Automatic ECG Diagnosis Using Convolutional Neural Network
Abstract
1. Introduction
2. ECG Signal and Dataset
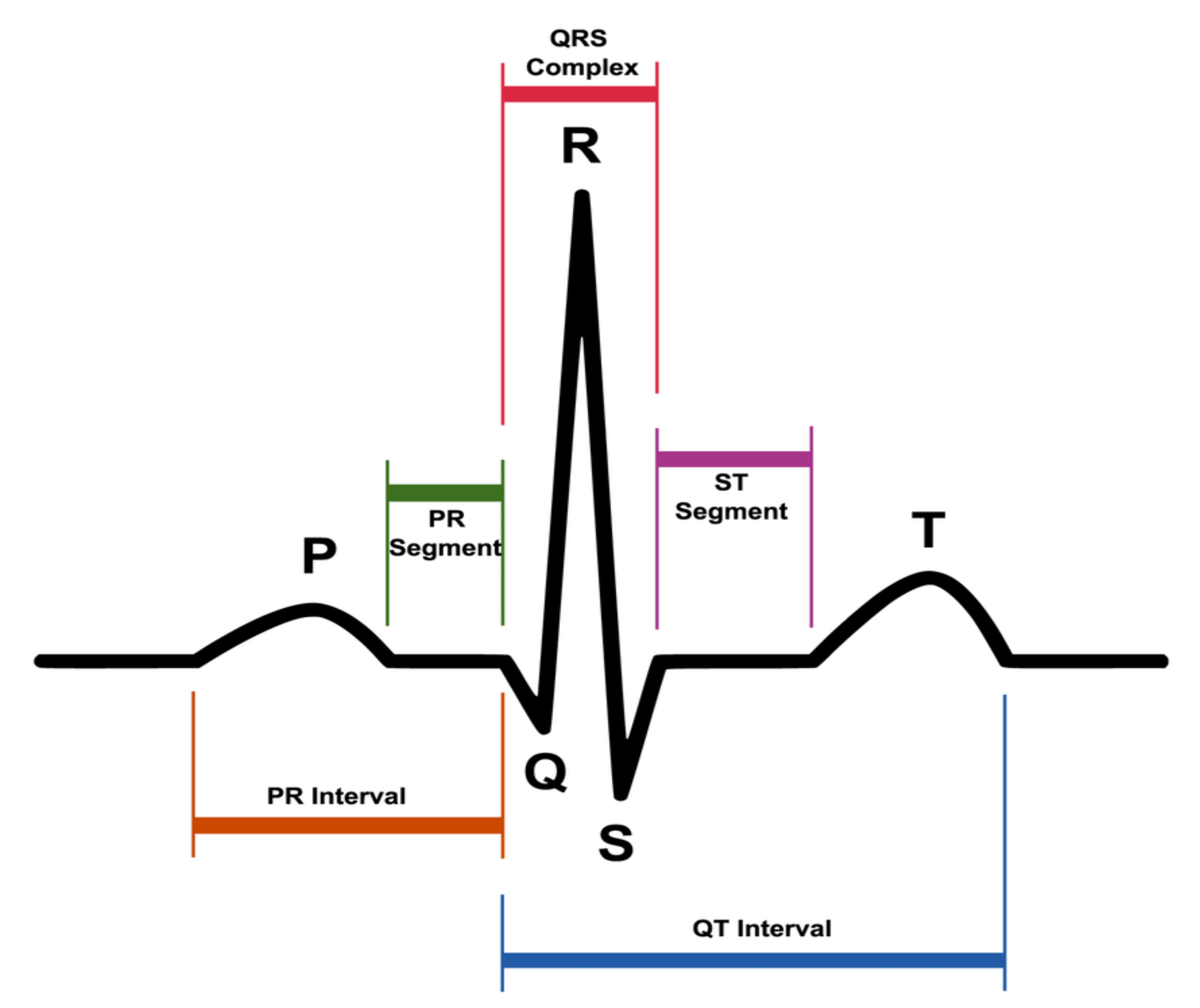
- P wave: the first wave that occurs in the ECG cycle, a small deflection that represents atrial depolarization or most commonly called “atrial contraction”;
- T wave: represents the depolarization of ventricles or most commonly called “ventricular relaxation”;
- Q, R, and S waves: together, these waves form the so-called QRS complex. The QRS complex represents the contraction of the ventricles or, technically speaking, the depolarization complex of the ventricles. In particular, the Q wave represents the depolarization of the interventricular septum, the R wave reflects the depolarization of the main mass of the ventricles, and the S wave is the final depolarization of the ventricles at the base of the heart.
- PR interval or PQ interval: the PR interval is a stretch formed by the P wave and the PR segment (rectilinear stretch) that begins with the P wave, that is, during the first deflection, and ends at the QRS complex. This interval indicates the time that the depolarization wave takes propagating from the atrial sinus node along the part of the electrical conduction system of the heart present on the myocardium;
- ST segment, i.e., the time between the end of the QRS complex and the start of the T wave;
- QT interval, i.e., the time between the beginning of the QRS complex and the end of the T wave, which is the electrocardiographic manifestation of ventricular depolarization and repolarization [23].
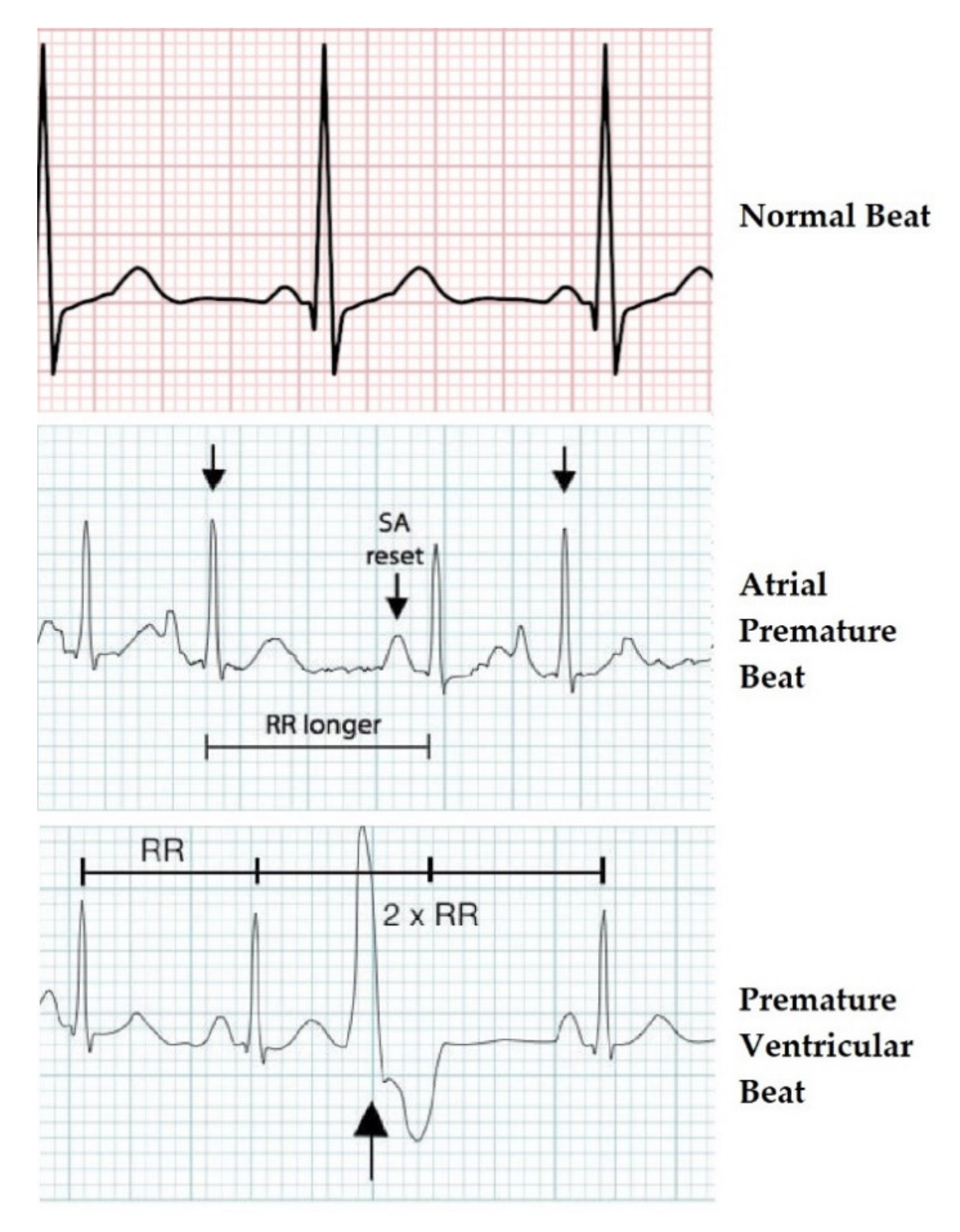
- Normal;
- Atrial premature beat;
- Premature ventricular contraction.
3. ECG Diseases Classification Based on CNN
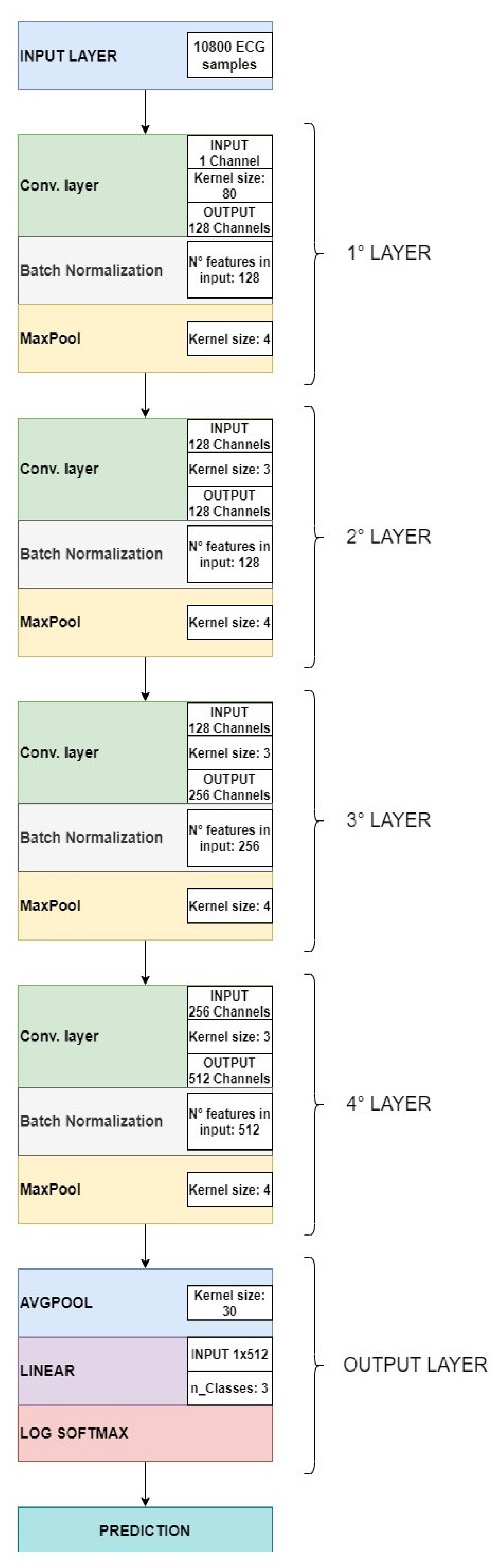
3.1. CNN General Characteristics and Architecture Adopted
- 1D convolution layers;
- Batch normalization layers;
- ReLU (Rectified Linear Units) layers;
- Pooling layers;
- Softmax.
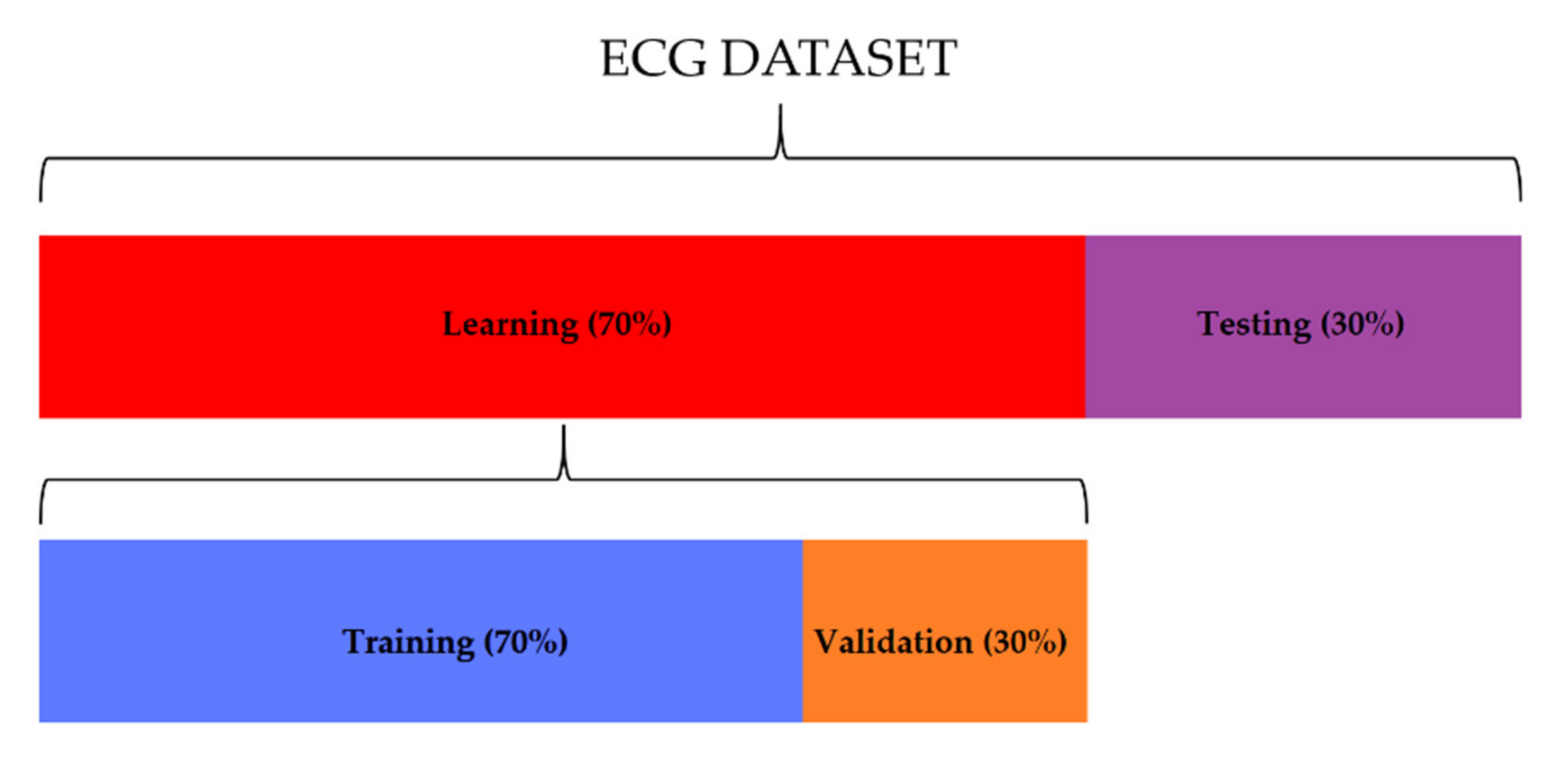
3.2. Training/Validation and Testing Dataset
- “Normal” class, containing 1421 ECG segments;
- “Premature ventricular contraction” class, containing 335 ECG segments;
- “Atrial premature beat” class, containing 133 ECG segments.
- Training/validation set, consisting of 995 segments for the “normal” class, 234 segments for the “premature ventricular contraction” class, and 93 segments for the “atrial premature beat” class. The 70% of this set was used for the training, and the other 30% was used for the testing;
- Testing set, consisting of 426 segments for the “normal” class, 101 segments for the “premature ventricular contraction” class, and 40 segments for the “atrial premature beat” class.
4. Methods
5. Performance Analysis
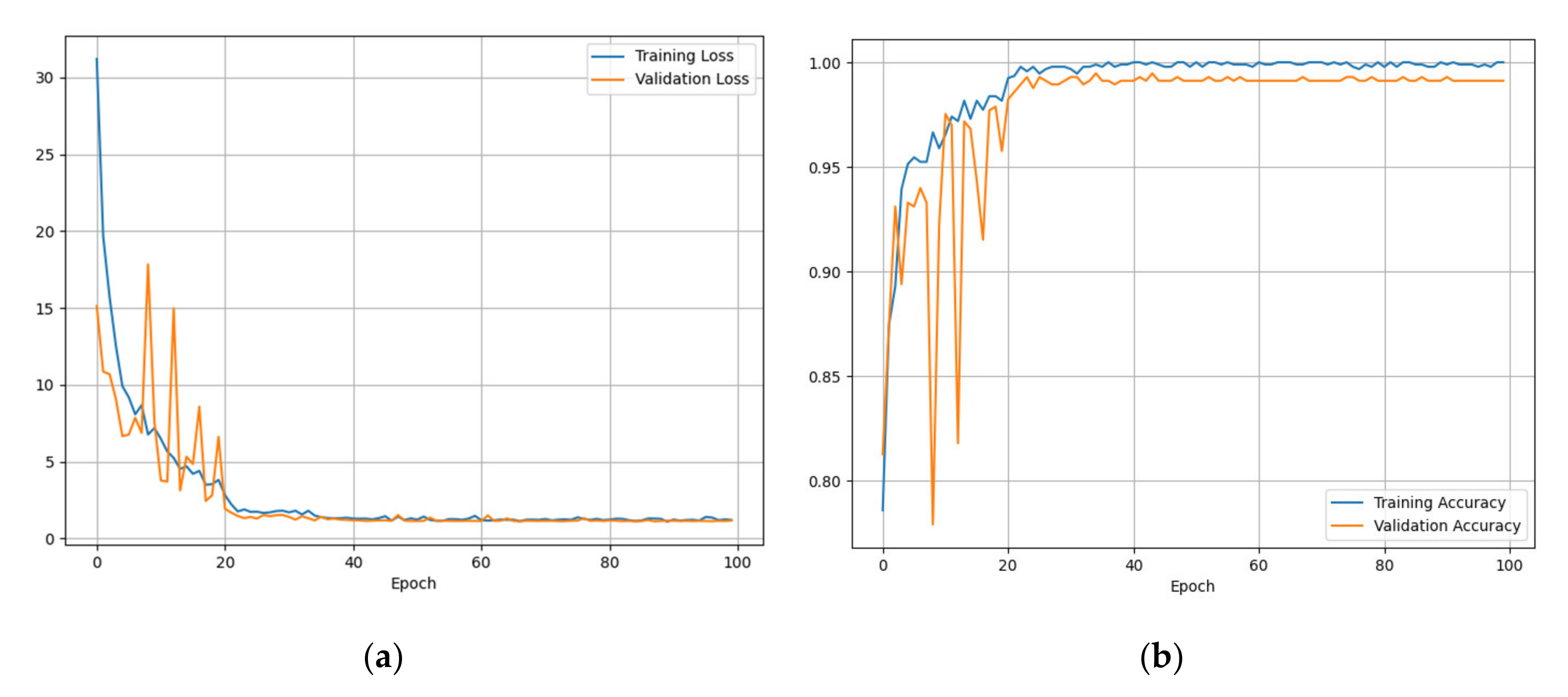
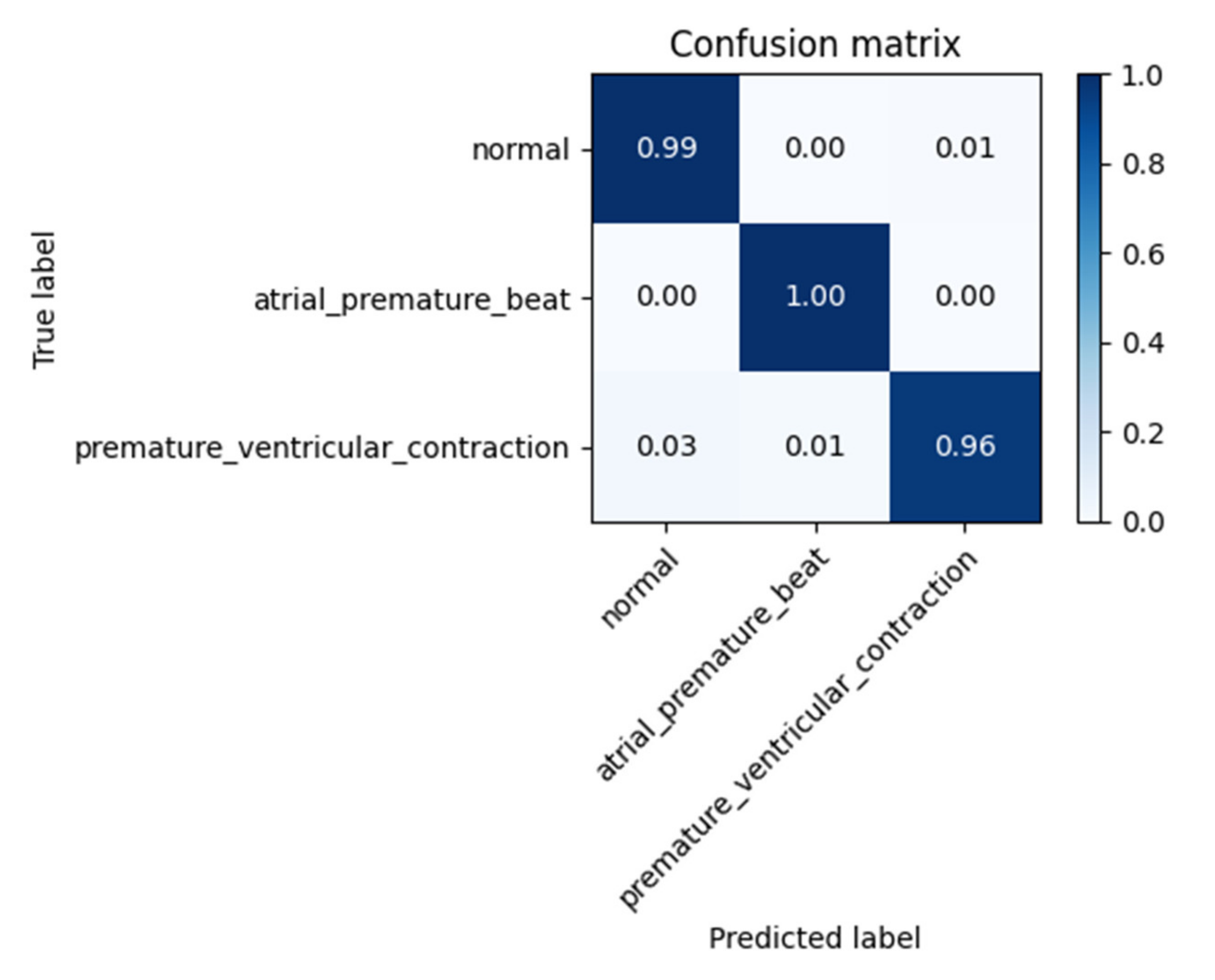
5.1. Test Results
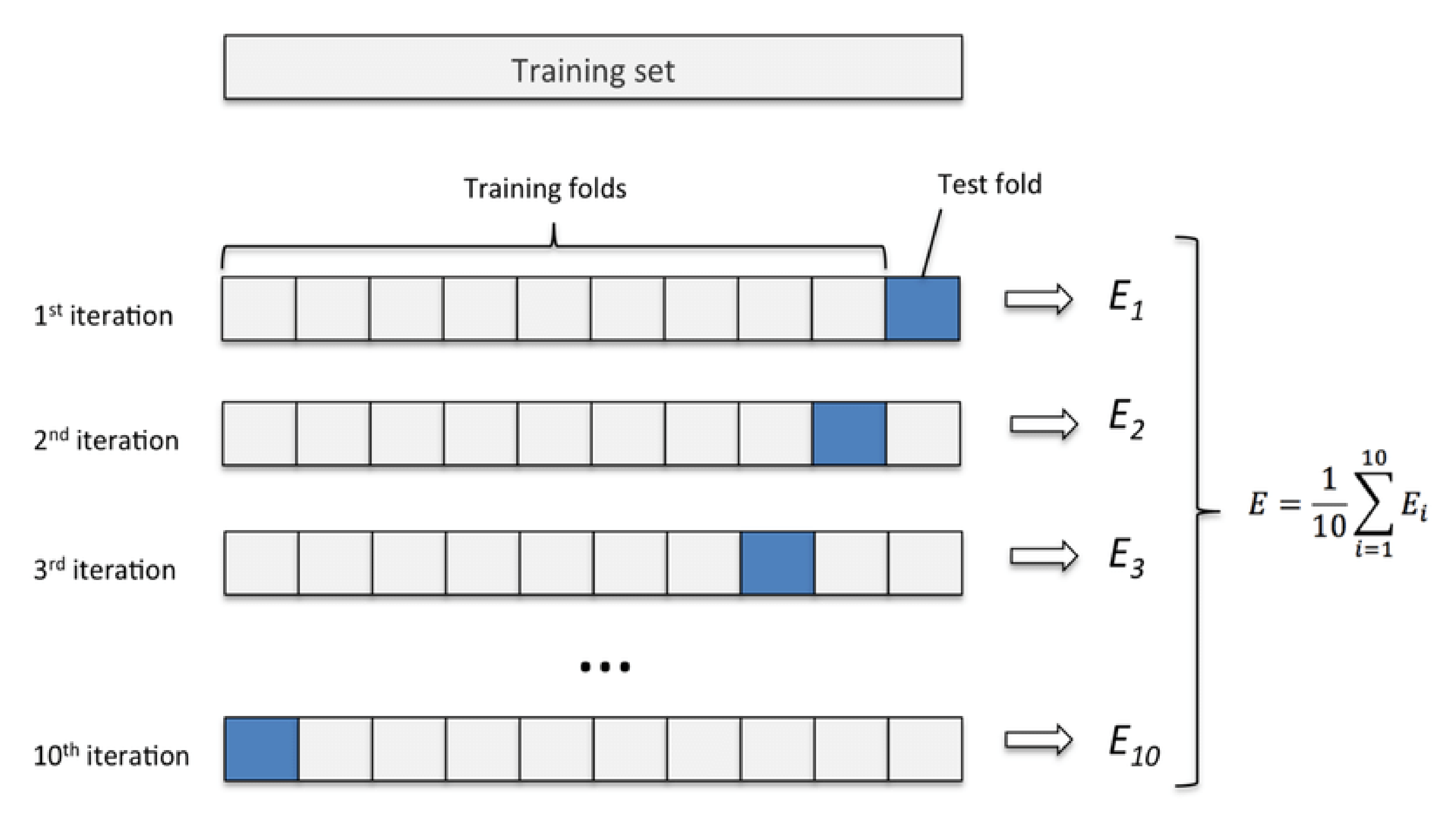
5.2. Cross-Validation Analysis
6. Discussion
7. Conclusions
- 98.33% mean accuracy;
- 98.33% sensitivity;
- 98.35% specificity;
- 1.65% false positive ratio;
- 1.66% false negative ratio;
- 98.33% F1 score.
Author Contributions
Funding
Conflicts of Interest
References
- Benjamin, E.J.; Blaha, M.J.; Chiuve, S.E.; Cushman, M.; Das, S.R.; Deo, R.; de Ferranti, S.D.; Floyd, J.; Fornage, M.; Gillespie, C.; et al. Heart disease and stroke statistics 2017 update: A report from the american heart association. Circulation 2017, 135, 146–603. [Google Scholar] [CrossRef]
- Mitra, M.; Samanta, R. Cardiac arrhythmia classification using neural networks with selected features. Proc. Technol. 2013, 10, 76–84. [Google Scholar] [CrossRef]
- Salerno, S.M.; Alguire, P.C.; Waxman, H.S. Competency in interpretation of 12-lead electrocardiograms: A summary and appraisal of published evidence. Ann. Intern. Med. 2003, 138, 751–760. [Google Scholar] [CrossRef]
- Beritelli, F.; Capizzi, G.; Lo Sciuto, G.; Napoli, C.; Woźniak, M. A novel training method to preserve generalization of RBPNN classifiers applied to ECG signals diagnosis. Neural Netw. 2018, 108, 131–138. [Google Scholar] [CrossRef]
- Beritelli, F.; Capizzi, G.; Lo Sciuto, G.; Scaglione, F.; Połap, D.; Woźniak, M. A Neural Network Pattern Recognition Approach to Automatic Rainfall Classification by Using Signal Strength in LTE/4G Networks. In Proceedings of the International Joint Conference on Rough Sets, Olsztyn, Poland, 3–7 July 2017. [Google Scholar]
- Beritelli, F.; Capizzi, G.; Lo Sciuto, G.; Napoli, C.; Scaglione, F. Rainfall estimation based on the intensity of the received signal in a LTE/4G mobile terminal by using a probabilistic neural network. IEEE Access 2018, 6, 30865–30873. [Google Scholar] [CrossRef]
- Avanzato, R.; Beritelli, F.; Di Franco, F.; Puglisi, V.F. A Convolutional Neural Networks approach to Audio Classification for Rainfall Estimation. In Proceedings of the 10th IEEE International Conference on Intelligent Data Acquisition and Advanced Computing Systems: Technology and Applications, Metz, France, 18–21 September 2019. [Google Scholar]
- Beritelli, F.; Spadaccini, A. A Statistical Approach to Biometric Identity Verification based on Heart Sounds. In Proceedings of the Fourth International Conference on Emerging Security Information, Systems and Technologies, Venice, Italy, 18–25 July 2010. [Google Scholar]
- Beritelli, F.; Spadaccini, A. The Role of Voice Activity Detection in Forensic Speaker Verification. In Proceedings of the 17th IEEE International Conference on Digital Signal Processing (DSP 2011), Corfu Island, Greece, 6–9 July 2011. [Google Scholar]
- Rajpurkar, P.; Hannun, A.Y.; Haghpanahi, M.; Bourn, C.; Ng, A.Y. Cardiologist-level arrhythmia detection with convolutional neural networks. arXiv 2017, arXiv:1707.01836. Available online: https://arxiv.org/abs/1707.01836 (accessed on 5 June 2020).
- Acharya, U.R.; Fujita, H.; Lih, O.S.; Adam, M.; Tan, J.H.; Chua, C.K. Automated detection of coronary artery disease using different durations of ECG segments with convolutional neural network. Knowl. Based Syst. 2017, 132, 62–71. [Google Scholar] [CrossRef]
- Kiranyaz, S.; Ince, T.; Gabbouj, M. Real-time patient-specific ECG classification by 1-D convolutional neural networks. IEEE Trans. Biomed. Eng. 2016, 63, 664–675. [Google Scholar] [CrossRef]
- Taji, B.; Chan, A.D.; Shirmohammadi, S. Classifying measured electrocardiogram signal quality using deep belief networks. In Proceedings of the IEEE International Instrumentation and Measurement Technology Conference (I2MTC), Turin, Italy, 22–25 May 2017. [Google Scholar]
- Acharya, U.R.; Fujita, H.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adam, M. Application of deep convolutional neural network for automated detection of myocardial infarction using ECG signals. Inf. Sci. 2017, 415, 190–198. [Google Scholar] [CrossRef]
- Savalia, S.; Emamian, V. Cardiac Arrhythmia classification by multi-layer perceptron and convolution neural networks. Bioengineering 2018, 5, 35. [Google Scholar] [CrossRef]
- Zubair, M.; Kim, J.; Yoon, C. An Automated ECG beat classification system using convolutional neural networks. In Proceedings of the 6th International Conference on IT Convergence and Security (ICITCS), Prague, Czech Republic, 26 September 2016; pp. 1–5. [Google Scholar]
- Li, D.; Zhang, J.; Zhang, Q.; Wei, X. Classification of ECG signals based on 1D convolution neural network. In Proceedings of the IEEE 19th International Conference on e-Health Networking, Applications and Services (Healthcom), Dalian, China, 12–15 October 2017. [Google Scholar]
- Baloglu, U.B.; Talo, M.; Yildirim, O.; Tan, R.S.; Acharya, U.R. Classification of myocardial infarction with multi-lead ECG signals and deep CNN. Pattern Recognit. Lett. 2019, 122, 23–30. [Google Scholar] [CrossRef]
- Tang, X.; Shu, L. Classification of electrocardiogram signals with RS and quantum neural networks. Int. J. Multimedia Ubiquitous Eng. 2014, 9, 363–372. [Google Scholar] [CrossRef]
- Moavenian, M.; Khorrami, H. A qualitative comparison of artificial neural networks and support vector machines in ECG arrhythmias classification. Expert Syst. Appl. 2010, 37, 3088–3093. [Google Scholar] [CrossRef]
- Jambukia, S.H.; Dabhi, V.K.; Prajapati, H.B. Classification of ECG signals using machine learning techniques: A survey. In Proceedings of the International Conference on Advances in Computer Engineering and Applications (ICACEA), Ghaziabad, India, 19–20 March 2015. [Google Scholar]
- Vijayavanan, M.; Rathikarani, V.; Dhanalakshmi, P. Automatic classification of ECG signal for heart disease diagnosis using morphological features. Int. J. Comput. Sci. Eng. Technol. (IJCSET) 2014, 5, 449–455. [Google Scholar]
- Sansone, M.; Fusco, R.; Pepino, A.; Sansone, C. Electrocardiogram pattern recognition and analysis based on artificial neural networks and support vector machines: A review. J. Healthc. Eng. 2013, 4, 465–504. [Google Scholar] [CrossRef]
- Jollis, J.G.; Granger, C.B.; Henry, T.D.; Antman, E.M.; Berger, P.B.; Moyer, P.H.; Pratt, F.D.; Rokos, I.C.; Acuña, A.R.; Roettig, M.L.; et al. Systems of care for st-segment–levation myocardial infarction: A report from the American heart association’s mission: Lifeline. Circ. Cardiovas. Qual. Outcomes 2012, 5, 423–428. [Google Scholar] [CrossRef]
- Wang, D.; Taubel, J.; Arezina, R. Comparison of six commonly used qt correction models and their parameter estimation methods. J. Biopharm. Stat. 2012, 22, 1148–1161. [Google Scholar] [CrossRef]
- Moody, G.B.; Mark, R.G.; Goldberger, A.L. Physionet: A web-based resource for the study of physiologic signals. IEEE Eng. Med. Biol. 2001, 20, 70–75. [Google Scholar] [CrossRef]
- Moody, G.B.; Mark, R.G. The impact of the MIT-BIH arrhythmia database. IEEE Eng. Med. Biol. 2001, 20, 45–50. [Google Scholar] [CrossRef]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. Physiobank, physiotoolkit, and physionet. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef]
- Dai, W.; Dai, C.; Qu, S.; Li, J.; Das, S. Very deep convolutional neural network for raw waveforms. In Proceedings of the IEEE International Conference on Acoustics Speech and Signal Processing (ICASSP), New Orleans, LA, USA, 5–9 March 2017. [Google Scholar]
- Avanzato, R.; Beritelli, F. An innovative acoustic rain gauge based on convolutional neural networks. Information 2020, 11, 183. [Google Scholar] [CrossRef]
- Beleites, C.; Salzer, R.; Sergo, V. Validation of soft classification models using partial class memberships: An extended concept of sensitivity & co. applied to grading of astrocytoma tissues. Chemometr. Intell. Lab. Syst. 2013, 122, 12–22. [Google Scholar]
- Scikit Learn. Machine Learning in Python. Available online: https://scikit-learn.org/stable/ (accessed on 14 May 2020).
- Sridhar, C.; Acharya, U.R.; Fujita, H.; Bairy, G.M. Automated diagnosis of coronary artery disease using nonlinear features extracted from ECG signals. In Proceedings of the IEEE International Conference on Systems, Man, and Cybernetics, SMC, Budapest, Hungary, 9–12 October 2016. [Google Scholar]
- Ranjan, R.; Arya, R.; Fernandes, S.L.; Sravya, E.; Jain, V. A fuzzy neural network approach for automatic k-complex detection in sleep EEG signal. Pattern Recognit. Lett. 2018, 115, 74–83. [Google Scholar] [CrossRef]
- Bukhari, S.H.R.; Rehmani, M.H.; Siraj, S. A Survey of channel bonding for wireless networks and guidelines of channel bonding for futuristic cognitive radio sensor networks. IEEE Commun. Surv. Tutor. 2016, 18, 924–948. [Google Scholar] [CrossRef]
- Beritelli, F.; Gallotta, A.; Rametta, C. A Dual Streaming approach for speech quality enhancement of VoIP service over 3G networks. In Proceedings of the IEEE International Conference on Digital Signal Processing (DSP), Santorini, Greece, 1–3 July 2013. [Google Scholar]
| Researcher | Preprocessing | Database | Classes | Model | Accuracy |
|---|---|---|---|---|---|
| Acharya et al. [14] | R-Peaks | MIT-BIH arrhythmia | 2 | 1-D CNN, 11-layer | 95.22% |
| Savalia et al. [15] | MIT-BIH arrhythmia and Keggar | 2 (MLP) 9 (CNN) | 1-D CNN, 5-layer | 88.7% | |
| Zubair et al. [16] | MIT-BIH arrhythmia | 5 | 1-D CNN, 4-layer | 92.7% | |
| Li et al. [17] | Wavelet transform | MIT-BIH arrhythmia | 5 | 1-D CNN, 6-layer | 97.5% |
| Baloglu et al. [18] | Wavelet transform | MIT-BIH arrhythmia | 12 lead ECG | 1-D CNN, 10-layer | 99.8% |
| Proposed method | MIT-BIH arrhythmia | 3 | 1-D CNN, 5-layer | 98.33% |
| INPUT | Vectors of 10,800 ECG Samples | ||
|---|---|---|---|
| LAYER 1 | Conv1D (1, 128, 80, 4): input 1 channels output 128 channels kernel_size 80 stride 4 | BatchNorm1D (128): N_features: 128 | MaxPool1D: kernel_size 4 |
| LAYER 2 | Conv1D (128, 128, 3): input 128 channels output 128 channels kernel_size 4 | BatchNorm1D (128): N_features: 128 | MaxPool1D: kernel_size 4 |
| LAYER 3 | Conv1D (128, 256, 3): input 128 channels output 256 channels kernel_size 4 | BatchNorm1D (256): N_features: 256 | MaxPool1D: kernel_size 4 |
| LAYER 4 | Conv1D (256, 512, 3): input 256 channels output 512 channels kernel_size 4 | BatchNorm1D (512): N_features: 512 | MaxPool1D: kernel_size 4 |
| OUTPUT LAYER | AvgPool1D (30): kernel_size 30 | Linear (512, num_classes): input 1 × 512output num_classes: 3 | Log Softmax |
| α | Class | TPR | TNR | FPR | FDR | F1 Score |
|---|---|---|---|---|---|---|
| 1 | Normal | 99.0% | 97.1% | 2.9% | 1% | 98.0% |
| 2 | Atrial premature beat | 100% | 99.0% | 1.0% | 0% | 99.5% |
| 3 | Premature ventricular contraction | 96.0% | 98.96% | 1.04% | 4% | 97.5% |
| Mean Accuracy | 98.33% | 98.33% | 98.35% | 1.65% | 1.66% | 98.33% |
| Method | FE | Model | ACC | TPR | TNR | FPR | FDR |
|---|---|---|---|---|---|---|---|
| Sridhar et al. [33] | DT | DWT | 96.56% | 90.87% | 98.45% | 9.13% | 1.55% |
| Ranjan et al. [34] | RP | FFNN | 87.66% | 94.04% | 76.21% | 5.96% | 23.79% |
| Acharya et al. [14] | RP | 11-layer CNN | 95.22% | 95,49% | 94.19% | ||
| Beritelli et al. [4] | PNN | 96.53% | 93.1% | 100% | |||
| Savalia et al. 1,2 [15] | MLP/5-layer CNN | 88.7%/83.5% | |||||
| Zubair et al. [16] | 4-layer CNN | 92.7% | |||||
| Li et al. [17] | Wavelet transform | 6-layer CNN | 97.5% | ||||
| Baloglu et al. [18] | Wavelet transform | 10-layer CNN | 99.8% | 99.5% | |||
| Proposed method | 5-layer CNN | 98.33% | 98.33% | 98.35% | 1.65% | 1.66% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Avanzato, R.; Beritelli, F. Automatic ECG Diagnosis Using Convolutional Neural Network. Electronics 2020, 9, 951. https://doi.org/10.3390/electronics9060951
Avanzato R, Beritelli F. Automatic ECG Diagnosis Using Convolutional Neural Network. Electronics. 2020; 9(6):951. https://doi.org/10.3390/electronics9060951
Chicago/Turabian StyleAvanzato, Roberta, and Francesco Beritelli. 2020. "Automatic ECG Diagnosis Using Convolutional Neural Network" Electronics 9, no. 6: 951. https://doi.org/10.3390/electronics9060951
APA StyleAvanzato, R., & Beritelli, F. (2020). Automatic ECG Diagnosis Using Convolutional Neural Network. Electronics, 9(6), 951. https://doi.org/10.3390/electronics9060951






