1. Introduction
Alzheimer’s disease (AD), a progressively prevalent brain disorder, has become the fourth leading cause of death in industrialized nations. Its primary symptoms, memory loss and cognitive impairment, are attributed to the death and damage of neurons associated with memory in the brain. The intermediate condition between normal brain function and AD is known as mild cognitive impairment (MCI), which is a state that often precedes the onset of AD. Studies indicate that a high likelihood of progressing to AD is observed in individuals with MCI [
1,
2]. As the disease advances and transitions through stages, increasing caution is exercised by clinicians when assessing patients. Distinguishing the specific symptomatic differences among these groups presents a significant challenge for researchers.
Indispensable standardized image data for this assessment process is provided by medical imaging modalities such as positron emission tomography (PET) [
3,
4,
5], magnetic resonance imaging (MRI) [
6], and computed tomography (CT) [
7]. Among these techniques, MRI is widely used in diagnosing a variety of conditions including brain tumors, neurological disorders, spinal cord injuries and abnormalities, and liver diseases due to its high safety and sensitivity. Its diverse imaging sequences offer distinct advantages for different pathologies, and MRI is especially prevalent in AD classification [
8,
9]. Features extracted from MRI, such as gray and white matter volumes, cortical thickness, and cerebrospinal fluid (CSF) levels, are known to aid in MCI and AD classification and diagnosis, as well as enabling disease staging. In recent years, great promise has been shown by convolutional neural networks (CNNs) in automatically analyzing brain MRI scans for the diagnosis of cognitive disorders [
10,
11,
12].
2. Related Works
In recent years, deep learning has been increasingly applied to Alzheimer’s disease (AD) classification, particularly through the use of multimodal neuroimaging and clinical data [
13,
14]. Early work mainly focused on conventional convolutional neural networks (CNNs) to extract informative patterns from brain MRI scans. Several recent reviews, including those by Agarwal et al. [
15] and Grueso et al. [
16], have outlined the development of these approaches and highlighted a clear shift from simple binary classification toward more clinically meaningful multi-class settings (AD/CN/MCI).
Building on this progression, later studies have reported notable improvements in classification performance. For instance, Oktavian et al. [
17] applied a ResNet-18 model and achieved an accuracy of 0.88 for three-class classification on the ADNI dataset, while Nasir et al. [
18] showed that ensemble CNN strategies could reach 0.90 accuracy. To ensure a fair and up-to-date comparison, we re-implemented the DenseNet-169 model proposed in [
19] using our own ADNI data split in 2025, obtaining an accuracy of 0.86. Under the same experimental conditions, our proposed HMFF surpasses these baselines, achieving an accuracy of 0.94 while also offering improved model interpretability.
Beyond architectural depth and ensemble size, recent investigations have emphasized the importance of efficient feature representation. DenseNet-based models have been reported to exhibit strong feature reuse and gradient propagation properties, with Shehri et al. [
19] identifying DenseNet-169 as a competitive feature extractor for AD classification. Nevertheless, direct comparison across studies remains challenging due to differences in datasets, preprocessing protocols, and evaluation splits—particularly when results are reported on heterogeneous sources such as Kaggle rather than clinically curated datasets. To ensure a fair and reproducible benchmark, the DenseNet-169 architecture was re-implemented in this study using an identical ADNI data split, allowing a controlled comparison under consistent experimental conditions.
Building upon prior methodologies, the present study emphasizes the optimization of methodological robustness, interpretable multimodal fusion, and clinical applicability. By integrating 3D DenseNet-121-based volumetric imaging features with structured clinical priors through a hierarchical fusion strategy, the proposed HMFF seeks to refine existing fusion strategies by addressing challenges such as overfitting risk, feature redundancy, and limited interpretability. This positioning enables the proposed approach to be evaluated as a practical technical validation, focusing not only on classification accuracy but also on statistical stability and generalizability, thereby providing a complementary contribution to multimodal AD staging research.
3. Methodology
This section details the research design, including the composition of the dataset, the standardized preprocessing workflow, and the proposed hierarchical fusion architecture.
3.1. Datasets
In the field of Alzheimer’s disease research, although various public datasets are accessible through platforms such as Kaggle and OASIS, this study chose the ADNI (Alzheimer’s Neuroimaging Project) dataset because of its higher data completeness and stronger pattern consistency. Specifically, this study included data from participants in the ADNI-1, ADNI-2, ADNI-3, and ADNI-4 cohorts, which included both T1-weighted MRI scans and core clinical scales, namely the Mini-Mental State Examination (MMSE) and the Clinical Dementia Assessment Scale (CDR). These two scales were chosen to ensure the completeness of data from different stages of ADNI and to minimize potential sampling bias due to missing values. During data processing, we implemented a broad inclusion strategy, excluding only instances with corrupted files, missing metadata, or incomplete diagnostic labels. This approach aimed to initially observe the model’s performance under different scanning conditions. This has been described as a limitation of this study in the discussion section. Detailed information on participants and scan distribution is shown in
Table 1.
The dataset was partitioned into training, validation, and test sets using a 60/20/20 ratio. To prevent data leakage, we strictly implemented subject-level splitting, utilizing Subject IDs as the primary basis for partitioning. Given that individual subjects in the ADNI dataset often have multiple scans across different time points (multiple sessions), we ensured that all historical images and corresponding clinical scales for a single subject were assigned exclusively to the same set (either training, validation, or test). This mechanism avoids information overlap across sets, which could otherwise lead to an overestimation of model performance. Furthermore, to address class imbalance, stratified sampling was employed during the allocation process to maintain consistent proportions of AD, MCI, and CN subjects across all subsets. This strategy aims to reduce bias in the evaluation process and enhance the reference value of the experimental results.
3.2. Data Preprocessing
A comprehensive and rigorous image preprocessing pipeline was implemented before model training to ensure consistent MRI data quality and enhance deep learning performance [
20]. The architecture is illustrated in
Figure 1. First, N4 bias field correction was applied to eliminate intensity inhomogeneities caused by magnetic field nonuniformity, thereby improving image contrast and detail clarity. This technique employs a multi-resolution strategy to balance correction effectiveness with computational efficiency, which strengthens the accuracy and stability of subsequent feature extraction.
To standardize the spatial alignment of all subjects’ scans, every MRI volume was registered to the MNI152 template coordinate space. The SyN nonlinear algorithm from the ANTs toolkit (ANTsPy version 0.5.4) was utilized during this registration to preserve anatomical integrity while compensating for variations in scan positioning and individual anatomy, thereby effectively reducing bias due to structural differences. Following this, skull stripping was performed to focus on intracranial tissue and remove non-brain structures. Precise brain tissue probability masks were generated by the ANTsNet (ANTsPyNet version 0.2.9) deep learning model and subsequently thresholder to exclude extracerebral regions, ensuring that the neural network concentrates on core brain features. Voxel intensities were then normalized via min-max scaling to the range [0, 1], a process that significantly mitigates intensity differences across scanners and subjects, accelerates model convergence, and stabilizes feature extraction. Finally, to address inconsistencies in image dimensions and resolution, all volumes were resampled to a uniform spatial resolution of 128 × 128 × 128 voxels using nearest-neighbor interpolation, which preserves structural details and original characteristics. This resampling step eliminated sampling-rate-induced variability in model performance and ensured data standardization.
The complete preprocessing architecture, which guarantees high-quality and consistent MRI inputs, facilitates effective feature learning by the deep learning model and improves both accuracy and generalizability in Alzheimer’s disease classification.
3.3. Proposed System
A hierarchical multi-modal fusion framework (HMFF) is proposed, which combines a 3D DenseNet-121 deep convolutional neural network with an XGBoost classifier. This is done to fully leverage both MRI and clinical data features to improve diagnostic accuracy and generalizability. The 3D DenseNet-121 architecture, with its dense connectivity design, enhances feature propagation and reuse, reduces parameter redundancy, and improves gradient flow, thereby enabling the deep network to capture more subtle and clinically relevant anatomical patterns. This network is optimized for 3D MRI volumes, taking single-channel inputs of size 128 × 128 × 128 voxels.
The proposed HMFF was implemented using Python 3.10 and PyTorch 2.4 on the Google (Google LLC, Mountain View, CA, USA) Colab platform, utilizing an NVIDIA T4 GPU (NVIDIA Corporation, Santa Clara, CA, USA). To ensure rigorous experimental reproducibility, we fixed the global random seed at 42 for all stages, including dataset partitioning, weight initialization, and the training process. Key libraries included XGBoost 2.1, SHAP 0.44, and Scikit-learn 1.5.
The full model framework is illustrated in
Figure 2. The 3D DenseNet-121 model is composed of multiple Dense Blocks, each containing several layers of 3D convolutions, batch normalization, and ReLU activations. Through dense connections, direct access to the feature maps of all preceding layers is provided. transition layers, which comprise a convolution and average pooling, are inserted between dense blocks to control the number of feature channels and reduce spatial dimensions. At the network’s end, a global average pooling layer is used to compress the 3D feature maps into a 1024-dimensional global feature vector, which is then fed into the subsequent fusion module.
Structured clinical scale data, including the Mini-Mental State Examination (MMSE) and Clinical Dementia Rating (CDR) scores, are processed by an XGBoost classifier. By employing the gradient-boosted tree algorithm, an ensemble of multiple decision trees is used to produce a probability distribution over the three diagnostic categories (AD, MCI, CN) [
21]. The tree-based structure of XGBoost effectively models nonlinear feature interactions and utilizes regularization to control model complexity.
During the feature fusion stage, the 1024-dimensional imaging feature vector extracted by the 3D DenseNet-121 is combined with the three-dimensional clinical probability vector generated by XGBoost. To enhance feature efficiency and interpretability, the SHAP (Shapley Additive Explanations) algorithm is applied exclusively on the validation set to perform feature attribution and to identify the 90 most discriminative imaging features from the high-dimensional representation. Importantly, this validation set is used solely for feature importance estimation and model selection, without any involvement in parameter optimization or weight updating.
The resulting 90 selected imaging features are concatenated with the clinical probability vector to form a 93-dimensional fused representation. This fused feature vector is then used to train and fine-tune a lightweight two-layer multilayer perceptron (MLP) classifier strictly on the training set, following the established training protocol. To prevent data leakage and ensure methodological rigor, an independent test set (comprising 20% of the dataset, subject-wise separated) is reserved exclusively for final performance evaluation. It is not involved in any feature selection, model training, or optimization procedures.
3.4. Training and Optimization
In clinical data processing, this study used XGBoost to build a three-class classification model, with input features including the MMSE total score and Global CDR score. Due to the low input dimensionality (containing only two features), the model complexity was relatively low; therefore, a predefined hyperparameter configuration (maximum depth 6, learning rate 0.3) was adopted, and multi-class log loss (mlogloss) was used as the optimization objective. To ensure reproducibility, the random seed was fixed at 42. During training and validation, the XGBoost model was trained only on the training set to maintain consistency with the training logic of deep image networks and strictly prevent data leakage. In the feature fusion stage, the three-dimensional probability dimension output by XGBoost was used as a high-order clinical feature and fused with 1024-dimensional image features. Subsequently, the fused vector was input into a fully connected PyTorch-based network (including Dropout layers) for processing. In the overall hierarchical architecture, XGBoost acts as a clinical representation extractor, working in conjunction with image features.
After independently training the XGBoost model on clinical data, the parameters of the 3D Dense-Net-121 and the final classifier were jointly optimized. A multi-class cross-entropy loss function was used, and an AdamW optimizer with a weight decay rate of 1 × 10
−5 was employed to suppress overfitting. The initial learning rate was set to 1 × 10
−4. Considering the limitations of computational resources and cloud GPU memory, the batch size was set to 6. Although the small batch size caused random gradient fluctuations in the early stages of training (
Figure 3), the model eventually achieved stable convergence. The training process lasted for 30 epochs, and an early stopping mechanism was introduced to prevent overfitting. Furthermore, we implemented a multi-stage feature selection strategy, using the SHAP algorithm as the core algorithm for evaluating feature importance. To ensure objectivity and prevent data leakage, we implemented a rigorous two-stage development protocol. First, the SHAP algorithm was applied to the validation set to identify the 90 most influential core features from the 1024-dimensional imaging vector by calculating their marginal contributions. Subsequently, a refined classifier (an improved multilayer perceptron) was constructed based on these 90 selected features and 3 clinical probabilities (forming a 93-dimensional vector). Crucially, the fine-tuning of this classifier (lasting 10 epochs) was performed strictly on the training set, while the validation set functioned solely as an unbiased mechanism for feature attribution. The final performance evaluation was then conducted during the testing phase using a completely independent, subject-level test set that was not involved in any feature selection or parameter optimization processes.
This framework combines the deep feature extraction capabilities of 3D Dense-Net-121 with the structured data modeling advantages of gradient boosting trees. The design aims to technically validate a three-class classification task for Alzheimer’s disease by leveraging the unique characteristics of each model through multimodal feature fusion. This method provides a practical reference for the fusion of multimodal information.
3.5. Ablation Study on the Hierarchical Multi-Modal Integration
To systematically quantify the contribution of each modality and to validate the effectiveness of the proposed hierarchical multimodal fusion strategy, an ablation study was conducted by comparing three configurations: (i) a clinical-only baseline, (ii) an imaging-only baseline, and (iii) the complete HMFF, as summarized in
Table 2.
The resulting 95% confidence intervals (CIs) for accuracy, recall, precision, and F1-score are now included, providing additional statistical rigor. When only clinical assessments were used, the model exhibited limited discriminative capability, achieving a mean accuracy of 0.71 (95% CI: [0.679, 0.741]) and an F1-score of 0.66. In contrast, the imaging-only configuration provided a strong baseline with a mean accuracy of 0.92 (95% CI: [0.902, 0.940]). While the performance gain of the proposed HMFF (mean accuracy: 0.94; 95% CI: [0.918, 0.951]) over the imaging-only baseline may appear incremental, these results should be interpreted with caution regarding their statistical robustness.
Despite the modest numerical increase, a 1000-iteration bootstrap analysis confirmed that the HMFF consistently outperformed the unimodal baselines with a p-value of less than 0.001, indicating that the improvement is statistically significant and not due to random fluctuations. These findings suggest that the hierarchical integration of clinical priors provides a stable, albeit subtle, enhancement in discriminative power, offering better classification stability across varying data distributions. This robust statistical foundation, rather than the raw percentage increase alone, substantiates the value of the proposed fusion framework in clinical staging tasks.
4. Experimental Results
We evaluated the HMFF using quantitative indicators, ablation experiments, and visualization analysis. These experiments aimed to assess the model’s accuracy and stability.
4.1. Model Evaluation
To determine the optimal feature subset for improving model performance, we evaluated the impact of systematically increasing the number of selected important features on model classification accuracy. In the early stages of training, accuracy was limited when using a small feature set (e.g., 30–50 features) because these features were insufficient to adequately represent complex pathological and imaging information. Accuracy improved significantly with increasing feature count, peaking at approximately 90 features, with an observed single-experiment accuracy of 0.93 (as shown in
Figure 4). To validate the statistical robustness of this configuration and compare it with the baseline, we performed a bootstrap analysis on the test set for 1000 iterations. The proposed HMFF model achieved a mean accuracy of 0.94, a 95% confidence interval of [0.918, 0.951], and a standard deviation of 0.0085. This was significantly higher than the baseline model using only imagery, which yielded a mean accuracy of 0.92 (95% CI: [0.902, 0.940]). All statistically significant
p-values were less than 0.001, indicating that the results were not due to random fluctuations. While the performance difference between 70 and 150 features was negligible (~0.001), 90 features were identified as the only point where accuracy, recall, precision, and F1 score all peaked simultaneously, confirming this as a stable optimal balance between information completeness and model complexity. As shown in
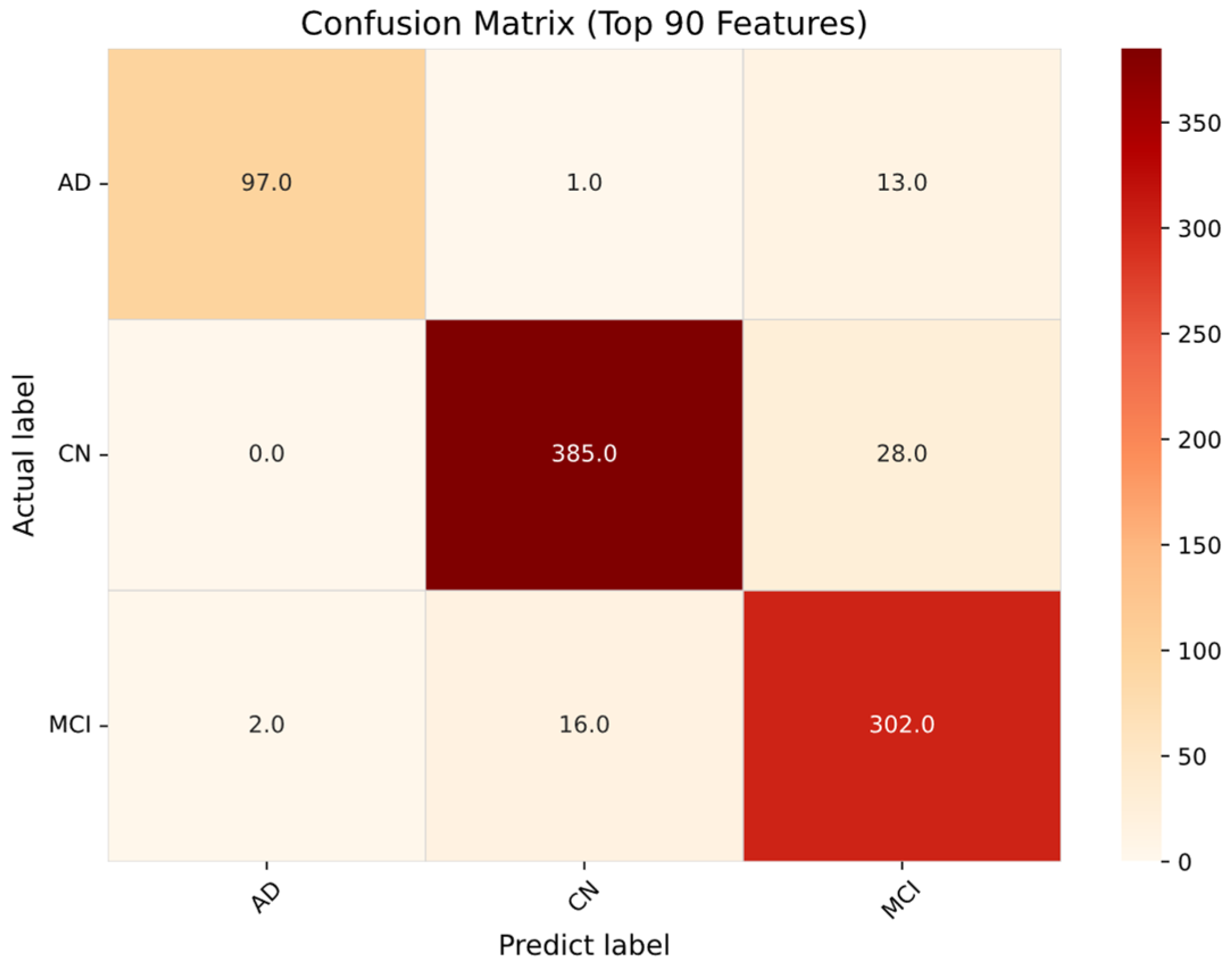
Figure 4, when the number of features exceeds 90, the accuracy tends to plateau or slightly decrease, indicating that the additional features introduce redundancy or noise, thus adversely affecting the stability and efficiency of the model. To visually demonstrate the model’s classification performance with the optimal configuration of 90 features,
Figure 5 shows a confusion matrix representing the raw predictive distribution on the independent test set. This matrix clearly shows the distribution of correct predictions across the three diagnostic categories (AD, MCI, and CN), confirming a high recognition rate for each category. These raw counts serve as the basis for calculating the class-wise metrics, while the final performance reported in
Table 2 reflects the aggregated mean results from 1000 bootstrap iterations to ensure statistical stability. This matrix shows that although the model exhibits strong discriminative power, most misclassifications occur in the MCI category, consistent with the clinical heterogeneity and high pathological similarity observed between early-stage AD and stable-stage MCI patients in the ADNI dataset.
For the quantitative assessment of the diagnostic performance and clinical utility of the three-way Alzheimer’s disease classification model, four standard evaluation metrics were employed:
Recall is a metric that quantifies the model’s ability to identify true positive cases, which provides critical insight into the risk of missed diagnoses in clinical decision-making. Precision evaluates the trustworthiness of positive predictions, balancing false positives and detection needs in practical settings. The F1-score, which harmonizes recall and precision, serves as a comprehensive metric under class-imbalance conditions, more fully reflecting the model’s classification strength across different disease states. Overall accuracy quantifies the proportion of correctly classified instances across all categories, which facilitates rapid comparison of global performance across architectures and feature sets.
For performance validation, these four metrics were computed for each feature subset (
Table 3), data split, and architectural variant to ensure reliability and generalizability [
22]. This multi-level experimental design not only quantifies the model’s differential ability to diagnose AD, MCI, and CN, but also probes its robustness and limitations under varying data distributions. It was demonstrated that judicious and precise feature selection (
Table 4) markedly improves all metrics—especially recall and F1-score—thereby enhancing clinical identification performance [
23]. Conversely, exceeding the optimal feature count can inflate model complexity, leading to metric plateauing or decline and revealing the loss of generalization under overparameterization.
4.2. Comparisons and Limitations
The comparative results summarized in
Table 5 indicate that the proposed Hierarchical Multi-Modal Fusion Framework (HMFF) achieves competitive and robust performance relative to recent learning-based approaches for Alzheimer’s disease classification on the ADNI dataset. These findings are consistent with prior studies demonstrating the effectiveness of multimodal deep learning in capturing complementary pathological information across different cognitive states [
24]. Previous work has demonstrated that integrating structural MRI with clinical assessments provides a reliable foundation for disease staging by jointly modeling neuroanatomical alterations and cognitive decline patterns [
25]. In addition, the importance of feature selection in neurodegenerative disease classification has been widely recognized, particularly for mitigating the curse of dimensionality and enhancing generalization when handling high-dimensional imaging features [
26].
Distinct from existing methods that primarily emphasize architectural depth or ensemble complexity, this study adopts a methodologically balanced fusion strategy that integrates a 3D DenseNet-121 imaging backbone with an XGBoost-based clinical modeling branch, augmented by a hierarchical feature selection mechanism. This design enables effective dimensionality reduction while preserving discriminative information, thereby enhancing classification stability and reducing the risk of overfitting. Moreover, convolutional neural network-based multimodal neuroimaging frameworks have been shown to benefit from imaging biomarkers in improving diagnostic accuracy [
25]; however, many such approaches remain limited in interpretability and reproducibility. By contrast, the proposed HMFF incorporates systematic feature refinement and model interpretability analysis, enhancing both generalizability and practical applicability under clinically realistic conditions.
The relatively robust performance of our 3D DenseNet-121 baseline compared to some earlier studies may be attributed to several practical factors. First, the adoption of a standardized image preprocessing workflow likely provided a cleaner and more consistent input for the network, reducing the impact of scanning artifacts. Furthermore, by re-evaluating these baseline architectures under identical experimental conditions and data splits, we aimed to establish a reliable benchmark for our fusion framework. These results suggest that when fundamental steps such as spatial normalization and skull stripping are carefully managed, standard deep learning architectures can achieve very steady and reliable performance for AD classification tasks.
Despite these advantages, several limitations remain. First, class imbalance within the dataset may introduce bias, particularly in the MCI category, where higher misclassification rates were observed. This outcome likely reflects the intrinsic clinical heterogeneity of MCI and the subtlety of early pathological changes, which remain challenging for automated models to distinguish reliably. Second, the current evaluation is restricted to the ADNI dataset, which may limit generalization to broader clinical populations. To address these limitations, future work will focus on validation using larger-scale, multi-center datasets with more balanced class distributions. Furthermore, the integration of additional complementary modalities—such as functional MRI and diffusion tensor imaging—represents a promising direction for capturing both structural and functional alterations, potentially further enhancing diagnostic accuracy and robustness.
Beyond data-driven improvements, model interpretability remains a core requirement for clinical application. It has been emphasized in previous studies that transparency and trustworthiness are crucial for the widespread acceptance of AI medical models. To address this, Grad-CAM visualization techniques were incorporated into this study, successfully revealing the model’s attention on key brain regions. Future work will further explore combining class activation mapping (CAM) with other interpretability methods. To further enhance model training stability and generalization ability, the exploration of batch re-normalization as an alternative to batch normalization is also proposed for future research. Although batch normalization is widely used to stabilize training, it may introduce statistical bias in small batch scenarios, which can affect model generalization. Batch re-normalization is expected to improve training stability by reducing reliance on batch statistics via the introduction of correction terms.
4.3. Grad-CAM Visualization Analysis
In the clinical application of deep learning models, model interpretability is a key factor for establishing trust among medical professionals and ensuring diagnostic reliability. To address this, Gradient-weighted Class Activation Mapping (Grad-CAM) technology is employed in this study, as shown in
Table 6, to perform visualization analysis on the 3D DenseNet-121 model. This analysis reveals the model’s decision basis and its focus areas in the Alzheimer’s disease three-class classification task. Grad-CAM generates heatmaps that reflect the model’s attention distribution by calculating the gradients of the target class with respect to the last convolutional layer’s feature maps [
27].
This technique can localize specific brain regions in MRI images where disease-related features are identified by the model, thereby providing an intuitive and quantitative basis for interpretation. The specific implementation process is as follows: first, 3D MRI images are inputted into the trained 3D DenseNet-121 model to obtain prediction probabilities for the AD, MCI, and CN classes; next, the feature maps from the final convolutional layer are extracted and class gradient weights are calculated; finally, a weighted combination of these weights and feature maps is performed, and the result is normalized and overlaid onto the original images to generate the visualization heatmaps.
Experimental results show that key brain regions highly associated with Alzheimer’s disease pathology are successfully identified by Grad-CAM as shown in
Table 6. For AD classification, the model’s attention is primarily focused on regions known to be vulnerable to the disease, such as the hippocampus, entorhinal cortex, and medial temporal lobe. For MCI classification, the attention distribution is observed to cover structures around the hippocampus and parts of the frontal lobe, while a relatively uniform attention distribution is presented for CN classification, which reflects the overall integrity of normal brain tissue.
These visualization results not only validate the consistency between the features learned by the model and known Alzheimer’s disease neuropathological mechanisms, but also provide clinicians with an intuitive tool to understand the model’s predictive logic. Through Grad-CAM analysis, the clinical applicability and diagnostic reliability of the proposed multimodal fusion architecture are further confirmed, which lays an important foundation for the deployment of AI-assisted Alzheimer’s disease diagnosis in practical medical settings.
5. Conclusions
In this study, a hierarchical multimodal fusion framework (HMFF) was optimized to facilitate the task of three-stage Alzheimer’s disease classification, encompassing cognitively normal (CN), mild cognitive impairment (MCI), and Alzheimer’s disease (AD) subjects. By systematically integrating volumetric MRI features with structured clinical priors, the proposed framework achieved a peak accuracy of 0.93 and an F1-score of 0.92 when an optimized subset of 90 core features was employed, substantially outperforming unimodal clinical and imaging baselines. Beyond quantitative performance gains, the proposed HMFF demonstrates methodological advantages in terms of feature efficiency and interpretability. Grad-CAM-based visualization analysis revealed that the learned representations consistently attend to clinically relevant neuroanatomical regions, such as the hippocampus and medial temporal lobe, thereby reinforcing the neurological plausibility of the model’s decision process. These results indicate that hierarchical feature refinement and multimodal integration can jointly enhance both diagnostic accuracy and model transparency.
Nevertheless, several limitations remain. The current framework was evaluated exclusively on the ADNI dataset, which may constrain its generalizability to broader clinical populations. In addition, the intrinsic heterogeneity of the MCI category continues to pose a challenge, as reflected by higher misclassification rates relative to CN and AD. Importantly, the objective of this work is not to present a finalized clinical diagnostic system, but rather to establish a technically validated and interpretable baseline for multimodal feature fusion in neurodegenerative disease staging. Future work will therefore focus on large-scale multicenter validation, the incorporation of additional biomarkers and imaging modalities, and the exploration of longitudinal disease modeling to further enhance robustness and clinical applicability in real-world settings.