rustad, T.; Dikiy, A. NMR approach for monitoring post-mortem changes in Atlantic salmon fillets stored at 0 and 4 degrees C. Food Chem. 2015, 184, 12–22. [Google Scholar] [CrossRef] [PubMed]
- Southam, A.D.; Easton, J.M.; Stentiford, G.D.; Ludwig, C.; Arvanitis, T.N.; Viant, M.R. Metabolic Changes in Flatfish Hepatic Tumours Revealed by NMR-Based Metabolomics and Metabolic Correlation Networks. J. Proteome Res. 2008, 7, 5277–5285. [Google Scholar] [CrossRef] [PubMed]
- The R Project for Statistical Computing. Available online: https://www.r-project.org/ (accessed on 18 October 2016).
- Date, Y.; Iikura, T.; Yamazawa, A.; Moriya, S.; Kikuchi, J. Metabolic sequences of anaerobic fermentation on glucose-based feeding substrates based on correlation analyses of microbial and metabolite profiling. J. Proteome Res. 2012, 11, 5602–5610. [Google Scholar] [CrossRef] [PubMed]
- Date, Y.; Sakata, K.; Kikuchi, J. Chemical profiling of complex biochemical mixtures from various seaweeds. Polym. J. 2012, 44, 888–894. [Google Scholar] [CrossRef]
- Google Charts. Available online: https://developers.google.com/chart/ (accessed on 18 October 2016).
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chikayama, E.; Yamashina, R.; Komatsu, K.; Tsuboi, Y.; Sakata, K.; Kikuchi, J.; Sekiyama, Y. FoodPro: A Web-Based Tool for Evaluating Covariance and Correlation NMR Spectra Associated with Food Processes. Metabolites 2016, 6, 36. https://doi.org/10.3390/metabo6040036
Chikayama E, Yamashina R, Komatsu K, Tsuboi Y, Sakata K, Kikuchi J, Sekiyama Y. FoodPro: A Web-Based Tool for Evaluating Covariance and Correlation NMR Spectra Associated with Food Processes. Metabolites. 2016; 6(4):36. https://doi.org/10.3390/metabo6040036
Chicago/Turabian StyleChikayama, Eisuke, Ryo Yamashina, Keiko Komatsu, Yuuri Tsuboi, Kenji Sakata, Jun Kikuchi, and Yasuyo Sekiyama. 2016. "FoodPro: A Web-Based Tool for Evaluating Covariance and Correlation NMR Spectra Associated with Food Processes" Metabolites 6, no. 4: 36. https://doi.org/10.3390/metabo6040036
APA StyleChikayama, E., Yamashina, R., Komatsu, K., Tsuboi, Y., Sakata, K., Kikuchi, J., & Sekiyama, Y. (2016). FoodPro: A Web-Based Tool for Evaluating Covariance and Correlation NMR Spectra Associated with Food Processes. Metabolites, 6(4), 36. https://doi.org/10.3390/metabo6040036






