Metabolomics Changes in Meat and Subcutaneous Fat of Male Cattle Submitted to Fetal Programming
Abstract
:1. Introduction
2. Methodology
2.1. Declaration of Ethics for the Use of Animals in Experimentation
2.2. Experimental Design
2.3. Tissue Collection
2.4. Tissue Homogenization and Metabolite Extraction
2.5. Preparation of Extraction Solvent
2.6. Targeted Metabolomics
2.7. Statistical Analysis
3. Results
3.1. Principal Component Analysis (PCA)
3.2. Heatmaps
3.3. Meat Metabolites
3.4. Subcutaneous Fat Metabolites
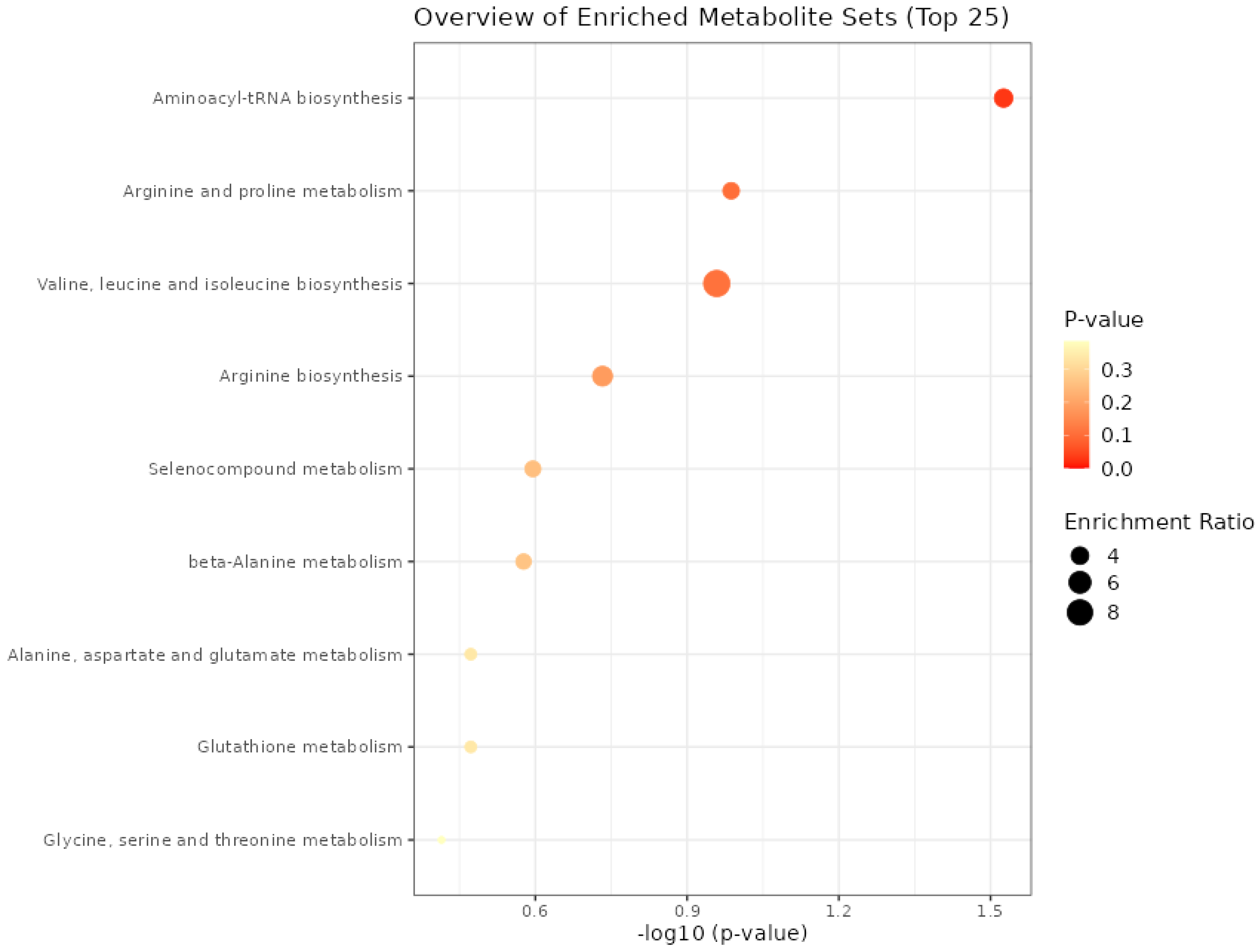
3.5. Enrichment Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nowacka-Woszuk, J. Nutrigenomics in Livestock—Recent Advances. J. Appl. Genet. 2020, 61, 93–103. [Google Scholar] [CrossRef]
- Kauffman, R.G. Meat Composition. In Handbook of Meat and Meat Processing, 2nd ed.; Routledge: London, UK, 2012; pp. 45–61. [Google Scholar] [CrossRef]
- Costa, T.C.; Gionbelli, M.P.; de Souza Duarte, M. Fetal Programming in Ruminant Animals: Understanding the Skeletal Muscle Development to Improve Meat Quality. Anim. Front. 2021, 11, 66–73. [Google Scholar] [CrossRef]
- Long, N.M.; Prado-Cooper, M.J.; Krehbiel, C.R.; Desilva, U.; Wettemann, R.P. Effects of Nutrient Restriction of Bovine Dams during Early Gestation on Postnatal Growth, Carcass and Organ Characteristics, and Gene Expression in Adipose Tissue and Muscle. J. Anim. Sci. 2010, 88, 3251–3261. [Google Scholar] [CrossRef]
- Emwas, A.H.M.; Al-Rifai, N.; Szczepski, K.; Alsuhaymi, S.; Rayyan, S.; Almahasheer, H.; Jaremko, M.; Brennan, L.; Lachowicz, J.I. You Are What You Eat: Application of Metabolomics Approaches to Advance Nutrition Research. Foods 2021, 10, 1249. [Google Scholar] [CrossRef]
- Werner, E.; Heilier, J.F.; Ducruix, C.; Ezan, E.; Junot, C.; Tabet, J.C. Mass Spectrometry for the Identification of the Discriminating Signals from Metabolomics: Current Status and Future Trends. J. Chromatogr. B 2008, 871, 143–163. [Google Scholar] [CrossRef]
- Griffin, J.L. The Cinderella Story of Metabolic Profiling: Does Metabolomics Get to Go to the Functional Genomics Ball? Philos. Trans. R. Soc. B Biol. Sci. 2005, 361, 147–161. [Google Scholar] [CrossRef]
- McCoard, S.A.; Stevens, D.R.; Whitney, T.R. Sustainable Sheep and Goat Production through Strategic Nutritional Management and Advanced Technologies. Anim. Agric. Sustain. Chall. Innov. 2020, 231–246. [Google Scholar] [CrossRef]
- Goldansaz, S.A.; Guo, A.C.; Sajed, T.; Steele, M.A.; Plastow, G.S.; Wishart, D.S. Livestock Metabolomics and the Livestock Metabolome: A Systematic Review. PLoS ONE 2017, 12, e0177675. [Google Scholar] [CrossRef] [PubMed]
- Picard, B.; Lebret, B.; Cassar-Malek, I.; Liaubet, L.; Berri, C.; Le Bihan-Duval, E.; Hocquette, J.F.; Renand, G. Recent Advances in Omic Technologies for Meat Quality Management. Meat Sci. 2015, 109, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Berri, C.; Picard, B.; Lebret, B.; Andueza, D.; Lefèvre, F.; Le Bihan-Duval, E.; Beauclercq, S.; Chartrin, P.; Vautier, A.; Legrand, I.; et al. Predicting the Quality of Meat: Myth or Reality? Foods 2019, 8, 436. [Google Scholar] [CrossRef] [PubMed]
- Carrillo, J.A.; He, Y.; Li, Y.; Liu, J.; Erdman, R.A.; Sonstegard, T.S.; Song, J. Integrated Metabolomic and Transcriptome Analyses Reveal Finishing Forage Affects Metabolic Pathways Related to Beef Quality and Animal Welfare. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Yu, K.; Matzapetakis, M.; Valent, D.; Saco, Y.; De Almeida, A.M.; Terré, M.; Bassols, A. Skeletal Muscle Metabolomics and Blood Biochemistry Analysis Reveal Metabolic Changes Associated with Dietary Amino Acid Supplementation in Dairy Calves. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Chen, C.; Xie, K.; Wang, J.; Pan, Z. Current State of Metabolomics Research in Meat Quality Analysis and Authentication. Foods 2021, 10, 2388. [Google Scholar] [CrossRef] [PubMed]
- Antonelo, D.; Gómez, J.F.M.; Cônsolo, N.R.B.; Beline, M.; Colnago, L.A.; Schilling, W.; Zhang, X.; Suman, S.P.; Gerrard, D.E.; Balieiro, J.C.C.; et al. Metabolites and Metabolic Pathways Correlated With Beef Tenderness. Meat Muscle Biol. 2020, 4, 19–20. [Google Scholar] [CrossRef]
- Zuo, H.; Wang, P.; Guo, Z.; Luo, X.; Zhang, Y.; Mao, Y. Metabolites Analysis on Water-Holding Capacity in Beef Longissimus Lumborum Muscle during Postmortem Aging. Metabolites 2022, 12, 242. [Google Scholar] [CrossRef]
- Bischof, G.; Witte, F.; Terjung, N.; Heinz, V.; Juadjur, A.; Gibis, M. Metabolic, Proteomic and Microbial Changes Postmortem and during Beef Aging. Crit. Rev. Food Sci. Nutr. 2022, 1–34. [Google Scholar] [CrossRef]
- Muroya, S.; Ueda, S.; Komatsu, T.; Miyakawa, T.; Ertbjerg, P. MEATabolomics: Muscle and Meat Metabolomics in Domestic Animals. Metabolites 2020, 10, 188. [Google Scholar] [CrossRef]
- Du, M.; Tong, J.; Zhao, J.; Underwood, K.R.; Zhu, M.; Ford, S.P.; Nathanielsz, P.W. Fetal Programming of Skeletal Muscle Development in Ruminant Animals. J. Anim. Sci. 2010, 88, E51–E60. [Google Scholar] [CrossRef]
- Du, M.; Zhao, J.X.; Yan, X.; Huang, Y.; Nicodemus, L.V.; Yue, W.; Mccormick, R.J.; Zhu, M.J. Fetal Muscle Development, Mesenchymal Multipotent Cell Differentiation, and Associated Signaling Pathways. J. Anim. Sci. 2011, 89, 583–590. [Google Scholar] [CrossRef]
- de Zawadzki, A.; Arrivetti, L.O.R.; Vidal, M.P.; Catai, J.R.; Nassu, R.T.; Tullio, R.R.; Berndt, A.; Oliveira, C.R.; Ferreira, A.G.; Neves-Junior, L.F.; et al. Mate Extract as Feed Additive for Improvement of Beef Quality. Food Res. Int. 2017, 99, 336–347. [Google Scholar] [CrossRef]
- Peng, D.Q.; Kim, S.J.; Lee, H.G. Metabolomics Analyses to Characterize Metabolic Alterations in Korean Native Calves by Oral Vitamin A Supplementation. Sci. Rep. 2020, 10, 8092. [Google Scholar] [CrossRef] [PubMed]
- Gómez, J.F.M.; Cônsolo, N.R.B.; Antonelo, D.S.; Beline, M.; Gagaoua, M.; Higuera-Padilla, A.; Colnago, L.A.; Gerrard, D.E.; Silva, S.L. Impact of Cattle Feeding Strategy on the Beef Metabolome. Metabolites 2022, 12, 640. [Google Scholar] [CrossRef] [PubMed]
- Cônsolo, N.R.B.; Buarque, V.L.M.; Silva, J.; Poleti, M.D.; Barbosa, L.C.G.S.; Higuera-Padilla, A.; Gómez, J.F.M.; Colnago, L.A.; Gerrard, D.E.; Saran Netto, A.; et al. Muscle and Liver Metabolomic Signatures Associated with Residual Feed Intake in Nellore Cattle. Anim. Feed Sci. Technol. 2021, 271, 114757. [Google Scholar] [CrossRef]
- Gandra, J.R.; Pedrini, C.A.; Cônsolo, N.R.B.; Acosta, A.P.; Seno, L.O.; Barbosa, L.C.G.S.; Noia, I.Z.; Buarque, V.L.M.; Padilla, A.R.H.; Colnago, L.A.; et al. Metabolome Fingerprints, Performance and Carcass Quality of Beef Calves Supplemented with Antibiotic Free Additive. Anim. Biotechnol. 2020, 33, 710–722. [Google Scholar] [CrossRef]
- Schalch Junior, F.J.; Polizel, G.H.G.; Cançado, F.A.C.Q.; Fernandes, A.C.; Mortari, I.; Pires, P.R.L.; Fukumasu, H.; Santana, M.H.d.A.; Saran Netto, A. Prenatal Supplementation in Beef Cattle and Its Effects on Plasma Metabolome of Dams and Calves. Metabolites 2022, 12, 347. [Google Scholar] [CrossRef] [PubMed]
- Polizel, G.H.G.; Cançado, F.A.C.Q.; Dias, E.F.F.; Fernandes, A.C.; Cracco, R.C.; Carmona, B.T.; Castellar, H.H.; Poleti, M.D.; de Almeida Santana, M.H. Effects of Different Prenatal Nutrition Strategies on the Liver Metabolome of Bulls and Its Correlation with Body and Liver Weight. Metabolites 2022, 12, 441. [Google Scholar] [CrossRef]
- Martin, D.E.; Jones, A.K.; Pillai, S.M.; Hoffman, M.L.; McFadden, K.K.; Zinn, S.A.; Govoni, K.E.; Reed, S.A. Maternal Restricted- And Overfeeding during Gestation Result in Distinct Lipid and Amino Acid Metabolite Profiles in the Longissimus Muscle of the Offspring. Front. Physiol. 2019, 10, 515. [Google Scholar] [CrossRef]
- Muroya, S.; Zhang, Y.; Kinoshita, A.; Otomaru, K.; Oshima, K.; Gotoh, Y.; Oshima, I.; Sano, M.; Roh, S.; Oe, M.; et al. Maternal Undernutrition during Pregnancy Alters Amino Acid Metabolism and Gene Expression Associated with Energy Metabolism and Angiogenesis in Fetal Calf Muscle. Metabolites 2021, 11, 582. [Google Scholar] [CrossRef]
- National Research Council. Nutrient Requirements of Beef Cattle: Seventh Revised Edition: Update 2000, 7th ed.; The National Academies Press: Washington, DC, USA, 2000. [Google Scholar]
- Ministério da Agricultura e Pecuária—Governo Federal Brasileiro DECRETO No 30.691, de 29 de MARÇO de 1952. Regulamento da Inspeção Industrial e Sanitária de Produtos de Origem Animal. 1952, pp. 1–44. Available online: https://www2.camara.leg.br/legin/fed/decret/1950-1959/decreto-30691-29-marco-1952-339586-normaatualizada-pe.pdf (accessed on 10 December 2023).
- Ludtke, C.B.; Ciocca, J.R.P.; Dandin, T.; Barbalho, P.C.; Vilela, J.A.; Ferrarini, C.; Abate Humanitário de Bovinos. AMA Editora e Treinamento Ltda, Rio de Janeiro. 2012; p. 148. Available online: https://www.gov.br/agricultura/pt-br/assuntos/producao-animal/arquivos-publicacoes-bem-estar-animal/programa-steps-2013-abate-humanitario-de-bovinos.pdf (accessed on 10 December 2023).
- Cracco, R.C.; Ruy, I.M.; Polizel, G.H.G.; Fernandes, A.C.; Furlan, É.; Baldin, G.C.; Santos, G.E.C.; Santana, M.H. de A. Evaluation of Maternal Nutrition Effects in the Lifelong Performance of Male Beef Cattle Offspring. Vet. Sci. 2023, 10, 443. [Google Scholar] [CrossRef]
- Zukunft, S.; Prehn, C.; Röhring, C.; Möller, G.; Hrabě de Angelis, M.; Adamski, J.; Tokarz, J. High-Throughput Extraction and Quantification Method for Targeted Metabolomics in Murine Tissues. Metabolomics 2018, 14, 18. [Google Scholar] [CrossRef]
- Geletu, U.S.; Usmael, M.A.; Mummed, Y.Y.; Ibrahim, A.M. Quality of Cattle Meat and Its Compositional Constituents. Vet. Med. Int. 2021, 2021, 7340495. [Google Scholar] [CrossRef] [PubMed]
- Olaoye, O.A. MiniReview Meat: An Overview of Its Composition, Biochemical Changes and Associated Microbial Agents. Int. Food Res. J. 2011, 18, 877–885. [Google Scholar]
- Kwon, H.; Ford, S.P.; Bazer, F.W.; Spencer, T.E.; Nathanielsz, P.W.; Nijland, M.J.; Hess, B.W.; Wu, G. Maternal Nutrient Restriction Reduces Concentrations of Amino Acids and Polyamines in Ovine Maternal and Fetal Plasma and Fetal Fluids. Biol. Reprod. 2004, 71, 901–908. [Google Scholar] [CrossRef] [PubMed]
- Hellmuth, C.; Uhl, O.; Kirchberg, F.F.; Harder, U.; Peissner, W.; Koletzko, B.; Nathanielsz, P.W. Influence of Moderate Maternal Nutrition Restriction on the Fetal Baboon Metabolome at 0.5 and 0.9 Gestation. Nutr. Metab. Cardiovasc. Dis. 2016, 26, 786–796. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Capillas, C.; Jiménez-Colmenero, F. Biogenic Amines in Meat and Meat Products. Crit. Rev. Food Sci. Nutr. 2010, 44, 489–599. [Google Scholar] [CrossRef] [PubMed]
- Hannun, Y.A.; Obeid, L.M. Principles of Bioactive Lipid Signalling: Lessons from Sphingolipids. Nat. Rev. Mol. Cell Biol. 2008, 9, 139–150. [Google Scholar] [CrossRef]
- Vesper, H.; Schmelz, E.M.; Nikolova-Karakashian, M.N.; Dillehay, D.L.; Lynch, D.V.; Merrill, A.H. Sphingolipids in Food and the Emerging Importance of Sphingolipids to Nutrition. J. Nutr. 1999, 129, 1239–1250. [Google Scholar] [CrossRef]
- Fischbeck, A.; Krüger, M.; Blaas, N.; Humpf, H.U. Analysis of Sphingomyelin in Meat Based on Hydrophilic Interaction Liquid Chromatography Coupled to Electrospray Lonization-Tandem Mass Spectrometry (HILIC-HPLC-ESI-MS/MS). J. Agric. Food Chem. 2009, 57, 9469–9474. [Google Scholar] [CrossRef]
- Muroya, S. An Insight into Farm Animal Skeletal Muscle Metabolism Based on a Metabolomics Approach. Meat Sci. 2023, 195, 108995. [Google Scholar] [CrossRef]
- Yu, Q.; Tian, X.; Shao, L.; Li, X.; Dai, R. Mitochondria Changes and Metabolome Differences of Bovine Longissimus Lumborum and Psoas Major during 24 h Postmortem. Meat Sci. 2020, 166, 108112. [Google Scholar] [CrossRef]
- Nagata, Y.; Partridge, T.A.; Matsuda, R.; Zammit, P.S. Entry of Muscle Satellite Cells into the Cell Cycle Requires Sphingolipid Signaling. J. Cell Biol. 2006, 174, 245–253. [Google Scholar] [CrossRef]
- Jairath, G.; Singh, P.K.; Dabur, R.S.; Rani, M.; Chaudhari, M. Biogenic Amines in Meat and Meat Products and Its Public Health Significance: A Review. J. Food Sci. Technol. 2015, 52, 6835–6846. [Google Scholar] [CrossRef]
- Shalaby, A.R. Significance of Biogenic Amines to Food Safety and Human Health. Food Res. Int. 1996, 29, 675–690. [Google Scholar] [CrossRef]
- Bardócz, S. Polyamines in Food and Their Consequences for Food Quality and Human Health. Trends Food Sci. Technol. 1995, 6, 341–346. [Google Scholar] [CrossRef]
- Halász, A.; Baráth, Á.; Simon-Sarkadi, L.; Holzapfel, W. Biogenic Amines and Their Production by Microorganisms in Food. Trends Food Sci. Technol. 1994, 5, 42–49. [Google Scholar] [CrossRef]
- Stadnik, J.; Dolatowski, Z.J. Corresponding Author-Adres Do Korespondencji: Dr Inż Biogenic Amines in Meat and Fermented Meat Products. Acta Sci. Pol. Technol. Aliment 2010, 9, 251–263. [Google Scholar]
- Cervelli, M.; Leonetti, A.; Duranti, G.; Sabatini, S.; Ceci, R.; Mariottini, P. Skeletal Muscle Pathophysiology: The Emerging Role of Spermine Oxidase and Spermidine. Med. Sci. 2018, 6, 14. [Google Scholar] [CrossRef]
- Ijaz, M.; Zhang, D.; Hou, C.; Mahmood, M.; Hussain, Z.; Zheng, X.; Li, X. Changes in Postmortem Metabolites Profile of Atypical and Typical DFD Beef. Meat Sci. 2022, 193, 108922. [Google Scholar] [CrossRef]
- Zou, Y.; Yang, Y.; Fu, X.; He, X.; Liu, M.; Zong, T.; Li, X.; Htet Aung, L.; Wang, Z.; Yu, T. The Regulatory Roles of Aminoacyl-TRNA Synthetase in Cardiovascular Disease. Mol. Ther. Nucleic Acids 2021, 25, 372–387. [Google Scholar] [CrossRef]
- Zhu, M.J.; Ford, S.P.; Means, W.J.; Hess, B.W.; Nathanielsz, P.W.; Du, M. Maternal Nutrient Restriction Affects Properties of Skeletal Muscle in Offspring. J. Physiol. 2006, 575, 241–250. [Google Scholar] [CrossRef]
- Suravajhala, P.; Kogelman, L.J.A.; Kadarmideen, H.N. Multi-Omic Data Integration and Analysis Using Systems Genomics Approaches: Methods and Applications in Animal Production, Health and Welfare. Genet. Sel. Evol. 2016, 48. [Google Scholar] [CrossRef] [PubMed]

| Metabolites | NP | PP | FP | p Value 1 |
|---|---|---|---|---|
| PC aa C26:0 | 0.65 ± 0.14 a | 0.78 ± 0.13 a | 0.36 ± 0.21 b | 0.003 |
| PC ae C34:2 | 3.17 ± 1.39 ab | 5.91 ± 2.89 a | 1.35 ± 1.71 b | 0.010 |
| PC ae C40:1 | 0.19 ± 0.13 ab | 0.41 ± 0.18 a | 0.14 ± 0.18 b | 0.012 |
| PC ae C34:1 | 6.33 ± 2.26 a | 4.70 ± 2.01 ab | 1.62 ± 2.06 b | 0.013 |
| PC aa C34:2 | 100.38 ± 60.14 a | 94.48 ± 33.05 a | 18.24 ± 26.06 b | 0.017 |
| PC ae C38:5 | 3.77 ± 1.87 ab | 5.03 ± 2.51 a | 1.33 ± 1.82 b | 0.017 |
| SM C18:0 | 10.25 ± 4.15 ab | 14.15 ± 3.40 a | 4.55 ± 5.69 b | 0.019 |
| Espermidine | 0.35 ± 0.09 b | 0.49 ± 0.13 a | 0.52 ± 0.06 a | 0.022 |
| Ala | 864.56 ± 118.87 b | 1061.35 ± 74.07 a | 981.71 ± 147.33 ab | 0.022 |
| SM (OH) C24:1 | 0.59 ± 0.17 a | 0.41 ± 0.16 ab | 0.22 ± 0.24 b | 0.024 |
| PC ae C36:5 | 3.77 ± 1.55 ab | 5.35 ± 2.66 a | 1.58 ± 2.09 b | 0.025 |
| PC aa C38:0 | 1.35 ± 0.55 a | 1.28 ± 0.38 a | 0.45 ± 0.66 b | 0.026 |
| PC ae C36:1 | 9.42 ± 3.51 ab | 11.49 ± 6.88 a | 2.82 ± 4.13 b | 0.027 |
| lisoPC a C26:1 | 0.29 ± 0.13 a | 0.30 ± 0.10 a | 0.10 ± 0.15 b | 0.031 |
| Thr | 61.53 ± 9.96 b | 80.00 ± 10.63 a | 78.21 ± 11.78 a | 0.032 |
| PC ae C40:6 | 1.23 ± 0.56 ab | 1.78 ± 0.79 a | 0.60 ± 0.88 b | 0.034 |
| PC aa C32:2 | 7.42 ± 3.39 ab | 10.42 ± 7.10 a | 1.91 ± 3.52 b | 0.035 |
| PC aa C42:0 | 0.06 ± 0.03 ab | 0.09 ± 0.04 a | 0.04 ± 0.03 b | 0.038 |
| PC aa C34:3 | 8.94 ± 4.69 a | 7.60 ± 3.76 ab | 2.12 ± 2.97 b | 0.040 |
| PC ae C32:1 | 1.89 ± 0.95 a | 1.60 ± 0.80 ab | 0.48 ± 0.59 b | 0.040 |
| PC ae C36:2 | 19.28 ± 11.68 a | 19.43 ± 5.84 a | 5.16 ± 8.17 b | 0.041 |
| PC aa C36:2 | 288.65 ± 162.68 a | 180.22 ± 118.81 ab | 64.00 ± 94.01 b | 0.043 |
| Arg | 45.40 ± 11.59 b | 59.03 ± 8.93 a | 59.62 ± 9.35 a | 0.044 |
| PC ae C44:6 | 0.11 ± 0.04 ab | 0.16 ± 0.05 a | 0.09 ± 0.04 b | 0.045 |
| PC ae C36:4 | 4.66 ± 3.02 ab | 7.54 ± 4.99 a | 1.29 ± 2.01 b | 0.049 |
| Metabolites | NP | PP | FP | p Value 1 |
|---|---|---|---|---|
| PC aa C36:0 | 2.06 ± 2.89 ab | 1.48 ± 1.01 b | 7.39 ± 5.11 a | 0.013 |
| PC ae C40:5 | 0.87 ± 0.84 b | 1.19 ± 0.77 ab | 2.32 ± 1.09 a | 0.016 |
| PC ae C36:0 | 1.64 ± 2.52 ab | 0.77 ± 0.49 b | 4.43 ± 2.88 a | 0.019 |
| PC ae C38:0 | 1.32 ± 1.51 ab | 1.08 ± 0.64 b | 3.24 ± 1.55 a | 0.027 |
| PC aa C38:3 | 7.23 ± 9.04 b | 7.83 ± 8.98 ab | 23.52 ± 14.35 a | 0.031 |
| PC aa C38:5 | 7.12 ± 8.26 b | 9.12 ± 8.92 ab | 19.84 ± 12.54 a | 0.032 |
| PC aa C36:1 | 43.79 ± 43.77 b | 35.70 ± 33.50 b | 143.48 ± 83.53 a | 0.034 |
| PC ae C30:2 | 0.40 ± 0.35 b | 0.41 ± 0.35 b | 1.11 ± 0.49 a | 0.042 |
| PC aa C40:6 | 2.65 ± 3.74 b | 2.92 ± 3.77 ab | 9.15 ± 8.32 a | 0.048 |
| lysoPC a C26:1 | 0.11 ± 0.10 b | 0.25 ± 0.15 a | 0.21 ± 0.07 ab | 0.049 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernandes, A.C.; Polizel, G.H.G.; Cracco, R.C.; Cançado, F.A.C.Q.; Baldin, G.C.; Poleti, M.D.; Ferraz, J.B.S.; Santana, M.H.d.A. Metabolomics Changes in Meat and Subcutaneous Fat of Male Cattle Submitted to Fetal Programming. Metabolites 2024, 14, 9. https://doi.org/10.3390/metabo14010009
Fernandes AC, Polizel GHG, Cracco RC, Cançado FACQ, Baldin GC, Poleti MD, Ferraz JBS, Santana MHdA. Metabolomics Changes in Meat and Subcutaneous Fat of Male Cattle Submitted to Fetal Programming. Metabolites. 2024; 14(1):9. https://doi.org/10.3390/metabo14010009
Chicago/Turabian StyleFernandes, Arícia Christofaro, Guilherme Henrique Gebim Polizel, Roberta Cavalcante Cracco, Fernando Augusto Correia Queiroz Cançado, Geovana Camila Baldin, Mirele Daiana Poleti, José Bento Sterman Ferraz, and Miguel Henrique de Almeida Santana. 2024. "Metabolomics Changes in Meat and Subcutaneous Fat of Male Cattle Submitted to Fetal Programming" Metabolites 14, no. 1: 9. https://doi.org/10.3390/metabo14010009
APA StyleFernandes, A. C., Polizel, G. H. G., Cracco, R. C., Cançado, F. A. C. Q., Baldin, G. C., Poleti, M. D., Ferraz, J. B. S., & Santana, M. H. d. A. (2024). Metabolomics Changes in Meat and Subcutaneous Fat of Male Cattle Submitted to Fetal Programming. Metabolites, 14(1), 9. https://doi.org/10.3390/metabo14010009








