Multi-Task Deep Learning for Lung Nodule Detection and Segmentation in CT Scans
Abstract
1. Introduction
1.1. Objectives
- Unify detection and segmentation in a single architecture that shares features while preserving task-specific heads.
- Adopt a 2.5D formulation that captures limited through-plane context while avoiding volumetric convolutions required by full-3D models.
- Improve sensitivity to small/low-contrast nodules and evaluate performance at the clinically relevant nodule level.
1.2. Contributions
- A data pipeline tailored for LUNA16 (HU normalisation, CLAHE enhancement, lung masking, and slice-level packaging) for stable 2.5D inputs;
- A 2.5D Mask R-CNN-based MTL model with anchors tuned for small nodules and an auxiliary RoI classifier to retain borderline true positives;
- A nodule-level evaluation protocol that merges slice-wise predictions across z, reporting Precision/Recall/-score alongside Dice/IoU;
- Evidence that the proposed design attains a favourable precision–recall/segmentation trade-off, while avoiding volumetric convolutions, reflecting a structural trade-off between through-plane context and architectural complexity.
2. Related Work
2.1. Lung Nodule Detection
2.2. Lung Nodule Segmentation
2.3. Multi-Task Learning Approaches
2.4. Datasets and Evaluation Metrics
2.5. Summary
3. Methodology
3.1. Data Collection
- Lung Masks: Binary segmentation masks identifying the lung parenchyma. These are provided as separate volumetric files aligned with each CT scan and are used to suppress non-lung regions during preprocessing.
- Nodule Annotations: Structured CSV files provide nodule locations in world coordinates (in millimetres), including centre coordinates and nodule diameter. These annotations are converted to voxel coordinates using the scan’s spatial metadata (origin and spacing). For segmentation supervision, approximate 2D masks are generated by projecting the annotated nodule diameter onto axial slices, resulting in simplified geometric masks rather than voxel-level delineations of true nodule morphology.
3.2. Preprocessing Pipeline
3.3. Multi-Task Learning Architecture
3.3.1. Shared Backbone and FPN
3.3.2. Detection and Segmentation Heads
3.3.3. Design Motivation and Model Evolution
- Anchor Generator Optimisation. The default RPN anchor scales were manually redefined as , , , , and pixels, each with aspect ratios . This reconfiguration improves anchor coverage for small nodules often under 10 mm in diameter, which are frequently missed under the default setup in LUNA16.
- Increased Proposal Count. The maximum number of RPN proposals was raised to 2000 during training and 300 during inference. This allows more candidate regions to be passed to the RoI heads, increasing the likelihood of capturing true nodules.
- Auxiliary RoI Classifier. A fully connected classifier was added after the RoI feature extraction stage to predict whether a proposal corresponds to a true nodule. During training, proposals with IoU to any ground truth were labelled positive, and a binary cross-entropy loss was added to the total loss to supervise this auxiliary classifier. The complete loss formulation is presented in Section 3.3.4. In inference, the auxiliary logits were fused with the primary classification scores to re-rank predictions. The auxiliary RoI classifier was designed as a complementary decision head that focuses on local RoI-level discrimination, while the primary classification head prioritises recall at the proposal level.
3.3.4. Loss Functions
3.4. Evaluation Metrics
3.4.1. Image Quality Assessment (IQA)
3.4.2. Detection and Segmentation Metrics
3.4.3. Nodule-Level Evaluation Strategy
3.5. Training Strategy
- Phase 1: Detection-Only Training. The mask head was frozen, and only the detection components were trained. This phase established strong detection weights without being influenced by segmentation gradients.
- Phase 2: Segmentation-Only Training. The detection branch was frozen, and the mask head was fine-tuned to learn instance masks based on stable RoI proposals.
- Phase 3: Joint Fine-Tuning. Both branches were unfrozen and jointly optimised, starting from previously trained weights. The number of epochs in each phase was iteratively adjusted based on validation trends to achieve optimal performance.
4. Results and Discussions
4.1. Image Enhancement and Quality Assessment
4.1.1. PSNR-Based Evaluation
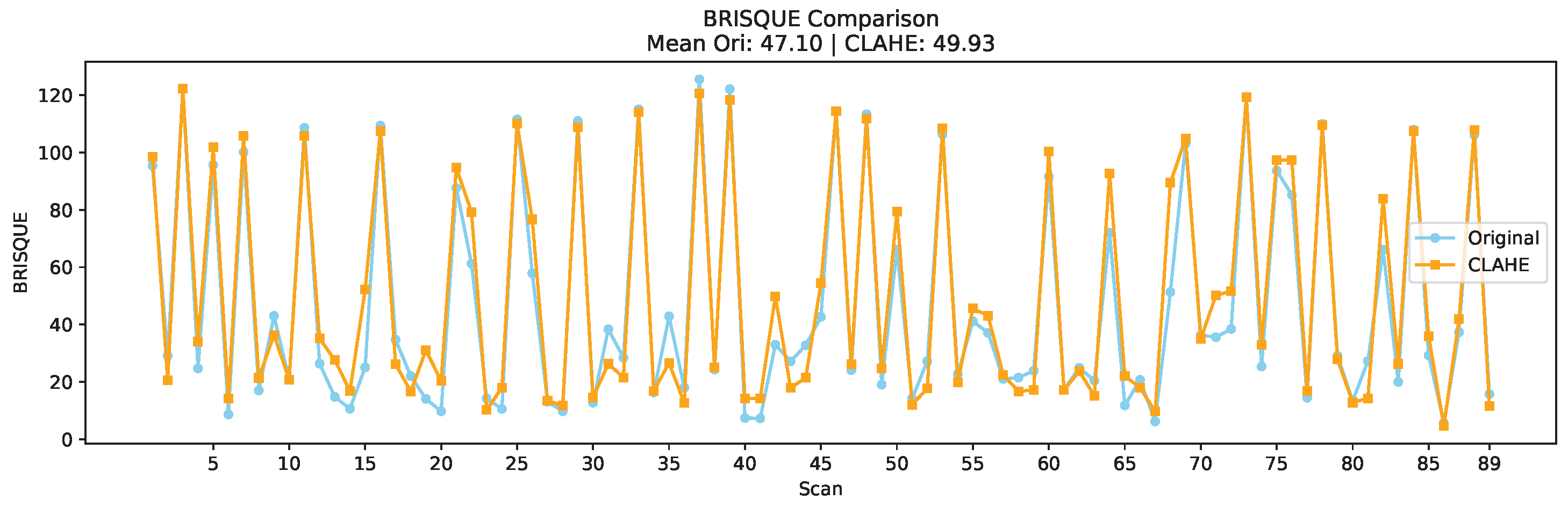
4.1.2. BRISQUE-Based Evaluation
4.1.3. Entropy-Based Evaluation
4.1.4. Paired t-Test Results and Conclusion
- BRISQUE: , → significant
- Entropy: , → significant
4.2. Preprocessing Visualisation
4.3. Multi-Task Model Performance
4.3.1. Detection and Segmentation Performance
4.3.2. Validation of Enhancement Effectiveness
4.3.3. Ablation Study
4.4. Limitations
4.5. Comparison with Published Baselines
4.6. Discussion and Implications
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cancer Research UK. Lung Cancer Statistics. 2025. Available online: https://www.cancerresearchuk.org/health-professional/cancer-statistics/statistics-by-cancer-type/lung-cancer#lung_stats1 (accessed on 5 June 2025).
- The National Lung Screening Trial Research Team. Reduced lung-cancer mortality with low-dose computed tomographic screening. N. Engl. J. Med. 2011, 365, 395–409. [Google Scholar] [CrossRef]
- MacMahon, H.; Naidich, D.P.; Goo, J.M.; Lee, K.S.; Leung, A.N.; Mayo, J.R.; Mehta, A.C.; Ohno, Y.; Powell, C.A.; Prokop, M.; et al. Guidelines for management of incidental pulmonary nodules detected on CT images: From the Fleischner Society 2017. Radiology 2017, 284, 228–243. [Google Scholar] [CrossRef] [PubMed]
- Riquelme, D.; Akhloufi, M.A. Deep learning for lung cancer nodules detection and classification in CT scans. AI 2020, 1, 28–67. [Google Scholar] [CrossRef]
- Marinakis, I.; Karampidis, K.; Papadourakis, G. Pulmonary Nodule Detection, Segmentation and Classification Using Deep Learning: A Comprehensive Literature Review. BioMedInformatics 2024, 4, 2043–2106. [Google Scholar] [CrossRef]
- Wang, Y.; Mustaza, S.M.; Ab-Rahman, M.S. Pulmonary Nodule Segmentation using Deep Learning: A Review. IEEE Access 2024, 12, 119039–119055. [Google Scholar] [CrossRef]
- Setio, A.A.A.; Traverso, A.; De Bel, T.; Berens, M.S.; Van Den Bogaard, C.; Cerello, P.; Chen, H.; Dou, Q.; Fantacci, M.E.; Geurts, B.; et al. Validation, comparison, and combination of algorithms for automatic detection of pulmonary nodules in computed tomography images: The LUNA16 challenge. Med. Image Anal. 2017, 42, 1–13. [Google Scholar] [CrossRef]
- El-Baz, A.; Beache, G.M.; Gimel farb, G.; Suzuki, K.; Okada, K.; Elnakib, A.; Soliman, A.; Abdollahi, B. Computer-aided diagnosis systems for lung cancer: Challenges and methodologies. Int. J. Biomed. Imaging 2013, 2013, 942353. [Google Scholar] [CrossRef]
- Zhang, G.; Jiang, S.; Yang, Z.; Gong, L.; Ma, X.; Zhou, Z.; Bao, C.; Liu, Q. Automatic nodule detection for lung cancer in CT images: A review. Comput. Biol. Med. 2018, 103, 287–300. [Google Scholar] [CrossRef]
- Zheng, L.; Lei, Y. A review of image segmentation methods for lung nodule detection based on computed tomography images. Matec Web Conf. 2018, 232, 02001. [Google Scholar] [CrossRef]
- Pehrson, L.M.; Nielsen, M.B.; Ammitzbøl Lauridsen, C. Automatic pulmonary nodule detection applying deep learning or machine learning algorithms to the LIDC-IDRI database: A systematic review. Diagnostics 2019, 9, 29. [Google Scholar] [CrossRef]
- Alshayeji, M.H.; Abed, S. Lung cancer classification and identification framework with automatic nodule segmentation screening using machine learning. Appl. Intell. 2023, 53, 19724–19741. [Google Scholar] [CrossRef]
- Luo, D.; He, Q.; Ma, M.; Yan, K.; Liu, D.; Wang, P. ECANodule: Accurate Pulmonary Nodule Detection and Segmentation with Efficient Channel Attention. In Proceedings of the 2023 International Joint Conference on Neural Networks (IJCNN), Gold Coast, Australia, 18–23 June 2023; IEEE: Piscataway, NJ, USA, 2023; pp. 1–8. [Google Scholar]
- Zhang, Y.; Yang, Q. A survey on multi-task learning. IEEE Trans. Knowl. Data Eng. 2021, 34, 5586–5609. [Google Scholar] [CrossRef]
- Thung, K.H.; Wee, C.Y. A brief review on multi-task learning. Multimed. Tools Appl. 2018, 77, 29705–29725. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, X.; Che, T.; Bao, G.; Li, S. Multi-task deep learning for medical image computing and analysis: A review. Comput. Biol. Med. 2023, 153, 106496. [Google Scholar] [CrossRef]
- Li, R.; Xiao, C.; Huang, Y.; Hassan, H.; Huang, B. Deep learning applications in computed tomography images for pulmonary nodule detection and diagnosis: A review. Diagnostics 2022, 12, 298. [Google Scholar] [CrossRef]
- Li, R.; Honarvar Shakibaei Asli, B. Multi-Task Deep Learning for Lung Nodule Detection and Segmentation in CT Scans: A Review. Electronics 2025, 14, 3009. [Google Scholar] [CrossRef]
- George, J.; Skaria, S. Using YOLO based deep learning network for real time detection and localization of lung nodules from low dose CT scans. In Medical Imaging 2018: Computer-Aided Diagnosis; SPIE: Bellingham, WA, USA, 2018; Volume 10575, pp. 347–355. [Google Scholar]
- Xie, H.; Yang, D.; Sun, N.; Chen, Z.; Zhang, Y. Automated pulmonary nodule detection in CT images using deep convolutional neural networks. Pattern Recognit. 2019, 85, 109–119. [Google Scholar] [CrossRef]
- Dou, Q.; Chen, H.; Yu, L.; Qin, J.; Heng, P.A. Multilevel contextual 3-D CNNs for false positive reduction in pulmonary nodule detection. IEEE Trans. Biomed. Eng. 2016, 64, 1558–1567. [Google Scholar] [CrossRef]
- Anirudh, R.; Thiagarajan, J.J.; Bremer, T.; Kim, H. Lung nodule detection using 3D convolutional neural networks trained on weakly labeled data. In Medical Imaging 2016: Computer-Aided Diagnosis; SPIE: Bellingham, WA, USA, 2016; Volume 9785, pp. 791–796. [Google Scholar]
- Paul, R.; Hawkins, S.H.; Schabath, M.B.; Gillies, R.J.; Hall, L.O.; Goldgof, D.B. Predicting malignant nodules by fusing deep features with classical radiomics features. J. Med. Imaging 2018, 5, 011021. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Zhang, H.; Sun, J.; Wang, S.; Zhang, Y. YOLO-MSRF for lung nodule detection. Biomed. Signal Process. Control 2024, 94, 106318. [Google Scholar] [CrossRef]
- Tang, H.; Li, Z.; Zhang, D.; He, S.; Tang, J. Divide-and-conquer: Confluent triple-flow network for RGB-T salient object detection. IEEE Trans. Pattern Anal. Mach. Intell. 2024, 47, 1958–1974. [Google Scholar] [CrossRef]
- Wang, C.Y.; Yeh, I.H.; Mark Liao, H.Y. YOLOv9: Learning what you want to learn using programmable gradient information. In Computer Vision—ECCV 2024, Proceedings of the 18th European Conference on Computer Vision, Milan, Italy, 29 September–4 October 2024; Springer: Cham, Switzerland, 2024; pp. 1–21. [Google Scholar]
- Tong, G.; Li, Y.; Chen, H.; Zhang, Q.; Jiang, H. Improved U-NET network for pulmonary nodules segmentation. Optik 2018, 174, 460–469. [Google Scholar] [CrossRef]
- Ma, X.; Song, H.; Jia, X.; Wang, Z. An improved V-Net lung nodule segmentation model based on pixel threshold separation and attention mechanism. Sci. Rep. 2024, 14, 4743. [Google Scholar] [CrossRef]
- Cao, H.; Liu, H.; Song, E.; Hung, C.C.; Ma, G.; Xu, X.; Jin, R.; Lu, J. Dual-branch residual network for lung nodule segmentation. Appl. Soft Comput. 2020, 86, 105934. [Google Scholar] [CrossRef]
- Wang, S.; Zhou, M.; Gevaert, O.; Tang, Z.; Dong, D.; Liu, Z.; Jie, T. A multi-view deep convolutional neural networks for lung nodule segmentation. In Proceedings of the 2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Jeju, Republic of Korea, 11–15 July 2017; IEEE: Piscataway, NJ, USA, 2017; pp. 1752–1755. [Google Scholar]
- Liu, C.; Liu, H.; Zhang, X.; Guo, J.; Lv, P. Multi-scale and multi-view network for lung tumor segmentation. Comput. Biol. Med. 2024, 172, 108250. [Google Scholar] [CrossRef]
- Tyagi, S.; Talbar, S.N. CSE-GAN: A 3D conditional generative adversarial network with concurrent squeeze-and-excitation blocks for lung nodule segmentation. Comput. Biol. Med. 2022, 147, 105781. [Google Scholar] [CrossRef]
- Usman, M.; Shin, Y.G. Deha-net: A dual-encoder-based hard attention network with an adaptive roi mechanism for lung nodule segmentation. Sensors 2023, 23, 1989. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Xu, L.; Ma, Z.; Zhao, Y.; Li, X. APU-Net: A U-Net Enhanced Network with Dynamic Feature Fusion and Pyramid Cross-Attention Mechanism for Polyp Segmentation. Digit. Signal Process. 2026, 172, 105879. [Google Scholar] [CrossRef]
- Hu, T.; Lan, Y.; Zhang, Y.; Xu, J.; Li, S.; Hung, C.C. A lung nodule segmentation model based on the transformer with multiple thresholds and coordinate attention. Sci. Rep. 2024, 14, 31743. [Google Scholar] [CrossRef] [PubMed]
- Yadav, D.P.; Sharma, B.; Webber, J.L.; Mehbodniya, A.; Chauhan, S. EDTNet: A spatial aware attention-based transformer for the pulmonary nodule segmentation. PLoS ONE 2024, 19, e0311080. [Google Scholar] [CrossRef]
- Tang, H.; Zhang, C.; Xie, X. Nodulenet: Decoupled false positive reduction for pulmonary nodule detection and segmentation. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2019: 22nd International Conference, Shenzhen, China, 13–17 October 2019, Proceedings, Part VI; Springer: Cham, Switzerland, 2019; pp. 266–274. [Google Scholar]
- Song, G.; Nie, Y.; Zhang, J.; Chen, G. Multi-task weakly-supervised learning model for pulmonary nodules segmentation and detection. In Proceedings of the 2020 International Conference on Innovation Design and Digital Technology (ICIDDT), Zhenjing, China, 5–6 December 2020; IEEE: Piscataway, NJ, USA, 2020; pp. 343–347. [Google Scholar]
- Nguyen, T.C.; Nguyen, T.P.; Cao, T.; Dao, T.T.P.; Ho, T.N.; Nguyen, T.V.; Tran, M.T. MANet: Multi-branch attention auxiliary learning for lung nodule detection and segmentation. Comput. Methods Programs Biomed. 2023, 241, 107748. [Google Scholar] [CrossRef]
- Armato, S.G., III; McLennan, G.; Bidaut, L.; McNitt-Gray, M.F.; Meyer, C.R.; Reeves, A.P.; Zhao, B.; Aberle, D.R.; Henschke, C.I.; Hoffman, E.A.; et al. The lung image database consortium (LIDC) and image database resource initiative (IDRI): A completed reference database of lung nodules on CT scans. Med. Phys. 2011, 38, 915–931. [Google Scholar] [CrossRef]
- Trial Summary—Learn-NLST—The Cancer Data Access System. Available online: https://cdas.cancer.gov/learn/nlst/trial-summary/ (accessed on 14 July 2025).
- Van Ginneken, B.; Armato, S.G., III; de Hoop, B.; van Amelsvoort-van de Vorst, S.; Duindam, T.; Niemeijer, M.; Murphy, K.; Schilham, A.; Retico, A.; Fantacci, M.E.; et al. Comparing and combining algorithms for computer-aided detection of pulmonary nodules in computed tomography scans: The ANODE09 study. Med. Image Anal. 2010, 14, 707–722. [Google Scholar] [CrossRef] [PubMed]
- ELCAP Public Lung Image Database. Available online: http://www.via.cornell.edu/lungdb.html (accessed on 14 July 2025).
- A.T.C. Tianchi Medical AI Competition Dataset. 2017. Available online: https://tianchi.aliyun.com/competition/entrance/231601/information (accessed on 14 July 2025).
- Hao, S.; Wang, P.; Hu, Y. Haze image recognition based on brightness optimization feedback and color correction. Information 2019, 10, 81. [Google Scholar] [CrossRef]
- Ni, Y.; Zi, D.; Chen, W.; Wang, S.; Xue, X. Egc-yolo: Strip steel surface defect detection method based on edge detail enhancement and multiscale feature fusion. J. Real-Time Image Process. 2025, 22, 65. [Google Scholar] [CrossRef]
- Lin, T.Y.; Dollár, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature pyramid networks for object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2117–2125. [Google Scholar]














| Type | Model/Method | Dataset | ACC (%) | Sens. (%) | AUC | Notes |
|---|---|---|---|---|---|---|
| 2D CNN-based | DetectNet [19] | LIDC-IDRI | 93.00 | 89.00 | – | -score = 90.96, single-stage |
| Faster R-CNN Variant [20] | LUNA16 | – | 86.42 | 95.4 | Deconv + dual RPN | |
| YOLO-MSRF [24] | LUNA16 | 95.41 | 94.02 | – | Single-stage, MSRF | |
| 3D CNN-based | Multilevel CNN [21] | LUNA16 | – | 94.40 | – | Multi-context fusion |
| Point-supervised [22] | LUNA16 | – | 80.00 | – | Weak supervision | |
| Hybrid | 3D CNN + VGG + Radiomics [23] | NLST | 76.79 | – | 78 | Feature-level fusion |
| Type | Model/Method | Dataset | DSC (%) | Sens. (%) | Notes |
|---|---|---|---|---|---|
| 2D CNN-based | Improved U-Net [27] | LUNA16 | 73.6 | – | Residual connections, batch normalisation, skip connections |
| 3D CNN-based | Improved Dig-CS-VNet [28] | LUNA16, LNDb | 94.9, 81.1 | 92.7, 76.9 | Feature separation, 3D attention blocks |
| ResNet-based | DB-ResNet [29] | LIDC-IDRI | 89.40 | – | Dual-branch structure, intensity pooling, modular ResBlocks |
| Multi-view CNN | MV-CNN [30] | LIDC-IDRI | 82.74 | – | Axial/coronal/sagittal branches, late fusion |
| Multi-scale + view | MSMV-Net [31] | LUNA16, MSD | 55.60, 59.94 | – | 2D view fusion, attention-weighted deep supervision |
| GAN-based | CSE-GAN [32] | LUNA16, ILND | 80.74, 76.36 | 85.46, 82.56 | 3D U-Net + CSE attention, sScE discriminator |
| Transformer-based | DEHA-Net [33] | LIDC-IDRI | 87.91 | 90.84 | Dual-encoder, transformer modules, hard attention |
| Type | Model/Method | Dataset | FROC (%) | DSC (%) | Key Features |
|---|---|---|---|---|---|
| Cascaded | NoduleNet [37] | LIDC-IDRI | +10.27% vs. single-task | 83.1 | Decoupled tasks, FP reduction, segmentation refinement module |
| Parallel (Weakly sup.) | Multi-branch U-net + ConvLSTM [38] | LIDC-IDRI | +6.89% vs. single-task | 82.26 | Sequential context, dynamic loss weighting, weak labels |
| Hybrid | ECANodule [13] | LIDC-IDRI | 91.1 | 83.4 | Dense skip connections, attention, OHEM training |
| Hybrid (Deep-attention) | MANet [39] | LIDC-IDRI | 88.11 | 82.74 | Deep supervision, multi-branch attention, boundary enhancement |
| Dataset | Modality | Annotation Type | Size | Notes |
|---|---|---|---|---|
| LIDC-IDRI [40] | CT, DX, CR | Bounding boxes, malignancy ratings (4 readers) | 1018 scans | Multi-reader annotated; basis for many derived datasets |
| LUNA16 [7] | CT | Filtered nodules ≥3 mm from LIDC-IDRI | 888 scans | High-quality subset of LIDC-IDRI; used for 10-fold CV |
| NLST [41] | CT, X-ray | Patient outcome, lesion info | 54,000+ participants | Large-scale clinical trial; mortality-focused |
| ANODE09 [42] | CT | True nodules and irrelevant findings | 55 scans | Partially annotated; used for CAD evaluation |
| ELCAP [43] | CT | Nodule locations | 50 scans | Early low-dose CT dataset; used for CAD benchmarking |
| Tianchi [44] | CT | 3D nodule boxes, clinical labels | 1000+ scans | 5–30 mm nodules annotated by 3 doctors |
| Metric | Description | Formula |
|---|---|---|
| Accuracy (ACC) | Correct predictions among all cases. | |
| Sensitivity (Recall) | True positive rate. | |
| Specificity | True negative rate. | |
| Precision (PPV) | Positive predictions that are correct. | |
| -Score | Harmonic mean of precision and recall. | |
| Dice Similarity Coefficient (DSC) | Similarity between prediction and GT. | |
| Intersection over Union (IoU) | Overlap ratio of prediction and GT. | |
| AUC | Area under ROC curve. | – |
| ROC Curve | True positive rate vs. false positive rate. | – |
| FROC Curve | Sensitivity vs. false positives per scan. | – |
| CPM | Average sensitivity at 7 FPs/scan. | – |
| Mean Average Precision (mAP) | Mean of AP over all classes. |
| Type | Metric | Description | Formula |
|---|---|---|---|
| FR | MSE | Calculates the mean squared error between images. A smaller MSE indicates higher image similarity. A lower value indicates better quality. | , where and are corresponding pixel values in the two images, and N is the total number of pixels. |
| SSIM | Assesses perceptual similarity based on luminance, contrast, and structure. | , where , are the mean intensities of images x and y, respectively; , are their standard deviations; is the cross-covariance. and are constants to stabilise the division. | |
| PSNR | Measures signal fidelity between two images. A higher value indicates better image quality. | , where is the maximum possible pixel value of the image. | |
| NR | PIQE | Evaluates perceptual quality based on block-level distortion. It analyses local blocks for artefacts and noise. Lower value indicates better quality. | , where is blockiness measure and is noise measure. |
| NIQE | Estimates deviation from natural image statistics. A lower value indicates better quality. | , where , and , are the mean vectors and covariance matrices of the test and natural. | |
| BRISQUE | Predicts image quality using natural scene statistics. A lower score means better quality. | , where and are local mean and standard deviation from MSCN coefficients. Features are modelled using AGGD, and quality is predicted via SVR. | |
| Entropy | Measures intensity distribution complexity. Higher values indicate greater contrast and detail. | , where is the normalized histogram probability of pixel value . |
| Scans (Nodule/Non-Nodule) | Samples Total | Pos. | Neg. | |
|---|---|---|---|---|
| Training | 70 (53/17) | 317 | 266 | 51 |
| Validation | 19 (14/5) | 75 | 60 | 15 |
| Epoch | TP | FP | FN | Precision | Recall | -Score | Dice | IoU |
|---|---|---|---|---|---|---|---|---|
| 6 | 46 | 11 | 17 | 0.81 | 0.73 | 0.77 | 0.46 | 0.30 |
| 15 | 34 | 45 | 29 | 0.43 | 0.54 | 0.48 | 0.76 | 0.64 |
| 19 | 39 | 10 | 24 | 0.80 | 0.62 | 0.70 | 0.81 | 0.70 |
| 35 | 17 | 0 | 46 | 1.00 | 0.27 | 0.43 | 0.82 | 0.72 |
| Input Type | Epoch | Precision | Recall | -Score | Dice | IoU |
|---|---|---|---|---|---|---|
| Original CT | 16 | 0.85 | 0.54 | 0.66 | 0.85 | 0.76 |
| CLAHE-enhanced CT | 19 | 0.80 | 0.62 | 0.70 | 0.81 | 0.70 |
| Model | TP | FP | FN | Precision | Recall | -Score | Dice | IoU |
|---|---|---|---|---|---|---|---|---|
| Baseline | ||||||||
| (3-slice, default) | 18 | 5 | 45 | 0.78 | 0.29 | 0.42 | 0.84 | 0.73 |
| Final Model | ||||||||
| (5-slice + anchor + aux) | 39 | 10 | 24 | 0.80 | 0.62 | 0.70 | 0.81 | 0.70 |
| Model | Task | Dataset | Metric(s) | Reported Value |
|---|---|---|---|---|
| Faster R-CNN Variant [20] | Detection (2D) | LUNA16 | Sens/AUC | 86.4/95.4% |
| YOLO-MSRF [24] | Detection (2D) | LUNA16 | Sens | 94.02% |
| Multilevel 3D CNN [21] | Detection (3D) | LUNA16 | Sens | 94.4% |
| Improved U-Net [27] | Segmentation (2D) | LUNA16 | Dice | 73.6% |
| DEHA-Net [33] | Segmentation (2D) | LIDC-IDRI | Dice | 87.91% |
| CSE-GAN [32] | Segmentation (3D) | LUNA16 | Dice | 80.7% |
| NoduleNet [37] | Cascaded (Detect→Seg) | LIDC-IDRI | FROC/Dice | +10.3% vs. single-task/83.1% |
| ECANodule [13] | Hybrid (Detect + Seg) | LIDC-IDRI | FROC/Dice | 91.1/83.4% |
| Ours (2.5D MTL) | MTL (Detect + Seg) | LUNA16 subset0 | -score/Dice | 0.70/0.81 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Li, R.; Honarvar Shakibaei Asli, B. Multi-Task Deep Learning for Lung Nodule Detection and Segmentation in CT Scans. Electronics 2026, 15, 736. https://doi.org/10.3390/electronics15040736
Li R, Honarvar Shakibaei Asli B. Multi-Task Deep Learning for Lung Nodule Detection and Segmentation in CT Scans. Electronics. 2026; 15(4):736. https://doi.org/10.3390/electronics15040736
Chicago/Turabian StyleLi, Runhan, and Barmak Honarvar Shakibaei Asli. 2026. "Multi-Task Deep Learning for Lung Nodule Detection and Segmentation in CT Scans" Electronics 15, no. 4: 736. https://doi.org/10.3390/electronics15040736
APA StyleLi, R., & Honarvar Shakibaei Asli, B. (2026). Multi-Task Deep Learning for Lung Nodule Detection and Segmentation in CT Scans. Electronics, 15(4), 736. https://doi.org/10.3390/electronics15040736






