Metabolomics Provide Sensitive Insights into the Impacts of Low Level Environmental Contamination on Fish Health—A Pilot Study
Abstract
1. Introduction
2. Results
2.1. Fish Morphology
2.2. Fish Liver Metabolomics
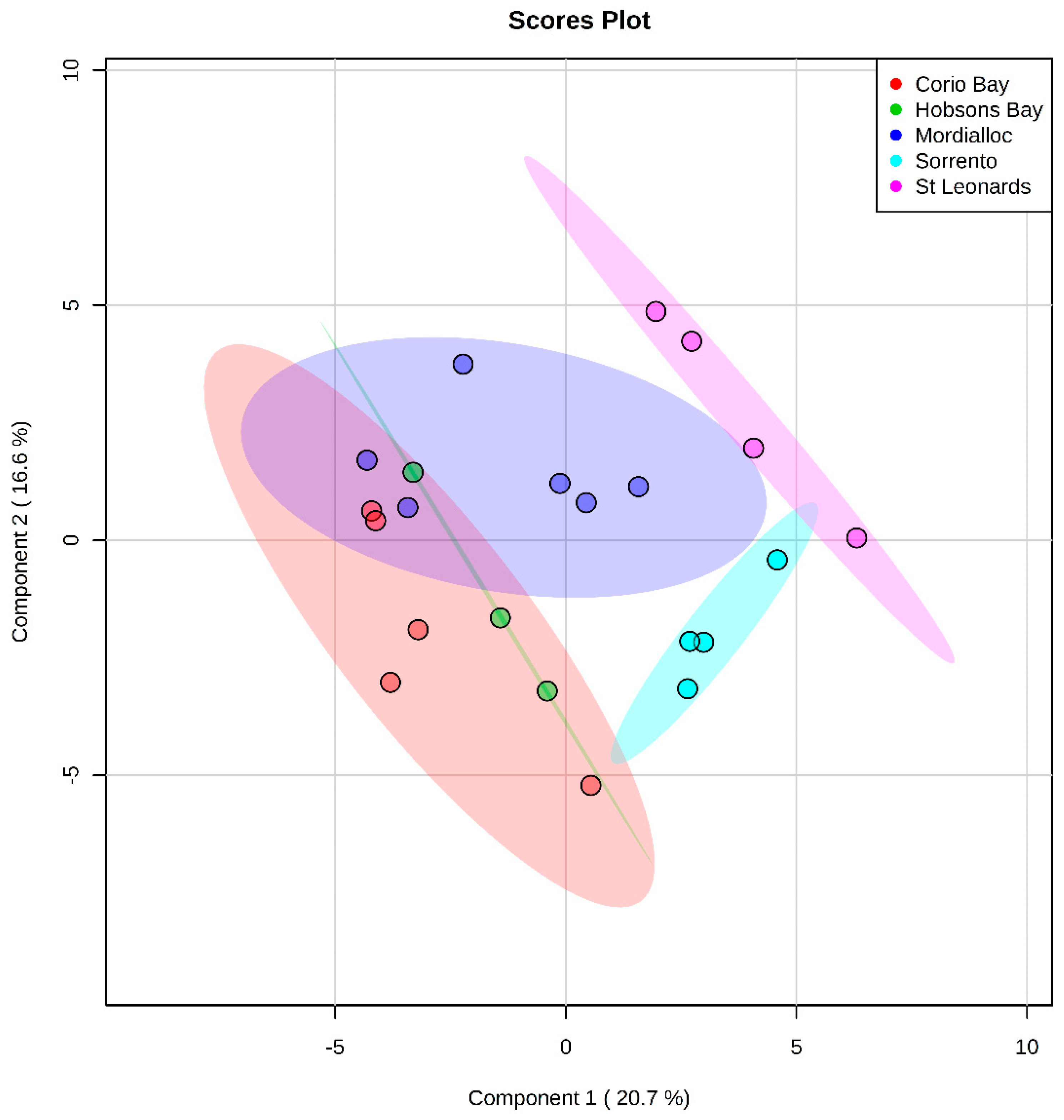
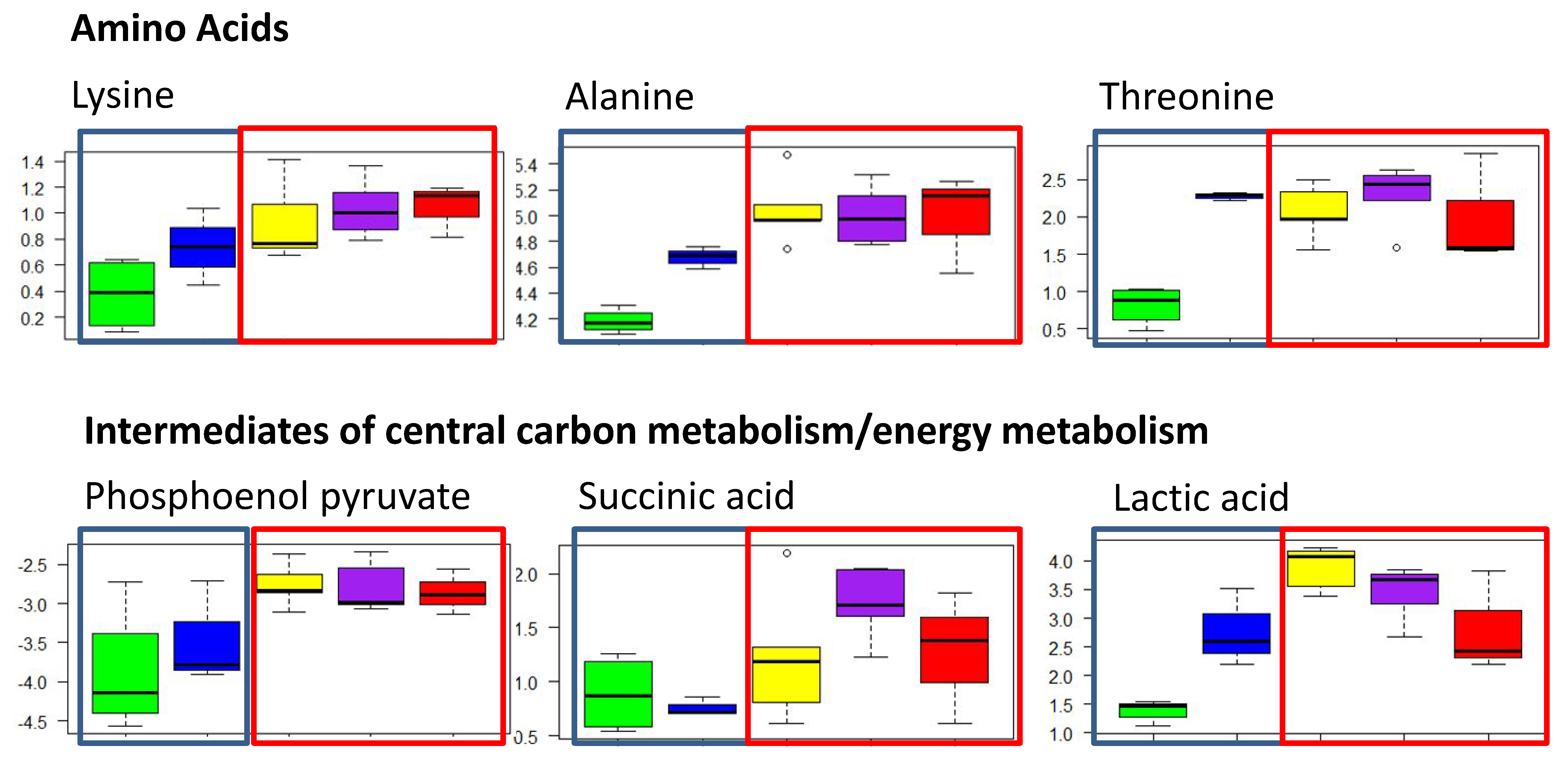
2.2.1. Polar Metabolites
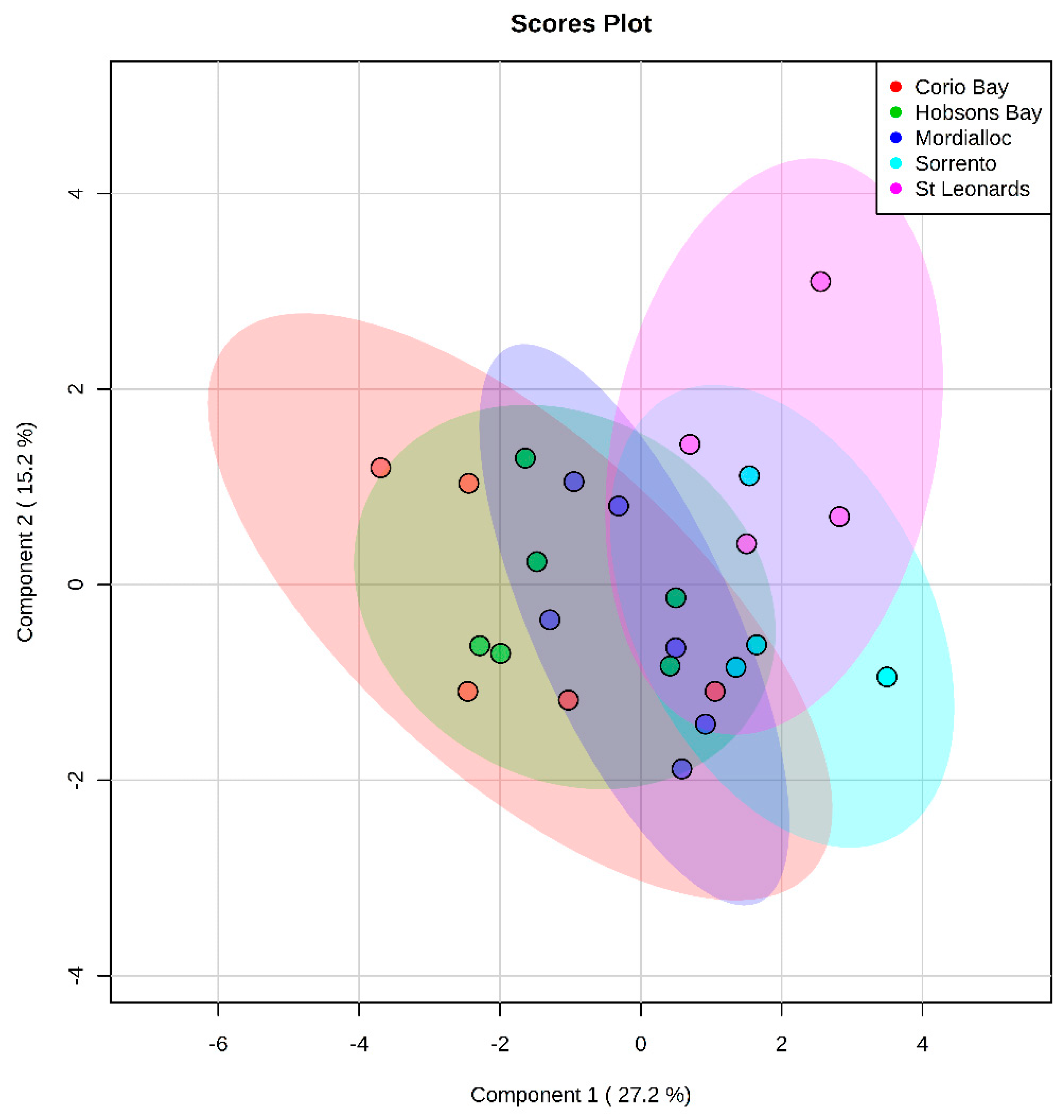
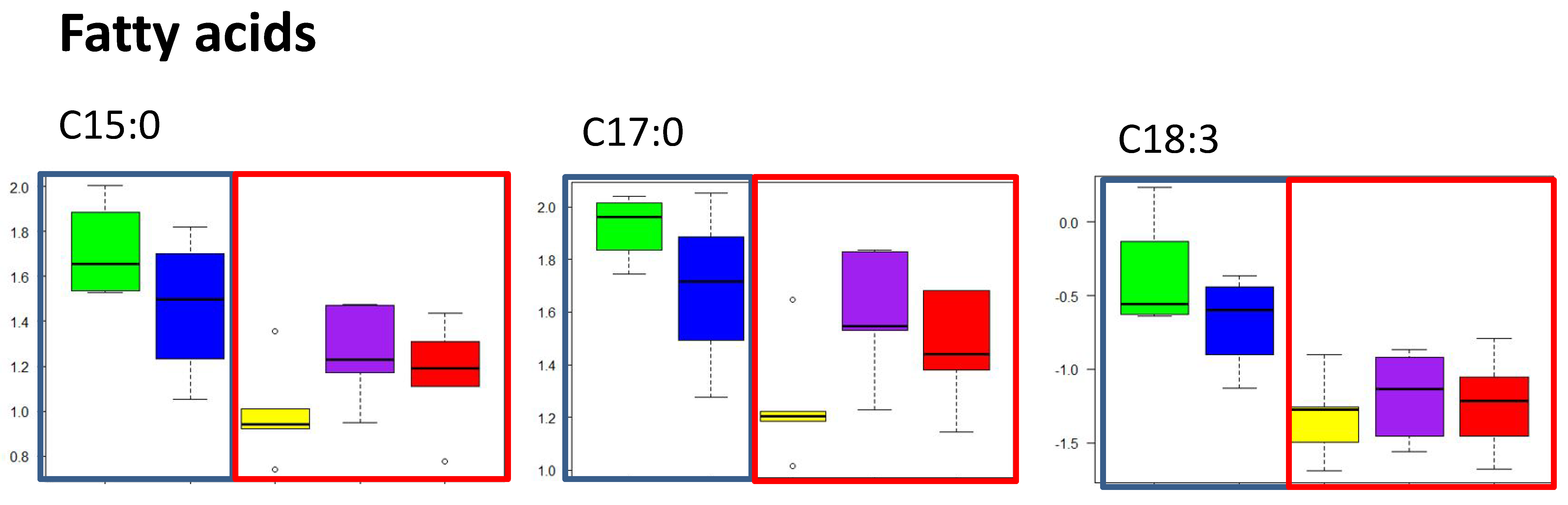
2.2.2. Free Fatty Acid Metabolites
3. Discussion
3.1. Effects of Contamination
3.2. Effect of Temperature and Diet
3.2.1. Temperature
3.2.2. Diet
3.3. Combining Traditional and Novel Biological Assessment Techniques in Understanding Fish Health
4. Materials and Methods
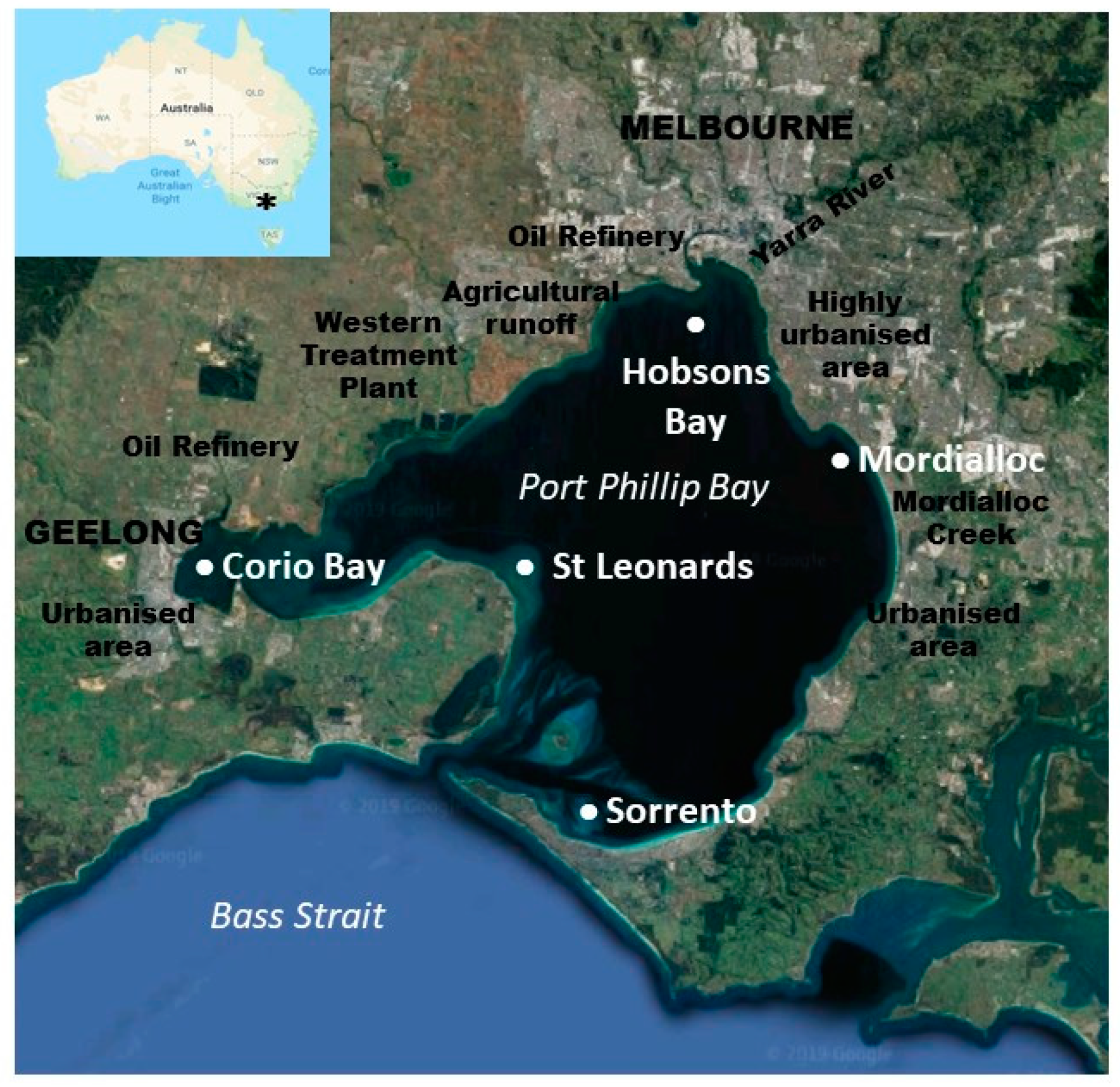
4.1. Fish Collection and Processing
4.2. Metabolomics
4.2.1. Polar Metabolites
4.2.2. Free Fatty Acid Metabolites
4.3. Statistical Analysis
4.3.1. Morphological Parameters
4.3.2. Fish Liver Metabolomics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Samuelsson, L.M.; Larsson, D.G.J. Contributions from metabolomics to fish research. Mol. Biosyst. 2008, 4, 974–979. [Google Scholar] [CrossRef] [PubMed]
- Southam, A.D.; Lange, A.; Hines, A.; Hill, E.M.; Katsu, Y.; Iguchi, T.; Tyler, C.R.; Viant, M.R. Metabolomics Reveals Target and Off-Target Toxicities of a Model Organophosphate Pesticide to Roach (Rutilus rutilus): Implications for Biomonitoring. Environ. Sci. Technol. 2011, 45, 3759–3767. [Google Scholar] [CrossRef] [PubMed]
- Viant, M.R. Metabolomics of aquatic organisms: The new omics on the block. Mar. Ecol. Prog. Ser. 2007, 332, 301–306. [Google Scholar] [CrossRef]
- Griffin, J.L.; Walker, L.A.; Shore, R.F.; Nicholson, J.K. Metabolic profiling of chronic cadmium exposure in the rat. Chem. Res. Toxicol. 2001, 14, 1428–1434. [Google Scholar] [CrossRef] [PubMed]
- Jones, O.A.H.; Murfitt, S.; Svendsen, C.; Turk, A.; Hazel, T.; Spurgeon, D.J.; Walker, L.A.; Shore, R.F.; Long, S.M.; Griffin, J.L. Comparisons of metabolic and physiological changes in rats following short term oral dosing with pesticides commonly found in food. Food Chem. Toxicol. 2013, 59, 438–445. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Huang, X.; Yang, M.; Wang, S.; Ren, S.; Li, H.; Peng, X. GC/MS-based metabolomics approach to identify biomarkers differentiating survivals from death in crucian carps infected by Edwardsiella tarda. Fish Shellfish Immunol. 2014, 39, 215–222. [Google Scholar] [CrossRef]
- Zhou, L.-F.; Zhao, B.-W.; Guan, N.-N.; Wang, W.-M.; Gao, Z.-X. Plasma metabolomics profiling for fish maturation in blunt snout bream. Metabolomics 2017, 13, 40. [Google Scholar] [CrossRef]
- Cipriano, R.C.; Smith, M.L.; Vermeersch, K.A.; Dove, A.D.M.; Styczynski, M.P. Differential metabolite levels in response to spawning-induced inappetence in Atlantic salmon Salmo salar. Comp. Biochem. Physiol. D 2015, 13, 52–59. [Google Scholar] [CrossRef]
- Van Scoy, A.R.; Anderson, B.S.; Philips, B.M.; Voorhees, J.; McCann, M.; De Haro, H.; Martin, M.J.; McCall, J.; Todd, C.R.; Crane, D.; et al. NMR-based characterization of the acute metabolic effects of weathered crude and dispersed oil in spawning top smelt and their embryos. Ecotoxicol. Environ. Saf. 2012, 78, 99–109. [Google Scholar] [CrossRef]
- Van Scoy, A.R.; Lin, C.Y.; Anderson, B.S.; Philips, B.M.; Martin, M.J.; McCall, J.; Todd, C.R.; Crane, D.; Sowby, M.L.; Viant, M.R.; et al. Metabolic responses produced by crude versus dispersed oil in Chinook salmon pre-smolts via NMR-based metabolomics. Ecotoxicol. Environ. Saf. 2010, 73, 710–717. [Google Scholar] [CrossRef]
- Williams, T.D.; Davies, I.M.; Wua, H.; Diab, A.M.; Webster, L.; Viant, M.R.; Chipman, J.K.; Leaver, M.J.; George, S.G.; Moffat, C.F.; et al. Molecular responses of European flounder (Platichthys flesus) chronically exposed to contaminated estuarine sediments. Chemosphere 2014, 108, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Brandão, F.; Cappello, T.; Raimundo, J.; Santos, M.A.; Maisano, M.; Mauceri, A.; Pacheco, M.; Pereira, P. Unravelling the mechanisms of mercury hepatotoxicity in wild fish (Liza aurata) through a triad approach: Bioaccumulation, metabolomic profiles and oxidative stress. Metallomics 2015, 7, 1352–1363. [Google Scholar] [CrossRef] [PubMed]
- Cappello, T.; Pereira, P.; Maisano, M.; Mauceri, A.; Pacheco, M.; Fasulo, S. Advances in understanding the mechanisms of mercury toxicity in wild golden grey mullet (Liza aurata) by 1H NMR-based metabolomics. Environ. Pollut. 2016, 219, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Southam, A.D.; Lange, A.; Al-Salhi, R.; Hill, E.M.; Tyler, C.R.; Viant, M.R. Distinguishing between the metabolome and xenobiotic exposome in environmental field samples analysed by direct-infusion mass spectrometry based metabolomics and lipidomics. Metabolomics 2014, 10, 1050–1058. [Google Scholar] [CrossRef] [PubMed]
- Rebelein, A.; Portner, H.O.; Bock, C. Untargeted metabolic profiling reveals distinct patterns of thermal sensitivity in two related notothenioids. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 2018, 217, 43–54. [Google Scholar] [CrossRef] [PubMed]
- Wei, F.F.; Sakata, K.; Asakura, T.; Date, Y.; Kikuchi, J. Systemic Homeostasis in Metabolome, Ionome, and Microbiome of Wild Yellowfin Goby in Estuarine Ecosystem. Sci. Rep. 2018, 8, 3478. [Google Scholar] [CrossRef]
- Wen, X.; Hu, Y.D.; Zhang, X.Y.; Wei, X.Z.; Wang, T.; Yin, S.W. Integrated application of multi-omics provides insights into cold stress responses in pufferfish Takifugu fasciatus. BMC Genom. 2019, 20, 563. [Google Scholar] [CrossRef]
- Spooner, D.; Walker, T.; Acevedo, S.; Morris, E. Port Phillip Bay Environmental Data Review—Biophysical Assessment of Climate Change; Fisheries Victoria Internal Report No 27; Department of Primary Industries: Queenscliff, Victoria, Australia, 2011. [Google Scholar]
- Phillips, D.J.H.; Richardson, B.J.; Murray, A.P.; Fabris, G.J. Trace metals, organochlorine and hydrocarbons in Port Phillip Bay, Victoria: A historical review. Mar. Pollut. Bull. 1992, 25, 200–217. [Google Scholar] [CrossRef]
- Sampson, J.; Easton, A.; Singh, M. Port Phillip Bay. In Estuaries of Australia in 2050 and Beyond; Springer: Dordrecht, The Netherlands, 2014; pp. 49–68. [Google Scholar]
- Gagnon, M.M.; Baker, J.K.; Long, S.M.; Hassell, K.L.; Pettigrove, V.J. Contaminant (PAHs, OCs, PCBs and trace metals) concentrations are declining in axial tissue of sand flathead (Platycephalus bassensis) collected from an urbanised catchment (Port Phillip Bay, Australia). Mar. Pollut. Bull. 2016, 109, 661–666. [Google Scholar] [CrossRef]
- Barbee, N.; Longmore, A.; Townsend, K.; Pettigrove, V.; Swearer, S. Science Knowledge Synthesis of Nutrient Cycling, Marine Pests and Pollutants: Informing the Development of the New Port Phillip Bay Environmental Management Plan; CAPIM Technical Report No. 60; Centre for Aquatic Pollution Identification and Management, University of Melbourne: Melbourne, Victoria, Austrralia, 2016. [Google Scholar]
- Sharp, S.; Sharley, D.; Pettigrove, V. Sediment Quality Monitoring Program. Final Summary Report; Centre for Aquatic Pollution Identification and Management: Parkville, Australia, 2013. [Google Scholar]
- ANZG Australian and New Zealand Guidelines for Fresh and Marine Water Quality. Australian and New Zealand Governments and Australian State and Territory Governments. Available online: http://www.waterquality.gov.au/anz-guidelines/guideline-values/default/sediment-quality-toxicants (accessed on 20 November 2019).
- Jordan, A.R. Reproductive biology, early life-history and settlement distribution of sand flathead (Platycephalus bassensis) in Tasmania. Mar. Freshw. Res. 2001, 52, 589–601. [Google Scholar] [CrossRef]
- Nicholson, G.J.; Theodoropoulos, T.; Fabris, G.J. Hydrocarbons, Pesticides, PCB and PAH in Port Phillip Bay (Victoria) Sand Flathead. Mar. Pollut. Bull. 1994, 28, 115–120. [Google Scholar] [CrossRef]
- Gagnon, M.M.; Holdway, D.A. EROD activity, serum SDH and PAH biliary metabolites in sand flathead (Platycephalus bassensis) collected in Port Phillip Bay, Australia. Mar. Pollut. Bull. 2002, 44, 230–237. [Google Scholar] [CrossRef]
- Verdouw, J.J.; Macleod, C.K.; Nowak, B.F.; Lyle, J.M. Implications of age, size and region on mercury contamination in estuarine fish species. Water Air Soil Pollut. 2011, 214, 297–306. [Google Scholar] [CrossRef]
- Jones, H.J.; Swadling, K.M.; Tracey, S.R.; Macleod, C.K. Long term trends of Hg uptake in resident fish from a polluted estuary. Mar. Pollut. Bull. 2013, 73, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Baker, J.K.; Long, S.M.; Hassell, K.L.; Pettigrove, V.J.; Gagnon, M.M. Health Status of Inhabiting an Industrialised and Urbanised Embayment, Port Phillip Bay, Victoria as Measured by Biomarkers of Exposure and Effects. PLoS ONE 2016, 11, e0164257. [Google Scholar] [CrossRef]
- Fu, D.; Bridle, A.; Leef, M.; Gagnon, M.M.; Hassell, K.L.; Nowak, B.F. Using a multi-biomarker approach to assess the effects of pollution on sand flathead (Platycephalus bassensis) from Port Phillip Bay, Victoria, Australia. Mar. Pollut. Bull. 2017, 119, 211–219. [Google Scholar] [CrossRef]
- Van der Oost, R.; Beyer, J.; Vermeulen, N.P. Fish bioaccumulation and biomarkers in environmental risk assessment: A review. Environ. Toxicol. Pharmacol. 2003, 13, 57–149. [Google Scholar] [CrossRef]
- Dahle, R.; Taranger, G.L.; Karlsen, Ø.; Kjesbu, O.S.; Norberg, B. Gonadal development and associated changes in liver size and sexual steroids during the reproductive cycle of captive male and female Atlantic cod (Gadus morhua L.). Comp. Biochem. Physiol. A Mol. Integr. Physiol. 2003, 136, 641–653. [Google Scholar] [CrossRef]
- Taranger, G.L.; Carrillo, M.; Schulz, R.W.; Fontaine, P.; Zanuy, S.; Felip, A.; Weltzien, F.A.; Dufour, S.; Karlsen, Ø.; Norberg, B.; et al. Control of puberty in farmed fish. Gen. Comp. Endocrinol. 2010, 165, 483–515. [Google Scholar] [CrossRef]
- Hassell, K.L. Aquatic Environmental Stress Research Group, School of Science, RMIT-University. Personal communication, 2019. [Google Scholar]
- Long, S.M.; Tull, D.L.; Jeppe, K.J.; De Souza, D.P.; Dayalan, S.; Pettigrove, V.J.; McConville, M.J.; Hoffmann, A.A. A multi-platform metabolomics approach demonstrates changes in energy metabolism and the transsulfuration pathway in Chironomus tepperi following exposure to zinc. Aquat. Toxicol. 2015, 162, 54–65. [Google Scholar] [CrossRef]
- Fernandes, C.; Fontaínhas-Fernandes, A.; Peixoto, F.; Salgado, M.A. Bioaccumulation of heavy metals in Liza saliens from the Esmoriz—Paramos coastal lagoon, Portugal. Ecotoxicol. Environ. Saf. 2007, 66, 426–431. [Google Scholar] [CrossRef] [PubMed]
- Finger, A.; Lavers, J.L.; Dann, P.; Nugegoda, D.; Orbell, J.D.; Robertson, B.; Scarpaci, C. The Little Penguin (Eudyptula minor) as an indicator of coastal trace metal pollution. Environ. Pollut. 2015, 205, 365–377. [Google Scholar] [CrossRef] [PubMed]
- Bratberg, M.; Olsvik, P.A.; Edvardsen, R.B.; Brekken, H.K.; Vadla, R.; Meier, S. Effects of oil pollution and persistent organic pollutants (POPs) on glycerophospholipids in liver and brain of male Atlantic cod (Gadus morhua). Chemosphere 2013, 90, 2157–2171. [Google Scholar] [CrossRef] [PubMed]
- Gagnon, M.M.; Holdway, D.A. EROD induction and biliary metabolite excretion following exposure to the water accommodated fraction of crude oil and to chemically dispersed crude oil. Arch. Environ. Contam. Toxicol. 2000, 38, 70–77. [Google Scholar] [CrossRef]
- Fernandes, C.; Fontaínhas-Fernandes, A.; Rocha, E.; Salgado, M.A. Monitoring pollution in Esmoriz–Paramos lagoon, Portugal: Liver histological and biochemical effects in Liza saliens. Environ. Monit. Assess. 2008, 145, 315–322. [Google Scholar] [CrossRef]
- Mathews, C.K.; van Holde, K.E.; Ahern, K.G. Biochemistry, 3rd ed.; Addison Wesley Longman, Inc.: San Francisco, CA, USA, 2000. [Google Scholar]
- De Boeck, G.; Vlaeminck, A.; Blust, R. Effects of Sublethal Copper Exposure on Copper Accumulation, Food Consumption, Growth, Energy Stores, and Nucleic Acid Content in Common Carp. Arch. Environ. Contam. Toxicol. 1997, 33, 415–422. [Google Scholar] [CrossRef]
- De Smet, H.; Blust, R. Stress Responses and Changes in Protein Metabolism in Carp Cyprinus carpio during Cadmium Exposure. Ecotoxicol. Environ. Saf. 2001, 48, 255–262. [Google Scholar] [CrossRef]
- Almeida, J.A.; Novelli, E.L.B.; Dal Pai Silva, M.; Alves, R., Jr. Environmental cadmium exposure and metabolic responses of the Nile tilapia Oreochromis niloticus. Environ. Pollut. 2011, 114, 169–175. [Google Scholar] [CrossRef]
- Burns, K.A.; Smith, J.L. Hydrocarbons in Victorian coastal ecosystems (Australia): Chronic petroleum inputs to Western Port and Port Phillip Bays. Arch. Environ. Contam. Toxicol. 1982, 11, 129–140. [Google Scholar] [CrossRef]
- Speranza, E.D.; Tatone, L.M.; Cappelletti, N.; Colombo, J.C. Cost-benefit of feeding on anthropogenic organic matter: Lipid changes in a detritivorous fish (Prochilodus lineatus). Ichthyol. Res. 2013, 60, 334–342. [Google Scholar] [CrossRef]
- Kowalczyk-Pecka, D.; Pecka, S.; Kowalczuk-Vasilev, E. Selected fatty acids as biomarkers of exposure to microdoses of molluscicides in snails Helix pomatia (Gastropoda Pulmonata). Environ. Pollut. 2017, 222, 138–145. [Google Scholar] [CrossRef]
- Kowalczyk-Pecka, D.; Stanisław Pecka, S.; Kowalczuk-Vasilev, E. Changes in fatty acid metabolism induced by varied micro-supplementation with zinc in snails Helix pomatia (Gastropoda Pulmonata). Ecotoxicol. Environ. Saf. 2017, 138, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Li, J.; Luo, T.; Fan, Y.-W.; Zhang, B.; Ye, J.; Ye, H.; Sun, Y.; Deng, Z.-Y. Predictable Effects of Dietary Lipid Sources on the Fatty Acids Compositions of Four 1-Year-Old Wild Freshwater Fish from Poyang Lake. J. Agric. Food Chem. 2013, 61, 210–218. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.H.; Jackson, J.R.; Twining, C.; Rudstam, L.G.; Zollweg-Horan, E.; Kraft, C.; Lawrence, P.; Kothapalli, K.; Wang, Z.; Brennan, J.T. Saturated Branched Chain, Normal Odd-Carbon-Numbered, and n-3 (Omega-3) Polyunsaturated Fatty Acids in Freshwater Fish in the Northeastern United States. J. Agric. Food Chem. 2016, 64, 7512–7519. [Google Scholar] [CrossRef] [PubMed]
- Budge, S.M.; Parrish, C.C.; McKenzie, C.H. Fatty acid composition of phytoplankton, settling particulate matter and sediments at a sheltered bivalve aquaculture site. Mar. Chem. 2001, 76, 285–303. [Google Scholar] [CrossRef]
- Bottino, N.R.; Gennity, J.; Lilly, M.L.; Simmons, E.; Finne, G. Seasonal and nutritional effects on the fatty acids of three species of shrimp, Penaeus setiferus, P. aztecus and P. duorarum. Aquacult 1980, 19, 139–148. [Google Scholar] [CrossRef]
- Sheridan, M.A. Regulation of lipid metabolism in poikilothermic vertebrates. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 1994, 107, 494–508. [Google Scholar] [CrossRef]
- Kattner, G.; Hagen, W.; Graeve, M.; Albers, C. Exceptional lipids and fatty acids in the pteropod Clione limacina (Gastropoda) from both polar oceans. Mar. Chem. 1998, 61, 219–228. [Google Scholar] [CrossRef]
- Gagnon, M.M.; Rawson, C.A. Bioindicator species for EROD activity measurements: A review with Australian fish as a case study. Ecol. Indic. 2017, 73, 166–180. [Google Scholar] [CrossRef]
- Biales, A.D.; Fritsch, E.B.; Connon, R.E. Real-world application of ‘omics endpoints Response. Environ. Toxicol. Chem. 2015, 24, 700–702. [Google Scholar] [CrossRef]
- Rochfort, S.J.; Ezernieks, V.; Maher, A.D.; Ingram, B.A.; Olsen, L. Mussel metabolomics—Species discrimination and provenance determination. Food Res. Int. 2013, 54, 1302–1312. [Google Scholar] [CrossRef]
- Australian Animal Welfare Strategy. Iki Jime Humane Killing of Fish. Available online: http://www.ikijime.com/wp-content/uploads/2013/07/ikijime_brch_offshore.pdf (accessed on 16 March 2017).
- Overgaard, A.J.; Weir, J.M.; De Souza, D.P.; Tull, D.; Haase, C.; Meikle, P.J.; Pociot, F. Lipidomic and metabolomic characterization of a genetically modified mouse model of the early stages of human type 1 diabetes pathogenesis. Metabolomics 2016, 12, 13. [Google Scholar] [CrossRef] [PubMed]
- Sumner, L.W.; Amberg, A.; Barrett, D.; Beale, M.H.; Beger, R.; Daykin, C.A.; Fan, T.W.-M.; Fiehn, O.; Goodacre, R.; Griffin, J.L.; et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 2007, 3, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Sinclair, G.M.; O’Brien, A.L.; Keough, M.; De Souza, D.P.; Dayalan, S.; Kanojia, K.; Kouremenos, K.; Tull, D.L.; Coleman, R.A.; Jones, O.A.H.; et al. Metabolite Changes in an Estuarine Annelid Following Sublethal Exposure to a Mixture of Zinc and Boscalid. Metabolites 2019, 9, 229. [Google Scholar] [CrossRef] [PubMed]
- Szyma’ nska, E.; Saccenti, E.; Smilde, A.K.; Westerhuis, J.A. Double-Check: Validation of Diagnostic Statistics for PLS-DA Models in Metabolomics Studies. Metabolomics 2012, 8, 3–16. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.G.; Psychogios, N.; Young, N.; Wishart, D.S. Metaboanalyst: A web server for metabolomic data analysis and interpretation. Nucleic Acids Res. 2009, 37, W652–W660. [Google Scholar] [CrossRef]
- Xia, J.G.; Sinelnikov, I.V.; Han, B.; Wishart, D.S. Metaboanalyst 3.0-making metabolomics more meaningful. Nucleic Acids Res. 2015, 43, W251–W257. [Google Scholar] [CrossRef]
| Site | N | As | Cu | Hg | Ni | Se | Zn |
|---|---|---|---|---|---|---|---|
| Sorrento | 4 | 7.30 ± 0.57 | 0.16 ± 0.03 | 0.14 ± 0.02 | 0.02 ± 0.00 | 0.45 ± 0.03 | 5.93 ± 0.85 |
| St Leonards | 4 | 7.87 ± 1.11 | 0.13 ± 0.01 | 0.10 ± 0.02 | 0.02 ± 0.00 | 0.44 ± 0.04 | 6.70 ± 0.45 |
| Corio Bay | 5 | 2.38 ± 0.57 | 0.10 ± 0.01 | 0.19 ± 0.03 | 0.01 ± 0.00 | 0.47 ± 0.02 | 7.16 ± 0.28 |
| Mordialloc | 5 | 6.40 ± 1.39 | 0.14 ± 0.00 | 0.29 ± 0.05 | <0.01 | 0.47 ± 0.02 | 6.34 ± 0.17 |
| Hobsons Bay | 5 | 2.89 ± 0.75 | 0.11 ± 0.01 | 0.20 ± 0.05 | <0.01 | 0.42 ± 0.02 | 6.46 ± 0.42 |
| Site | Type | N | Standard Length (mm) | Carcass Weight (g) | CF 1 | LSI 2 |
|---|---|---|---|---|---|---|
| Sorrento | R | 4 | 216.25 ± 3.47ab | 76.80 ± 4.11a | 0.76 ± 0.04a | 0.80 ± 0.04a |
| St. Leonards | R | 4 | 210.75 ± 3.47ab | 77.35 ± 6.66a | 0.80 ± 0.04a | 0.88 ± 0.06ab |
| Corio Bay | U | 5 | 225.40 ± 5.91a | 98.48 ± 9.21a | 0.85 ± 0.04a | 1.21 ± 0.09cb |
| Mordialloc | U | 6 | 200.67 ± 6.41b | 78.74 ± 8.4a | 0.87 ± 0.02a | 1.87 ± 0.23d |
| Hobsons Bay | U | 3 | 226.67 ± 3.71a | 97.07 ± 6.25a | 0.83 ± 0.02a | 1.59 ± 0.09cd |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Long, S.M.; Tull, D.L.; De Souza, D.P.; Kouremenos, K.A.; Dayalan, S.; McConville, M.J.; Hassell, K.L.; Pettigrove, V.J.; Gagnon, M.M. Metabolomics Provide Sensitive Insights into the Impacts of Low Level Environmental Contamination on Fish Health—A Pilot Study. Metabolites 2020, 10, 24. https://doi.org/10.3390/metabo10010024
Long SM, Tull DL, De Souza DP, Kouremenos KA, Dayalan S, McConville MJ, Hassell KL, Pettigrove VJ, Gagnon MM. Metabolomics Provide Sensitive Insights into the Impacts of Low Level Environmental Contamination on Fish Health—A Pilot Study. Metabolites. 2020; 10(1):24. https://doi.org/10.3390/metabo10010024
Chicago/Turabian StyleLong, Sara M., Dedreia L. Tull, David P. De Souza, Konstantinos A. Kouremenos, Saravanan Dayalan, Malcolm J. McConville, Kathryn L. Hassell, Vincent J. Pettigrove, and Marthe Monique Gagnon. 2020. "Metabolomics Provide Sensitive Insights into the Impacts of Low Level Environmental Contamination on Fish Health—A Pilot Study" Metabolites 10, no. 1: 24. https://doi.org/10.3390/metabo10010024
APA StyleLong, S. M., Tull, D. L., De Souza, D. P., Kouremenos, K. A., Dayalan, S., McConville, M. J., Hassell, K. L., Pettigrove, V. J., & Gagnon, M. M. (2020). Metabolomics Provide Sensitive Insights into the Impacts of Low Level Environmental Contamination on Fish Health—A Pilot Study. Metabolites, 10(1), 24. https://doi.org/10.3390/metabo10010024








