Perturbations of Lipid Metabolism Indexed by Lipidomic Biomarkers
Abstract
:1. Introduction
2. The Integrated Lipidome Knowledge Base
3. Development of Lipidomics for Biomedical Applications
4. Why Serum Analysis Can Serve for Investigations of Liver Function
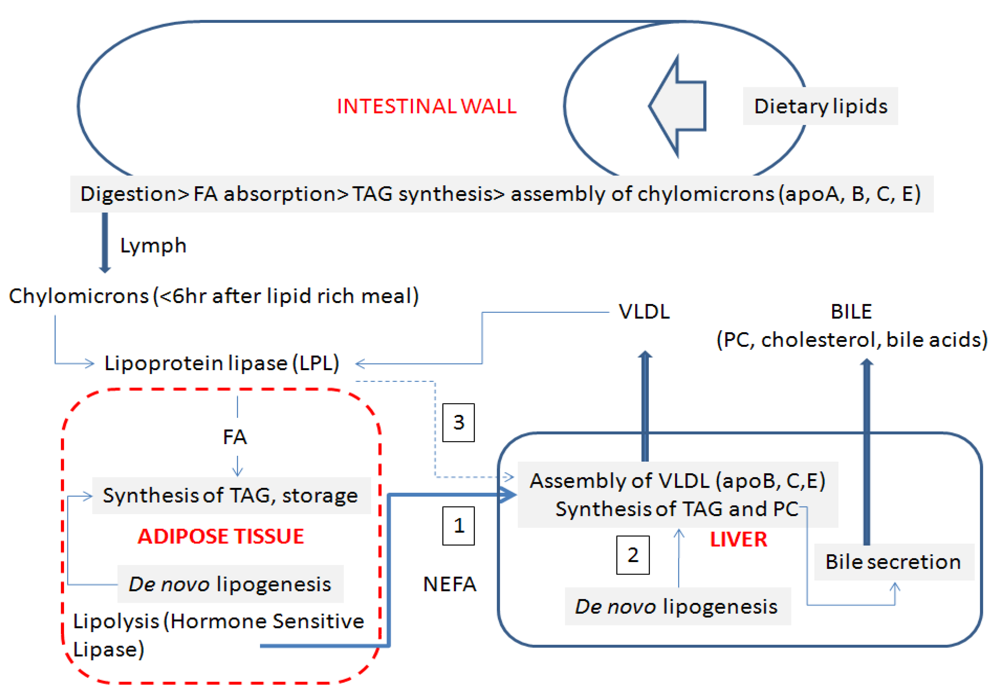
5. Clinical Investigations of Fatty Liver
5.1. Serum Glycerides
5.2. Serum VLDL Reflects a Deficit of Phosphatidylcholine Synthesis in Liver
5.3. Measurement of de Novo Lipogenesis from Serum Lipidomics
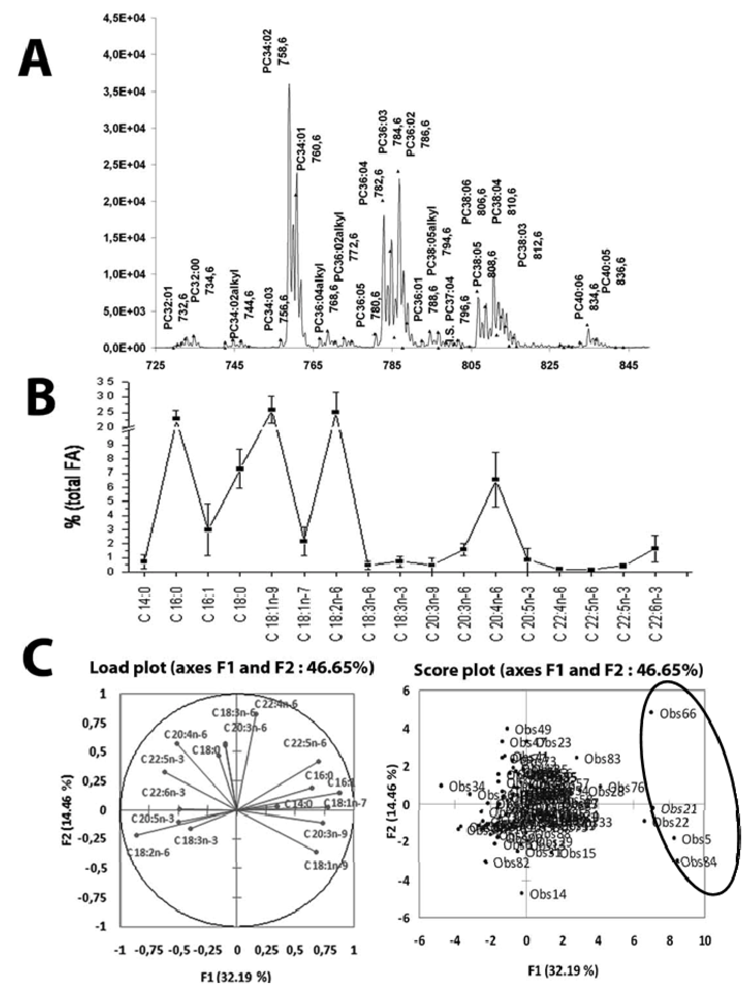
5.3.1. Biomarker Selection
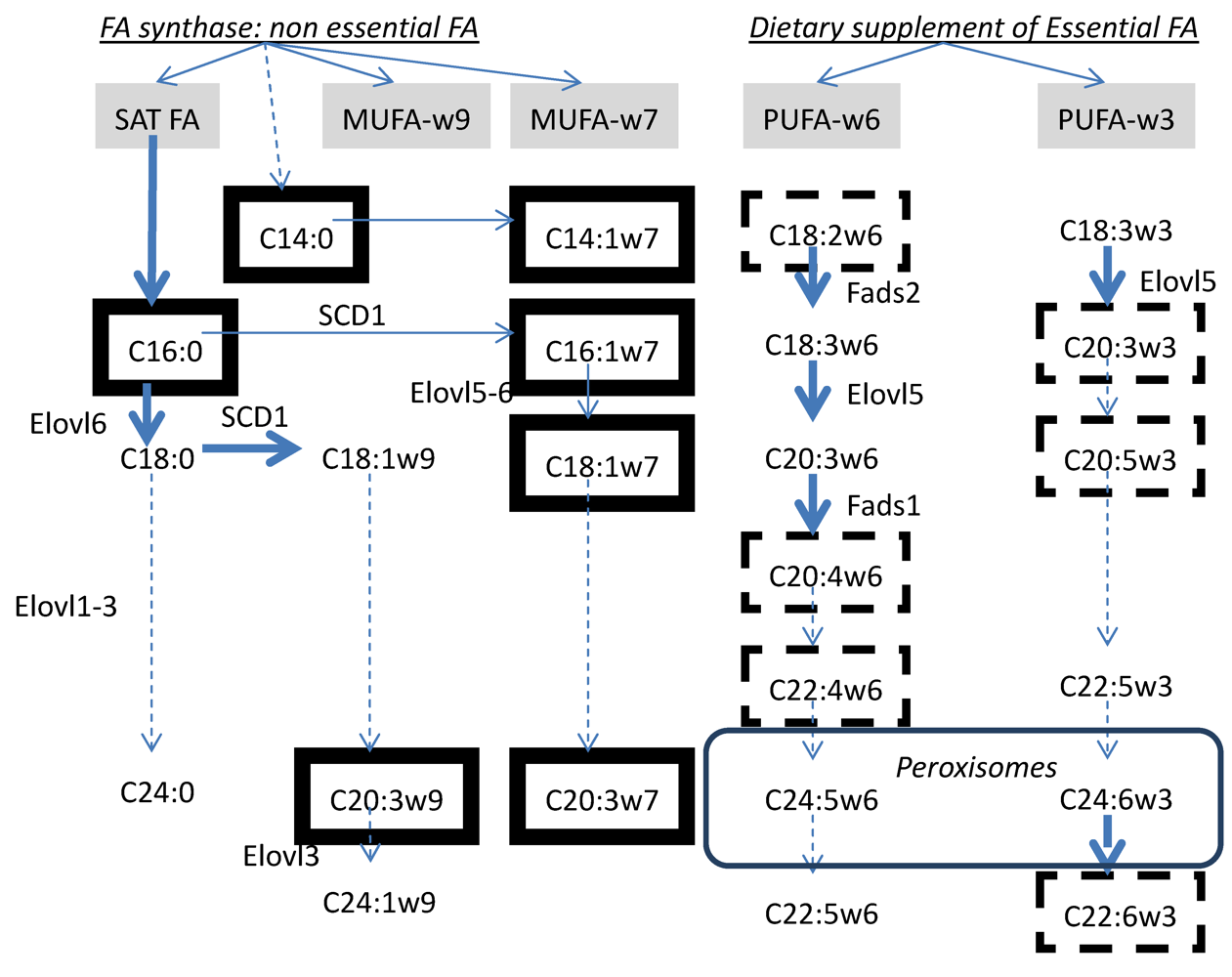
5.3.2. Sources of Fatty Acids Stored in Liver and Secreted via VLDL Lipoprotein as Triglycerides
6. Hepatitis Virus C Infection Perturbs Lipogenesis
7. NASH and NAFLD have Distinct Lipidomic Profiles
8. Lipidomic Application in Polycystic Ovarian Syndrome
9. Conclusions
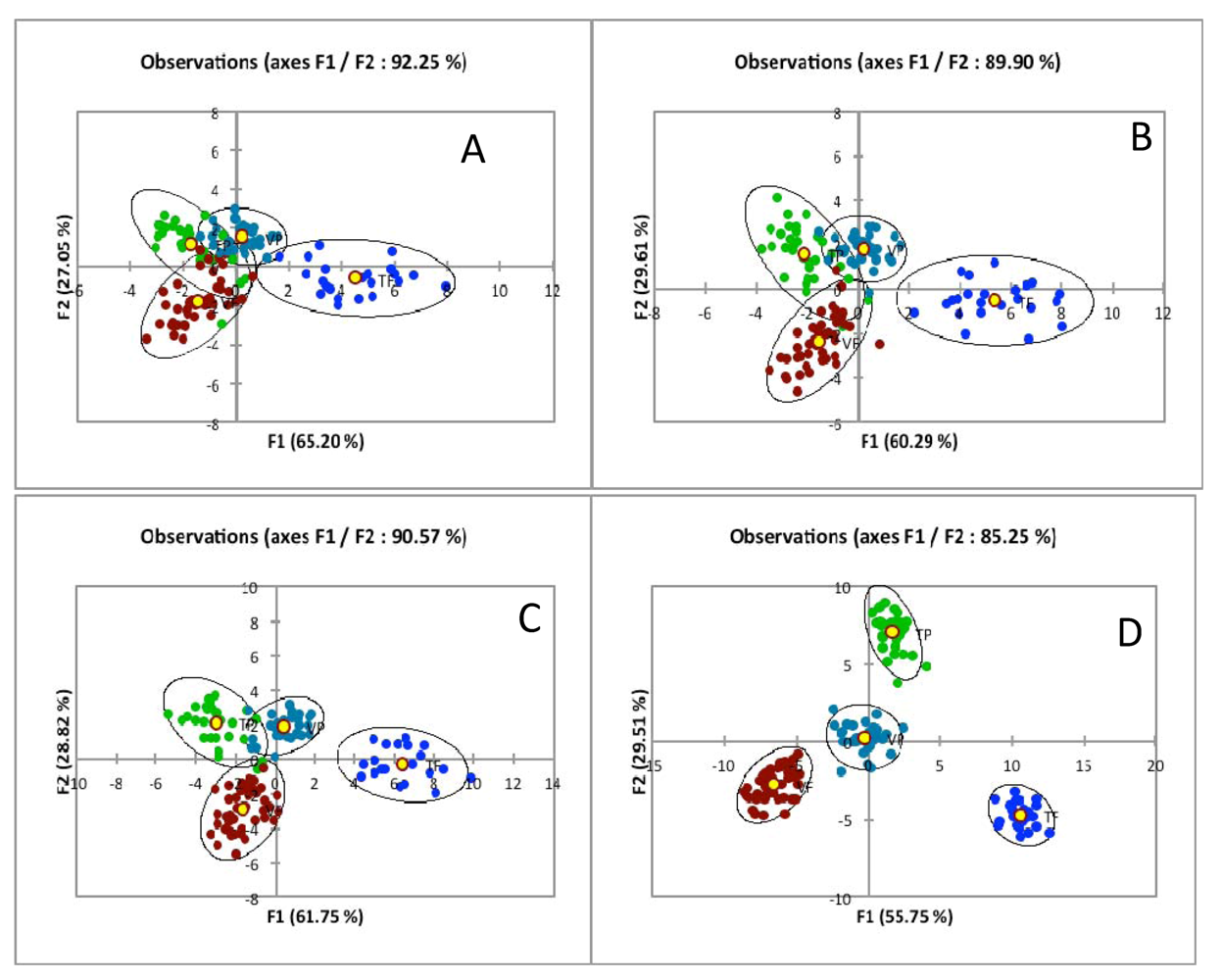
Conflict of Interest
References
- Lei, Z.; Huhman, D.V.; Sumner, L.W. Mass Spectrometry Strategies in Metabolomics. J. Biol. Chem. 2011, 286, 25435–25442. [Google Scholar]
- Schwudke, D.; Schuhmann, K.; Herzog, R.; Bornstein, S.R.; Shevchenko, A. Shotgun Lipidomics on High Resolution Mass Spectrometers. Cold Spring Harbor Perspective. Biol. 2011, 3. [Google Scholar]
- Kanehisa, M.; Goto, S.; Furumichi, M.; Tanabe, M.; Hirakawa, M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 2010, 38, D355–D360. [Google Scholar]
- Quehenberger, O.; Armando, A.M.; Brown, A.H.; Milne, S.B.; Myers, D.S.; Merrill, A.H.; Bandyopadhyay, S.; Jones, K.N.; Kelly, S.; Shaner, R.L.; et al. Lipidomics reveals a remarkable diversity of lipids in human plasma. J. Lipid Res. 2010, 51, 3299–3305. [Google Scholar] [CrossRef]
- Wiest, M.M.; Watkins, S.M. Biomarker discovery using high-dimensional lipid analysis. Curr. Opin. Lipidol. 2007, 18, 181–186. [Google Scholar] [CrossRef]
- Donnelly, K.L.; Smith, C.I.; Schwarzenberg, S.J.; Jessurun, J.; Boldt, M.D.; Parks, E.J. Sources of fatty acids stored in liver and secreted via lipoproteins in patients with nonalcoholic fatty liver disease. J. Clin. Invest. 2005, 115, 1343–1351. [Google Scholar]
- Mazer, N.A.; Carey, M.C. Mathematical model of biliary lipid secretion—A quantitative analysis of physiological and biochemical data from man and other species. J. Lipid Res. 1984, 25, 932–953. [Google Scholar]
- Garcia-Monzon, C.; Lo Iacono, O.; Mayoral, R.; Gonzalez-Rodriguez, A.; Miquilena-Colina, M.E.; Lozano-Rodriguez, T.; Garcia-Pozo, L.; Vargas-Castrillon, J.; Casado, M.; Bosca, L.; Valverde, A.M.; et al. Hepatic insulin resistance is associated with increased apoptosis and fibrogenesis in nonalcoholic steatohepatitis and chronic hepatitis C. J. Hepatol. 2011, 54, 142–152. [Google Scholar] [Green Version]
- Zhang, W.; Kudo, H.; Kawai, K.; Fujisaka, S.; Usui, I.; Sugiyama, T.; Tsukada, K.; Chen, N.; Takahara, T. Tumor necrosis factor-alpha accelerates apoptosis of steatotic hepatocytes from a murine model of non-alcoholic fatty liver disease. Biochem. Biophys. Res. Commun. 2010, 391, 1731–1736. [Google Scholar]
- Ibrahim, S.H.; Kohli, R.; Gores, G.J. Mechanisms of Lipotoxicity in NAFLD and Clinical Implications. J. Pediat. Gastroenterol. Nutr. 2011, 53, 131–140. [Google Scholar]
- Wree, A.; Kahraman, A.; Gerken, G.; Canbay, A. Obesity Affects the Liver—The Link between Adipocytes and Hepatocytes. Digestion 2011, 83, 124–133. [Google Scholar] [CrossRef]
- Puri, P.; Wiest, M.M.; Cheung, O.; Mirshahi, F.; Sargeant, C.; Min, H.-K.; Contos, M.J.; Sterling, R.K.; Fuchs, M.; Zhou, H.; et al. The Plasma Lipidomic Signature of Nonalcoholic Steatohepatitis. Hepatology 2009, 50, 1827–1838. [Google Scholar]
- Hashimoto, E.; Tokushige, K. Prevalence, gender, ethnic variations, and prognosis of NASH. J. Gastroenterol. 2011, 46, 63–69. [Google Scholar] [CrossRef]
- Serfaty, L.; Lemoine, M. Definition and natural history of metabolic steatosis: clinical aspects of NAFLD, NASH and cirrhosis. Diabetes Metab. 2008, 34, 634–637. [Google Scholar] [CrossRef]
- Bellentani, S.; Saccoccio, G.; Masutti, F.; Croce, L.S.; Brandi, G.; Sasso, F.; Cristanini, G.; Tiribelli, C. Prevalence of and risk factors for hepatic steatosis in northern Italy. Ann. Int. Med. 2000, 132, 112–117. [Google Scholar]
- Meikle, P.J.; Christopher, M.J. Lipidomics is providing new insight into the metabolic syndrome and its sequelae. Curr. Opin. Lipidol. 2011, 22, 210–215. [Google Scholar] [CrossRef]
- Jamjute, P.; Ahmad, A.; Ghosh, T.; Banfield, P. Liver function test and pregnancy. J. Matern. Fetal Neonatal Med. 2009, 22, 274–283. [Google Scholar] [CrossRef]
- Boullart, A.C.I.; de Graaf, J.; Stalenhoef, A.F. Serum triglycerides and risk of cardiovascular disease. Biochim. Biophys. Acta 2011. [Google Scholar] [CrossRef]
- Debois, D.; Bralet, M.-P.; Le Naour, F.; Brunelle, A.; Laprevote, O. In situ lipidomic analysis of nonalcoholic fatty liver by cluster TOF-SIMS imaging. Anal. Chem. 2009, 81, 2823–2831. [Google Scholar] [CrossRef]
- Cao, J.; Zhou, Y.; Peng, H.; Huang, X.; Stahler, S.; Suri, V.; Qadri, A.; Gareski, T.; Jones, J.; Hahm, S.; et al. Targeting Acyl-CoA:diacylglycerol acyltransferase 1 (DGAT1) with small molecule inhibitors for the treatment of metabolic diseases. J. Biol. Chem. 2011. [Google Scholar] [CrossRef]
- Cole, L.K.; Vance, J.E.; Vance, D.E. Phosphatidylcholine biosynthesis and lipoprotein metabolism. Biochim. Biophys. Acta 2011. [Google Scholar] [CrossRef]
- Zeisel, S.H. Gene response elements, genetic polymorphisms and epigenetics influence the human dietary requirement for choline. Iubmb. Life 2007, 59, 380–387. [Google Scholar] [CrossRef]
- Leitch, C.A.; Jones, P.J.H. Measurement of human lipogenesis using deuterium incorporation. J. Lipid Res. 1993, 34, 157–163. [Google Scholar]
- Perez-Echarri, N.; Perez-Matute, P.; Marcos-Gomez, B.; Marti, A.; Alfredo Martinez, J.; Moreno-Allaga, M.J. Down-regulation in muscle and liver lipogenic genes: EPA ethyl ester treatment in lean and overweight (high-fat-fed) rats. J. Nutr. Biochem. 2009, 20, 705–714. [Google Scholar] [CrossRef]
- Puri, P.; Baillie, R.A.; Wiest, M.M.; Mirshahi, F.; Choudhury, J.; Cheung, O.; Sargeant, C.; Contos, M.J.; Sanyal, A.J. A lipidomic analysis of nonalcoholic fatty liver disease. Hepatology 2007, 46, 1081–1090. [Google Scholar] [CrossRef]
- Liang, F.; Kume, S.; Koya, D. SIRT1 and insulin resistance. Nat. Rev. Endocrinol. 2009, 5, 367–373. [Google Scholar] [CrossRef]
- Antonio Chavez, J.; Summers, S.A. Lipid oversupply, selective insulin resistance, and lipotoxicity: Molecular mechanisms. Biochim. Et Biophys. Acta-Mol. Cell Biol. Lipids 2010, 1801, 252–265. [Google Scholar]
- Das, S.; Behera, M.K.; Misra, S.; Baliarsihna, A.K. beta-Cell function and insulin resistance in pregnancy and their relation to fetal development. Metab. Syndr. Relat. Disord. 2010, 8, 25–31. [Google Scholar] [CrossRef]
- Skrzypski, J.; Bellenger, S.; Bellenger, J.; Sinclair, A.; Poisson, J.-P.; Tessier, C.; Rialland, M.; Narce, M. Revisiting delta-6 desaturase regulation by C18 unsaturated fatty acids, depending on the nutritional status. Biochimie 2009, 91, 1443–1449. [Google Scholar] [CrossRef]
- Bartenschlager, R.; Penin, F.; Lohmann, V.; Andre, P. Assembly of infectious hepatitis C virus particles. Trends Microbiol. 2011, 19, 95–103. [Google Scholar] [CrossRef]
- Stepanova, M.; Rafiq, N.; Younossi, Z.M. Components of metabolic syndrome are independent predictors of mortality in patients with chronic liver disease: A population-based study. Gut 2010, 59, 1410–1415. [Google Scholar] [CrossRef]
- Syed, G.H.; Amako, Y.; Siddiqui, A. Hepatitis C virus hijacks host lipid metabolism. Trends Endocrinol. Metab. 2010, 21, 33–40. [Google Scholar] [CrossRef]
- Bernsmeier, C.; Heim, M.H. Insulin resistance in chronic hepatitis C: Mechanisms and clinical relevance. Swiss Med. Weekly 2009, 139, 678–684. [Google Scholar]
- Aragones, G.; Alonso-Villaverde, C.; Oliveras-Ferraros, C.; Beltran-Debon, R.; Rull, A.; Rodriguez-Sanabria, F.; Camps, J.; Vazquez Martin, A.; Menendez, J.A.; Joven, J. Infection with HIV and HCV enhances the release of fatty acid synthase into circulation: Evidence for a novel indicator of viral infection. BMC Gastroenterol. 2010, 10, 92. [Google Scholar] [CrossRef]
- Lima-Cabello, E.; Garcia-Mediavilla, M.V.; Miquilena-Colina, M.E.; Vargas-Castrillon, J.; Lozano-Rodriguez, T.; Fernandez-Bermejo, M.; Olcoz, J.L.; Gonzalez-Gallego, J.; Garcia-Monzon, C.; Sanchez-Campos, S. Enhanced expression of pro-inflammatory mediators and liver X-receptor-regulated lipogenic genes in non-alcoholic fatty liver disease and hepatitis C. Clin. Sci. 2011, 120, 239–250. [Google Scholar]
- Banaszewska, B.; Pawelczyk, L.; Spaczynski, R.Z.; Duleba, A.J. Effects of simvastatin and metformin on polycystic ovary syndrome after six months of treatment. J. Clin. Endocrinol. Metab. 2011, 96, 3493–3501. [Google Scholar] [CrossRef]
- Cress, A.; Banaszewska, B.; Pawelczyk, L.; Spaczynski, R.Z.; Duleba, A.J. A Combination of simvastatin and metformin reduces elevated levels of cytokines in women with PCOS. Endocrine Rev. 2010, 31, 1. [Google Scholar] [CrossRef]
- Wolf, C.; Quinn, P.J. Lipidomics in Diagnosis of lipidoses. Sub Cell. Biochem. 2008, 567–588. [Google Scholar] [CrossRef]
- Wolf, C.; Quinn, P.J. Lipidomics: Practical aspects and applications. Prog. Lipid Res. 2008, 47, 15–36. [Google Scholar] [CrossRef]
- Muhlhausler, B.S.; Cook-Johnson, R.; James, M.; Miljkovic, D.; Duthoit, E.; Gibson, R. Opposing effects of omega-3 and omega-6 long chain polyunsaturated Fatty acids on the expression of lipogenic genes in omental and retroperitoneal adipose depots in the rat. J. Nutr. Metab. 2010. [Google Scholar] [CrossRef]
- DeFina, L.F.; Marcoux, L.G.; Devers, S.M.; Cleaver, J.P.; Willis, B.L. Effects of omega-3 supplementation in combination with diet and exercise on weight loss and body composition. Am. J. Clin. Nutr. 2011, 93, 455–462. [Google Scholar] [CrossRef]
- Guillou, H.; Zadravec, D.; Martin, P.G.P.; Jacobsson, A. The key roles of elongases and desaturases in mammalian fatty acid metabolism: Insights from transgenic mice. Prog. Lipid Res. 2010, 49, 186–199. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lamaziere, A.; Wolf, C.; Quinn, P.J. Perturbations of Lipid Metabolism Indexed by Lipidomic Biomarkers. Metabolites 2012, 2, 1-18. https://doi.org/10.3390/metabo2010001
Lamaziere A, Wolf C, Quinn PJ. Perturbations of Lipid Metabolism Indexed by Lipidomic Biomarkers. Metabolites. 2012; 2(1):1-18. https://doi.org/10.3390/metabo2010001
Chicago/Turabian StyleLamaziere, Antonin, Claude Wolf, and Peter J. Quinn. 2012. "Perturbations of Lipid Metabolism Indexed by Lipidomic Biomarkers" Metabolites 2, no. 1: 1-18. https://doi.org/10.3390/metabo2010001
APA StyleLamaziere, A., Wolf, C., & Quinn, P. J. (2012). Perturbations of Lipid Metabolism Indexed by Lipidomic Biomarkers. Metabolites, 2(1), 1-18. https://doi.org/10.3390/metabo2010001



