Bioinformatics Analysis Confirms the Target Protein Underlying Mitotic Catastrophe of 4T1 Cells under Combinatorial Treatment of PGV-1 and Galangin
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cells Culture
2.2. PGV-1 and Galangin
2.3. Sample Preparation
2.4. Cytotoxicity Assay
2.5. MayGrünwald-Giemsa Staining
2.6. Senescence-Associated β-Galactosidase Assay
2.7. Statistical Analysis
2.8. Target Prediction of Galangin and PGV-1 in TNBC
2.9. UALCAN Analysis
2.10. Oncolnc Database Analysis
2.11. Molecular Docking
3. Results
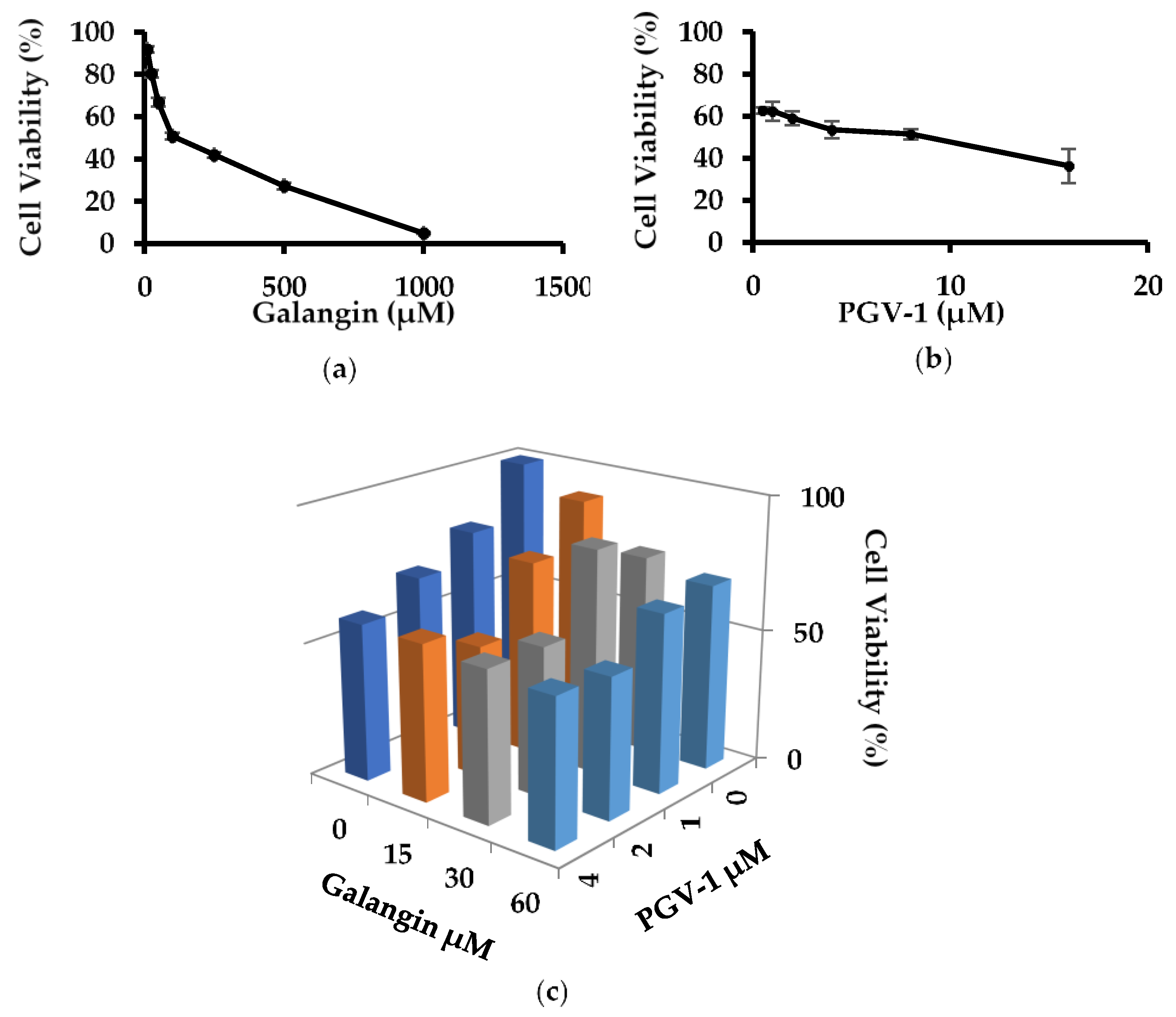
3.1. Anti-Proliferative Activity of Galangin and PGV-1 in TNBC
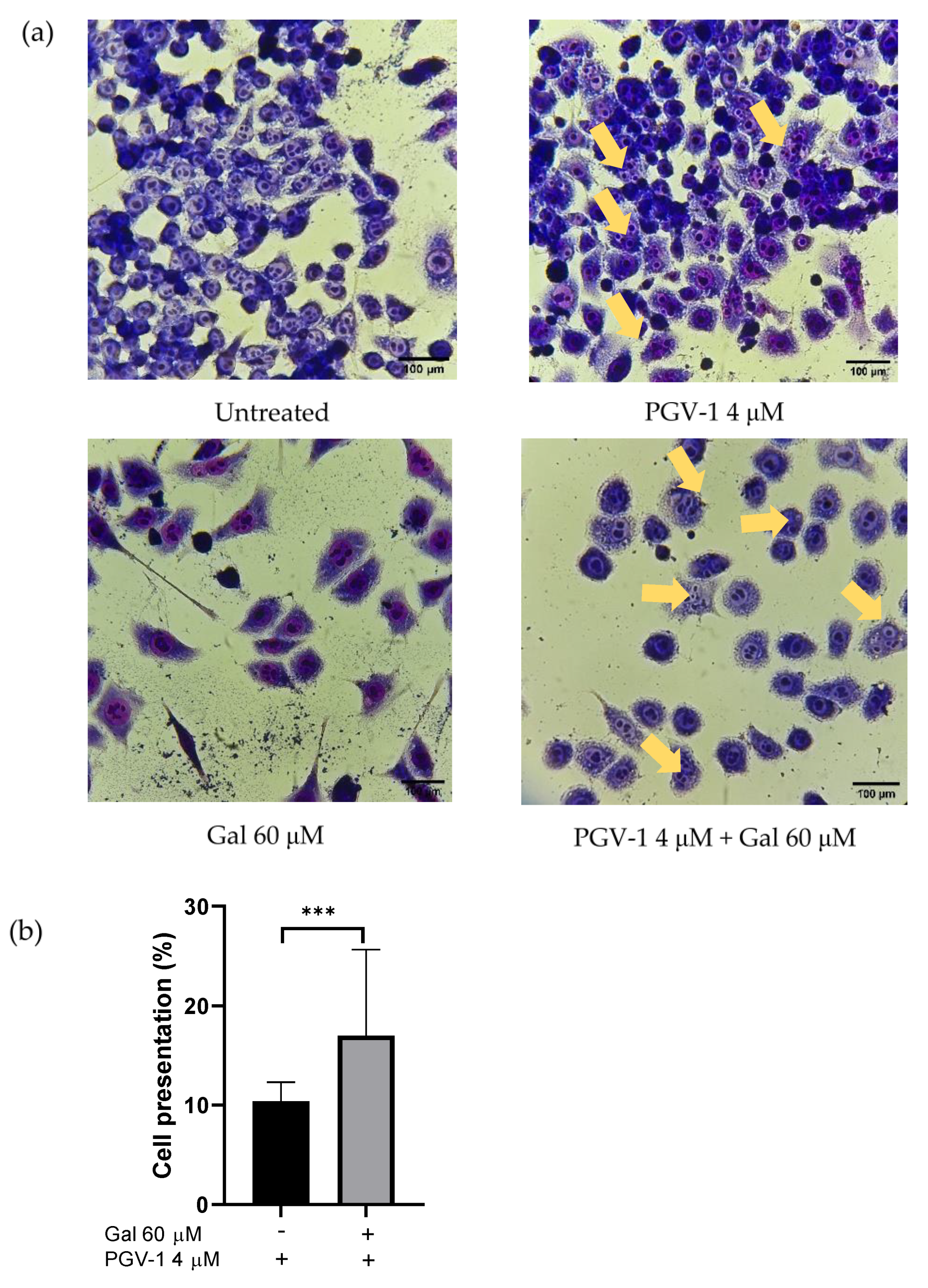
3.2. Effect of Galangin and PGV-1 on Cell Cycle
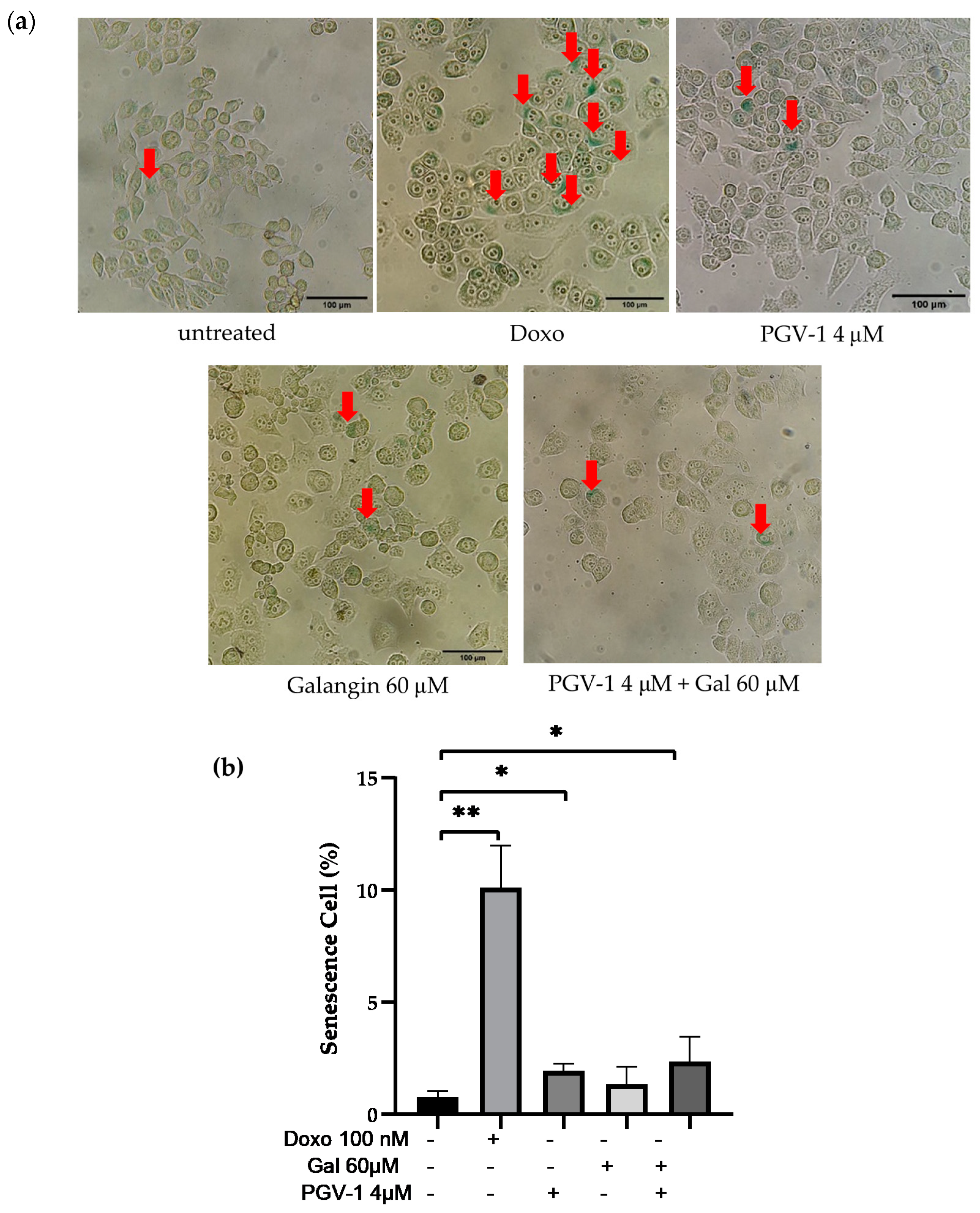
3.3. Galangin and Its Combination with PGV-1 Induces Cell Senescence
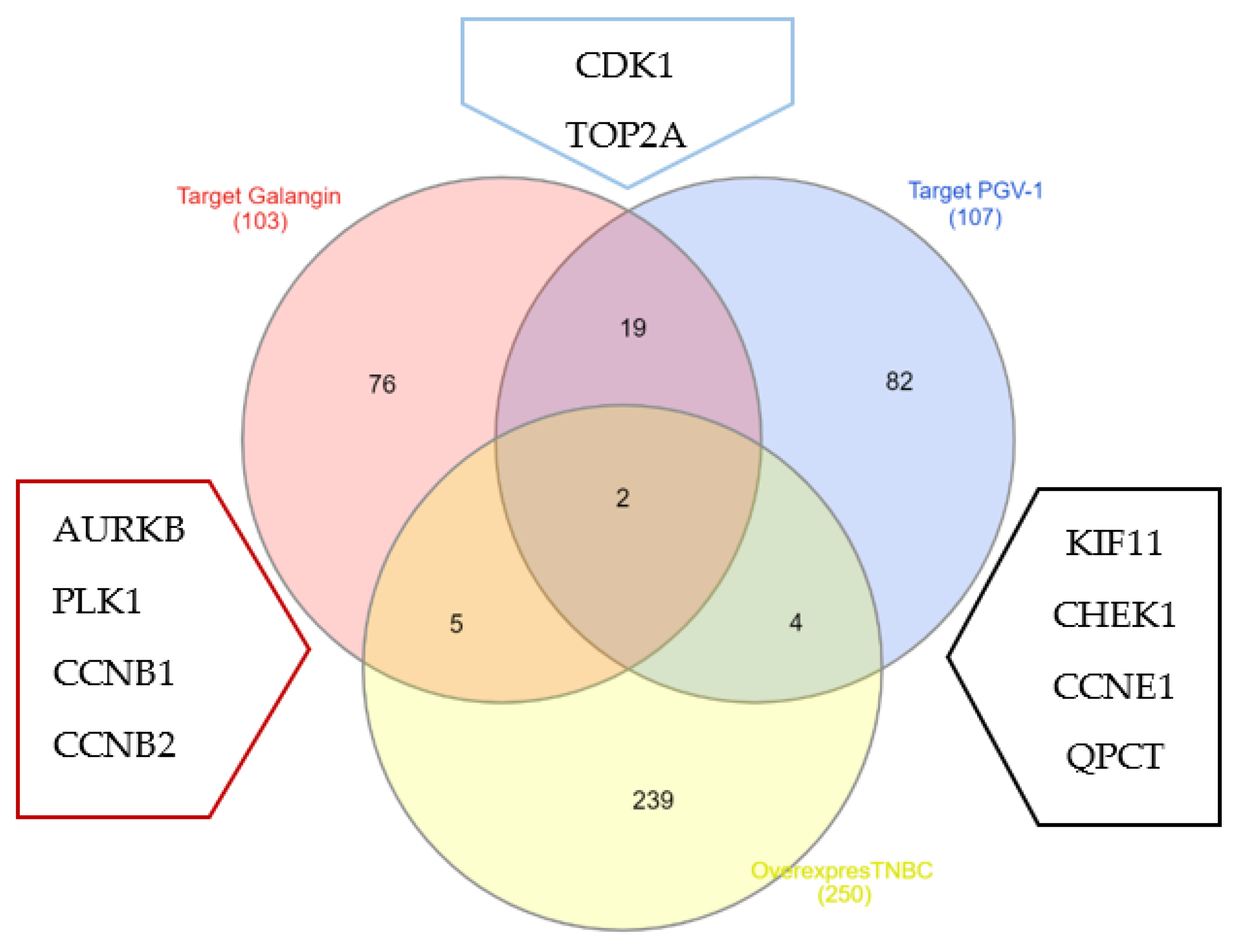
3.4. Galangin and PGV-1 Target Proteins Prediction on TNBC
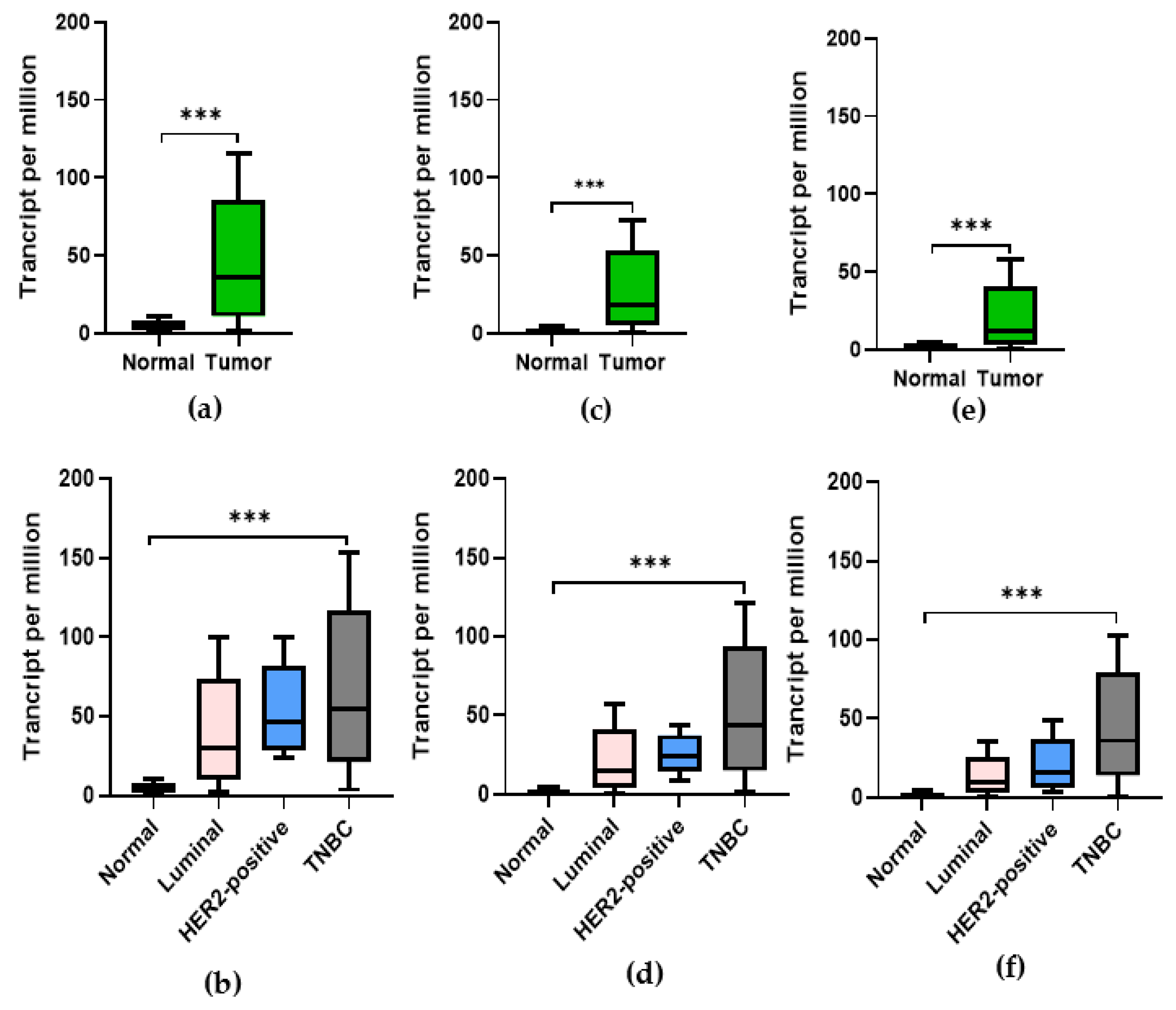
3.5. Expression of CDK1, PLK-1, and AURKB in Breast Cancer
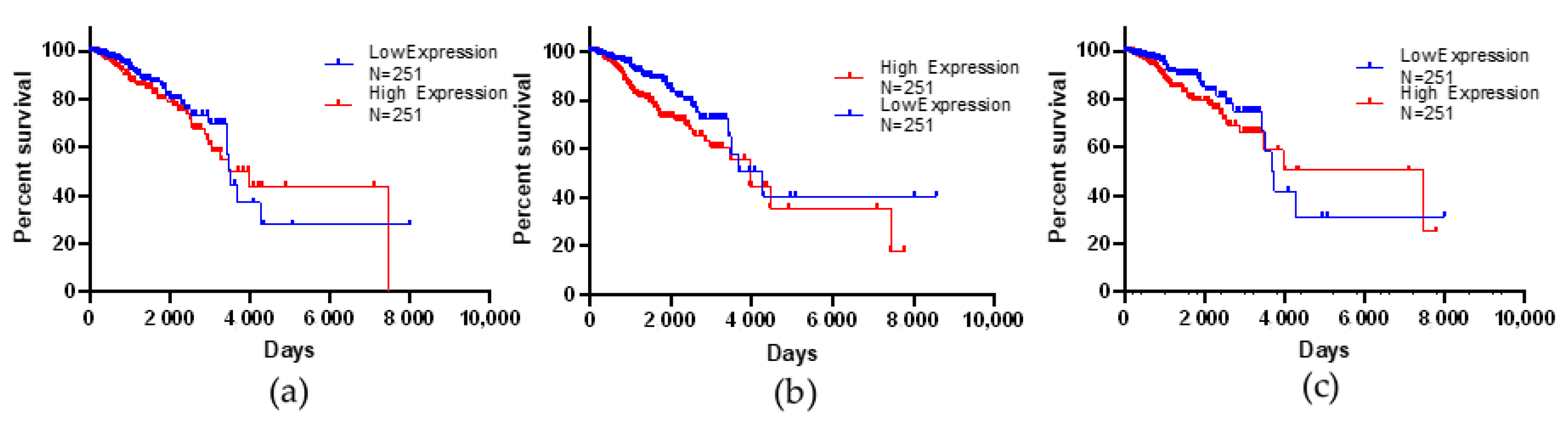
3.6. Kaplan–Meier Survival Analysis
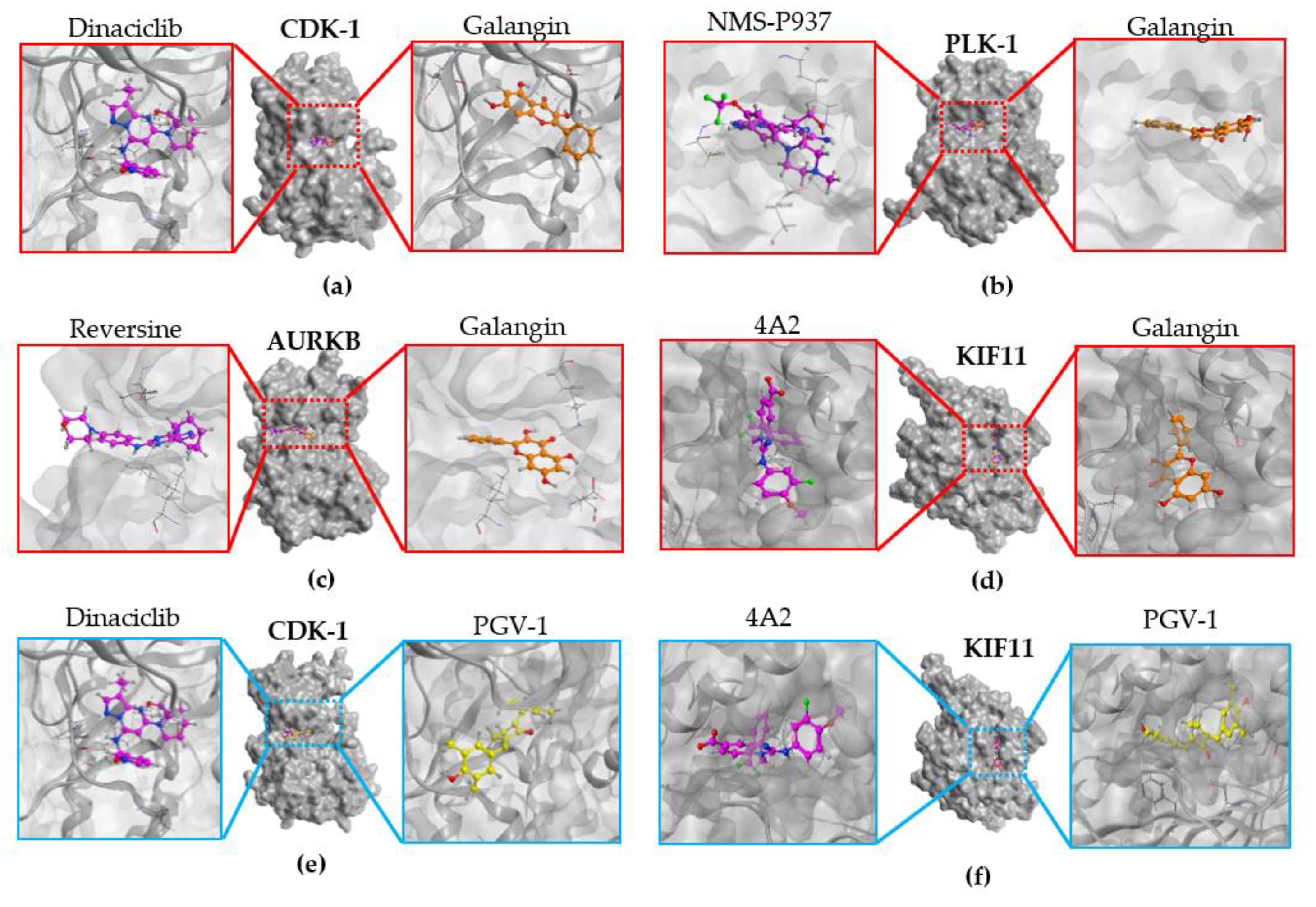
3.7. Molecular Docking
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Desai, A.; Qazi, G.; Ganju, R.; El-Tamer, M.; Singh, J.; Saxena, A.; Bedi, Y.; Taneja, S.; Bhat, H. Medicinal Plants and Cancer Chemoprevention. Curr. Drug Metab. 2008, 9, 581–591. [Google Scholar] [CrossRef] [Green Version]
- Zou, Y.; Li, R.; Kuang, D.; Zuo, M.; Li, W.; Tong, W.; Jiang, L.; Zhou, M.; Chen, Y.; Gong, W.; et al. Galangin Inhibits Cholangiocarcinoma Cell Growth and Metastasis through Downregulation of MicroRNA-21 Expression. BioMed Res. Int. 2020, 2020, 5846938. [Google Scholar] [CrossRef] [PubMed]
- Ren, K.; Zhang, W.; Wu, G.; Ren, J.; Lu, H.; Li, Z.; Han, X. Synergistic anti-cancer effects of galangin and berberine through apoptosis induction and proliferation inhibition in oesophageal carcinoma cells. Biomed. Pharmacother. 2016, 84, 1748–1759. [Google Scholar] [CrossRef]
- Liu, D.; You, P.; Luo, Y.; Yang, M.; Liu, Y. Galangin Induces Apoptosis in MCF-7 Human Breast Cancer Cells Through Mitochondrial Pathway and Phosphatidylinositol 3-Kinase/Akt Inhibition. Pharmacology 2018, 102, 58–66. [Google Scholar] [CrossRef]
- Song, W.; Yan, C.-Y.; Zhou, Q.-Q.; Zhen, L.-L. Galangin potentiates human breast cancer to apoptosis induced by TRAIL through activating AMPK. Biomed. Pharmacother. 2017, 89, 845–856. [Google Scholar] [CrossRef]
- Tolomeo, M.; Grimaudo, S.; Di Cristina, A.; Pipitone, R.M.; Dusonchet, L.; Meli, M.; Crosta, L.; Gebbia, N.; Invidiata, F.P.; Titone, L.; et al. Galangin increases the cytotoxic activity of imatinib mesylate in imatinib-sensitive and imatinib-resistant Bcr-Abl expressing leukemia cells. Cancer Lett. 2008, 265, 289–297. [Google Scholar] [CrossRef]
- Ravichandra, V.; Hanumantharayappa, B.; Papasani, V. Evaluation of Cardio Protective Activity of Galangin against Doxorubicin Induced Cardiomyopathy. Int. J. Pharm. Pharm. Sci. 2014, 6, 86–90. [Google Scholar]
- Nur, F.N.F.; Nugraheni, N.; Salsabila, I.A.; Haryanti, S.; Da’I, M.; Meiyanto, E.; Ahlina, F.N. Revealing the Reversal Effect of Galangal (Alpinia galanga L.) Extract against Oxidative Stress in Metastatic Breast Cancer Cells and Normal Fibroblast Cells Intended as a Co- Chemotherapeutic and Anti-Ageing Agent. Asian Pac. J. Cancer Prev. 2020, 21, 107–117. [Google Scholar] [CrossRef] [Green Version]
- Meiyanto, E.; Putri, H.; Larasati, Y.A.; Utomo, R.Y.; Jenie, R.I.; Ikawati, M.; Lestari, B.; Yoneda-Kato, N.; Nakamae, I.; Kawaichi, M.; et al. Anti-proliferative and Anti-metastatic Potential of Curcumin Analogue, Pentagamavunon-1 (PGV-1), Toward Highly Metastatic Breast Cancer Cells in Correlation with ROS Generation. Adv. Pharm. Bull. 2019, 9, 445–452. [Google Scholar] [CrossRef] [Green Version]
- Meiyanto, E.; Husnaa, U.; Kastian, R.F.; Putri, H.; Larasati, Y.A.; Khumaira, A.; Pamungkas, D.D.P.; Jenie, R.I.; Kawaichi, M.; Lestari, B.; et al. The Target Differences of Anti-Tumorigenesis Potential of Curcumin and its Analogues against HER-2 Positive and Triple-Negative Breast Cancer Cells. Adv. Pharm. Bull. 2020, 11, 188–196. [Google Scholar] [CrossRef]
- Lestari, B.; Nakamae, I.; Yoneda-Kato, N.; Morimoto, T.; Kanaya, S.; Yokoyama, T.; Shionyu, M.; Shirai, T.; Meiyanto, E.; Kato, J.-Y. Pentagamavunon-1 (PGV-1) inhibits ROS metabolic enzymes and suppresses tumor cell growth by inducing M phase (prometaphase) arrest and cell senescence. Sci. Rep. 2019, 9, 14867. [Google Scholar] [CrossRef] [Green Version]
- Fragomeni, S.M.; Sciallis, A.; Jeruss, J.S. Molecular Subtypes and Local-Regional Control of Breast Cancer. Surg. Oncol. Clin. N. Am. 2018, 27, 95–120. [Google Scholar] [CrossRef]
- Chalakur-Ramireddy, N.K.; Pakala, S.B. Combined drug therapeutic strategies for the effective treatment of Triple Negative Breast Cancer. Biosci. Rep. 2018, 38, BSR20171357. [Google Scholar] [CrossRef] [Green Version]
- Hermawan, A.; Fitriasari, A.; Junedi, S.; Ikawati, M. Pgv-0 and Pgv-1 Increased Apoptosis Induction Of Doxorubicin on Mcf-7 Breast Cancer Cells. Pharmacon 2011, 12, 55–59. [Google Scholar]
- Meiyanto, E.; Septisetyani, E.P.; Larasati, Y.A.; Kawaichi, M. Curcumin Analog Pentagamavunon-1 (PGV-1) Sensitizes Widr Cells to 5-Fluorouracil through Inhibition of NF-κB Activation. Asian Pac. J. Cancer Prev. 2018, 19, 49–56. [Google Scholar] [CrossRef]
- SwissTargetPrediction. Available online: http://www.swisstargetprediction.ch/ (accessed on 17 May 2021).
- Daina, A.; Michielin, O.; Zoete, V. SwissTargetPrediction: Updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 2019, 47, W357–W364. [Google Scholar] [CrossRef] [Green Version]
- Interactivenn. Available online: http://www.interactivenn.net/website (accessed on 21 May 2021).
- Ualcan. Available online: http://ualcan.path.uab.edu (accessed on 21 May 2021).
- Oncolnc. Available online: http://www.oncolnc.org/ (accessed on 21 May 2021).
- Anaya, J. OncoLnc: Linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. PeerJ Comput. Sci. 2016, 2, e67. [Google Scholar] [CrossRef] [Green Version]
- Baran, V.; Brzakova, A.; Rehák, P.; Kovarikova, V.; Solc, P. PLK1 regulates spindle formation kinetics and APC/C activation in mouse zygote. Zygote 2015, 24, 338–345. [Google Scholar] [CrossRef]
- Otto, T.; Sicinski, P. Cell Cycle Proteins as Promising Targets in Cancer Therapy. Nat. Rev. Cancer 2017, 17, 93–115. [Google Scholar] [CrossRef] [Green Version]
- Schatten, H. Mitosis. In Brenner’s Encyclopedia of Genetics; Academic Press: Waltham, MA, USA, 2013; pp. 448–451. ISBN 978-0-08-096156-9. [Google Scholar]
- Gong, D.; Ferrell, J.E. The Roles of Cyclin A2, B1, and B2 in Early and Late Mitotic Events. Mol. Biol. Cell 2010, 21, 3149–3161. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wendorff, T.; Schmidt, B.H.; Heslop, P.; Austin, C.A.; Berger, J.M. The Structure of DNA-Bound Human Topoisomerase II Alpha: Conformational Mechanisms for Coordinating Inter-Subunit Interactions with DNA Cleavage. J. Mol. Biol. 2012, 424, 109–124. [Google Scholar] [CrossRef] [Green Version]
- Fang, Y.; Zhang, X. Targeting NEK2 as a promising therapeutic approach for cancer treatment. Cell Cycle 2016, 15, 895–907. [Google Scholar] [CrossRef] [Green Version]
- Swarnkar, S.; Avchalumov, Y.; Raveendra, B.L.; Grinman, E.; Puthanveettil, S.V. Kinesin Family of Proteins Kif11 and Kif21B Act as Inhibitory Constraints of Excitatory Synaptic Transmission Through Distinct Mechanisms. Sci. Rep. 2018, 8, 17419. [Google Scholar] [CrossRef]
- Patil, M.; Pabla, N.; Dong, Z. Checkpoint kinase 1 in DNA damage response and cell cycle regulation. Cell. Mol. Life Sci. 2013, 70, 4009–4021. [Google Scholar] [CrossRef] [Green Version]
- Siu, K.T.; Rosner, M.R.; Minella, A.C. An integrated view of cyclin E function and regulation. Cell Cycle 2012, 11, 57–64. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Sun, B.; Zhu, F. Molecular characterization of glutaminyl-peptide cyclotransferase(QPCT)in Scylla paramamosain and its role in Vibrio alginolyticus and white spot syndrome virus (WSSV) infection. Fish Shellfish. Immunol. 2018, 78, 299–309. [Google Scholar] [CrossRef]
- Murray, T.J.; Yang, X.; Sherr, D.H. Growth of a human mammary tumor cell line is blocked by galangin, a naturally occurring bioflavonoid, and is accompanied by down-regulation of cyclins D3, E, and A. Breast Cancer Res. 2006, 8, R17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Denisenko, T.V.; Sorokina, I.V.; Gogvadze, V.; Zhivotovsky, B. Mitotic catastrophe and cancer drug resistance: A link that must to be broken. Drug Resist. Updat. 2016, 24, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Mc Gee, M.M. Targeting the Mitotic Catastrophe Signaling Pathway in Cancer. Mediat. Inflamm. 2015, 2015, 146282. [Google Scholar] [CrossRef] [Green Version]
- Vakifahmetoglu-Norberg, H.; Zhivotovsky, B. The unpredictable caspase-2: What can it do? Trends Cell Biol. 2010, 20, 150–159. [Google Scholar] [CrossRef] [PubMed]
- Vitale, I.; Galluzzi, L.; Castedo, M.; Kroemer, G. Mitotic catastrophe: A mechanism for avoiding genomic instability. Nat. Rev. Mol. Cell Biol. 2011, 12, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Masawang, K.; Pedro, M.; Cidade, H.; Reis, R.M.; Neves, M.P.; Corrêa, A.G.; Sudprasert, W.; Bousbaa, H.; Pinto, M.M. Evaluation of 2′,4′-dihydroxy-3,4,5-trimethoxychalcone as antimitotic agent that induces mitotic catastrophe in MCF-7 breast cancer cells. Toxicol. Lett. 2014, 229, 393–401. [Google Scholar] [CrossRef]
- García, I.A.; Garro, C.; Fernandez, E.; Soria, G. Therapeutic opportunities for PLK1 inhibitors: Spotlight on BRCA1-deficiency and triple negative breast cancers. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2020, 821, 111693. [Google Scholar] [CrossRef]

| Compound | IC50 (μM) |
|---|---|
| Galangin | 120 |
| PGV-1 | 8 |
| Galangin (μM) | PGV-1 (μM) | ||
|---|---|---|---|
| 1 | 2 | 4 | |
| 15 | −0.13 | 0.03 | −0.24 |
| 30 | 0.12 | 0.10 | 0.04 |
| 60 | 0.56 | 0.20 | 0.06 |
| Protein | Role | Function | Reference |
|---|---|---|---|
| PLK-1 | Protein Kinase | chromosome segregation, spindle assembly and cytokinesis | [22] |
| AURKB | Protein Kinase | chromosome-microtubule attachment | [23] |
| CDK1 | Protein Kinase | G2/M regulator | [24] |
| CCNB1 | Subunit regulator CDK1 | G2/M regulator | [25] |
| CCNB2 | Subunit regulator CDK1 | G2/M regulator | [25] |
| TOP2A | Enzim Katalis | Chromosome separation and DNA replication | [26] |
| NEK2 | Protein Kinase | Chromosome duplication and separation | [27] |
| Protein | Role | Function | Reference |
|---|---|---|---|
| CDK1 | Protein Kinase | G2/M regulation | [24] |
| TOP2A | Enzim Katalis | Chromosome separation and DNA replication | [26] |
| KIF11 | Enzim Katalis | Forms bioplar mitotic spindles | [28] |
| CHK1 | Protein Kinase | DDR mediator | [29] |
| CCNE1 | Protein Kinase | G1/S regulator | [30] |
| QPCT | Enzim | Biosynthesis of piroglutamil peptida | [27] |
| No. | Protein | PDB ID | Native Ligand | Galangin | ||
|---|---|---|---|---|---|---|
| RMSD (Å) | Docking Score (kcal/mol) | RMSD (Å) | Docking Score (kcal/mol) | |||
| 1 | PLK-1 | 2YAC | 0.8535 | −13.9847 | 0.7247 | −13.5717 |
| 2 | AURKB | 2VGO | 1.3148 | −7.8545 | 1.7724 | −9.0003 |
| 3 | CDK-1 | 6GU6 | 1.2543 | −10.8300 | 1.6107 | −12.2080 |
| 4 | KIF11 | 3ZCW | 0.8005 | −13.6193 | 1.1531 | −13.4846 |
| No. | Protein | PDB ID | Native Ligand | PGV-1 | ||
|---|---|---|---|---|---|---|
| RMSD (Å) | Docking Score (kcal/mol) | RMSD (Å) | Docking Score (kcal/mol) | |||
| 1 | CDK-1 | 6GU6 | 1.2543 | −10.8300 | 1.2612 | −14.0391 |
| 2 | KIF11 | 3ZCW | 0.8005 | −13.6193 | 0.8661 | −13.3464 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hasbiyani, N.A.F.; Wulandari, F.; Nugroho, E.P.; Hermawan, A.; Meiyanto, E. Bioinformatics Analysis Confirms the Target Protein Underlying Mitotic Catastrophe of 4T1 Cells under Combinatorial Treatment of PGV-1 and Galangin. Sci. Pharm. 2021, 89, 38. https://doi.org/10.3390/scipharm89030038
Hasbiyani NAF, Wulandari F, Nugroho EP, Hermawan A, Meiyanto E. Bioinformatics Analysis Confirms the Target Protein Underlying Mitotic Catastrophe of 4T1 Cells under Combinatorial Treatment of PGV-1 and Galangin. Scientia Pharmaceutica. 2021; 89(3):38. https://doi.org/10.3390/scipharm89030038
Chicago/Turabian StyleHasbiyani, Nurul Awali Fauziyah, Febri Wulandari, Eri Prasetyo Nugroho, Adam Hermawan, and Edy Meiyanto. 2021. "Bioinformatics Analysis Confirms the Target Protein Underlying Mitotic Catastrophe of 4T1 Cells under Combinatorial Treatment of PGV-1 and Galangin" Scientia Pharmaceutica 89, no. 3: 38. https://doi.org/10.3390/scipharm89030038
APA StyleHasbiyani, N. A. F., Wulandari, F., Nugroho, E. P., Hermawan, A., & Meiyanto, E. (2021). Bioinformatics Analysis Confirms the Target Protein Underlying Mitotic Catastrophe of 4T1 Cells under Combinatorial Treatment of PGV-1 and Galangin. Scientia Pharmaceutica, 89(3), 38. https://doi.org/10.3390/scipharm89030038






