Purification and Biochemical Characterization of Taxadiene Synthase from Bacillus koreensis and Stenotrophomonas maltophilia
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.1.1. Isolation and Screening for TDS Producing Endophytic Bacteria from Different Plants
2.1.2. Taxadiene Synthase Activity and Protein Concentration
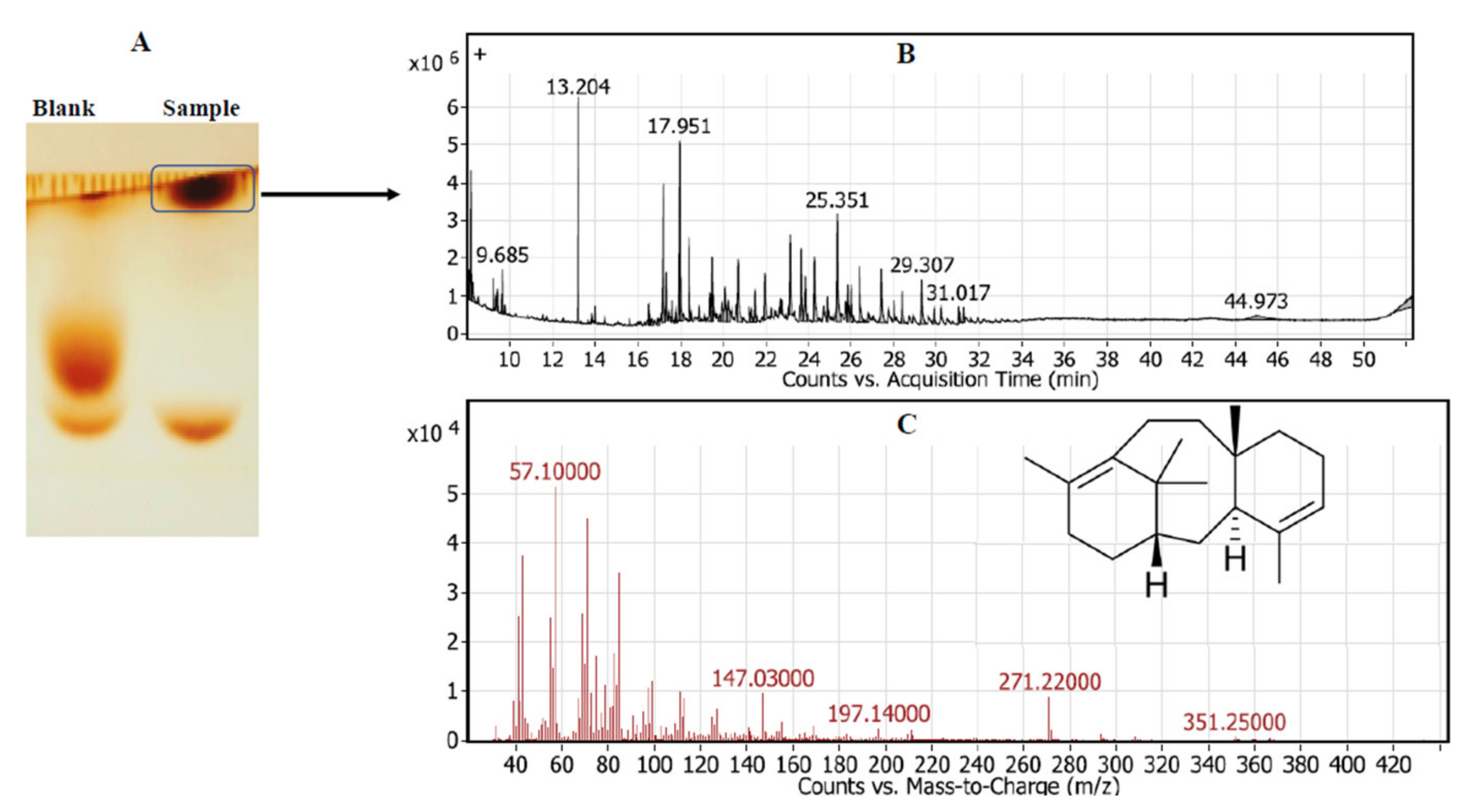
2.1.3. Chemical Identity of Taxadiene
2.1.4. Molecular Detection of TDS by PCR
2.1.5. Molecular Identification of the Potent TDS Producing Bacterial Isolates
2.1.6. Bioprocess Optimization of the Potent Bacterial Isolates for Maximizing Their TDS Yield by Two-Factorial Plackett-Burman Design and Faced Central Composite Design (FCCD)
2.1.7. Purification and Molecular Subunit Structure of TDS from the Most Potent Bacteria
2.1.8. Biochemical Properties of the Purified TDS from the Potent Bacterial Isolate
3. Results
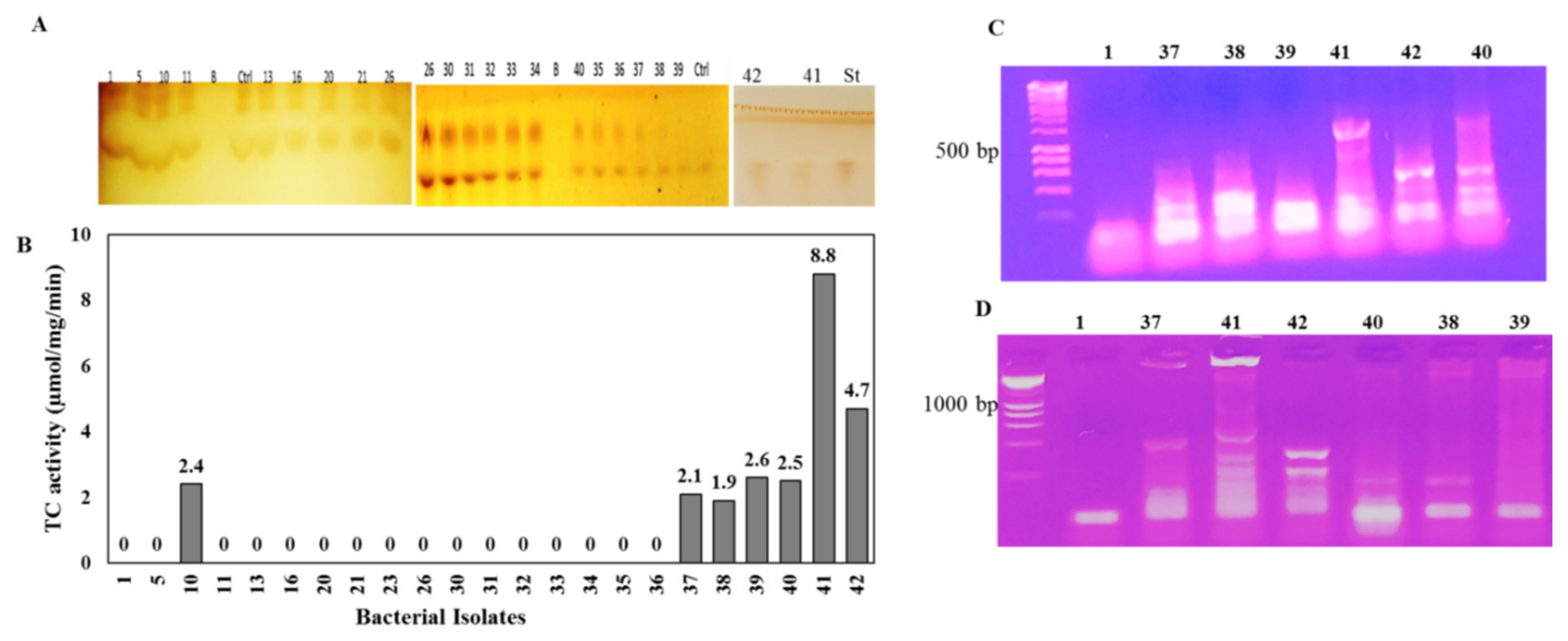
3.1. Screening for the Potent Taxadiene Synthase Producing Bacterial Isolates
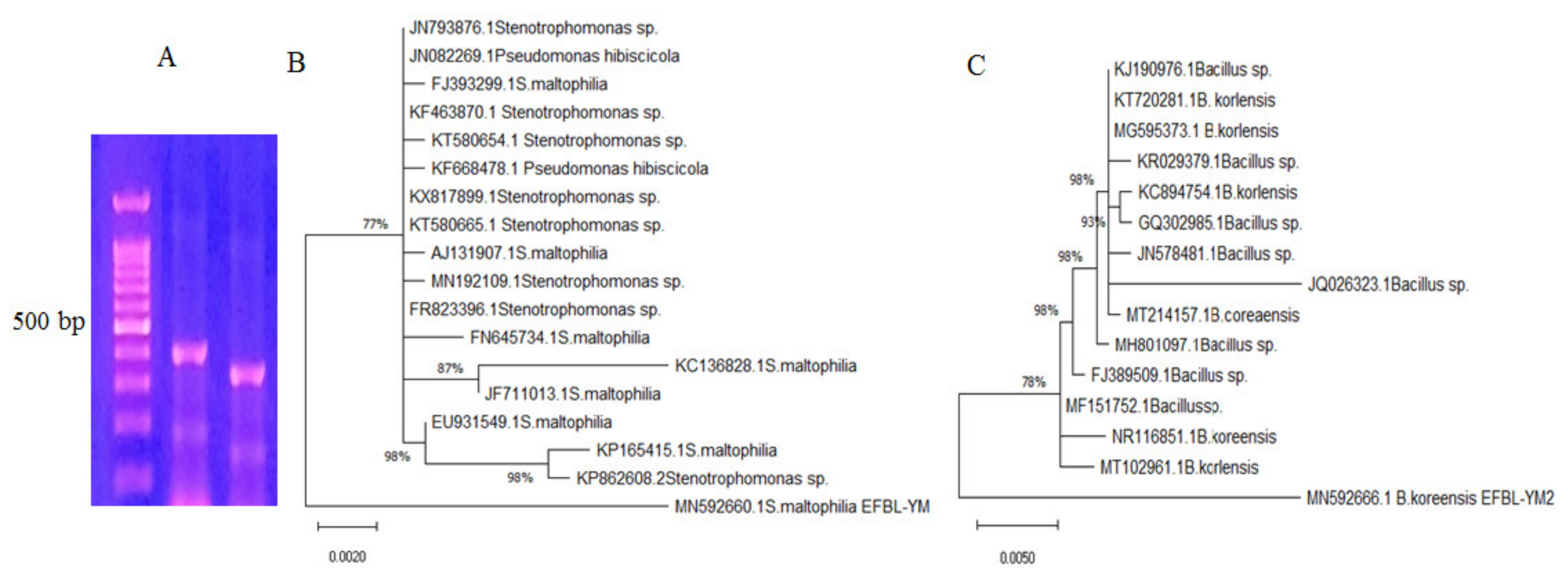
3.2. Molecular Identification of the Potent TDS Producing Bacterial Isolates (# 41 and 42)
3.3. Bioprocess Optimization of TDS Productivity of B. koreensis and S. maltophilia by Response Surface Methodology
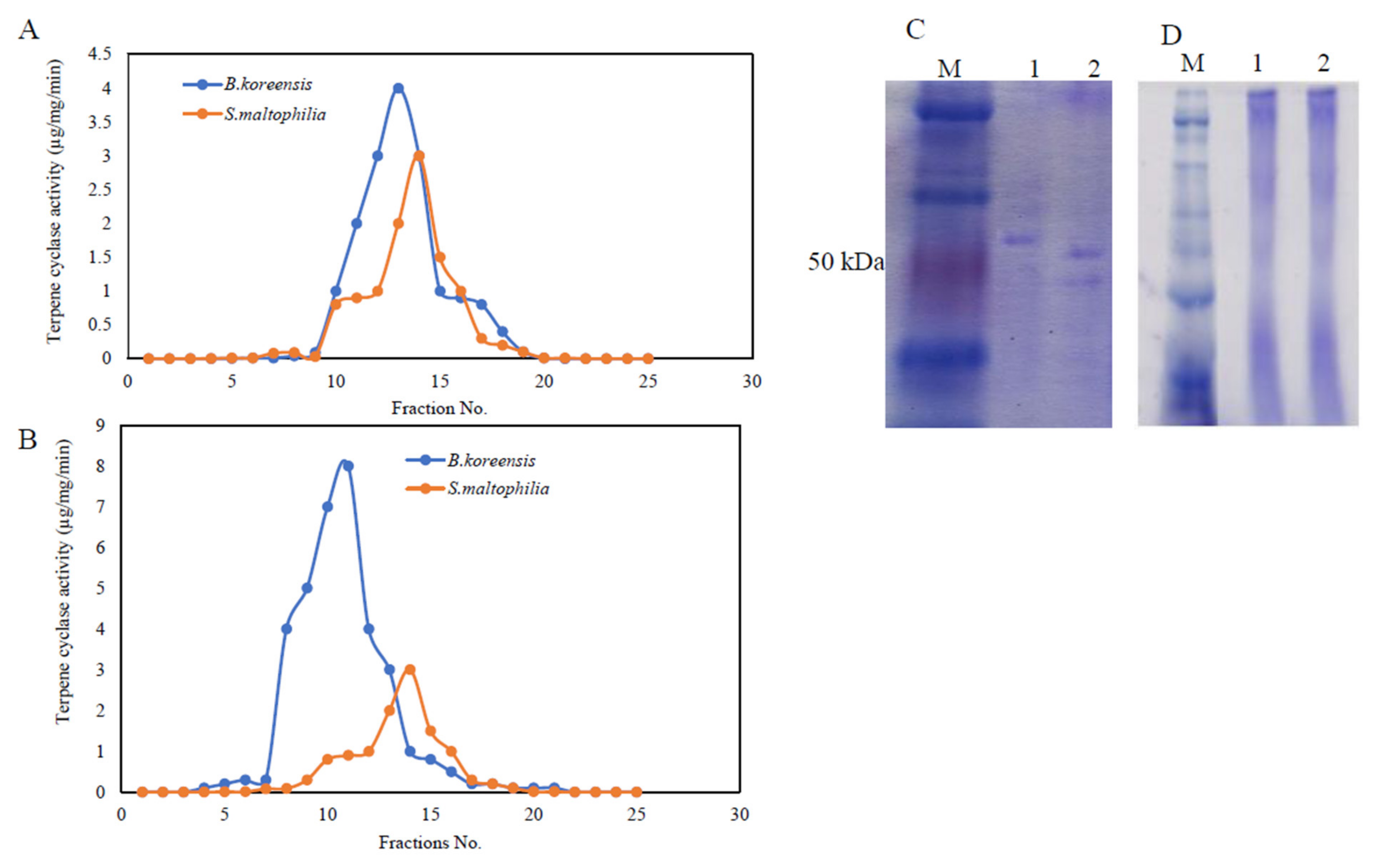
3.4. Purification, and Molecular Subunit Structure of TDS from S. maltophilia and B. koreensis
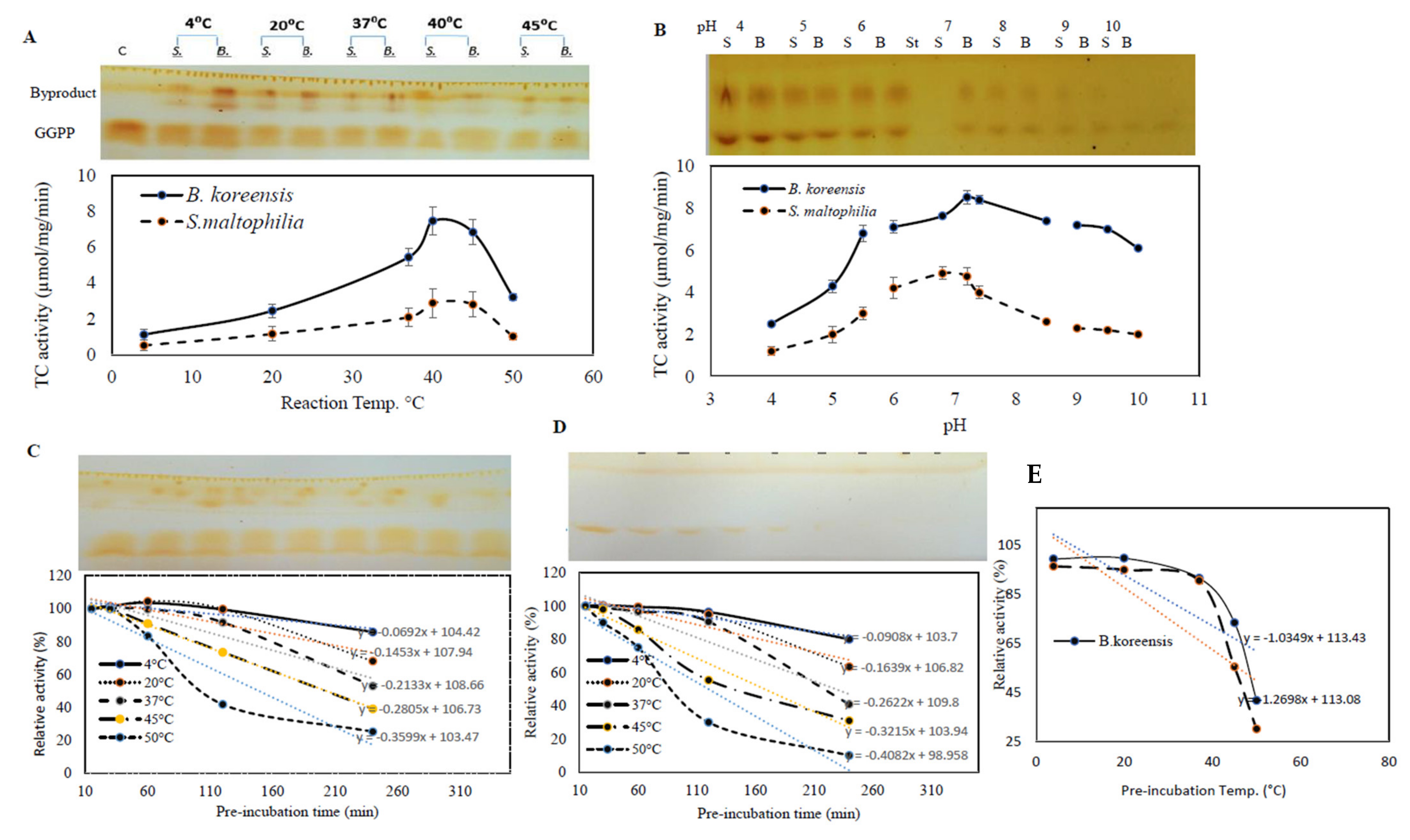
3.5. The Biochemical Properties of Purified TDS from Both Bacterial Isolates
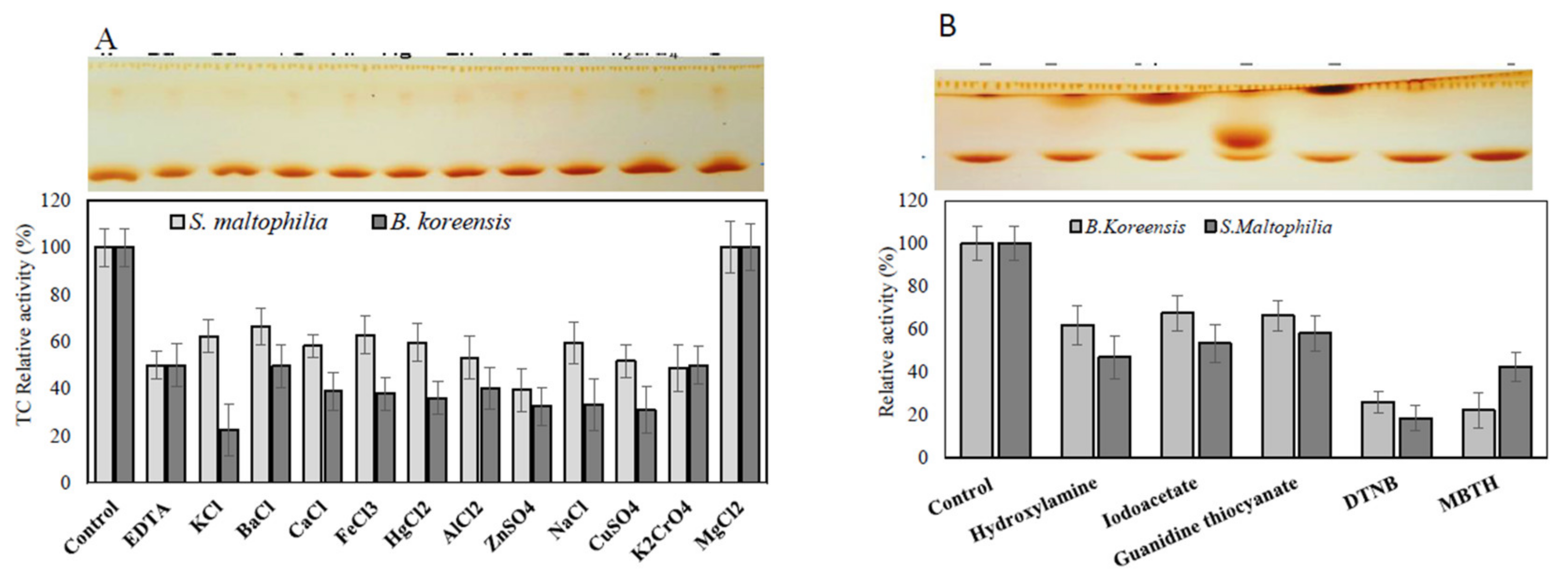
3.6. Effect of Inhibitors and Amino Acid Suicide Analogues on the Activity of TDS
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Suffness, M. Taxol: Science and Applications; CRC Press: Boca Raton, FL, USA, 1995; ISBN 9780849383823. [Google Scholar]
- Wani, M.C.; Taylor, H.L.; Wall, M.E.; Coggon, P.; McPhail, A.T. Plant Antitumor Agents. VI. The Isolation and Strcture of Taxol, a Novel Antileukemic and Antitumo Agent from Taxus bretvifolia. J. Am. Chem. Soc. 1971, 19, 2325–2327. [Google Scholar] [CrossRef]
- Hezari, M.; Lewis, N.G.; Croteau, R. Purification and Characterization of Taxa-4(5),11(12)-diene Synthase from Pacific Yew (Taxus Brevifolia) that Catalyzes the First Committed Step of Taxol Biosynthesis. Archiv. Biochem. Biophys. 1995, 322, 437–444. [Google Scholar] [CrossRef] [PubMed]
- Schiff, P.B.; Fant, J.; Horwitz, S.B. Promotion of microtubule assembly in vitro by taxol. Nature 1979, 277, 665–667. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; El-Sayed, M.T.; Rady, A.M.; Zein, N.; Enan, G.; Shindia, A.; El-Hefnawy, S.; Sitohy, M.; Sitohy, B. Exploiting the Biosynthetic Potency of Taxol from Fungal Endophytes of Conifers Plants; Genome Mining and Metabolic Manipulation. Molecules 2020, 25, 3000. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Ali, D.M.I.; Yassin, M.A.; Zayed, R.A.; Ali, G.S. Sterol inhibitor “Fluconazole” enhance the Taxol yield and molecular expression of its encoding genes cluster from Aspergillus flavipes. Process Biochem. 2019, 76, 55–67. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Fathalla, M.; Yassin, M.A.; Zein, N.; Morsy, S.; Sitohy, M.; Sitohy, B. Conjugation of Aspergillus flavipes taxol with porphyrin increases the anticancer activity of taxol and ameliorates its cytotoxic effects. Molecules 2020, 25, 263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Sayed, A.S.A.; Hassan, M.N.; Nada, H.M.S. Purification, immobilization, and biochemical characterization of l-arginine deiminase from thermophilic Aspergillus fumigatus KJ434941: Anticancer activity in vitro. Biotechnol. Progress 2015, 31, 396–405. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Mohamed, N.Z.; Safan, S.; Yassin, M.A.; Shaban, L.; Shindia, A.A.; Shad Ali, G.; Sitohy, M.Z. Restoring the Taxol biosynthetic machinery of Aspergillus terreus by Podocarpus gracilior Pilger microbiome, with retrieving the ribosome biogenesis proteins of WD40 superfamily. Sci. Rep. 2019, 9, 11534. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Sayed, A.S.A.; Abdel-Ghany, S.E.; Ali, G.S. Genome editing approaches: Manipulating of lovastatin and taxol synthesis of filamentous fungi by CRISPR/Cas9 system. Appl. Microbiol. Biotechnol. 2017, 101, 3953–3976. [Google Scholar] [CrossRef] [PubMed]
- Kusari, S.; Singh, S.; Jayabaskaran, C. Rethinking production of Taxol® (paclitaxel) using endophyte biotechnology. Trends Biotechnol. 2014, 32, 304–311. [Google Scholar] [CrossRef] [PubMed]
- Kusari, S.; Hertweck, C.; Spiteller, M. Chemical Ecology of Endophytic Fungi: Origins of Secondary Metabolites. Chem. Biol. 2012, 19, 792–798. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kusari, S.; Zühlke, S.; Spiteller, M. Effect of artificial reconstitution of the interaction between the plant camptotheca acuminata and the fungal endophyte fusarium solani on camptothecin biosynthesis. J. Nat. Prod. 2011, 74, 764–775. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; El Sayed, M.T.; Nada, H.S.; Hassan, A.E.; Yousef, E.K. Production and Characterization of Taxol as Anticancer Agent from Aspergillus terreus. J. Pure Appl. Microbiol. 2019, 13, 2055–2063. [Google Scholar] [CrossRef] [Green Version]
- El enshasy, H.A.; El-Baz, A.F.; Ammar, E.M. Simultaneous production and decomposition of different rifamycins during Amycolatopsis mediterranei in shake flask and stirred tank bioreactor. Commun. Current Res. Educ. Topics Trends Appl. Microbiol. 2005, 315–321. [Google Scholar]
- El-Sayed, A.S.; Shindia, A.A.; Zaher, Y. L-Amino acid oxidase from filamentous fungi: Screening and optimization. Annal. Microbiol. 2012, 62, 773–784. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Safan, S.; Mohamed, N.Z.; Shaban, L.; Ali, G.S.; Sitohy, M.Z. Induction of Taxol biosynthesis by Aspergillus terreus, endophyte of Podocarpus gracilior Pilger, upon intimate interaction with the plant endogenous microbes. Process Biochem. 2018, 71, 31–40. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Shindia, A.A.; Abouzeid, A.; Koura, A.; Hassanein, S.E.; Ahmed, R.M. Triggering the biosynthetic machinery of Taxol by Aspergillus flavipes via cocultivation with Bacillus subtilis: Proteomic analyses emphasize the chromatin remodeling upon fungal-bacterial interaction. Environ. Sci. Pollut. Res. 2021, 28, 39866–39881. [Google Scholar] [CrossRef] [PubMed]
- Dickschat, J.S. Bacterial Diterpene Biosynthesis. Angew. Chem. Int. Ed. 2019, 58, 15964–15976. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Tran, W.; Taatjes, C.A.; Alonso-Gutierrez, J.; Lee, T.S.; Gladden, J.M. Rapid discovery and functional characterization of terpene synthases from four endophytic xylariaceae. PLoS ONE 2016, 11, e0146983. [Google Scholar] [CrossRef] [Green Version]
- Prisic, S.; Xu, J.; Coates, R.M.; Peters, R.J. Probing the role of the DXDD motif in class II diterpene cyclases. ChemBioChem 2007, 8, 869–874. [Google Scholar] [CrossRef] [PubMed]
- Christianson, D.W. Structural and Chemical Biology of Terpenoid Cyclases. Chem. Rev. 2017, 117, 11570–11648. [Google Scholar] [CrossRef] [Green Version]
- Köksal, M.; Jin, Y.; Coates, R.M.; Croteau, R.; Christianson, D.W. Taxadiene synthase structure and evolution of modular architecture in terpene biosynthesis. Nature 2011, 469, 116–122. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, Q.; Roessner, C.A.; Croteau, R.; Scott, A.I. Engineering Escherichia coli for the synthesis of taxadiene, a key intermediate in the biosynthesis of taxol. Bioorg. Medic. Chem 2001, 9, 2237–2242. [Google Scholar] [CrossRef]
- DeJong, J.H.M.; Liu, Y.; Bollon, A.P.; Long, R.M.; Jennewein, S.; Williams, D.; Croteau, R.B. Genetic engineering of taxol biosynthetic genes in Saccharomyces cerevisiae. Biotechnol. Bioengin 2006, 93, 212–224. [Google Scholar] [CrossRef] [PubMed]
- Abdallah, I.I.; Pramastya, H.; Van Merkerk, R.; Sukrasno; Quax, W.J. Metabolic engineering of bacillus subtilis toward taxadiene biosynthesis as the first committed step for taxol production. Front. Microbiol. 2019, 10, 218. [Google Scholar] [CrossRef] [PubMed]
- ElMekawy, A.; Hegab, H.M.; El-Baz, A.; Hudson, S.M. Kinetic properties and role of bacterial chitin deacetylase in the bioconversion of chitin to chitosan. Recent Pat. Biotechnol. 2013, 7, 234–241. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Akbar, A.; Iqrar, I.; Ali, R.; Norman, D.; Brennan, M.; Ali, G.S. A glucanolytic Pseudomonas sp. associated with Smilax bona-nox L. displays strong activity against Phytophthora parasitica. Microbiol. Res. 2018, 207, 140–152. [Google Scholar] [CrossRef] [PubMed]
- El-Baz, A.F.; Shetaia, M.Y.; Elkhouli, R.R. Xylitol production by candida tropicalis under different statistically optimized growth conditions. Afr. J. Biotechnol. 2011, 10, 15353–15363. [Google Scholar] [CrossRef]
- Heinig, U.; Jennewein, S. Getting to the bottom of Taxol biosynthesis by fungi. Fungal Div. 2013, 60, 161–170. [Google Scholar] [CrossRef] [Green Version]
- Artz, J.D.; Wernimont, A.K.; Dunford, J.E.; Schapira, M.; Dong, A.; Zhao, Y.; Lew, J.; Russell, R.G.G.; Ebetino, F.H.; Oppermann, U. Molecular characterization of a novel geranylgeranyl pyrophosphate synthase from Plasmodium Parasites. J. Biolog. Chem. 2011, 286, 3315–3322. [Google Scholar] [CrossRef] [Green Version]
- El-Sayed, A.S.A.; Ali, G.S. Aspergillus flavipes is a novel efficient biocontrol agent of Phytophthora parasitica. Biol. Control 2020, 140, 104072. [Google Scholar] [CrossRef]
- Hassan, A.; Sorour, N.M.; El-Baz, A.; Shetaia, Y. Simple synthesis of bacterial cellulose/magnetite nanoparticles composite for the removal of antimony from aqueous solution. Int. J. Env. Sci. Technol. 2019, 16, 1433–1448. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Shindia, A.A.; Ali, G.S.; Yassin, M.A.; Hussein, H.; Awad, S.A.; Ammar, H.A. Production and bioprocess optimization of antitumor Epothilone B analogue from Aspergillus fumigatus, endophyte of Catharanthus roseus, with response surface methodology. Enzym. Microb. Technol. 2021, 143, 109718. [Google Scholar] [CrossRef]
- El-Mekawy, A.; Hudson, S.; El-Baz, A.F.; Hamza, H.; El-Halafawy, K. Preparation of Chitosan Films Mixed with Super-absorbent Polymer and Evaluation of Its Haemostatic and Antibacterial Activities. J. Appl. Polymer Sci. 2010, 116, 3489–3496. [Google Scholar]
- Edgar, R.C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 2004, 5, 113. [Google Scholar] [CrossRef] [Green Version]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Molec. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [Green Version]
- Maamoun, H.S.; Rabie, G.H.; Shaker, I.; Alaidaroos, B.A.; El-Sayed, A.S.A. Biochemical and kinetic properties of tyrosinase from Aspergillus terreus and Penicillium copticola; Undecanoic acid is a potent enzyme inhibitor from Aspergillus flavus, endophyte of Moringa oleifera. Molecules 2021, 26, 1309. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; George, N.M.; Yassin, M.A.; Alaidaroos, B.A.; Bolbol, A.A.; Mohamed, M.S.; Rady, A.M.; Aziz, S.W.; Zayed, R.A.; Sitohy, M.Z. Purification and Characterization of Ornithine Decarboxylase from Aspergillus terreus; Kinetics of Inhibition by Various Inhibitors. Molecules 2019, 24, 2756. [Google Scholar] [CrossRef] [Green Version]
- El-Sayed, A.S.A.; Shindia, A.A.; AbouZaid, A.A.; Yassin, A.M.; Ali, G.S.; Sitohy, M.Z. Biochemical characterization of peptidylarginine deiminase-like orthologs from thermotolerant Emericella dentata and Aspergillus nidulans. Enzym. Microb. Technol 2019, 124, 41–53. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Shindia, A.A.; Zeid, A.A.A.; Yassin, A.M.; Sitohy, M.Z.; Sitohy, B. Aspergillus nidulans thermostable arginine deiminase-Dextran conjugates with enhanced molecular stability, proteolytic resistance, pharmacokinetic properties and anticancer activity. Enzym. Microb. Technol. 2019, 131, 109432. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Abdel-Azeim, S.; Ibrahim, H.M.; Yassin, M.A.; Abdel-Ghany, S.E.; Esener, S.; Ali, G.S. Biochemical stability and molecular dynamic characterization of Aspergillus fumigatus cystathionine γ-lyase in response to various reaction effectors. Enzym. Microb. Technol. 2015, 81, 31–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- El-Baz, A.F.; Shetaia, Y.M.; Elkhouli, R.R. Kinetic behavior of Candida tropicalis during xylitol production using semisynthetic and hydrolysate based media. Afr. J. Biotechnol. 2011, 10, 16617–16625. [Google Scholar] [CrossRef] [Green Version]
- El-Baz, A.F.; Mohamed Sorour, N.; Shetaia, Y.M. Trichosporon jirovecii-mediated synthesis of cadmium sulfide nanoparticles. J. Basic Microbiol. 2016, 56, 520–530. [Google Scholar] [CrossRef]
- El-Sayed, A.S.; Khalaf, S.A.; Aziz, H.A. Characterization of homocysteine γ-lyase from submerged and solid cultures of Aspergillus fumigatus ASH (JX006238). J. Microbiol. Biotechnol. 2013, 23, 499–510. [Google Scholar] [CrossRef] [Green Version]
- El-Sayed, A.S.; Shindia, A.A. Characterization and immobilization of purified Aspergillus flavipes l-methioninase: Continuous production of methanethiol. J. Appl. Microbiol. 2011, 111, 54–69. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Hassan, A.E.A.; Shindia, A.A.; Mohamed, S.G.; Sitohy, M.Z. Aspergillus flavipes methionine γ-lyase-dextran conjugates with enhanced structural, proteolytic stability and anticancer efficiency. J. Mol. Catal. B Enzym. 2016, 133, S15–S24. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Ibrahim, H.; Sitohy, M.Z. Co-immobilization of PEGylated Aspergillus flavipes l-methioninase with glutamate dehydrogenase: A novel catalytically stable anticancer consortium. Enzym. Microb. Technol. 2014, 54, 59–69. [Google Scholar] [CrossRef] [PubMed]
- El-Sayed, A.S.A.; Ruff, L.E.; Ghany, S.E.A.; Ali, G.S.; Esener, S. Molecular and Spectroscopic Characterization of Aspergillus flavipes and Pseudomonas putida L-Methionine γ-Lyase in Vitro. Appl. Biochem. Biotechnol. 2017, 181, 1513–1532. [Google Scholar] [CrossRef]
- Kusari, S.; Lamshöft, M.; Spiteller, M. Aspergillus fumigatus Fresenius, an endophytic fungus from Juniperus communis L. Horstmann as a novel source of the anticancer pro-drug deoxypodophyllotoxin. J. Appl. Microbiol. 2009, 107, 1019–1030. [Google Scholar] [CrossRef] [PubMed]
- Netzker, T.; Fischer, J.; Weber, J.; Mattern, D.J.; König, C.C.; Valiante, V.; Schroeckh, V.; Brakhage, A.A. Microbial communication leading to the activation of silent fungal secondary metabolite gene clusters. Front. Microbiol. 2015, 6, 299. [Google Scholar] [CrossRef] [PubMed]
- Schroeckh, V.; Scherlach, K.; Nützmann, H.-W.; Shelest, E.; Schmidt-Heck, W.; Schuemann, J.; Martin, K.; Hertweck, C.; Brakhage, A.A. Intimate bacterial-fungal interaction triggers biosynthesis of archetypal polyketides in Aspergillus nidulans. Proc. Nat. Acad. Sci. USA 2009, 106, 14558–14563. [Google Scholar] [CrossRef] [Green Version]
- Soliman, S.S.M.; Raizada, M.N. Interactions between Co-Habitating fungi Elicit Synthesis of Taxol from an Endophytic Fungus in Host Taxus Plants. Front. Microbiol. 2013, 4, 3. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soliman, S.; Tang, Y. Natural and engineered production of taxadiene with taxadiene synthase. Biotechnol. Bioeng. 2015, 112, 229–235. [Google Scholar] [CrossRef]
- Pollak, F.C.; Berger, R.G. Geosmin and Related Volatiles in Bioreactor-Cultured Streptomyces citreus CBS 109.60. Appl. Environ. Microbiol. 1996, 62, 1295–1299. [Google Scholar] [CrossRef] [Green Version]
- Izaguirre, G.; Hwang, C.J.; Krasner, S.W.; McGuire, M.J. Geosmin and 2-Methylisoborneol from Cyanobacteria in Three Water Supply Systems. Appl. Environ. Microbiol 1982, 43, 708–714. [Google Scholar] [CrossRef] [Green Version]
- Dickschat, J.S.; Nawrath, T.; Thiel, V.; Kunze, B.; Müller, R.; Schulz, S. Biosynthesis of the off-flavor 2-methylisoborneol by the myxobacterium Nannocystis Exedens. Angew. Chem. 2007, 46, 8287–8290. [Google Scholar] [CrossRef] [PubMed]
- Yamada, Y.; Kuzuyama, T.; Komatsu, M.; Shin-ya, K.; Omura, S.; Cane, D.E.; Ikeda, H. Terpene synthases are widely distributed in bacteria. Proc. Nat. Acad. Sci. USA 2015, 112, 857–862. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Whittington, D.A.; Wise, M.L.; Urbansky, M.; Coates, R.M.; Croteau, R.B. Christianson, D.W. Bornyl diphosphate synthase: Structure and strategy for carbocation manipulation by a terpenoid cyclase. Proc. Nat. Acad. Sci. USA 2002, 99, 15375–15380. [Google Scholar] [CrossRef] [Green Version]
- Zhang, P.; Zhou, P.-P.; Yu, L.-J. An endophytic taxol-producing fungus from Taxus x media, Aspergillus candidus MD3. FEMS Microbiol. Lett. 2009, 293, 155–159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.; Wang, T.T.; Xu, Q.L.; Xiong, Y.; Zhang, L.; Han, H.; Xu, K.; Guo, W.J.; Xu, Q.; Tan, R.X. Genome Mining and Comparative Biosynthesis of Meroterpenoids from Two Phylogenetically Distinct Fungi. Angew. Chem Int. Ed. 2018, 57, 8184–8188. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Zhou, P.-P.; Yu, L.-J. An Endophytic Taxol-Producing Fungus from Taxus media, Cladosporium cladosporioides MD2. Curr. Microbiol. 2016, 59, 227–232. [Google Scholar] [CrossRef]
- Staniek, A.; Woerdenbag, H.; Kayser, O. Taxomyces andreanae: A Presumed Paclitaxel Producer Demystified? Planta Medica 2009, 75, 1561–1566. [Google Scholar] [CrossRef] [PubMed]
- Williams, D.C.; Carroll, B.J.; Jin, Q.; Rithner, C.D.; Lenger, S.R.; Floss, H.G.; Coates, R.M.; Williams, R.M.; Croteau, R. Intramolecular proton transfer in the cyclization of geranylgeranyl diphosphate to the taxadiene precursor of taxol catalyzed by recombinant taxadiene synthase. Chem. Biol. 2000, 7, 969–977. [Google Scholar] [CrossRef] [Green Version]
- Exposito, O.; Bonfill, M.; Moyano, E.; Onrubia, M.; Mirjalili, M.; Cusido, R.; Palazon, J. Biotechnological Production of Taxol and Related Taxoids: Current State and Prospects. Anti-Cancer Agents Med. Chem. 2009, 9, 109–121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Croteau, R.; Ketchum, R.E.B.; Long, R.M.; Kaspera, R.; Wildung, M.R. Taxol biosynthesis and molecular genetics. Phytochem. Rev. Proc. Phytochem. Soc. Eur. 2006, 5, 75–97. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Sayed, A.S.; Shouman, S.A.; Nassrat, H.M. Pharmacokinetics, immunogenicity and anticancer efficiency of Aspergillus flavipes l-methioninase. Enzym. Microb. Technol. 2012, 51, 200–210. [Google Scholar] [CrossRef]
- El-Sayed, A.S.A.; Khalaf, S.A.; Azez, H.A.; Sitohy, B.; El-Baz, A.F. Production, bioprocess optimization and anticancer activity of Camptothecin from Aspergillus terreus and Aspergillus flavus, endophytes of Ficus elastica. Process Biochem. 2021, 107, 59–73. [Google Scholar] [CrossRef]
- El-Sayed, A.S. Microbial l-methioninase: Production, molecular characterization, and therapeutic applications. Appl. Microbiol. Biotechnol. 2010, 86, 445–467. [Google Scholar] [CrossRef] [PubMed]
- El-Sayed, A.S.A.; Yassin, M.A.; Ibrahim, H. Coimmobilization of l -methioninase and glutamate dehydrogenase: Novel approach for l -homoalanine synthesis. Biotechnol. Appl. Biochem. 2015, 62, 514–522. [Google Scholar] [CrossRef] [PubMed]
| Code | Variables | Level | |
|---|---|---|---|
| −1 | 1 | ||
| X1 | Yeast extract (g/L) | 1 | 4 |
| X2 | Peptone (g/L) | 1 | 5 |
| X3 | Phenylalanine (g/L) | 1 | 4 |
| X4 | Tyrosine (g/L) | 1 | 4 |
| X5 | Tryptophan (g/L) | 1 | 4 |
| X6 | Glycine (g/L) | 1 | 4 |
| X7 | Valine (g/L) | 1 | 4 |
| X8 | Methionine (g/L) | 1 | 4 |
| X9 | Asparagine (g/L) | 1 | 4 |
| X10 | Cysteine (g/L) | 1 | 4 |
| X11 | Glutamic acid (g/L) | 1 | 4 |
| X12 | Beef extract (g/L) | 1 | 4 |
| X13 | Sucrose (g/L) | 1 | 5 |
| X14 | Glucose (g/L) | 1 | 5 |
| X15 | Xylose, (g/L) | 1 | 5 |
| X16 | NaCl (g/L) | 2 | 6 |
| X17 | Ammonium nitrate (g/L) | 1 | 4 |
| X18 | pH | 6 | 8 |
| X19 | Incubation period (day) | 3 | 6 |
| g/L | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Run | X1 | X2 | X3 | X4 | X5 | X6 | X7 | X8 | X9 | X10 | X11 | X12 | X13 | X14 | X15 | x16 | X17 | X18 | X19 | Predicted TDS Activity | |
| 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | 2.26 | 3.465 |
| 2 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1.8 | 2.85 |
| 3 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 2.27 | 3.55 |
| 4 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 3.78 | 5.85 |
| 5 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | 0.8 | 1.25 |
| 6 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | −1 | 6.98 | 10.92 |
| 7 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | 6.78 | 10.52 |
| 8 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 6.09 | 9.55 |
| 9 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | 7.08 | 10.95 |
| 10 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | 5.60 | 8.765 |
| 11 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 5.03 | 7.86 |
| 12 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 5.67 | 8.85 |
| 13 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | 2.58 | 3.95 |
| 14 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 11.72 | 18.3 |
| 15 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | 2.12 | 3.3 |
| 16 | −1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | 1 | 1 | 1 | −1 | 1 | 1.18 | 1.75 |
| 17 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 4.07 | 6.35 |
| 18 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | 5.56 | 8.68 |
| 19 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | 6.56 | 10.25 |
| 20 | 1 | 1 | −1 | −1 | 1 | 1 | 1 | 1 | −1 | 1 | −1 | 1 | −1 | −1 | −1 | −1 | 1 | 1 | −1 | 10.46 | 16.35 |
| Step | B. koreensis | S. maltophilia | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Total Protein (mg) | Total Activity (µmol/min) | Specific Activity (µmol/mg/min) | Yield (%) | Purification Fold | Total Protein (mg) | Total Activity (µmol/min) | Specific Activity (µmol/mg/min) | Yield (%) | Purification Fold | |
| Crude enzyme | 413.6 | 177.6 | 0.43 | 100 | 1 | 650 | 78.2 | 0.13 | 100 | 1 |
| Acetone precipitate | 87.5 | 60.9 | 0.7 | 34.3 | 1.6 | 586 | 39.5 | 0.54 | 52.6 | 4.48 |
| Sephadex-G200 | 32.3 | 40.6 | 1.26 | 22.9 | 2.9 | 23.9 | 30.5 | 1.29 | 50.0 | 10.6 |
| DEAE-Sepharose | 0.5 | 3.42 | 6.84 | 19.2 | 15.9 | 5.6 | 9.8 | 1.7 | 16.9 | 14.6 |
| °C | B. koreensis | S. maltophilia | ||||
|---|---|---|---|---|---|---|
| T1/2 (h) * | Kr (min) ** | Tm (°C) *** | T1/2 (h) * | Kr (min) ** | Tm (°C) *** | |
| 4 | 136.0 | 0.02 × 10−3 | 61.3 | 98 | 0.03 × 10−3 | 49.7 |
| 20 | 66.4 | 0.1 × 10−3 | 68 | 0.14 × 10−3 | ||
| 37 | 46 | 0.15 × 10−3 | 48 | 0.24 × 10−3 | ||
| 45 | 23 | 0.24 × 10 −3 | 24 | 0.30 × 10−3 | ||
| 50 | 10 | 0.29 × 10−3 | 11 | 0.32 × 10−3 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
El-Sayed, A.S.A.; Fathalla, M.; Shindia, A.A.; Rady, A.M.; El-Baz, A.F.; Morsy, Y.; Sitohy, B.; Sitohy, M. Purification and Biochemical Characterization of Taxadiene Synthase from Bacillus koreensis and Stenotrophomonas maltophilia. Sci. Pharm. 2021, 89, 48. https://doi.org/10.3390/scipharm89040048
El-Sayed ASA, Fathalla M, Shindia AA, Rady AM, El-Baz AF, Morsy Y, Sitohy B, Sitohy M. Purification and Biochemical Characterization of Taxadiene Synthase from Bacillus koreensis and Stenotrophomonas maltophilia. Scientia Pharmaceutica. 2021; 89(4):48. https://doi.org/10.3390/scipharm89040048
Chicago/Turabian StyleEl-Sayed, Ashraf S. A., Maher Fathalla, Ahmed A. Shindia, Amgad M. Rady, Ashraf F. El-Baz, Yara Morsy, Basel Sitohy, and Mahmoud Sitohy. 2021. "Purification and Biochemical Characterization of Taxadiene Synthase from Bacillus koreensis and Stenotrophomonas maltophilia" Scientia Pharmaceutica 89, no. 4: 48. https://doi.org/10.3390/scipharm89040048
APA StyleEl-Sayed, A. S. A., Fathalla, M., Shindia, A. A., Rady, A. M., El-Baz, A. F., Morsy, Y., Sitohy, B., & Sitohy, M. (2021). Purification and Biochemical Characterization of Taxadiene Synthase from Bacillus koreensis and Stenotrophomonas maltophilia. Scientia Pharmaceutica, 89(4), 48. https://doi.org/10.3390/scipharm89040048







