Machine Learning Algorithm to Predict Obstructive Coronary Artery Disease: Insights from the CorLipid Trial
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Population and Eligibility Criteria
2.2. Study Outcomes
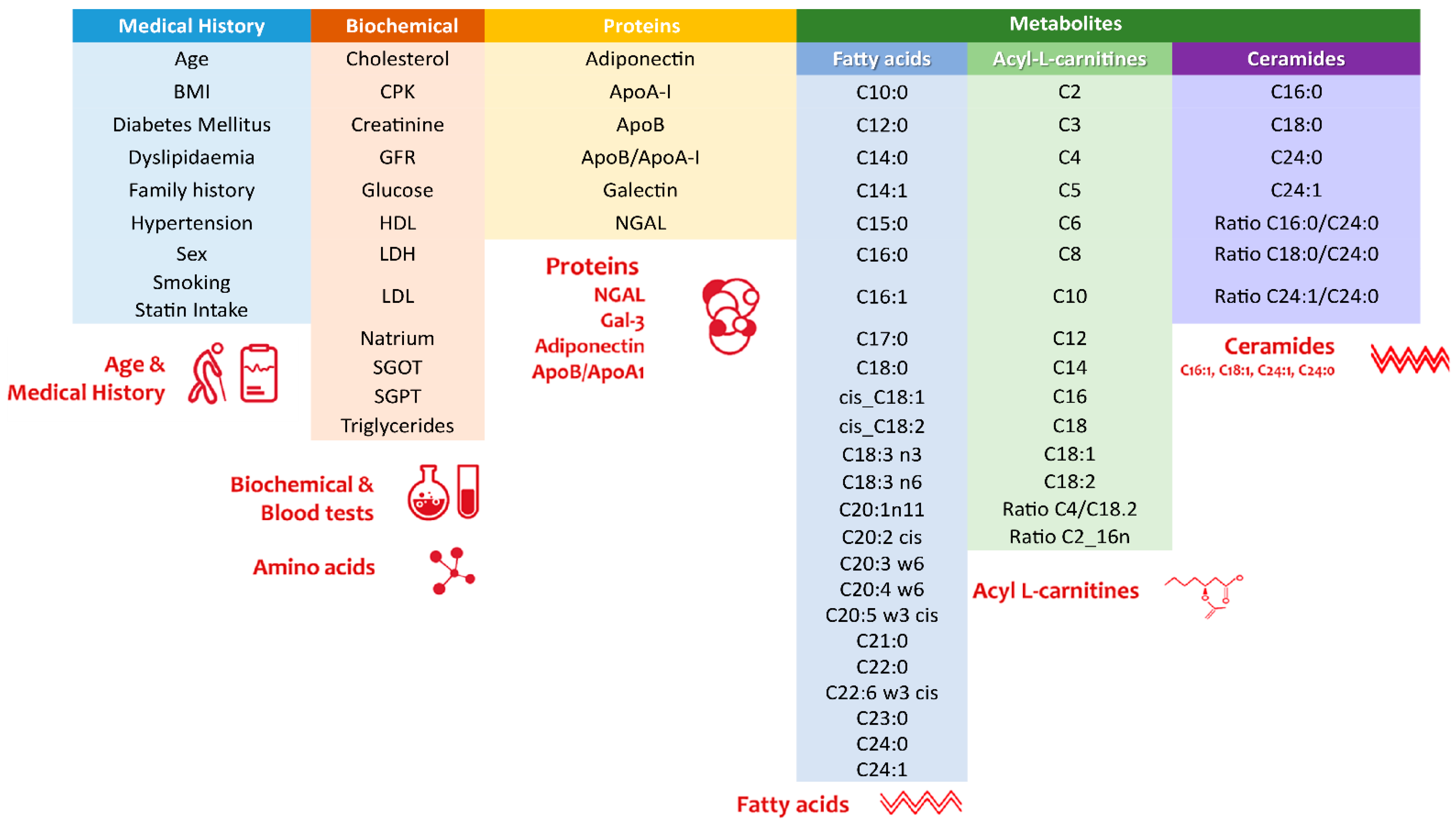
2.3. Metabolic Marker Analyses
2.4. Angiographic Analyses
2.5. Statistical Considerations
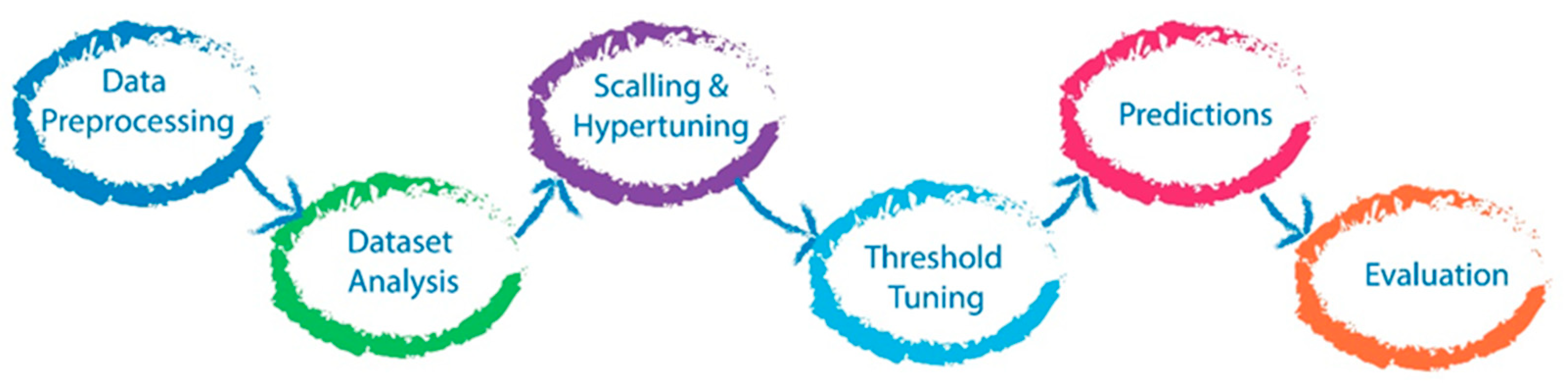
2.6. Machine Learning Algorithm
2.7. Prediction Model Evaluation
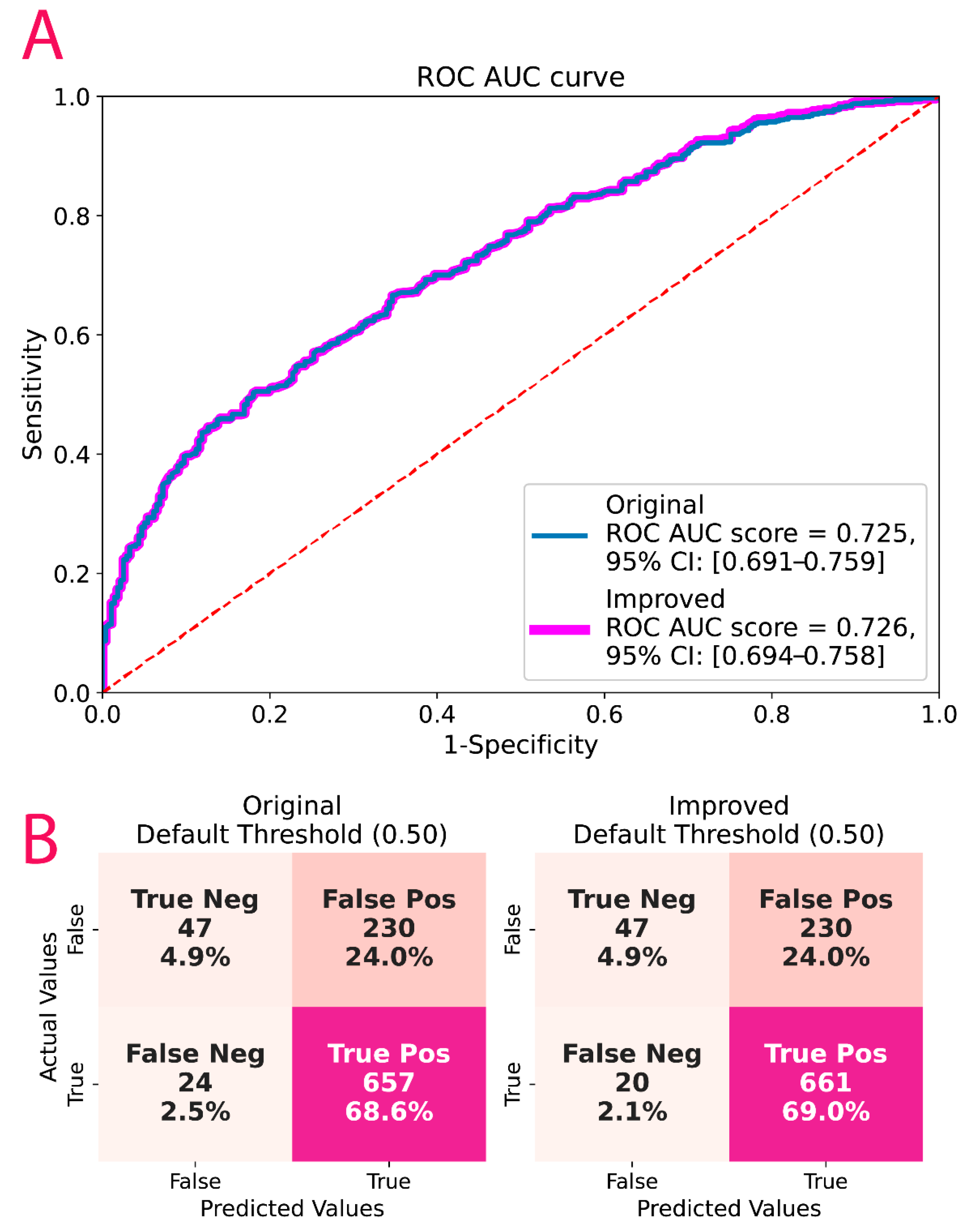
2.8. Post-Hoc Model Correction
2.9. Data Scaling
2.10. Hyperparameter Tuning
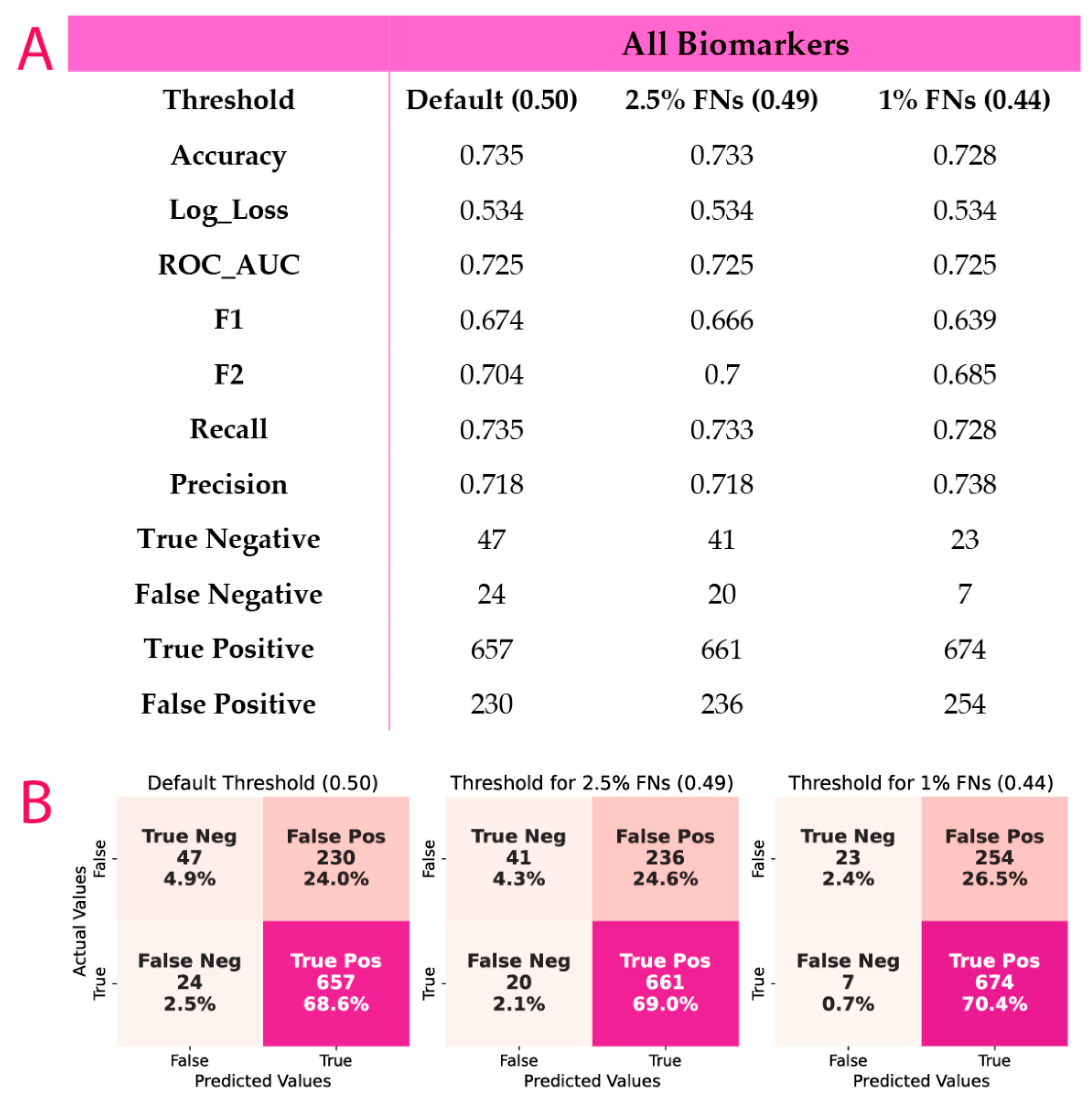
2.11. Probability Threshold Tuning
2.12. Code Development
3. Results
3.1. Baseline Characteristics
3.2. Descriptive Analyses of Categorical and Continuous Variables According to CAD Subgroups
3.3. Metabolite Analyses According to SYNTAX Score Groups
3.4. ML Results
3.5. Post-Hoc Model Correction
4. Discussion
4.1. Metabolites in Cardiovascular Diseases
4.2. Coronary Artery Disease Prediction
4.3. Limitations, Strengths and Further Research
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Nowbar, A.N.; Gitto, M.; Howard, J.P.; Francis, D.P.; Al-Lamee, R. Mortality From Ischemic Heart Disease. Circ. Cardiovasc. Qual. Outcomes 2019, 12, e005375. [Google Scholar] [CrossRef] [PubMed]
- Mozaffarian, D.; Wilson, P.W.; Kannel, W.B. Beyond Established and Novel Risk Factors. Circulation 2008, 117, 3031–3038. [Google Scholar] [CrossRef] [PubMed]
- Vizirianakis, I.S.; Chatzopoulou, F.; Papazoglou, A.S.; Karagiannidis, E.; Sofidis, G.; Stalikas, N.; Stefopoulos, C.; Kyritsis, K.A.; Mittas, N.; Theodoroula, N.F.; et al. The GEnetic Syntax Score: A genetic risk assessment implementation tool grading the complexity of coronary artery disease—Rationale and design of the GESS study. BMC Cardiovasc. Disord. 2021, 21, 284. [Google Scholar] [CrossRef]
- Leon-Mimila, P.; Wang, J.; Huertas-Vazquez, A. Relevance of Multi-Omics Studies in Cardiovascular Diseases. Front. Cardiovasc. Med. 2019, 6, 91. [Google Scholar] [CrossRef] [PubMed]
- Griffin, J.L.; Atherton, H.J.; Shockcor, J.P.; Atzori, L. Metabolomics as a tool for cardiac research. Nat. Rev. Cardiol. 2011, 8, 630–643. [Google Scholar] [CrossRef]
- Müller, J.; Bertsch, T.; Volke, J.; Schmid, A.; Klingbeil, R.; Metodiev, Y.; Karaca, B.; Kim, S.-H.; Lindner, S.; Schupp, T.; et al. Narrative review of metabolomics in cardiovascular disease. J. Thorac. Dis. 2021, 13, 2532–2550. [Google Scholar] [CrossRef]
- Pomyen, Y.; Wanichthanarak, K.; Poungsombat, P.; Fahrmann, J.; Grapov, D.; Khoomrung, S. Deep metabolome: Applications of deep learning in metabolomics. Comput. Struct. Biotechnol. J. 2020, 18, 2818–2825. [Google Scholar] [CrossRef]
- Iliou, A.; Mikros, E.; Karaman, I.; Elliott, F.; Griffin, J.L.; Tzoulaki, I.; Elliott, P. Metabolic phenotyping and cardiovascular disease: An overview of evidence from epidemiological settings. Heart 2021, 107, 1123–1129. [Google Scholar] [CrossRef]
- Sen, P.; Lamichhane, S.; Mathema, V.B.; McGlinchey, A.; Dickens, A.M.; Khoomrung, S.; Orešič, M. Deep learning meets metabolomics: A methodological perspective. Brief. Bioinform. 2020, 22, 1531–1542. [Google Scholar] [CrossRef]
- Krittanawong, C.; Johnson, K.; Rosenson, R.S.; Wang, Z.; Aydar, M.; Baber, U.; Min, J.K.; Tang, W.H.W.; Halperin, J.L.; Narayan, S.M. Deep learning for cardiovascular medicine: A practical primer. Eur. Heart J. 2019, 40, 2058–2073. [Google Scholar] [CrossRef]
- Goldstein, B.A.; Navar, A.M.; Carter, R.E. Moving beyond regression techniques in cardiovascular risk prediction: Applying machine learning to address analytic challenges. Eur. Heart J. 2017, 38, 1805–1814. [Google Scholar] [CrossRef] [PubMed]
- Mittas, N.; Chatzopoulou, F.; Kyritsis, K.A.; Papagiannopoulos, C.I.; Theodoroula, N.F.; Papazoglou, A.S.; Karagiannidis, E.; Sofidis, G.; Moysidis, D.V.; Stalikas, N.; et al. A Risk-Stratification Machine Learning Framework for the Prediction of Coronary Artery Disease Severity: Insights from the GESS Trial. Front. Cardiovasc. Med. 2022, 8, 812182. [Google Scholar] [CrossRef] [PubMed]
- Qiao, H.Y.; Li, J.H.; Schoepf, U.J.; Bayer, R.R.; Tinnefeld, F.C.; Di Jiang, M.; Yang, F.; Guo, B.J.; Zhou, C.S.; Ge, Y.Q.; et al. Prognostic Implication of CT-FFR Based Functional SYNTAX Score in Patients with de Novo Three-Vessel Disease. Eur. Heart J. Cardiovasc. Imaging 2020, 22, 1434–1442. [Google Scholar] [CrossRef] [PubMed]
- Schwalm, J.; Di, S.; Sheth, T.; Natarajan, M.K.; O’Brien, E.; McCready, T.; Petch, J. A machine learning–based clinical decision support algorithm for reducing unnecessary coronary angiograms. Cardiovasc. Digit. Health J. 2022, 3, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Karagiannidis, E.; Sofidis, G.; Papazoglou, A.S.; Deda, O.; Panteris, E.; Moysidis, D.V.; Stalikas, N.; Kartas, A.; Papadopoulos, A.; Stefanopoulos, L.; et al. Correlation of the severity of coronary artery disease with patients’ metabolic profile- rationale, design and baseline patient characteristics of the CorLipid trial. BMC Cardiovasc. Disord. 2021, 21, 79. [Google Scholar] [CrossRef] [PubMed]
- Sianos, G.; Morel, M.-A.; Kappetein, A.P.; Morice, M.-C.; Colombo, A.; Dawkins, K.; van den Brand, M.; Van Dyck, N.; Russell, M.E.; Mohr, F.W.; et al. The SYNTAX Score: An Angiographic Tool Grading the Complexity of Coronary Artery Disease. EuroIntervention 2005, 1, 219–227. [Google Scholar]
- Collet, J.-P.; Thiele, H.; Barbato, E.; Barthélémy, O.; Bauersachs, J.; Bhatt, D.L.; Dendale, P.; Dorobantu, M.; Edvardsen, T.; Folliguet, T.; et al. 2020 ESC Guidelines for the Management of Acute Coronary Syndromes in Patients Presenting without Persistent ST-Segment Elevation: The Task Force for the Management of Acute Coronary Syndromes in Patients Presenting without Persistent ST-Segment Elevation of the European Society of Cardiology (ESC). Eur. Heart J. 2021, 42, 1289–1367. [Google Scholar] [CrossRef]
- Freund, Y.; Schapire, R.E. A Decision-Theoretic Generalization of On-Line Learning and an Application to Boosting. J. Comput. Syst. Sci. 1997, 55, 119–139. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R.; Friedman, J. Boosting and Additive Trees. In The Elements of Statistical Learning: Data Mining, Inference, and Prediction; Hastie, T., Tibshirani, R., Friedman, J., Eds.; Springer Series in Statistics; Springer: New York, NY, USA, 2009; pp. 337–387. ISBN 978-0-387-84858-7. [Google Scholar]
- Mason, L.; Baxter, J.; Bartlett, P.L.; Frean, M.R. Boosting Algorithms as Gradient Descent. In Proceedings of the Advances in Neural Information Processing Systems 12 (NIPS 1999), Denver, CO, USA, 29 November–4December 1999; MIT Press: Cambridge, MA, USA, 1999; p. 7. [Google Scholar]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; Association for Computing Machinery: New York, NY, USA, 2016; pp. 785–794. [Google Scholar]
- Pavlou, M.; Ambler, G.; Seaman, S.; De Iorio, M.; Omar, R.Z. Review and Evaluation of Penalised Regression Methods for Risk Prediction in Low-Dimensional Data with Few Events. Stat. Med. 2016, 35, 1159–1177. Available online: https://pubmed.ncbi.nlm.nih.gov/26514699/ (accessed on 7 July 2022). [CrossRef]
- Liapikos, T.; Zisi, C.; Kodra, D.; Kademoglou, K.; Diamantidou, D.; Begou, O.; Pappa-Louisi, A.; Theodoridis, G. Quantitative structure retention relationship (QSRR) modelling for Analytes’ retention prediction in LC-HRMS by applying different Machine Learning algorithms and evaluating their performance. J. Chromatogr. B 2022, 1191, 123132. [Google Scholar] [CrossRef]
- Elbarouni, B.; Goodman, S.G.; Yan, R.T.; Welsh, R.C.; Kornder, J.M.; DeYoung, J.P.; Wong, G.C.; Rose, B.; Grondin, F.R.; Gallo, R.; et al. Validation of the Global Registry of Acute Coronary Event (GRACE) risk score for in-hospital mortality in patients with acute coronary syndrome in Canada. Am. Heart J. 2009, 158, 392–399. [Google Scholar] [CrossRef]
- Diamond, G.A.; Forrester, J.S. Analysis of Probability as an Aid in the Clinical Diagnosis of Coronary-Artery Disease. N. Engl. J. Med. 1979, 300, 1350–1358. [Google Scholar] [CrossRef]
- Al’Aref, S.J.; Maliakal, G.; Singh, G.; van Rosendael, A.R.; Ma, X.; Xu, Z.; Alawamlh, O.A.H.; Lee, B.; Pandey, M.; Achenbach, S.; et al. Machine Learning of Clinical Variables and Coronary Artery Calcium Scoring for the Prediction of Obstructive Coronary Artery Disease on Coronary Computed Tomography Angiography: Analysis from the CONFIRM Registry. Eur. Heart J. 2020, 41, 359–367. [Google Scholar] [CrossRef]
- Johnson, M.K.; Kuhn, M. Feature Engineering and Selection: A Practical Approach for Predictive Models; CRC Press: Boca Raton, FL, USA, 2019. [Google Scholar]
- Liebal, U.W.; Phan, A.N.T.; Sudhakar, M.; Raman, K.; Blank, L.M. Machine Learning Applications for Mass Spectrometry-Based Metabolomics. Metabolites 2020, 10, 243. [Google Scholar] [CrossRef]
- Acharjee, A.; Ament, Z.; West, J.A.; Stanley, E.; Griffin, J.L. Integration of metabolomics, lipidomics and clinical data using a machine learning method. BMC Bioinform. 2016, 17, 37–49. [Google Scholar] [CrossRef]
- Poss, A.M.; Maschek, J.A.; Cox, J.E.; Hauner, B.J.; Hopkins, P.N.; Hunt, S.C.; Holland, W.L.; Summers, S.A.; Playdon, M.C. Machine learning reveals serum sphingolipids as cholesterol-independent biomarkers of coronary artery disease. J. Clin. Investig. 2020, 130, 1363–1376. [Google Scholar] [CrossRef]
- Cui, S.; Li, L.; Zhang, Y.; Lu, J.; Wang, X.; Song, X.; Liu, J.; Li, K. Machine Learning Identifies Metabolic Signatures that Predict the Risk of Recurrent Angina in Remitted Patients after Percutaneous Coronary Intervention: A Multicenter Prospective Cohort Study. Adv. Sci. 2021, 8, 2003893. [Google Scholar] [CrossRef]
- Fernández Peralbo, M.A.; Priego-Capote, F.; Galache-Osuna, J.G.; Luque de Castro, M.D. Targeted Analysis of Omega-6-Derived Eicosanoids in Human Serum by SPE-LC-MS/MS for Evaluation of Coronary Artery Disease. Electrophoresis 2013, 34, 2901–2909. [Google Scholar] [CrossRef]
- Jung, S.; Ahn, E.; Koh, S.B.; Lee, S.-H.; Hwang, G.-S. Purine Metabolite-Based Machine Learning Models for Risk Prediction, Prognosis, and Diagnosis of Coronary Artery Disease. Biomed. Pharmacother. 2021, 139, 111621. [Google Scholar] [CrossRef]
- Cavus, E.; Karakas, M.; Ojeda, F.M.; Kontto, J.; Veronesi, G.; Ferrario, M.M.; Linneberg, A.; Jørgensen, T.; Meisinger, C.; Thorand, B.; et al. Association of Circulating Metabolites with Risk of Coronary Heart Disease in a European Population: Results from the Biomarkers for Cardiovascular Risk Assessment in Europe (BiomarCaRE) Consortium. JAMA Cardiol. 2019, 4, 1270–1279. [Google Scholar] [CrossRef]
- Deda, O.; Panteris, E.; Meikopoulos, T.; Begou, O.; Mouskeftara, T.; Karagiannidis, E.; Papazoglou, A.S.; Sianos, G.; Theodoridis, G.; Gika, H. Correlation of Serum Acylcarnitines with Clinical Presentation and Severity of Coronary Artery Disease. Biomolecules 2022, 12, 354. [Google Scholar] [CrossRef]
- Karagiannidis, E.; Moysidis, D.V.; Papazoglou, A.S.; Panteris, E.; Deda, O.; Stalikas, N.; Sofidis, G.; Kartas, A.; Bekiaridou, A.; Giannakoulas, G.; et al. Prognostic significance of metabolomic biomarkers in patients with diabetes mellitus and coronary artery disease. Cardiovasc. Diabetol. 2022, 21, 70. [Google Scholar] [CrossRef]
- Karagiannidis, E.; Papazoglou, A.; Stalikas, N.; Deda, O.; Panteris, E.; Begou, O.; Sofidis, G.; Moysidis, D.; Kartas, A.; Chatzinikolaou, E.; et al. Serum Ceramides as Prognostic Biomarkers of Large Thrombus Burden in Patients with STEMI: A Micro-Computed Tomography Study. J. Pers. Med. 2021, 11, 89. [Google Scholar] [CrossRef]
- Papazoglou, A.S.; Stalikas, N.; Moysidis, D.V.; Otountzidis, N.; Kartas, A.; Karagiannidis, E.; Giannakoulas, G.; Sianos, G. CERT2 ceramide- and phospholipid-based risk score and major adverse cardiovascular events: A systematic review and meta-analysis. J. Clin. Lipidol. 2022, 16, 272–276. [Google Scholar] [CrossRef]
- Vignoli, A.; Tenori, L.; Giusti, B.; Takis, P.G.; Valente, S.; Carrabba, N.; Balzi, D.; Barchielli, A.; Marchionni, N.; Gensini, G.F.; et al. NMR-based metabolomics identifies patients at high risk of death within two years after acute myocardial infarction in the AMI-Florence II cohort. BMC Med. 2019, 17, 3. [Google Scholar] [CrossRef]
- Hilvo, M.; Wallentin, L.; Ghukasyan Lakic, T.; Held, C.; Kauhanen, D.; Jylhä, A.; Lindbäck, J.; Siegbahn, A.; Granger, C.B.; Koenig, W.; et al. Prediction of Residual Risk by Ceramide-Phospholipid Score in Patients with Stable Coronary Heart Disease on Optimal Medical Therapy. J. Am. Heart Assoc. 2020, 9, e015258. [Google Scholar] [CrossRef]
- Orlenko, A.; Kofink, D.; Lyytikäinen, L.-P.; Nikus, K.; Mishra, P.; Kuukasjärvi, P.; Karhunen, P.J.; Kähönen, M.; Laurikka, J.O.; Lehtimäki, T.; et al. Model Selection for Metabolomics: Predicting Diagnosis of Coronary Artery Disease Using Automated Machine Learning. Bioinformatics 2020, 36, 1772–1778. [Google Scholar] [CrossRef]
- Cui, H.; Shu, S.; Li, Y.; Yan, X.; Chen, X.; Chen, Z.; Hu, Y.; Chang, Y.; Hu, Z.; Wang, X.; et al. Plasma Metabolites–Based Prediction in Cardiac Surgery–Associated Acute Kidney Injury. J. Am. Heart Assoc. 2021, 10, e021825. [Google Scholar] [CrossRef]
- Akella, A.; Akella, S. Machine learning algorithms for predicting coronary artery disease: Efforts toward an open source solution. Future Sci. OA 2021, 7, FSO698. [Google Scholar] [CrossRef]
- Krittanawong, C.; Virk, H.U.H.; Bangalore, S.; Wang, Z.; Johnson, K.W.; Pinotti, R.; Zhang, H.; Kaplin, S.; Narasimhan, B.; Kitai, T.; et al. Machine Learning Prediction in Cardiovascular Diseases: A Meta-Analysis. Sci. Rep. 2020, 10, 16057. [Google Scholar] [CrossRef]
- Krittanawong, C.; Zhang, H.; Wang, Z.; Aydar, M.; Kitai, T. Artificial Intelligence in Precision Cardiovascular Medicine. J. Am. Coll. Cardiol. 2017, 69, 2657–2664. [Google Scholar] [CrossRef] [PubMed]
- Kigka, V.I.; Georga, E.I.; Sakellarios, A.I.; Tachos, N.S.; Andrikos, I.; Tsompou, P.; Rocchiccioli, S.; Pelosi, G.; Parodi, O.; Michalis, L.K.; et al. A Machine Learning Approach for the Prediction of the Progression of Cardiovascular Disease Based on Clinical and Non-Invasive Imaging Data. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 18–21 July 2018; Volume 2018, pp. 6108–6111. [Google Scholar] [CrossRef]
- Achenbach, S.; Fuchs, F.; Goncalves, A.; Kaiser-Albers, C.; Ali, Z.A.; Bengel, F.M.; Dimmeler, S.; Fayad, Z.A.; Mebazaa, A.; Meder, B.; et al. Non-Invasive Imaging as the Cornerstone of Cardiovascular Precision Medicine. Eur. Heart J. Cardiovasc. Imaging 2022, 23, 465–475. [Google Scholar] [CrossRef] [PubMed]
- Begou, O.A.; Deda, O.; Karagiannidis, E.; Sianos, G.; Theodoridis, G.; Gika, H.G. Development and Validation of a RPLC-MS/MS Method for the Quantification of Ceramides in Human Serum. J. Chromatogr. B 2021, 1175, 122734. [Google Scholar] [CrossRef] [PubMed]
- Meikopoulos, T.; Deda, O.; Karagiannidis, E.; Sianos, G.; Theodoridis, G.; Gika, H. A HILIC-MS/MS Method Development and Validation for the Quantitation of 13 Acylcarnitines in Human Serum. Anal. Bioanal. Chem. 2022, 414, 3095–3108. [Google Scholar] [CrossRef]
- Mouskeftara, T.; Goulas, A.; Ioannidou, D.; Ntenti, C.; Agapakis, D.; Assimopoulou, A.; Gika, H. A Study of Blood Fatty Acids Profile in Hyperlipidemic and Normolipidemic Subjects in Association with Common PNPLA3 and ABCB1 Polymorphisms. Metabolites 2021, 11, 90. [Google Scholar] [CrossRef]
| For 958 CORLIPID Patients | N | N % | |
|---|---|---|---|
| Sex | Female | 255 | 26.6% |
| Male | 703 | 73.4% | |
| Hypertension | No | 398 | 41.5% |
| Yes | 560 | 58.5% | |
| Diabetes mellitus | No | 642 | 67.0% |
| Yes | 316 | 33.0% | |
| Dyslipidaemia | No | 594 | 62.0% |
| Yes | 363 | 37.9% | |
| Family history | No | 788 | 82.3% |
| Yes | 169 | 17.6% | |
| Smoking | No | 535 | 55.8% |
| Yes | 423 | 44.2% | |
| Statin administration | No | 487 | 50.8% |
| Yes | 455 | 47.5% | |
| Age group | 65< | 504 | 52.6% |
| 65> | 452 | 47.2% | |
| Previous stroke | No | 929 | 97.0% |
| Yes | 28 | 2.9% | |
| Peripheral vascular disease | No | 914 | 95.4% |
| Yes | 43 | 4.5% | |
| Aortic aneurysms | No | 928 | 96.9% |
| Yes | 29 | 3.0% | |
| Chronic pulmonary obstructive disease | No | 904 | 94.4% |
| Yes | 54 | 5.6% | |
| Autoimmune disease | No | 941 | 98.2% |
| Yes | 17 | 1.8% | |
| Atrial fibrillation | No | 858 | 89.6% |
| Yes | 100 | 10.4% | |
| ACS | No | 425 | 44.4% |
| Yes | 533 | 55.6% | |
| CAD groups | NSTEMI | 170 | 17.7% |
| STEMI | 222 | 23.2% | |
| Unstable angina | 141 | 14.7% | |
| Stable angina | 425 | 44.4% | |
| Syntax score groups | 0 | 277 | 28.9% |
| 1–22 | 471 | 49.2% | |
| <22 | 210 | 21.9% | |
| Median | ↓95.0% CIs | ↑95.0% CIs | |
|---|---|---|---|
| Age | 65 | 65 | 66 |
| Syntax score | 10.0 | 9.0 | 12.0 |
| Body mass index | 28.00 | 27.80 | 28.40 |
| Total cholesterol | 159.0 | 156.0 | 163.0 |
| Triglycerides | 125 | 122 | 130 |
| High-density lipoprotein | 40 | 39 | 41 |
| Low-density lipoprotein | 88 | 85 | 92 |
| High-sensitivity troponin T | 35.0 | 30.0 | 46.0 |
| Low ventricular ejection fraction (%) | 55 | 55 | 60 |
| CAD Groups | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NSTEMI(α) | STEMI(β) | Unstable Angina(γ) | Stable Angina(δ) | (Pair) p-Value * | ||||||
| N | % | N | % | N | % | N | % | |||
| Sex | Female | 41 | 24.10 | 45 | 20.30 | 43 | 30.50 | 126 | 29.60 | 0.063 |
| Male | 129 | 75.90 | 177 | 79.70 | 98 | 69.50 | 299 | 70.40 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Hypertension | No | 63 | 37.10 | 129 | 58.10 | 57 | 40.40 | 149 | 35.10 | 0.005 (β-α), <0.001 (β-γ), <0.001 (β-δ), |
| Yes | 107 | 62.90 | 93 | 41.90 | 84 | 59.60 | 276 | 64.90 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Diabetes mellitus | No | 111 | 65.30 | 160 | 72.10 | 86 | 61.00 | 285 | 67.10 | 0.164 |
| Yes | 59 | 34.70 | 62 | 27.90 | 55 | 39.00 | 140 | 32.90 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Dyslipidemia | No | 104 | 61.20 | 166 | 74.80 | 92 | 65.20 | 232 | 54.60 | 0.045 (β-α), <0.001 (β-δ), |
| Yes | 65 | 38.20 | 56 | 25.20 | 49 | 34.80 | 193 | 45.40 | ||
| Total | 169 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Family history | No | 133 | 78.20 | 169 | 76.10 | 121 | 85.80 | 365 | 85.90 | 0.012(δ-β) |
| Yes | 37 | 21.80 | 53 | 23.90 | 19 | 13.50 | 60 | 14.10 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 140 | 100.00 | 425 | 100.00 | ||
| Smoking | No | 78 | 45.90 | 94 | 42.30 | 77 | 54.60 | 286 | 67.30 | <0.001(δ-α), <0.001(δ-β) |
| Yes | 92 | 54.10 | 128 | 57.70 | 64 | 45.40 | 139 | 32.70 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Age (groups) | 65< | 93 | 54.70 | 143 | 64.40 | 67 | 47.50 | 201 | 47.30 | 0.013 (β-γ), <0.001 (β-δ), |
| 65> | 76 | 44.70 | 79 | 35.60 | 73 | 51.80 | 224 | 52.70 | ||
| Total | 169 | 100.00 | 222 | 100.00 | 140 | 100.00 | 425 | 100.00 | ||
| Previous stroke | No | 166 | 97.60 | 214 | 96.40 | 138 | 97.90 | 411 | 96.70 | 0.602 |
| Yes | 4 | 2.40 | 8 | 3.60 | 2 | 1.40 | 14 | 3.30 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 140 | 100.00 | 425 | 100.00 | ||
| Peripheral vascular disease | No | 160 | 94.10 | 215 | 96.80 | 133 | 94.30 | 406 | 95.50 | 0.53 |
| Yes | 10 | 5.90 | 7 | 3.20 | 8 | 5.70 | 18 | 4.20 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 141 | 100.00 | 424 | 100.00 | ||
| Aortic aneurysms | No | 167 | 98.20 | 220 | 99.10 | 141 | 100.00 | 400 | 94.10 | 0.003 (γ-δ), 0.003 (β-δ), 0.016 (α-δ) |
| Yes | 2 | 1.20 | 2 | 0.90 | 0 | 0.00 | 25 | 5.90 | ||
| Total | 169 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Chronic pulmonary obstructive disease | No | 158 | 92.90 | 213 | 95.90 | 134 | 95.00 | 399 | 93.90 | 0.574 |
| Yes | 12 | 7.10 | 9 | 4.10 | 7 | 5.00 | 26 | 6.10 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Autoimmune disease | No | 167 | 98.20 | 219 | 98.60 | 137 | 97.20 | 418 | 98.40 | 0.758 |
| Yes | 3 | 1.80 | 3 | 1.40 | 4 | 2.80 | 7 | 1.60 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Atrial fibrillation | No | 155 | 91.20 | 208 | 93.70 | 127 | 90.10 | 368 | 86.60 | 0.03 (δ-β) |
| Yes | 15 | 8.80 | 14 | 6.30 | 14 | 9.90 | 57 | 13.40 | ||
| Total | 170 | 100.00 | 222 | 100.00 | 141 | 100.00 | 425 | 100.00 | ||
| Known CAD | No | 138 | 81.20 | 201 | 90.50 | 121 | 85.80 | 321 | 75.50 | 0.318 |
| Yes | 9 | 5.30 | 8 | 3.60 | 6 | 4.30 | 26 | 6.10 | ||
| Total | 147 | 100.00 | 209 | 100.00 | 127 | 100.00 | 347 | 100.00 | ||
| eGFR < 60 | No | 132 | 77.60 | 191 | 86.00 | 121 | 85.80 | 374 | 88.00 | <0.001 (δ-α) |
| Yes | 38 | 22.40 | 29 | 13.10 | 19 | 13.50 | 41 | 9.60 | ||
| Total | 170 | 100.00 | 220 | 100.00 | 140 | 100.00 | 415 | 100.00 | ||
| CAD Groups | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| NSTEMI(α) | STEMI(β) | Unstable Angina(γ) | Stable Angina(δ) | p-Value * (Pair) | |||||
| Mean | ±SD | Mean | ±SD | Mean | ±SD | Mean | ±SD | ||
| BMI | 27.84 | 4.33 | 28.74 | 4.64 | 28.35 | 4.87 | 28.73 | 4.54 | 0.189 |
| Grace Score | 123 | 41 | 125 | 37 | 96 | 32 | 89 | 25 | <0.001 (δ-α), <0.001 (δ-β), <0.001 (γ-α), <0.001 (γ-β), |
| eGFR | 88.2 | 40.17 | 98.1 | 38.42 | 92.93 | 33.41 | 93.17 | 32.66 | 0.086 |
| Total glucose | 122.15 | 59 | 134.72 | 67.83 | 117.38 | 42.05 | 115.37 | 57.85 | 0.002 (δ-β), 0.032 (α-β) |
| Creatinine | 1.3 | 1.38 | 1.04 | 0.6 | 1.01 | 0.79 | 1.02 | 0.87 | 0.076 |
| Cholesterol | 162.9 | 46.1 | 168.9 | 45.3 | 162.1 | 39.1 | 163.1 | 41.8 | 0.648 |
| Triglycerides | 158 | 128 | 158 | 190 | 147 | 72 | 144 | 119 | 0.159 |
| High-density lipoprotein | 40 | 13 | 39 | 10 | 42 | 12 | 45 | 14 | <0.001 (β-δ), <0.001 (α-δ) |
| Low-density lipoprotein | 92 | 39 | 101 | 39 | 91 | 34 | 90 | 35 | 0.024 (γ-β) |
| High-sensitivity troponin T | 564.5 | 936 | 2442.30 | 2675.80 | 106.1 | 397.6 | 38.5 | 159.9 | <0.001 (δ-α), <0.001 (δ-β), <0.001 (δ-γ,) <0.001 (γ-α), <0.001 (γ-β), <0.001 (α-β), |
| Serum glutamic-oxaloacetic transaminase | 42.2 | 60.8 | 172 | 508.2 | 24 | 16.9 | 22.1 | 17.3 | <0.001 (δ-α), <0.001 (δ-β), <0.001 (γ-α), <0.001 (γ-β), <0.001 (α-β), |
| Serum glutamic pyruvic transaminase | 297.5 | 3372.40 | 78.9 | 341.8 | 26.7 | 25.3 | 24.3 | 33.3 | 0.017 (δ-α), <0.001 (δ-β), <0.001 (γ-β), <0.001 (α-β), |
| Lactate dehydrogenase | 308 | 165 | 629 | 601 | 211 | 66 | 222 | 120 | <0.001 (δ-α), <0.001 (δ-β), <0.001 (γ-α), <0.001 (γ-β), <0.001 (α-β), |
| Creatine phosphokinase | 317 | 693 | 1166 | 1763 | 113 | 159 | 114 | 131 | <0.001 (δ-α), <0.001 (δ-β), <0.001 (γ-α), <0.001 (γ-β), <0.001 (α-β), |
| Low ventricular ejection fraction (%) | 0.5 | 0.11 | 0.44 | 0.1 | 0.54 | 0.1 | 0.56 | 0.09 | <0.001 (δ-α), <0.001 (δ-β), 0.015 (γ-α), <0.001 (γ-β), <0.001 (α-β), |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Panteris, E.; Deda, O.; Papazoglou, A.S.; Karagiannidis, E.; Liapikos, T.; Begou, O.; Meikopoulos, T.; Mouskeftara, T.; Sofidis, G.; Sianos, G.; et al. Machine Learning Algorithm to Predict Obstructive Coronary Artery Disease: Insights from the CorLipid Trial. Metabolites 2022, 12, 816. https://doi.org/10.3390/metabo12090816
Panteris E, Deda O, Papazoglou AS, Karagiannidis E, Liapikos T, Begou O, Meikopoulos T, Mouskeftara T, Sofidis G, Sianos G, et al. Machine Learning Algorithm to Predict Obstructive Coronary Artery Disease: Insights from the CorLipid Trial. Metabolites. 2022; 12(9):816. https://doi.org/10.3390/metabo12090816
Chicago/Turabian StylePanteris, Eleftherios, Olga Deda, Andreas S. Papazoglou, Efstratios Karagiannidis, Theodoros Liapikos, Olga Begou, Thomas Meikopoulos, Thomai Mouskeftara, Georgios Sofidis, Georgios Sianos, and et al. 2022. "Machine Learning Algorithm to Predict Obstructive Coronary Artery Disease: Insights from the CorLipid Trial" Metabolites 12, no. 9: 816. https://doi.org/10.3390/metabo12090816
APA StylePanteris, E., Deda, O., Papazoglou, A. S., Karagiannidis, E., Liapikos, T., Begou, O., Meikopoulos, T., Mouskeftara, T., Sofidis, G., Sianos, G., Theodoridis, G., & Gika, H. (2022). Machine Learning Algorithm to Predict Obstructive Coronary Artery Disease: Insights from the CorLipid Trial. Metabolites, 12(9), 816. https://doi.org/10.3390/metabo12090816











