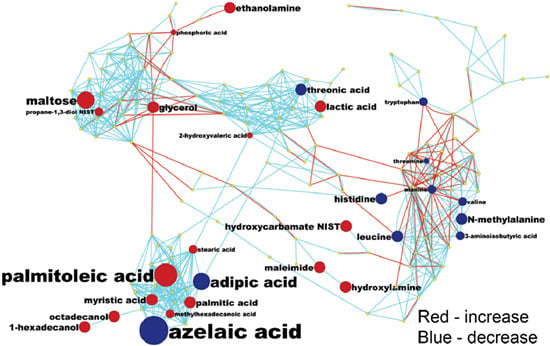
Systemic Metabolomic Changes in Blood Samples of Lung Cancer Patients Identified by Gas Chromatography Time-of-Flight Mass Spectrometry
Abstract
:1. Introduction
2. Results and Discussion
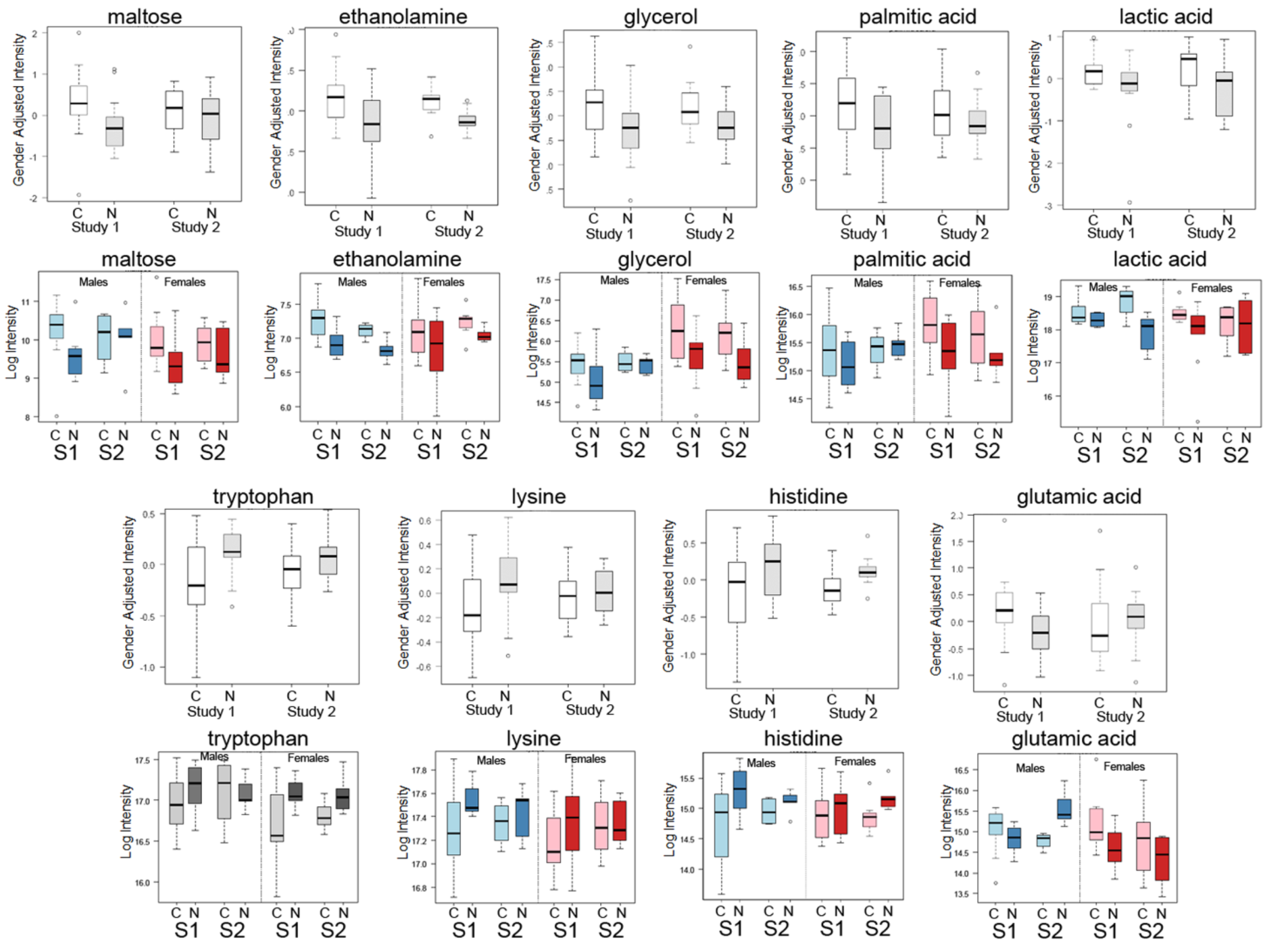
2.1. Differential Analysis Results of Cancer Cases and Controls
| Type and Stage | Female | Male | Total | Smoking History | ||
|---|---|---|---|---|---|---|
| (A) Study 1 (38 Samples) | ||||||
| Lung cancer | NSCLC adenocarcinoma stage unknown | 8 | 10 | 18 | Current or former smokers | |
| Control | 12 | 8 | 20 | Current or former smokers | ||
| Average age (cancer) | 62 (SE 2.38) (range 53–72) | 67 (SE 3.66) (range 50–85) | ||||
| Average age (control) | 64 (SE 2.71) (range 49–80) | 66 (SE 2.65) (range 58–82) | ||||
| (B) Study 2 (22 Samples) | ||||||
| Lung cancer | 7 | 4 | 11 | |||
| NSCLC Stage I-IIB | 1 | 1 | 1 former smoker | |||
| NSCLC Stage IIIA-IV | 2 | 2 | 4 | 1 never smoker 1 former smoker | ||
| SCLC | 3 | 3 | 1 unknown, 1 former smoker 1 current smoker | |||
| Mesothelioma | 1 | 1 | 1 former smoker | |||
| Secondary 2nd metastasis to lung | 1 | 1 | 1 former smoker | |||
| other | 1 | 1 | 1 former smoker | |||
| Control | 6 | 5 | 11 | unknown | ||
| Average age (cancer) | 67 (SE 4.2) (range 47–76) | 67 (SE 2.66) (range 61–73) | 11 | |||
| Average age (control) | 54 (SE 2.64) (range 44–61) | 69 (SE 3.79) (range 61–83) | 11 | |||
| Metabolite | Mean Cancer | Mean Control | Fold (Cancer/Control) | Raw p-Value |
|---|---|---|---|---|
| (A) Study 1 (FHCRC) | ||||
| Maltose | 1298 | 780 | 1.664 | 0.013 |
| Ethanolamine | 156,214 | 123,699 | 1.263 | 0.016 |
| Glycerol | 66,463 | 49,062 | 1.355 | 0.023 |
| Palmitic acid | 53,763 | 41,293 | 1.302 | 0.047 |
| Lactic acid | 380,753 | 301,909 | 1.261 | 0.055 |
| Tryptophan | 121,513 | 143,383 | 0.847 | 0.005 |
| Lysine | 159,156 | 179,325 | 0.888 | 0.042 |
| Histidine | 30,526 | 37,025 | 0.824 | 0.036 |
| Glutamicacid | 39,179 | 27,794 | 1.410 | 0.026 |
| (B) Study 2 (UC Davis) | ||||
| Maltose | 1061 | 989 | 1.074 | 0.819 |
| Ethanolamine | 150,655 | 127,546 | 1.181 | 0.006 |
| Glycerol | 67,557 | 47,052 | 1.436 | 0.315 |
| Palmitic acid | 50,740 | 43,659 | 1.162 | 0.797 |
| Lactic acid | 381,850 | 296,663 | 1.287 | 0.108 |
| Tryptophan | 126,621 | 139,426 | 0.908 | 0.391 |
| Lysine | 167,528 | 172,015 | 0.974 | 0.636 |
| Histidine | 31,053 | 36,840 | 0.843 | 0.047 |
| Glutamicacid | 31,486 | 34,887 | 0.903 | 0.914 |
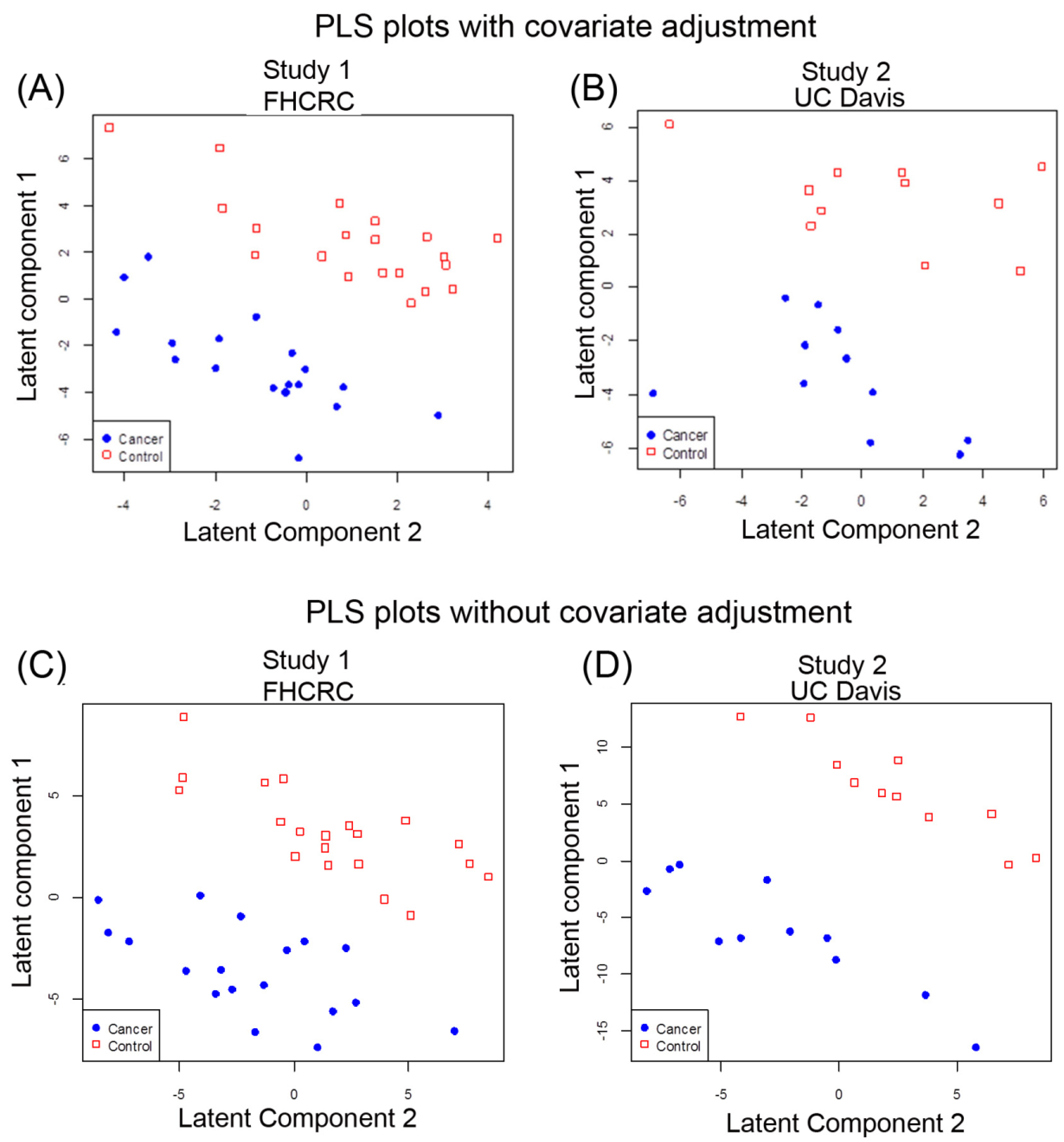
2.2. Multivariate Analysis of Data by PLS-LDA
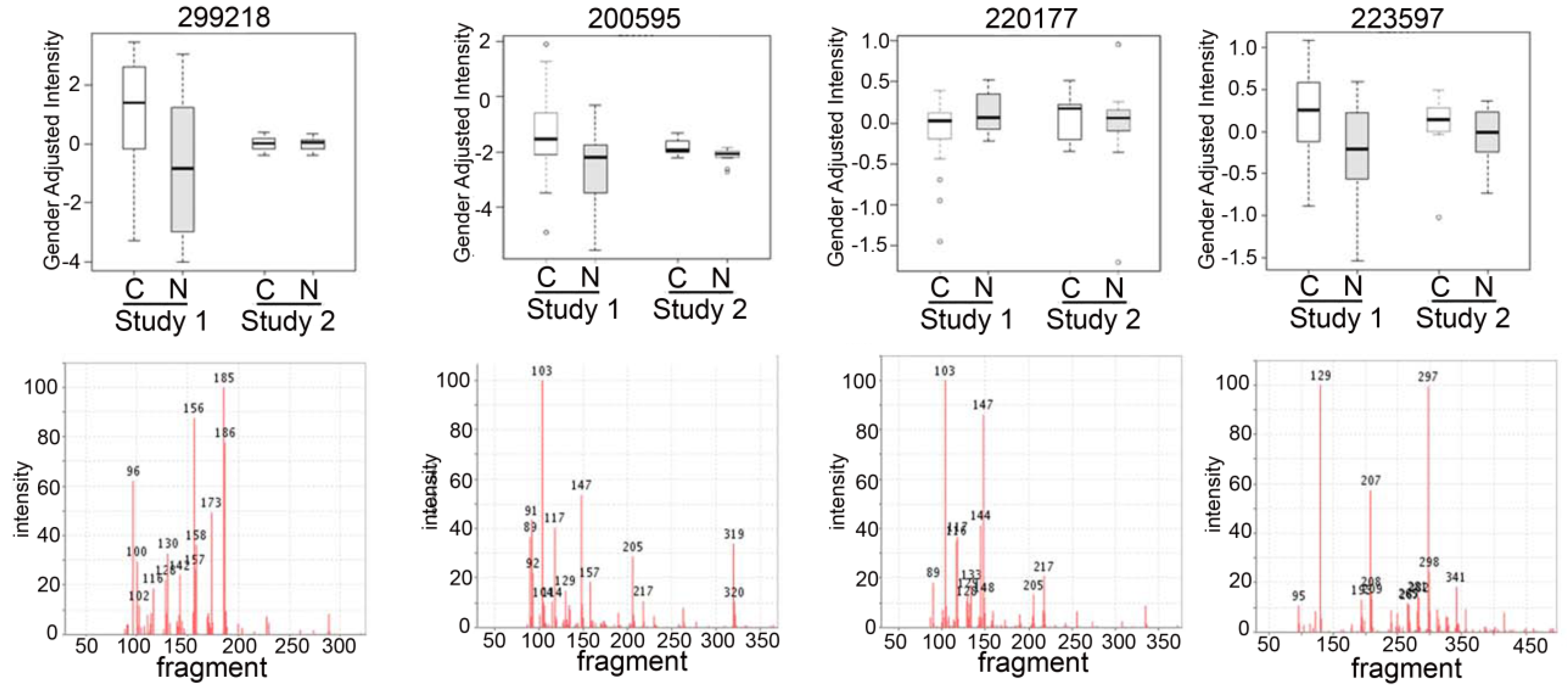
2.3. Detection of Unknown Metabolites
2.4. Discussion
2.4.1. Systemic Metabolic Changes in Blood from Lung Cancer Patients
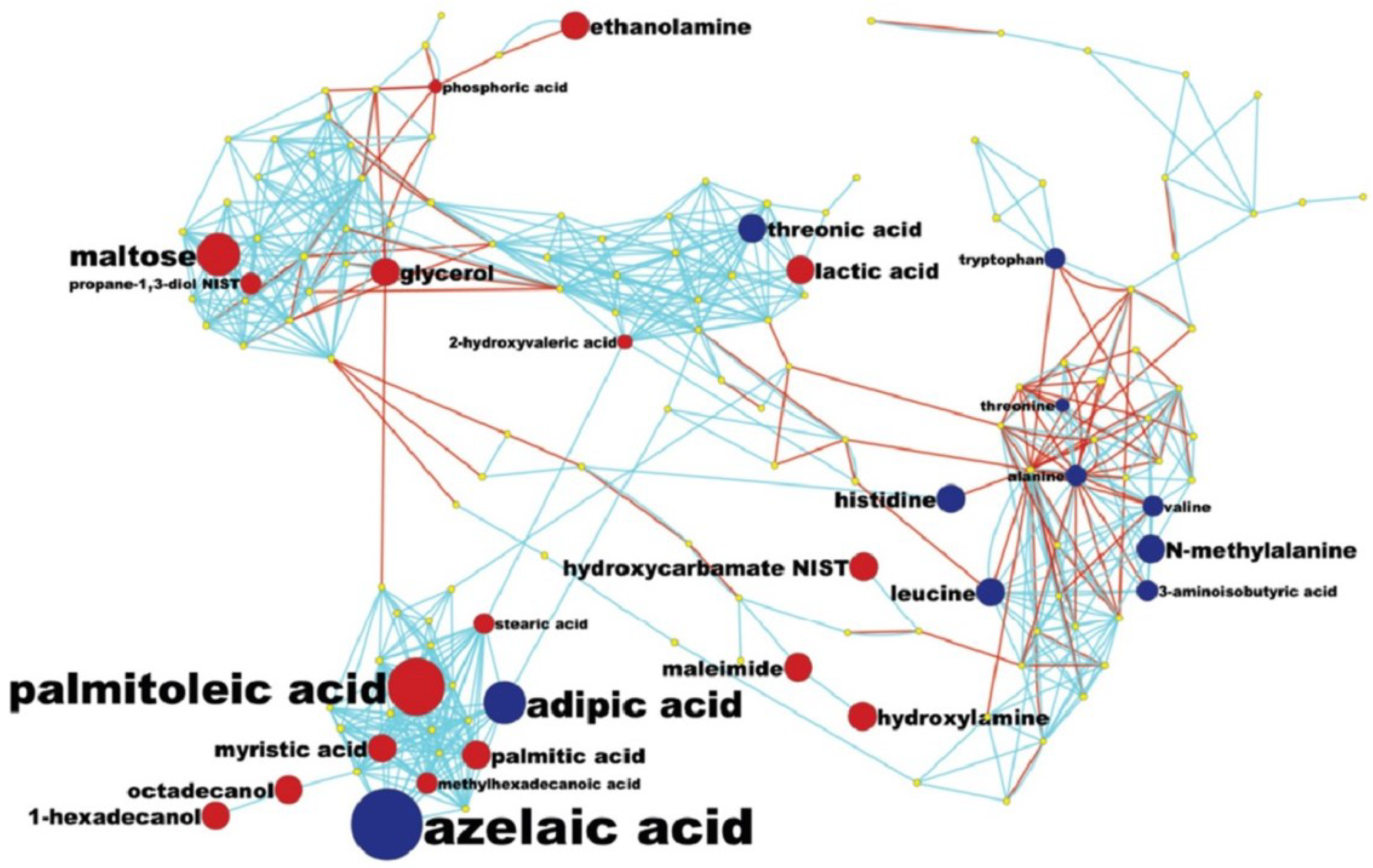
2.4.2. Pathway Analysis and Overall Metabolic Effect on Blood Metabolites
2.4.3. Metabolomic Biomarker Potential for Lung Cancer Detection-Clinical Use
3. Experimental Section
3.1. Patient Samples
3.2. Non-Targeted Metabolomics Analysis by ALEX-CIS-GC/TOF MS
3.3. Raw Data Processing and Chemometrics
3.4. Differential Analysis and Partial Least Squares Analysis
3.5. MetaMapp Mapping of Identified Compounds
4. Conclusions
Acknowledgments
Author Contributions
Supplementary Materials
Conflicts of Interest
References
- Siegel, R.; Ma, J.; Zou, Z.; Jemal, A. Cancer statistics, 2014. CA Cancer J. Clin. 2014, 64, 9–29. [Google Scholar] [CrossRef] [PubMed]
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA Cancer J. Clin. 2011, 61, 69–90. [Google Scholar] [CrossRef] [PubMed]
- Molina, J.R.; Yang, P.; Cassivi, S.D.; Schild, S.E.; Adjei, A.A. Non-small cell lung cancer: Epidemiology, risk factors, treatment, and survivorship. Mayo Clin. Proc. 2008, 83, 584–594. [Google Scholar] [CrossRef] [PubMed]
- National Lung Screening Trial Research, T.; Aberle, D.R.; Berg, C.D.; Black, W.C.; Church, T.R.; Fagerstrom, R.M.; Galen, B.; Gareen, I.F.; Gatsonis, C.; Goldin, J.; et al. The national lung screening trial: Overview and study design. Radiology 2011, 258, 243–253. [Google Scholar]
- Goulart, B.H.; Bensink, M.E.; Mummy, D.G.; Ramsey, S.D. Lung cancer screening with low-dose computed tomography: Costs, national expenditures, and cost-effectiveness. J. Natl. Compr. Canc. Netw. 2012, 10, 267–275. [Google Scholar] [PubMed]
- Louie, A.V.; Rodrigues, G.; Olsthoorn, J.; Palma, D.; Yu, E.; Yaremko, B.; Ahmad, B.; Aivas, I.; Gaede, S. Inter-observer and intra-observer reliability for lung cancer target volume delineation in the 4D-CT era. Radiother. Oncol.: J. Eur. Soc. Ther. Radiol. Oncol. 2010, 95, 166–171. [Google Scholar] [CrossRef]
- Bigbee, W.L.; Gopalakrishnan, V.; Weissfeld, J.L.; Wilson, D.O.; Dacic, S.; Lokshin, A.E.; Siegfried, J.M. A multiplexed serum biomarker immunoassay panel discriminates clinical lung cancer patients from high-risk individuals found to be cancer-free by ct screening. J. Thorac. Oncol. 2012, 7, 698–708. [Google Scholar] [CrossRef] [PubMed]
- Tsay, J.C.; DeCotiis, C.; Greenberg, A.K.; Rom, W.N. Current readings: Blood-based biomarkers for lung cancer. Semin. Thorac. Cardiovasc. Surg. 2013, 25, 328–334. [Google Scholar] [CrossRef] [PubMed]
- Deberardinis, R.J.; Sayed, N.; Ditsworth, D.; Thompson, C.B. Brick by brick: Metabolism and tumor cell growth. Curr. Opin. Genet. Dev. 2008, 18, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Sreekumar, A.; Poisson, L.M.; Rajendiran, T.M.; Khan, A.P.; Cao, Q.; Yu, J.; Laxman, B.; Mehra, R.; Lonigro, R.J.; Li, Y.; et al. Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature 2009, 457, 910–914. [Google Scholar] [CrossRef] [PubMed]
- Denkert, C.; Budczies, J.; Kind, T.; Weichert, W.; Tablack, P.; Sehouli, J.; Niesporek, S.; Konsgen, D.; Dietel, M.; Fiehn, O. Mass spectrometry-based metabolic profiling reveals different metabolite patterns in invasive ovarian carcinomas and ovarian borderline tumors. Cancer Res. 2006, 66, 10795–10804. [Google Scholar] [CrossRef] [PubMed]
- Urayama, S.; Zou, W.; Brooks, K.; Tolstikov, V. Comprehensive mass spectrometry based metabolic profiling of blood plasma reveals potent discriminatory classifiers of pancreatic cancer. Rapid Commun. Mass Spectrom. 2010, 24, 613–620. [Google Scholar] [CrossRef] [PubMed]
- Fan, L.; Zhang, W.; Yin, M.; Zhang, T.; Wu, X.; Zhang, H.; Sun, M.; Li, Z.; Hou, Y.; Zhou, X.; et al. Identification of metabolic biomarkers to diagnose epithelial ovarian cancer using a uplc/qtof/ms platform. Acta Oncol. 2012, 51, 473–479. [Google Scholar] [CrossRef] [PubMed]
- Slupsky, C.M.; Steed, H.; Wells, T.H.; Dabbs, K.; Schepansky, A.; Capstick, V.; Faught, W.; Sawyer, M.B. Urine metabolite analysis offers potential early diagnosis of ovarian and breast cancers. Clin. Cancer Res. 2010, 16, 5835–5841. [Google Scholar] [CrossRef] [PubMed]
- Woo, H.M.; Kim, K.M.; Choi, M.H.; Jung, B.H.; Lee, J.; Kong, G.; Nam, S.J.; Kim, S.; Bai, S.W.; Chung, B.C. Mass spectrometry based metabolomic approaches in urinary biomarker study of women's cancers. Clin. Chim. Acta 2009, 400, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Aa, J.; Xu, J.; Sun, M.; Qian, S.; Cheng, L.; Yang, S.; Shi, R. Metabolomic phenotype of gastric cancer and precancerous stages based on gas chromatography time-of-flight mass spectrometry. J. Gastroenterol. Hepatol. 2011, 26, 1290–1297. [Google Scholar] [CrossRef] [PubMed]
- Maeda, J.; Higashiyama, M.; Imaizumi, A.; Nakayama, T.; Yamamoto, H.; Daimon, T.; Yamakado, M.; Imamura, F.; Kodama, K. Possibility of multivariate function composed of plasma amino acid profiles as a novel screening index for non-small cell lung cancer: A case control study. BMC Cancer 2010, 10, 690. [Google Scholar] [CrossRef] [PubMed]
- Amon, L.M.; Pitteri, S.J.; Li, C.I.; McIntosh, M.; Ladd, J.J.; Disis, M.; Porter, P.; Wong, C.H.; Zhang, Q.; Lampe, P.; et al. Concordant release of glycolysis proteins into the plasma preceding a diagnosis of er+ breast cancer. Cancer Res. 2012, 72, 1935–1942. [Google Scholar] [CrossRef] [PubMed]
- Buckendahl, A.C.; Budczies, J.; Fiehn, O.; Darb-Esfahani, S.; Kind, T.; Noske, A.; Weichert, W.; Sehouli, J.; Braicu, E.; Dietel, M.; et al. Prognostic impact of amp-activated protein kinase expression in ovarian carcinoma: Correlation of protein expression and gc/tof-ms-based metabolomics. Oncol. Rep. 2011, 25, 1005–1012. [Google Scholar] [PubMed]
- Suhre, K.; Shin, S.Y.; Petersen, A.K.; Mohney, R.P.; Meredith, D.; Wagele, B.; Altmaier, E.; CardioGram; Deloukas, P.; Erdmann, J.; et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature 2011, 477, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Serkova, N.J.; Glunde, K. Metabolomics of cancer. Methods Mol. Biol. 2009, 520, 273–295. [Google Scholar] [PubMed]
- Lane, A.N.; Fan, T.W.; Bousamra, M., 2nd; Higashi, R.M.; Yan, J.; Miller, D.M. Stable sotope-resolved metabolomics (SIRM) in cancer research with clinical application to nonsmall cell lung cancer. OMICS 2011, 15, 173–182. [Google Scholar] [CrossRef] [PubMed]
- Mardia, K.V.; Kent, J.T.; Bibby, J.M. Multivariate Analysis; Academic Press: London, UK; New York, NY, USA, 1979; p. 15. [Google Scholar]
- Wold, S. Exponentially weighted moving principal components-analysis and projections to latent structures. Chemometr. Intell. Lab. Syst. 1994, 23, 149–161. [Google Scholar] [CrossRef]
- Kind, T.; Wohlgemuth, G.; Lee do, Y.; Lu, Y.; Palazoglu, M.; Shahbaz, S.; Fiehn, O. Fiehnlib: Mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Anal. Chem. 2009, 81, 10038–10048. [Google Scholar] [CrossRef] [PubMed]
- Kumari, S.; Stevens, D.; Kind, T.; Denkert, C.; Fiehn, O. Applying in-silico retention index and mass spectra matching for identification of unknown metabolites in accurate mass gc-tof mass spectrometry. Anal. Chem. 2011, 83, 5895–5902. [Google Scholar] [CrossRef] [PubMed]
- Kind, T.; Fiehn, O. Seven golden rules for heuristic filtering of molecular formulas obtained by accurate mass spectrometry. BMC Bioinf. 2007, 8, 105. [Google Scholar] [CrossRef]
- Wen, T.; Gao, L.; Wen, Z.; Wu, C.; Tan, C.S.; Toh, W.Z.; Ong, C.N. Exploratory investigation of plasma metabolomics in human lung adenocarcinoma. Mol. Biosyst. 2013, 9, 2370–2378. [Google Scholar] [CrossRef] [PubMed]
- Cascino, A.; Muscaritoli, M.; Cangiano, C.; Conversano, L.; Laviano, A.; Ariemma, S.; Meguid, M.M.; Rossi Fanelli, F. Plasma amino acid imbalance in patients with lung and breast cancer. Anticancer Res. 1995, 15, 507–510. [Google Scholar] [PubMed]
- Miyagi, Y.; Higashiyama, M.; Gochi, A.; Akaike, M.; Ishikawa, T.; Miura, T.; Saruki, N.; Bando, E.; Kimura, H.; Imamura, F.; et al. Plasma free amino acid profiling of five types of cancer patients and its application for early detection. PLoS ONE 2011, 6, e24143. [Google Scholar] [CrossRef] [PubMed]
- Rossi Fanelli, F.; Cangiano, C.; Ceci, F.; Cellerino, R.; Franchi, F.; Menichetti, E.T.; Muscaritoli, M.; Cascino, A. Plasma tryptophan and anorexia in human cancer. Eur. J. Cancer Clin. Oncol. 1986, 22, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Rossi Fanelli, F.; Cangiano, C.; Muscaritoli, M.; Conversano, L.; Torelli, G.F.; Cascino, A. Tumor-induced changes in host metabolism: A possible marker of neoplastic disease. Nutrition 1995, 11, 595–600. [Google Scholar] [PubMed]
- Rocha, C.M.; Carrola, J.; Barros, A.S.; Gil, A.M.; Goodfellow, B.J.; Carreira, I.M.; Bernardo, J.; Gomes, A.; Sousa, V.; Carvalho, L.; et al. Metabolic signatures of lung cancer in biofluids: NMR-based metabonomics of blood plasma. J. Proteome Res. 2011, 10, 4314–4324. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.S.; Pinsky, P.F.; Cramer, D.W.; Ransohoff, D.F.; Hartge, P.; Pfeiffer, R.M.; Urban, N.; Mor, G.; Bast, R.C., Jr.; Moore, L.E.; et al. A framework for evaluating biomarkers for early detection: Validation of biomarker panels for ovarian cancer. Cancer Prev. Res. 2011, 4, 375–383. [Google Scholar] [CrossRef]
- Cramer, D.W.; Bast, R.C., Jr.; Berg, C.D.; Diamandis, E.P.; Godwin, A.K.; Hartge, P.; Lokshin, A.E.; Lu, K.H.; McIntosh, M.W.; Mor, G.; et al. Ovarian cancer biomarker performance in prostate, lung, colorectal, and ovarian cancer screening trial specimens. Cancer Prev. Res. 2011, 4, 365–374. [Google Scholar] [CrossRef]
- Wedge, D.C.; Allwood, J.W.; Dunn, W.; Vaughan, A.A.; Simpson, K.; Brown, M.; Priest, L.; Blackhall, F.H.; Whetton, A.D.; Dive, C.; et al. Is serum or plasma more appropriate for intersubject comparisons in metabolomic studies? An assessment in patients with small-cell lung cancer. Anal. Chem. 2011, 83, 6689–6697. [Google Scholar] [CrossRef] [PubMed]
- Fan, T.W.; Lane, A.N.; Higashi, R.M.; Farag, M.A.; Gao, H.; Bousamra, M.; Miller, D.M. Altered regulation of metabolic pathways in human lung cancer discerned by 13C stable isotope-resolved metabolomics (SIRM). Mol. Cancer 2009, 8, 41. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S.; Sato, Y.; Furumichi, M.; Tanabe, M. Kegg for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 2012, 40, D109–D114. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S.; Furumichi, M.; Tanabe, M.; Hirakawa, M. Kegg for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 2010, 38, D355–D360. [Google Scholar] [CrossRef] [PubMed]
- Barupal, D.K.; Haldiya, P.K.; Wohlgemuth, G.; Kind, T.; Kothari, S.L.; Pinkerton, K.E.; Fiehn, O. Metamapp: Mapping and visualizing metabolomic data by integrating information from biochemical pathways and chemical and mass spectral similarity. BMC Bioinf. 2012, 13, 99. [Google Scholar] [CrossRef]
- Pisters, P.W.; Pearlstone, D.B. Protein and amino acid metabolism in cancer cachexia: Investigative techniques and therapeutic interventions. Crit. Rev. Clin. Lab Sci. 1993, 30, 223–272. [Google Scholar] [CrossRef] [PubMed]
- Sozzi, G.; Boeri, M.; Rossi, M.; Verri, C.; Suatoni, P.; Bravi, F.; Roz, L.; Conte, D.; Grassi, M.; Sverzellati, N.; et al. Clinical utility of a plasma-based mirna signature classifier within computed tomography lung cancer screening: A correlative mild trial study. J. Clin. Oncol. 2014, 32, 768–773. [Google Scholar] [CrossRef] [PubMed]
- Weckwerth, W.; Wenzel, K.; Fiehn, O. Process for the integrated extraction, identification and quantification of metabolites, proteins and rna to reveal their co-regulation in biochemical networks. Proteomics 2004, 4, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Fiehn, O. Extending the breadth of metabolite profiling by gas chromatography coupled to mass spectrometry. Trends Anal. Chem. 2008, 27, 261–269. [Google Scholar] [CrossRef]
- Kind, T.; Tolstikov, V.; Fiehn, O.; Weiss, R.H. A comprehensive urinary metabolomic approach for identifying kidney cancer. Anal. Biochem. 2007, 363, 185–195. [Google Scholar] [CrossRef]
- Scholz, M.; Fiehn, O. Setupx–A public study design database for metabolomic projects. Pac. Symp. Biocomput. 2007, 363, 169–180. [Google Scholar]
- Boulesteix, A.L.; Strimmer, K. Partial least squares: A versatile tool for the analysis of high-dimensional genomic data. Brief Bioinform 2007, 8, 32–44. [Google Scholar] [CrossRef] [PubMed]
- R Development Core Team. The R Stats Package. Available online: https://stat.ethz.ch/R-manual/R-patched/library/stats/html/00Index.html (accessed on 31 March 2015).
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Miyamoto, S.; Taylor, S.L.; Barupal, D.K.; Taguchi, A.; Wohlgemuth, G.; Wikoff, W.R.; Yoneda, K.Y.; Gandara, D.R.; Hanash, S.M.; Kim, K.; et al. Systemic Metabolomic Changes in Blood Samples of Lung Cancer Patients Identified by Gas Chromatography Time-of-Flight Mass Spectrometry. Metabolites 2015, 5, 192-210. https://doi.org/10.3390/metabo5020192
Miyamoto S, Taylor SL, Barupal DK, Taguchi A, Wohlgemuth G, Wikoff WR, Yoneda KY, Gandara DR, Hanash SM, Kim K, et al. Systemic Metabolomic Changes in Blood Samples of Lung Cancer Patients Identified by Gas Chromatography Time-of-Flight Mass Spectrometry. Metabolites. 2015; 5(2):192-210. https://doi.org/10.3390/metabo5020192
Chicago/Turabian StyleMiyamoto, Suzanne, Sandra L. Taylor, Dinesh K. Barupal, Ayumu Taguchi, Gert Wohlgemuth, William R. Wikoff, Ken Y. Yoneda, David R. Gandara, Samir M. Hanash, Kyoungmi Kim, and et al. 2015. "Systemic Metabolomic Changes in Blood Samples of Lung Cancer Patients Identified by Gas Chromatography Time-of-Flight Mass Spectrometry" Metabolites 5, no. 2: 192-210. https://doi.org/10.3390/metabo5020192
APA StyleMiyamoto, S., Taylor, S. L., Barupal, D. K., Taguchi, A., Wohlgemuth, G., Wikoff, W. R., Yoneda, K. Y., Gandara, D. R., Hanash, S. M., Kim, K., & Fiehn, O. (2015). Systemic Metabolomic Changes in Blood Samples of Lung Cancer Patients Identified by Gas Chromatography Time-of-Flight Mass Spectrometry. Metabolites, 5(2), 192-210. https://doi.org/10.3390/metabo5020192