Comprehensive Analysis of 13C6 Glucose Fate in the Hypoxia-Tolerant Blind Mole Rat Skin Fibroblasts
Abstract
:1. Introduction
2. Results
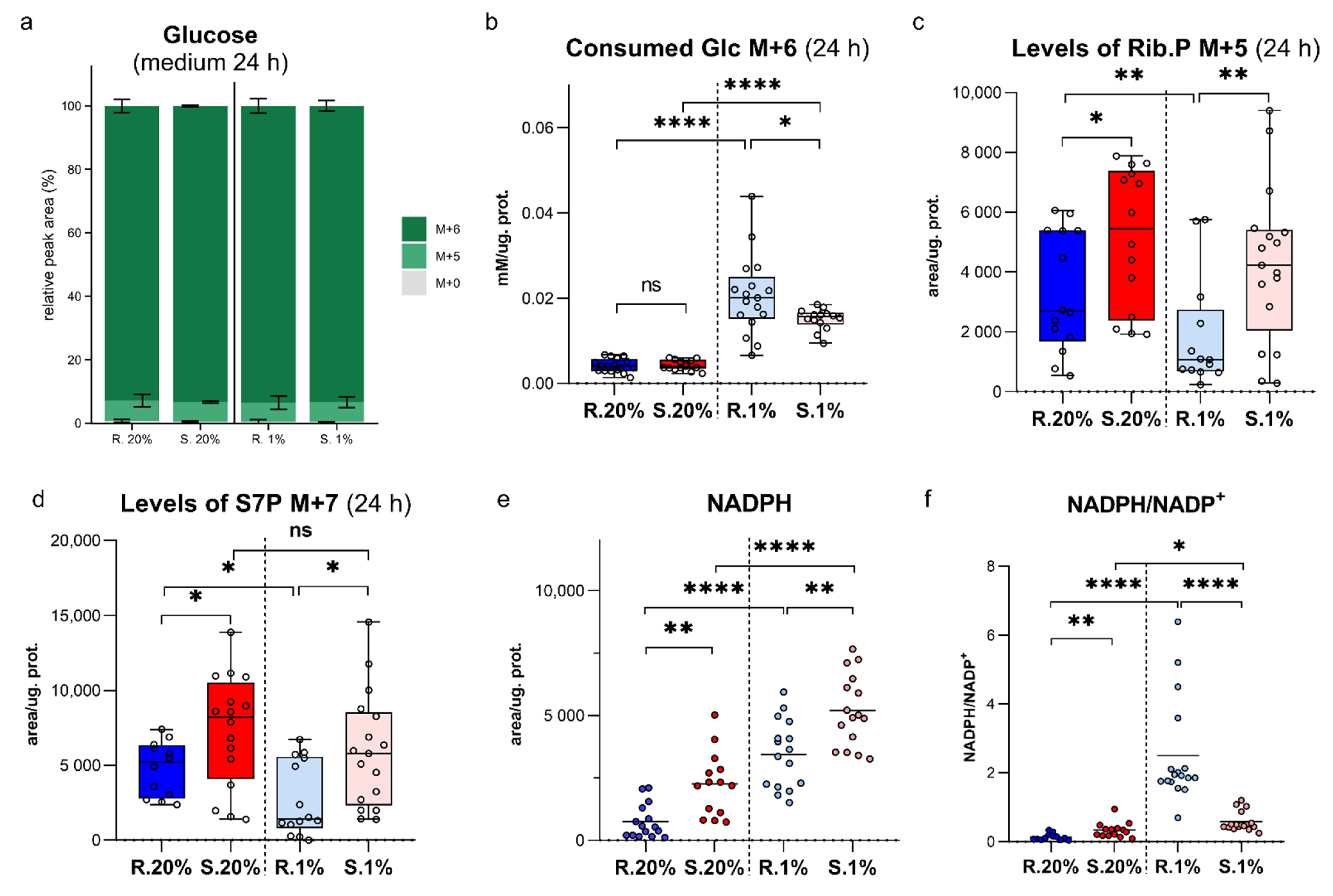
2.1. An Upregulated Pentose Phosphate Pathway (PPP) in Spalax Cells Provides Higher Levels of NADPH
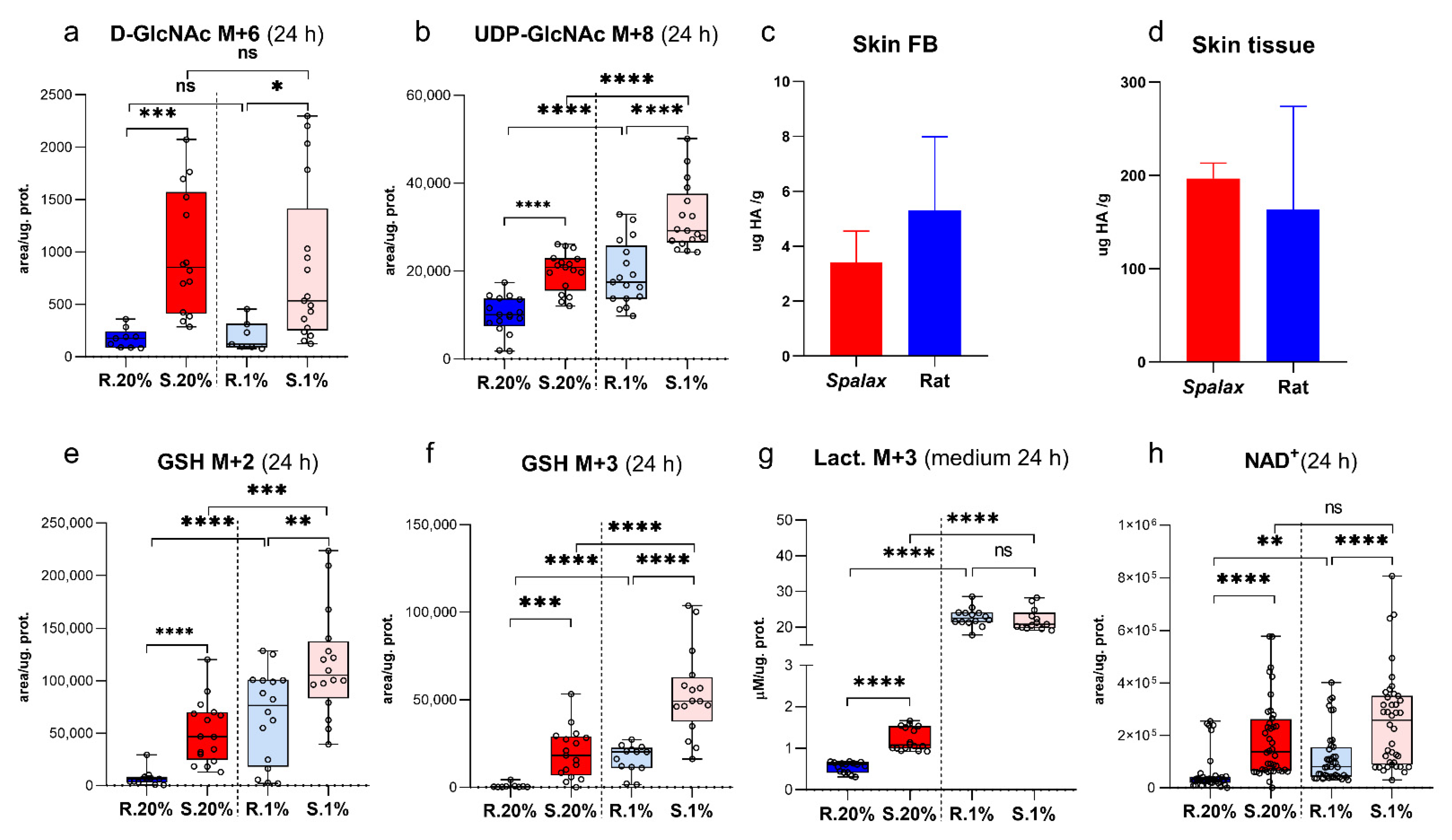
2.2. Spalax Cells Branched Much More Consumed Glucose to Hexosamine Biosynthetic Pathway (HBP), but Not to Hyaluronic Acid Production
2.3. Spalax Cells Massively Redirect Glucose Carbons to the Synthesis of the Tripeptide Glu-Cys-Gly, Glutathione (GSH)
2.4. Upregulated Lactate Production in Spalax Cells Maintains NAD+ Pool for Glycolytic Machinery and Rejects Glucose-Derived Carbons from Downstream Processing in the TCA Cycle
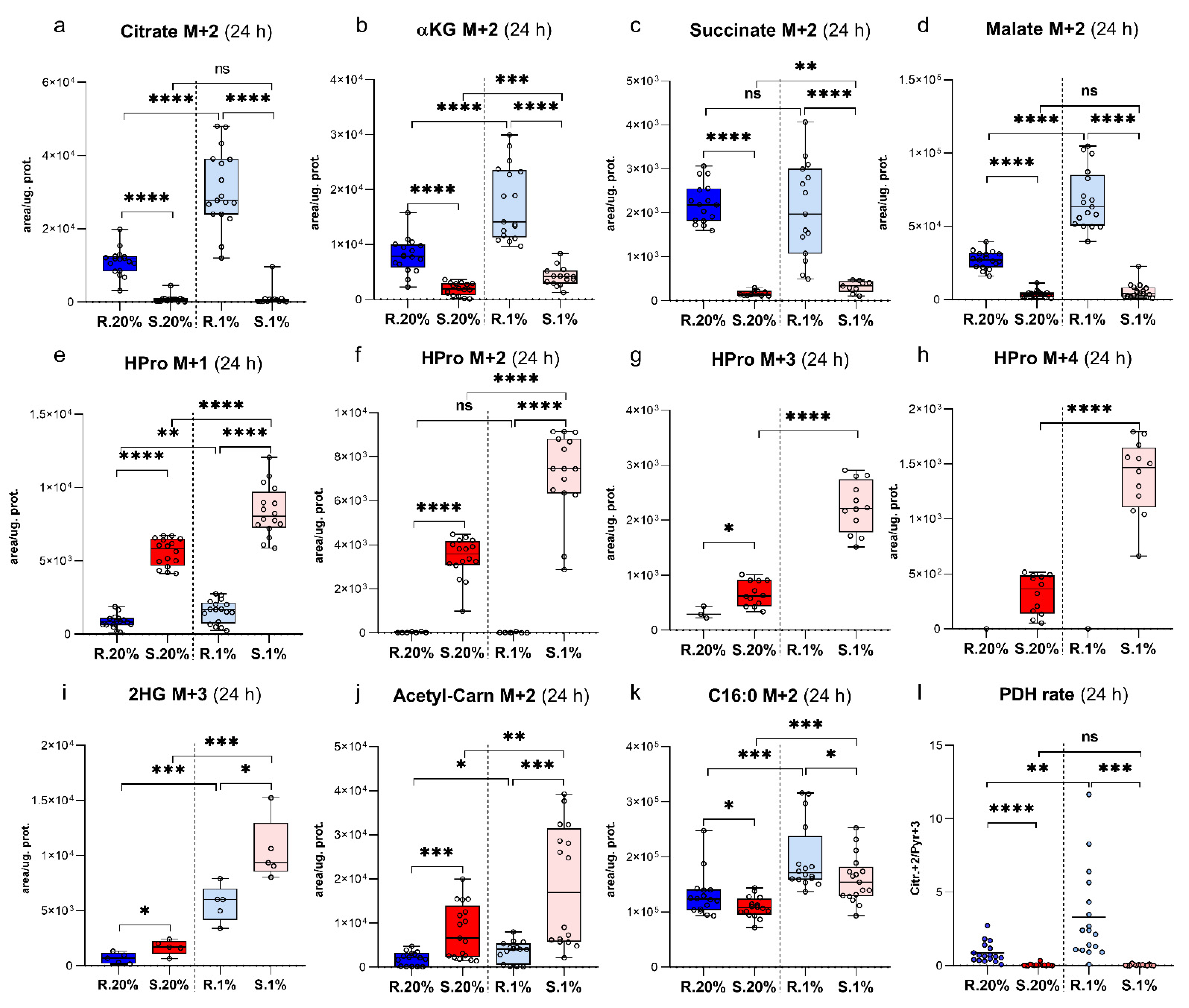
2.5. Spalax Cells Metabolized Less Glc-Originated Pyruvate Entering via Pyruvate Dehydrogenase Complex (PDC) Than the Rat Cells, While Pyruvate Carboxylase (PC)-Mediated Anaplerosis Is More Pronounced in Spalax
2.6. Succinate Is Essential for Hypoxic Spalax Cells
2.7. A Large Part of Glc-Originated Carbons Was Diverted to the Metabolism along with the Axis αKG→Glu→Pro→HPro Instead of Processing in the TCA Cycle Reactions
2.8. Spalax Cells Possess Upregulated Flux of PC-Metabolized Glc Carbons into 2 Hydroxyglutarate (2HG)
2.9. The Newly-Produced Ac-CoA Forwards to Acetyl Carnitine (ALCAR) Synthesis More Than to De-Novo Fatty Acids Production in Spalax Cells
2.10. Hypoxic Spalax Cells Significantly Amplified the Harboring of Glc-Derived Carbons That Entered to Mitochondria as HPro
2.11. Spalax Cells Express Higher Levels of Hypoxia- Inducible Factor-1a (HIF-1α) under Normoxia and Hypoxia Compared to the Rat Cells
2.12. Estimation of the Mitochondrial Aerobic Metabolism (MAM) and the Glycolytic Function (GF) in Spalax and Rat Cells Using the Seahorse Platform
3. Discussion
4. Methods
4.1. General Experimental Design
4.2. Animals
4.3. Cell Culture
4.4. Experiment Schedule, Extraction, and LC-MS Analysis
4.5. 13C6 Glucose Tracing Experiment Design
4.6. LC-MS Metabolomics Analysis
4.7. Oxygen Consumption Rate (OCR) and Extracellular Acidification Rate (ECAR) Measurements
4.8. Hyaluronic Acid Assay
4.9. Western Blot
4.10. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ivy, C.M.; Sprenger, R.J.; Bennett, N.C.; van Jaarsveld, B.; Hart, D.W.; Kirby, A.M.; Yaghoubi, D.; Storey, K.B.; Milsom, W.K.; Pamenter, M.E. The hypoxia tolerance of eight related African mole-rat species rivals that of naked mole-rats, despite divergent ventilatory and metabolic strategies in severe hypoxia. Acta Physiol. 2020, 228, e13436. [Google Scholar] [CrossRef]
- Staples, J.F.; Buck, L.T. Matching cellular metabolic supply and demand in energy-stressed animals. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 2009, 153, 95–105. [Google Scholar] [CrossRef]
- Fahlman, A.; Moore, M.J.; Garcia-Parraga, D. Respiratory function and mechanics in pinnipeds and cetaceans. J. Exp. Biol. 2017, 220, 1761–1773. [Google Scholar] [CrossRef] [Green Version]
- Shams, I.; Avivi, A.; Nevo, E. Oxygen and carbon dioxide fluctuations in burrows of subterranean blind mole rats indicate tolerance to hypoxic-hypercapnic stresses. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 2005, 142, 376–382. [Google Scholar] [CrossRef]
- Tacutu, R.; Craig, T.; Budovsky, A.; Wuttke, D.; Lehmann, G.; Taranukha, D.; Costa, J.; Fraifeld, V.E.; de Magalhaes, J.P. Human Ageing Genomic Resources: Integrated databases and tools for the biology and genetics of ageing. Nucleic Acids Res. 2013, 41, D1027–D1033. [Google Scholar] [CrossRef]
- Caballero, B.; Tomas-Zapico, C.; Vega-Naredo, I.; Sierra, V.; Tolivia, D.; Hardeland, R.; Rodriguez-Colunga, M.J.; Joel, A.; Nevo, E.; Avivi, A.; et al. Antioxidant activity in Spalax ehrenbergi: A possible adaptation to underground stress. J. Comp. Physiol. A Neuroethol. Sens. Neural Behav. Physiol. 2006, 192, 753–759. [Google Scholar] [CrossRef]
- Manov, I.; Hirsh, M.; Iancu, T.C.; Malik, A.; Sotnichenko, N.; Band, M.; Avivi, A.; Shams, I. Pronounced cancer resistance in a subterranean rodent, the blind mole-rat, Spalax: In vivo and in vitro evidence. BMC Biol. 2013, 11, 91. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Galkin, A.; Moncada, S. S-nitrosation of mitochondrial complex I depends on its structural conformation. J. Biol. Chem. 2007, 282, 37448–37453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bruick, R.K.; McKnight, S.L. A conserved family of prolyl-4-hydroxylases that modify HIF. Science 2001, 294, 1337–1340. [Google Scholar] [CrossRef] [Green Version]
- Paton, C.M.; Ntambi, J.M. Biochemical and physiological function of stearoyl-CoA desaturase. Am. J. Physiol. Endocrinol. Metab. 2009, 297, E28–E37. [Google Scholar] [CrossRef] [Green Version]
- Rich, P.R. The molecular machinery of Keilin’s respiratory chain. Biochem. Soc. Trans. 2003, 31, 1095–1105. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Jackson, R.M. Reactive species mechanisms of cellular hypoxia-reoxygenation injury. Am. J. Physiol. Cell Physiol. 2002, 282, C227–C241. [Google Scholar] [CrossRef] [Green Version]
- Kraut, J.A.; Madias, N.E. Lactic acidosis. N. Engl. J. Med. 2015, 372, 1078–1079. [Google Scholar] [CrossRef]
- Oikari, S.; Kettunen, T.; Tiainen, S.; Hayrinen, J.; Masarwah, A.; Sudah, M.; Sutela, A.; Vanninen, R.; Tammi, M.; Auvinen, P. UDP-sugar accumulation drives hyaluronan synthesis in breast cancer. Matrix Biol. 2018, 67, 63–74. [Google Scholar] [CrossRef] [Green Version]
- Yoo, J.; Mashalidis, E.H.; Kuk, A.C.Y.; Yamamoto, K.; Kaeser, B.; Ichikawa, S.; Lee, S.Y. GlcNAc-1-P-transferase-tunicamycin complex structure reveals basis for inhibition of N-glycosylation. Nat. Struct. Mol. Biol. 2018, 25, 217–224. [Google Scholar] [CrossRef]
- Lu, S.C. Regulation of glutathione synthesis. Mol. Asp. Med. 2009, 30, 42–59. [Google Scholar] [CrossRef] [Green Version]
- Valvona, C.J.; Fillmore, H.L.; Nunn, P.B.; Pilkington, G.J. The Regulation and Function of Lactate Dehydrogenase A: Therapeutic Potential in Brain Tumor. Brain Pathol. 2016, 26, 3–17. [Google Scholar] [CrossRef] [Green Version]
- Kraut, J.A.; Madias, N.E. Lactic acidosis. N. Engl. J. Med. 2014, 371, 2309–2319. [Google Scholar] [CrossRef]
- Owen, O.E.; Kalhan, S.C.; Hanson, R.W. The key role of anaplerosis and cataplerosis for citric acid cycle function. J. Biol. Chem. 2002, 277, 30409–30412. [Google Scholar] [CrossRef] [Green Version]
- Peuhkurinen, K.J.; Hassinen, I.E. Pyruvate carboxylation as an anaplerotic mechanism in the isolated perfused rat heart. Biochem. J. 1982, 202, 67–76. [Google Scholar] [CrossRef] [Green Version]
- Buescher, J.M.; Antoniewicz, M.R.; Boros, L.G.; Burgess, S.C.; Brunengraber, H.; Clish, C.B.; DeBerardinis, R.J.; Feron, O.; Frezza, C.; Ghesquiere, B.; et al. A roadmap for interpreting (13)C metabolite labeling patterns from cells. Curr. Opin. Biotechnol. 2015, 34, 189–201. [Google Scholar] [CrossRef]
- Keuper, M.; Jastroch, M.; Yi, C.X.; Fischer-Posovszky, P.; Wabitsch, M.; Tschop, M.H.; Hofmann, S.M. Spare mitochondrial respiratory capacity permits human adipocytes to maintain ATP homeostasis under hypoglycemic conditions. FASEB J. 2014, 28, 761–770. [Google Scholar] [CrossRef]
- Young, J.D. Metabolic flux rewiring in mammalian cell cultures. Curr. Opin. Biotechnol. 2013, 24, 1108–1115. [Google Scholar] [CrossRef] [Green Version]
- Swarup, A.; Lu, J.; DeWoody, K.C.; Antoniewicz, M.R. Metabolic network reconstruction, growth characterization and 13C-metabolic flux analysis of the extremophile Thermus thermophilus HB8. Metab. Eng. 2014, 24, 173–180. [Google Scholar] [CrossRef]
- Antoniewicz, M.R. A guide to (13)C metabolic flux analysis for the cancer biologist. Exp. Mol. Med. 2018, 50, 19. [Google Scholar] [CrossRef] [Green Version]
- Wang, G.L.; Semenza, G.L. Purification and characterization of hypoxia-inducible factor 1. J. Biol. Chem. 1995, 270, 1230–1237. [Google Scholar] [CrossRef] [Green Version]
- Lahiri, S.; Roy, A.; Baby, S.M.; Hoshi, T.; Semenza, G.L.; Prabhakar, N.R. Oxygen sensing in the body. Prog. Biophys. Mol. Biol. 2006, 91, 249–286. [Google Scholar] [CrossRef]
- Semenza, G.L. Oxygen sensing, hypoxia-inducible factors, and disease pathophysiology. Annu. Rev. Pathol. 2014, 9, 47–71. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.H.; Bae, H.C.; Noh, K.H.; Song, K.H.; Ye, S.K.; Mao, C.P.; Lee, K.M.; Wu, T.C.; Kim, T.W. Gain of HIF-1alpha under normoxia in cancer mediates immune adaptation through the AKT/ERK and VEGFA axes. Clin. Cancer Res. 2015, 21, 1438–1446. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhong, H.; De Marzo, A.M.; Laughner, E.; Lim, M.; Hilton, D.A.; Zagzag, D.; Buechler, P.; Isaacs, W.B.; Semenza, G.L.; Simons, J.W. Overexpression of hypoxia-inducible factor 1alpha in common human cancers and their metastases. Cancer Res. 1999, 59, 5830–5835. [Google Scholar]
- Shams, I.; Avivi, A.; Nevo, E. Hypoxic stress tolerance of the blind subterranean mole rat: Expression of erythropoietin and hypoxia-inducible factor 1 alpha. Proc. Natl. Acad. Sci. USA 2004, 101, 9698–9703. [Google Scholar] [CrossRef] [Green Version]
- Semenza, G.L.; Jiang, B.H.; Leung, S.W.; Passantino, R.; Concordet, J.P.; Maire, P.; Giallongo, A. Hypoxia response elements in the aldolase A, enolase 1, and lactate dehydrogenase A gene promoters contain essential binding sites for hypoxia-inducible factor 1. J. Biol. Chem. 1996, 271, 32529–32537. [Google Scholar] [CrossRef] [Green Version]
- Wolters-Eisfeld, G.; Mercanoglu, B.; Strohmaier, A.; Guengoer, C.; Izbicki, J.R.; Bockhorn, M. Hypoxia induced HIF1a-mediated O-GalNAc glycosylation of cytosolic O-GlcNAcylated proteins to regulate signaling pathways in pancreatic cancer. J. Clin. Oncol. 2017, 35, e15739. [Google Scholar] [CrossRef]
- Kim, J.W.; Tchernyshyov, I.; Semenza, G.L.; Dang, C.V. HIF-1-mediated expression of pyruvate dehydrogenase kinase: A metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 2006, 3, 177–185. [Google Scholar] [CrossRef] [Green Version]
- Macklin, P.S.; Yamamoto, A.; Browning, L.; Hofer, M.; Adam, J.; Pugh, C.W. Recent advances in the biology of tumour hypoxia with relevance to diagnostic practice and tissue-based research. J. Pathol. 2020, 250, 593–611. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mohlin, S.; Wigerup, C.; Jogi, A.; Pahlman, S. Hypoxia, pseudohypoxia and cellular differentiation. Exp. Cell Res. 2017, 356, 192–196. [Google Scholar] [CrossRef]
- Harwood, K.R.; Hanover, J.A. Nutrient-driven O-GlcNAc cycling—Think globally but act locally. J. Cell Sci. 2014, 127, 1857–1867. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tian, X.; Azpurua, J.; Hine, C.; Vaidya, A.; Myakishev-Rempel, M.; Ablaeva, J.; Mao, Z.; Nevo, E.; Gorbunova, V.; Seluanov, A. High-molecular-mass hyaluronan mediates the cancer resistance of the naked mole rat. Nature 2013, 499, 346–349. [Google Scholar] [CrossRef] [Green Version]
- Chandel, N.S.; Maltepe, E.; Goldwasser, E.; Mathieu, C.E.; Simon, M.C.; Schumacker, P.T. Mitochondrial reactive oxygen species trigger hypoxia-induced transcription. Proc. Natl. Acad. Sci. USA 1998, 95, 11715–11720. [Google Scholar] [CrossRef] [Green Version]
- De Groot, H.; Littauer, A. Hypoxia, reactive oxygen, and cell injury. Free Radic. Biol. Med. 1989, 6, 541–551. [Google Scholar] [CrossRef]
- Pompella, A.; Visvikis, A.; Paolicchi, A.; De Tata, V.; Casini, A.F. The changing faces of glutathione, a cellular protagonist. Biochem. Pharmacol. 2003, 66, 1499–1503. [Google Scholar] [CrossRef]
- Gorres, K.L.; Raines, R.T. Prolyl 4-hydroxylase. Crit. Rev. Biochem. Mol. Biol. 2010, 45, 106–124. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, R.; Leung, I.K.; Tian, Y.M.; Abboud, M.I.; Ge, W.; Domene, C.; Cantrelle, F.X.; Landrieu, I.; Hardy, A.P.; Pugh, C.W.; et al. Structural basis for oxygen degradation domain selectivity of the HIF prolyl hydroxylases. Nat. Commun. 2016, 7, 12673. [Google Scholar] [CrossRef] [PubMed]
- Hewitson, K.S.; Granatino, N.; Welford, R.W.; McDonough, M.A.; Schofield, C.J. Oxidation by 2-oxoglutarate oxygenases: Non-haem iron systems in catalysis and signalling. Philos. Trans. A Math. Phys. Eng. Sci. 2005, 363, 807–828. [Google Scholar] [CrossRef]
- Xu, W.; Yang, H.; Liu, Y.; Yang, Y.; Wang, P.; Kim, S.H.; Ito, S.; Yang, C.; Wang, P.; Xiao, M.T.; et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell 2011, 19, 17–30. [Google Scholar] [CrossRef] [Green Version]
- Zhao, S.; Lin, Y.; Xu, W.; Jiang, W.; Zha, Z.; Wang, P.; Yu, W.; Li, Z.; Gong, L.; Peng, Y.; et al. Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1alpha. Science 2009, 324, 261–265. [Google Scholar] [CrossRef] [Green Version]
- Bailey, P.S.J.; Nathan, J.A. Metabolic Regulation of Hypoxia-Inducible Transcription Factors: The Role of Small Molecule Metabolites and Iron. Biomedicines 2018, 6, 60. [Google Scholar] [CrossRef] [Green Version]
- Koivunen, P.; Lee, S.; Duncan, C.G.; Lopez, G.; Lu, G.; Ramkissoon, S.; Losman, J.A.; Joensuu, P.; Bergmann, U.; Gross, S.; et al. Transformation by the (R)-enantiomer of 2-hydroxyglutarate linked to EGLN activation. Nature 2012, 483, 484–488. [Google Scholar] [CrossRef]
- Weerapana, E.; Imperiali, B. Asparagine-linked protein glycosylation: From eukaryotic to prokaryotic systems. Glycobiology 2006, 16, 91R–101R. [Google Scholar] [CrossRef] [Green Version]
- Eletto, D.; Chevet, E.; Argon, Y.; Appenzeller-Herzog, C. Redox controls UPR to control redox. J. Cell Sci. 2014, 127, 3649–3658. [Google Scholar] [CrossRef] [Green Version]
- Ferrer, C.M.; Lynch, T.P.; Sodi, V.L.; Falcone, J.N.; Schwab, L.P.; Peacock, D.L.; Vocadlo, D.J.; Seagroves, T.N.; Reginato, M.J. O-GlcNAcylation regulates cancer metabolism and survival stress signaling via regulation of the HIF-1 pathway. Mol. Cell 2014, 54, 820–831. [Google Scholar] [CrossRef] [Green Version]
- Achouri, Y.; Noel, G.; Vertommen, D.; Rider, M.H.; Veiga-Da-Cunha, M.; Van Schaftingen, E. Identification of a dehydrogenase acting on D-2-hydroxyglutarate. Biochem. J. 2004, 381, 35–42. [Google Scholar] [CrossRef] [Green Version]
- Struys, E.A. D-2-Hydroxyglutaric aciduria: Unravelling the biochemical pathway and the genetic defect. J. Inherit. Metab. Dis. 2006, 29, 21–29. [Google Scholar] [CrossRef]
- Parker, A.; Engel, P.C. Preliminary evidence for the existence of specific functional assemblies between enzymes of the beta-oxidation pathway and the respiratory chain. Biochem. J. 2000, 345 Pt 3, 429–435. [Google Scholar] [CrossRef] [PubMed]
- Toplak, M.; Brunner, J.; Schmidt, J.; Macheroux, P. Biochemical characterization of human D-2-hydroxyglutarate dehydrogenase and two disease related variants reveals the molecular cause of D-2-hydroxyglutaric aciduria. Biochim. Biophys. Acta Proteins Proteom. 2019, 1867, 140255. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.S.; Leissing, T.M.; Chowdhury, R.; Hopkinson, R.J.; Schofield, C.J. 2-Oxoglutarate-Dependent Oxygenases. Annu. Rev. Biochem. 2018, 87, 585–620. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Zeng, J.; Geng, P.; Fang, C.; Wang, Y.; Sun, M.; Wang, C.; Wang, J.; Yin, P.; Hu, C.; et al. Global Metabolic Profiling Identifies a Pivotal Role of Proline and Hydroxyproline Metabolism in Supporting Hypoxic Response in Hepatocellular Carcinoma. Clin. Cancer Res. 2018, 24, 474–485. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stetten, M.R. Some aspects of the metabolism of hydroxyproline, studied with the aid of isotopic nitrogen. J. Biol. Chem. 1949, 181, 31–37. [Google Scholar] [CrossRef]
- Valle, D.; Goodman, S.I.; Harris, S.C.; Phang, J.M. Genetic evidence for a common enzyme catalyzing the second step in the degradation of proline and hydroxyproline. J. Clin. Investig. 1979, 64, 1365–1370. [Google Scholar] [CrossRef]
- Glaysher, S.; Cree, I.A. Isolation and culture of colon cancer cells and cell lines. Methods Mol. Biol. 2011, 731, 135–140. [Google Scholar] [CrossRef]
- Mackay, G.M.; Zheng, L.; van den Broek, N.J.; Gottlieb, E. Analysis of Cell Metabolism Using LC-MS and Isotope Tracers. Methods Enzymol. 2015, 561, 171–196. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.; Malik, A.; Bicker, A.; Poetzsch, G.; Avivi, A.; Shams, I.; Hankeln, T. Hypoxia tolerance, longevity and cancer-resistance in the mole rat Spalax—A liver transcriptomics approach. Sci. Rep. 2017, 7, 14348. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Miskevich, D.; Chaban, A.; Dronina, M.; Abramovich, I.; Gottlieb, E.; Shams, I. Comprehensive Analysis of 13C6 Glucose Fate in the Hypoxia-Tolerant Blind Mole Rat Skin Fibroblasts. Metabolites 2021, 11, 734. https://doi.org/10.3390/metabo11110734
Miskevich D, Chaban A, Dronina M, Abramovich I, Gottlieb E, Shams I. Comprehensive Analysis of 13C6 Glucose Fate in the Hypoxia-Tolerant Blind Mole Rat Skin Fibroblasts. Metabolites. 2021; 11(11):734. https://doi.org/10.3390/metabo11110734
Chicago/Turabian StyleMiskevich, Dmitry, Anastasia Chaban, Maria Dronina, Ifat Abramovich, Eyal Gottlieb, and Imad Shams. 2021. "Comprehensive Analysis of 13C6 Glucose Fate in the Hypoxia-Tolerant Blind Mole Rat Skin Fibroblasts" Metabolites 11, no. 11: 734. https://doi.org/10.3390/metabo11110734





