Comparative Chemistry of Aspergillus oryzae (RIB40) and A. flavus (NRRL 3357)
Abstract
:1. Introduction
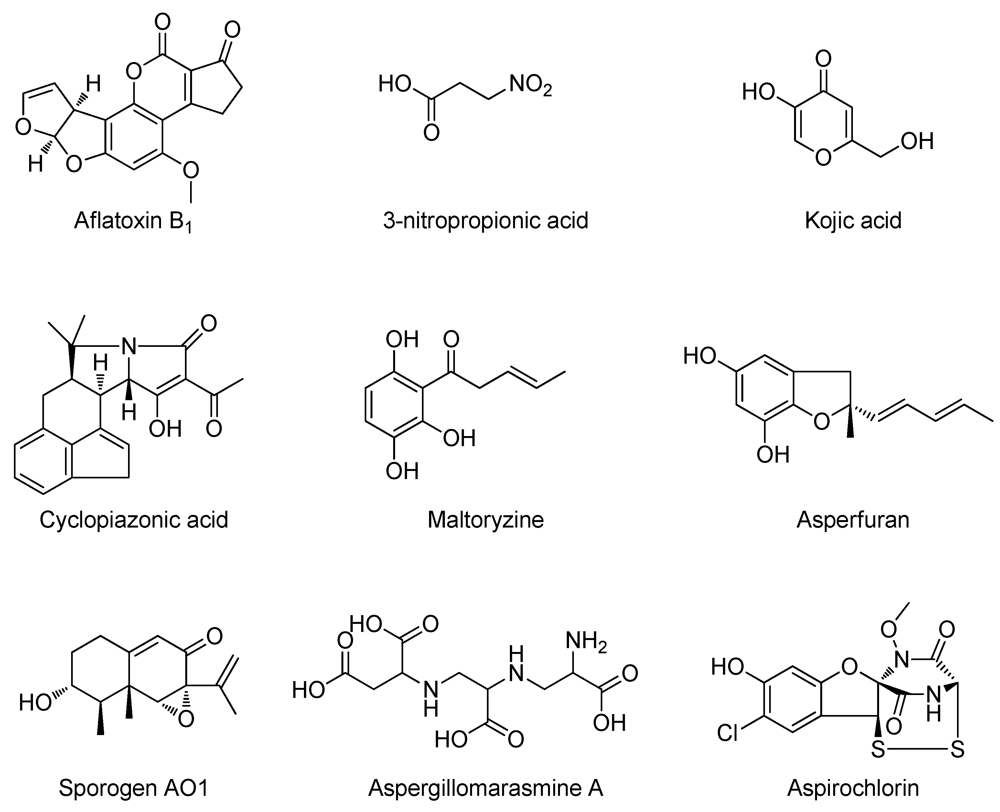
2. Results and Discussion
2.1. De-Replication of A. oryzae RIB40
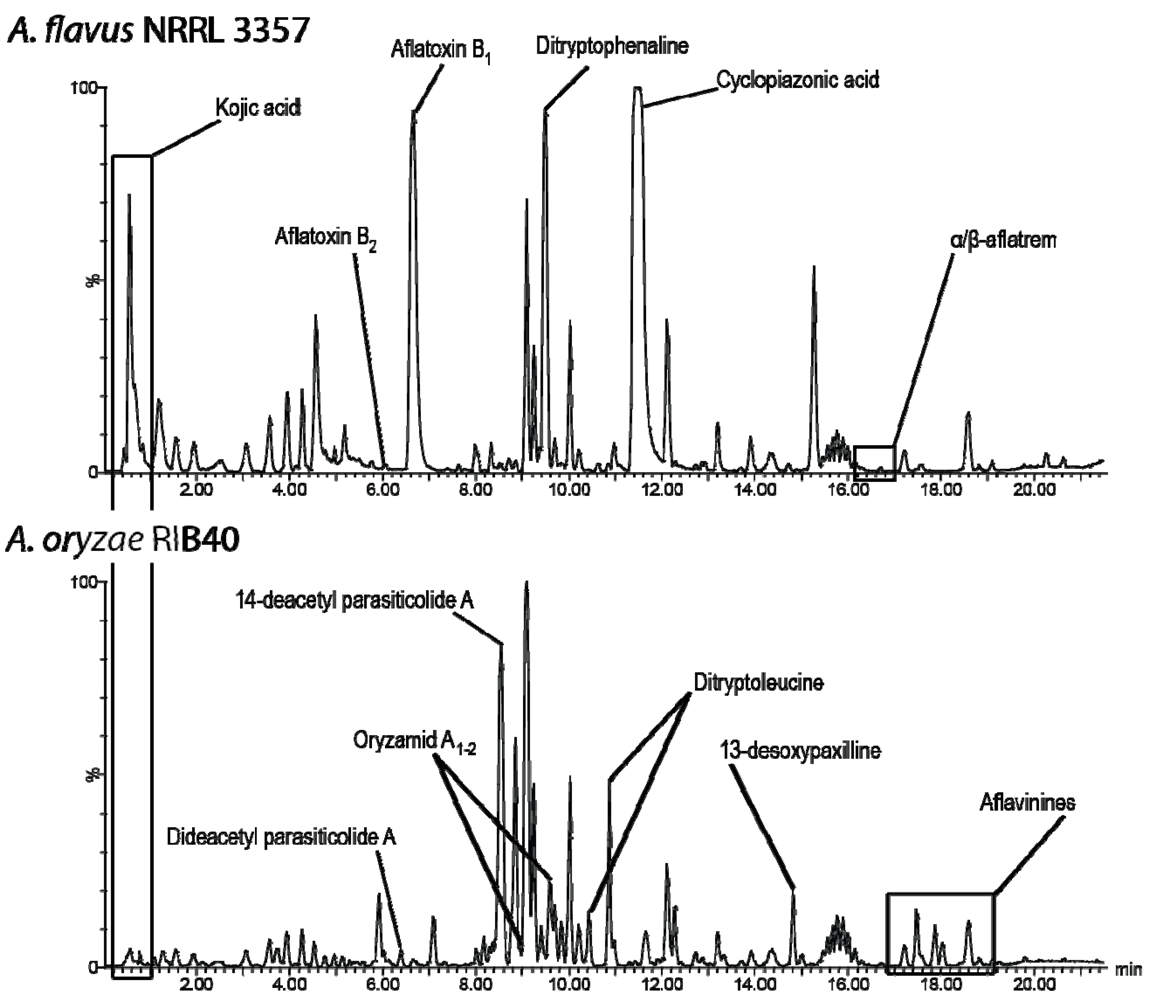
| Metabolite | A. flavus NRRL 3357 | A. oryzae RIB40 |
|---|---|---|
| Kojic acid | + | + |
| Aflatoxin | + | - |
| Aflavinines | (+) | + |
| Aflatrem | + | 13-desoxypaxilline |
| Miyakamides | (+) | Oryzamides * |
| Aspirochlorine | - | + |
| Cyclopiazonic acid | + | - |
| Ditryptophenaline | + | Ditryptoleucine * |
| Parasiticolide A | - | 14-deacetyl parasiticolide A * |
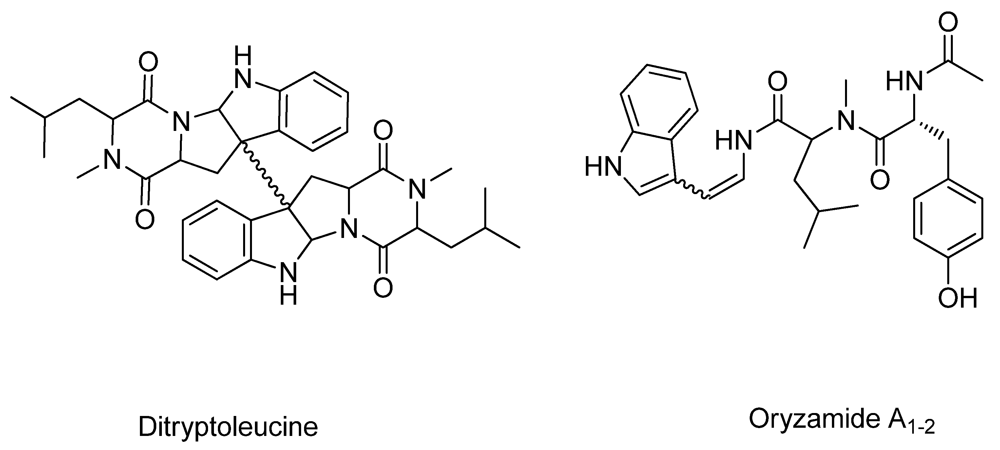
2.2. New Metabolites to A. oryzae RIB40
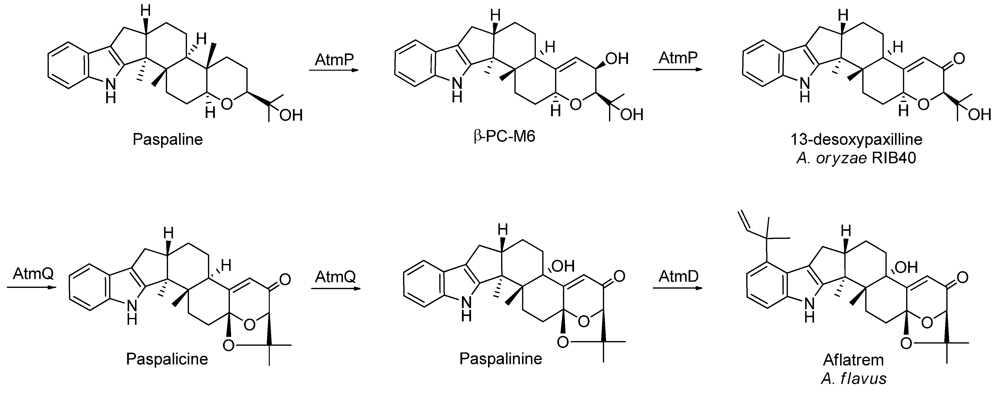
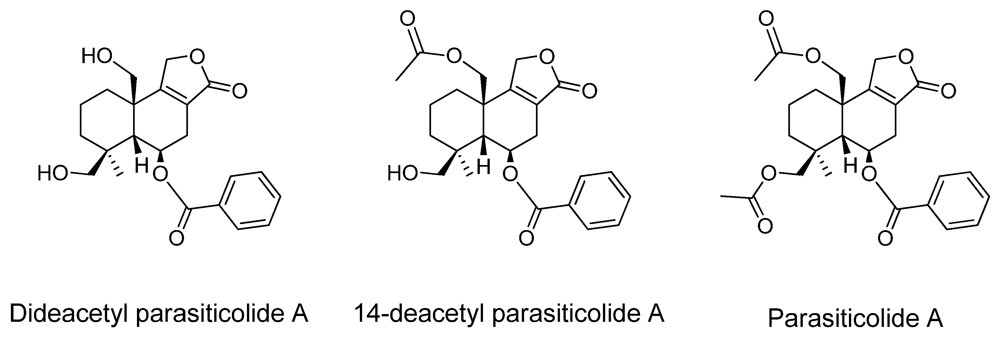
3. Experimental
3.1. Fungal Material and Fermentation
3.2. Extraction and Isolation of Pure Compounds
3.3. Marfeys Method
3.4. Selective Extraction of Sclerotium Metabolites
3.5. HPLC-DAD-TOF Method
3.6. NMR Instrumentation
3.7. MS/MS Method Used for Aflatrem and Parasiticolide Screening
| Compound | Scan event | RT(min) | Ion type | Transition ( m/z) a | Cone (V) | Collision energy (eV) |
|---|---|---|---|---|---|---|
| Paxilline | 1 | 4.0 | Quantifier | 436 → 130 | 2525 | 3030 |
| Qualifier | 436 → 182 | |||||
| Paspalinine | 2 | 4.3 | Quantifier | 434 → 130 | 2525 | 2020 |
| Qualifier | 434 → 376 | |||||
| 13-desoxypaxilline | 3 | 4.8 | Quantifier | 420 → 182 | 2525 | 3030 |
| Qualifier | 420 → 130 | |||||
| Aflatrem | 4 | 5.2 | Quantifier | 502 → 198 | 2525 | 2020 |
| Qualifier | 502 → 445 | |||||
| Paspaline | 5 | 5.5 | Quantifier | 422 → 130 | 2525 | 2020 |
| Qualifier | 422 → 275 | |||||
| Dideacetyl-parasiticolide A | 1 | 7.0 | Quantifier | 387 → 217 | 3030 | 4040 |
| Qualifier | 387 → 189 | |||||
| 14-deacetyl parasiticolide A | 2 | 8.7 | Quantifier | 429 → 217 | 3030 | 4040 |
| Qualifier | 429 → 189 | |||||
| Parasiticolide A | 3 | 10.4 | Quantifier | 488 → 229 | 3030 | 3030 |
| Qualifier | 488 → 247 |
4. Conclusions
Supplementary Materials
Acknowledgments
Conflict of Interest
References and Notes
- Machida, M.; Yamada, O.; Gomi, K. Genomics of Aspergillus oryzae: Learning from the history of koji mold and exploration of its future. DNA Res. 2008, 15, 173–183. [Google Scholar] [CrossRef]
- Punt, P.J.; Biezen, N.V.; Conesa, A.; Albers, A.; Mangnus, J.; van den Hondel, C. Filamentous fungi as cell factories for heterologous protein production. Trends Biotechnol. 2002, 20, 200–206. [Google Scholar] [CrossRef]
- Meyer, V. Genetic engineering of filamentous fungi—Progress, obstacles and future trends. Biotechnol. Adv. 2008, 26, 177–185. [Google Scholar] [CrossRef]
- Fisch, K.M.; Bakeer, W.; Yakasai, A.A.; Song, Z.; Pedrick, J.; Wasil, Z.; Bailey, A.M.; Lazarus, C.M.; Simpson, T.J.; Cox, R.J. Rational domain swaps decipher programming in fungal highly reducing polyketide synthases and resurrect an extinct metabolite. J. Am. Chem. Soc. 2011, 133, 16335–16641. [Google Scholar]
- Geiser, D.M.; Pitt, J.I.; Taylor, J.W. Cryptic speciation and recombination in the aflatoxin-producing fungus Aspergillus flavus. Proc. Natl. Acad. Sci. USA 1998, 95, 388–393. [Google Scholar]
- Geiser, D.M.; Dorner, J.W.; Horn, B.W.; Taylor, J.W. The phylogenetics of mycotoxin and sclerotium production in Aspergillus flavus and Aspergillus oryzae. Fungal Genet. Biol. 2000, 31, 169–179. [Google Scholar] [CrossRef]
- Abe, K.; Gomi, K.; Hasegawa, F.; Machida, M. Impact of Aspergillus oryzae genomics on industrial production of metabolites. Mycopathologia 2006, 162, 143–153. [Google Scholar] [CrossRef]
- Kobayashi, T.; Abe, K.; Asai, K.; Gomi, K.; Juvvadi, P.R.; Kato, M.; Kitamoto, K.; Takeuchi, M. Genomics of Aspergillus oryzae. Biosci. Biotechnol. Biochem. 2007, 71, 646–670. [Google Scholar] [CrossRef]
- Iwasaki, T.; Kosikowski, F.V. Production of beta-nitropropionic acid in foods. J. Food Sci. 1973, 38, 1162–1165. [Google Scholar] [CrossRef]
- Orth, R. Mycotoxins of Aspergillus oryzae strains for use in food-industry as starters and enzyme producing molds. Annales de la Nutrition et de l’Alimentation 1977, 31, 617–624. [Google Scholar]
- Manabe, M; Tanaka, K.; Goto, T.; Matsuura, S. Production capabilities of kojic acid and aflatixin by koji mold. In Toxigenic Fungi—Their Toxins and Health Hazards; Kurata, H., Ueno, Y., Eds.; Elsevier: Amsterdam, The Netherlands, 1984. [Google Scholar]
- Bentley, R. From miso, sake and shoyu to cosmetics: A century of science for kojic acid. Nat. Prod. Rep. 2006, 23, 1046–1062. [Google Scholar] [CrossRef]
- Pfefferle, W.; Anke, H.; Bross, M.; Steffan, B.; Vianden, R.; Steglich, W. Asperfuran, a novel antifungal metabolite from Aspergillus oryzae. J. Antibiot. 1990, 43, 648–654. [Google Scholar] [CrossRef]
- Tanaka, S.; Wada, K.; Katayama, M.; Marumo, S. Isolation of sporogen AO1, a sporogenic substance, from Aspergillus oryzae. Agric. Biol. Chem. 1984, 48, 3189–3191. [Google Scholar] [CrossRef]
- Tanaka, S.; Wada, K.; Marumo, S.; Hattori, H. Structure of sporogen AO1, a sporogenic substance of Aspergillus oryzae. Tetrahedron Lett. 1984, 25, 5907–5910. [Google Scholar] [CrossRef]
- Iizuka, H.; Iida, M. Maltoryzine, a new toxic metabolite produced by a strain of Aspergillus oryzae var. microsporus isolated from poisonous malt sprout. Nature 1962, 196, 681–682. [Google Scholar] [CrossRef]
- Barbier, M.; Vetter, W.; Bogdanov, D.; Lederer, E. Synthese und eigenschafgen eines analogen des lycomarasmins und der aspergillomarasmine. Annalen der Chemie-Justus Liebig 1963, 668, 132. [Google Scholar] [CrossRef]
- Robert, M.; Barbier, M.; Lederer, E.; Roux, L.; Bieman, K.; Vetter, W. Two new natural phytotoxins—Aspergillomarasmines A and B and their identity to lycomarasmine and its derivatives. Bulletin de la Societe Chimique de France 1962, 187–188. [Google Scholar]
- Monti, F.; Ripamonti, F.; Hawser, S.P.; Islam, K. Aspirochlorine: A highly selective and potent inhibitor of fungal protein synthesis. J. Antibiot. 1999, 52, 311–318. [Google Scholar] [CrossRef]
- Sakata, K.; Masago, H.; Sakurai, A.; Takahashi, N. Isolation of aspirochlorine (=antibiotic A30641) posessing a novel dithiodiketopiperazine structure from Aspergillus flavus. Tetrahedron Lett. 1982, 23, 2095–2098. [Google Scholar] [CrossRef]
- Sakata, K.; Kuwatsuka, T.; Sakurai, A.; Takahashi, N.; Tamura, G. Isolation of aspirochlorine (=antibiotic A30641) as a true anti-microbial constituent of the antibiotic, oryzachlorin, from Aspergillus oryzae. Agric. Biol. Chem. 1983, 47, 2673–2674. [Google Scholar] [CrossRef]
- Sakata, K.; Maruyama, M.; Uzawa, J.; Sakurai, A.; Lu, H.S.M.; Clardy, J. Structural revision of aspirochlorine (=antibiotic A30641), a novel epidithiopiperazine-2,5-dione produced by Aspergillus spp. Tetrahedron Lett. 1987, 28, 5607–5610. [Google Scholar]
- Klausmeyer, P.; McCloud, T.G.; Tucker, K.D.; Cardellina, J.H.; Shoemaker, R.H. Aspirochlorine class compounds from Aspergillus flavus inhibit azole-resistant Candida albicans. J. Nat. Prod. 2005, 68, 1300–1302. [Google Scholar] [CrossRef]
- Barbesgaard, P.; Heldt-Hansen, H.P.; Diderichsen, B. On the safety of Aspergillus oryzae: A review. Appl. Microbiol. Biotechnol. 1992, 36, 569–572. [Google Scholar]
- Tanaka, K.; Goto, T.; Manabe, M.; Matsuura, S. Traditional Japanese fermented foods free from mycotoxin contamination. Jpn. Agric. Res. Quart. 2002, 36, 45–50. [Google Scholar]
- Varga, J.; Frisvad, J.C.; Samson, R.A. Two new aflatoxin producing species, and an overview of Aspergillus section Flavi. Stud. Mycol. 2011, 69, 57–80. [Google Scholar] [CrossRef]
- Rokas, A.; Payne, G.; Fedorova, N.D.; Baker, S.E.; Machida, M.; Yu, J.; Georgianna, D.R.; Dean, R.A.; Bhatnagar, D.; Cleveland, T.E. What can comparative genomics tell us about species concepts in the genus Aspergillus? Mycol. 2007, 59, 11–17. [Google Scholar]
- Machida, M.; Asai, K.; Sano, M.; Tanaka, T.; Kumagai, T.; Terai., G.; Kusumoto, K.-I.; Arima, T.; Akita, O.; Kashiwagi, Y.; et al. Genome sequencing and analysis of Aspergillus oryzae. Nature 2005, 438, 1157–1161. [Google Scholar] [CrossRef]
- Machida, M.; Terabayashi, Y.; Sano, M.; Yamane, N.; Tamano, K.; Payne, G.A.; Yu, J.; Cleveland, T.E.; Nierman, W.C. Genomics of industrial aspergilli and comparison with toxigenic relatives. Food Addit. Contam. Part A 2008, 25, 1147–1151. [Google Scholar] [CrossRef]
- Payne, G.A.; Nierman, W.C.; Wortman, J.R.; Pritchard, B.L.; Brown, D.; Dean, R.A.; Bhatnagar, D.; Cleveland, T.E.; Machida, M.; Yu, J. Whole genome comparison of Aspergillus flavus and A. oryzae. Med. Mycol. 2006, 44, S9–S11. [Google Scholar] [CrossRef]
- Yu, J.; Payne, G.A.; Nierman, W.C.; Machida, M.; Bennett, J.W.; Campbell, B.C.; Robens, J.F.; Bhatnagar, D.; Dean, R.A.; Cleveland, T.E. Aspergillus flavus genomics as a tool for studying the mechanism of aflatoxin formation. Food Addit. Contam. Part A 2008, 25, 1152–1157. [Google Scholar] [CrossRef]
- Laatsch, H. AntiBase 2010. Available online: http://www.wiley-vch.de/stmdata/antibase2010.php (accessed on 27 December 2011).
- Cleveland, T.E.; Yu, J.; Fedorova, N.; Bhatnagar, D.; Payne, G.A.; Nierman, W.C.; Bennett, J.W. Potential of Aspergillus flavus genomics for applications in biotechnology. Trends Biotechnol. 2009, 27, 151–157. [Google Scholar] [CrossRef]
- Lee, Y.-H.; Tominaga, M.; Hayashi, R.; Sakamoto, K.; Yamada, O.; Akita, O. Aspergillus oryzae strains with a large deletion of the aflatoxin biosynthetic homologous gene cluster differentiated by chromosomal breakage. Appl. Microbiol. Biotechnol. 2006, 72, 339–345. [Google Scholar] [CrossRef]
- Tominaga, M.; Lee, Y.H.; Hayashi, R.; Suzuki, Y.; Yamada, O.; Sakamoto, K.; Gotoh, K.; Akita, O. Molecular analysis of an inactive aflatoxin biosynthesis gene cluster in Aspergillus oryzae RIB strains. Appl. Environ. Microbiol. 2006, 72, 484–490. [Google Scholar]
- Tokuoka, M.; Seshime, Y.; Fujii, I.; Kitamoto, K.; Takahashi, T.; Koyama, Y. Identification of a novel polyketide synthase-nonribosomal peptide synthetase (PKS-NRPS) gene required for the biosynthesis of cyclopiazonic acid in Aspergillus oryzae. Fungal Genet. Biol. 2008, 45, 1608–1615. [Google Scholar] [CrossRef]
- Chang, P.-K.; Horn, B.W.; Dorner, J.W. Clustered genes involved in cyclopiazonic acid production are next to the aflatoxin biosynthesis gene cluster in Aspergillus flavus. Fungal Genet. Biol. 2009, 46, 176–182. [Google Scholar] [CrossRef]
- Nicholson, M.J.; Koulman, A.; Monahan, B.J.; Pritchard, B.L.; Payne, G.A.; Scott, B. Identification of two aflatrem biosynthetic gene loci in Aspergillus flavus and metabolic engineering in Penicillium paxilli to elucidate gene function. Appl. Environ. Microbiol. 2009, 75, 7469–7481. [Google Scholar]
- Shwab, E.K.; Bok, J.W.; Tribus, M.; Galehr, J.; Graessle, S.; Keller, N.P. Histone deacetylase activity regulates chemical diversity in Aspergillus. Eukaryot. Cell 2007, 6, 1656–1664. [Google Scholar] [CrossRef]
- Shwab, E.K.; Keller, N.P. Regulation of secondary metabolite production in filamentous ascomycetes. Mycol. Res. 2008, 112, 225–230. [Google Scholar] [CrossRef]
- Williams, R.B.; Henrikson, J.C.; Hoover, A.R.; Lee, A.E.; Cichewicz, R.H. Epigenetic remodeling of the fungal secondary metabolome. Org. Biomol. Chem. 2008, 6, 1895–1897. [Google Scholar] [CrossRef]
- Henrikson, J.C.; Hoover, A.R.; Joyner, P.M.; Cichewicz, R.H. A chemical epigenetics approach for engineering the in situ biosynthesis of a cryptic natural product from Aspergillus niger. Org. Biomol. Chem. 2009, 7, 435–438. [Google Scholar]
- Yakasai, A.A.; Davison, J.; Wasil, Z.; Halo, L.M.; Butts, C.P.; Lazarus, C.M.; Bailey, A.M.; Simpson, T.J.; Cox, R.J. Nongenetic reprogramming of a fungal highly reducing polyketide synthase. J. Am. Chem. Soc. 2011, 133, 10990–10998. [Google Scholar]
- Frisvad, J.C.; Thrane, U. Standardized high-performance liquid-chromatography of 182 mycotoxins and other fungal metabolites based on alkylphenone retention indexes and UV-Vis spectra (Diode-Array Detection). J. Chromatogr. A 1987, 404, 195–214. [Google Scholar] [CrossRef]
- Smedsgaard, J. Micro-scale extraction procedure for standardized screening of fungal metabolite production in cultures. J. Chromatogr. A 1997, 760, 264–270. [Google Scholar] [CrossRef]
- Wicklow, D.T.; Cole, R.J. Tremorgenic indole metabolites and aflatoxins in sclerotia of Aspergillus flavus—An evolutionary perspective. Can. J. Bot. 1982, 60, 525–528. [Google Scholar] [CrossRef]
- Gloer, J.B.; Tepaske, M.R.; Sima, J.S.; Wicklow, D.T.; Dowd, P.F. Antiinsectan aflavinine derivatives from the sclerotia of Aspergillus flavus. J. Org. Chem. 1988, 53, 5457–5460. [Google Scholar]
- Gloer, J.B.; Rinderknecht, B.L.; Wicklow, D.T.; Dowd, P.F. Nominine—A new insecticidal indole diterpene from the sclerotia of Aspergillus nomius. J. Org. Chem. 1989, 54, 2530–2532. [Google Scholar]
- Staub, G.M.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. Aspernomine—A cytotoxic antiinsectan metabolite with a novel ring-system from the sclerotia of Aspergillus nomius. J. Am. Chem. Soc. 1992, 114, 1015–1017. [Google Scholar] [CrossRef]
- Staub, G.M.; Gloer, K.B.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. New paspalinine derivatives with antiinsectan activity from the sclerotia of Aspergillus nomius. Tetrahedron Lett. 1993, 34, 2569–2572. [Google Scholar]
- Tepaske, M.R.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. The structure of tubingensin B—A cytotoxic carbazole alkaloid from the sclerotia of Aspergillus tubingensis. Tetrahedron Lett. 1989, 30, 5965–5968. [Google Scholar]
- Tepaske, M.R.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. 3 new aflavinines from the sclerotia of Aspergillus tubingensis. Tetrahedron 1989, 45, 4961–4968. [Google Scholar] [CrossRef]
- Tepaske, M.R.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. Aflavazole—A new antiinsectan carbazole metabolite from the sclerotia of Aspergillus flavus. J. Org. Chem. 1990, 55, 5299–5301. [Google Scholar] [CrossRef]
- Tepaske, M.R.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. Aflavarin and beta-aflatrem—New anti-insectan metabolites from the sclerotia of Aspergillus flavus. J. Nat. Prod. 1992, 55, 1080–1086. [Google Scholar] [CrossRef]
- Raper, K.B.; Fennell, D.I. The genus Aspergillus; Williams & Wilkins: Baltimore, MD, USA, 1965. [Google Scholar]
- Wicklow, D.T.; Mcalpin, C.E.; Yeoh, Q.L. Diversity of Aspergillus oryzae genotypes (RFLP) isolated from traditional soy sauce production within Malaysia and Southeast Asia. Mycoscience 2007, 48, 373–380. [Google Scholar] [CrossRef]
- Jin, F.J.; Takahashi, T.; Utsushikawa, M.; Furukido, T.; Nishida, M.; Ogawa, M.; Tokuoka, M. A trial of minimization of chromosome 7 in Aspergillus oryzae by multiple chromosomal deletions. Mol. Genet. Genomics 2010, 283, 1–12. [Google Scholar]
- Wilson, B.J. Toxins other than aflatoxins produced by Aspergillus flavus. Bacteriol. Rev. 1966, 30, 478–484. [Google Scholar]
- Springer, J.P.; Clardy, J.C.; Wells, J.M.; Cole, R.J.; Kirksey, J.W. Structure of paxilline, a tremorgenic metabolite of Penicillium paxilli Bainier. Tetrahedron Lett. 1975, 2531–2534. [Google Scholar]
- Longland, C.L.; Dyer, J.L.; Michelangeli, F. The mycotoxin paxilline inhibits the cerebellar inositol 1,4, 5-trisphosphate receptor. Eur. J. Pharmacol. 2000, 408, 219–225. [Google Scholar] [CrossRef]
- Bilmen, J.G.; Wootton, L.L.; Michelangeli, F. The mechanism of inhibition of the sarco/endoplasmic reticulum Ca2+ ATPase by paxilline. Arch. Biochem. Biophys. 2002, 406, 55–64. [Google Scholar] [CrossRef]
- Sabater-Vilar, M.; Nijmeijer, S.; Fink-Gremmels, J. Genotoxicity assessment of five tremorgenic mycotoxins (fumitremorgen B, paxilline, penitrem A, verruculogen, and verrucosidin) produced by molds isolated from fermented meats. J. Food Protec. 2003, 66, 2123–2129. [Google Scholar]
- Sheehan, J.J.; Benedetti, B.L.; Barth, A.L. Anticonvulsant effects of the BK-channel antagonist paxilline. Epilepsia 2009, 50, 711–720. [Google Scholar] [CrossRef]
- Wilson, B.J.; Wilson, C.H. Toxin from Aspergillus flavus—Production on food materials of substance causing tremors in mice. Science 1964, 144, 177–178. [Google Scholar]
- Gallagher, R.T.; Wilson, B.J. Aflatrem, the tremorgenic mycotoxin from Aspergillus flavus. Mycopathologia 1978, 66, 183–185. [Google Scholar] [CrossRef]
- Gallagher, R.T.; Clardy, J.C.; Wilson, B.J. Aflatrem, a tremorgenic toxin from Aspergillus flavus. Tetrahedron Lett. 1980, 21, 239–242. [Google Scholar] [CrossRef]
- Cole, R.J.; Dorner, J.W.; Springer, J.P.; Cox, R.H. Indole metabolites from a strain of Aspergillus flavus. J. Agric. Food Chem. 1981, 29, 293–295. [Google Scholar] [CrossRef]
- Steyn, P.S.; Vleggaar, R. Tremorgenic mycotoxins. Fortschritte der Chemie Organischer Naturstoffe 1985, 48, 1–80. [Google Scholar] [CrossRef]
- Bills, G.F.; Giacobbe, R.A.; Lee, S.H.; Peláez, F.; Tkacz, J.S. Tremorgenic mycotoxins, paspalitrem A and C, from a tropical Phomopsis. Mycol. Res. 1992, 96, 977–983. [Google Scholar] [CrossRef]
- Laakso, J.A.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. A new penitrem analog with antiinsectan activity from the sclerotia of Aspergillus sulphureus. J. Agric. Food Chem. 1993, 41, 973–975. [Google Scholar]
- Shiomi, K.; Hatae, K.; Yamaguchi, Y.; Masuma, R.; Tomoda, H.; Kobayashi, S.; Omura, S. New antibiotics miyakamides produced by a fungus. J. Antibiot. 2002, 55, 952–961. [Google Scholar] [CrossRef]
- Fukuyama, K.; Kawai, H.; Tsukihara, T.; Tsukihara, K.; Katsube, Y.; Hamasaki, T.; Hatsuda, Y.; Kuwano, H. Structure-analysis of a bromo derivative of parasiticolide A by X-ray-diffraction method. Bull. Chem. Soc. Jpn. 1975, 48, 2949–2950. [Google Scholar] [CrossRef]
- Ishikawa, Y.; Morimoto, K.; Hamasaki, T. Flavoglaucin, a metabolite of Eurotium chevalieri, its antioxidation and synergism with tocopherol. J. Am. Oil Chem. Soc. 1984, 61, 1864–1868. [Google Scholar] [CrossRef]
- Gould, R.O.; Simpson, T.J.; Walkinshaw, M.D. Isolation and X-ray crystal structures of astellolides A and B, sesquiterpenoid metabolites of Aspergillus variecolor. Tetrahedron Lett. 1981, 22, 1047–1050. [Google Scholar] [CrossRef]
- Pildain, M.B.; Frisvad, J.C.; Vaamonde, G.; Cabral, D.; Varga, J.; Samson, R.A. Two novel aflatoxin-producing Aspergillus species from Argentinean peanuts. Int. J. Syst. Evol. Microbiol. 2008, 58, 725–735. [Google Scholar]
- Ayer, W.A.; Trifonov, L.S. Drimane sesquiterpene lactones from Peniophora polygonia. J. Nat. Prod. 1992, 55, 1454–1461. [Google Scholar] [CrossRef]
- Hamasaki, T.; Kuwano, H.; Isono, K.; Hatsuda, Y.; Fukuyama, K.; Tsukihara, T.; Katsube, Y. New metabolite, parasiticolide A, from Aspergillus parasiticus. Agric. Biol. Chem. 1975, 39, 749–751. [Google Scholar] [CrossRef]
- Springer, J.P.; Buchi, G.; Kobbe, B.; Demain, A.L.; Clardy, J.C. The structure of ditryptophenaline—New metabolite of Aspergillus flavus. Tetrahedron Lett. 1997, 27, 2403–2406. [Google Scholar]
- Barrow, C.J.; Cai, P.; Snyder, J.K.; Sedlock, D.M.; Sun, H.H.; Cooper, R. WIN 64821, a new competitive antagonist to substance-P, isolated from an Aspergillus species—Structure determination and solution conformation. J. Org. Chem. 1993, 58, 6016–6021. [Google Scholar]
- Oleynek, J.J.; Sedlock, D.M.; Barrow, C.J.; Appell, K.C.; Casiano, F.; Haycock, D.; Ward, S.J.; Kaplita, P.; Gillum, A.M. WIN-64821, a novel neurokinin antagonist produced by an Aspergillus sp .2. Biological-activity. J. Antibiot. 1994, 47, 391–398. [Google Scholar]
- Barrow, C.J.; Sedlock, D.M. 1′-(2-Phenyl-ethylene)-ditryptophenaline, a new dimeric diketopiperazine from Aspergillus flavus. J. Nat. Prod. 1994, 57, 1239–1244. [Google Scholar] [CrossRef]
- Movassaghi, M.; Schmidt, M.; Ashenhurst, J. Concise total synthesis of (+)-WIN 64821 and (−)-ditryptophenaline. Angew. Chem. Int. Ed. Engl. 2008, 47, 1485–1487. [Google Scholar]
- Arai, K.; Kimura, K.; Mushiroda, T.; Yamamoto, Y. Structures of fructigenines A and B, new alkaloids isolated from Penicillium fructigenum Takeuchi. Chem. Pharm. Bull. 1989, 37, 2937–2939. [Google Scholar] [CrossRef]
- Frisvad, J.C.; Samson, R.A. Polyphasic taxonomy of Penicillium subgenus Penicillium. A guide to identification of the food and air-borne terverticillate Penicillia and their mycotoxins. Stud. Mycol. 2004, 49, 1–173. [Google Scholar]
- Frisvad, J.C.; Smedsgaard, J.; Larsen, T.O.; Samson, R.A. Mycotoxins, drugs and other extrolites produced by species in Penicillium subgenus Penicillium. Stud. Mycol. 2004, 49, 201–241. [Google Scholar]
- Frisvad, J.C.; Skouboe, P.; Samson, R.A. Taxonomic comparison of three different groups of aflatoxin producers and a new efficient producer of aflatoxin B1, sterigmatocystin and 3-O-methylsterigmatocystin, Aspergillus rambellii sp nov. Syst. Appl. Microbiol. 2005, 28, 442–453. [Google Scholar] [CrossRef]
- Nielsen, K.F.; Mogensen, J.M.; Johansen, M.; Larsen, T.O.; Frisvad, J.C. Review of secondary metabolites and mycotoxins from the Aspergillus niger group. Anal. Bioanal. Chem. 2009, 395, 1225–1242. [Google Scholar] [CrossRef]
- Raper, K.B.; Thom, C. Manual of the Penicillia; Williams and Wilkins: Baltimore, MD, USA, 1949. [Google Scholar]
- Medina, A.; Gonzalez, G.; Saez, J.M. Bee pollen, a substrate that stimulates ochratoxin A production by Aspergillus ochraceus. Syst. Appl. Microbiol. 2004, 27, 261–267. [Google Scholar] [CrossRef]
- Samson, R.A.; Hoekstra, E.S.; Frisvad, J.C. Introduction to Food- and Airborne Fungi. 7; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 2004. [Google Scholar]
Supplementary Files
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Rank, C.; Klejnstrup, M.L.; Petersen, L.M.; Kildgaard, S.; Frisvad, J.C.; Held Gotfredsen, C.; Ostenfeld Larsen, T. Comparative Chemistry of Aspergillus oryzae (RIB40) and A. flavus (NRRL 3357). Metabolites 2012, 2, 39-56. https://doi.org/10.3390/metabo2010039
Rank C, Klejnstrup ML, Petersen LM, Kildgaard S, Frisvad JC, Held Gotfredsen C, Ostenfeld Larsen T. Comparative Chemistry of Aspergillus oryzae (RIB40) and A. flavus (NRRL 3357). Metabolites. 2012; 2(1):39-56. https://doi.org/10.3390/metabo2010039
Chicago/Turabian StyleRank, Christian, Marie Louise Klejnstrup, Lene Maj Petersen, Sara Kildgaard, Jens Christian Frisvad, Charlotte Held Gotfredsen, and Thomas Ostenfeld Larsen. 2012. "Comparative Chemistry of Aspergillus oryzae (RIB40) and A. flavus (NRRL 3357)" Metabolites 2, no. 1: 39-56. https://doi.org/10.3390/metabo2010039
APA StyleRank, C., Klejnstrup, M. L., Petersen, L. M., Kildgaard, S., Frisvad, J. C., Held Gotfredsen, C., & Ostenfeld Larsen, T. (2012). Comparative Chemistry of Aspergillus oryzae (RIB40) and A. flavus (NRRL 3357). Metabolites, 2(1), 39-56. https://doi.org/10.3390/metabo2010039





