1. Introduction
Myocardial computed tomography perfusion (CTP) imaging plays a critical role in the evaluation of coronary artery disease by providing quantitative information on myocardial blood flow [
1,
2,
3]. However, acquiring high-quality CTP images often requires repeated scanning across multiple cardiac cycles, resulting in substantial patient radiation exposure [
4]. In clinical practice, reducing the current or voltage of the X-ray tube is a common strategy to reduce the radiation dose. However, this increases the noise level in the reconstructed images [
5]. This increased noise not only degrades visual quality but also negatively affects quantitative perfusion measurements, potentially leading to diagnostic inaccuracies [
6].
Traditional denoising approaches for low-dose CT, including linear filtering [
7], nonlinear diffusion [
8,
9], and iterative reconstruction algorithms [
10], have been widely used to suppress noise in low-dose acquisitions. Although these methods can effectively reduce noise, they often blur fine anatomical and structural details and introduce reconstruction artifacts. Such issues are critical in myocardial CTP imaging, where preserving perfusion details is essential. An alternative approach, compressed sensing [
11], has achieved popularity by utilizing sparsity constraints in transform domains to reconstruct high-quality images from severely undersampled or low-dose datasets. Although compressed sensing can improve noise suppression while preserving edges, it is computationally expensive, sensitive to parameter tuning, and may introduce incoherent artifacts if the sparsity assumptions are incorrect. More advanced statistical model-based iterative reconstruction methods, such as Adaptive Statistical Iterative Reconstruction (ASIR) and Advanced Modelled Iterative Reconstruction (ADMIRE), can achieve improved noise-resolution trade-offs [
12,
13]. However, they require proprietary scanner software, involve high computational costs, and may still produce texture inconsistencies compared to standard-dose images [
14].
In recent years, deep learning-based image denoising has been popular as a powerful alternative, utilizing convolutional neural networks (CNNs) to learn a direct mapping from noisy to high-quality images [
15,
16]. Supervised training with paired low-dose and standard-dose data has demonstrated substantial improvements in noise reduction while preserving anatomical details in CT, MRI, and PET modalities [
17,
18]. Several architectures, such as DnCNN [
15], UNet [
19,
20], and GAN-based networks [
21], have been adapted for specific anatomical applications and have reported superior performance compared to traditional techniques. However, most studies in this domain focus on non-medical natural images or pay little attention to general CT or thoracic imaging, with limited work addressing the unique challenges of low-dose myocardial CTP, including temporal consistency requirements, dynamic contrast enhancement, and preservation of small anatomical structures [
22].
To address these challenges, we present a DnCNN-inspired denoising framework specifically tuned for myocardial CTP imaging. By learning from domain-specific data, the network is optimized to suppress noise while preserving delicate myocardial structures and safeguarding perfusion-related intensity patterns. The proposed model incorporates a variance-stabilizing transformation to unify the mixed Poisson–Gaussian noise distribution, followed by its corresponding inverse transform after reconstruction to preserve intensity fidelity. To enhance denoising efficiency, the input volumes were spatially downsampled, and multi-channel feature maps were generated to match the architecture requirements of FFDNet-style processing. The model is trained on synthetically generated low-dose images derived from high-dose porcine datasets, enabling controlled noise modelling and the availability of ground truth. To our knowledge, this is the first controlled low-dose simulation and denoising pipeline designed explicitly for dynamic myocardial CTP, evaluated using both reference-based and no-reference IQA metrics. Quantitative evaluation using both reference-based metrics (MSE, PSNR, SSIM) and no-reference image quality measures (NIQE, FID, KID) demonstrates that the proposed approach achieves superior noise suppression with minimal loss of diagnostic detail, providing a potential pathway toward safer low-dose myocardial CTP imaging protocol without compromising clinical utility.
2. Related Work
Iterative reconstruction (IR) has served as a fundamental technique in CT reconstruction, providing superior noise reduction compared to conventional filtered back projection (FBP). Unlike FBP, which applies a single-pass analytical reconstruction, IR repeatedly refines the image estimate using projection data to minimize reconstruction error. This iterative process reduces noise, improves contrast, and enhances overall image quality [
23]. Over the years, IR has evolved into several well-known variants, including statistical IR, adaptive statistical IR, model-based IR and hybrid approaches [
24,
25,
26]. These methods aim to balance image quality and computational efficiency, producing diagnostic-quality images at lower radiation doses. However, IR is constrained by two key factors: high computational cost and reliance on unprocessed projection data or sinogram.
Despite its effectiveness, iterative reconstruction faces practical challenges in ultra-low-dose CT settings. At extremely low photon counts, noise characteristics change significantly, reducing the ability of IR algorithms to maintain structural detail [
27]. The statistical models underlying IR can fail to converge optimally when photon statistics are poor, leading to image artifacts and degraded diagnostic performance. Furthermore, IR methods depend on sinogram data, which is limited in availability and computationally expensive. These constraints have encouraged researchers to explore alternative strategies that can simultaneously suppress noise and preserve structure more efficiently and adaptively. Following IR, compressed sensing (CS) emerged as a significant advancement in CT reconstruction. CS exploits sparsity priors to enable accurate image recovery from undersampled projection data, reducing radiation dose and acquisition requirements. However, CS reconstructions are limited by high computational cost and dependence on careful parameter tuning, which have constrained their widespread clinical adoption [
28].
The shortcomings of iterative reconstruction have influenced the development of learning-based denoising techniques. Advances in machine learning have opened a new direction for CT denoising. Learning-based methods leverage large datasets to learn a mapping from noisy to clean images, enabling networks to distinguish structured anatomical signals from unwanted noise [
29]. Unlike fixed analytical filters, deep learning models adapt their processing to the specific characteristics of the input data. Approaches span both the reconstruction domain, where models process raw projection data [
30], and the image domain, where denoising is applied to reconstructed images [
21]. This adaptability has allowed deep learning-based methods to outperform traditional approaches in many benchmark studies.
Reconstruction domain denoising holds promise for tackling noise at its source, directly in the sinogram or projection space. By operating before image formation, these methods can suppress noise without introducing certain artifacts that arise post-reconstruction. Examples include deep networks that predict and subtract noise from sinograms or reconstruct cleaner images through learned forward and inverse transforms [
31]. However, applying deep learning in this domain faces significant barriers. Access to raw projection data is typically restricted to CT scanner manufacturers due to proprietary formats and data privacy concerns. This limits the availability of large-scale datasets required for practical training and has slowed broader adoption compared to image-domain techniques, where reconstructed images are more readily accessible.
Several notable deep learning approaches have targeted the reconstruction domain. Yin et al. [
31] employed a ResNet-based architecture across three stages: sinogram, FBP, and image domains, progressively removing noise at each stage. Ernest et al. [
32] used a UNet model to directly denoise sinogram data, reporting measurable improvements in PSNR and SSIM. Ramon et al. [
33] adopted a 3D convolutional autoencoder to predict high-dose myocardial CTP images from low-dose inputs. Yuan et al. [
30] proposed a UNet approach that interpolates sparse sinograms before reconstruction to improve fidelity, while these methods demonstrate strong technical results, their reliance on projection data has limited scalability and clinical translation, especially in multi-center or multi-vendor settings.
Image-domain denoising has therefore become the most widely studied and implemented approach for low-dose CT (LDCT) imaging, primarily because reconstructed CT images are readily available for both training and evaluation. By operating directly on pixel or voxel intensities, these methods integrate easily with existing clinical workflows. They can be applied retrospectively to archived datasets or in real time during routine practice. Early convolutional neural network (CNN) methods, such as the patch-based approach proposed by Chen et al. [
16], demonstrated measurable improvements in PSNR and SSIM compared to classical algorithms like BM3D and K-SVD, but often failed to preserve delicate anatomical structures. This motivated enhancements such as residual and deconvolution layers [
18], which offered better structural recovery but still faced limitations when relying solely on pixel-wise loss functions.
To address these shortcomings, subsequent research focused on structural and edge preservation. For example, You et al. [
34] proposed a 3D Structurally Sensitive Loss (SSL) combining SSIM, adversarial, and
terms, while Gholizadeh et al. [
35] leveraged dilated convolutions to capture broader contextual information while preserving edge details. ResNet-based models with fused attention mechanisms [
36] integrate spatial and channel attention to enhance detail preservation while maintaining computational efficiency. UNet variants leverage hierarchical feature extraction and skip connections for stable reconstruction performance. GAN-based approaches [
21,
37] achieve superior perceptual quality and texture reproduction, though often at the cost of modest trade-offs in quantitative metrics such as PSNR. These innovations improved perceptual quality but sometimes showed variable performance across datasets, highlighting the domain-dependence of learned models. Other approaches, like Kadimesetty et al. [
38], applied CNN-based denoising directly to perfusion map reconstruction, bypassing the need for separate preprocessing, while this streamlined method eliminates intermediate denoising steps, it does not guarantee alignment with high-dose reference standards when manual validation is necessary.
In recent years, advanced architectures have extended the capabilities of image-domain LDCT denoising. The DnCNN model [
15], adapted for CT imaging, has achieved competitive performance by modelling residual noise rather than directly estimating the clean signal. However, in the context of myocardial CTP imaging, DnCNN presents two limitations. First, to the best of our knowledge, it has not yet been explored for myocardial CTP, with prior applications focusing primarily on low-dose abdominal and lung CT [
39,
40]. Second, its original formulation is tailored to Gaussian noise only, whereas LDCT data typically contain a
blend of Poisson and Gaussian noise, with Poisson noise, arising from the stochastic nature of X-ray photon emission [
41,
42], being dominant and signal-dependent, and Gaussian noise primarily from detector electronics, being signal-independent. This mismatch makes the default DnCNN suboptimal for realistic LDCT scenarios. FFDNet [
43], an extension of DnCNN by the same group of authors, introduces noise level maps to handle more general noise, providing a potential pathway for signal-dependent Poisson noise. Yet, its efficacy on Poisson-dominated LDCT data remains untested. To bridge this gap while leveraging the power of the learning models, we propose a
modified DnCNN designed to learn and remove blended Gaussian–Poisson noise patterns in low-dose cardiac CTP.
3. Proposed Method
3.1. Model Summary
We propose a deep learning framework inspired by DnCNN-FFDNet for denoising low-dose CT images corrupted by blended Poisson–Gaussian noise. Our proposed model works in several stages as described below.
Stage 1: Simulating Poisson–Gaussian Noise. In the first stage, we generate the simulated low-dose CT image from high-dose CT images. We also generate a noise map based on the simulated noise to feed into the FFDNet model. Equation (
1) illustrates how simulated noise is generated. Equation (
2) defines the process for generating the noise map.
Stage 2: Variance-Stabilizing Transform (VST) Preprocessing. The second stage employs a Generalized Anscombe Transform (GAT) [
44] to convert signal-dependent Poisson noise into approximately signal-independent Gaussian noise, stabilizing the noise variance. This transformation is essential because it prepares the data for CNN-based Gaussian denoising, as models like FFDNet/DnCNN perform optimally with Gaussian noise. By preprocessing the input data with GAT, we create a noise-level map compatible with FFDNet, thereby enhancing denoising performance and ensuring more effective training [
45]. The generation of the noise level map is defined in Equation (
2).
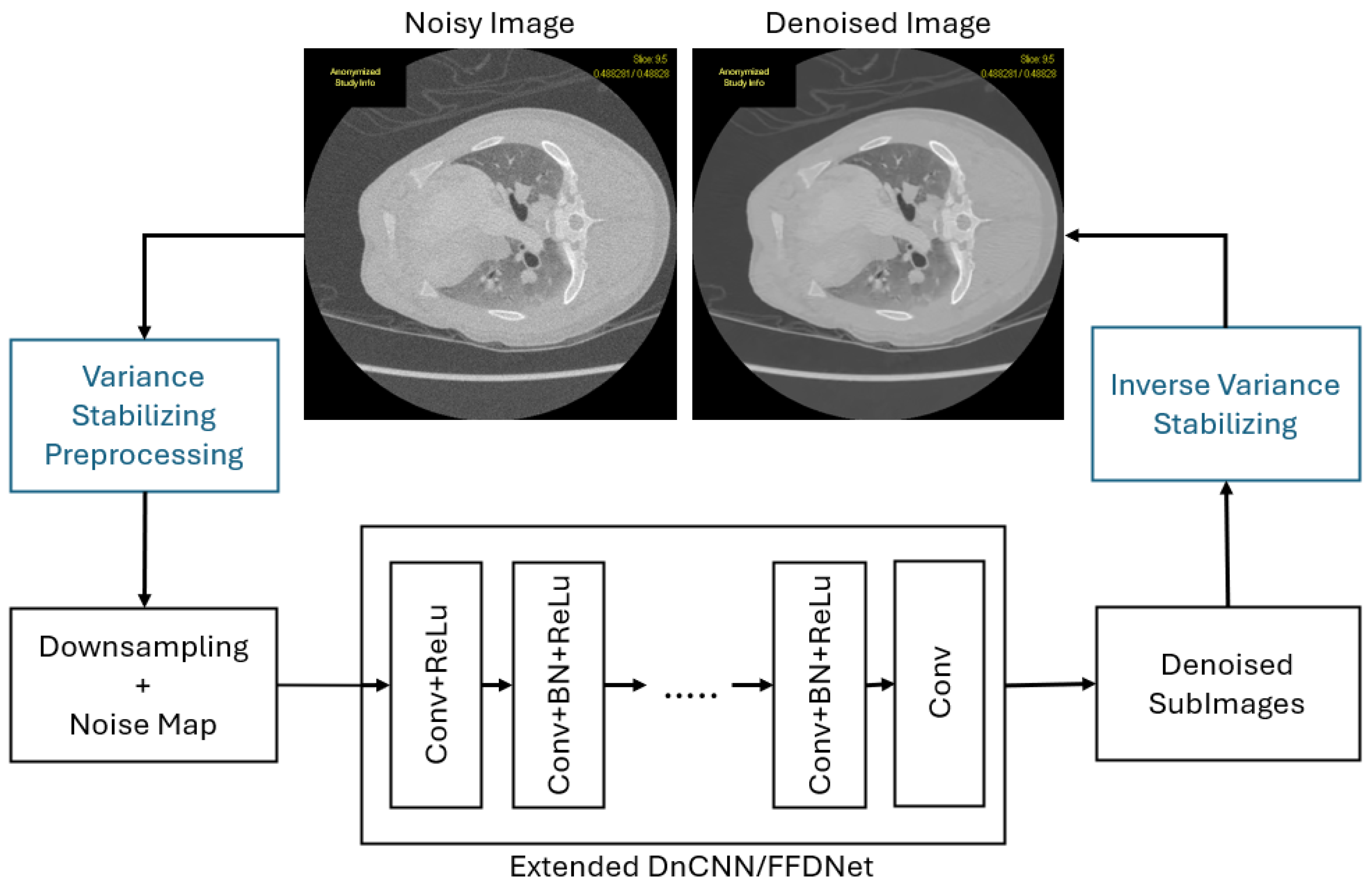
Stage 3: Noise-Adaptive CNN Denoising. The third stage employs a DnCNN-inspired model, with added improvements from FFDNet, enhanced with dilated convolutions for multi-scale feature extraction and residual learning (DnCNN-style). The network takes the simulated low-dose image and the noise level map as input, enabling adaptive denoising. The residual learning strategy of DnCNN predicts the noise component, which is subtracted from the input to produce the denoised output in the VST domain. After denoising, the Inverse Generalized Anscombe Transform (iGAT) reconstructs the final denoised CT image in the original intensity domain, with optional bias correction for improved accuracy.
Figure 1 illustrates the schematic diagram of the proposed denoising architecture.
3.2. Data Description
Data Acquisition: Our study utilized porcine cardiac CT perfusion (CTP) datasets acquired at three tube-current settings: high-dose (80 mA), mid-dose (40 mA), and low-dose (20 mA). Dose level classification depends on subject size, exposure duration, and tube current. For example, a 20 mA scan with a 0.35 s gantry rotation, 80 kilovolts (kV) and 20 perfusion acquisitions corresponds to a practical exposure of 20 × 0.35 s × 20. While 40 mA is typically considered a low dose in humans, the same current may be mid- or high-dose in animal studies.
Data Splitting: For training, we have 4 myocardial CTP studies with 2240 images each, a total of 8960 images to train the model. For testing, we have 8 myocardial CTP studies available, which were scanned with different tube current settings: 80 mA for high-dose radiation and 20 mA for low-dose radiation.
Table 1 shows the study names and our given aliases to those studies, along with their scan dose.
3.3. Training Procedure
Noise Simulation: To create realistic low-dose training pairs, we formed three dose combinations: (80 mA, 20 mA), (80 mA, 40 mA), and (40 mA, 20 mA). In each pair, the higher-dose image was designated as the reference, while the lower-dose image defined the target noise characteristics. Controlled Poisson–Gaussian noise was simulated by modelling quantum noise as a Poisson process (signal-dependent) and electronic noise as additive Gaussian noise (signal-independent). The Poisson component was scaled according to the photon flux ratio between dose levels, and the Gaussian variance was estimated from detector noise measurements. Noise was applied to the high-dose image such that the resulting synthetic image matched the noise statistics of the corresponding low-dose acquisition. This process ensured that training pairs reflected realistic mixed-noise conditions.
Variance Stabilizing Transform (VST) Preprocessing: Because the Poisson component is signal-dependent, a generalized Anscombe transform (GAT) was applied to all simulated noisy images to stabilize variance and approximate the noise distribution as additive white Gaussian noise (AWGN). This transformation facilitated the use of Gaussian-denoising networks, as their performance is tested only with Gaussian noise. Along with the transformed noisy image, a noise-level map was generated to capture spatially varying noise levels. This map was provided as an auxiliary input to the FFDNet network, enabling noise-adaptive processing. This improvement over the DnCNN network allows it to use a fixed noise level during training.
Network Training: We employed a DnCNN-inspired architecture adapted from FFDNet to leverage both residual learning and noise-level conditioning. The network consisted of sequential convolutional layers with batch normalization and ReLU activation, dilated convolutions for multi-scale feature capture, and skip connections for improved gradient flow. Residual learning was implemented such that the network predicted the noise component, which was then subtracted from the input to obtain the denoised image in the VST domain. The inverse GAT was subsequently applied to return the image to its original intensity scale. The primary loss function was mean squared error (MSE) between the predicted and ground-truth high-dose images in the VST domain. Training was conducted on NVIDIA GPU hardware. Realistic Poisson–Gaussian blended noise simulation, variance stabilization, and noise-adaptive residual learning ensured that the network was explicitly trained to address the spatially varying, mixed-noise characteristics of low-dose CT while preserving diagnostically relevant myocardial structures.
The dataset was derived from four CTP studies, each contributing 2240 images, resulting in 8960 image pairs. Of these, 7960 pairs were randomly selected for training and 1000 for validation to avoid bias toward any particular study. Each noisy image was accompanied by a noise-level map, serving as an auxiliary input to the FFDNet architecture and enabling adaptive denoising. The network operated by downsampling noisy inputs, processing them through the multi-scale architecture, and reconstructing denoised sub-images, which were then combined into a clean output, which was finally transformed back to the original intensity domain via the inverse Anscombe transform. Training was performed with a batch size of 16, corresponding to 498 iterations per epoch. A maximum of 50 epochs was allowed, but early stopping with a tolerance of 8 epochs was applied; training converged after 12 epochs when the validation loss plateaued. Optimization used Stochastic Gradient Descent (SGD) with an initial learning rate of and momentum of 0.8. We used FFDNet’s default downsampling. The average inference latency (conducted in Core i5-4570 CPU @ 3.20 GHz (4CPUs)) was 1.2 s per image, corresponding to a throughput of approximately 0.83 frames per second.
3.4. Novelty and Contributions
The objective of this work is not to introduce a new network topology, but to adapt and specialize a proven denoising architecture for the unique challenges of dynamic low-dose myocardial CTP imaging. Although convolutional denoising networks such as DnCNN have been widely studied for static natural and medical images, their direct application to dynamic myocardial CTP is challenging due to mixed signal-dependent and independent noise characteristics, strong temporal dependencies, and the need to preserve perfusion dynamics. The novelty of this work lies in the following contributions:
First adaptation of DnCNN/FFDNet to dynamic myocardial CTP: To the best of our knowledge, this work presents the first systematic adaptation and evaluation of a DnCNN/FFDNet-based denoising framework for dynamic myocardial CT perfusion sequences rather than static CT images.
Noise-adaptive denoising pipeline tailored to myocardial CTP: We introduce a noise-aware preprocessing strategy based on the Generalized Anscombe Transform (GAT) to stabilize the combined Poisson–Gaussian noise inherent to low-dose CTP, enabling effective use of Gaussian-noise–oriented denoising models while respecting CT noise physics.
Controlled high-dose porcine simulation framework: A controlled porcine CTP dataset is used to generate paired high-dose and synthetically degraded low-dose sequences, enabling rigorous reference-based evaluation under realistic motion and perfusion conditions.
Comprehensive quantitative and perceptual evaluation: We present a systematic assessment combining reference-based metrics (MSE, PSNR, SSIM), no-reference perceptual metrics (FID, KID, NIQE) retrained for the myocardial CTP domain, temporal consistency analysis, and expert radiologist assessment. To the best of our knowledge, such retraining and application of no-reference IQAs has not previously been reported for dynamic myocardial CTP denoising.
4. Results
4.1. Reference-Based Image Quality Assessment (IQA)
Reference-based image quality metrics are a conventional means of assessing reconstructed image quality. These metrics are beneficial when a reference image is available, enabling the reconstructed image to be quantitatively compared against the reference to obtain an objective quality score. In our study, high-dose (80 mA) images were available and subsequently simulated to represent low-dose (20 mA) acquisitions. After reconstructing these low-dose images to approximate mA quality, we compared them against the original high-dose (80 mA) reference images to evaluate reconstruction performance.
4.1.1. Means Squared Error (MSE)
MSE measures the average squared difference between corresponding pixels. Equation (
3) defines Mean Squared Error, where
X is the reference image and
Y is the reconstructed image.
Table 2 presents the Mean Squared Error (MSE) values for all simulated test cases, comparing the simulated noisy low-dose images (20 mA) with their corresponding reconstructed outputs. The MSE values presented here are the averages across all MSEs calculated for each image in each test case. Across all test cases, the reconstructed images consistently achieve significantly lower MSE values, reducing the error by more than four times compared to the original noisy inputs. For example, in test case STC1, the MSE drops from 37.50 ± 0.87 to 7.62 ± 3.57, and in STC5, it is reduced from 35.41 ± 0.72 to 6.08 ± 2.17. This substantial and consistent reduction in MSE clearly demonstrates the effectiveness of the reconstruction process in recovering image fidelity and minimizing deviation from the high-quality reference (80 mA), thereby confirming the enhanced accuracy of the reconstructed images.
4.1.2. Peak Signal-to-Noise Ratio (PSNR)
PSNR is derived from MSE and quantifies the ratio of the maximum possible pixel value to the noise power. Equation (
4) defines PSNR, where
denotes the maximum possible intensity value of the image.
Table 3 presents the Peak Signal-to-Noise Ratio (PSNR) values for the simulated test cases, comparing the original noisy images acquired at a low dose (20 mA) with their corresponding reconstructed outputs. In all cases, the reconstructed images demonstrate a substantial improvement in PSNR, with increases of over 7–8 dB relative to the noisy inputs. For example, STC2 improves from 32.45 ± 0.09 to 40.59 ± 2.11 dB, and STC5 from 32.64 ± 0.09 to 40.51 ± 1.30 dB. Since PSNR is a widely used metric for quantifying image quality, where higher values indicate greater similarity to the reference image, these consistent gains confirm that the reconstruction process effectively suppresses noise and preserves structural detail, resulting in images that are significantly closer in quality to high-dose reference scans.
4.1.3. Structural Similarity Index (SSIM)
SSIM measures perceptual similarity by comparing luminance, contrast, and structure. Equation (
5) defines SSIM, where
and
are the means,
and
the variances, and
the covariance between images.
Table 4 presents the Structural Similarity Index (SSIM) values for the simulated test cases, comparing the noisy input images acquired at low dose (20 mA) with the corresponding reconstructed outputs. In every case, the reconstructed images exhibit a significant increase in SSIM, improving from values in the range of 0.63–0.68 (noisy) to 0.87–0.90. For instance, STC2 shows an increase from 0.63 ± 0.006 to 0.90 ± 0.002, and STC7 from 0.67 ± 0.008 to 0.89 ± 0.003. These consistently high SSIM values for the reconstructed images indicate strong structural resemblance to the reference 80 mA images. Given that SSIM is specifically designed to evaluate perceived image quality by comparing local structures, contrast, and luminance, the results clearly demonstrate that the reconstruction method effectively restores structural integrity and preserves critical anatomical details degraded in low-dose acquisitions.
4.2. No Reference Image Quality Assessments (IQA)
To further assess the proposed method’s capacity to generate high-quality HDCT from LDCT acquisitions, we employed no-reference image quality assessment (NR-IQA) metrics, also known as blind image quality metrics. This choice reflects the practical reality that, during clinical acquisition of low-dose scans, a corresponding high-dose reference is typically unavailable for direct comparison. Consequently, NR-IQA provides a viable framework to objectively evaluate model performance in scenarios where reference images are absent, thereby offering insights into the method’s applicability to real-world clinical workflows. It is important to note that blind image quality metrics rely on a global reference model, whereas reference-based metrics use a signal-dependent reference for evaluation.
Although FID/KID and NIQE were initially developed using statistical properties of natural scene images, their sensitivity to perceptual attributes such as smoothness, sharpness, and structural consistency makes them suitable for relative, no-reference image quality assessment in medical imaging. For FID and KID, a reference distribution is still required; therefore, we computed domain-specific feature statistics using a set of high-dose CT images, rather than relying on generic ImageNet-based statistics. Similarly, NIQE was re-estimated on the same high-dose dataset to create a medical-domain NIQE model, which was then applied to evaluate the reconstructed low-dose CT images.
It is worth noting that no-reference image quality metrics such as FID, KID, and NIQE should be interpreted as complementary statistical descriptors of feature-space similarity, rather than as substitutes for established reference-based image quality metrics. To our knowledge, these no-reference metrics have not previously been applied or reported in the myocardial CT perfusion literature. In this work, we therefore also aimed to demonstrate that these models can be retrained in this domain and used reliably to assess reconstructed CT perfusion images in the absence of a reference, while their behaviour correlates with reference-based IQAs in our experiments, a comprehensive study of their reliability and task-level diagnostic relevance is beyond the scope of this work.
4.2.1. FID
Fréchet Inception Distance (FID) measures the distance between the feature distributions of generated images and those of high-quality real images. Lower FID indicates better perceptual similarity. Equation (
6) defines Fréchet Inception Distance (FID),
where:
- -
, are the mean and covariance of reference image features,
- -
, are the mean and covariance of generated image features,
- -
denotes the trace of a matrix, i.e., the sum of the diagonal elements:
, where denotes the diagonal elements of matrix A.
Table 5 presents the Fréchet Inception Distance (FID) scores for each test case, comparing noisy low-dose CT images with their corresponding reconstructed versions. In all cases, the reconstructed images exhibit noticeably lower FID values, indicating a closer match to the distribution of the high-dose reference images. Since lower FID values indicate greater similarity in the feature distributions of the test and reference images, these results quantitatively confirm that the reconstruction method improves image realism and perceptual fidelity.
4.2.2. KID
Kernel Inception Distance (KID) estimates the squared Maximum Mean Discrepancy (MMD) between feature representations of generated and real images using a polynomial kernel. Unlike FID, KID has an unbiased estimator and is more robust on smaller sample sizes. Lower KID indicates better similarity. Equation (
7) defines Kernel Inception Distance (KID).
where:
- -
are features of real images,
- -
are features of generated images,
- -
denotes the feature mapping induced by the polynomial kernel,
- -
n and m are the number of real and generated samples, respectively.
Table 6 presents the Kernel Inception Distance (KID) mean and standard deviation values computed for each test case, comparing the noisy low-dose acquisitions with the corresponding reconstructed outputs. In every case, the reconstructed images achieve significantly lower KID scores, both in mean and variability, indicating a closer match to the reference distribution and greater perceptual similarity. These consistent improvements across all test cases confirm the effectiveness of the reconstruction process in producing images with statistical properties that better resemble those of high-dose reference scans. As KID is specifically designed to measure distribution-level similarity in the deep feature space, the results provide strong evidence of enhanced image quality and perceptual fidelity.
4.2.3. NIQE
Naturalness Image Quality Evaluator (NIQE) quantifies the statistical deviation of the image from natural scene statistics. Lower NIQE scores indicate higher perceptual quality. NIQE is defined by Equation (
8).
where:
- -
, are the mean and covariance of features from natural pristine images,
- -
, are the mean and covariance of features from the distorted image,
- -
T indicates transpose, and −1 denotes matrix inverse.
Table 7 presents the NIQE (Natural Image Quality Evaluator) scores computed for each test case under evaluation. Like other quantitative parameters used in this chapter, this score was calculated per image for one data set, and the average was then taken; while NIQE was initially developed based on statistical regularities observed in natural images, its ability to assess perceptual quality such as image smoothness, sharpness, and structural regularity makes it a useful no-reference metric, even in medical imaging contexts, when used for relative comparisons. In this study, we retrained NIQE on domain-specific data and then applied it consistently across both noisy and reconstructed CT images. As presented in
Table 7, the reconstructed images consistently yield lower NIQE scores compared to their noisy counterparts, indicating improved perceptual quality as perceived by the metric. When interpreted alongside other objective metrics such as PSNR, SSIM, MSE, FID, and KID; and subjective metrics such as AUC and expert ratings, NIQE further supports the conclusion that our proposed model significantly enhances image quality, particularly in terms of noise suppression and structural coherence.
4.2.4. SSIM
The Structural Similarity Index (SSIM) is traditionally defined for evaluating spatial similarity between two images as defined in Equation (
5). To extend this concept to dynamic imaging, the
temporal SSIM (tSSIM) measures structural consistency across the time dimension of a reconstructed CT perfusion sequence. Mathematically, for a temporal sequence
consisting of
T frames, temporal SSIM is computed as the average SSIM between consecutive time frames:
where
denotes the standard structural similarity index between frame
and the subsequent frame
.
In this work, temporal SSIM was computed by calculating SSIM between consecutive reconstructed frames along the time dimension of the CT perfusion sequence and averaging the resulting values. A higher temporal SSIM indicates improved preservation of temporal consistency, reduced flickering, and more physiologically coherent myocardial perfusion dynamics, reflecting the reconstruction method’s ability to maintain structural stability over time. The resulting tSSIM values summarized in
Table 8 demonstrate consistently high temporal fidelity across all test cases, indicating that the proposed reconstruction approach effectively preserves frame-to-frame structural coherence in myocardial CT perfusion imaging.
4.3. Subjective Inspection
In addition to objective quality metrics, we provide a subjective visual example in
Figure 2 illustrating the model’s effectiveness in correcting artifacts and restoring image quality degraded by simulated conditions. Visual inspection against the reference standard reveals that our method has enhanced capability to reproduce fine textures and anatomical structures.
The myocardial and pulmonary regions from
Figure 2 were magnified and are presented in
Figure 3. A comparative inspection of the three images clearly demonstrates that the noise present in the 20 mA acquisition (middle) is substantially reduced in the reconstructed 80 mA image (right). The myocardial structures appear well preserved, while the pulmonary vessels are more distinctly delineated, indicating effective noise suppression and structural recovery achieved by the proposed reconstruction method. This is comparable with the original 80 mA scan (right).
4.4. Comparative Performance
Although a comparative study is informative, in our case it is not an exact like-for-like evaluation because the reference studies available for comparison and listed in
Table 9 were conducted on a different dataset—the
2016 NIH-AAPM-Mayo Clinic Low Dose CT Grand Challenge dataset, where
quarter-dose refers to ¼th of the high-dose scan, which is specific to the Mayo acquisition protocol and cannot be directly translated to our experimental setting. Furthermore, the myocardial CTP imaging data used in our study are inherently susceptible to both noise and residual cardiac motion, unlike the only noise present in low-dose CT data used in works listed in
Table 9. In our porcine myocardial CTP scans, the full dose is 80 mA, and the low dose is 20 mA, which likewise represents a quarter of the reference dose. Thus, the comparison is provided only to offer a relative performance context rather than a strict one-to-one benchmark.
Despite this limitation, the quantitative performance of our proposed method compares favourably with several recent deep learning-based denoising approaches. Yang et al. [
21] proposed a GAN with Wasserstein distance and perceptual loss for LDCT denoising; Ma et al. [
37] introduced a hybrid-loss GAN designed to capture low-dose noise statistics better; Zhang et al. [
46] developed a highly efficient transformer architecture (Hformer) tailored for LDCT restoration; and Zhang et al. [
47] presented an adaptive dual-branch CNN using multi-scale residual attention. These prior GAN, CNN, and transformer-based methods typically reported PSNR values in the range of
–
and SSIM values of
–
on the
NIH-AAPM-Mayo dataset [
21,
37,
46,
47]. In contrast, our model achieves a mean PSNR of
39.96 dB and SSIM of
0.8906 on dynamically acquired myocardial CTP data under realistic Poisson–Gaussian noise conditions. These values indicate performance that is competitive with, or, in the majority of cases,
exceeds, that of several state-of-the-art approaches, despite the added complexity of perfusion imaging, where temporal contrast changes and cardiac motion produce substantially more challenging noise characteristics than in static abdominal or thoracic CT. Please note that the results for the comparative methods were taken from their respective publications, as reported. We did not re-implement or reproduce them.
4.5. Effect of GAT
A key contribution of the proposed method is the use of the Generalized Anscombe Transform (GAT) to stabilize the signal-dependent Poisson noise present in low-dose CT perfusion data, approximating it as signal-independent Gaussian noise. This enables a more effective application of denoising models such as DnCNN/FFDNet, which were initially designed under Gaussian noise assumptions. To assess the impact of variance stabilization, we compare the baseline DnCNN model applied with and without GAT.
As shown in
Table 10, applying the baseline model without GAT improves image quality relative to the noisy inputs; however, the gains remain limited due to the mismatch between the noise statistics and the model assumptions. In contrast, incorporating GAT consistently yields substantially lower MSE, higher PSNR, and improved SSIM across all test cases. These results demonstrate that variance stabilization is essential for effectively denoising low-dose myocardial CT perfusion images with Gaussian noise-oriented deep learning models.
4.6. Expert’s Scoring
A qualitative assessment was conducted by an expert radiologist to evaluate the denoising performance relative to the corresponding high-dose (80 mA) reference images. Two clinically relevant criteria were considered: (i) whether the reconstructed images provide sufficient anatomical detail to support confident diagnosis, and (ii) whether the reconstructed images can be considered diagnostically equivalent to high-dose acquisitions. Each criterion was scored on a five-point Likert scale, where 1 indicates minimal confidence and 5 indicates full confidence.
As summarized in
Table 11, the reconstructed images consistently received high scores across all test cases. For diagnostic sufficiency, a uniform score of 4.0 was assigned for all STCs, indicating consistent confidence in the preservation of diagnostically relevant detail. For diagnostic equivalence to high-dose imaging, the mean score was 4.06, with most cases rated at 4.0 or higher. These results suggest that the proposed denoising approach yields reconstructions that are not only diagnostically adequate but also closely comparable to high-dose CT perfusion images, supporting its potential clinical utility.
5. Limitations
Despite the encouraging results, several limitations of this study should be acknowledged. First, the proposed method focuses on domain adaptation and noise modelling rather than architectural novelty; the backbone network builds upon the established DnCNN/FFDNet method. A full architectural ablation isolating the individual contributions of dilated convolutions and noise-level maps was therefore not conducted, as the effectiveness of these components has been well documented in prior literature. Instead, this work emphasizes their integration and validation within the specific context of dynamic myocardial CT perfusion imaging.
Second, although a controlled porcine myocardial perfusion dataset enables reliable paired low-dose and reference evaluations, its size remains limited and does not include human clinical cases. While porcine CTP closely mimics human myocardial perfusion dynamics and is commonly used in translational imaging research, future validation in larger human datasets will be necessary to establish its generalizability and clinical applicability fully.
Third, quantitative evaluation primarily relies on established reference-based metrics (PSNR, SSIM) and domain-specific-trained no-reference perceptual metrics (NIQE, FID, KID). Although confidence intervals are reported to reflect variability, formal paired statistical hypothesis testing was not performed, as the consistently non-overlapping confidence ranges across dose levels already indicate statistically meaningful improvements. Additional task-based statistical analyses may further strengthen clinical interpretation.
Finally, while temporal stability was assessed using temporal SSIM to capture frame-to-frame consistency, more advanced temporal or task-driven metrics, as well as fully spatiotemporal (2.5D or 3D) network architectures, were beyond the scope of this feasibility study and represent important directions for future work.
6. Conclusions
This study presented a deep learning-based denoising approach for low-dose myocardial CTP imaging, addressing the challenge of preserving diagnostic quality while minimizing radiation exposure. The model was trained on synthetically generated low-dose data, created by applying a blended Poisson–Gaussian noise model to high-dose (80 mA) reference scans, enabling controlled training with ground truth for evaluation.
On synthetically degraded images, the proposed network achieved substantial improvements in quantitative metrics, including MSE, PSNR, and SSIM, demonstrating its ability to suppress noise and restore fine anatomical details. Evaluation on real low-dose (20 mA) animal scans, using no-reference IQA metrics such as NIQE, FID, KID, and subjective inspection, confirmed the method’s robustness and generalizability.
These findings highlight the potential of the proposed denoising framework to approximate high-dose image quality from low-dose acquisitions, offering a pathway to safer myocardial CTP studies without compromising clinical utility. This work was not intended as a benchmarking study against all state-of-the-art denoisers, but rather as a feasibility investigation focused on domain-specific noise modelling, radiation dose reduction potential, and generalization to real data.